

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141323.18 - phase: 0
(1638 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
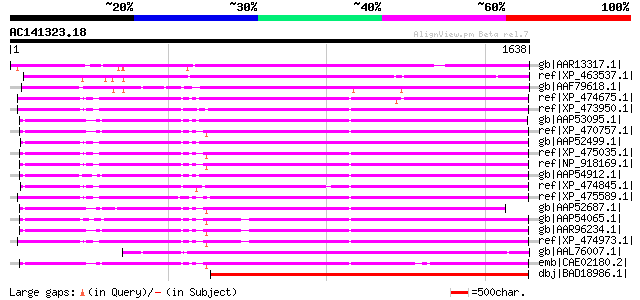
Score E
Sequences producing significant alignments: (bits) Value
gb|AAR13317.1| gag-pol polyprotein [Phaseolus vulgaris] 1356 0.0
ref|XP_463537.1| putative polyprotein [Oryza sativa (japonica cu... 1098 0.0
gb|AAF79618.1| F5M15.26 [Arabidopsis thaliana] gi|25402907|pir||... 1047 0.0
ref|XP_474675.1| OSJNBa0027O01.4 [Oryza sativa (japonica cultiva... 956 0.0
ref|XP_473950.1| OSJNBa0053K19.16 [Oryza sativa (japonica cultiv... 955 0.0
gb|AAP53095.1| putative retroelement [Oryza sativa (japonica cul... 954 0.0
ref|XP_470757.1| putative gag-pol precursor [Oryza sativa] gi|18... 954 0.0
gb|AAP52499.1| putative gag-pol precursor [Oryza sativa (japonic... 952 0.0
ref|XP_475035.1| OSJNBa0042F21.5 [Oryza sativa (japonica cultiva... 952 0.0
ref|NP_918169.1| putative gag-pol polyprotein [Oryza sativa (jap... 952 0.0
gb|AAP54912.1| gag-pol precursor [Oryza sativa (japonica cultiva... 944 0.0
ref|XP_474845.1| OSJNBa0035O13.3 [Oryza sativa (japonica cultiva... 932 0.0
ref|XP_475589.1| putative polyprotein [Oryza sativa (japonica cu... 925 0.0
gb|AAP52687.1| putative gag-pol precursor [Oryza sativa (japonic... 922 0.0
gb|AAP54065.1| putative gypsy-type retrotransposon [Oryza sativa... 919 0.0
gb|AAR96234.1| putative polyprotein [Oryza sativa (japonica cult... 917 0.0
ref|XP_474973.1| OSJNBb0015G09.7 [Oryza sativa (japonica cultiva... 915 0.0
gb|AAL76007.1| prpol [Zea mays] 912 0.0
emb|CAE02180.2| OSJNBa0080E14.11 [Oryza sativa (japonica cultiva... 902 0.0
dbj|BAD18986.1| GAG-POL precursor [Vitis vinifera] 900 0.0
>gb|AAR13317.1| gag-pol polyprotein [Phaseolus vulgaris]
Length = 1859
Score = 1356 bits (3510), Expect = 0.0
Identities = 729/1762 (41%), Positives = 1063/1762 (59%), Gaps = 180/1762 (10%)
Query: 4 QIAQLLEQNEALLASVET-------------------------------IQQVQQQEKTD 32
+IAQL EQN+ LL +E +Q +Q K
Sbjct: 140 EIAQLKEQNKRLLDRLEQSEREGHSRAPSPSPFQSGTRTIAQAIPHTSLVQHTRQSAKPV 199
Query: 33 SHHGDLDEPEPQPFSTEIWNAPVPDNFKPPQLPTFDGRSDPSEHVTVFNTRMSVYGVADS 92
+ + ++ P+ PF+ +I P+PD ++ + +DG +DP EH+ +F T+M++Y +
Sbjct: 200 TPN-EVANPKGHPFTDDIIATPLPDKWRGLTINLYDGSTDPDEHLNIFRTQMTLYTTDRT 258
Query: 93 LKCKLLAGTLADAALRWYMSLPRFSIVGYQDLTKKFTQQFSGTRHRKVLSTSLFNVRQGP 152
+ CK+ +L + L W+ LP SI + L KFT Q++ +R + S SL NV+Q
Sbjct: 259 VWCKVFPTSLREGPLGWFSDLPPNSIASFDALELKFTTQYATSRPHRTSSMSLLNVKQER 318
Query: 153 NESLREYLARFNDSTIKVSNPNQEVFVGAFQNGLRAGQFNESLAQKPADSMEEIIARAEC 212
ESLR ++ RF+ + + N N E+ + + + G+F ESL ++P +M+E+ RA
Sbjct: 319 GESLRTFMNRFSKVCMNIRNLNPEIAMHHLVSAILPGRFTESLIKRPPCNMDELRTRATK 378
Query: 213 YVKGEESNAEKRARDVKEKGSSGGERRNHYVPPNRDRGTFKKPYERNQPRYVPE--HFTP 270
+++ EE R + +S G R PP T ER++P P ++TP
Sbjct: 379 FMQIEEHIDYHRKTYAENTDNSKGIR-----PP-----TIPTDRERHRPNRGPRFHNYTP 428
Query: 271 LNTRPEKILKEVFESKIIPPPPFNRSKTM-GQDKNAWCKYHLIEGHNTDDCVHLKREIEK 329
L K+L E + ++IP S+T D + C+YH GH T+ C LK +IE+
Sbjct: 429 LIVPRGKVLDEALQIELIPT--LRPSQTPPNADTSKRCQYHRNYGHTTEGCQALKDKIEE 486
Query: 330 LLLNGKLRGYAKE--------------KHHSERREDKP---------------------- 353
L+ G LR + K + S RR+D+
Sbjct: 487 LVQAGHLRKFVKTTITAPRSPQRDHDPRERSGRRDDRTRDNHYRSSRRKRSESPIRRTRP 546
Query: 354 ---NPE------------------------PKHTLHTISGGFAGGGESSNLRKKYVRQVM 386
+PE P ++ I+GGFAGGG S++ RKK++R +
Sbjct: 547 KSESPERRSRTKQKVRTVINTIAGPVSLGQPPQEINYIAGGFAGGGCSNSARKKHLRAIQ 606
Query: 387 LLGDSPSSPKKY-PHVTFSSEDFGRVLPHDDDPLVISVQLFNWEIKRVLIDTGSSADVLY 445
+ +P+ + + P +TF+ EDF + P DDP+VI+V++ + I +VL+D GSS D+LY
Sbjct: 607 SVHSTPTQRRPHIPPITFTDEDFTAIDPSQDDPMVITVEIDKFAIAKVLVDQGSSVDILY 666
Query: 446 FDAFSKMGLSEEQLQPFNGTLSGFTGEKVHVRGYVTLKTIFGTGDQQTSIKVRYLVINSP 505
++ F KM + E ++QP+N + GF+ E+V +G++ L T FG +I +RYL++N+
Sbjct: 667 WETFKKMKIPEAEIQPYNEQIVGFSRERVDTKGFIDLYTTFGDDYLSKTINIRYLLVNAN 726
Query: 506 SSYNVIIGRPSINLLDAFVSTKHLLMKYPLDSGRVGVVHGDQKIARECYHASLRL----- 560
+SYN+++GRPSIN L A VST HL MK+P +G + VH DQK ARECY ASL++
Sbjct: 727 TSYNILLGRPSINRLKAIVSTPHLAMKFPSVNGDIATVHIDQKTARECYVASLKVEPTRR 786
Query: 561 -----------------------KKHKEFEVQVIDLDPREDFPQERIEPIEEVKDVVIGP 597
++ + V ++DLDPR D P R+E E+++ + +
Sbjct: 787 LYTTSAERTTERRGRSTERRSRGRESRRHLVALVDLDPRLDDP--RMEAGEDLQPIFLRD 844
Query: 598 LPHQITKIGTSLSGLEEEILVQLLRENVDLFAWAPSDMPGIDIGVACHHLAVRTSVKPVV 657
+ T +GTSL + E + + L +N DLFAW +DMPG+ V H L+V T +P+
Sbjct: 845 KDRK-TYMGTSLKPDDRETIGKTLTKNADLFAWTAADMPGVKSDVITHRLSVYTEARPIA 903
Query: 658 QRKRKMGEEKRKAVDEEVKKLQEAHFICEIKYPTWLANTVLVKKASGKWRMCVDYTDLNM 717
Q+KRK+GEE+RKA EE KL +A FI + Y TWLAN V+VKK +GKWRMCVDYTDLN
Sbjct: 904 QKKRKLGEERRKAAREETDKLIQAGFIQKAHYTTWLANVVMVKKTNGKWRMCVDYTDLNK 963
Query: 718 ACPKDPYPLPSIDHLIDNASGYKTLSFMDAYSGYNQIKMDPLDAPKTAFMTNQKNYHYRV 777
ACPKD YPLP+ID L+D A+G++ LSF+DAYSGYNQI+M D KTAF T+ N+ Y V
Sbjct: 964 ACPKDSYPLPTIDRLVDGAAGHQILSFLDAYSGYNQIQMYHRDREKTAFRTDSDNFFYEV 1023
Query: 778 MSFGLRNAGATFQRSMDTIFSNQIGRNLEVYIDDLVVKTSEKQSHSVDLKEIFQQIRKFS 837
M FGL+NAGAT+QR MD +F + IGRN+EVY+DD+VVK+ + H DLKE+FQ +R++
Sbjct: 1024 MPFGLKNAGATYQRLMDHVFHDMIGRNVEVYVDDIVVKSDSCEQHVSDLKEVFQALRQYR 1083
Query: 838 MRLNPTKCTFGVQAGKFLGFLLTKKGIEANPDKCQAIINMRNPCNIREVQQLTGRLAALS 897
MRLNP KC FGV+ GKFLGF+LT +GIEANP+KC+AI MR+P ++E+Q+L RL +LS
Sbjct: 1084 MRLNPEKCAFGVEGGKFLGFMLTHRGIEANPEKCKAITEMRSPKGLKEIQRLVSRLTSLS 1143
Query: 898 RFLSCAGDKAFAFFATIKKKEEFEWNQECDEAFGKIKQFLTTPPILHRPTKGAGLFLYLS 957
RF+ ++ +KK +FEW EC++ F ++K FL +PP++ +P + +YL+
Sbjct: 1144 RFVPKLAERTRPIIKLLKKTSKFEWTDECEQNFQQLKAFLASPPVIQKPNAREPIVVYLA 1203
Query: 958 VSENALSSVLVEESDEREKPIYFVSRVLKGAELRYQKIEKLALAVIITARKLRPYFQSHK 1017
VS A+SS LV+E E+P+YFVSRVL AE RYQ +EK+A A++ITAR++R YFQ+HK
Sbjct: 1204 VSNEAVSSALVQEIKAEERPVYFVSRVLHDAETRYQMVEKVAFALVITARRMRMYFQNHK 1263
Query: 1018 VVIRTNYPVKQILGKLDLAGRMLSWSVELSEYDIQFAPRNNIKSQVLADFVVEFTSPIQE 1077
V++RTNYP+ +IL K DLAGRM+ W+VELSE+ I++ PR IKSQ LADF E T + E
Sbjct: 1264 VIVRTNYPIMKILTKPDLAGRMIGWAVELSEFHIEYQPRGAIKSQALADFTAELTPYLTE 1323
Query: 1078 AVPHVWLLSVDGSSNIKGSGAGIILEGPGDLLIEQSLKFDFKASNNQAEYEALIAGMLLA 1137
P W L VDGSSN + SGAG++LEGPG++++EQ++KF+FK SNNQAEYEA+IAG+ LA
Sbjct: 1324 RTPR-WTLYVDGSSNSRSSGAGVVLEGPGEIVVEQAMKFEFKTSNNQAEYEAIIAGLHLA 1382
Query: 1138 QEMGAKNLRARSDSQLMTNQISGEYQTKDQQLSKYLARVRKLAGDFQFFEAIYVPRESNS 1197
E+ N+ +SDS+L+ Q++GEY+ ++ L +Y V+ L F+ +V RE+N+
Sbjct: 1383 IELEVTNITCKSDSRLVVGQLTGEYEVRETLLQQYFHFVKNLLNRFKEISFQHVRRENNT 1442
Query: 1198 RADLLAKLASTKKPGNNRTVIQEVISAPSTDEKAVFELNQEPEGWMTPLLKFLTGSFVAK 1257
RAD L++LA+ KK G +R+ I ++ PS + + +P WMTP+ ++LT
Sbjct: 1443 RADALSRLATLKKKGAHRSAIHVTLAKPSVGTEECMATDTQP-NWMTPIKQYLTDGVCDP 1501
Query: 1258 NDEYAQLVRRRATKFVVIAGKLYKRGRASPLLRCLGEGETELVLLEVHEGVCGSHIGGRS 1317
+ E + ++ +A ++++I LY+RG + PLL+CLG + V+ E+HEG+CG+H G R+
Sbjct: 1502 HLE--KTMKLQAARYILIGEDLYRRGYSRPLLKCLGPEQVTYVMTELHEGICGTHSGART 1559
Query: 1318 LAAKLLRAGYYWPRMAHDCCEFVKKCDKCQRFSDKKNAPANELTSVFSPWPFHKWGVDIV 1377
++AK+LRAGYYWP + DC E+ G+DI+
Sbjct: 1560 MSAKILRAGYYWPTLQGDCTEY---------------------------------GMDII 1586
Query: 1378 GPFPQAPGQLKFLIVDVDYFTKWVEAEAVSKITAERVVKFYWKKIICHFGLPKYIVTDNG 1437
GPF GQ KFL+V +DYFTKW+EAE ++ ITA V F WK I+C FGLP+ I+TDNG
Sbjct: 1587 GPFTPGKGQCKFLLVGIDYFTKWIEAEPLTAITARNVQSFVWKNIVCRFGLPQIIITDNG 1646
Query: 1438 TQFASSKVVNFCKQLGIETKFVSVIHPQANGQAESANKMIVNDIKKKLEDAKGLWAEQLH 1497
QF + F ++L I+ SV HPQ NGQAE+ANK+I+N++KK+L +KG W E+L
Sbjct: 1647 RQFTDRGLAEFYEKLHIKHITSSVEHPQTNGQAEAANKVILNELKKRLGPSKGNWTEELL 1706
Query: 1498 EVLWSYHTTPHSTTGETPFTMVYGADAMLPVEIDTPTWRREHFSEESNEVGIRCTMDMID 1557
EVLW+Y TP STT ETP+++ YG +AM+PVEI P+ RR+ + N+ + +D+I+
Sbjct: 1707 EVLWAYRCTPQSTTQETPYSLTYGTEAMIPVEIGEPSLRRQTLDLDLNKESLLVGLDLIN 1766
Query: 1558 EVREAAHIREFAAKQRAARRYNSKVIPRSMKEGDLVLK-QVVAPTRIGKLLPSWKGPYRV 1616
E+R+ IRE A K RAARRYNSKV PRS ++GDLV + + A GK +W+GP+R+
Sbjct: 1767 ELRDKCKIREEACKIRAARRYNSKVKPRSYQKGDLVWRMRSDARKDGGKFSSNWEGPFRI 1826
Query: 1617 KEKLQHGAYKLEELSGNPVPRT 1638
GAY LE LSG PRT
Sbjct: 1827 SNTATGGAYYLEYLSGKSAPRT 1848
>ref|XP_463537.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 2001
Score = 1098 bits (2841), Expect = 0.0
Identities = 618/1661 (37%), Positives = 951/1661 (57%), Gaps = 79/1661 (4%)
Query: 45 PFSTEIWNAPVPDNFKPPQLPTFDGRSDPSEHVTVFNTRMSVYGVADSLKCKLLAGTLAD 104
P S + AP P NFK + + G +DP+E++ V+ T + G D+ K K+L L
Sbjct: 342 PLSQNLQMAPWPINFKLSNITKYKGDTDPNEYLRVYETAVEAAGGDDTTKAKILPTMLEG 401
Query: 105 AALRWYMSLPRFSIVGYQDLTKKFTQQFSGTRHRKVLSTSLFNVRQGPNESLREYLARFN 164
AL WY ++P +I ++ + F F G + L+ ++Q P E+LR ++ +F+
Sbjct: 402 VALSWYTTIPPMTIYSWEHMRDTFRAGFIGAYEEPKEADDLYAMKQLPGETLRSFIVKFS 461
