

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144481.4 + phase: 0
(152 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
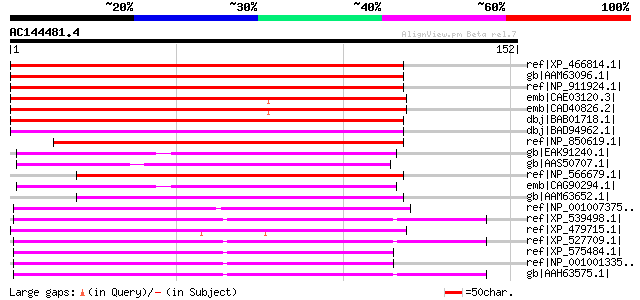
Score E
Sequences producing significant alignments: (bits) Value
ref|XP_466814.1| glycolipid transfer protein-like [Oryza sativa ... 193 6e-49
gb|AAM63096.1| unknown [Arabidopsis thaliana] gi|28973539|gb|AAO... 187 4e-47
ref|NP_911924.1| hypothetical protein [Oryza sativa (japonica cu... 187 6e-47
emb|CAE03120.3| OJ000114_01.1 [Oryza sativa (japonica cultivar-g... 103 8e-22
emb|CAD40826.2| OSJNBa0006B20.21 [Oryza sativa (japonica cultiva... 103 8e-22
dbj|BAB01718.1| unnamed protein product [Arabidopsis thaliana] 96 2e-19
dbj|BAD94962.1| hypothetical protein [Arabidopsis thaliana] gi|4... 90 1e-17
ref|NP_850619.1| glycolipid transfer protein-related [Arabidopsi... 84 7e-16
gb|EAK91240.1| hypothetical protein CaO19.6327 [Candida albicans... 83 1e-15
gb|AAS50707.1| ABL064Wp [Ashbya gossypii ATCC 10895] gi|45185166... 80 1e-14
ref|NP_566679.1| glycolipid transfer protein-related [Arabidopsi... 79 2e-14
emb|CAG90294.1| unnamed protein product [Debaryomyces hansenii C... 79 3e-14
gb|AAM63652.1| unknown [Arabidopsis thaliana] 79 3e-14
ref|NP_001007375.1| hypothetical protein LOC492502 [Danio rerio]... 78 5e-14
ref|XP_539498.1| PREDICTED: similar to phosphoinositol 4-phospha... 77 6e-14
ref|XP_479715.1| hypothetical protein [Oryza sativa (japonica cu... 77 6e-14
ref|XP_527709.1| PREDICTED: similar to phosphoinositol 4-phospha... 77 8e-14
ref|XP_575484.1| PREDICTED: similar to phosphoinositol 4-phospha... 77 1e-13
ref|NP_001001335.1| pleckstrin homology domain containing, famil... 76 1e-13
gb|AAH63575.1| PLEKHA9 protein [Homo sapiens] gi|4050073|gb|AAC9... 72 2e-12
>ref|XP_466814.1| glycolipid transfer protein-like [Oryza sativa (japonica
cultivar-group)] gi|47847777|dbj|BAD21554.1| glycolipid
transfer protein-like [Oryza sativa (japonica
cultivar-group)] gi|47847652|dbj|BAD22518.1| glycolipid
transfer protein-like [Oryza sativa (japonica
cultivar-group)]
Length = 202
Score = 193 bits (491), Expect = 6e-49
Identities = 91/118 (77%), Positives = 106/118 (89%)
Query: 1 MAIVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMD 60
MA+VKSDIGGNI+RLE+KY S+P+K+ LYS+VQ EV+ KTAK SSSCT GLLWLTRAMD
Sbjct: 46 MALVKSDIGGNITRLENKYSSDPSKYEQLYSMVQEEVQNKTAKGSSSCTNGLLWLTRAMD 105
Query: 61 FLVAVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVV 118
FLV +FRNL+EH DW+MSQACTDSY KTLKKWHGWLASS+ TVAM LAP+R+KFMEV+
Sbjct: 106 FLVELFRNLLEHQDWTMSQACTDSYTKTLKKWHGWLASSSFTVAMKLAPNREKFMEVI 163
>gb|AAM63096.1| unknown [Arabidopsis thaliana] gi|28973539|gb|AAO64094.1| unknown
protein [Arabidopsis thaliana]
