

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144503.9 + phase: 0 /pseudo
(346 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
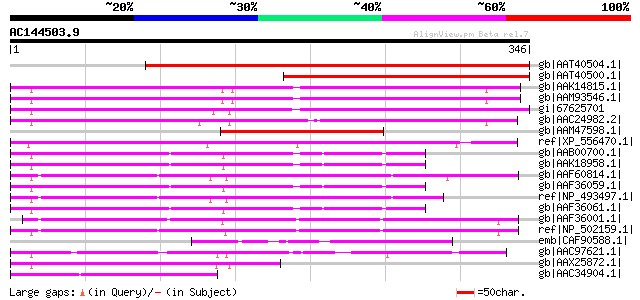
Score E
Sequences producing significant alignments: (bits) Value
gb|AAT40504.1| putative polyprotein [Solanum demissum] 309 9e-83
gb|AAT40500.1| putative reverse transcriptase [Solanum demissum] 206 8e-52
gb|AAK14815.1| polyprotein [Schistosoma japonicum] 150 7e-35
gb|AAM93546.1| polyprotein [Schistosoma japonicum] 150 7e-35
gi|67625701 TPA: endonuclease-reverse transcriptase [Schistosoma... 126 8e-28
gb|AAC24982.2| reverse transcriptase [synthetic construct] 124 4e-27
gb|AAM47598.1| NBS/LRR resistance protein-like protein [Capsicum... 119 1e-25
ref|XP_556470.1| ENSANGP00000028171 [Anopheles gambiae str. PEST... 112 2e-23
gb|AAB00700.1| Hypothetical protein C34D4.5 [Caenorhabditis eleg... 105 2e-21
gb|AAK18958.1| Hypothetical protein F56C9.2 [Caenorhabditis eleg... 102 2e-20
gb|AAF60814.1| Hypothetical protein Y58G8A.2 [Caenorhabditis ele... 101 3e-20
gb|AAF36059.1| Hypothetical protein Y75D11A.4 [Caenorhabditis el... 100 6e-20
ref|NP_493497.1| predicted CDS, reverse transcriptase family mem... 100 6e-20
gb|AAF36061.1| Hypothetical protein Y76B12C.5 [Caenorhabditis el... 97 5e-19
gb|AAF36001.1| Hypothetical protein Y71F9AL.3 [Caenorhabditis el... 94 5e-18
ref|NP_502159.1| predicted CDS, reverse transcriptase family mem... 94 5e-18
emb|CAF90588.1| unnamed protein product [Tetraodon nigroviridis] 93 1e-17
gb|AAC97621.1| ORF MSV061 putative LINE reverse transcriptase, s... 90 9e-17
gb|AAX25872.1| unknown [Schistosoma japonicum] 88 4e-16
gb|AAC34904.1| reverse transcriptase [Limulus polyphemus] gi|751... 85 3e-15
>gb|AAT40504.1| putative polyprotein [Solanum demissum]
Length = 832
Score = 309 bits (791), Expect = 9e-83
Identities = 151/257 (58%), Positives = 189/257 (72%), Gaps = 1/257 (0%)
Query: 91 YEGASTSVRTQDGTTEDFPITIGVHQGSTLSPYLFTLVLDVLTEHIQELAPRCMLFAD-V 149
Y A T VRT G +E FP+ IG+HQGS LSP+LF LV+D LT IQE P CMLFAD +
Sbjct: 370 YYTAKTRVRTVGGDSEHFPVEIGLHQGSVLSPFLFALVMDELTRSIQETVPWCMLFADDI 429
Query: 150 VLVGESREEVNGRLETWRQALEAYGFRLSRSTTEYMECNFSGRRSRSTLEVKVGDHIIPQ 209
VL+ E+R+ VN RLE WRQ LE+ GFRLSR+ TEY+ C FS + +EV++ +IP+
Sbjct: 430 VLIDETRDRVNARLEVWRQTLESKGFRLSRTKTEYLGCKFSDGLDETDVEVRLAAQVIPK 489
Query: 210 VTRFKYLGSFV*NDGEIEADVSHRIQAEWLKWRRASGVLCDKKVPLKLKGKFYRTAVRPA 269
F+YLG+ + G+I+ DV+HR+ A W+KWR ASGVLCDKK+ KLKGKFYR VRPA
Sbjct: 490 KESFRYLGAVIQGSGDIDDDVTHRVGAAWMKWRLASGVLCDKKISPKLKGKFYRVVVRPA 549
Query: 270 LLYGTECWAVKSQHENQVSVAEMRMLRWMSRKTRHDRIKNDTIRERVGVAPIVEKLVENR 329
LLYG ECW VK+ H +++ VAEMRMLRWM TR D+I+N+ IRE+VGVA +V+KL E R
Sbjct: 550 LLYGAECWPVKNAHVHKMHVAEMRMLRWMCGHTRSDKIRNEVIREKVGVASVVDKLREAR 609
Query: 330 LR*FGHVERRPVDAVVR 346
LR FGHV+RR DA VR
Sbjct: 610 LRWFGHVKRRSADAPVR 626
>gb|AAT40500.1| putative reverse transcriptase [Solanum demissum]
Length = 213
Score = 206 bits (524), Expect = 8e-52
Identities = 97/164 (59%), Positives = 123/164 (74%)
Query: 183 EYMECNFSGRRSRSTLEVKVGDHIIPQVTRFKYLGSFV*NDGEIEADVSHRIQAEWLKWR 242
EY+ C FS + +EV++ +IP+ FKYLG+ + G+I+ DV+HR+ A W+KWR
Sbjct: 7 EYLGCKFSDVLDETDVEVRLAAQVIPKKESFKYLGAVIQGSGDIDDDVTHRVGAAWMKWR 66
Query: 243 RASGVLCDKKVPLKLKGKFYRTAVRPALLYGTECWAVKSQHENQVSVAEMRMLRWMSRKT 302
ASGVLCDKK+PLKLKGKFYR VRPALLYG ECW VK+ H +++ VAEMRMLRWM T
Sbjct: 67 LASGVLCDKKIPLKLKGKFYRVVVRPALLYGAECWPVKNAHVHKMHVAEMRMLRWMCGHT 126
