

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144481.6 - phase: 0
(754 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
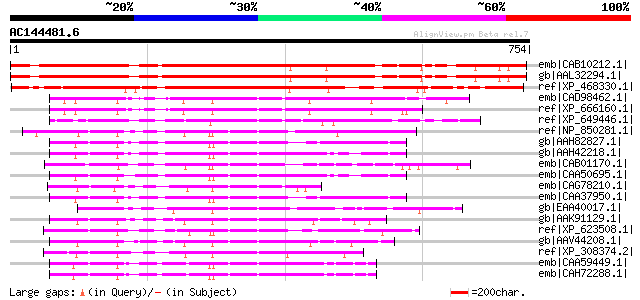
Score E
Sequences producing significant alignments: (bits) Value
emb|CAB10212.1| kinesin like protein [Arabidopsis thaliana] gi|7... 823 0.0
gb|AAL32294.1| phragmoplast-associated kinesin-related protein 2... 823 0.0
ref|XP_468330.1| putative phragmoplast-associated kinesin [Oryza... 634 e-180
emb|CAD98462.1| kinesin-like boursin, possible [Cryptosporidium ... 160 1e-37
ref|XP_666160.1| kinesin-like boursin [Cryptosporidium hominis] ... 158 5e-37
ref|XP_649446.1| kinesin family protein [Entamoeba histolytica H... 156 2e-36
ref|NP_850281.1| kinesin motor protein-related [Arabidopsis thal... 152 3e-35
gb|AAH82827.1| LOC397908 protein [Xenopus laevis] 151 7e-35
gb|AAH42218.1| MGC52588 protein [Xenopus laevis] 150 1e-34
emb|CAB01170.1| Hypothetical protein F23B12.8 [Caenorhabditis el... 150 1e-34
emb|CAA50695.1| kinesin like protein [Xenopus laevis] gi|2497521... 150 1e-34
emb|CAG78210.1| unnamed protein product [Yarrowia lipolytica CLI... 150 1e-34
emb|CAA37950.1| kinesine [Xenopus laevis] gi|119217|sp|P28025|EG... 149 3e-34
gb|EAA40017.1| GLP_572_50389_48461 [Giardia lamblia ATCC 50803] 148 6e-34
gb|AAK91129.1| KRP120-2 [Daucus carota] 148 7e-34
ref|XP_623508.1| PREDICTED: similar to kinesin-like boursin [Api... 148 7e-34
gb|AAV44208.1| putative kinesin [Oryza sativa (japonica cultivar... 147 1e-33
ref|XP_308374.2| ENSANGP00000019061 [Anopheles gambiae str. PEST... 146 2e-33
emb|CAA59449.1| kinesin-related protein [Homo sapiens] gi|170662... 146 3e-33
emb|CAH72288.1| kinesin family member 11 [Homo sapiens] gi|55959... 146 3e-33
>emb|CAB10212.1| kinesin like protein [Arabidopsis thaliana]
gi|7268138|emb|CAB78475.1| kinesin like protein
[Arabidopsis thaliana] gi|7428717|pir||B71405 probable
kinesin - Arabidopsis thaliana
Length = 959
Score = 823 bits (2126), Expect = 0.0
Identities = 465/781 (59%), Positives = 557/781 (70%), Gaps = 85/781 (10%)
Query: 1 MAPTPSSS-SKQIHTTHLITPRSKHRLNFNGVKPAPTPPHPHPSPHPNFNNKDSPPEHPI 59
MAPTPSSS S Q T + TP++K RLNF+ +P+P+ + SPPEHP+
Sbjct: 1 MAPTPSSSRSNQTQYTLIRTPQTKQRLNFHS-----------KTPNPDGSKDPSPPEHPV 49
Query: 60 EVIARIRDYPDRKDKPLSVLQASSNSRSIRVKADFGYRDFTLDGVSVSEEEELDLFYKKF 119
EVI RIRDYPDRK+K S+LQ +++++++RV+AD GYRDFTLDGVS SE+E L+ FYKKF
Sbjct: 50 EVIGRIRDYPDRKEKSPSILQVNTDNQTVRVRADVGYRDFTLDGVSFSEQEGLEEFYKKF 109
Query: 120 VESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQAGIVYRALRDILGDGDTDSEGSDG 179
+E RI GVK+G+KCTIMMYGPTG+GKSHTMFGC K+ GIVYR+LRDILGD D D
Sbjct: 110 IEERIKGVKVGNKCTIMMYGPTGAGKSHTMFGCGKEPGIVYRSLRDILGDSDQD------ 163
Query: 180 DSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGGFGFGWSKSNASKVKLEVMG 239
GV TFVQVTVLE+YNEEIYDLLSTN G GW K ++KV+LEVMG
Sbjct: 164 --------GV-TFVQVTVLEVYNEEIYDLLSTNSSNN----LGIGWPKGASTKVRLEVMG 210
Query: 240 KKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSRSHCMVILDVPTVGGRLMLV 299
KKAKNA++ISG EAGKISKEI KVEKRRIVKSTLCN+RSSRSHC++ILDVPTVGGRLMLV
Sbjct: 211 KKAKNASFISGTEAGKISKEIVKVEKRRIVKSTLCNERSSRSHCIIILDVPTVGGRLMLV 270
Query: 300 DMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDS 359
DMAGSENI+QAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDS
Sbjct: 271 DMAGSENIDQAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDS 330
Query: 360 FEDDKSKILMILCASPDPKEIHKTISTLEYGAKAKCIVRGPHTPVKE----EDSSSTVIL 415
FEDDKSKILMILCASPDPKE+HKT+ TLEYGAKAKCIVRG HTP K+ ++S+S VIL
Sbjct: 331 FEDDKSKILMILCASPDPKEMHKTLCTLEYGAKAKCIVRGSHTPNKDKYGGDESASAVIL 390
Query: 416 GSRIAAMDEFIMKLQMENKLREKERNEAHKKLMKKEEEIAALR---TKVETAPASEEEIN 472
GSRIAAMDEFI+KLQ E K +EKERNEA K+L KKEEE+AALR T+ E +EEEI
Sbjct: 391 GSRIAAMDEFIIKLQSEKKQKEKERNEAQKQLKKKEEEVAALRSLLTQREACATNEEEIK 450
Query: 473 LKVNERTRHLRQELEKKLEECQRMTNEFVELERKRMEERILQQQEEVEILRKRLEEIELQ 532
KVNERT+ L+ EL+KKLEEC+RM EFVE+ER+RMEERI+QQQEE+E++R+RLEEIE+
Sbjct: 451 EKVNERTQLLKSELDKKLEECRRMAEEFVEMERRRMEERIVQQQEELEMMRRRLEEIEV- 509
Query: 533 LCSSSKQERKDENESKEMEPNGFMRKLLSVYKSTDDLGMVKSMDLDMDDQEPFLAREVIV 592
E + N E +GF ++L S+Y S DD GMVKSMDLDM D EP V
Sbjct: 510 -------EFRRSNGGSVDETSGFAKRLRSLY-SDDDPGMVKSMDLDMGDPEPVKQVWGAV 561
Query: 593 GMQG---ISPNQPCPNTLKNGVQEDAYVCAPNFGQKACLSTVYEEEGEEEAEQDHDKVEE 649
Q IS N N L+ E+ + + + CLSTV+EEE E
Sbjct: 562 SHQSSNTISSN--FTNLLQPKPSEN--MLTQMYPDRVCLSTVFEEE------------EV 605
Query: 650 DEEVEKEVIEEKRVCSVVNKSPKIED--------YTGADKENNGSNRLLRIHNIFTLCGN 701
+EE EK ++E+K +C + P + + GAD + + S+R LRI NIFTLCGN
Sbjct: 606 EEEEEKVIVEDKSICLITTPMPSLNSEGLGKENCFNGADDKESASSRRLRIQNIFTLCGN 665
Query: 702 QRELSQYG------TPIPTKKRSDESF---DFKCSPVKSSEKKDSVLRVSNKEN--LEAY 750
QRELSQ+ I + + D F K + E K++ + V +EN L+ Y
Sbjct: 666 QRELSQHSGQEEDQANIASPDKKDNQFFSITNKAEALAVEEAKENNISVDQRENGQLDIY 725
Query: 751 V 751
V
Sbjct: 726 V 726
>gb|AAL32294.1| phragmoplast-associated kinesin-related protein 2 [Arabidopsis
thaliana] gi|16973451|gb|AAL32293.1|
phragmoplast-associated kinesin-related protein 2
[Arabidopsis thaliana] gi|18414189|ref|NP_567426.1|
phragmoplast-associated kinesin-related protein 2
(PAKRP2) [Arabidopsis thaliana]
Length = 869
Score = 823 bits (2126), Expect = 0.0
Identities = 465/781 (59%), Positives = 557/781 (70%), Gaps = 85/781 (10%)
Query: 1 MAPTPSSS-SKQIHTTHLITPRSKHRLNFNGVKPAPTPPHPHPSPHPNFNNKDSPPEHPI 59
MAPTPSSS S Q T + TP++K RLNF+ +P+P+ + SPPEHP+
Sbjct: 1 MAPTPSSSRSNQTQYTLIRTPQTKQRLNFHS-----------KTPNPDGSKDPSPPEHPV 49
Query: 60 EVIARIRDYPDRKDKPLSVLQASSNSRSIRVKADFGYRDFTLDGVSVSEEEELDLFYKKF 119
EVI RIRDYPDRK+K S+LQ +++++++RV+AD GYRDFTLDGVS SE+E L+ FYKKF
Sbjct: 50 EVIGRIRDYPDRKEKSPSILQVNTDNQTVRVRADVGYRDFTLDGVSFSEQEGLEEFYKKF 109
Query: 120 VESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQAGIVYRALRDILGDGDTDSEGSDG 179
+E RI GVK+G+KCTIMMYGPTG+GKSHTMFGC K+ GIVYR+LRDILGD D D
Sbjct: 110 IEERIKGVKVGNKCTIMMYGPTGAGKSHTMFGCGKEPGIVYRSLRDILGDSDQD------ 163
Query: 180 DSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGGFGFGWSKSNASKVKLEVMG 239
GV TFVQVTVLE+YNEEIYDLLSTN G GW K ++KV+LEVMG
Sbjct: 164 --------GV-TFVQVTVLEVYNEEIYDLLSTNSSNN----LGIGWPKGASTKVRLEVMG 210
Query: 240 KKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSRSHCMVILDVPTVGGRLMLV 299