Query: 165 DSTIKVSNPNQEVFVGAFQNGLRAGQFNESLAQKPADSMEEIIARAECYVKGEESNAEKR 224
++ + + E+ + A + L G LA+ + +++ R E + +GEE +
Sbjct: 462 RVRCQIRHVDDEMLIAAAKRALLPGPLRFDLARNRPKTTKDLFERMESFARGEEDELRVQ 521
Query: 225 ARDV--------KEKGSSGGERRNHYVPPNRDRGTFKKPYERNQPRYVPEHFTPLNTRPE 276
+ K K S GE + + FK Y ++Q + V N E
Sbjct: 522 EEEAVLLGKKQSKNKQISQGEEQKGE-NTGKPWKKFKYDYRQDQKKQVNFIGDGSNNERE 580
Query: 277 KILKEVFESKIIPPPPFNRSKTMGQ------------------------DKNAWCKYHLI 312
K + ++ K GQ D+ +C+ H
Sbjct: 581 KGKHQWDNTRRGRNNWGQSGKGRGQWWNSGRGRGRWWNNERGKGRENKPDQTKFCQTHGP 640
Query: 313 EGHNTDDCV-----------HLKREIEKL--LLNGKLRGYAKEKHHSERREDKPN----P 355
GH+T++C H E ++ LL + Y + K+ R + N P
Sbjct: 641 GGHSTEECYSKFCHIHGPGGHSTEECRQMTHLLEKHVNRY-ENKYEGARDQRGQNAIEAP 699
Query: 356 E----------PKHTLHTISGGFAGGGESSNLRKKYVRQVMLLGDS-PSSPKKYPH--VT 402
+ PK ++ I+GG + G ES RK YVRQV +G S S+P Y ++
Sbjct: 700 QVMKIEAIEEVPKRVINAITGGSSLGVESKRQRKAYVRQVNHVGTSYQSNPPVYSKTVIS 759
Query: 403 FSSEDFGRVLPHDDDPLVISVQLFNWEIKRVLIDTGSSADVLYFDAFSKMGLSEEQLQPF 462
F ED +L DPLV+SV++ E++RVLID GSSADVL++DAF KM + E++L
Sbjct: 760 FGPEDAEGILFPHQDPLVVSVEIAQCEVQRVLIDGGSSADVLFYDAFKKMQIPEDRLTNA 819
Query: 463 NGTLSGFTGEKVHVRGYVTLKTIFGTGDQQTSIKVRYLVINSPSSYNVIIGRPSINLLDA 522
L GF G++VH G ++L+ +FG G ++ + V++ P YN I+GR +IN+ +A
Sbjct: 820 GVPLQGFGGQQVHAIGKISLQVVFGKGTNVRKEEIVFDVVDMPYQYNAILGRSTINIFEA 879
Query: 523 FVSTKHLLMKYPLDSGRVGVVHGDQKIARECYHASLRLKKHKEFEVQVIDLDPREDFPQE 582
+ ++ MK P G + V G+Q AR+ K V V+D E +
Sbjct: 880 IIHHNYICMKLPGLRGVI-TVRGEQLAARKYELQGTPSVKG----VHVVDQKQGEYIKIQ 934
Query: 583 RIEPIEEVKDVVIGPL-PHQITKIGTSLSGLEEEILVQLLRENVDLFAWAPSDMPGIDIG 641
+ P + K V + P + IG +L EE ++++++EN+ +FAW+P ++ G+D
Sbjct: 935 KPIPEGKTKKVQLDEHNPGKFILIGENLEKHIEEEILKVVKENMAVFAWSPDELQGVDRS 994
Query: 642 VACHHLAVRTSVKPVVQRKRKMGEEKRKAVDEEVKKLQEAHFICEIKYPTWLANTVLVKK 701
+ H+LA+++ KP Q+ R+M ++++A E++KL +A I E+ +P WLAN VLVKK
Sbjct: 995 LIEHNLAIKSGYKPKKQKLRRMSTDRQQAAKIELEKLLKAKVIREVMHPEWLANPVLVKK 1054
Query: 702 ASGKWRMCVDYTDLNMACPKDPYPLPSIDHLIDNASGYKTLSFMDAYSGYNQIKMDPLDA 761
A+GKWRMC+D+TDLN ACPKD +PLP ID L+D +G + +SF+DAYSGY+Q+ M D
Sbjct: 1055 ANGKWRMCIDFTDLNKACPKDDFPLPRIDQLVDATAGCELMSFLDAYSGYHQVFMVKEDE 1114
Query: 762 PKTAFMTNQKNYHYRVMSFGLRNAGATFQRSMDTIFSNQIGRNLEVYIDDLVVKTSEKQS 821
KT+F+T Y + M FGL+NAGATF R + + + Q+GRN+E YIDD+VVK+ + +
Sbjct: 1115 EKTSFITPFGTYCFIRMPFGLKNAGATFARLIGKVLAKQLGRNVEAYIDDIVVKSKQAFT 1174
Query: 822 HSVDLKEIFQQIRKFSMRLNPTKCTFGVQAGKFLGFLLTKKGIEANPDKCQAIINMRNPC 881
H DL+E F+ +RK S++LNP KC FGV+AGK LGFL++K+GIEANPDK AI M P
Sbjct: 1175 HGKDLQETFENLRKCSVKLNPEKCVFGVRAGKLLGFLVSKRGIEANPDKIAAIHQMEPPK 1234
Query: 882 NIREVQQLTGRLAALSRFLSCAGDKAFAFFATIKKKEEFEWNQECDEAFGKIKQFLTTPP 941
N REVQ+LTGR+A+LSRFLS + +K FF T++ FEW EC +AF +K++L P
Sbjct: 1235 NTREVQRLTGRMASLSRFLSKSAEKGLPFFKTLRGANTFEWTAECQQAFDDLKKYLHEMP 1294
Query: 942 ILHRPTKGAGLFLYLSVSENALSSVLVEESDEREKPIYFVSRVLKGAELRYQKIEKLALA 1001
L P KG L +Y++ + +S+VLV+E + R+ P+YFVS L+G + RY ++EKL A
Sbjct: 1295 TLASPPKGQPLLMYVAATPATVSAVLVQEEENRQVPVYFVSEALQGPKTRYSEVEKLIYA 1354
Query: 1002 VIITARKLRPYFQSHKVVIRTNYPVKQILGKLDLAGRMLSWSVELSEYDIQFAPRNNIKS 1061
+++ +RKLR YF SH + I + YP+ ++L ++AGR+ W++EL +D+++ R IKS
Sbjct: 1355 IVMASRKLRHYFLSHDITIPSAYPIGEVLTNKEVAGRIAKWAMELLPFDLKYISRTAIKS 1414
Query: 1062 QVLADFVVEFTS---PIQEAVPHVWLLSVDGSSNIKGSGAGIILEGPGDLLIEQSLKFDF 1118
QVLADFV E+T QE V W++ DG+ N G+GA +++ P ++ S++ F
Sbjct: 1415 QVLADFVAEWTPNEVEQQEEVKKPWIVFSDGACNAAGAGAAAVVKTPMKQTLKYSVQLVF 1474
Query: 1119 KASNNQAEYEALIAGMLLAQEMGAKNLRARSDSQLMTNQISGEYQTKDQQLSKYLARVRK 1178
++NN AEYE ++ M A+ +GA+ L ++DS+L+ S ++ K++ ++KYL R
Sbjct: 1475 PSTNNTAEYEGVLLAMRKARALGARRLIVKTDSKLVAGHFSKSFEAKEETMAKYLEEARL 1534
Query: 1179 LAGDFQFFEAIYVPRESNSRADLLAKLASTKKPGNNRTVIQEVISAPSTDEKAVFELNQE 1238
F + RE N AD LAK A+T +P N ++I+ PS ++K V + +E
Sbjct: 1535 NEKHFLGITVKAITREENGEADELAKAAATGQPLENS--FFDIITQPSYEKKEVACIQRE 1592
Query: 1239 PEGWMTPLLKFLTGSFVAKNDEYAQLVRRRATKFVVIAGKLYKRGRASPLLRCLGEGETE 1298
+ W P+LK+L + + + +E A+ ++ + K+ V+ G+LYK G +PLL+C+ E
Sbjct: 1593 GD-WREPILKYLVSAQLPEKEEEAKRIQLMSKKYKVVEGQLYKSGVTAPLLKCVTREEGM 1651
Query: 1299 LVLLEVHEGVCGSHIGGRSLAAKLLRAGYYWPRMAHDCCEFVKKCDKCQRFSDKKNAPAN 1358
+++E+HEG+CG+H S+A+K++R G YWP + D +++K C CQ+F AP
Sbjct: 1652 KMVVEIHEGLCGAHQAPWSVASKVIRQGIYWPTIMKDTEKYIKTCKACQKFGPMTKAPPK 1711
Query: 1359 ELTSVFSPWPFHKWGVDIVGPFPQAPGQLKFLIVDVDYFTKWVEAEAVSKITAERVVKFY 1418
EL + WPF++WG+DIVGP P+A G L+F+IV ++YF++W+EAEAV++IT+ V KF
Sbjct: 1712 ELQPIPPVWPFYRWGIDIVGPLPRAKGDLRFVIVAIEYFSRWIEAEAVARITSAAVQKFV 1771
Query: 1419 WKKIICHFGLPKYIVTDNGTQFASSKVVNFCKQLGIETKFVSVIHPQANGQAESANKMIV 1478
WK IIC FG+PK IV DNG QF S K + CK L ++ F SV HPQ NG E AN I+
Sbjct: 1772 WKNIICRFGIPKEIVCDNGKQFESGKFQDMCKGLNLQINFASVGHPQTNGVVERANGKIM 1831
Query: 1479 NDIKKKLE-DAKGLWAEQLHEVLWSYHTTPHSTTGETPFTMVYGADAMLPVEIDTPTWRR 1537
IKK+LE AKG W E L VLW+ TT +TG TPF +VYG +AM P E+ + R
Sbjct: 1832 EAIKKRLEGSAKGKWPEDLLSVLWALRTTVVRSTGMTPFRLVYGDEAMTPSEVGAHSPRM 1891
Query: 1538 EHFSEESNEVGIRCTMDMIDEVREAAHIREFAAKQRAARRYNSKVIPRSMKEGDLVLKQV 1597
++ +E G T++M+DE+R A + + + YN KV R ++EGDLVLK+V
Sbjct: 1892 --IFDQKDEEGREITLEMLDEIRVEALEKMASYTEGTKSYYNQKVKTRPIEEGDLVLKKV 1949
Query: 1598 VAPTRIGKLLPSWKGPYRVKEKLQHGAYKLEELSGNPVPRT 1638
+ +GKL W+GP+ VK+K + GA+KL L G + T
Sbjct: 1950 LNEVAVGKLESKWEGPFIVKKKTETGAFKLAYLDGEELKHT 1990
>gb|AAF79618.1| F5M15.26 [Arabidopsis thaliana] gi|25402907|pir||H86337 protein
F5M15.26 [imported] - Arabidopsis thaliana
Length = 1838
Score = 1047 bits (2707), Expect = 0.0
Identities = 615/1678 (36%), Positives = 923/1678 (54%), Gaps = 123/1678 (7%)
Query: 38 LDEPEPQPFSTEIWNAPVPDNFKPPQLPTFDGRSDPSEHVTVFNTRMSVYGVA----DSL 93
L+E + PF+ I N + K +L +++GR+DP E +T FN ++ + D+
Sbjct: 196 LEETQQTPFTRRITNVSIRGAQKI-KLESYNGRNDPKEFLTSFNVAINRAELTIDNFDAG 254
Query: 94 KCKLLAGTLADAALRWYMSLPRFSIVGYQDLTKKFTQQFSGTRHRKVLSTSLFNVRQGPN 153
+C++ L A W+ L SI + LT F + ++ + + L+++ QG
Sbjct: 255 RCQIFIEHLTGPAHNWFSRLKPNSIDSFHQLTSSFLKHYAPLIENQTSNADLWSISQGAK 314
Query: 154 ESLREYLARFNDSTIKVSNPNQEVFVGAFQNGLRAGQFNESLAQKPADSMEEIIARAECY 213
ESLR ++ RF ++ P++ V +F + + ++E+ + RA +
Sbjct: 315 ESLRSFVDRFKLVVTNITVPDEAAIVALRNAVWYDSRFRDDITLHAPSTLEDALHRASRF 374
Query: 214 VKGEESNAEKRARDVKEKGSSGGERRNHYVPPNRDRGTFKK-PYERNQPRYVPEHFTPLN 272
++ EE EK + + P +D K P + N+PR +
Sbjct: 375 IELEE-----------EKLILARKHNSTKTPACKDAVVIKVGPDDSNEPRQHLDRNPSAG 423
Query: 273 TRPEKILKEVFESKIIPPPPFNRSKTMGQDKNAWCKYHLIEGHNTDDCVHLKR------- 325
+P L P + R +C+YH H+T++C L+
Sbjct: 424 RKPTSFLVSTETPDAKPWNKYIRDADSPAAGPMYCEYHKSRAHSTENCRFLQGLLMAKYK 483
Query: 326 ------EIEKLLLNGK--LRGYAKEKHHSERREDKPNPEPK------------------- 358
E ++ +N K R + + + P P +
Sbjct: 484 SGGITIECDRPPINNKNQRRNETTARQYLNDQTKPPTPAEQGIITSADDPAAKRQRNGKA 543
Query: 359 --------HTLHTISGGFAGGGESSNLRKKYVRQVMLLGDSPSS------PKKYPHVTFS 404
+H I GG +S K+Y ++ ++ PSS P + ++F+
Sbjct: 544 IAAEPVVVRQVHVIMGGLQNCSDSVRSIKQYRKKAEMVVAWPSSTSTTRNPNQSAPISFT 603
Query: 405 SEDF-GRVLPHDDDPLVISVQLFNWEIKRVLIDTGSSADVLYFDAFSKMGLSEEQLQPFN 463
D G PH+D PLV+ + + + + RVLIDTGSS D+++ D + M +++ Q++P +
Sbjct: 604 DVDLEGLDTPHND-PLVVELIISDSRVTRVLIDTGSSVDLIFKDVLTAMNITDRQIKPVS 662
Query: 464 GTLSGFTGEKVHVRGYVTLKTIFGTGDQQTSIKVRYLVINSPSSYNVIIGRPSINLLDAF 523
L+GF G+ V G + L G V+++VI P+ YNVI+G P I+ + A
Sbjct: 663 KPLAGFDGDFVMTIGTIKLPIFVGG----LIAWVKFVVIGKPAVYNVILGTPWIHQMQAI 718
Query: 524 VSTKHLLMKYPLDSGRVGVVHGDQKIARECYHASLRLKKHKEFEVQVIDLDPREDFPQER 583
ST H +K+P +G + + +E S ++ + +++++D +
Sbjct: 719 PSTYHQCVKFPTHNGIFTL-----RAPKEAKTPSRSYEESELCRTEMVNIDESD------ 767
Query: 584 IEPIEEVKDVVIGPLPHQITKIGTSLSGLEEEILVQLLRENVDLFAWAPSDMPGIDIGVA 643
P + +G +S L+ LL+ N FAW+ DM GID +
Sbjct: 768 ---------------PTRCVGVGAEISPSIRLELIALLKRNSKTFAWSIEDMKGIDPAIT 812
Query: 644 CHHLAVRTSVKPVVQRKRKMGEEKRKAVDEEVKKLQEAHFICEIKYPTWLANTVLVKKAS 703
H L V + KPV Q++RK+G E+ +AV+EEV+KL +A I E+KYP WLAN V+VKK +
Sbjct: 813 AHELNVDPTFKPVKQKRRKLGPERARAVNEEVEKLLKAGQIIEVKYPEWLANPVVVKKKN 872
Query: 704 GKWRMCVDYTDLNMACPKDPYPLPSIDHLIDNASGYKTLSFMDAYSGYNQIKMDPLDAPK 763
GKWR+CVDYTDLN ACPKD YPLP ID L++ SG LSFMDA+SGYNQI M D K
Sbjct: 873 GKWRVCVDYTDLNKACPKDSYPLPHIDRLVEATSGNGLLSFMDAFSGYNQILMHKDDQEK 932
Query: 764 TAFMTNQKNYHYRVMSFGLRNAGATFQRSMDTIFSNQIGRNLEVYIDDLVVKTSEKQSHS 823
T+F+T++ Y Y+VMSFGL+NAGAT+QR ++ + ++QIGR +EVYIDD++VK+ + + H
Sbjct: 933 TSFVTDRGTYCYKVMSFGLKNAGATYQRFVNKMLADQIGRTVEVYIDDMLVKSLKPEDHV 992
Query: 824 VDLKEIFQQIRKFSMRLNPTKCTFGVQAGKFLGFLLTKKGIEANPDKCQAIINMRNPCNI 883
L + F + + M+LNPTKCTFGV +G+FLG+++TK+GIEANP + +AI+ + +P N
Sbjct: 993 EHLSKCFDVLNTYGMKLNPTKCTFGVTSGEFLGYVVTKRGIEANPKQIRAILELPSPRNA 1052