Query: 303 RHDRIKNDTIRERVGVAPIVEKLVENRLR*FGHVERRPVDAVVR 346
R D+I+N+ IRE+VGVA +V+KL E RLR FGHV+RR DA VR
Sbjct: 127 RSDKIRNEVIREKVGVASVVDKLREARLRWFGHVKRRSADAPVR 170
>gb|AAK14815.1| polyprotein [Schistosoma japonicum]
Length = 1091
Score = 150 bits (378), Expect = 7e-35
Identities = 106/358 (29%), Positives = 175/358 (48%), Gaps = 22/358 (6%)
Query: 1 KLWERVIERRLRK--EARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVFIDL 58
K+ +I RRL K E + +NQ GF PGR ++ I+ LR+V+E T ++ VF+DL
Sbjct: 653 KILASIILRRLTKAHEEQTRENQGGFRPGRGCIDQIFTLRQVLEHRHTFRRPTIAVFLDL 712
Query: 59 EKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQGS 118
+ +D + R+++W+ L KGV YI I+ +Y + VR + + + GV Q
Sbjct: 713 KAAFDSIDRKVMWQCLSLKGVPEKYINLIQALYSNTTCRVRAYGRLSSELTTSSGVRQAC 772
Query: 119 TLSPYLFTLVLDVLTEHIQELA--PRCMLFA-----------DVVLVGESREEVNGRLET 165
LSP+LF ++D+L E + P LF D+VL+ E +++ L T
Sbjct: 773 PLSPFLFNFIIDILLELTLSSSDFPGVDLFPGDKLTDLEYADDIVLLSEDADKMQDFLTT 832
Query: 166 WRQALEAYGFRLSRSTTEYMECNFSGRRSRSTLEVKVGDHIIPQVTRFKYLGSFV*NDGE 225
+ G R S S + + ++ S ++ +G I V RF YLGS + +G
Sbjct: 833 LNMNVSMLGMRFSPSKCKMLLQDW----LNSAPKLVIGRETIECVNRFTYLGSLISPNGL 888
Query: 226 IEADVSHRIQAEWLKWRRASGVLCDKKVPLKLKGKFYRTAVRPALLYGTECWAVKSQHEN 285
+ ++S RI + + + + L KG+ Y AVR L YG E W ++ +
Sbjct: 889 VSDEISARIHKARSAFANLRHLWRRRDIRLMTKGRVYCAAVRSVLPYGCETWPLRVEDIR 948
Query: 286 QVSVAEMRMLRWMSRKTRHDRIKNDTIRERV--GVAPIVEKLVE-NRLR*FGHVERRP 340
++ V + R LR ++R +R+ N +R RV ++++V +RLR GHV R P
Sbjct: 949 RILVFDHRCLRNIARVCWDNRVSNAWVRNRVLGKYGKSIDEVVNLHRLRWLGHVLRMP 1006
>gb|AAM93546.1| polyprotein [Schistosoma japonicum]
Length = 976
Score = 150 bits (378), Expect = 7e-35
Identities = 106/358 (29%), Positives = 175/358 (48%), Gaps = 22/358 (6%)
Query: 1 KLWERVIERRLRK--EARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVFIDL 58
K+ +I RRL K E + +NQ GF PGR ++ I+ LR+V+E T ++ VF+DL
Sbjct: 538 KILASIILRRLTKAHEEQTRENQGGFRPGRGCIDQIFTLRQVLEHRHTFRRPTIAVFLDL 597
Query: 59 EKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQGS 118
+ +D + R+++W+ L KGV YI I+ +Y + VR + + + GV Q
Sbjct: 598 KAAFDSIDRKVMWQCLSLKGVPEKYINLIQALYSNTTCRVRAYGRLSSELTTSSGVRQAC 657
Query: 119 TLSPYLFTLVLDVLTEHIQELA--PRCMLFA-----------DVVLVGESREEVNGRLET 165
LSP+LF ++D+L E + P LF D+VL+ E +++ L T
Sbjct: 658 PLSPFLFNFIIDILLELTLSSSDFPGVDLFPGDKLTDLEYADDIVLLSEDADKMQDFLTT 717
Query: 166 WRQALEAYGFRLSRSTTEYMECNFSGRRSRSTLEVKVGDHIIPQVTRFKYLGSFV*NDGE 225
+ G R S S + + ++ S ++ +G I V RF YLGS + +G
Sbjct: 718 LNMNVSMLGMRFSPSKCKMLLQDW----LNSAPKLVIGRETIECVNRFTYLGSLISPNGL 773
Query: 226 IEADVSHRIQAEWLKWRRASGVLCDKKVPLKLKGKFYRTAVRPALLYGTECWAVKSQHEN 285
+ ++S RI + + + + L KG+ Y AVR L YG E W ++ +
Sbjct: 774 VSDEISARIHKARSAFANLRHLWRRRDIRLMTKGRVYCAAVRSVLPYGCETWPLRVEDIR 833
Query: 286 QVSVAEMRMLRWMSRKTRHDRIKNDTIRERV--GVAPIVEKLVE-NRLR*FGHVERRP 340
++ V + R LR ++R +R+ N +R RV ++++V +RLR GHV R P
Sbjct: 834 RILVFDHRCLRNIARVCWDNRVSNAWVRNRVLGKYGKSIDEVVNLHRLRWLGHVLRMP 891
>gi|67625701 TPA: endonuclease-reverse transcriptase [Schistosoma mansoni]
Length = 992
Score = 126 bits (317), Expect = 8e-28
Identities = 95/360 (26%), Positives = 163/360 (44%), Gaps = 19/360 (5%)
Query: 1 KLWERVIERRLRK--EARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVFIDL 58
K++ RV+ R++ +A++ D Q GF R + I LR ++E+ L++ FID
Sbjct: 574 KVFNRVLLNRMKDSVDAQLRDQQAGFRKDRSCTDQIATLRIIVEQSIEWNSSLYINFIDY 633
Query: 59 EKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQGS 118