KKAKNA++ISG EAGKISKEI KVEKRRIVKSTLCN+RSSRSHC++ILDVPTVGGRLMLV
Sbjct: 211 KKAKNASFISGTEAGKISKEIVKVEKRRIVKSTLCNERSSRSHCIIILDVPTVGGRLMLV 270
Query: 300 DMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDS 359
DMAGSENI+QAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDS
Sbjct: 271 DMAGSENIDQAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDS 330
Query: 360 FEDDKSKILMILCASPDPKEIHKTISTLEYGAKAKCIVRGPHTPVKE----EDSSSTVIL 415
FEDDKSKILMILCASPDPKE+HKT+ TLEYGAKAKCIVRG HTP K+ ++S+S VIL
Sbjct: 331 FEDDKSKILMILCASPDPKEMHKTLCTLEYGAKAKCIVRGSHTPNKDKYGGDESASAVIL 390
Query: 416 GSRIAAMDEFIMKLQMENKLREKERNEAHKKLMKKEEEIAALR---TKVETAPASEEEIN 472
GSRIAAMDEFI+KLQ E K +EKERNEA K+L KKEEE+AALR T+ E +EEEI
Sbjct: 391 GSRIAAMDEFIIKLQSEKKQKEKERNEAQKQLKKKEEEVAALRSLLTQREACATNEEEIK 450
Query: 473 LKVNERTRHLRQELEKKLEECQRMTNEFVELERKRMEERILQQQEEVEILRKRLEEIELQ 532
KVNERT+ L+ EL+KKLEEC+RM EFVE+ER+RMEERI+QQQEE+E++R+RLEEIE+
Sbjct: 451 EKVNERTQLLKSELDKKLEECRRMAEEFVEMERRRMEERIVQQQEELEMMRRRLEEIEV- 509
Query: 533 LCSSSKQERKDENESKEMEPNGFMRKLLSVYKSTDDLGMVKSMDLDMDDQEPFLAREVIV 592
E + N E +GF ++L S+Y S DD GMVKSMDLDM D EP V
Sbjct: 510 -------EFRRSNGGSVDETSGFAKRLRSLY-SDDDPGMVKSMDLDMGDPEPVKQVWGAV 561
Query: 593 GMQG---ISPNQPCPNTLKNGVQEDAYVCAPNFGQKACLSTVYEEEGEEEAEQDHDKVEE 649
Q IS N N L+ E+ + + + CLSTV+EEE E
Sbjct: 562 SHQSSNTISSN--FTNLLQPKPSEN--MLTQMYPDRVCLSTVFEEE------------EV 605
Query: 650 DEEVEKEVIEEKRVCSVVNKSPKIED--------YTGADKENNGSNRLLRIHNIFTLCGN 701
+EE EK ++E+K +C + P + + GAD + + S+R LRI NIFTLCGN
Sbjct: 606 EEEEEKVIVEDKSICLITTPMPSLNSEGLGKENCFNGADDKESASSRRLRIQNIFTLCGN 665
Query: 702 QRELSQYG------TPIPTKKRSDESF---DFKCSPVKSSEKKDSVLRVSNKEN--LEAY 750
QRELSQ+ I + + D F K + E K++ + V +EN L+ Y
Sbjct: 666 QRELSQHSGQEEDQANIASPDKKDNQFFSITNKAEALAVEEAKENNISVDQRENGQLDIY 725
Query: 751 V 751
V
Sbjct: 726 V 726
>ref|XP_468330.1| putative phragmoplast-associated kinesin [Oryza sativa (japonica
cultivar-group)] gi|47847808|dbj|BAD21583.1| putative
phragmoplast-associated kinesin [Oryza sativa (japonica
cultivar-group)] gi|47497096|dbj|BAD19147.1| putative
phragmoplast-associated kinesin [Oryza sativa (japonica
cultivar-group)]
Length = 915
Score = 634 bits (1634), Expect = e-180
Identities = 399/771 (51%), Positives = 482/771 (61%), Gaps = 97/771 (12%)
Query: 3 PTPSSSSKQIHTTHLITPRSKHRLNFNGVKPAPTPPHPHPSPHPNFNNKDSPPEHPIEVI 62
P P + + TT L TP SKHRL+F PA TP + EHP+EVI
Sbjct: 10 PGPPPTPQAAMTTPLKTPASKHRLHF----PAMTPRNGGGG-----GAAAGGTEHPVEVI 60
Query: 63 ARIRDYPDRKDKPLSVLQASSNSRSIRVKADFG-YRDFTLDGVSVSEEEELDLFYKKFVE 121
RIR+ S L+ + ++RV+ D G RDFTLDGVSVSEEE+L+ FY++FV
Sbjct: 61 GRIRNLAAGAGGA-SALEIAGGGTAVRVRGDAGGCRDFTLDGVSVSEEEDLEGFYRRFVR 119
Query: 122 SRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQAGIVYRALRDIL-----GDGDTDSEG 176
SRI GV++G KCT+M+YGPTGSGKSHTMFGC+KQ GIVYRALRDIL G G G
Sbjct: 120 SRIEGVRVGAKCTVMVYGPTGSGKSHTMFGCAKQPGIVYRALRDILEGGGGGGGGVSGGG 179
Query: 177 SDGDS----SKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGGFGFGWSKSNASK 232
+GD GF +G+ FVQV VLEIYNEEIYDLL +G +K NA K
Sbjct: 180 GEGDGRGEDDAGFGMGL--FVQVAVLEIYNEEIYDLLVGSGAN----------AKGNAPK 227
Query: 233 VKLEVMGKKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSRSHCMVILDVPTV 292
+LEVMGKKAKNATYISGNEAGKIS+E+ KVEKRRIVKSTLCN+RSSRSHCM+ILDVP+V
Sbjct: 228 ARLEVMGKKAKNATYISGNEAGKISREVAKVEKRRIVKSTLCNERSSRSHCMIILDVPSV 287
Query: 293 GGRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKL 352
GGRLMLVDMAGSENIE AGQTGFEAKMQTAKINQGN ALKRVVESIANGDSHVPFRDSKL
Sbjct: 288 GGRLMLVDMAGSENIEAAGQTGFEAKMQTAKINQGNTALKRVVESIANGDSHVPFRDSKL 347
Query: 353 TMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAKAKCIVRGPHTPV---KEEDS 409
TMLLQDSFEDDKSKILMILCASPDPKE+HKT+STLEYGAKAKCI+R H K
Sbjct: 348 TMLLQDSFEDDKSKILMILCASPDPKELHKTVSTLEYGAKAKCIIRAAHAATPRDKMSSE 407
Query: 410 SSTVILGSRIAAMDEFIMKLQMENKLREKERNEAHKKLMKKEEEIAALRTKV-----ETA 464
S+ +L SRI AM++FI LQ ENKLREKERNEA L KKEEE+A LR K+ + A
Sbjct: 408 ESSTMLNSRIVAMNQFIYNLQKENKLREKERNEAQSVLRKKEEELAQLRAKLKLIEGQGA 467
Query: 465 PASEEEINLKVNERTRHLRQELEKKLEECQRMTNEFVELERKRMEERILQQQEEVEILRK 524
A EEEIN KV E+T+ LR EL K MEE++L+QQ+E+ L++
Sbjct: 468 AAKEEEINSKVMEKTQSLRTELMK-------------------MEEKMLRQQQELLALQQ 508
Query: 525 RLEEIELQLCSSSKQERKDENESKEMEPNGFMRKLLSVYKSTD-DLGMVKSMDLDMDDQE 583
RL+E+E +++ +++ + +L + D + M S+D DM DQ
Sbjct: 509 RLKEVE-----------REKPVQQDIIGGRLLARLSEMSARADQSMSMDMSIDFDMGDQP 557
Query: 584 PFLAREVI---VGMQGISPNQ-----PCPNTLKNGVQEDAYVCAPNFGQKACLSTVYEEE 635
+VI QG +Q C + ++ QED V + +K LSTV+ EE
Sbjct: 558 AAQDVKVIKEDTRKQGQIWSQANTAGSCTSAVE---QEDDVVRLSGYPEKVVLSTVF-EE 613
Query: 636 GEEEAEQDHDKVEEDEEVEKEVIEEKRVCSVVNKSPKIEDYTGADKENNGSNRLLRIHNI 695
G+EE ++D +EEV KEV+EE S V + P E A + N RI NI
Sbjct: 614 GDEEEDKDSG---VEEEVCKEVVEE----SYVMQQPLAEPEDPATRNN-------RIQNI 659
Query: 696 FTLCGNQRELSQYGTPIPTKKRSDESFDFKCSPVKSSEKKDSVLRVSNKEN 746
F LCGN REL++ K DE+ + K+ RV EN
Sbjct: 660 FRLCGNHRELAKKVQSPAKKAFGDENNEPAKQTFGDENKQQPAKRVFGDEN 710
>emb|CAD98462.1| kinesin-like boursin, possible [Cryptosporidium parvum]
Length = 1184
Score = 160 bits (406), Expect = 1e-37
Identities = 179/684 (26%), Positives = 307/684 (44%), Gaps = 120/684 (17%)
Query: 59 IEVIARIRDYPDRKDKPLS---VLQASSNSRSIRVKAD--------FGYRDFTLDGVSVS 107
++VI R R +++ K S VLQ +S+ I V + + FT DGV S
Sbjct: 18 VKVIVRCRPLTEQEKKDPSNSNVLQVKPDSKEIVVSHQSLSRKFDSYSTKLFTFDGVCGS 77
Query: 108 EEEELDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQ------------ 155
+ +LF K++V ++ V LG CTI YG TG+GK++TM G K+
Sbjct: 78 FTSQRELF-KQYVVPIVDEVLLGFNCTIFAYGQTGTGKTYTMEGDMKEYLESSNLELTEH 136
Query: 156 AGIVYRALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGG 215
AGI+ RA++ I +S+ ++ GVR V+ LEIYNEE+ DLLS
Sbjct: 137 AGIIPRAVQLIFER--LESQYTE--------YGVR----VSYLEIYNEELSDLLSDE--- 179
Query: 216 GGGGGFGFGWSKSNASKVKLEVMGKKAKNATYISG---NEAGKISKEIQKVEKRRIVKST 272
K N ++ ++ GK+ N + N+A I + ++R T
Sbjct: 180 -----------KLNL-RIYDDIAGKRGLNVDRLEEIPVNKAQDILNILSTAVRKRRTAET 227
Query: 273 LCNDRSSRSHCMVILDVPTVG-----------GRLMLVDMAGSENIEQAGQTGFEAKMQT 321
L N SSRSHC+ + + T G+L LVD+AGSENI+++G + + +
Sbjct: 228 LLNKSSSRSHCIFTITIHTKETNIDGEDVLKVGKLNLVDLAGSENIQRSGANAVKDRAKE 287
Query: 322 A-KINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEI 380
A INQ + L RV+ ++ S+VP+RDSKLT LLQDS ++K +I + +