gi|28393611|gb|AAO42225.1| unknown protein [Arabidopsis
thaliana] gi|2459429|gb|AAB80664.1| expressed protein
[Arabidopsis thaliana] gi|25408274|pir||H84745
hypothetical protein At2g33470 [imported] - Arabidopsis
thaliana gi|42571029|ref|NP_973588.1| glycolipid
transfer protein-related [Arabidopsis thaliana]
gi|18403285|ref|NP_565766.1| glycolipid transfer
protein-related [Arabidopsis thaliana]
Length = 202
Score = 187 bits (475), Expect = 4e-47
Identities = 87/118 (73%), Positives = 100/118 (84%)
Query: 1 MAIVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMD 60
M +VKSDIGGNI+RLE YLS+P KF LY+ VQVE+E+K AK SSSCT GLLWLTRAMD
Sbjct: 46 MTLVKSDIGGNITRLEKNYLSDPDKFKYLYTFVQVEIESKIAKGSSSCTNGLLWLTRAMD 105
Query: 61 FLVAVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVV 118
FLV +FRNL+ H DWSM QAC DSY KTLKKWHGWLASST ++A+ LAPDRKKFM+V+
Sbjct: 106 FLVELFRNLVAHQDWSMPQACADSYQKTLKKWHGWLASSTFSMALKLAPDRKKFMDVI 163
>ref|NP_911924.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|22296432|dbj|BAC10199.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 202
Score = 187 bits (474), Expect = 6e-47
Identities = 84/118 (71%), Positives = 107/118 (90%)
Query: 1 MAIVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMD 60
M+IVK+DIGGNI+RLE+KY S+P+K+ L+S+V+VE+ +KTAK+SSSCT GLLWLTRAMD
Sbjct: 46 MSIVKNDIGGNITRLETKYASDPSKYEQLHSMVKVEISSKTAKSSSSCTNGLLWLTRAMD 105
Query: 61 FLVAVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVV 118
FLVA+F NL++H DW MSQAC+D+Y KTLKKWHGWLASS+ +VA+ LAPDRKKFME++
Sbjct: 106 FLVALFHNLVQHPDWQMSQACSDAYSKTLKKWHGWLASSSFSVAIKLAPDRKKFMEII 163
>emb|CAE03120.3| OJ000114_01.1 [Oryza sativa (japonica cultivar-group)]
Length = 276
Score = 103 bits (257), Expect = 8e-22
Identities = 48/120 (40%), Positives = 77/120 (64%), Gaps = 1/120 (0%)
Query: 1 MAIVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMD 60
MA+++ D+ NI RL+ YL +P+K+ L +++ EV+ TA+ SC +LWLTR+MD
Sbjct: 105 MAVLRLDVQRNIERLQELYLLDPSKYYNLEEILEKEVDEGTARKVDSCARAILWLTRSMD 164
Query: 61 FLVAVFRNLIEHADWS-MSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVVI 119
F +A+ + L E +D +Q +Y TLK WHGW++S+ +AM L PDRK F+ +++
Sbjct: 165 FTIALLQRLEEDSDQKCFAQLVESAYMVTLKPWHGWISSAAYKIAMKLIPDRKMFINLLV 224
>emb|CAD40826.2| OSJNBa0006B20.21 [Oryza sativa (japonica cultivar-group)]
gi|50924476|ref|XP_472598.1| OSJNBa0006B20.21 [Oryza
sativa (japonica cultivar-group)]
Length = 266
Score = 103 bits (257), Expect = 8e-22
Identities = 48/120 (40%), Positives = 77/120 (64%), Gaps = 1/120 (0%)
Query: 1 MAIVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMD 60
MA+++ D+ NI RL+ YL +P+K+ L +++ EV+ TA+ SC +LWLTR+MD
Sbjct: 105 MAVLRLDVQRNIERLQELYLLDPSKYYNLEEILEKEVDEGTARKVDSCARAILWLTRSMD 164
Query: 61 FLVAVFRNLIEHADWS-MSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVVI 119