EK +D V R LWK L GV + I++ Y+G + + T+ F + GV QG
Sbjct: 634 EKAFDSVDRTTLWKLLRHYGVPEKIVNIIRNSYDGLNCQIVHGGQLTDSFEVKTGVRQGC 693
Query: 119 TLSPYLFTLVLDVLTE-------HIQELAPRCML----FA-DVVLVGESREEVNGRLETW 166
LSP+LF LV+D + + H + R L FA D+ L+ ++++++ + +
Sbjct: 694 LLSPFLFLLVIDWIMKTSTSGGMHGIQWTGRMQLDDLDFADDLALLSQTQQQMQEKTTSV 753
Query: 167 RQALEAYGFRLSRSTTEYMECNFSGRRSRSTLEVKVGDHIIPQVTRFKYLGSFV*NDGEI 226
A A G +++ ++ + N + T + + + V F YLGS + G
Sbjct: 754 AAASAAVGLNINKGKSKTLRYN-----TICTNPITLDGEALEDVEIFTYLGSIIDEHGGS 808
Query: 227 EADVSHRIQAEWLKWRRASGVLCDKKVPLKLKGKFYRTAVRPALLYGTECWAVKSQHENQ 286
+ADV RI + + + K++ K + + T V+ LLYG E W +
Sbjct: 809 DADVRARIGKARAAYLQLKNIWSSKQLSTNTKVRIFNTNVKTVLLYGAETWRTTKAIIQK 868
Query: 287 VSVAEMRMLRWMSRKTRHDRIKNDTIRERVGVAPIVEKLVENRLR*FGHVERRPVDAVVR 346
+ V LR + R D I N + E P E++ + R + H R+ + V R
Sbjct: 869 IQVFINSCLRKILRIRWPDTISNKLLWETTNQIPAEEEIRKKRWKWIWHTLRKSPNCVTR 928
>gb|AAC24982.2| reverse transcriptase [synthetic construct]
Length = 1016
Score = 124 bits (311), Expect = 4e-27
Identities = 99/357 (27%), Positives = 167/357 (46%), Gaps = 23/357 (6%)
Query: 1 KLWERVIERRLRK--EARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVFIDL 58
KL I RRL K E + Q GF GR ++ I+ LR+++E T ++ +VF+D+
Sbjct: 578 KLLASXILRRLFKTRERLTREEQAGFRSGRGCIDHIFTLRQMLEHRHTYRRPTIVVFLDI 637
Query: 59 EKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQGS 118
+D + R +LW L KKGV +I +K +Y S VR + + F + GV QG
Sbjct: 638 RAAFDSLDRTVLWDCLLKKGVPEKFINILKALYTNTSGRVRAYNHLSPLFHSSSGVRQGC 697
Query: 119 TLSPYLF---------TLVLDVLTEHIQELAPRCML----FADVVLVGESREEVNGRLET 165
+SP+LF T ++DV + L +L D+VL+ ++ + + L
Sbjct: 698 PISPFLFNFAIDDILETALMDVSNGGVDMLPGERLLDLEYADDIVLLCDNAQGMQSALNQ 757
Query: 166 WRQALEAYGFRLSRSTTEYMECNFSGRRSRSTLEVKVGDHIIPQVTRFKYLGSFV*NDGE 225
++ YG + S + + ++ TL+ G+ I V +F YLGS++ G
Sbjct: 758 LAISVRRYGMCFAPSKCKVLLQDWQDSHPVLTLD---GEQ-IEVVEKFVYLGSYISAGGG 813
Query: 226 IEADVSHRIQAEWLKWRRASGVLCDKKVPLKLKGKFYRTAVRPALLYGTECWAVKSQHEN 285
+ ++ RI + + + V L +KG+ Y +VR LLY E W ++ +
Sbjct: 814 VSDEIDARIMKARAAYANLGHLWRLRDVSLAVKGRIYNASVRAVLLYACETWPLRVEDVR 873
Query: 286 QVSVAEMRMLRWMSRKTRHDRIKNDTIRERV----GVAPIVEKLVENRLR*FGHVER 338
++SV + R LR ++ + N +R RV I ++++RLR GHV R
Sbjct: 874 RLSVFDHRCLRRIADIQWQHHVSNAEVRHRVFGRRDDNSIGVTILKHRLRWLGHVLR 930
>gb|AAM47598.1| NBS/LRR resistance protein-like protein [Capsicum annuum]
Length = 122
Score = 119 bits (298), Expect = 1e-25
Identities = 58/110 (52%), Positives = 80/110 (72%), Gaps = 1/110 (0%)
Query: 141 PRCMLFAD-VVLVGESREEVNGRLETWRQALEAYGFRLSRSTTEYMECNFSGRRSRSTLE 199
P CMLFAD VVL+ E+R VN +LE WRQ LE+ GFR+SR+ TEY+EC F+ R + +
Sbjct: 10 PWCMLFADDVVLIDETRGGVNDKLELWRQTLESKGFRVSRTKTEYVECKFNDVRRENEVV 69
Query: 200 VKVGDHIIPQVTRFKYLGSFV*NDGEIEADVSHRIQAEWLKWRRASGVLC 249
V++ + + +FKYLGS + ++GEI+ DVSHRI A W+KW+ ASGV+C
Sbjct: 70 VRLEAQEVKKRDKFKYLGSVIQSNGEIDEDVSHRIGAGWMKWKLASGVMC 119
>ref|XP_556470.1| ENSANGP00000028171 [Anopheles gambiae str. PEST]
gi|55238617|gb|EAL39934.1| ENSANGP00000028171 [Anopheles
gambiae str. PEST]
Length = 777
Score = 112 bits (279), Expect = 2e-23
Identities = 93/364 (25%), Positives = 158/364 (42%), Gaps = 33/364 (9%)
Query: 1 KLWERVIERRL--RKEARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVFIDL 58
K++ +++ RL E V + Q GF G+ T + I+ +R+++E+ K D + +FID
Sbjct: 420 KIFSLILQDRLVPHVEEIVGNYQRGFRNGKSTTDQIFTMRQILEKMAEYKNDTYHLFIDF 479