Sbjct: 288 AGMINQSLLTLGRVINALVEHSSYVPYRDSKLTRLLQDSL-GGRTKTCIIATITASSIYL 346
Query: 381 HKTISTLEYGAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENK-----L 435
+T++TL+Y +AK I + PV + + V++ +++ +LQ + L
Sbjct: 347 EETLNTLDYAHRAKNI---KNMPVVNQKMTKKVMIREMNCEIEKLKQELQCNREKNGVYL 403
Query: 436 REKERNEAHKKLMKKEEEIAALRTKVETAPASEEEINLKVNERTRHLRQE-LEKKLEECQ 494
+ NE KL + EI + ++++ A +E+ VN T L ++ L + +
Sbjct: 404 PLSQFNEMESKLQSQANEIIEMESELQNQHALYKEMESTVNSLTDQLNEKSLRVRAGDFA 463
Query: 495 RM-TNEFVELERKRMEERILQQQEEVEILRK-----------------RLEEIELQLCSS 536
+ ++ +L ++ ++ +Q + +E + K +L +++ QL S
Sbjct: 464 NIHVSKHAKLYHEKYKDLFMQMNQSLENIGKLHRKIIGSELFQSSQNTQLFKLQQQLISD 523
Query: 537 SKQERKDENESKEMEPNGFMRKLLSVYKSTDDLGMVKSMDLDMDDQEPFLAREVIVGMQG 596
+ K E E+ N + LL +K+ L V+ + M D L VI +
Sbjct: 524 IQLTEKKTRECLEILYNDLNQDLLVHWKNKSQLSHVEI--IGMIDTGKKLCNNVISMLS- 580
Query: 597 ISPNQPCPNTLKNGVQEDAYVCAPNFGQ--KACLSTVYE---------EEGEEEAEQDHD 645
++L+ + E+ Y N + K CL T E +G + D +
Sbjct: 581 --------DSLEQSLTEELYAGMNNAKENLKKCLKTQDEITTRIKDEIRKGLVDCSTDTE 632
Query: 646 KVEED--EEVEKEVIEEKRVCSVV 667
K+ D ++E+ + +R S+V
Sbjct: 633 KISMDITNQLERLKLSNERCQSLV 656
>ref|XP_666160.1| kinesin-like boursin [Cryptosporidium hominis]
gi|54657099|gb|EAL35931.1| kinesin-like boursin
[Cryptosporidium hominis]
Length = 1184
Score = 158 bits (400), Expect = 5e-37
Identities = 165/618 (26%), Positives = 279/618 (44%), Gaps = 110/618 (17%)
Query: 59 IEVIARIRDYPDRKDKPLS---VLQASSNSRSIRVKAD--------FGYRDFTLDGVSVS 107
++VI R R +++ K S VLQ +S+ I V + + FT DGV S
Sbjct: 18 VKVIVRCRPLTEQEKKDPSNSNVLQVKPDSKEIVVSHQSLSRKFDSYSTKLFTFDGVCGS 77
Query: 108 EEEELDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQ------------ 155
+ +LF K++V ++ V LG CTI YG TG+GK++TM G K+
Sbjct: 78 FTSQRELF-KQYVVPIVDEVLLGFNCTIFAYGQTGTGKTYTMEGDMKEYLESSNLELTEH 136
Query: 156 AGIVYRALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGG 215
AGI+ RA++ I +S+ ++ GVR V+ LEIYNEE+ DLLS
Sbjct: 137 AGIIPRAVQLIFER--LESQYTE--------YGVR----VSYLEIYNEELSDLLSDE--- 179
Query: 216 GGGGGFGFGWSKSNASKVKLEVMGKKAKNATYISG---NEAGKISKEIQKVEKRRIVKST 272
K N ++ ++ GK+ N + N+A I + ++R T
Sbjct: 180 -----------KLNL-RIYDDIAGKRGLNVDRLEEIPVNKAQDILNILSTAVRKRRTAET 227
Query: 273 LCNDRSSRSHCMVILDVPTVG-----------GRLMLVDMAGSENIEQAGQTGFEAKMQT 321
L N SSRSHC+ + + T G+L LVD+AGSENI+++G + + +
Sbjct: 228 LLNKSSSRSHCIFTITIHTKETNIDGEDVLKVGKLNLVDLAGSENIQRSGANAVKDRAKE 287
Query: 322 A-KINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEI 380
A INQ + L RV+ ++ S+VP+RDSKLT LLQDS ++K +I + +
Sbjct: 288 AGMINQSLLTLGRVINALVEHSSYVPYRDSKLTRLLQDSL-GGRTKTCIIATITASSIYL 346
Query: 381 HKTISTLEYGAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENK-----L 435
+T++TL+Y +AK I + PV + + V++ +++ +LQ + L
Sbjct: 347 EETLNTLDYAHRAKNI---KNMPVVNQKMTKKVMIREMNCEIEKLKQELQCNREKNGVYL 403
Query: 436 REKERNEAHKKLMKKEEEIAALRTKVETAPASEEEINLKVNERTRHLRQE-LEKKLEECQ 494
+ NE KL + EI + ++++ A +E+ VN T L ++ L + +
Sbjct: 404 PLSQFNEMESKLQSQANEIIEMESELQNQHALYKEMESTVNSLTDQLNEKSLRVRAGDFA 463
Query: 495 RM-TNEFVELERKRMEERILQQQEEVEILRK-----------------RLEEIELQLCSS 536
+ ++ +L ++ ++ +Q + +E + K +L +++ QL S
Sbjct: 464 NIHVSKHAKLYHEKYKDLFMQMNQSLENIGKLHRKIIGSELFQSSQNTQLFKLQQQLISD 523
Query: 537 SKQERKDENESKEMEPNGFMRKLLSVYKSTDDL------GMVKS--------MDLDMDDQ 582
+ K E E+ N + LL +K+ L GM+ + + + D
Sbjct: 524 IQLTEKKTRECLEIIYNDLNQDLLVHWKNKSQLSHGEIIGMIDTGKKLCNNVISMLSDSL 583
Query: 583 EPFLAREVIVGMQGISPN 600
E L E+ GM N
Sbjct: 584 EQSLTEELYAGMNNAKEN 601
>ref|XP_649446.1| kinesin family protein [Entamoeba histolytica HM-1:IMSS]
gi|56465885|gb|EAL44056.1| kinesin family protein
[Entamoeba histolytica HM-1:IMSS]
Length = 863
Score = 156 bits (395), Expect = 2e-36
Identities = 168/657 (25%), Positives = 304/657 (45%), Gaps = 100/657 (15%)
Query: 59 IEVIARIRDYPDRKDKPLSVLQASSNSRSIRVKADFGYRDFTLDGVSVSEEEELDLFYKK 118
I+V R R P ++++ ++ +Q + + + K + D V S+ + + F
Sbjct: 3 IKVYVRCR--PGKENEHIANIQMDNQNLIVSGKK------YEFDQVFSSQVNQ-NQFCDS 53
Query: 119 FVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQAGIVYRALRDILGDGDTDSEGSD 178
+E I V G CT YG TG+GK++TM G + G++ R + ++ +
Sbjct: 54 ALEGFIGKVIDGYNCTFFAYGQTGTGKTYTMEGEEENEGVIPRVINELFMTLEKR----- 108
Query: 179 GDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGGFGFGWSKSNASKVKLEVM 238
G+R ++VT +EIYNE++YDLLS K A K + E
Sbjct: 109 ---------GLRYRMRVTHVEIYNEKVYDLLSDE-------------RKELAIKERKEGN 146
Query: 239 GKKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSRSHCMVILDVP-------- 290
G ++AT ++ + + + + K +R +T N SSRSHC+ + V
Sbjct: 147 GASPESATELTVTKEN-VHQILAKSSGQRRTAATDINTNSSRSHCIFTVTVQIIRDSDFE 205
Query: 291 ---TVGGRLMLVDMAGSENIEQAGQTGFEAK-MQTAKINQGNIALKRVVESIANGDSHVP 346
V GR+ VD+AGSEN ++AG G + K ++ INQ +AL RV+ ++ GDS++P
Sbjct: 206 GDFVVPGRIHCVDLAGSENSKRAGIIGDKVKQLEGMAINQSLLALNRVIIGVSKGDSYIP 265
Query: 347 FRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAKAKCIVRGPHTPVKE 406
FR S LT +LQD+ S M+ SP +++ +T+STLEY + K I +TP +
Sbjct: 266 FRSSPLTRILQDAL-GGASITAMVATISPAQEDLEETLSTLEYAKRVKTI---KNTPKQN 321
Query: 407 EDSSSTVILGSRIAAMDEFIMKLQMENKLREKERNEAHKKLMKKEE--------EIAAL- 457
+ L ++ + + KL E K EK E ++++ E E+ AL
Sbjct: 322 ASVRKDIFLQKKLDKIIN-LKKLITEGKRNEKPLTEEEIEILQMEVAKVNQQNLEMQALF 380
Query: 458 -RTKVETAPASEE------EINLKVNERTRHLRQELEKKLEECQRMTNEFVELE--RKRM 508
+ K E + A ++ I+LK + L+ L K ++ + +T + +LE +R
Sbjct: 381 AQAKGEVSAAQDQLKSAMNYIHLK-RQNEIELKTMLGKVVKSIEILTKQSRKLEDIYERN 439
Query: 509 EERILQQQEEVEILRKRLEEIELQLCSSSKQERKDENESKEMEPNGFMRKLLSVYKSTDD 568
E+I Q + ++ +++ EIE ++ ++E+K+ S + N + +D
Sbjct: 440 NEKIKQNDDIIKQIQENENEIEKKITEQIEEEKKEIQMSWNVIENKI--------EESDK 491
Query: 569 LGMVKSMDLDMDDQEPFLAREVIVGMQGISPNQPCPNTLKNGVQEDAYVCAPNFGQKACL 628
+++ ++ + +Q+ ++ ++ + M+ C LK + V +
Sbjct: 492 KSIIEGINFEEFNQQ-WIYKDTLGRMEIF-----CKEGLKQEKENKENV----------I 535
Query: 629 STVYEEEGEEEAEQDHDKVEEDEEVEKEVIEEKRVCSVVNKSPK--IEDYTGADKEN 683
+ E+E E Q+ K ED E +KE+ E++ +VN+S K IE+ KE+
Sbjct: 536 KILKEKENINEEFQEKLKTIEDIE-QKEIKEKEEEMEIVNESLKGVIEELKKKKKES 591
Score = 40.0 bits (92), Expect = 0.28
Identities = 48/185 (25%), Positives = 86/185 (45%), Gaps = 23/185 (12%)
Query: 376 DPKEIHKTISTLEYGAKA---KCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFI---MKL 429
+ KEI + + +E + K I+ G + E+ + I + M+ F +K