F +A+ + L E +D +Q +Y TLK WHGW++S+ +AM L PDRK F+ +++
Sbjct: 165 FTIALLQRLEEDSDQKCFAQLVESAYMVTLKPWHGWISSAAYKIAMKLIPDRKMFINLLV 224
>dbj|BAB01718.1| unnamed protein product [Arabidopsis thaliana]
Length = 233
Score = 95.9 bits (237), Expect = 2e-19
Identities = 43/118 (36%), Positives = 73/118 (61%)
Query: 1 MAIVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMD 60
MA+++ DI NI RLE + S+P ++ L +++ E + +++ SC+ LWLTRAMD
Sbjct: 73 MAVLRHDIDQNIQRLEKMWESDPLVYSNLVEILRKEAKEGSSRKPKSCSRAALWLTRAMD 132
Query: 61 FLVAVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVV 118
F +A+ + L++ +M QA + Y T+K WHGW++S+ VA+ L P+ F+ V+
Sbjct: 133 FTLALLQRLVKDMSQNMEQAIEECYNLTIKPWHGWISSAAFKVALKLVPNNNTFINVL 190
>dbj|BAD94962.1| hypothetical protein [Arabidopsis thaliana]
gi|45773920|gb|AAS76764.1| At1g21360 [Arabidopsis
thaliana]
Length = 223
Score = 90.1 bits (222), Expect = 1e-17
Identities = 45/118 (38%), Positives = 65/118 (54%)
Query: 1 MAIVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMD 60
MA+++ DI NI RLE Y ++ ++ L +++ E E T+K +SC L WLTR MD
Sbjct: 63 MAVLRQDIDQNIQRLEKFYETDSCVYSNLAEILKKEKEEGTSKMVASCGRALFWLTRTMD 122
Query: 61 FLVAVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVV 118
F + R L + M + + Y TLK HGW+AS+ V + L PD K FME +
Sbjct: 123 FTAGLLRLLSKEMSSKMEELVEECYMTTLKPHHGWIASAAFKVCLKLVPDNKTFMEAI 180
>ref|NP_850619.1| glycolipid transfer protein-related [Arabidopsis thaliana]
Length = 149
Score = 84.0 bits (206), Expect = 7e-16
Identities = 37/105 (35%), Positives = 64/105 (60%)
Query: 14 RLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMDFLVAVFRNLIEHA 73
RLE + S+P ++ L +++ E + +++ SC+ LWLTRAMDF +A+ + L++
Sbjct: 2 RLEKMWESDPLVYSNLVEILRKEAKEGSSRKPKSCSRAALWLTRAMDFTLALLQRLVKDM 61
Query: 74 DWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVV 118
+M QA + Y T+K WHGW++S+ VA+ L P+ F+ V+
Sbjct: 62 SQNMEQAIEECYNLTIKPWHGWISSAAFKVALKLVPNNNTFINVL 106
>gb|EAK91240.1| hypothetical protein CaO19.6327 [Candida albicans SC5314]
Length = 197
Score = 83.2 bits (204), Expect = 1e-15
Identities = 45/114 (39%), Positives = 62/114 (53%), Gaps = 4/114 (3%)
Query: 3 IVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMDFL 62
+VKSD+ GNI+++ +K L +PA L LV E +TKT A T GLLWL+R + F
Sbjct: 48 VVKSDMTGNITKIRNKLLEDPANSATLQDLVLTEAKTKTKTA----TQGLLWLSRGLQFT 103
Query: 63 VAVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFME 116
R ++ ++ TD+Y KTL K+HG L +AM P RK F E
Sbjct: 104 AQAMRETVDAPGKELTVTFTDAYTKTLSKFHGMLVKPVFKLAMKACPYRKDFFE 157
>gb|AAS50707.1| ABL064Wp [Ashbya gossypii ATCC 10895] gi|45185166|ref|NP_982883.1|
ABL064Wp [Eremothecium gossypii]
Length = 196
Score = 79.7 bits (195), Expect = 1e-14
Identities = 42/112 (37%), Positives = 62/112 (54%), Gaps = 4/112 (3%)