Query: 59 EKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQGS 118
+ YD + R L+ A+ G+ IR ++ + VR + F T G+ QG
Sbjct: 480 KAAYDSIARVKLYDAMSSFGIPAKLIRLVRMTMTNVTCQVRVDGKLSGPFATTKGLRQGD 539
Query: 119 TLSPYLFTLVLD--------VLTEHIQELAPRCMLFA-DVVLVGESREEVNGRLETWRQA 169
L+ LF L L+ T I + + + +A D+ ++G V + QA
Sbjct: 540 GLACLLFNLALERAIRDSRVETTGTIFYKSTQILAYADDIDIIGLRLSYVAEAYQGIEQA 599
Query: 170 LEAYGFRLSRSTTEYMECNFS----GRRSRSTLEVKVGDHIIPQVTRFKYLGSFV*NDGE 225
E G +++ + T+ M + + +V++G+ V F YLGS V ND
Sbjct: 600 AENLGLQINEAKTKLMVATSADLPINNPNLRRRDVQIGERTFEVVPEFTYLGSKVSNDNS 659
Query: 226 IEADVSHRIQAEWLKWRRASGVLCDKKVPLKLKGKFYRTAVRPALLYGTECWAVKSQHEN 285
+E ++ R+ A + K + + K Y T + P L Y +E W + E
Sbjct: 660 MEVELRARMLAANRSFYSLKKQFTSKNLSRRTKLGLYSTYIVPVLTYASETWTLSKSDEA 719
Query: 286 QVSVAEMRMLR-----------WMSRKTRHDRIKNDTIRERVGVAPIVEKLVENRLR*FG 334
++ E +MLR W SR ND + E G +V+++ RLR G
Sbjct: 720 LLAAFERKMLRRILGPVCVEGQWRSR-------YNDELYEMYGDLTVVQRIKLARLRWAG 772
Query: 335 HVER 338
HV R
Sbjct: 773 HVVR 776
>gb|AAB00700.1| Hypothetical protein C34D4.5 [Caenorhabditis elegans]
gi|17539028|ref|NP_501121.1| predicted CDS, reverse
transcriptase family member (4H911) [Caenorhabditis
elegans] gi|7497012|pir||T29286 hypothetical protein
C34D4.5 - Caenorhabditis elegans
Length = 624
Score = 105 bits (263), Expect = 2e-21
Identities = 90/291 (30%), Positives = 142/291 (47%), Gaps = 22/291 (7%)
Query: 1 KLWERVIERRLRK--EARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVFIDL 58
K++ ++ RRL E+ + ++Q GF GR + I L +IE+ + L L+FID
Sbjct: 205 KMFSSILTRRLTPSLESYLDESQNGFRKGRCCADNIQSLTMLIEKCNEFQLPLLLLFIDY 264
Query: 59 EKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQGS 118
+ +D++ + +LEK G A + I++M EG + D + + GV QG
Sbjct: 265 QTAFDKIGHSAVVSSLEKAGADPAMRKMIQEMMEGGQAEISVHDKKLK-VNLRTGVRQGD 323
Query: 119 TLSPYLFTLVLD-VLTEHIQELAP----------RCMLFA-DVVLVGESREEVNGRLETW 166
+ SP LF+ L +LT+ E A R + FA DVVL+ + EEV RLE
Sbjct: 324 SASPALFSAALQAILTDCDNEFAGVGIKVEGRHIRRLEFADDVVLICSTPEEVQERLEIL 383
Query: 167 RQALEAYGFRLSRSTTEYMECNFSGRRSRSTLEVKVGDHIIPQVTRFKYLGSFV*NDGEI 226
+ YG ++++S T ++ F RS G IIP V +YLG ++ G I
Sbjct: 384 DRISSIYGLKINQSKTVLLKNKF----CRSQDVFFNGSPIIP-VPGCRYLGRWIDISGSI 438
Query: 227 EADVSHRIQAEWLKWRRASGVLCDKKVPLKLKGKFYRTAVRPALLYGTECW 277
+ ++S RI+A W VL + +P K + ++ V PALLY +E W
Sbjct: 439 DEEISRRIRAGWGALVGIKEVL--RIMPNKERIILFKQNVLPALLYASETW 487
>gb|AAK18958.1| Hypothetical protein F56C9.2 [Caenorhabditis elegans]
gi|17553662|ref|NP_498615.1| predicted CDS, reverse
transcriptase family member (3I419) [Caenorhabditis
elegans] gi|7504398|pir||T16474 hypothetical protein
F56C9.2 - Caenorhabditis elegans
Length = 772
Score = 102 bits (254), Expect = 2e-20
Identities = 88/291 (30%), Positives = 142/291 (48%), Gaps = 22/291 (7%)
Query: 1 KLWERVIERRLRK--EARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVFIDL 58
K++ ++ RRL E+ + ++Q GF GR + I L +IE+ + L L+FID
Sbjct: 353 KMFSSILTRRLTPTLESYLDESQNGFRKGRCCADNIQSLTMLIEKCNEFQLPLLLLFIDY 412
Query: 59 EKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQGS 118
+ +D++ + +LEK G A + I++M +G + D + + GV QG
Sbjct: 413 QTAFDKIGHSAVVSSLEKAGADPAMRKMIQEMMDGGQAEITVHDKKLK-VNLCTGVRQGD 471
Query: 119 TLSPYLFTLVLD-VLTEHIQELAP----------RCMLFA-DVVLVGESREEVNGRLETW 166
+ SP LF+ L +LT+ E A R + FA DVVL+ + EV RLE
Sbjct: 472 SASPALFSAALQAILTDCDNEFAGVGINVEGRHIRRLEFADDVVLICSTPGEVQERLEIL 531
Query: 167 RQALEAYGFRLSRSTTEYMECNFSGRRSRSTLEVKVGDHIIPQVTRFKYLGSFV*NDGEI 226