Sbjct: 471 EKKEIQMSWNVIENKIEESDKKSIIEG----INFEEFNQQWIYKDTLGRMEIFCKEGLKQ 526
Query: 430 QMENK------LREKER-NEAHKKLMKKEEEIAALRTKVETAPASEEEINLKVNERTRHL 482
+ ENK L+EKE NE ++ +K E+I K + EEE+ + VNE + +
Sbjct: 527 EKENKENVIKILKEKENINEEFQEKLKTIEDIEQKEIKEK-----EEEMEI-VNESLKGV 580
Query: 483 RQELEKKLEECQRMTNEFVELERKRMEERILQQQEEVEILRKRLEEIELQLCSSSKQERK 542
+EL+KK +E Q ++ E KR E+R + E ++K++ E + L S +
Sbjct: 581 IEELKKKKKESQEFEEIIIQNEVKRNEKRKEMIKSLNENIQKQINENKKILIQSIENMGN 640
Query: 543 DENES 547
+ N +
Sbjct: 641 EINNT 645
>ref|NP_850281.1| kinesin motor protein-related [Arabidopsis thaliana]
Length = 1039
Score = 152 bits (385), Expect = 3e-35
Identities = 159/623 (25%), Positives = 277/623 (43%), Gaps = 88/623 (14%)
Query: 19 TPRSKHRLNFNGVKPAPTP------PHPHPSPHPNFNNKDSPPEHPIEVIARIRDYPDRK 72
TP R + GV P+P P P N ++D+ E ++VI R + + +
Sbjct: 4 TPEVVSRKSGVGVIPSPAPFLTPRLERRRPDSFSNRLDRDNK-EVNVQVILRCKPLSEEE 62
Query: 73 DKPL--SVLQASSNSRSIRVKADFGYRD----FTLDGVSVSEEEELDLFYKKFVESRING 126
K V+ + R + V + F D V + ++ + Y + + ++
Sbjct: 63 QKSSVPRVISCNEMRREVNVLHTIANKQVDRLFNFDKVFGPKSQQRSI-YDQAIAPIVHE 121
Query: 127 VKLGDKCTIMMYGPTGSGKSHTMFGCSK--------QAGIVYRALRDILGDGDTDSEGSD 178
V G CT+ YG TG+GK++TM G + +AG++ RA+R I DT E +
Sbjct: 122 VLEGFSCTVFAYGQTGTGKTYTMEGGMRKKGGDLPAEAGVIPRAVRHIF---DT-LEAQN 177
Query: 179 GDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGGFGFGWSKSNASKVKLEV- 237
D S ++VT LE+YNEE+ DLL+ + S+S+ K + +
Sbjct: 178 ADYS----------MKVTFLELYNEEVTDLLAQDDS-----------SRSSEDKQRKPIS 216
Query: 238 MGKKAKNATYISGNE------AGKISKEIQKVEKRRIVKSTLCNDRSSRSHCMVILDV-- 289
+ + K + + G E A I +++ +R TL N RSSRSH + + V
Sbjct: 217 LMEDGKGSVVLRGLEEEVVYSANDIYALLERGSSKRRTADTLLNKRSSRSHSVFTITVHI 276
Query: 290 --PTVG-------GRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESIAN 340
++G G+L LVD+AGSENI ++G A+ + +IN+ + L RV+ ++
Sbjct: 277 KEESMGDEELIKCGKLNLVDLAGSENILRSGARDGRAR-EAGEINKSLLTLGRVINALVE 335
Query: 341 GDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAKAKCIVRGP 400
SHVP+RDSKLT LL+DS K+K +I SP + +T+STL+Y +AK I P
Sbjct: 336 HSSHVPYRDSKLTRLLRDSL-GGKTKTCIIATISPSAHSLEETLSTLDYAYRAKNIKNKP 394
Query: 401 HTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENKLR-EKERNEAHKKLMKKEEEIAALRT 459
K L + D ++ +M+ +R +++N + + +E +
Sbjct: 395 EANQK---------LSKAVLLKDLYLELERMKEDVRAARDKNGVYIAHERYTQEEVEKKA 445
Query: 460 KVETAPASEEEINLK---------VNERTRHLRQELEKKLEECQRMTNEFVE--LERKRM 508
++E E E+NL + E + ++E L++C+R + + L+ K
Sbjct: 446 RIERIEQLENELNLSESEVSKFCDLYETEKEKLLDVESDLKDCKRNLHNSNKDLLDLKEN 505
Query: 509 EERILQQQEEVEILRKRLEEIELQLCSSSKQERKDENESKEMEPNGFMRKLLSVYKSTDD 568
+++ + +E E++ R++ E L +K R D + + F R +D+
Sbjct: 506 YIQVVSKLKEKEVIVSRMKASETSLIDRAKGLRCDLQHASNDINSLFTRLDQKDKLESDN 565
Query: 569 LGMVKSMDLDMDDQEPFLAREVI 591
M+ +D L R V+
Sbjct: 566 QSMLLKFGSQLDQNLKDLHRTVL 588
>gb|AAH82827.1| LOC397908 protein [Xenopus laevis]
Length = 1067
Score = 151 bits (382), Expect = 7e-35
Identities = 160/556 (28%), Positives = 251/556 (44%), Gaps = 94/556 (16%)
Query: 59 IEVIARIRDYP--DRKDKPLSVLQASSNSRSIRVKAD-----FGYRDFTLDGVSVSEEEE 111
I+V+ R R + +RK SVL+ S + + V+ G + +T D V ++
Sbjct: 19 IQVVVRCRPFNQLERKASSHSVLECDSQRKEVYVRTGGINDKLGKKTYTFDMVFGPAAKQ 78
Query: 112 LDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQ-----------AGIVY 160
+D+ Y+ V ++ V +G CTI YG TG+GK+ TM G AGI+
Sbjct: 79 IDV-YRSVVCPILDEVIMGYNCTIFAYGQTGTGKTFTMEGERSSDEEFTWEQDPLAGIIP 137
Query: 161 RALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGG 220
R L I + SE G V+V++LEIYNEE++DLLS + G
Sbjct: 138 RTLHQIF---EKLSEN-----------GTEFSVKVSLLEIYNEELFDLLSPSPDVG---- 179
Query: 221 FGFGWSKSNASKVKLEVMGKKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSR 280
+ G K IS + ++ +++ RR STL N SSR
Sbjct: 180 -----ERLQMFDDPRNKRGVIIKGLEEISVHNKDEVYHILERGAARRKTASTLMNAYSSR 234
Query: 281 SHCMVILDV----PTVGG-------RLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNI 329
SH + + + TV G +L LVD+AGSENI ++G A+ + INQ +
Sbjct: 235 SHSVFAVTIHMKETTVDGEELVKIGKLNLVDLAGSENIGRSGAVDKRAR-EAGNINQSLL 293
Query: 330 ALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEY 389
L RV+ ++ H+P+R+SKLT +LQDS ++K +I SP + +T+STL+Y
Sbjct: 294 TLGRVITALVERTPHIPYRESKLTRILQDSL-GGRTKTSIIATVSPASINLEETVSTLDY 352
Query: 390 GAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENKLREKERNEAHKKLMK 449
+AK I+ P K + K KE E ++L
Sbjct: 353 ANRAKSIMNKPEVNQK-------------------------LTKKALIKEYTEEIERL-- 385
Query: 450 KEEEIAALRTKVETAPASE--EEINLKVNERTRHLRQELEK--KLEECQRMTNEFVELER 505
+ E+AA R K +SE E++ KV + + + EK +EE + +E +
Sbjct: 386 -KRELAAAREKNGVYLSSENYEQLQGKVLSQEEMITEYTEKITAMEEELKSISELFADNK 444
Query: 506 KRMEE--RILQ-QQEEVEILRKRLEEIELQLCSSSKQERKDENESKEMEPNGFMRKLLSV 562
K +EE ILQ +++E+E + L+E + QL S E K++ +G KLLS
Sbjct: 445 KELEECTTILQCKEKELEETQNHLQESKEQLAQESFVVSAFETTEKKL--HGTANKLLST 502
Query: 563 YKST--DDLGMVKSMD 576
+ T D G+ + +D
Sbjct: 503 VRETTRDVSGLHEKLD 518
>gb|AAH42218.1| MGC52588 protein [Xenopus laevis]
Length = 1067
Score = 150 bits (380), Expect = 1e-34
Identities = 155/552 (28%), Positives = 249/552 (45%), Gaps = 86/552 (15%)
Query: 59 IEVIARIRDYP--DRKDKPLSVLQASSNSRSIRVKAD-----FGYRDFTLDGVSVSEEEE 111
I+V+ R R + +RK SVL+ S + + V+ G + +T D V ++
Sbjct: 19 IQVVVRCRPFNQLERKASSHSVLECESQRKEVCVRTGGINDKLGKKTYTFDMVFGPAAKQ 78
Query: 112 LDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQ-----------AGIVY 160
+D+ Y+ V ++ V +G CTI YG TG+GK+ TM G AGI+
Sbjct: 79 IDV-YRSVVCPILDEVIMGYNCTIFAYGQTGTGKTFTMEGERSSDEEFTWEQDPLAGIIP 137
Query: 161 RALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGG 220
R L I + SE G V+V++LEIYNEE++DLLS + G
Sbjct: 138 RTLHQIF---EKLSEN-----------GTEFSVKVSLLEIYNEELFDLLSPSPDVG---- 179
Query: 221 FGFGWSKSNASKVKLEVMGKKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSR 280
+ G K IS + ++ + +++ +R STL N SSR
Sbjct: 180 -----ERLQMFDDPRNKRGVIIKGLEEISVHNKDEVYQILERGAAKRKTASTLMNAYSSR 234
Query: 281 SHCMVILDV----PTVGG-------RLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNI 329
SH + + + T+ G +L LVD+AGSENI ++G A+ + INQ +
Sbjct: 235 SHSVFSVTIHMKETTIDGEELVKIGKLNLVDLAGSENIGRSGAVDKRAR-EAGNINQSLL 293
Query: 330 ALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEY 389
L RV+ ++ H+P+R+SKLT +LQDS ++K +I SP + +T+STLEY