Query: 3 IVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMDFL 62
+V+ D+ GNI++L ++ LS+P + L LV E A+ S + + GLLWLTR + F
Sbjct: 49 VVQKDLTGNITKLRNRQLSHPGESATLQELVIAE----RAQGSKTASEGLLWLTRGLQFT 104
Query: 63 VAVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKF 114
R ++H + +S+ TD+Y KTL K HG L +AM P RK F
Sbjct: 105 AQALRETLDHPELELSKTFTDAYGKTLTKHHGMLVRPVFKLAMKACPYRKDF 156
>ref|NP_566679.1| glycolipid transfer protein-related [Arabidopsis thaliana]
Length = 144
Score = 79.3 bits (194), Expect = 2e-14
Identities = 34/98 (34%), Positives = 60/98 (60%)
Query: 21 SNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMDFLVAVFRNLIEHADWSMSQA 80
S+P ++ L +++ E + +++ SC+ LWLTRAMDF +A+ + L++ +M QA
Sbjct: 4 SDPLVYSNLVEILRKEAKEGSSRKPKSCSRAALWLTRAMDFTLALLQRLVKDMSQNMEQA 63
Query: 81 CTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVV 118
+ Y T+K WHGW++S+ VA+ L P+ F+ V+
Sbjct: 64 IEECYNLTIKPWHGWISSAAFKVALKLVPNNNTFINVL 101
>emb|CAG90294.1| unnamed protein product [Debaryomyces hansenii CBS767]
gi|50426473|ref|XP_461833.1| unnamed protein product
[Debaryomyces hansenii]
Length = 197
Score = 78.6 bits (192), Expect = 3e-14
Identities = 42/114 (36%), Positives = 62/114 (53%), Gaps = 4/114 (3%)
Query: 3 IVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMDFL 62
+V+ D+ GNI+++ +K L++PA L LV E TKT A T GLLWL+R + F
Sbjct: 48 VVQKDMTGNITKIRTKLLADPAGSGTLQDLVLSEANTKTKTA----TQGLLWLSRGLQFT 103
Query: 63 VAVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFME 116
R +++ ++ TD+Y KTL ++HG L +AM P RK F E
Sbjct: 104 SQAMRETVDNPSKELAVTFTDAYSKTLSQYHGMLVKPIFKLAMKACPYRKDFFE 157
>gb|AAM63652.1| unknown [Arabidopsis thaliana]
Length = 144
Score = 78.6 bits (192), Expect = 3e-14
Identities = 34/98 (34%), Positives = 59/98 (59%)
Query: 21 SNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMDFLVAVFRNLIEHADWSMSQA 80
S+P ++ L +++ E + +++ SC+ LWLTRAMDF +A+ + L++ +M QA
Sbjct: 4 SDPLVYSNLVEILRKEAKEGSSRKPKSCSRAALWLTRAMDFTLALLQRLVKDMSQNMEQA 63
Query: 81 CTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVV 118
+ Y T+K WHGW++S+ VA+ L P F+ V+
Sbjct: 64 IEECYNLTIKPWHGWISSAAFKVALKLVPKNNTFINVL 101
>ref|NP_001007375.1| hypothetical protein LOC492502 [Danio rerio]
gi|55250041|gb|AAH85465.1| Zgc:101886 [Danio rerio]
Length = 549
Score = 77.8 bits (190), Expect = 5e-14
Identities = 43/119 (36%), Positives = 67/119 (56%), Gaps = 1/119 (0%)
Query: 2 AIVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMDF 61
A VK D GNI +++ K +S+P F L S+V EV+T+ A+ +S T LLWL R + F
Sbjct: 385 APVKIDFVGNIKKIQQKVVSDPESFPTLQSIVLHEVKTEVAQVRNSATEALLWLKRGLKF 444
Query: 62 LVAVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVVIS 120
L F + I + A ++Y KTL+++HGW+ +A+ AP + FM ++S