+ YG ++++S T ++ F RS + G IIP V +YLG ++ G I
Sbjct: 532 DRISSNYGLKINQSKTVLLKNKF----CRSQDVLFNGSPIIP-VPGCRYLGRWIDISGSI 586
Query: 227 EADVSHRIQAEWLKWRRASGVLCDKKVPLKLKGKFYRTAVRPALLYGTECW 277
+ ++S RI+A W VL + +P K + ++ V PALLY +E W
Sbjct: 587 DEEISRRIRAGWGALVGIKEVL--RIMPNKERIILFKQNVLPALLYASETW 635
>gb|AAF60814.1| Hypothetical protein Y58G8A.2 [Caenorhabditis elegans]
gi|17566042|ref|NP_503166.1| predicted CDS, reverse
transcriptase family member (5A739) [Caenorhabditis
elegans]
Length = 769
Score = 101 bits (252), Expect = 3e-20
Identities = 89/365 (24%), Positives = 163/365 (44%), Gaps = 29/365 (7%)
Query: 1 KLWERVIERRLRK---EARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVFID 57
K + + + R+RK EA+ + Q GF T++ I+ ++R++E R + L LVFID
Sbjct: 355 KTFTKCVLNRIRKTLEEAQPVE-QAGFRRSFSTIDHIHSVQRLLEVGREYQIPLTLVFID 413
Query: 58 LEKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQG 117
+K +D V + +W +L+++GV+ YI +++ Y ST+ T P+ GV QG
Sbjct: 414 FQKAFDSVEPQDVWDSLQQQGVQRGYINLLQECYTDCSTTF-TPFHKNVTVPVIRGVRQG 472
Query: 118 STLSPYLFTLVLDVL----------------TEHIQELAPRC--MLFA-DVVLVGESREE 158
+SP LF+ L+ + TE I+ + FA D+VLV +
Sbjct: 473 DPISPNLFSACLEHVFRQLNWKHFKGDERYETEGIRVNGQNLTNLRFADDIVLVAHNPRT 532
Query: 159 VNGRLETWRQALEAYGFRLSRSTTEYMECNFSGRRSRSTLEVKVGDHIIPQVTRFKYLGS 218
+ L + + G +++ T+ + F+ +R + II V + YLG
Sbjct: 533 ASQMLTELVEKCSSVGLKINTGKTKVLRNRFAYKRKVEIRCPNTTNIIIDDVNEYIYLGR 592
Query: 219 FV*NDGEIEADVSHRIQAEWLKWRRASGVLCDKKVPLKLKGKFYRTAVRPALLYGTECWA 278
+ + + ++ R +A W + P +L+ + + V PAL YG+E W
Sbjct: 593 QINDSNNLLPELHRRRRAAWAAFTNIKSTTDQITCP-RLRANLFDSTVLPALTYGSEAWT 651
Query: 279 VKSQHENQVSVA----EMRMLRWMSRKTRHDRIKNDTIRERVGVAPIVEKLVENRLR*FG 334
+ +V V E +++ + R I + +RE+ + + + + +L G
Sbjct: 652 FTKELAERVRVTHAALERKLVGLTLTEQRERNIHREEVREKSKLRDPLIHIKKKKLGWAG 711
Query: 335 HVERR 339
HV RR
Sbjct: 712 HVARR 716
>gb|AAF36059.1| Hypothetical protein Y75D11A.4 [Caenorhabditis elegans]
gi|17570529|ref|NP_508323.1| predicted CDS, reverse
transcriptase family member (XC378) [Caenorhabditis
elegans]
Length = 480
Score = 100 bits (249), Expect = 6e-20
Identities = 88/291 (30%), Positives = 141/291 (48%), Gaps = 22/291 (7%)
Query: 1 KLWERVIERRLRK--EARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVFIDL 58
K++ ++ RRL E+ + ++Q GF GR + I L +IE+ + L L+FID
Sbjct: 61 KMFSGILTRRLTPTLESYLDESQNGFRKGRCCADNIQSLTMLIEKCNEFQLPLLLLFIDY 120
Query: 59 EKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQGS 118
+ +D++ + +LE+ G A + I++M E + D + + GV QG
Sbjct: 121 QTAFDKISHSSVVSSLERAGADPAMRKMIQEMMERGQAEITVHDKKLK-VNLCTGVRQGD 179
Query: 119 TLSPYLFTLVLD-VLTEHIQELAP----------RCMLFA-DVVLVGESREEVNGRLETW 166
+ SP LF+ L +LT+ ELA R + FA DVVL+ + EEV RLE
Sbjct: 180 SASPALFSAALQAILTDCDNELAGVGISVEGRHIRRLEFADDVVLICSTPEEVQERLEIL 239
Query: 167 RQALEAYGFRLSRSTTEYMECNFSGRRSRSTLEVKVGDHIIPQVTRFKYLGSFV*NDGEI 226
+ YG ++ +S T ++ F RS + G IIP V +YLG ++ G I
Sbjct: 240 DRISSYYGLKIDQSKTVLLKNKF----CRSQDVLFNGSPIIP-VPGCRYLGRWIDISGSI 294
Query: 227 EADVSHRIQAEWLKWRRASGVLCDKKVPLKLKGKFYRTAVRPALLYGTECW 277
+ ++S RI+A W VL + +P K ++ V PALLY ++ W
Sbjct: 295 DEEISRRIRAGWGALVGIKEVL--RIMPNKENIILFKQNVLPALLYASKTW 343
>ref|NP_493497.1| predicted CDS, reverse transcriptase family member (1O881)
[Caenorhabditis elegans]
Length = 835
Score = 100 bits (249), Expect = 6e-20
Identities = 82/311 (26%), Positives = 144/311 (45%), Gaps = 25/311 (8%)