Query: 884 REVQQLTGRLAALSRFLSCAGDKAFAFFATIKKKEEFEWNQECDEAFGKIKQFLTTPPIL 943
REVQ+LTGR+AAL+RF+S + DK F+ +K++ +F+W+++ +EAF K+K +L+TPPIL
Sbjct: 1053 REVQRLTGRIAALNRFISRSTDKCLPFYNLLKRRAQFDWDKDSEEAFEKLKDYLSTPPIL 1112
Query: 944 HRPTKGAGLFLYLSVSENALSSVLVEESDEREKPIYFVSRVLKGAELRYQKIEKLALAVI 1003
+P G L+LY++VS++A+SSVLV E ++PI++ S+ L AE RY IEK ALAV+
Sbjct: 1113 VKPEVGETLYLYIAVSDHAVSSVLVREDRGEQRPIFYTSKSLVEAETRYPVIEKAALAVV 1172
Query: 1004 ITARKLRPYFQSHKVVIRTNYPVKQILGKLDLAGRMLSWSVELSEYDIQFAPRNNIKSQV 1063
+ARKLRPYFQSH + + T+ P++ L +GRM W+VELSEYDI F PR +KSQV
Sbjct: 1173 TSARKLRPYFQSHTIAVLTDQPLRVALHSPSQSGRMTKWAVELSEYDIDFRPRPAMKSQV 1232
Query: 1064 LADFVVEFTSPIQEAVPHV-------WLLSVDGSSNIKGSGAGIILEGPGDLLIEQSLKF 1116
LADF++E P+Q A V W L VDGSS+ +GSG GI L P ++EQS +
Sbjct: 1233 LADFLIEL--PLQSAERAVSGNRGEEWSLYVDGSSSARGSGIGIRLVSPTAEVLEQSFRL 1290
Query: 1117 DFKASNNQAEYEALIAGMLLAQEMGAKNLRARSDSQLMTNQISGEYQTKDQQLSKYLARV 1176
F A+NN AEYE LIAG+ LA M + A +DSQL+ Q+SGEY+ K++++ YL V
Sbjct: 1291 RFVATNNVAEYEVLIAGLRLAAGMQITTIHAFTDSQLIAGQLSGEYEAKNEKMDAYLKIV 1350
Query: 1177 RKLAGDFQFFEAIYVPRESNSRADLLAKLASTKKPGNNRTVIQEVISAPSTDEKAVFEL- 1235
+ + DF+ F+ +PR N+ AD LA LA T R + E I PS D E+
Sbjct: 1351 QLMTKDFENFKLSKIPRGDNAPADALAALALTSDSDLRRIIPVESIDKPSIDSTDAVEIV 1410
Query: 1236 ----------NQEPEGWMTPLLKFLTGSFVAKNDEYAQLVRRRATKFVVIAGKLYKRGRA 1285
+P W + +L+ + + A+ +R +A K+ ++ L K
Sbjct: 1411 NTIRSSNAPDPADPTDWRVEIRDYLSDGTLPSDKWTARRLRIKAAKYTLMKEHLLKVSAF 1470
Query: 1286 SPLLRCLGEGETELVLLEVHEGVCGSHIGGRSLAAKLLRAGYYWPRMAHDCCEFVKKCDK 1345
+L CL E ++ E HEG G+H GGR+LA KL + G+YWP M DC F KC++
Sbjct: 1471 GAMLNCLHGTEINEIMKETHEGAAGNHSGGRALALKLKKLGFYWPTMISDCKTFTAKCEQ 1530
Query: 1346 CQRFSDKKNAPANELTSVFSPWPFHKWGVDIVGPFPQAPGQLKFLIVDVDYFTKWVEAEA 1405
CQR + + P L + +P+PF +W +DIVGP P A Q +F++V DYFTKWVEAE+
Sbjct: 1531 CQRHAPTIHQPTELLRAGVAPYPFMRWAMDIVGPMP-ASRQKRFILVMTDYFTKWVEAES 1589
Query: 1406 VSKITAERVVKFYWKKIICHFGLPKYIVTDNGTQFASSKVVNFCKQLGIETKFVSVIHPQ 1465
+ I A V F WK IIC GLP I+TDNG+QF S NFC I + +PQ
Sbjct: 1590 YATIRANDVQNFVWKFIICRHGLPYEIITDNGSQFISLSFENFCASWKIRLNKSTPRYPQ 1649
Query: 1466 ANGQAESANKMIVNDIKKKLEDAKGLWAEQLHEVLWSYHTTPHSTTGETPFTMVYGADAM 1525
NGQAE+ NK I++ +KK+L++ KG WA++L VLWSY TTP S T +TPF YG +AM
Sbjct: 1650 GNGQAEATNKTILSGLKKRLDEKKGAWADELDGVLWSYRTTPRSATDQTPFAHAYGMEAM 1709
Query: 1526 LPVEIDTPTWRREHF--SEESNEVGIRCTMDMIDEVREAAHIREFAAKQRAARRYNSKVI 1583
P E+ + RR + E N+ + +D ++E+R AA R + AA+ YN KV
Sbjct: 1710 APAEVGYSSLRRSMMVKNPELNDRMMLDRLDDLEEIRNAALCRIQNYQLAAAKHYNQKVH 1769
Query: 1584 PRSMKEGDLVLKQV---VAPTRIGKLLPSWKGPYRVKEKLQHGAYKLEELSGNPVPRT 1638
R GDLVL++V A GKL +W+G Y+V + ++ G Y+L +SG VPRT
Sbjct: 1770 NRHFDVGDLVLRKVFENTAEINAGKLGANWEGSYQVSKIVRPGDYELLTMSGTAVPRT 1827
>ref|XP_474675.1| OSJNBa0027O01.4 [Oryza sativa (japonica cultivar-group)]
gi|38346188|emb|CAD39529.2| OSJNBa0027O01.4 [Oryza sativa
(japonica cultivar-group)]
Length = 2013
Score = 956 bits (2470), Expect = 0.0
Identities = 574/1653 (34%), Positives = 868/1653 (51%), Gaps = 91/1653 (5%)
Query: 25 VQQQEKTDSHHG----DLDEPEPQPFSTEIWNAPVPDNFKPPQLPTFDGRSDPSEHVTVF 80
+Q++ HH D D F+ ++ P FKP + +DG ++P +TV+
Sbjct: 396 IQRRAARGHHHSPNRHDDDLDGVAAFTDDLRRVDWPAGFKPTGIEKYDGTTNPESWLTVY 455
Query: 81 NTRMSVYGVADSLKCKLLAGTLADAALRWYMSLPRFSIVGYQDLTKKFTQQFSGTRHRKV 140
+ G + L LAD+A W LPR +I + +L F F GT R
Sbjct: 456 GLAIRAAGGDNKAMANYLPVALADSARSWLHGLPRGTIGSWAELRDHFIANFQGTFERPG 515
Query: 141 LSTSLFNVRQGPNESLREYLARFNDSTIKVSNPNQEVFVGAFQNGLRAGQFNESLAQKPA 200
L+NV Q ESLR+Y+ RF++ K+S+ +V + AF G+R + +KP
Sbjct: 516 TQYDLYNVIQKSGESLRDYIRRFSEQRNKISDITDDVIIAAFTKGIRHEELVGKFGRKPP 575
Query: 201 DSMEEIIARAECYVKGEESNAEKRARDVKEKGSSGGERRNHYVPPNRDRGTFKKPYERNQ 260
+++ + +A Y K E++ K+ G S ++N P G+ + +
Sbjct: 576 RTVKLMFEKANEYAKAEDAVTAS-----KQSGPSW--KQNKGTPATGGGGSNNHKDRKRK 628
Query: 261 PRYVPEHFTPLNTRPEKILKEVFESKIIPPPPFNRSKTMGQDKNAWCKYHLIEGHNTDDC 320
P E+ + +R T + N+ C +H H DC
Sbjct: 629 PA------------------ELVATASHSSRQRSRVNTFDKIMNSQCPHHPNSNHVAKDC 670
Query: 321 VHLKREIEKLLLNGKLRGYAKEKHHSERREDKPNP-----EPKHTLHTISGGFAGGGESS 375
K+ ++ + + +E+ S +++D P K H G A ES
Sbjct: 671 FVYKQFADQYTKTAR-KNSDEEQSTSRKKDDGDTPAGFQDHRKELNHIFGGPLAY--ESK 727
Query: 376 NLRKKYVRQVMLLGDSPSSPKKYPHVTFS---SEDFGRVLPHDDDPLVISVQLFNWEIKR 432
+K R++ + ++ + S+ RV+ PLV+ + N +++R
Sbjct: 728 RKQKLTEREINAVQPDTPQYLRWSEIAIKFDRSDHPDRVVHPGRYPLVLDPVVRNVKLRR 787
Query: 433 VLIDTGSSADVLYFDAFSKMGLSEEQLQPFNGTLSG-FTGEKVHVRGYVTLKTIFGTGDQ 491
LID GS+ ++L+ M + +L+P N G G G +TL FGT +
Sbjct: 788 TLIDGGSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPLGQITLPVTFGTREN 847
Query: 492 QTSIKVRYLVINSPSSYNVIIGRPSINLLDAFVSTKHLLMKYPLDSGRVGVVHGDQKIAR 551
+ + + V + ++Y+ I+GRP++ A +++MK P G + + D K A
Sbjct: 848 FRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMKMPGPRGVLSL-RSDIKQAV 906
Query: 552 ECYHASLRLKKHKEFEVQVIDLDPREDFPQERIEPIEEVKDVVIGPLPH-QITKIGTSLS 610
C S + + +E RED E G +P +I+K G S +
Sbjct: 907 TCDKESCDMAQTREMA------SAREDIRLAAATASE-------GEVPATKISKSGESEA 953
Query: 611 GLEEEIL----VQLLRENVDLFAWAPSDMPGIDIGVACHHLAVRTSVKPVVQRKRKMGEE 666
++ L N D+FAW PSDMPGI V H L V+ KP+ QR R+ ++
Sbjct: 954 KTKKIPLDPSDPTKTANNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQD 1013
Query: 667 KRKAVDEEVKKLQEAHFICEIKYPTWLANTVLVKKASGKWRMCVDYTDLNMACPKDPYPL 726
++ A+ EE+ KL A FI E+ +P WLAN VLV+K +G+WRMCVDYTDLN +CPKDP+ L
Sbjct: 1014 RKDAIKEELTKLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGL 1073
Query: 727 PSIDHLIDNASGYKTLSFMDAYSGYNQIKMDPLDAPKTAFMTNQKNYHYRVMSFGLRNAG 786
P ID ++D+ +G + LSF+D YSGY+QI++ D KT+F+T Y Y M FGL+NAG
Sbjct: 1074 PRIDQVVDSTAGCELLSFLDCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMPFGLKNAG 1133
Query: 787 ATFQRSMDTIFSNQIGRNLEVYIDDLVVKTSEKQSHSVDLKEIFQQIRKFSMRLNPTKCT 846
AT+QR + FS QIGRN+E Y+DD+VVKT +K DL+E F IR F M+LNP KCT
Sbjct: 1134 ATYQRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCT 1193
Query: 847 FGVQAGKFLGFLLTKKGIEANPDKCQAIINMRNPCNIREVQQLTGRLAALSRFLSCAGDK 906
FGV +GK LGF+++ +GI+ANP+K AI+NM+ P ++VQ+LTG +AALSRF+S G++
Sbjct: 1194 FGVPSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGER 1253
Query: 907 AFAFFATIKKKEEFEWNQECDEAFGKIKQFLTTPPILHRPTKGAGLFLYLSVSENALSSV 966
FF +KK ++F+W E +AF K+ LT PP+L P L LY+S + +S+V
Sbjct: 1254 GMPFFKLLKKTDDFQWGPEAQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTV 1313
Query: 967 LVEESDER------EKPIYFVSRVLKGAELRYQKIEKLALAVIITARKLRPYFQSHKVVI 1020
LV E +E ++PIYFVS VL ++ RY +++KL ++IT RKL YFQ H V +
Sbjct: 1314 LVVEREEEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTV 1373
Query: 1021 RTNYPVKQILGKLDLAGRMLSWSVELSEYDIQFAPRNNIKSQVLADFVVEFTSPIQEAVP 1080
T++P+ IL ++ GR+ W++EL DI F PR +IKSQ LADFV E+T QE P
Sbjct: 1374 VTSFPLGDILHNREVNGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTE-CQEDTP 1432
Query: 1081 ----HVWLLSVDGSSNIKGSGAGIILEGPGDLLIEQSLKFDFKASNNQAEYEALIAGMLL 1136
W + DGS + G+GAG++L P + L F AS+N AEYEAL+ G+ +
Sbjct: 1433 AENMEHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRI 1492
Query: 1137 AQEMGAKNLRARSDSQLMTNQISGEYQTKDQQLSKYLARVRKLAGDFQFFEAIYVPRESN 1196
A +G K L R DSQL+ NQ+ E+ D + Y VRKL F E +V R +N
Sbjct: 1493 AISLGIKRLIVRGDSQLVVNQVMKEWSYLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNN 1552
Query: 1197 SRADLLAKLASTKKPGNNRTVIQ----------EVISAPSTDEKAVFELNQEPEGWMTPL 1246
AD LA S ++ + ++ + T + A+ E++ W PL
Sbjct: 1553 EAADRLANFGSKREAAPSDVFVEHLYTPTVPHKDTTQVAGTHDAAMVEVD-----WREPL 1607
Query: 1247 LKFLTGSFVAKNDEYAQLVRRRATKFVVIAGKLYKRGRASPLLRCLGEGETELVLLEVHE 1306
++FLT + ++ + A+ + RR+ +V+ +LYK+ + L RC+ E +L ++H
Sbjct: 1608 IRFLTSQELPQDKDEAERISRRSKLYVLHEAELYKKSPSGILQRCVSLEEGRQLLKDIHS 1667
Query: 1307 GVCGSHIGGRSLAAKLLRAGYYWPRMAHDCCEFVKKCDKCQRFSDKKNAPANELTSVFSP 1366
G+CG+H R++ K R G++WP D + V+ C+ CQ F+ + + PA EL ++
Sbjct: 1668 GICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLS 1727
Query: 1367 WPFHKWGVDIVGPFPQAPGQLKFLIVDVDYFTKWVEAEAVSKITAERVVKFYWKKIICHF 1426
WPF WG+D+VGPF +A G L V +D F+KW+EA+ V ITA+ F+ I+ F
Sbjct: 1728 WPFAVWGLDMVGPFKKAVGGYTHLFVAIDKFSKWIEAKPVVTITADNARDFF-INIVHRF 1786
Query: 1427 GLPKYIVTDNGTQFASSKVVNFCKQLGIETKFVSVIHPQANGQAESANKMIVNDIKKKLE 1486
G+P I+TDNGTQF +FC+ GI+ + SV HP +NGQ E AN MI+ IK ++
Sbjct: 1787 GVPNRIITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVF 1846
Query: 1487 DA----KGLWAEQLHEVLWSYHTTPHSTTGETPFTMVYGADAMLPVEIDTPTWRREHFSE 1542
D G W +QL VLWS TTP TG++PF +VYGA+AMLP E++ + R +F E
Sbjct: 1847 DRLKPYAGKWVQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFRE 1906
Query: 1543 ESNEVGIRCTMDMIDEVREAAHIREFAAKQRAARRYNSKVIPRSMKEGDLVLKQVVAPTR 1602
E E + ++EVREAA IR Q R +N V R+ GDLVL+++