Sbjct: 294 TLGRVITALVERAPHIPYRESKLTRILQDSL-GGRTKTSIIATVSPASINLEETMSTLEY 352
Query: 390 GAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMEN--KLREKERNEAHKKL 447
++AK I+ P K + I + + + +N L + + K+
Sbjct: 353 ASRAKNIMNKPEVNQKLTKKALIKEYTEEIERLKRELATAREKNGVYLSNENYEQLQGKV 412
Query: 448 MKKEEEIAALRTKVETAPASEEEINLKVNERTRHLRQELEKKLEECQRMTNEFVELERKR 507
+ +EE I K+ A EEEI +R L + +K+LEEC
Sbjct: 413 LSQEEVITEYSEKI---AAMEEEI-----KRIGELFADNKKELEEC-------------- 450
Query: 508 MEERILQ-QQEEVEILRKRLEEIELQLCSSSKQERKDENESKEMEPNGFMRKLLSVYKST 566
ILQ +++E+E + L+E + QL + E K++ +G KLLS + T
Sbjct: 451 --TTILQCKEKELEATQNNLQESKEQLAQEAFVVSAMETTEKKL--HGTANKLLSTVRET 506
Query: 567 --DDLGMVKSMD 576
D G+ + +D
Sbjct: 507 TRDVSGLHEKLD 518
>emb|CAB01170.1| Hypothetical protein F23B12.8 [Caenorhabditis elegans]
gi|3875617|emb|CAB01153.1| Hypothetical protein F23B12.8
[Caenorhabditis elegans] gi|23491748|dbj|BAC19818.1|
kinesin like protein KLP-14 [Caenorhabditis elegans]
gi|17557420|ref|NP_506582.1| BiMC Kinesin related,
kinesin-like protein (108.3 kD) (bmk-1) [Caenorhabditis
elegans] gi|9945020|gb|AAG03081.1| bimC kinesin BMK-1
[Caenorhabditis elegans] gi|7499724|pir||T20621
hypothetical protein F23B12.8 - Caenorhabditis elegans
Length = 958
Score = 150 bits (379), Expect = 1e-34
Identities = 173/672 (25%), Positives = 277/672 (40%), Gaps = 123/672 (18%)
Query: 51 KDSPPEHPIEVIARIRDY--PDRKDKPLSVLQASSNSRSIRVKA-DFGYRDFTLDGVSVS 107
K S P + V RIR +R +K +V++ ++I +K FG T D +
Sbjct: 6 KHSEPTSNLRVAVRIRPMNGTERSEKCTNVVKVDKGKQAIELKGKSFGPFFRTYDPDTTQ 65
Query: 108 EEEELDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQA---------GI 158
EE Y V S+I V G CT+ YG TG+GK+ TM G A GI
Sbjct: 66 EE-----IYSDLVSSQIKKVIAGFNCTVFAYGQTGTGKTFTMEGGRTDAKSSTDDPTTGI 120
Query: 159 VYRALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGG 218
+ RA+ DI + C ++V+ +E+YNEE++DLL+
Sbjct: 121 IPRAVEDIFEQLER-------------CGCEEYSLRVSYIELYNEELFDLLA-------- 159
Query: 219 GGFGFGWSKSNASKVKLEVMGKKAKNATYISGNEA------GKISKEIQKVEKRRIVKST 272
S N + +L + K +SG E + K +Q ++R +T
Sbjct: 160 -------STDNEDRERLRIFDDPNKKGVIVSGVEEVPVRNRSDVFKLLQLGAEKRRTAAT 212
Query: 273 LCNDRSSRSHCMVILDV-----PTVG------GRLMLVDMAGSENIEQAGQTGFEAKMQT 321
L N SSRSH + +++V T G G+L LVD+AGSENI ++G G AK +
Sbjct: 213 LMNMHSSRSHSLFMVNVVIRENTTTGEELVKQGKLNLVDLAGSENIGRSGAQGNRAK-EA 271
Query: 322 AKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIH 381
INQ + L RV+ + H+P+R+SKLT LLQDS + +I SP
Sbjct: 272 GSINQSLLTLGRVIRLLTTNGQHIPYRESKLTRLLQDSL-GGSTITSLIATLSPSSSNFE 330
Query: 382 KTISTLEYGAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENKLREKERN 441
++ STLEY +A I P + ++ K KE +
Sbjct: 331 ESQSTLEYAMRAANIKNKP-------------------------VCNTKLSKKTILKEYS 365
Query: 442 EAHKKLMKKEEEIAALRTKVETAPASEEEINLKVNERTRHLRQELEKKLEECQRMTNEFV 501
+ +KL +++ A + V + S +E K +E+ + L Q L+ ++ + T
Sbjct: 366 DEIEKL-RRDLRAAREKNGVIISQESHDEFQ-KNSEKVQELEQHLDNAVDRLRIFTE--- 420
Query: 502 ELERKRMEERILQQQEEVEILRKRLEEIELQLCSSSKQERKDENE--SKEMEPNGFMRKL 559
++ M+E+ Q E L RL + ++ + ++E D N+ +K +E G M
Sbjct: 421 --DQMHMDEQYRQLYERKGELEDRLRD-RIREMAVKEKELADTNDVLNKHIEVIGLMH-- 475
Query: 560 LSVYKSTDDL----GMVKSMDLDM-------DDQEPFLAREVIV---GMQGISPNQPCPN 605
S K+ D L G M D+ D+ E R IV + +S +
Sbjct: 476 TSALKAFDQLTTSQGAADEMQTDLRAFWRKVDEMERADDRNKIVIDHFSKKMSDFIDTTS 535
Query: 606 TLKNGVQEDAYVCAPNFGQKACLSTVYEEEGEEEAEQDHDKVE--------EDEEVEKEV 657
L N ++ D + + + C E + E +E + E EK+V
Sbjct: 536 KLTNSLKADGDLLSKKLSENICPHVKSMESAHRDIESTAMDMETHAICIMKKSSEAEKKV 595
Query: 658 IEEKRVCSVVNK 669
I+ S+ N+
Sbjct: 596 IDNMTAISMENR 607
>emb|CAA50695.1| kinesin like protein [Xenopus laevis]
gi|2497521|sp|Q91783|EG52_XENLA Kinesin-related motor
protein Eg5 2
Length = 1067
Score = 150 bits (379), Expect = 1e-34
Identities = 153/552 (27%), Positives = 248/552 (44%), Gaps = 86/552 (15%)
Query: 59 IEVIARIRDYP--DRKDKPLSVLQASSNSRSIRVKAD-----FGYRDFTLDGVSVSEEEE 111
I+V+ R R + +RK SVL+ S + + V+ G + +T D V ++
Sbjct: 19 IQVVVRCRPFNQLERKASSHSVLECESQRKEVCVRTGEVNDKLGKKTYTFDMVFGPAAKQ 78
Query: 112 LDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQ-----------AGIVY 160
+D+ Y+ V ++ V +G CTI YG TG+GK+ TM G AGI+
Sbjct: 79 IDV-YRSVVCPILDEVIMGYNCTIFAYGQTGTGKTFTMEGERSSDEEFTWEQDPLAGIIP 137
Query: 161 RALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGG 220
R L I K +G V+V++LEIYNEE++DLLS + G
Sbjct: 138 RTLHQIF--------------EKLSEIGTEFSVKVSLLEIYNEELFDLLSPSPDVG---- 179
Query: 221 FGFGWSKSNASKVKLEVMGKKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSR 280
+ G K IS + ++ + +++ +R STL N SSR
Sbjct: 180 -----ERLQMFDDPRNKRGVIIKGLEEISVHNKDEVYQILERGAAKRKTASTLMNAYSSR 234
Query: 281 SHCMVILDV----PTVGG-------RLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNI 329
SH + + + T+ G +L LVD+AGSENI ++G A+ + INQ +
Sbjct: 235 SHSVFSVTIHMKETTIDGEELVKIGKLNLVDLAGSENIGRSGAVDKRAR-EAGNINQSLL 293
Query: 330 ALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEY 389
L RV+ ++ H+P+R+SKLT +LQDS ++K +I SP + +T+STL+Y
Sbjct: 294 TLGRVITALVERAPHIPYRESKLTRILQDSL-GGRTKTSIIATVSPASINLEETMSTLDY 352
Query: 390 GAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMEN--KLREKERNEAHKKL 447
++AK I+ P K + I + + + +N L + + K+
Sbjct: 353 ASRAKNIMNKPEVNQKLTKKALIKEYTEEIERLKRELATAREKNGVYLSNENYEQLQGKV 412
Query: 448 MKKEEEIAALRTKVETAPASEEEINLKVNERTRHLRQELEKKLEECQRMTNEFVELERKR 507
+ +EE I K+ A EEEI +R L + +K+LEEC
Sbjct: 413 LSQEEMITEYSEKI---AAMEEEI-----KRIGELFADNKKELEEC-------------- 450
Query: 508 MEERILQ-QQEEVEILRKRLEEIELQLCSSSKQERKDENESKEMEPNGFMRKLLSVYKST 566
ILQ +++E+E + L+E + QL + E K++ +G KLLS + T
Sbjct: 451 --TTILQCKEKELEATQNNLQESKEQLAQEAFVVSAMETTEKKL--HGTANKLLSTVRET 506
Query: 567 --DDLGMVKSMD 576
D G+ + +D
Sbjct: 507 TRDVSGLHEKLD 518
>emb|CAG78210.1| unnamed protein product [Yarrowia lipolytica CLIB99]
gi|50555986|ref|XP_505401.1| hypothetical protein
[Yarrowia lipolytica]
Length = 929
Score = 150 bits (379), Expect = 1e-34
Identities = 133/441 (30%), Positives = 212/441 (47%), Gaps = 81/441 (18%)
Query: 55 PEHPIEVIARIRDYPDRKD-KPLSVLQASSNSRSIRVKADFGY-----RDFTLDGVSVSE 108
P ++V+ R R +R+ + SV+ +S + + + G + +T D V E
Sbjct: 22 PSTGMKVLVRCRGRNERETTENSSVVVKTSGHKGREITIEGGPVAHTGKTYTFDRVFGPE 81
Query: 109 EEELDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFG---------CSKQAGIV 159