Query: 1 KLWERVIERRLRK---EARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVFID 57
K + + + R+RK EA+ + Q GF T++ I+ ++R++E R + L LVFID
Sbjct: 474 KTFTKCVLNRIRKTLEEAQPVE-QAGFRRSFSTIDHIHSVQRLLEVGREYQIPLTLVFID 532
Query: 58 LEKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQG 117
+K +D V + +W +L+++GV+ YI +++ Y ST+ T P+T GV QG
Sbjct: 533 FQKDFDSVEPQDVWDSLQQQGVQRGYINLLQECYTDCSTTF-TPFHKNVTVPVTRGVRQG 591
Query: 118 STLSPYLFTLVLDVL----------------TEHIQELAPRC--MLFA-DVVLVGESREE 158
+SP LF+ L+ + TE I+ + FA D+VLV +
Sbjct: 592 DPISPNLFSACLEHVFRQLNWKHFKGDERYETEGIRVNGQNLTNLRFADDIVLVAHNPRT 651
Query: 159 VNGRLETWRQALEAYGFRLSRSTTEYMECNFSGRRSRSTLEVKVGDHIIPQVTRFKYLGS 218
V+ L + + G +++ T+ + FS +R + II V + YLG
Sbjct: 652 VSQMLTELVEKCSSVGLKINTGKTKVLRNRFSYKRKVEIRCPNTTNIIIDDVNEYIYLGR 711
Query: 219 FV*NDGEIEADVSHRIQAEWLKWRRASGVLCDKKVPLKLKGKFYRTAVRPALLYGTECWA 278
+ + + A++ R +A W + + P +L+ + + V PAL YG+E W
Sbjct: 712 QINDSNNLLAELHRRRRAAWAAFTNIKSTMDQITCP-RLRANLFDSTVLPALTYGSEAWT 770
Query: 279 VKSQHENQVSV 289
+ +V V
Sbjct: 771 FTKELAERVRV 781
>gb|AAF36061.1| Hypothetical protein Y76B12C.5 [Caenorhabditis elegans]
gi|17544228|ref|NP_500154.1| predicted CDS, reverse
transcriptase family member (4C744) [Caenorhabditis
elegans]
Length = 938
Score = 97.4 bits (241), Expect = 5e-19
Identities = 87/292 (29%), Positives = 143/292 (48%), Gaps = 23/292 (7%)
Query: 1 KLWERVIERRLRK--EARVTDNQFG-FMPGRLTMEAIYLLRRVIERYRTDKKDLHLVFID 57
K++ ++ RRL E+ + ++Q F GR + I L +IE+ + L L+FID
Sbjct: 604 KMFSSILTRRLTPTLESYLDESQTKVFRKGRCCADNIQSLTMLIEKCNEFQLPLLLLFID 663
Query: 58 LEKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQG 117
+ +D++ + +LE+ G A + I+++ EG + D + + G+ QG
Sbjct: 664 YQTAFDKISHSSVVSSLERAGADPAMRKMIQELMEGGQAEITVHDKKLK-VNLCTGIRQG 722
Query: 118 STLSPYLFTLVLD-VLTEHIQELAP----------RCMLFA-DVVLVGESREEVNGRLET 165
+ SP LF+ L +LT+ ELA R + FA DVVL+ + EEV RLE
Sbjct: 723 DSASPALFSAALQAILTDCDNELAGVGISVEGRHIRRLEFADDVVLICSTPEEVQERLEI 782
Query: 166 WRQALEAYGFRLSRSTTEYMECNFSGRRSRSTLEVKVGDHIIPQVTRFKYLGSFV*NDGE 225
+ YG ++++S T ++ F RS + G IIP V +YLG ++ G
Sbjct: 783 LDRISSNYGLKINQSKTVLLKNKF----CRSQDILFNGSPIIP-VPGCRYLGRWIDISGS 837
Query: 226 IEADVSHRIQAEWLKWRRASGVLCDKKVPLKLKGKFYRTAVRPALLYGTECW 277
I+ ++S RI+A W VL + +P K + ++ V PALLY +E W
Sbjct: 838 IDEEISRRIRAGWGALVGIKEVL--RIMPNKERIILFKQNVLPALLYASETW 887
>gb|AAF36001.1| Hypothetical protein Y71F9AL.3 [Caenorhabditis elegans]
gi|17510457|ref|NP_491073.1| reverse transcriptase
family member (1D477) [Caenorhabditis elegans]
Length = 423
Score = 94.4 bits (233), Expect = 5e-18
Identities = 85/356 (23%), Positives = 149/356 (40%), Gaps = 29/356 (8%)
Query: 9 RRLRKEARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVFIDLEKTYDRVPRE 68
RR EA+ + Q GF T++ I+ L+R++E R + L LVFID +K +D V +
Sbjct: 2 RRSLDEAQPVE-QAGFRRSFSTIDHIHSLQRLLEVGREYQIPLTLVFIDFKKAFDSVEHQ 60
Query: 69 ILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQGSTLSPYLFTLV 128
+WK+L+++G AYI +K+ Y+ +T+ P+T GV QG +SP LF+
Sbjct: 61 AIWKSLDEQGADGAYIDLLKECYKNCTTNFAPFHRPV-TVPVTKGVRQGDPISPNLFSAC 119
Query: 129 LDVLTEHIQELA--------------------PRCMLFA-DVVLVGESREEVNGRLETWR 167
L+ + + + P + FA D+VL+ + L+
Sbjct: 120 LEHVFRKLSWIELKGEAEDYDTIPGMRVNGRNPTNLRFADDIVLIANHPNTASKMLQELV 179
Query: 168 QALEAYGFRLSRSTTEYMECNFSGRRSRSTLEVKVGDHIIPQVTRFKYLGSFV*NDGEIE 227
Q G ++ T+ + F+ S+ + V + YLG + +
Sbjct: 180 QKCSEVGLEINTGKTKVLRNRFAD-PSKVYFGSPSPTTQLDDVDEYIYLGRQINAQNNLM 238