Sbjct: 1907 ERYEEDRVDDLHRLEEVREAALIRSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTTRD 1966
Query: 1603 IGKLLPSWKGPYRVKEKLQHGAYKLEELSGNPV 1635
KL P W+GP+ + E + G+Y+L+ G V
Sbjct: 1967 RHKLSPLWEGPFIISEVTRPGSYRLKREDGTLV 1999
>ref|XP_473950.1| OSJNBa0053K19.16 [Oryza sativa (japonica cultivar-group)]
gi|38344177|emb|CAE03508.2| OSJNBa0053K19.16 [Oryza
sativa (japonica cultivar-group)]
Length = 2010
Score = 955 bits (2469), Expect = 0.0
Identities = 568/1643 (34%), Positives = 860/1643 (51%), Gaps = 71/1643 (4%)
Query: 25 VQQQEKTDSHHG----DLDEPEPQPFSTEIWNAPVPDNFKPPQLPTFDGRSDPSEHVTVF 80
+Q++ HH D D F+ ++ P FKP + +DG ++P +TV+
Sbjct: 393 IQRRAARGHHHSPNRHDDDLDGVAAFTDDLRRVDWPAGFKPTGIEKYDGTTNPESWLTVY 452
Query: 81 NTRMSVYGVADSLKCKLLAGTLADAALRWYMSLPRFSIVGYQDLTKKFTQQFSGTRHRKV 140
+ G + L LAD+A W LPR +I + +L F F GT R
Sbjct: 453 GLAIRAAGGDNKAMANYLPVALADSARSWLHGLPRGTIGSWAELRDHFIANFQGTFERPG 512
Query: 141 LSTSLFNVRQGPNESLREYLARFNDSTIKVSNPNQEVFVGAFQNGLRAGQFNESLAQKPA 200
L+NV Q ESLR+Y+ RF++ K+S+ +V + AF G+R + +KP
Sbjct: 513 TQYDLYNVIQKSGESLRDYIRRFSEQRNKISDITDDVIIAAFTKGIRHEELVGKFGRKPP 572
Query: 201 DSMEEIIARAECYVKGEESNAEKRARDVKEKGSSGGERRNHYVPPNRDRGTFKKPYERNQ 260
+++ + +A Y K E++ K+ G S ++N P G+ + +
Sbjct: 573 MTVKLMFEKANEYAKAEDAVTAS-----KQSGPSW--KQNKGTPATGGGGSNNHKDRKRK 625
Query: 261 PRYVPEHFTPLNTRPEKILKEVFESKIIPPPPFNRSKTMGQDKNAWCKYHLIEGHNTDDC 320
P E+ + +R T + N+ C +H H DC
Sbjct: 626 PA------------------ELVATASHSSRQRSRVNTFDKIMNSQCSHHPNSNHVAKDC 667
Query: 321 VHLKREIEKLLLNGKLRGYAKEKHHSERREDKPNP-----EPKHTLHTISGGFAGGGESS 375
K+ ++ + + +E+ S +++D P K H G A ES
Sbjct: 668 FVYKQFADQYTKTAR-KNSDEEQSTSRKKDDGDTPAGFQDHRKELNHIFGGPLAY--ESK 724
Query: 376 NLRKKYVRQVMLLGDSPSSPKKYPHVTFS---SEDFGRVLPHDDDPLVISVQLFNWEIKR 432
+K R++ + ++ + S+ RV+ PLV+ + N +++R
Sbjct: 725 RKQKLTEREINAVQPDTPQYLRWSEIAIKFDRSDHPDRVVHPGRYPLVLDPVVRNVKLRR 784
Query: 433 VLIDTGSSADVLYFDAFSKMGLSEEQLQPFNGTLSG-FTGEKVHVRGYVTLKTIFGTGDQ 491
LID GS+ ++L+ M + +L+P N G G G +TL FGT +
Sbjct: 785 TLIDGGSALNILFAKTLDDMQIPHSELKPSNAPFHGVIPGLSATPLGQITLPVTFGTREN 844
Query: 492 QTSIKVRYLVINSPSSYNVIIGRPSINLLDAFVSTKHLLMKYPLDSGRVGVVHGDQKIAR 551
+ + + V + ++Y+ I+GRP++ A +++MK P G + + D K A
Sbjct: 845 FRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMKMPGPRGVLSL-RSDIKQAV 903
Query: 552 ECYHASLRLKKHKEFEVQVIDLDPREDFPQERIEPIEEVKDVVIGPLPHQITKIGTSLSG 611
C S + + +E D+ E EP ++ G + KI S
Sbjct: 904 TCDKESCDMAQTREMASAREDIRLAAATASEGEEPATKISKS--GESEAKTKKIPLDPSD 961
Query: 612 LEEEILVQLLRENVDLFAWAPSDMPGIDIGVACHHLAVRTSVKPVVQRKRKMGEEKRKAV 671
+ N D+FAW PSDMPGI V H L V+ KP+ QR R+ ++++ A+
Sbjct: 962 PAKTA------NNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAI 1015
Query: 672 DEEVKKLQEAHFICEIKYPTWLANTVLVKKASGKWRMCVDYTDLNMACPKDPYPLPSIDH 731
EE+ KL A FI E+ +P WLAN VLV+K +G+WRMCVDYTDLN +CPKDP+ LP ID
Sbjct: 1016 KEELTKLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPRIDQ 1075
Query: 732 LIDNASGYKTLSFMDAYSGYNQIKMDPLDAPKTAFMTNQKNYHYRVMSFGLRNAGATFQR 791
++D+ +G + LSF+D YSGY+QI++ D KT+F+T Y Y M FGL+NAGAT+QR
Sbjct: 1076 VVDSTAGRELLSFLDCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMPFGLKNAGATYQR 1135
Query: 792 SMDTIFSNQIGRNLEVYIDDLVVKTSEKQSHSVDLKEIFQQIRKFSMRLNPTKCTFGVQA 851
+ FS QIGRN+E Y+DD+VVKT +K DL+E F IR F M+LNP KCTFGV +
Sbjct: 1136 MIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGVPS 1195
Query: 852 GKFLGFLLTKKGIEANPDKCQAIINMRNPCNIREVQQLTGRLAALSRFLSCAGDKAFAFF 911
GK +GF+++ +GI+ANP+K AI+NM+ P ++VQ+LTG +AALSRF+S G++ FF
Sbjct: 1196 GKLVGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFF 1255
Query: 912 ATIKKKEEFEWNQECDEAFGKIKQFLTTPPILHRPTKGAGLFLYLSVSENALSSVLVEES 971
+KK + F+W E +AF K+ LT PP+L P L LY+S + +S+VLV E
Sbjct: 1256 KLLKKTDNFQWGPEAQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTVLVVER 1315
Query: 972 DER------EKPIYFVSRVLKGAELRYQKIEKLALAVIITARKLRPYFQSHKVVIRTNYP 1025
+E ++PIYFVS VL ++ RY +++KL ++IT RKL YFQ H V + T++P
Sbjct: 1316 EEEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSFP 1375
Query: 1026 VKQILGKLDLAGRMLSWSVELSEYDIQFAPRNNIKSQVLADFVVEFTSPIQEAVP----H 1081
+ IL + GR+ W++EL DI F PR +IKSQ LADFV E+T QE P
Sbjct: 1376 LGDILHNREANGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTE-CQEDTPAENME 1434
Query: 1082 VWLLSVDGSSNIKGSGAGIILEGPGDLLIEQSLKFDFKASNNQAEYEALIAGMLLAQEMG 1141
W + DGS + G+GAG++L P + L F AS+N AEYEAL+ G+ +A +G
Sbjct: 1435 HWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLG 1494
Query: 1142 AKNLRARSDSQLMTNQISGEYQTKDQQLSKYLARVRKLAGDFQFFEAIYVPRESNSRADL 1201
K L R DSQL+ NQ+ E+ D + Y VRKL F E +V R +N AD
Sbjct: 1495 IKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADR 1554
Query: 1202 LAKLASTKKPGNNRTVIQEVIS-----APSTDEKAVFELNQEPEGWMTPLLKFLTGSFVA 1256
LA S ++ + ++ + + +T ++ W PL++FLT +
Sbjct: 1555 LANFGSKREVAPSDVFVEHLYTPTVPHKDTTQVAGTHDVAMVETDWREPLIRFLTSQELP 1614
Query: 1257 KNDEYAQLVRRRATKFVVIAGKLYKRGRASPLLRCLGEGETELVLLEVHEGVCGSHIGGR 1316
++ + A+ + RR+ +V+ +LYK+ + L RC+ E +L ++H G+CG+H R
Sbjct: 1615 QDKDEAERISRRSKLYVMHEAELYKKSPSGILQRCVSLEEGRQLLQDIHSGICGNHAAAR 1674
Query: 1317 SLAAKLLRAGYYWPRMAHDCCEFVKKCDKCQRFSDKKNAPANELTSVFSPWPFHKWGVDI 1376
++ K R G++WP D + V+ C+ CQ F+ + + PA EL ++ WPF WG+D+
Sbjct: 1675 TIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWGLDM 1734
Query: 1377 VGPFPQAPGQLKFLIVDVDYFTKWVEAEAVSKITAERVVKFYWKKIICHFGLPKYIVTDN 1436
VGPF +A G L V +D F+KW+EA+ V ITA+ F+ I+ FG+P I+TDN
Sbjct: 1735 VGPFKKAVGGYTHLFVAIDKFSKWIEAKPVVTITADNARDFF-INIVHRFGVPNRIITDN 1793
Query: 1437 GTQFASSKVVNFCKQLGIETKFVSVIHPQANGQAESANKMIVNDIKKKLEDA----KGLW 1492
GTQF +FC+ GI+ + SV HP +NGQ E AN MI+ IK ++ D G W
Sbjct: 1794 GTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPYAGKW 1853
Query: 1493 AEQLHEVLWSYHTTPHSTTGETPFTMVYGADAMLPVEIDTPTWRREHFSEESNEVGIRCT 1552
+QL VLWS TTP TG++PF +VYGA+AMLP E++ + R +F EE E
Sbjct: 1854 VQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFREERYEEDRVDD 1913
Query: 1553 MDMIDEVREAAHIREFAAKQRAARRYNSKVIPRSMKEGDLVLKQVVAPTRIGKLLPSWKG 1612
+ ++EVREAA IR Q R +N V R+ GDLVL+++ KL P W+G
Sbjct: 1914 LHRLEEVREAALIRSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTTRDRHKLSPLWEG 1973
Query: 1613 PYRVKEKLQHGAYKLEELSGNPV 1635
P+ + E + G+Y+L+ G V
Sbjct: 1974 PFIISEVTRPGSYRLKREDGTLV 1996
>gb|AAP53095.1| putative retroelement [Oryza sativa (japonica cultivar-group)]
gi|37533012|ref|NP_920808.1| putative retroelement [Oryza
sativa (japonica cultivar-group)]
gi|19881590|gb|AAM00991.1| Putative retroelement [Oryza
sativa]
Length = 2017
Score = 954 bits (2465), Expect = 0.0
Identities = 577/1641 (35%), Positives = 859/1641 (52%), Gaps = 93/1641 (5%)
Query: 32 DSHHGDLDEPEPQPFSTEIWNAPVPDNFKPPQLPTFDGRSDPSEHVTVFNTRMSVYGVAD 91
D H DLD F+ ++ P FKP + +DG ++P +TV+ + G
Sbjct: 413 DRHDDDLDGVAA--FTDDLRRVDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDS 470
Query: 92 SLKCKLLAGTLADAALRWYMSLPRFSIVGYQDLTKKFTQQFSGTRHRKVLSTSLFNVRQG 151
L LAD+A W LPR +I + +L F F GT R L+NV Q
Sbjct: 471 KAMANYLPVALADSARSWLHGLPRGTIESWAELRDHFIANFQGTFERPGTQYDLYNVIQK 530
Query: 152 PNESLREYLARFNDSTIKVSNPNQEVFVGAFQNGLRAGQFNESLAQKPADSMEEIIARAE 211
ESLR+Y+ RF++ K+S+ +V + AF G+R + +KP +++ + +A
Sbjct: 531 SGESLRDYIRRFSEQRNKISDITDDVIIAAFTKGIRHEELVGKFGRKPPRTVKLMFEKAN 590
Query: 212 CYVKGEESNAEKRARDVKEK------GSSGGERRNHYVPPNRDRGTFKKPYERNQPRYVP 265
Y K E++ + K + GG NH +DR
Sbjct: 591 EYAKAEDAVTASKQSGPSWKPNKGTPATGGGGSNNH-----KDR---------------- 629
Query: 266 EHFTPLNTRPEKILKEVFESKIIPPPPFNRSKTMGQDKNAWCKYHLIEGHNTDDCVHLKR 325
+PE+++ S +R T + N+ C +H H DC K+
Sbjct: 630 ------KRKPEELVATATHSS----RQRSRVNTFDKIMNSQCPHHPNSNHVAKDCFVYKQ 679
Query: 326 EIEKLLLNGKLRGYAKEKHHSERREDKPNP-----EPKHTLHTISGGFAGGGESSNLRKK 380
E+ + + +E+ S +++D P K H G A ES +K
Sbjct: 680 FAEQYTKTTR-KNSDEEQSTSRKKDDGDTPAGFQDHRKELNHIFGGPLAY--ESKRKQKL 736
Query: 381 YVRQVMLLGDSPSSPKKYPHVTFS---SEDFGRVLPHDDDPLVISVQLFNWEIKRVLIDT 437
R++ + ++ + S+ RV+ PLV+ + N +++R LID
Sbjct: 737 TEREINAVQPDTPQYLRWSEIAIKFDRSDHPDRVVHPGRYPLVLDPVVRNVKLRRTLIDG 796
Query: 438 GSSADVLYFDAFSKMGLSEEQLQPFNGTLSG-FTGEKVHVRGYVTLKTIFGTGDQQTSIK 496
GS+ ++L+ M + +L+P N G G G +TL FGT + +
Sbjct: 797 GSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPLGQITLPVTFGTRENFRTEN 856
Query: 497 VRYLVINSPSSYNVIIGRPSINLLDAFVSTKHLLMKYPLDSGRVGVVHGDQKIARECYHA 556
+ + V + ++Y+ I+GRP++ A +++MK P G + + D K A C
Sbjct: 857 ISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMKLPGPRGVLSL-RSDIKQAVTCDKE 915
Query: 557 SLRLKKHKEFEVQVIDLDPREDFPQERIEPIEEVKDVVIGPLPH-QITKIGTSLSGLEEE 615
S + + +E RED E G +P +I+K G S + ++
Sbjct: 916 SCDMAQTREMA------SAREDIRLAAATASE-------GEVPATKISKSGESEAKTKKI 962
Query: 616 IL----VQLLRENVDLFAWAPSDMPGIDIGVACHHLAVRTSVKPVVQRKRKMGEEKRKAV 671