++ +F + V S ++ + G CTI YG TG+GK++TM G AGIV
Sbjct: 82 SDQGMIF--EAVSSSLDEMLQGYNCTIFAYGQTGTGKTYTMTGDFNLDERGEAVSNAGIV 139
Query: 160 YRALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGG 219
RAL ++ GS G++S V+++ +E+YNEE+ DLLS+ G
Sbjct: 140 PRALVELF----KRLSGSAGENS----------VKLSYVELYNEELRDLLSSQG------ 179
Query: 220 GFGFGWSKSNASKVKLEVMGKKAKNATYISGNEAGKIS------KEIQKVEKRRIVKSTL 273
KL + + K T + G E + K +Q+ RR V +T
Sbjct: 180 -----------DTKKLRIFEEPGKKGTVVQGLEEAYVRSCTEAMKVLQEGFTRRQVAATK 228
Query: 274 CNDRSSRSHCMVILDVPTVG-----------GRLMLVDMAGSENIEQAGQTGFEAKMQTA 322
CND SSRSH ++ + + T G+L LVD+AGSEN+ ++G A+ +
Sbjct: 229 CNDMSSRSHSVLTITLSTKEYTADGQEYLRTGKLNLVDLAGSENVGRSGAENMRAR-EAG 287
Query: 323 KINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHK 382
INQ + L RV+ S+ +G H+P+R+SKLT LLQ+S ++K ++I SP I +
Sbjct: 288 SINQSLLTLGRVINSLVDGTLHIPYRESKLTRLLQESL-GGRTKTVIIATVSPARVSIDE 346
Query: 383 TISTLEYGAKAKCIVRGPHTPVKEEDSSSTVILG------SRIAAMDEFIMK-----LQM 431
TISTLEY +AK I ++PV E +S V + +R+ A E K L
Sbjct: 347 TISTLEYSHRAKNI---KNSPVVNETTSKQVFIKDYVDEIARLRADLESTRKGQGVFLSQ 403
Query: 432 ENKLREKERNEAHKKLMKKEE 452
E+ + NE+HK + +++
Sbjct: 404 ESYQLLLDENESHKVTINEQK 424
>emb|CAA37950.1| kinesine [Xenopus laevis] gi|119217|sp|P28025|EG51_XENLA
Kinesin-related motor protein Eg5 1
Length = 1060
Score = 149 bits (377), Expect = 3e-34
Identities = 159/556 (28%), Positives = 251/556 (44%), Gaps = 94/556 (16%)
Query: 59 IEVIARIRDYP--DRKDKPLSVLQASSNSRSIRVKAD-----FGYRDFTLDGVSVSEEEE 111
I+V+ R R + +RK SVL+ S + + V+ G + +T D V ++
Sbjct: 12 IQVVVRCRPFNQLERKASSHSVLECDSQRKEVYVRTGEVNDKLGKKTYTFDMVFGPAAKQ 71
Query: 112 LDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQ-----------AGIVY 160
+++ Y+ V ++ V +G CTI YG TG+GK+ TM G AGI+
Sbjct: 72 IEV-YRSVVCPILDEVIMGYNCTIFAYGQTGTGKTFTMEGERSSDEEFTWEQDPLAGIIP 130
Query: 161 RALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGG 220
R L I + SE G V+V++LEIYNEE++DLLS + G
Sbjct: 131 RTLHQIF---EKLSEN-----------GTEFSVKVSLLEIYNEELFDLLSPSPDVG---- 172
Query: 221 FGFGWSKSNASKVKLEVMGKKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSR 280
+ G K IS + ++ +++ RR STL N SSR
Sbjct: 173 -----ERLQMFDDPRNKRGVIIKGLEEISVHNKDEVYHILERGAARRKTASTLMNAYSSR 227
Query: 281 SHCMVILDV----PTVGG-------RLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNI 329
SH + + + TV G +L LVD+AGSENI ++G A+ + INQ +
Sbjct: 228 SHSVFSVTIHMKETTVDGEELVKIGKLNLVDLAGSENIGRSGAVDKRAR-EAGNINQSLL 286
Query: 330 ALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEY 389
L RV+ ++ H+P+R+SKLT +LQDS ++K +I SP + +T+STL+Y
Sbjct: 287 TLGRVITALVERTPHIPYRESKLTRILQDSL-GGRTKTSIIATVSPASINLEETVSTLDY 345
Query: 390 GAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENKLREKERNEAHKKLMK 449
+AK I+ P K + K KE E ++L
Sbjct: 346 ANRAKSIMNKPEVNQK-------------------------LTKKALIKEYTEEIERL-- 378
Query: 450 KEEEIAALRTKVETAPASE--EEINLKVNERTRHLRQELEK--KLEECQRMTNEFVELER 505
+ E+AA R K +SE E++ KV + + + EK +EE + +E +
Sbjct: 379 -KRELAAAREKNGVYLSSENYEQLQGKVLSQEEMITEYTEKITAMEEELKSISELFADNK 437
Query: 506 KRMEE--RILQ-QQEEVEILRKRLEEIELQLCSSSKQERKDENESKEMEPNGFMRKLLSV 562
K +EE ILQ +++E+E + L+E + QL S E K++ +G KLLS
Sbjct: 438 KELEECTTILQCKEKELEETQNHLQESKEQLAQESFVVSAFETTEKKL--HGTANKLLST 495
Query: 563 YKST--DDLGMVKSMD 576
+ T D G+ + +D
Sbjct: 496 VRETTRDVSGLHEKLD 511
>gb|EAA40017.1| GLP_572_50389_48461 [Giardia lamblia ATCC 50803]
Length = 642
Score = 148 bits (374), Expect = 6e-34
Identities = 157/579 (27%), Positives = 256/579 (44%), Gaps = 81/579 (13%)
Query: 99 FTLDGVSVSEEEELDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQAGI 158
FT D V S ++++F + V I+G G T+ YG TGSGK+HTM G G+
Sbjct: 68 FTYDAVYPSNSTQVEVFDES-VREMIDGCLEGYNATVFAYGQTGSGKTHTMMGQKDNPGM 126
Query: 159 VYRALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGG 218
+ A + I D ++ D + V+ + +EIYNE++ DLL+
Sbjct: 127 IPLAFQRIF---DFIAQAKDD----------QFLVRASFVEIYNEDLKDLLT-------- 165
Query: 219 GGFGFGWSKSNASKVKLE---VMGKKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCN 275
A+ ++L+ V G K+ + ++ I K IQK + R V +TL N
Sbjct: 166 ----------GATHLQLKEDPVKGVFIKDLSEHPVSDERHIDKLIQKGNESRAVAATLMN 215
Query: 276 DRSSRSHCM--VILDVPTV--------GGRLMLVDMAGSENIEQAGQTGFEAKMQTAKIN 325
SSRSH + V+L+ TV G+L LVD+AGSE E+ G TG K + AKIN
Sbjct: 216 ATSSRSHSIFQVVLERMTVIDGRECIRVGKLNLVDLAGSERQEKTGATGDRLK-EAAKIN 274
Query: 326 QGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTIS 385
L V+ + G H+P+RDSKLT LLQDS SK LM++ SP +T+S
Sbjct: 275 LSLTTLGCVISKLVEGSKHIPYRDSKLTRLLQDSL-GGNSKTLMVVAVSPASTNYDETMS 333
Query: 386 TLEYGAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKL--QMENKLREKERNEA 443
TL Y +AK I P +D ++I M ++ KL Q+ +++
Sbjct: 334 TLRYADRAKQIKNKPRINEDPKD--------AQIREMRNYVTKLEAQLAEIMQQANAGSG 385
Query: 444 HKKLMKKEEEIAALRTKVETAPASEEEINLKVNERTRHLRQELEKKLEECQRMTNEFVEL 503
+ K+ + T ++E N+ Q L K L++ ++ ++VE
Sbjct: 386 SEVEDKEAYDGEGNMGAGFTGYTADEMANV----------QSLRKNLDKTKKKRVKYVE- 434
Query: 504 ERKRMEERILQQQEEVEILRKRLEEIELQLCSSSKQERKDENESKEMEPNGFMRKLLSVY 563
+RK EE + EE+ L + +++E Q+ S ++ ++ + +K+ + L+
Sbjct: 435 QRKENEEAV--SAEELATLEEEQKKLEEQIKESERKAKERQLMAKK------IAALIEAN 486
Query: 564 KSTDDLGMVKSMDLDMDDQEPFLAREVIVGM----QGISPNQPCPNTLKNGVQEDAYVCA 619
KS V + + D AR +V + + + ++E
Sbjct: 487 KSKMVDKKVLENEERLKDAAIREARNALVAQKKEAERLKKELIEAEQQRKQLEEQCTTAL 546
Query: 620 PNFGQKACLSTVYEEEGEEEAEQDHDKVEEDEEVEKEVI 658
Q Y+E+ E E+ H+ VE D+ E+E+I
Sbjct: 547 DQAQQLELRLNEYKEQLAERREELHN-VEADQAKEREII 584
>gb|AAK91129.1| KRP120-2 [Daucus carota]
Length = 1045
Score = 148 bits (373), Expect = 7e-34
Identities = 155/545 (28%), Positives = 249/545 (45%), Gaps = 86/545 (15%)
Query: 59 IEVIARIRDYPDRKDKPLS--VLQASSNSRSIRVKADFGY----RDFTLDGVSVSEEEEL 112
++VI R R + + K + V+ + N R + + R F D V ++
Sbjct: 53 VQVIVRCRPLSEDEIKAHTPVVITCTENRREVCAVQNIASKQIDRSFMFDKVFGPASQQK 112
Query: 113 DLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQ--------AGIVYRALR 164
DL Y++ V + V G CTI YG TG+GK++TM G ++ AG++ RA++
Sbjct: 113 DL-YEQAVSPIVYEVLEGYNCTIFAYGQTGTGKTYTMEGGGRKKNGEFPSDAGVIPRAVK 171
Query: 165 DILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGGFGFG 224
I E + + S ++VT LE+YNEEI DLL+ F
Sbjct: 172 QIFNI----LESQNAEYS----------MKVTFLELYNEEITDLLAPEE---------FS 208