Query: 228 ADVSHRIQAEWLKWRRASGVLCDKKVPLKLKGKFYRTAVRPALLYGTECWAVKSQHENQV 287
++ R +A W + D K++ + + V PAL YG+E W +V
Sbjct: 239 PEIHRRRRAAWAAFNGIKNT-ADSITDKKIRANLFDSIVLPALTYGSEAWTFTKALSERV 297
Query: 288 SVAEMRMLRWMSRKTRHDRIKNDTIRERVGVAPIVEK----LVENRLR*FGHVERR 339
+ + R + T + + D RE + +V + + +L GHV RR
Sbjct: 298 RITHASLERRLVGITLTQQRERDLHREDIRTMSLVRDPLNFVKKRKLGWAGHVARR 353
>ref|NP_502159.1| predicted CDS, reverse transcriptase family member (4M184)
[Caenorhabditis elegans] gi|7499626|pir||T21233
hypothetical protein F22B3.6 - Caenorhabditis elegans
Length = 553
Score = 94.4 bits (233), Expect = 5e-18
Identities = 86/367 (23%), Positives = 157/367 (42%), Gaps = 32/367 (8%)
Query: 1 KLWERVIERRLRK---EARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVFID 57
K++ + + R+R+ EA+ + Q GF T++ I+ L+R++E R + L LVFID
Sbjct: 121 KVFTKCLLNRMRRSLDEAQPVE-QAGFRRSFSTIDHIHSLQRLLEVGREYQIPLTLVFID 179
Query: 58 LEKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQG 117
+K +D V + +WK+L+++G AYI +K+ Y+ +T+ T P+T GV QG
Sbjct: 180 FKKAFDSVEHQAIWKSLDEQGADGAYIDLLKECYKNCTTNF-TPFHRPVAVPVTKGVRQG 238
Query: 118 STLSPYLFTLVLDVLTEHIQELAPR--------------------CMLFA-DVVLVGESR 156
+SP LF+ L+ + + + + + FA D+VL+
Sbjct: 239 DPISPNLFSACLEHVFRKLSWIELKGEAEDYDTIPGMRVNGRNLTSLRFADDIVLIANHP 298
Query: 157 EEVNGRLETWRQALEAYGFRLSRSTTEYMECNFSGRRSRSTLEVKVGDHIIPQVTRFKYL 216
+ L+ Q G ++ T+ + F+ S+ + V + YL
Sbjct: 299 NTASKMLQELVQKCSEVGLEINTGKTKVLRNRFAD-PSKVYFSSPSPTTQLDDVDEYIYL 357
Query: 217 GSFV*NDGEIEADVSHRIQAEWLKWRRASGVLCDKKVPLKLKGKFYRTAVRPALLYGTEC 276
G + + ++ R +A W + D K++ + + V PAL YG+E
Sbjct: 358 GRQINAQNNLMPEIHRRRRAAWAAFNGIKNT-TDSITDKKIRANLFDSIVLPALTYGSEA 416
Query: 277 WAVKSQHENQVSVAEMRMLRWMSRKTRHDRIKNDTIRERVGVAPIVEK----LVENRLR* 332
W +V + + R + T + + D RE + +V + + +L
Sbjct: 417 WTFTKALSERVRITHASLERRLVGITLTQQRERDLHREDIRTMSLVRDPLNFVKKRKLGW 476
Query: 333 FGHVERR 339
GHV RR
Sbjct: 477 AGHVARR 483
>emb|CAF90588.1| unnamed protein product [Tetraodon nigroviridis]
Length = 183
Score = 93.2 bits (230), Expect = 1e-17
Identities = 63/175 (36%), Positives = 97/175 (55%), Gaps = 18/175 (10%)
Query: 122 PYLFTLVLDVLTEHIQELAPRCMLFA-DVVLVGESREEVNGRLETWRQALEAYGFRLSRS 180
P L +V+D LT+ +++ +P +FA D+V+ SRE+V +LE R ALE+ S
Sbjct: 19 PLLVAMVMDRLTDEVRQESPWTTMFAGDIVMC--SREQVEEKLEERRFALES-------S 69
Query: 181 TTEYMECNFSGRRSRSTLEVKVGDHIIPQVTRFKYLGSFV*NDGEIEADVSHRIQAEWLK 240
TE E + SG E+K G+ + K LGS V GE +V R+QA W
Sbjct: 70 KTEN-ERDLSGSVRLQGEEIKKGEDL-------KNLGSTVQTSGECGKEVKKRVQAGWNW 121
Query: 241 WRRASGVLCDKKVPLKLKGKFYRTAVRPALLYGTECWAVKSQHENQVSVAEMRML 295
W + SGV+CD+ V K+K K +T RPA+++G E ++ + E ++ +AEM+ L
Sbjct: 122 WGKVSGVMCDRGVSAKIKRKVDKTVARPAIIFGLETVPLRKRQEAELELAEMKAL 176
>gb|AAC97621.1| ORF MSV061 putative LINE reverse transcriptase, similar to
Caenorhabditis elegans GB:U00063 [Melanoplus sanguinipes
entomopoxvirus] gi|11361248|pir||T28222 hypothetical
protein ORF61 - Melanoplus sanguinipes entomopoxvirus
gi|9631308|ref|NP_048132.1| ORF MSV061 putative LINE
reverse transcriptase, similar to Caenorhabditis elegans
GB:U00063 [Melanoplus sanguinipes entomopoxvirus]
Length = 531
Score = 90.1 bits (222), Expect = 9e-17
Identities = 87/352 (24%), Positives = 158/352 (44%), Gaps = 46/352 (13%)
Query: 1 KLWERVIERRLRK--EARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVFIDL 58
KL+ +++ RL+ E +++ Q GF R + IY+L +I R KK+ + FID
Sbjct: 184 KLYSKILNNRLKPIMENLISEEQSGFRKNRSIKDNIYILHEII---RNRKKETYFAFIDF 240