L N D+FAW PSDMPGI V H L V+ KP+ QR R+ ++++ A+
Sbjct: 963 PLDPSDSTKTANNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAI 1022
Query: 672 DEEVKKLQEAHFICEIKYPTWLANTVLVKKASGKWRMCVDYTDLNMACPKDPYPLPSIDH 731
EE+ KL A FI E+ +P WLAN VLV+K +G+WRMCVDYTDLN CPKDP+ LP ID
Sbjct: 1023 KEELTKLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKCCPKDPFGLPRIDQ 1082
Query: 732 LIDNASGYKTLSFMDAYSGYNQIKMDPLDAPKTAFMTNQKNYHYRVMSFGLRNAGATFQR 791
++D+ +G + LSF+D YSGY+QI++ D KT+F+T Y Y M FGL+NAGAT+QR
Sbjct: 1083 VVDSTAGCELLSFLDCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMPFGLKNAGATYQR 1142
Query: 792 SMDTIFSNQIGRNLEVYIDDLVVKTSEKQSHSVDLKEIFQQIRKFSMRLNPTKCTFGVQA 851
+ FS QIGRN+E Y+DD+VVKT +K DL+E F IR F M+LNP KCTFGV +
Sbjct: 1143 MIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGVPS 1202
Query: 852 GKFLGFLLTKKGIEANPDKCQAIINMRNPCNIREVQQLTGRLAALSRFLSCAGDKAFAFF 911
GK LGF+++ +GI+ANP+K AI+NM+ P ++VQ+LTG +AALSRF+S G++ FF
Sbjct: 1203 GKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFF 1262
Query: 912 ATIKKKEEFEWNQECDEAFGKIKQFLTTPPILHRPTKGAGLFLYLSVSENALSSVLVEES 971
+KK ++F W E +AF K+ LT PP+L P L LY+S + +S+VLV E
Sbjct: 1263 KLLKKTDDFHWGPEAQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTVLVVER 1322
Query: 972 DER------EKPIYFVSRVLKGAELRYQKIEKLALAVIITARKLRPYFQSHKVVIRTNYP 1025
+E ++PIYFVS VL ++ RY +++KL ++IT RKL YFQ H V + T++P
Sbjct: 1323 EEDGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHLVTVVTSFP 1382
Query: 1026 VKQILGKLDLAGRMLSWSVELSEYDIQFAPRNNIKSQVLADFVVEFTSPIQEAVP----H 1081
+ IL + GR+ W++EL DI F PR +IKSQ LADFV E+T QE P
Sbjct: 1383 LGDILHNREANGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTE-CQEDTPAEKVE 1441
Query: 1082 VWLLSVDGSSNIKGSGAGIILEGPGDLLIEQSLKFDFKASNNQAEYEALIAGMLLAQEMG 1141
W + DGS + G+GAG++L P + L F AS+N AEYEAL+ G+ +A +G
Sbjct: 1442 HWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLG 1501
Query: 1142 AKNLRARSDSQLMTNQISGEYQTKDQQLSKYLARVRKLAGDFQFFEAIYVPRESNSRADL 1201
K L R DSQL+ NQ+ E+ D + Y VRKL F E +V R +N AD
Sbjct: 1502 IKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNKAADR 1561
Query: 1202 LAKLASTKKPGNNRTVIQEVISAPSTDEK------AVFELNQEPEGWMTPLLKFLTGSFV 1255
LA S ++ + V E + AP+ K ++ W PL++FLT +
Sbjct: 1562 LANFGSKREVAPS-DVFVEHLYAPTVPHKDTTQVAGTHDVAMVEADWREPLIRFLTSQEL 1620
Query: 1256 AKNDEYAQLVRRRATKFVVIAGKLYKRGRASPLLRCLGEGETELVLLEVHEGVCGSHIGG 1315
++ + A+ + RR+ +V+ +LYK+ + L RC+ E +L ++H G+CG+H
Sbjct: 1621 PQDKDEAERISRRSRLYVMHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNHAAA 1680
Query: 1316 RSLAAKLLRAGYYWPRMAHDCCEFVKKCDKCQRFSDKKNAPANELTSVFSPWPFHKWGVD 1375
R++ K R G++WP D + V+ C+ CQ F+ + + PA EL ++ WPF WG+D
Sbjct: 1681 RTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWGLD 1740
Query: 1376 IVGPFPQAPGQLKFLIVDVDYFTKWVEAEAVSKITAERVVKFYWKKIICHFGLPKYIVTD 1435
+VGPF +A G L V +D F+KW+EA+ V ITA+ F+ I+ FG+P I+TD
Sbjct: 1741 MVGPFKKAFGGYTHLFVAIDKFSKWIEAKPVVTITADNARDFF-INIVHRFGVPNRIITD 1799
Query: 1436 NGTQFASSKVVNFCKQLGIETKFVSVIHPQANGQAESANKMIVNDIKKKLEDA----KGL 1491
NG QF +FC+ GI+ + SV HP +NGQ E AN MI+ IK ++ D G
Sbjct: 1800 NGRQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPYAGK 1859
Query: 1492 WAEQLHEVLWSYHTTPHSTTGETPFTMVYGADAMLPVEIDTPTWRREHFSEESNEVGIRC 1551
W EQL VLWS TTP TG++PF +VYGA+AMLP E++ + R +F EE E
Sbjct: 1860 WVEQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFREERYEEDRVD 1919
Query: 1552 TMDMIDEVREAAHIREFAAKQRAARRYNSKVIPRSMKEGDLVLKQVVAPTRIGKLLPSWK 1611
+ ++EVREAA I+ Q R +N V R+ GDLVL+++ KL P W+
Sbjct: 1920 DLHRLEEVREAALIQSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTTRDRHKLSPLWE 1979
Query: 1612 GPYRVKEKLQHGAYKLEELSG 1632
GP+ + E + G+Y+L+ G
Sbjct: 1980 GPFIISEVTRPGSYRLKREDG 2000
>ref|XP_470757.1| putative gag-pol precursor [Oryza sativa] gi|18071370|gb|AAL58229.1|
putative gag-pol precursor [Oryza sativa]
Length = 2026
Score = 954 bits (2465), Expect = 0.0
Identities = 575/1646 (34%), Positives = 865/1646 (51%), Gaps = 97/1646 (5%)
Query: 32 DSHHGDLDEPEPQPFSTEIWNAPVPDNFKPPQLPTFDGRSDPSEHVTVFNTRMSVYGVAD 91
D H DLD F+ ++ P FKP + +DG ++P +TV+ + G
Sbjct: 413 DRHDDDLDGVAA--FTDDLRRVDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDS 470
Query: 92 SLKCKLLAGTLADAALRWYMSLPRFSIVGYQDLTKKFTQQFSGTRHRKVLSTSLFNVRQG 151
L LAD+A W LPR +I + +L F F GT R L+NV Q
Sbjct: 471 KAMANYLPVALADSARSWLHGLPRGTIGSWAELRDHFIANFQGTFERPGTQFDLYNVVQK 530
Query: 152 PNESLREYLARFNDSTIKVSNPNQEVFVGAFQNGLRAGQFNESLAQKPADSMEEIIARAE 211
ESLR+Y+ RF++ K+S+ +V + AF G+R + +KP +++ + +A
Sbjct: 531 SGESLRDYIRRFSEQRNKISDITDDVIIAAFTKGIRHEELVGKFGRKPPRTVKLMFEKAN 590
Query: 212 CYVKGEES-NAEKRA----RDVKEKGSSGGERRNHYVPPNRDRGTFKKPYERNQPRYVPE 266
Y K E++ A K++ + K+ ++GG N++ +DR
Sbjct: 591 EYAKAEDAVTASKQSGPSWKPKKDTPATGGGGSNNH----KDR----------------- 629
Query: 267 HFTPLNTRPEKILKEVFESKIIPPPPFNRSKTMGQDKNAWCKYHLIEGHNTDDCVHLKRE 326
+PE+++ S +R T + N+ C +H H DC K+
Sbjct: 630 -----KRKPEELVATAIHSS----RQRSRVNTFDKIMNSQCPHHPNSNHVAKDCFVYKQF 680
Query: 327 IEKLLLNGKLRGYAKEKHHSERREDKPNP-----EPKHTLHTISGGFAGGGESSNLRKKY 381
E+ + + +E+ S +++D P K H G A ES +K
Sbjct: 681 AEQYTKTTR-KNSDEEQSTSRKKDDGDTPVGFQDHRKELNHIFGGPLAY--ESKRKQKLT 737
Query: 382 VRQVMLLGDSPSSPKKYPHVTFS---SEDFGRVLPHDDDPLVISVQLFNWEIKRVLIDTG 438
R++ + ++ + S+ RV+ PLV+ + N +++R LID G
Sbjct: 738 EREINAVQPDTPQYLRWSEIAIKFDRSDHPDRVVHPGRYPLVLDPVVRNVKLRRTLIDGG 797
Query: 439 SSADVLYFDAFSKMGLSEEQLQPFNGTLSG-FTGEKVHVRGYVTLKTIFGTGDQQTSIKV 497
S+ ++L+ M + +L+P N G G G +TL FGT + + +
Sbjct: 798 SALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPLGQITLPVTFGTRENFRTENI 857
Query: 498 RYLVINSPSSYNVIIGRPSINLLDAFVSTKHLLMKYPLDSGRVGVVHGDQKIARECYHAS 557
+ V + ++Y+ I+GRP++ A +++MK P G + + D K A C S
Sbjct: 858 SFEVADFETAYHAILGRPALAKFMAVPHYTYMMMKMPGPRGVLSL-RSDIKQAVTCDKES 916
Query: 558 LRLKKHKEFEVQVIDLDPREDFPQERIEPIEEVKDVVIGPLPHQITKIGTSLSGLEEEIL 617
+ + +E RED E G +P TK TS SG E
Sbjct: 917 CDMAQTREMA------SAREDIRLAAATASE-------GEVP--ATK--TSKSGESEAKT 959
Query: 618 VQL---------LRENVDLFAWAPSDMPGIDIGVACHHLAVRTSVKPVVQRKRKMGEEKR 668
++ N D+FAW PSDMPGI V H L V+ KP+ QR R+ ++++
Sbjct: 960 KKIPLDPSNPTKTANNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRK 1019
Query: 669 KAVDEEVKKLQEAHFICEIKYPTWLANTVLVKKASGKWRMCVDYTDLNMACPKDPYPLPS 728
A+ EE+ KL A FI E+ +P WLAN VLV+K +G+WRMCVDYTDLN +CPKDP+ LP
Sbjct: 1020 DAIKEELTKLLAAGFIKEVHHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPR 1079
Query: 729 IDHLIDNASGYKTLSFMDAYSGYNQIKMDPLDAPKTAFMTNQKNYHYRVMSFGLRNAGAT 788
ID ++D+ +G + LSF+D YSGY+QI++ D KT+F+T Y Y M FGL+NAGAT
Sbjct: 1080 IDQVVDSTAGCELLSFLDCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMPFGLKNAGAT 1139
Query: 789 FQRSMDTIFSNQIGRNLEVYIDDLVVKTSEKQSHSVDLKEIFQQIRKFSMRLNPTKCTFG 848
+QR + FS QIGRN+E Y+DD+VVKT +K DL+E F IR F M+LNP KCTFG
Sbjct: 1140 YQRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFG 1199
Query: 849 VQAGKFLGFLLTKKGIEANPDKCQAIINMRNPCNIREVQQLTGRLAALSRFLSCAGDKAF 908
V +GK LGF+++ +GI+ANP+K AI+NM+ P ++VQ+LTG +AALSRF+S G++
Sbjct: 1200 VPSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGM 1259
Query: 909 AFFATIKKKEEFEWNQECDEAFGKIKQFLTTPPILHRPTKGAGLFLYLSVSENALSSVLV 968
FF +KK ++F+W E +AF K+ LT PP+L P L LY+S + +S+VLV
Sbjct: 1260 PFFKLLKKTDDFQWGPEAQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTVLV 1319
Query: 969 EESDER------EKPIYFVSRVLKGAELRYQKIEKLALAVIITARKLRPYFQSHKVVIRT 1022
E +E ++PIYFVS VL ++ RY +++KL ++IT RKL YFQ H V + T
Sbjct: 1320 VEREEEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVT 1379
Query: 1023 NYPVKQILGKLDLAGRMLSWSVELSEYDIQFAPRNNIKSQVLADFVVEFTSPIQEAVP-- 1080
++P+ IL + GR+ W++EL DI F PR +IKSQ LADFV E+T QE P
Sbjct: 1380 SFPLGDILHNREANGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTE-CQEDTPAE 1438
Query: 1081 --HVWLLSVDGSSNIKGSGAGIILEGPGDLLIEQSLKFDFKASNNQAEYEALIAGMLLAQ 1138
W + DGS + G+GAG++L P + L F AS+N AEYEAL+ G+ +A
Sbjct: 1439 KMEHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAI 1498
Query: 1139 EMGAKNLRARSDSQLMTNQISGEYQTKDQQLSKYLARVRKLAGDFQFFEAIYVPRESNSR 1198
+G K L R DSQL+ NQ+ E+ D + Y VRKL F E +V R +N
Sbjct: 1499 SLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEA 1558
Query: 1199 ADLLAKLASTKKPGNNRTVIQEVIS-----APSTDEKAVFELNQEPEGWMTPLLKFLTGS 1253
AD LA S ++ + ++ + + +T ++ W PL++FLT
Sbjct: 1559 ADRLANFGSKREVAPSDVFVEHLYTPTVPHKDTTQVAGTHDVAMVEADWREPLIRFLTSQ 1618
Query: 1254 FVAKNDEYAQLVRRRATKFVVIAGKLYKRGRASPLLRCLGEGETELVLLEVHEGVCGSHI 1313
+ ++ + A+ + RR+ +V+ +LYK+ + L RC+ E +L ++H G+CG+H
Sbjct: 1619 ELPQDKDEAERISRRSKLYVMHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNHA 1678