Sbjct: 445 L-KEFLSEINTGVKDVQGALYNAYGKTLRQYHGWVVRGVFALALRAAPSYEGFMAALVS 502
>ref|XP_539498.1| PREDICTED: similar to phosphoinositol 4-phosphate adaptor protein-2
[Canis familiaris]
Length = 648
Score = 77.4 bits (189), Expect = 6e-14
Identities = 44/142 (30%), Positives = 76/142 (52%), Gaps = 2/142 (1%)
Query: 2 AIVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMDF 61
A VK D+ GNI ++ KY++N +F L +V EVE A+ +S T LLWL R + F
Sbjct: 440 APVKMDLVGNIKKVNQKYITNKEEFTTLQKIVLHEVEADVAQVRNSATEALLWLKRGLKF 499
Query: 62 LVAVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVVISM 121
L F +++ + + A ++Y KTL++ HGW+ +A+ AP + F+ +++
Sbjct: 500 LKG-FLTEVKNGEKDIQTALNNAYGKTLRQHHGWVVRGVFALALRAAPSYEDFV-AALTI 557
Query: 122 LTLSNFVLAFLLSLKRITSFWL 143
+ AF + ++R S +L
Sbjct: 558 KEGDHQKAAFSVGMQRDLSLYL 579
>ref|XP_479715.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|42408369|dbj|BAD09520.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
gi|42408243|dbj|BAD09400.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 235
Score = 77.4 bits (189), Expect = 6e-14
Identities = 39/129 (30%), Positives = 73/129 (56%), Gaps = 10/129 (7%)
Query: 1 MAIVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLT---- 56
+ +++ DI N+ RL+ +P+K++ L ++V EVE T+K ++SCT +LWL
Sbjct: 65 LLVLRQDIQQNVQRLQDVLARDPSKYSSLTAIVTEEVEEGTSKKANSCTRAILWLASAVL 124
Query: 57 -----RAMDFLVAVFRNLIEHADW-SMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPD 110
R+++F + L+ D S+ + +Y TLK WHGW++S+ VA L P+
Sbjct: 125 RILPIRSINFSKHLLEGLLNTCDQSSLREIVEKAYITTLKPWHGWISSAAYRVAQKLIPE 184
Query: 111 RKKFMEVVI 119
++ F+ +++
Sbjct: 185 KEIFIALLM 193
>ref|XP_527709.1| PREDICTED: similar to phosphoinositol 4-phosphate adaptor protein-2
[Pan troglodytes]
Length = 634
Score = 77.0 bits (188), Expect = 8e-14
Identities = 44/142 (30%), Positives = 76/142 (52%), Gaps = 2/142 (1%)
Query: 2 AIVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMDF 61
A VK D+ GNI ++ KY++N +F L +V EVE A+ +S T LLWL R + F
Sbjct: 369 APVKMDLVGNIKKVNQKYITNKEEFTTLQKIVLHEVEADVAQVRNSATEALLWLKRGLKF 428
Query: 62 LVAVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVVISM 121
L F +++ + + A ++Y KTL++ HGW+ +A+ AP + F+ +++
Sbjct: 429 LKG-FLTEVKNGEKDIQTALNNAYGKTLRQHHGWVVRGVFALALRAAPSYEDFV-AALTV 486
Query: 122 LTLSNFVLAFLLSLKRITSFWL 143
+ AF + ++R S +L
Sbjct: 487 KEGDHQKEAFSIGMQRDLSLYL 508
>ref|XP_575484.1| PREDICTED: similar to phosphoinositol 4-phosphate adaptor protein-2
[Rattus norvegicus]
Length = 520
Score = 76.6 bits (187), Expect = 1e-13
Identities = 39/114 (34%), Positives = 63/114 (55%), Gaps = 1/114 (0%)
Query: 2 AIVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMDF 61
A VK D+ GNI ++ KY++N +F L +V EVE A+ +S T LLWL R + F