Query: 59 EKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQGS 118
+K +D V REILW + G I AIK +YE V+ ++ I G+ QG
Sbjct: 241 KKAFDNVDREILWNIMHLHGYPPHLINAIKSLYENTRIIVK-----SKVIKINKGIKQGC 295
Query: 119 TLSPYLFTLVLD-VLTEHIQ------ELAPRC----MLFA-DVVLVGESREEVNGRLETW 166
+S LF + +D ++ E Q EL +L+A D+V++ ++ + ++
Sbjct: 296 AVSLSLFNIYIDHIMNEWKQHNIKGIELNDNTNLNYILYADDLVIISTNKNDFIHGIDML 355
Query: 167 RQALEAYGFRLSRSTTEYMECNFSGRRSRSTLEVKVGDHIIPQVTRFKYLGSFV*NDGEI 226
+ Y +S T+ M F G + +E+ + D++I QV F YLG
Sbjct: 356 NNLSKKYKLPISFEKTKVMA--FKGTET-INIEIYIDDNLIQQVDCFTYLG--------- 403
Query: 227 EADVSHRIQAEWLKWRRASGVLCD-------KKVPLKLKGKFYRTAVRPALLYGTECWAV 279
++S+ + L +C+ KV +K+ Y+ P + +
Sbjct: 404 -YEISYTNKINLLYKLNQFDKICNLIINSLKDKVDIKIIIHVYKILAVPIFTSRFIFYEL 462
Query: 280 KSQHENQVSVAEMRMLRWMSRKTRHDRIKNDTIRERVGVAPIVEKLVENRLR 331
S+ EN ++ EM L + +++D IR+ + V +V+K+ E++ +
Sbjct: 463 SSEEENIITRHEMHFL----KNILDFNLEDDIIRQILDVYKLVDKINESKYK 510
>gb|AAX25872.1| unknown [Schistosoma japonicum]
Length = 416
Score = 87.8 bits (216), Expect = 4e-16
Identities = 60/195 (30%), Positives = 92/195 (46%), Gaps = 15/195 (7%)
Query: 1 KLWERVIERRLRK--EARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVFIDL 58
K+ +I RL K E + +NQ GF PGR ++ I+ +R+V+E ++ +VF+DL
Sbjct: 214 KILASIIIGRLTKTRELQTRENQAGFRPGRGCIDHIFTIRQVLEHRHAYRRPTMVVFLDL 273
Query: 59 EKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQGS 118
+ +D V RE+LW+ L KGV YI +K V + +F + GV Q
Sbjct: 274 KAAFDSVDREVLWQCLSLKGVPQKYINLVKLFTRTLPGRVTAYGELSSEFLTSSGVRQEC 333
Query: 119 TLSPYLFTLVLDVLTEHI--------QELAPRCML-----FADVVLVGESREEVNGRLET 165
LSP+LF V+D+L + + P L D+VL GE +++ L T
Sbjct: 334 PLSPFLFNFVIDMLLDITLSSSDFSGVDFLPGASLTDLEYADDIVLFGEDADKMQSLLTT 393
Query: 166 WRQALEAYGFRLSRS 180
+G R S S
Sbjct: 394 LSNNASMFGMRFSPS 408
>gb|AAC34904.1| reverse transcriptase [Limulus polyphemus] gi|7511751|pir||T30935
reverse transcriptase - Atlantic horseshoe crab
retrotransposon R (fragment)
Length = 1168
Score = 85.1 bits (209), Expect = 3e-15
Identities = 47/138 (34%), Positives = 75/138 (54%), Gaps = 1/138 (0%)
Query: 1 KLWERVIERRLRKEARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVFIDLEK 60
+L+ R++ RRL + ++ Q GF+P + I+LL+ + R + K L L +DL K
Sbjct: 525 RLYTRILARRLERAVQINPRQRGFVPQAGCRDNIFLLQSAMRRAKR-KGTLALGLLDLSK 583
Query: 61 TYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQGSTL 120
+D V + L +LE+ V ++R ++DMY G STS R +T + GV QG +
Sbjct: 584 AFDTVGHKHLLTSLERFAVHPHFVRIVEDMYSGCSTSFRVGSQSTRPIVLMRGVKQGDPM 643
Query: 121 SPYLFTLVLDVLTEHIQE 138
SP LF + LD L ++E
Sbjct: 644 SPILFNIALDPLLRQLEE 661
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.326 0.140 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 555,017,708
Number of Sequences: 2540612
Number of extensions: 21964071
Number of successful extensions: 59671
Number of sequences better than 10.0: 1215
Number of HSP's better than 10.0 without gapping: 760
Number of HSP's successfully gapped in prelim test: 455
Number of HSP's that attempted gapping in prelim test: 58275
Number of HSP's gapped (non-prelim): 1335
length of query: 346
length of database: 863,360,394
effective HSP length: 129
effective length of query: 217
effective length of database: 535,621,446
effective search space: 116229853782
effective search space used: 116229853782
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 75 (33.5 bits)
Medicago: description of AC144503.9