Query: 1314 GGRSLAAKLLRAGYYWPRMAHDCCEFVKKCDKCQRFSDKKNAPANELTSVFSPWPFHKWG 1373
R++ K R G++WP D + V+ C+ CQ F+ + + PA EL ++ WPF WG
Sbjct: 1679 AARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWG 1738
Query: 1374 VDIVGPFPQAPGQLKFLIVDVDYFTKWVEAEAVSKITAERVVKFYWKKIICHFGLPKYIV 1433
+D+VGPF +A G L V +D F+KW+EA+ V ITA+ F+ I+ FG+P I+
Sbjct: 1739 LDMVGPFKKAVGGYTHLFVAIDKFSKWIEAKPVVTITADNAPDFF-INIVHRFGVPNRII 1797
Query: 1434 TDNGTQFASSKVVNFCKQLGIETKFVSVIHPQANGQAESANKMIVNDIKKKLEDA----K 1489
TDNG QF + C+ GI+ + SV HP +NGQ E AN MI+ IK ++ D
Sbjct: 1798 TDNGRQFTGGVFKDCCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPYA 1857
Query: 1490 GLWAEQLHEVLWSYHTTPHSTTGETPFTMVYGADAMLPVEIDTPTWRREHFSEESNEVGI 1549
G W EQL VLWS TTP TG++PF +VYGA+AMLP E++ + R +F EE E
Sbjct: 1858 GKWVEQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFREERYEEDR 1917
Query: 1550 RCTMDMIDEVREAAHIREFAAKQRAARRYNSKVIPRSMKEGDLVLKQVVAPTRIGKLLPS 1609
+ ++EVREAA I+ Q R +N V R+ GDLVL+++ KL P
Sbjct: 1918 VDDLHRLEEVREAALIQSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTTRDRHKLSPL 1977
Query: 1610 WKGPYRVKEKLQHGAYKLEELSGNPV 1635
W+GP+ + + G+Y+L+ G V
Sbjct: 1978 WEGPFIISAVTRPGSYRLKREDGTLV 2003
>gb|AAP52499.1| putative gag-pol precursor [Oryza sativa (japonica cultivar-group)]
gi|37531820|ref|NP_920212.1| putative gag-pol precursor
[Oryza sativa (japonica cultivar-group)]
gi|22128689|gb|AAM92802.1| putative gag-pol precursor
[Oryza sativa (japonica cultivar-group)]
Length = 2017
Score = 952 bits (2461), Expect = 0.0
Identities = 566/1634 (34%), Positives = 860/1634 (51%), Gaps = 77/1634 (4%)
Query: 34 HHGDLDEPEPQPFSTEIWNAPVPDNFKPPQLPTFDGRSDPSEHVTVFNTRMSVYGVADSL 93
H DLD F+ ++ P FKP + +DG ++P +TV+ + G +
Sbjct: 415 HDDDLDGVAA--FTDDLRRVDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDNKA 472
Query: 94 KCKLLAGTLADAALRWYMSLPRFSIVGYQDLTKKFTQQFSGTRHRKVLSTSLFNVRQGPN 153
L LAD+A W LPR +I + +L F F GT R L+NV Q
Sbjct: 473 MANYLPVALADSARSWLHGLPRGTIGSWAELRDHFIANFQGTFERPGTQYDLYNVIQKSG 532
Query: 154 ESLREYLARFNDSTIKVSNPNQEVFVGAFQNGLRAGQFNESLAQKPADSMEEIIARAECY 213
ESLR+Y+ RF++ K+S+ +V + AF G+R + +KP +++ + +A Y
Sbjct: 533 ESLRDYIRRFSEQRNKISDITDDVIIAAFTKGIRHEELVGKFGRKPPRTVKLMFEKANEY 592
Query: 214 VKGEESNAEKRARDVKEKGSSGGERRNHYVPPNRDRGTFKKPYERNQPRYVPEHFTPLNT 273
K E++ K+ G S ++N P G+ + +P
Sbjct: 593 AKAEDAVTAS-----KQSGPSW--KQNKGTPATGGGGSNNHKDRKRKPA----------- 634
Query: 274 RPEKILKEVFESKIIPPPPFNRSKTMGQDKNAWCKYHLIEGHNTDDCVHLKREIEKLLLN 333
E+ + +R T + N+ C +H H DC K+ ++
Sbjct: 635 -------ELVATASHSSRQRSRVNTFDKIMNSQCPHHPNSNHVAKDCFVYKQFADQYTKT 687
Query: 334 GKLRGYAKEKHHSERREDKPN-----PEPKHTLHTISGGFAGGGESSNLRKKYVRQVMLL 388
+ + +E+ S +++D K H G A ES +K R++ +
Sbjct: 688 AR-KNSDEEQSTSRKKDDGDTLAGFQDHRKELNHIFGGPLAY--ESKRKQKLTEREINAV 744
Query: 389 GDSPSSPKKYPHVTFS---SEDFGRVLPHDDDPLVISVQLFNWEIKRVLIDTGSSADVLY 445
++ + S+ RV+ PLV+ + N +++R LID GS+ ++L+
Sbjct: 745 QPDTPQYLRWSEIAIKFDRSDHPDRVVHPGRYPLVLDPVVRNVKLRRTLIDGGSALNILF 804
Query: 446 FDAFSKMGLSEEQLQPFNGTLSG-FTGEKVHVRGYVTLKTIFGTGDQQTSIKVRYLVINS 504
M + +L+P N G G G +TL FGT + + + + V +
Sbjct: 805 AKTLDDMQIPRSELKPSNAPFHGVIPGLSATPLGQITLPVTFGTRENFRTENISFEVADF 864
Query: 505 PSSYNVIIGRPSINLLDAFVSTKHLLMKYPLDSGRVGVVHGDQKIARECYHASLRLKKHK 564
++Y+ I+GRP++ A +++MK P G + + D K A C S + + +
Sbjct: 865 ETAYHAILGRPALAKFMAVPHYTYMMMKMPGPRGVLSL-RSDIKQAVTCDKESCDMAQTR 923
Query: 565 EFEVQVIDLDPREDFPQERIEPIEEVKDVVIGPLPH-QITKIGTSLSGLEEEIL----VQ 619
E RED E G +P +I+K G S + ++ L
Sbjct: 924 EMA------SAREDIRLAAATASE-------GEVPATKISKSGESEAKTKKIPLDPSDPT 970
Query: 620 LLRENVDLFAWAPSDMPGIDIGVACHHLAVRTSVKPVVQRKRKMGEEKRKAVDEEVKKLQ 679
N D+FAW PSDMPGI V H L V+ KP+ QR R+ ++++ A+ EE+ KL
Sbjct: 971 KTANNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAIKEELTKLL 1030
Query: 680 EAHFICEIKYPTWLANTVLVKKASGKWRMCVDYTDLNMACPKDPYPLPSIDHLIDNASGY 739
A FI E+ +P WLAN VLV+K +G+WRMCVDYTDLN +CPKDP+ LP ID ++D+ +G
Sbjct: 1031 AAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPRIDQVVDSTAGC 1090
Query: 740 KTLSFMDAYSGYNQIKMDPLDAPKTAFMTNQKNYHYRVMSFGLRNAGATFQRSMDTIFSN 799
+ LSF+D YSGY+QI++ D KT+F+T Y Y M FGL+NAGAT+QR + FS
Sbjct: 1091 ELLSFLDCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMPFGLKNAGATYQRMIQRCFST 1150
Query: 800 QIGRNLEVYIDDLVVKTSEKQSHSVDLKEIFQQIRKFSMRLNPTKCTFGVQAGKFLGFLL 859
QIGRN+E Y+DD+VVKT +K +DL+E F IR F M+LNP KCTFGV +GK LGF++
Sbjct: 1151 QIGRNVEAYVDDVVVKTKQKDDLILDLEETFASIRAFRMKLNPEKCTFGVPSGKLLGFMV 1210
Query: 860 TKKGIEANPDKCQAIINMRNPCNIREVQQLTGRLAALSRFLSCAGDKAFAFFATIKKKEE 919
+ +GI+ANP+K AI+NM+ P ++VQ+LTG +AALSRF+S G++ FF +KK ++
Sbjct: 1211 SHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKLLKKTDD 1270
Query: 920 FEWNQECDEAFGKIKQFLTTPPILHRPTKGAGLFLYLSVSENALSSVLVEESDER----- 974
F+W E +AF K+ LT PP+L P L LY+S + +S+VLV E +E
Sbjct: 1271 FQWGPEAQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTVLVVEREEEGHVQK 1330
Query: 975 -EKPIYFVSRVLKGAELRYQKIEKLALAVIITARKLRPYFQSHKVVIRTNYPVKQILGKL 1033
++PIYFVS VL ++ RY +++KL ++IT RKL YFQ H V + T++P+ IL
Sbjct: 1331 VQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSFPLGDILHNR 1390
Query: 1034 DLAGRMLSWSVELSEYDIQFAPRNNIKSQVLADFVVEFTSPIQEAV---PHVWLLSVDGS 1090
+ GR+ W++EL D+ F PR +IKSQ LADFV E+T ++ W + DGS
Sbjct: 1391 EANGRIAKWALELMSLDLSFKPRISIKSQALADFVAEWTECQEDTTVKKMEHWTMHFDGS 1450
Query: 1091 SNIKGSGAGIILEGPGDLLIEQSLKFDFKASNNQAEYEALIAGMLLAQEMGAKNLRARSD 1150
+ G+GAG++L P + L F AS+N AEYEAL+ G+ +A +G K L R D
Sbjct: 1451 KRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIKRLIVRGD 1510
Query: 1151 SQLMTNQISGEYQTKDQQLSKYLARVRKLAGDFQFFEAIYVPRESNSRADLLAKLASTKK 1210
SQL+ NQ+ E+ D + Y VRKL F E +V R N AD LA S ++
Sbjct: 1511 SQLVVNQVMKEWSCLDDNMKAYRQEVRKLEDKFDGLELSHVLRHDNEAADRLANFGSKRE 1570
Query: 1211 PGNNRTVIQEVIS-----APSTDEKAVFELNQEPEGWMTPLLKFLTGSFVAKNDEYAQLV 1265
+ ++ + + +T + ++ W PL++FLT + ++ + A+ +
Sbjct: 1571 VAPSDVFVEHLYTPTVPHKDTTQAAGIHDVAMVETDWREPLIRFLTSQELPQDKDEAERI 1630
Query: 1266 RRRATKFVVIAGKLYKRGRASPLLRCLGEGETELVLLEVHEGVCGSHIGGRSLAAKLLRA 1325
RR+ +V+ +LYK+ + L RC+ E +L ++H G+CG+H R++ K R
Sbjct: 1631 SRRSKLYVMHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNHAAARTIVGKAYRQ 1690
Query: 1326 GYYWPRMAHDCCEFVKKCDKCQRFSDKKNAPANELTSVFSPWPFHKWGVDIVGPFPQAPG 1385
G++WP D + V+ C+ CQ F+ + + PA EL ++ WPF WG+D+VGPF +A G
Sbjct: 1691 GFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWGLDMVGPFKKAVG 1750
Query: 1386 QLKFLIVDVDYFTKWVEAEAVSKITAERVVKFYWKKIICHFGLPKYIVTDNGTQFASSKV 1445
L V +D F+KW+EA+ V ITA+ F+ I+ FG+P I+TDNGTQF
Sbjct: 1751 GYTHLFVAIDKFSKWIEAKPVVTITADNARDFF-INIVHRFGVPNRIITDNGTQFTGGVF 1809
Query: 1446 VNFCKQLGIETKFVSVIHPQANGQAESANKMIVNDIKKKLEDA----KGLWAEQLHEVLW 1501
+FC+ GI+ + SV HP +NGQ E AN MI+ IK ++ D G W +QL VLW
Sbjct: 1810 KDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPYAGKWVQQLPSVLW 1869
Query: 1502 SYHTTPHSTTGETPFTMVYGADAMLPVEIDTPTWRREHFSEESNEVGIRCTMDMIDEVRE 1561
S TTP TG++PF +VYGA+AMLP E++ + R +F EE E + ++EVRE
Sbjct: 1870 SLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFREERYEEDRVDDLHRLEEVRE 1929
Query: 1562 AAHIREFAAKQRAARRYNSKVIPRSMKEGDLVLKQVVAPTRIGKLLPSWKGPYRVKEKLQ 1621
AA IR Q R +N V R+ GDLVL+++ KL P W+GP+ + E +
Sbjct: 1930 AALIRSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTTRDRHKLSPLWEGPFIISEVTR 1989
Query: 1622 HGAYKLEELSGNPV 1635
G+Y+L+ G V
Sbjct: 1990 PGSYRLKREDGTLV 2003
>ref|XP_475035.1| OSJNBa0042F21.5 [Oryza sativa (japonica cultivar-group)]
gi|38347303|emb|CAE02298.2| OSJNBa0042F21.5 [Oryza sativa
(japonica cultivar-group)]
Length = 1950
Score = 952 bits (2461), Expect = 0.0
Identities = 572/1641 (34%), Positives = 858/1641 (51%), Gaps = 87/1641 (5%)
Query: 32 DSHHGDLDEPEPQPFSTEIWNAPVPDNFKPPQLPTFDGRSDPSEHVTVFNTRMSVYGVAD 91
D H DLD F+ ++ P FKP + +DG ++P +TV+ + G +
Sbjct: 346 DRHDDDLDGVAA--FTDDLRRVDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDN 403
Query: 92 SLKCKLLAGTLADAALRWYMSLPRFSIVGYQDLTKKFTQQFSGTRHRKVLSTSLFNVRQG 151
L LAD+A W LPR +I + +L F F GT R L+NV Q
Sbjct: 404 KAMANYLPVALADSARSWLHGLPRGTIGSWAELRDHFIANFQGTFERPGTQYDLYNVIQK 463
Query: 152 PNESLREYLARFNDSTIKVSNPNQEVFVGAFQNGLRAGQFNESLAQKPADSMEEIIARAE 211
ESLR+Y+ RF++ K+S+ +V + AF G+R + +KP +++ + +A
Sbjct: 464 SGESLRDYIRRFSEQRNKISDITDDVIIAAFTKGIRHEELVGKFGRKPPRTVKLMFEKAN 523
Query: 212 CYVKGEESNAEKRARDVKEKGSSGGERRNHYVPPNRDRGTFKKPYERNQPRYVPEHFTPL 271
Y K E++ K+ G S ++N P G+ + +P
Sbjct: 524 EYTKAEDAVTAS-----KQSGPSW--KQNKGTPATGGGGSNNHKDRKRKPA--------- 567
Query: 272 NTRPEKILKEVFESKIIPPPPFNRSKTMGQDKNAWCKYHLIEGHNTDDCVHLKREIEKLL 331
E+ + +R T + N+ C +H H DC K+ ++
Sbjct: 568 ---------ELVATASHSSRQRSRVNTFDKIMNSQCPHHPNSNHVAKDCFVYKQFADQYT 618