Query: 225 WSKSNASKVKLEVMGKKAKNATYISGNE------AGKISKEIQKVEKRRIVKSTLCNDRS 278
+ SK + +M + K ++ G E A +I K ++K +R TL N +S
Sbjct: 209 KFIEDKSKKPIALM-EDGKGGVFVRGLEEEIVCTANEIYKILEKGSAKRRTAETLLNKQS 267
Query: 279 SRSHCMVILDVPTVG-----------GRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQG 327
SRSH + + + G+L LVD+AGSENI ++G A+ + +IN+
Sbjct: 268 SRSHSIFSITIHIKECTPEGEEMIKCGKLNLVDLAGSENISRSGAREGRAR-EAGEINKS 326
Query: 328 NIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTL 387
+ L RV+ ++ HVP+RDSKLT LL+DS K+K +I SP + +T+STL
Sbjct: 327 LLTLGRVINALVEHSGHVPYRDSKLTRLLRDSL-GGKTKTCIIATISPSVYSLEETLSTL 385
Query: 388 EYGAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENKLR-EKER---NEA 443
+Y +AK I P K S+ L S I + + + + +N + K+R +EA
Sbjct: 386 DYAHRAKNIKNKPEINQKMMKSAMIKDLYSEIDRLKQEVFSAREKNGIYIPKDRYLQDEA 445
Query: 444 HKKLMKKEEEIAALRTKVETAPASEEEINLKVNERTRHLRQELEKKLEECQRMTNE---- 499
KK M E+I + E+ E+ N + + L EL KLE+ ++ +E
Sbjct: 446 DKKAM--AEKIERMELDFESRDKQFMELQGLHNSQLQ-LTAELSDKLEKTEKKLHETEHA 502
Query: 500 FVELERKRMEERILQQQEEVEI----------------LRKRLEEIELQLCS-SSKQERK 542
V+LE + + +++E I LR LE L + + +K ERK
Sbjct: 503 LVDLEERHRQANATIKEKEYLISNLIKSERSLIERAFELRAELESAALDVSNLFTKIERK 562
Query: 543 DENES 547
D+ E+
Sbjct: 563 DKIEN 567
>ref|XP_623508.1| PREDICTED: similar to kinesin-like boursin [Apis mellifera]
Length = 732
Score = 148 bits (373), Expect = 7e-34
Identities = 158/587 (26%), Positives = 265/587 (44%), Gaps = 107/587 (18%)
Query: 49 NNKDSPPEHPIEVIARIR--DYPDRKDKPLSVLQASSNSRSIRVKA--DFGYRDFTLDGV 104
N K +H I+V R+R + ++ K ++V+ SN I + D + FT D V
Sbjct: 6 NMKKEKKQH-IQVFVRVRPTNNVEKIGKSITVVDVQSNKEVIIRERPHDKFTKKFTFDKV 64
Query: 105 SVSEEEELDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQ--------- 155
+ +++ + Y V + V G CT+ YG TG+GK+ TM G
Sbjct: 65 FGTNAKQIQV-YNAVVSPLLEEVLAGYNCTVFAYGQTGTGKTFTMEGTDNDPSLHWQTDS 123
Query: 156 -AGIVYRALRDILGDGDTDSEGSDGDSSKGFCLGVRTF-VQVTVLEIYNEEIYDLLSTNG 213
AGI+ R+L + + L V+ + V+ + LE+YNEEI+DLLS
Sbjct: 124 TAGIIPRSLSHLFDELRV--------------LEVQEYSVRASFLELYNEEIFDLLS--- 166
Query: 214 GGGGGGGFGFGWSKSNASKVKLEVMGKKAKNATYISGNEAGKISKE------IQKVEKRR 267
S +A+K+++ K K A + G E I + +QK ++R
Sbjct: 167 ------------SSDDAAKIRIYEDPTK-KGAVIVHGLEEMTIHNKNEVFNILQKGSEKR 213
Query: 268 IVKSTLCNDRSSRSHCMVILDVP----TVGG-------RLMLVDMAGSENIEQAGQTGFE 316
+TL N SSRSH + + V T+ G +L LVD+AGSEN+ ++G
Sbjct: 214 QTAATLMNAHSSRSHTIFSITVHIKENTIDGEELLKTGKLNLVDLAGSENVGRSGAVDRR 273
Query: 317 AKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPD 376
A+ + INQ + L RV+ ++A HVP+R+SKLT LLQ+S +++ +I SP
Sbjct: 274 AR-EAGNINQSLLTLGRVITALAEKTPHVPYRESKLTRLLQESL-GGRTRTSIIATISPA 331
Query: 377 PKEIHKTISTLEYGAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENKLR 436
+ +T+STL+Y +A+ I P K F K ++ +
Sbjct: 332 SINLEETLSTLDYAHRARNITNRPEINQK-------------------FSKKALLQEYIE 372
Query: 437 EKERNEAHKKLMKKEEEIAALRTKVETAPASEEEINLKVNERTRHLRQELE--KKLEECQ 494
E ER +KK+ R + P S E+ + +++ + ++L K LEEC
Sbjct: 373 EIER-------LKKDLIACRERNGIYLTPDSYNEMQSLIEFQSKEIEEKLNHIKALEECM 425
Query: 495 RMTNE-FVELERKRMEERILQQQEEVEILRKRLEEIELQLCSSSKQ----ERKDENESKE 549
+ F+EL+ K E Q E+ ++ +LE L S+S + ER+ E +
Sbjct: 426 NSKEQIFIELKSKNSE-----QAYELLNVKNKLESTVNTLMSTSSRLAILEREKEEQKHL 480
Query: 550 MEPNGFMRK-LLSVYKSTDDLGMVKSMDLDMDDQEPFLAREVIVGMQ 595
+E + + LLS ++ D+ + + D++ + F R++ +G Q
Sbjct: 481 VEKHAYTENILLSQVQTVLDVANIVTSDVNKLHDKIF--RKMQIGQQ 525
>gb|AAV44208.1| putative kinesin [Oryza sativa (japonica cultivar-group)]
Length = 1056
Score = 147 bits (372), Expect = 1e-33
Identities = 155/542 (28%), Positives = 243/542 (44%), Gaps = 72/542 (13%)
Query: 59 IEVIARIRDYPDRKDKPLSVLQASSNSRSIRVKADFGY------RDFTLDGVSVSEEEEL 112
++VI R R D + K + + S N R V A R F D V ++
Sbjct: 50 VQVILRCRPMSDEETKSNTPVVISCNERRREVAATQIIANKQIDRTFAFDKVFGPASKQK 109
Query: 113 DLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQ----------AGIVYRA 162
DLF ++ + +N V G CTI YG TG+GK++TM G + AG++ RA
Sbjct: 110 DLF-EQSISPIVNEVLEGYNCTIFAYGQTGTGKTYTMEGGGTRKTKNGELPTDAGVIPRA 168
Query: 163 LRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGGFG 222
+R I D + C ++VT LE+YNEEI DLL+
Sbjct: 169 VRQIF------------DILEAQC--AEYSMKVTFLELYNEEITDLLAPEEPK------- 207
Query: 223 FGWSKSNASKVKLEVMGKKAKNATYISGNE------AGKISKEIQKVEKRRIVKSTLCND 276
F + +K + +M + K ++ G E AG+I K + K +R TL N
Sbjct: 208 FPIVPEDKTKKPIALM-EDGKGGVFVRGLEEEVVYSAGEIYKILDKGSAKRRTAETLLNK 266
Query: 277 RSSRSHCMVILDVPTVG-----------GRLMLVDMAGSENIEQAGQTGFEAKMQTAKIN 325
+SSRSH + + + G+L LVD+AGSENI ++G A+ + +IN
Sbjct: 267 QSSRSHSIFSITIHIKELTHEGEEMIKIGKLNLVDLAGSENISRSGARDGRAR-EAGEIN 325
Query: 326 QGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTIS 385
+ + L RV+ ++ HVP+RDSKLT LL+DS K+K +I SP + +T+S
Sbjct: 326 KSLLTLGRVINALVEHSGHVPYRDSKLTRLLRDSL-GGKTKTCIIATISPSVYCLEETLS 384
Query: 386 TLEYGAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENKL---REKE-RN 441
TL+Y +AK I P + S+ L S I + + + + +N + RE+ +
Sbjct: 385 TLDYAHRAKNIKNKPEVNQRMMKSAVIKDLYSEIDRLKQEVFAAREKNGIYIPRERYLQE 444
Query: 442 EAHKKLMKKEEEIAALRTKVETAPASEEEINLKVNERTRHLRQELEKKLEECQR----MT 497
EA KK M E+I L +E E+ ++ + + L EL +KL + Q+
Sbjct: 445 EAEKKAM--TEKIERLGADLEARDKQLVELK-ELYDAEQLLSAELSEKLGKTQKDLEDTK 501
Query: 498 NEFVELERKRMEERILQQQEEVEILRKRLEEIELQLCSSSKQERKDENESKEMEPNGFMR 557
N +LE K E +++E I E L C+ + E E+ + +G
Sbjct: 502 NVLHDLEEKYNEAESTIKEKEYVIFNLLKSEKSLVDCA---YNLRAELENAAADVSGLFS 558
Query: 558 KL 559
K+
Sbjct: 559 KI 560
>ref|XP_308374.2| ENSANGP00000019061 [Anopheles gambiae str. PEST]
gi|55245420|gb|EAA04655.2| ENSANGP00000019061 [Anopheles
gambiae str. PEST]
Length = 550
Score = 146 bits (369), Expect = 2e-33
Identities = 142/514 (27%), Positives = 236/514 (45%), Gaps = 85/514 (16%)
Query: 49 NNKDSPPEHP-IEVIARIRDYPDRKD--KPLSVLQASSNSRSIRVKADFG----YRDFTL 101
NN + P + ++V R+R R+ + V++ SN R +++K+++ + FT
Sbjct: 9 NNANKPKSNQNVQVYVRVRPTNAREKLIRSQEVVEVVSN-RELQLKSNYTDSRTSKKFTF 67
Query: 102 DGVSVSEEEELDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQ------ 155
D ++ ++ Y+ V I V G CT+ YG TG+GK+HTM G +Q
Sbjct: 68 DRTFAPNSKQHEV-YQAVVAPYIEEVLSGFNCTVFAYGQTGTGKTHTMVGEEEQNLSAAW 126
Query: 156 -----AGIVYRALRDILGD-GDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLL 209
GI+ RA+ + + T+ E S ++++ LE+YNEE+ DLL