Sbjct: 356 APVKMDLVGNIKKVNQKYITNKEEFTTLQKIVLHEVEADVARVRNSATEALLWLKRGLKF 415
Query: 62 LVAVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFM 115
L F +++ + + A ++Y KTL++ HGW+ +A+ AP + F+
Sbjct: 416 LKG-FLTEVKNGEKDIQTALNNAYGKTLRQHHGWVVRGVFALALRAAPSYEDFV 468
>ref|NP_001001335.1| pleckstrin homology domain containing, family A (phosphoinositide
binding specific) member 8 [Mus musculus]
gi|30481662|gb|AAH52360.1| Pleckstrin homology domain
containing, family A (phosphoinositide binding specific)
member 8 [Mus musculus]
Length = 474
Score = 76.3 bits (186), Expect = 1e-13
Identities = 39/114 (34%), Positives = 63/114 (55%), Gaps = 1/114 (0%)
Query: 2 AIVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMDF 61
A VK D+ GNI ++ KY++N +F L +V EVE A+ +S T LLWL R + F
Sbjct: 310 APVKMDLVGNIKKVNQKYITNKEEFTTLQKIVLHEVEADVAQVRNSATEALLWLKRGLKF 369
Query: 62 LVAVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFM 115
L F +++ + + A ++Y KTL++ HGW+ +A+ AP + F+
Sbjct: 370 LKG-FLTEVKNGEKDIQTALNNAYGKTLRQHHGWVVRGVFALALRAAPSYEDFV 422
>gb|AAH63575.1| PLEKHA9 protein [Homo sapiens] gi|4050073|gb|AAC97956.1| putative
glycolipid transfer protein [Homo sapiens]
gi|7705684|ref|NP_056983.1| pleckstrin homology domain
containing, family A (phosphoinositide binding specific)
member 9 [Homo sapiens]
Length = 391
Score = 72.4 bits (176), Expect = 2e-12
Identities = 42/142 (29%), Positives = 74/142 (51%), Gaps = 2/142 (1%)
Query: 2 AIVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMDF 61
A VK D+ NI ++ KY++N +F L +V EVE A+ +S T LLWL R + F
Sbjct: 239 APVKMDLVENIKKVNQKYITNKEEFTTLQKIVLHEVEADVAQVRNSATEALLWLKRGLKF 298
Query: 62 LVAVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVVISM 121
L F +++ + + A ++Y KTL++ HGW+ +A+ P + F+ +++
Sbjct: 299 LKG-FLTEVKNGEKDIQTALNNAYGKTLRQHHGWVVRGVFALALRATPSYEDFV-AALTV 356
Query: 122 LTLSNFVLAFLLSLKRITSFWL 143
+ AF + ++R S +L
Sbjct: 357 KEGDHRKEAFSIGMQRDLSLYL 378
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.327 0.134 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 222,854,450
Number of Sequences: 2540612
Number of extensions: 6826124
Number of successful extensions: 25807
Number of sequences better than 10.0: 187
Number of HSP's better than 10.0 without gapping: 100
Number of HSP's successfully gapped in prelim test: 87
Number of HSP's that attempted gapping in prelim test: 25643
Number of HSP's gapped (non-prelim): 188
length of query: 152
length of database: 863,360,394
effective HSP length: 128
effective length of query: 24
effective length of database: 538,162,058
effective search space: 12915889392
effective search space used: 12915889392
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 67 (30.4 bits)
Medicago: description of AC144481.4