Query: 332 LNGKLRGYAKEKHHSERREDKPNP-----EPKHTLHTISGGFAGGGESSNLRKKYVRQVM 386
+ + +E+ S +++D P K H G A ES +K R++
Sbjct: 619 KTAR-KNSDEEQSTSRKKDDGDTPAGFQDHRKELNHIFGGPLAY--ESKRKQKLTEREIN 675
Query: 387 LLGDSPSSPKKYPHVTFS---SEDFGRVLPHDDDPLVISVQLFNWEIKRVLIDTGSSADV 443
+ ++ + S+ RV+ PLV+ + N +++R LID GS+ ++
Sbjct: 676 AVQPDTPQYLRWSEIAIKFDRSDHPDRVVHPGRYPLVLDPVVRNVKLRRTLIDGGSALNI 735
Query: 444 LYFDAFSKMGLSEEQLQPFNGTLSG-FTGEKVHVRGYVTLKTIFGTGDQQTSIKVRYLVI 502
L+ M + +L+P N G G G +TL FGT + + + + V
Sbjct: 736 LFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPLGQITLPVTFGTRENFRTENISFEVA 795
Query: 503 NSPSSYNVIIGRPSINLLDAFVSTKHLLMKYPLDSGRVGVVHGDQKIARECYHASLRLKK 562
+ ++Y+ I+GRP++ A +++MK P G + + D K A C S + +
Sbjct: 796 DFETAYHAILGRPALAKFMAVPHYTYMMMKMPGPRGVLSL-RSDIKQAVTCDKESCDMAQ 854
Query: 563 HKEFEVQVIDLDPREDFPQERIEPIEEVKDVVIGPLPHQITKIGTSLSGLEEEILVQL-- 620
+E RED E G +P TK TS SG E ++
Sbjct: 855 TREMA------SAREDIRLAAATASE-------GEIP--ATK--TSKSGESEAKTKKIPL 897
Query: 621 -------LRENVDLFAWAPSDMPGIDIGVACHHLAVRTSVKPVVQRKRKMGEEKRKAVDE 673
N D+FAW PSDMPGI V H L V+ KP+ QR R+ ++++ A+ E
Sbjct: 898 DPSDPTKTANNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAIKE 957
Query: 674 EVKKLQEAHFICEIKYPTWLANTVLVKKASGKWRMCVDYTDLNMACPKDPYPLPSIDHLI 733
E+ KL A FI E+ +P WLAN VLV+K +G+WRMCVDYTDLN +CPKDP+ LP ID ++
Sbjct: 958 ELTKLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPRIDQVV 1017
Query: 734 DNASGYKTLSFMDAYSGYNQIKMDPLDAPKTAFMTNQKNYHYRVMSFGLRNAGATFQRSM 793
D+ +G + LSF+D YSGY+QI++ D KT+F+T Y Y M FGL+NAGAT+QR +
Sbjct: 1018 DSTAGCELLSFLDCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMPFGLKNAGATYQRMI 1077
Query: 794 DTIFSNQIGRNLEVYIDDLVVKTSEKQSHSVDLKEIFQQIRKFSMRLNPTKCTFGVQAGK 853
FS QIGRN+E Y+DD+VVKT +K DL+E F IR F M+LNP KCTFGV +GK
Sbjct: 1078 QRCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGVPSGK 1137
Query: 854 FLGFLLTKKGIEANPDKCQAIINMRNPCNIREVQQLTGRLAALSRFLSCAGDKAFAFFAT 913
LGF+++ +GI+ANP+K AI+NM+ P ++VQ+LTG +AALSRF+S G++ FF
Sbjct: 1138 LLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKL 1197
Query: 914 IKKKEEFEWNQECDEAFGKIKQFLTTPPILHRPTKGAGLFLYLSVSENALSSVLVEESDE 973
+KK + F+W E +AF K+ LT PPIL P L LY+S + +S+VLV E +E
Sbjct: 1198 LKKTDNFQWGPEAQKAFEDFKKLLTEPPILASPHPQEPLLLYVSATSQVVSTVLVVEREE 1257
Query: 974 R------EKPIYFVSRVLKGAELRYQKIEKLALAVIITARKLRPYFQSHKVVIRTNYPVK 1027
++PIYFVS VL ++ RY +++KL ++IT RKL YFQ H V + T++P+
Sbjct: 1258 EGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSFPLG 1317
Query: 1028 QILGKLDLAGRMLSWSVELSEYDIQFAPRNNIKSQVLADFVVEFTSPIQEAVP----HVW 1083
IL + GR+ W++EL DI F PR +IKSQ LADFV E+T QE P W
Sbjct: 1318 DILHNREANGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTE-CQEDTPAENMEHW 1376
Query: 1084 LLSVDGSSNIKGSGAGIILEGPGDLLIEQSLKFDFKASNNQAEYEALIAGMLLAQEMGAK 1143
+ DGS + G+GAG++L P + L F AS+N AEYEAL+ G+ +A +G K
Sbjct: 1377 TMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIK 1436
Query: 1144 NLRARSDSQLMTNQISGEYQTKDQQLSKYLARVRKLAGDFQFFEAIYVPRESNSRADLLA 1203
L R DSQL+ NQ+ E+ D + Y VRKL F E +V R +N AD LA
Sbjct: 1437 RLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLA 1496
Query: 1204 KLASTKKPGNNRTVIQEVIS-----APSTDEKAVFELNQEPEGWMTPLLKFLTGSFVAKN 1258
S ++ + ++ + + +T + ++ W P ++FLT + ++
Sbjct: 1497 NFGSKREMAPSDVFVEHLYTPTVPHKDTTQDADTHDVALVEADWREPFIRFLTSQELPQD 1556
Query: 1259 DEYAQLVRRRATKFVVIAGKLYKRGRASPLLRCLGEGETELVLLEVHEGVCGSHIGGRSL 1318
+ A+ + RR+ + + +LYK+ + L RC+ E +L ++H G+CG+H R++
Sbjct: 1557 KDEAERISRRSKLYAMHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNHAAARTI 1616
Query: 1319 AAKLLRAGYYWPRMAHDCCEFVKKCDKCQRFSDKKNAPANELTSVFSPWPFHKWGVDIVG 1378
K R G++WP D + V+ C+ CQ F+ + + PA EL ++ WPF WG+D+VG
Sbjct: 1617 VGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWGLDMVG 1676
Query: 1379 PFPQAPGQLKFLIVDVDYFTKWVEAEAVSKITAERVVKFYWKKIICHFGLPKYIVTDNGT 1438
PF +A G L V +D F+KW+EA+ V ITA+ F+ I+ FG+P I+TDNGT
Sbjct: 1677 PFKKAVGGYTHLFVAIDKFSKWIEAKPVVTITADNARDFF-INIVHRFGVPNRIITDNGT 1735
Query: 1439 QFASSKVVNFCKQLGIETKFVSVIHPQANGQAESANKMIVNDIKKKLEDA----KGLWAE 1494
QF +FC+ GI+ + SV HP +NGQ E AN MI+ IK ++ D G W +
Sbjct: 1736 QFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPYAGKWVQ 1795
Query: 1495 QLHEVLWSYHTTPHSTTGETPFTMVYGADAMLPVEIDTPTWRREHFSEESNEVGIRCTMD 1554
QL VLWS TTP TG++PF +VYGA+AMLP E++ + R +F EE E +
Sbjct: 1796 QLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFREERYEEDRVDDLH 1855
Query: 1555 MIDEVREAAHIREFAAKQRAARRYNSKVIPRSMKEGDLVLKQVVAPTRIGKLLPSWKGPY 1614
++E REAA IR Q R +N V R+ GDLVL+++ KL P W+GP+
Sbjct: 1856 RLEEAREAALIRSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTTRDRHKLSPLWEGPF 1915
Query: 1615 RVKEKLQHGAYKLEELSGNPV 1635
+ E + G+Y+L+ G V
Sbjct: 1916 IISEVTRPGSYRLKREDGTLV 1936
>ref|NP_918169.1| putative gag-pol polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 2012
Score = 952 bits (2460), Expect = 0.0
Identities = 572/1641 (34%), Positives = 857/1641 (51%), Gaps = 87/1641 (5%)
Query: 32 DSHHGDLDEPEPQPFSTEIWNAPVPDNFKPPQLPTFDGRSDPSEHVTVFNTRMSVYGVAD 91
D H DLD F+ ++ P FKP + +DG ++P +TV+ + G +
Sbjct: 408 DRHDDDLDGVAA--FTDDLRRVDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDN 465
Query: 92 SLKCKLLAGTLADAALRWYMSLPRFSIVGYQDLTKKFTQQFSGTRHRKVLSTSLFNVRQG 151
L LAD+A W LPR +I + +L F F GT R L+NV Q
Sbjct: 466 KAMANYLPVALADSARSWLHGLPRGTIGSWAELRDHFIANFQGTFERPGTQYDLYNVIQK 525
Query: 152 PNESLREYLARFNDSTIKVSNPNQEVFVGAFQNGLRAGQFNESLAQKPADSMEEIIARAE 211
ESLR+Y+ RF++ K+S+ +V + AF G+R + +KP +++ + +A
Sbjct: 526 SGESLRDYIRRFSEQRNKISDITDDVIIAAFTKGIRHEELVGKFGRKPPRTVKLMFEKAN 585
Query: 212 CYVKGEESNAEKRARDVKEKGSSGGERRNHYVPPNRDRGTFKKPYERNQPRYVPEHFTPL 271
Y K E++ K+ G S ++N P G+ + +P
Sbjct: 586 EYAKAEDAVTAS-----KQSGPSW--KQNKGTPATGGGGSNNHKDRKRKPA--------- 629
Query: 272 NTRPEKILKEVFESKIIPPPPFNRSKTMGQDKNAWCKYHLIEGHNTDDCVHLKREIEKLL 331
E+ + +R T + N+ C +H H DC K+ ++
Sbjct: 630 ---------ELVATASHSSRQRSRVNTFDKIMNSQCPHHPNSNHVAKDCFVYKQFADQYT 680
Query: 332 LNGKLRGYAKEKHHSERREDKPNP-----EPKHTLHTISGGFAGGGESSNLRKKYVRQVM 386
+ + +E+ S +++D P K H G A ES +K R++
Sbjct: 681 KTAR-KNTDEEQSTSRKKDDGDTPAGFQDHRKELNHIFGGPLAY--ESKRKQKLTEREIN 737
Query: 387 LLGDSPSSPKKYPHVTFS---SEDFGRVLPHDDDPLVISVQLFNWEIKRVLIDTGSSADV 443
+ ++ + S+ RV+ PLV+ + N +++R LID GS+ ++
Sbjct: 738 AVQPDTPQYLRWSEIAIKFDRSDHPDRVVHPGRYPLVLDPVVRNVKLRRTLIDGGSALNI 797
Query: 444 LYFDAFSKMGLSEEQLQPFNGTLSG-FTGEKVHVRGYVTLKTIFGTGDQQTSIKVRYLVI 502
L+ M + +L+P N G G G +TL FGT + + + + V
Sbjct: 798 LFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPLGQITLPVTFGTRENFRTENISFEVA 857
Query: 503 NSPSSYNVIIGRPSINLLDAFVSTKHLLMKYPLDSGRVGVVHGDQKIARECYHASLRLKK 562
+ ++Y+ I+GRP++ A +++MK P G + + D K A C S + +
Sbjct: 858 DFETAYHAILGRPALAKFMAVPHYTYMMMKMPGPRGVLSL-RSDVKQAVTCDKESCDMAQ 916
Query: 563 HKEFEVQVIDLDPREDFPQERIEPIEEVKDVVIGPLPHQITKIGTSLSGLEEEILVQL-- 620
+E RED E G +P TK TS SG E ++
Sbjct: 917 TREMA------SAREDIRLAAATASE-------GEIP--ATK--TSKSGESEAKTKKIPL 959
Query: 621 -------LRENVDLFAWAPSDMPGIDIGVACHHLAVRTSVKPVVQRKRKMGEEKRKAVDE 673
N D+FAW PSDMPGI V H L V+ KP+ QR R+ ++++ A+ E
Sbjct: 960 DPSDPTKTANNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAIKE 1019
Query: 674 EVKKLQEAHFICEIKYPTWLANTVLVKKASGKWRMCVDYTDLNMACPKDPYPLPSIDHLI 733
E+ KL A FI E+ +P WLAN VLV+K +G+W MCVDYTDLN +CPKDP+ LP ID ++
Sbjct: 1020 ELTKLLAAGFIKEVLHPDWLANPVLVRKKTGQWLMCVDYTDLNKSCPKDPFGLPRIDQVV 1079
Query: 734 DNASGYKTLSFMDAYSGYNQIKMDPLDAPKTAFMTNQKNYHYRVMSFGLRNAGATFQRSM 793
D+ +G + LSF+D YSGY+QI++ D KT+F+T Y Y M FGL+NAGAT+QR +
Sbjct: 1080 DSTAGCELLSFLDCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMPFGLKNAGATYQRMI 1139
Query: 794 DTIFSNQIGRNLEVYIDDLVVKTSEKQSHSVDLKEIFQQIRKFSMRLNPTKCTFGVQAGK 853
FS QIGRN+E Y+DD+VVKT +K DL+E F IR F M+LNP KCTFGV +GK
Sbjct: 1140 QRCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGVPSGK 1199
Query: 854 FLGFLLTKKGIEANPDKCQAIINMRNPCNIREVQQLTGRLAALSRFLSCAGDKAFAFFAT 913
LGF+++ +GI+ANP+K AI+NM++P ++VQ+LTG +AALSRF+S G++ FF
Sbjct: 1200 LLGFMVSHRGIQANPEKVTAILNMKSPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKL 1259
Query: 914 IKKKEEFEWNQECDEAFGKIKQFLTTPPILHRPTKGAGLFLYLSVSENALSSVLVEESDE 973
+KK + F W E +AF K+ LT PP+L P L LY+S + +S+VLV E +E
Sbjct: 1260 LKKTDSFRWGPEAQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTVLVVEREE 1319
Query: 974 R------EKPIYFVSRVLKGAELRYQKIEKLALAVIITARKLRPYFQSHKVVIRTNYPVK 1027
++PIYFVS VL ++ RY +++KL ++IT RKL YFQ H V + T++P+
Sbjct: 1320 EGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSFPLG 1379
Query: 1028 QILGKLDLAGRMLSWSVELSEYDIQFAPRNNIKSQVLADFVVEFTSPIQEAVP----HVW 1083
IL + GR+ W++EL DI F PR +IKSQ LADFV E+T QE P W