Sbjct: 127 EDDTQTGIIPRAVNHLFDELRMTELEFS---------------MRISYLELYNEELCDLL 171
Query: 210 STNGGGGGGGGFGFGWSKSNASKVKLEVMGK-KAKNATYISGNEA------GKISKEIQK 262
ST+ +K+ + + K + + G E + K + K
Sbjct: 172 STD------------------DTIKIRIFDDVQKKGSVIVQGLEEIPVHSKDDVYKLLAK 213
Query: 263 VEKRRIVKSTLCNDRSSRSHCM--VILDVPTVG---------GRLMLVDMAGSENIEQAG 311
++RR STL N +SSRSH + +I+ + G G+L LVD+AGSENI +AG
Sbjct: 214 GQERRKTASTLMNAQSSRSHTIFSIIVHIKENGIDGEEMLKIGKLNLVDLAGSENISKAG 273
Query: 312 QTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMIL 371
+T INQ + L RV+ ++ H+P+R+SKLT LLQ+S ++K +I
Sbjct: 274 NEKGIRTRETVNINQSLLTLGRVITALVEKTPHIPYRESKLTRLLQESL-GGRTKTSIIA 332
Query: 372 CASPDPKEIHKTISTLEYGAKAKCIVRGPH--------TPVKEEDSSSTVILGSRIAAMD 423
SP K+ +T+STLEY +AK I P T +KE + +AA D
Sbjct: 333 TVSPGNKDFEETLSTLEYAHRAKNIQNKPEANQKLSKKTVIKEYTEEIDRLKRDLMAARD 392
Query: 424 EFIMKLQMENKLREKERNE-AHKKLMKKEEEIAALRTKVETAPASEEEINLKVNERTRHL 482
+ + L E ++E A K+L K I AL+ + A E+ + ER L
Sbjct: 393 KNGIYLAEETYNEMVYKSEAATKELNDKLALIKALKEDLARKEAIFNEVACSLAEREEQL 452
Query: 483 RQ---ELEKKLEECQRMTNEFVELERKRMEERIL 513
R+ +L + E + +R+ +E++++
Sbjct: 453 RRTADDLGQTRAELSNTKKNLSKTKRRYVEKKVI 486
>emb|CAA59449.1| kinesin-related protein [Homo sapiens]
gi|1706622|sp|P52732|KIF11_HUMAN Kinesin-like protein
KIF11 (Kinesin-related motor protein Eg5) (Kinesin-like
spindle protein HKSP) (Thyroid receptor interacting
protein 5) (TRIP5) (Kinesin-like protein 1)
Length = 1057
Score = 146 bits (368), Expect = 3e-33
Identities = 148/506 (29%), Positives = 239/506 (46%), Gaps = 63/506 (12%)
Query: 59 IEVIARIRDY--PDRKDKPLSVLQASSNSRSIRVK----ADFGYRD-FTLDGVSVSEEEE 111
I+V+ R R + +RK S+++ + + V+ AD R +T D V + ++
Sbjct: 19 IQVVVRCRPFNLAERKASAHSIVECDPVRKEVSVRTGGLADKSSRKTYTFDMVFGASTKQ 78
Query: 112 LDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQ-----------AGIVY 160
+D+ Y+ V ++ V +G CTI YG TG+GK+ TM G AGI+
Sbjct: 79 IDV-YRSVVCPILDEVIMGYNCTIFAYGQTGTGKTFTMEGERSPNEEYTWEEDPLAGIIP 137
Query: 161 RALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGG 220
R L I + TD+ G V+V++LEIYNEE++DLL+ +
Sbjct: 138 RTLHQIF-EKLTDN-------------GTEFSVKVSLLEIYNEELFDLLNPSSD------ 177
Query: 221 FGFGWSKSNASKVKLEVMGKKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSR 280
+ + K V+ K + T + +E +I ++K +R +TL N SSR
Sbjct: 178 VSERLQMFDDPRNKRGVIIKGLEEITVHNKDEVYQI---LEKGAAKRTTAATLMNAYSSR 234
Query: 281 SHCMVILDV----PTVGG-------RLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNI 329
SH + + + T+ G +L LVD+AGSENI ++G A+ + INQ +
Sbjct: 235 SHSVFSVTIHMKETTIDGEELVKIGKLNLVDLAGSENIGRSGAVDKRAR-EAGNINQSLL 293
Query: 330 ALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEY 389
L RV+ ++ HVP+R+SKLT +LQDS +++ +I SP + +T+STLEY
Sbjct: 294 TLGRVITALVERTPHVPYRESKLTRILQDSL-GGRTRTSIIATISPASLNLEETLSTLEY 352
Query: 390 GAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENKLREKERN--EAHKKL 447
+AK I+ P K + I + + + +N + E N KL
Sbjct: 353 AHRAKNILNKPEVNQKLTKKALIKEYTEEIERLKRDLAAAREKNGVYISEENFRVMSGKL 412
Query: 448 MKKEEEIAALRTKVETAPASEEEINLKVNERTRHLRQELEKKLEECQRMTNEFVELERKR 507
+EE+I L +E A EEE+N +V E + EL++ + Q T E +E +K
Sbjct: 413 TVQEEQIVEL---IEKIGAVEEELN-RVTELFMDNKNELDQCKSDLQNKTQE-LETTQKH 467
Query: 508 MEERILQQQEEVEILRKRLEEIELQL 533
++E LQ +E E + LE E +L
Sbjct: 468 LQETKLQLVKE-EYITSALESTEEKL 492
>emb|CAH72288.1| kinesin family member 11 [Homo sapiens] gi|55959216|emb|CAI13671.1|
kinesin family member 11 [Homo sapiens]
gi|13699824|ref|NP_004514.2| kinesin family member 11
[Homo sapiens] gi|1171153|gb|AAA86132.1| kinesin-like
spindle protein HKSP
Length = 1056
Score = 146 bits (368), Expect = 3e-33
Identities = 148/506 (29%), Positives = 239/506 (46%), Gaps = 63/506 (12%)
Query: 59 IEVIARIRDY--PDRKDKPLSVLQASSNSRSIRVK----ADFGYRD-FTLDGVSVSEEEE 111
I+V+ R R + +RK S+++ + + V+ AD R +T D V + ++
Sbjct: 19 IQVVVRCRPFNLAERKASAHSIVECDPVRKEVSVRTGGLADKSSRKTYTFDMVFGASTKQ 78
Query: 112 LDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQ-----------AGIVY 160
+D+ Y+ V ++ V +G CTI YG TG+GK+ TM G AGI+
Sbjct: 79 IDV-YRSVVCPILDEVIMGYNCTIFAYGQTGTGKTFTMEGERSPNEEYTWEEDPLAGIIP 137
Query: 161 RALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGG 220
R L I + TD+ G V+V++LEIYNEE++DLL+ +
Sbjct: 138 RTLHQIF-EKLTDN-------------GTEFSVKVSLLEIYNEELFDLLNPSSD------ 177
Query: 221 FGFGWSKSNASKVKLEVMGKKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSR 280
+ + K V+ K + T + +E +I ++K +R +TL N SSR
Sbjct: 178 VSERLQMFDDPRNKRGVIIKGLEEITVHNKDEVYQI---LEKGAAKRTTAATLMNAYSSR 234
Query: 281 SHCMVILDV----PTVGG-------RLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNI 329
SH + + + T+ G +L LVD+AGSENI ++G A+ + INQ +
Sbjct: 235 SHSVFSVTIHMKETTIDGEELVKIGKLNLVDLAGSENIGRSGAVDKRAR-EAGNINQSLL 293
Query: 330 ALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEY 389
L RV+ ++ HVP+R+SKLT +LQDS +++ +I SP + +T+STLEY
Sbjct: 294 TLGRVITALVERTPHVPYRESKLTRILQDSL-GGRTRTSIIATISPASLNLEETLSTLEY 352
Query: 390 GAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENKLREKERN--EAHKKL 447
+AK I+ P K + I + + + +N + E N KL
Sbjct: 353 AHRAKNILNKPEVNQKLTKKALIKEYTEEIERLKRDLAAAREKNGVYISEENFRVMSGKL 412
Query: 448 MKKEEEIAALRTKVETAPASEEEINLKVNERTRHLRQELEKKLEECQRMTNEFVELERKR 507
+EE+I L +E A EEE+N +V E + EL++ + Q T E +E +K
Sbjct: 413 TVQEEQIVEL---IEKIGAVEEELN-RVTELFMDNKNELDQCKSDLQNKTQE-LETTQKH 467
Query: 508 MEERILQQQEEVEILRKRLEEIELQL 533
++E LQ +E E + LE E +L
Sbjct: 468 LQETKLQLVKE-EYITSALESTEEKL 492
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.312 0.131 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,316,052,143
Number of Sequences: 2540612
Number of extensions: 62276561
Number of successful extensions: 560538
Number of sequences better than 10.0: 10861
Number of HSP's better than 10.0 without gapping: 1753
Number of HSP's successfully gapped in prelim test: 9575
Number of HSP's that attempted gapping in prelim test: 462508
Number of HSP's gapped (non-prelim): 54115
length of query: 754
length of database: 863,360,394
effective HSP length: 136
effective length of query: 618
effective length of database: 517,837,162
effective search space: 320023366116
effective search space used: 320023366116
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 79 (35.0 bits)
Medicago: description of AC144481.6