Sbjct: 1380 DILHNREANGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTE-CQEDTPAENMEHW 1438
Query: 1084 LLSVDGSSNIKGSGAGIILEGPGDLLIEQSLKFDFKASNNQAEYEALIAGMLLAQEMGAK 1143
+ DGS + G+GAG++L P + L F AS+N AEYEAL+ G+ +A +G K
Sbjct: 1439 TMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIK 1498
Query: 1144 NLRARSDSQLMTNQISGEYQTKDQQLSKYLARVRKLAGDFQFFEAIYVPRESNSRADLLA 1203
L R DSQL+ NQ+ E+ D + Y VRKL F E +V R +N AD LA
Sbjct: 1499 RLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLA 1558
Query: 1204 KLASTKKPGNNRTVIQEVIS-----APSTDEKAVFELNQEPEGWMTPLLKFLTGSFVAKN 1258
S ++ + ++ + S +T ++ W PL++FLT + ++
Sbjct: 1559 NFGSKREAAPSDVFVEHLYSPTVPHKDATQAAGAHDVAMVEADWREPLIRFLTSQELPQD 1618
Query: 1259 DEYAQLVRRRATKFVVIAGKLYKRGRASPLLRCLGEGETELVLLEVHEGVCGSHIGGRSL 1318
+ A+ + RR+ +V+ +LYK+ + L RC+ E +L ++H G+CG+H R++
Sbjct: 1619 KDEAERISRRSKLYVLHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNHAAARTI 1678
Query: 1319 AAKLLRAGYYWPRMAHDCCEFVKKCDKCQRFSDKKNAPANELTSVFSPWPFHKWGVDIVG 1378
K R G++WP D + V+ C+ CQ F+ + + PA EL ++ WPF WG+D+VG
Sbjct: 1679 VGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWGLDMVG 1738
Query: 1379 PFPQAPGQLKFLIVDVDYFTKWVEAEAVSKITAERVVKFYWKKIICHFGLPKYIVTDNGT 1438
PF +A G L V +D F+KW+EA+ V ITA+ F+ I+ FG+P I+TDNGT
Sbjct: 1739 PFKKAVGGYTHLFVAIDKFSKWIEAKPVVTITADNARDFF-INIVHRFGVPNRIITDNGT 1797
Query: 1439 QFASSKVVNFCKQLGIETKFVSVIHPQANGQAESANKMIVNDIKKKLEDA----KGLWAE 1494
QF +FC+ GI+ + SV HP +NGQ E AN MI+ IK ++ D G W +
Sbjct: 1798 QFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPYAGKWVQ 1857
Query: 1495 QLHEVLWSYHTTPHSTTGETPFTMVYGADAMLPVEIDTPTWRREHFSEESNEVGIRCTMD 1554
QL VLWS TT TG++PF +VYGA+AMLP E++ + R +F EE E +
Sbjct: 1858 QLPSVLWSLRTTSSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFREERYEEDRVDDLH 1917
Query: 1555 MIDEVREAAHIREFAAKQRAARRYNSKVIPRSMKEGDLVLKQVVAPTRIGKLLPSWKGPY 1614
++E REAA IR Q R +N V R+ GDLVL+++ KL P W+GP+
Sbjct: 1918 RLEEAREAALIRSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTTRDRHKLSPLWEGPF 1977
Query: 1615 RVKEKLQHGAYKLEELSGNPV 1635
+ E + G+Y+L+ G V
Sbjct: 1978 IISEVTRPGSYRLKREDGTLV 1998
>gb|AAP54912.1| gag-pol precursor [Oryza sativa (japonica cultivar-group)]
gi|37536646|ref|NP_922625.1| gag-pol precursor [Oryza
sativa (japonica cultivar-group)]
gi|13876521|gb|AAK43497.1| gag-pol precursor [Oryza
sativa (japonica cultivar-group)]
Length = 2017
Score = 944 bits (2440), Expect = 0.0
Identities = 564/1637 (34%), Positives = 861/1637 (52%), Gaps = 79/1637 (4%)
Query: 32 DSHHGDLDEPEPQPFSTEIWNAPVPDNFKPPQLPTFDGRSDPSEHVTVFNTRMSVYGVAD 91
D H DLD F+ ++ P FKP + +DG ++P +T++ + G
Sbjct: 413 DRHDDDLDGVAA--FTDDLRRVDWPAGFKPTGIEKYDGTTNPESWLTIYGLAIRAAGGDS 470
Query: 92 SLKCKLLAGTLADAALRWYMSLPRFSIVGYQDLTKKFTQQFSGTRHRKVLSTSLFNVRQG 151
L LAD+A W LPR +I + +L F F GT L+NV Q
Sbjct: 471 KAMANYLPVALADSARSWLHGLPRGTIGSWAELRDHFIANFQGTFEHPGTQYDLYNVIQK 530
Query: 152 PNESLREYLARFNDSTIKVSNPNQEVFVGAFQNGLRAGQFNESLAQKPADSMEEIIARAE 211
ESLR+Y+ RF++ K+S+ +V + AF G+R + +KP +++ + +A
Sbjct: 531 SGESLRDYIRRFSEQRNKISDITDDVIIAAFTKGIRHEELVGKFGRKPPKTVKLMFEKAN 590
Query: 212 CYVKGEESNAEKRARDVKEKGSSGGERRNHYVPPNRDRGTFKKPYERNQPRYVPEHFTPL 271
Y K E++ V SG + + P + G +R +
Sbjct: 591 EYAKAEDA--------VTASKQSGPSWKPNKGTPAKGGGGSNNHKDRKR----------- 631
Query: 272 NTRPEKILKEVFESKIIPPPPFNRSKTMGQDKNAWCKYHLIEGHNTDDCVHLKREIEKLL 331
+PE+++ S +R T + N+ C +H H DC ++ E+
Sbjct: 632 --KPEELVATATHSS----RQHSRVNTFDKIMNSQCPHHPNSNHVAKDCFVYRQFAEQYT 685
Query: 332 LNGKLRGYAKEKHHSERREDKPNP-----EPKHTLHTISGGFAGGGESSNLRKKYVRQVM 386
+ + +E+ S +++D P K H G A ES +K R++
Sbjct: 686 KTTR-KNSDEEQSTSRKKDDGDTPAGFQDHRKELNHIFGGPLAY--ESKRKQKLTEREIN 742
Query: 387 LLGDSPSSPKKYPHVTFS---SEDFGRVLPHDDDPLVISVQLFNWEIKRVLIDTGSSADV 443
+ ++ + S+ RV+ PLV+ + N +++R LID GS+ ++
Sbjct: 743 AVQPDTPQYLRWSEIAIKFDRSDHPDRVVHPGRYPLVLDPVVRNVKLRRTLIDGGSALNI 802
Query: 444 LYFDAFSKMGLSEEQLQPFNGTLSG-FTGEKVHVRGYVTLKTIFGTGDQQTSIKVRYLVI 502
L+ M + +L+P N G G G +TL FGT + + + + V
Sbjct: 803 LFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPLGQITLPVTFGTRENFRTENISFEVD 862
Query: 503 NSPSSYNVIIGRPSINLLDAFVSTKHLLMKYPLDSGRVGVVHGDQKIARECYHASLRLKK 562
+ ++Y+ I+GRP++ A +++MK P G + + D K A C S + +
Sbjct: 863 DFETAYHAILGRPALAKFMAVPHYTYMMMKMPGPRGVLSL-RSDIKQAVTCDKESCDMAQ 921
Query: 563 HKEFEVQVIDLDPREDFPQERIEPIEEVKDVVIGPLPH-QITKIGTSLSGLEEEIL---- 617
+E RED E G +P +I+K G S + ++ L
Sbjct: 922 TREMA------SAREDIRLAAATASE-------GEVPATKISKSGESEAKTKKIPLDPSD 968
Query: 618 VQLLRENVDLFAWAPSDMPGIDIGVACHHLAVRTSVKPVVQRKRKMGEEKRKAVDEEVKK 677
N D+FAW PSDMPGI V H L V+ KP+ QR R+ ++++ A+ EE+ K
Sbjct: 969 STKTANNKDIFAWKPSDMPGIPREVIEHSLYVKEDAKPIKQRLRRFAQDRKDAIKEELTK 1028
Query: 678 LQEAHFICEIKYPTWLANTVLVKKASGKWRMCVDYTDLNMACPKDPYPLPSIDHLIDNAS 737
L A FI E+ +P WLAN VLV+K +G+WRMCVDYTDLN++CPKDP+ LP ID ++D+ +
Sbjct: 1029 LLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNISCPKDPFGLPRIDQVVDSTA 1088
Query: 738 GYKTLSFMDAYSGYNQIKMDPLDAPKTAFMTNQKNYHYRVMSFGLRNAGATFQRSMDTIF 797
G + LSF+D Y GY+QI++ D KT+F+T Y Y M FGL+NAGAT+QR + F
Sbjct: 1089 GCELLSFLDCYLGYHQIRLKESDCLKTSFITPFGAYCYVTMPFGLKNAGATYQRMIQRCF 1148
Query: 798 SNQIGRNLEVYIDDLVVKTSEKQSHSVDLKEIFQQIRKFSMRLNPTKCTFGVQAGKFLGF 857
S QIGRN+E Y+DD+VVKT +K DL+E F IR F M+LNP KCTFGV +GK LGF
Sbjct: 1149 STQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGVPSGKLLGF 1208
Query: 858 LLTKKGIEANPDKCQAIINMRNPCNIREVQQLTGRLAALSRFLSCAGDKAFAFFATIKKK 917
+++ +GI+ANP+K AI+NM+ P ++VQ+LTG +AALSRF+S G++ FF +KK
Sbjct: 1209 MVSHRGIQANPEKITAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKLLKKT 1268
Query: 918 EEFEWNQECDEAFGKIKQFLTTPPILHRPTKGAGLFLYLSVSENALSSVLVEESDER--- 974
++F+W + +AF K+ LT PP+L P L LY++ + +S+VLV E +E
Sbjct: 1269 DDFQWGPDAQKAFEDFKKLLTEPPVLASPHPQEPLLLYIAAASQVVSTVLVVEREEEGHV 1328
Query: 975 ---EKPIYFVSRVLKGAELRYQKIEKLALAVIITARKLRPYFQSHKVVIRTNYPVKQILG 1031
++PIYFVS VL ++ RY +++KL ++IT RKL YFQSH V + T++ + IL
Sbjct: 1329 QKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQSHSVTVVTSFSLGDILH 1388
Query: 1032 KLDLAGRMLSWSVELSEYDIQFAPRNNIKSQVLADFVVEFTSPIQEAVP----HVWLLSV 1087
+ GR+ W++EL DI F PR +IKSQ LADFV E+T QE P W +
Sbjct: 1389 NREANGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTE-CQEDTPAEKMEHWTMHF 1447
Query: 1088 DGSSNIKGSGAGIILEGPGDLLIEQSLKFDFKASNNQAEYEALIAGMLLAQEMGAKNLRA 1147
DGS + G+GAG++L P + L F AS+N AEYEAL+ G+ +A +G K L
Sbjct: 1448 DGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIKRLIV 1507
Query: 1148 RSDSQLMTNQISGEYQTKDQQLSKYLARVRKLAGDFQFFEAIYVPRESNSRADLLAKLAS 1207
R DSQL+ NQ+ E+ D + Y VRKL F E +V R +N AD LA S
Sbjct: 1508 RGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLANFGS 1567
Query: 1208 TKKPGNNRTVIQEVIS-----APSTDEKAVFELNQEPEGWMTPLLKFLTGSFVAKNDEYA 1262
++ + ++ + + +T ++ W PL++FLT + ++ + A
Sbjct: 1568 KREVAPSDVFVEHLYTPTVPHKDTTQIAGTHDVALVEADWREPLIRFLTSQELPQDKDEA 1627
Query: 1263 QLVRRRATKFVVIAGKLYKRGRASPLLRCLGEGETELVLLEVHEGVCGSHIGGRSLAAKL 1322
+ + RR+ +V+ +LYK+ + L RC+ E +L ++H G+CG+H R++ K
Sbjct: 1628 ERISRRSKLYVMHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNHAAARTIVGKA 1687
Query: 1323 LRAGYYWPRMAHDCCEFVKKCDKCQRFSDKKNAPANELTSVFSPWPFHKWGVDIVGPFPQ 1382
R G++WP D + V+ C+ CQ F+ + + PA EL ++ WPF WG+D+VGPF +
Sbjct: 1688 YRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWGLDMVGPFKK 1747
Query: 1383 APGQLKFLIVDVDYFTKWVEAEAVSKITAERVVKFYWKKIICHFGLPKYIVTDNGTQFAS 1442
A G L V +D F+KW+EA+ V ITA+ F+ I+ FG+P I+TDNG QF
Sbjct: 1748 AVGGYTHLFVAIDKFSKWIEAKPVVTITADNARDFF-INIVHRFGVPNRIITDNGRQFTG 1806
Query: 1443 SKVVNFCKQLGIETKFVSVIHPQANGQAESANKMIVNDIKKKLEDA----KGLWAEQLHE 1498
+FC+ GI+ + SV HP +NGQ E AN MI+ IK ++ D G W +QL
Sbjct: 1807 GVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPYAGKWVKQLPS 1866
Query: 1499 VLWSYHTTPHSTTGETPFTMVYGADAMLPVEIDTPTWRREHFSEESNEVGIRCTMDMIDE 1558
VLWS TTP TG++PF +VYGA+AMLP E++ + R ++ EE E + ++E
Sbjct: 1867 VLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNYREERYEEDRVDDLHRLEE 1926
Query: 1559 VREAAHIREFAAKQRAARRYNSKVIPRSMKEGDLVLKQVVAPTRIGKLLPSWKGPYRVKE 1618
VREAA I+ Q R +N V R+ GDLVL+++ KL P W+GP+ + +
Sbjct: 1927 VREAALIQSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTTRDRHKLSPLWEGPFIISD 1986
Query: 1619 KLQHGAYKLEELSGNPV 1635
+ G+Y+L+ G V
Sbjct: 1987 VTRPGSYRLKREDGTLV 2003
>ref|XP_474845.1| OSJNBa0035O13.3 [Oryza sativa (japonica cultivar-group)]
gi|21741410|emb|CAD40114.1| OSJNBa0035O13.3 [Oryza sativa
(japonica cultivar-group)]
Length = 2008
Score = 932 bits (2409), Expect = 0.0
Identities = 563/1638 (34%), Positives = 846/1638 (51%), Gaps = 90/1638 (5%)
Query: 34 HHGDLDEPEPQPFSTEIWNAPVPDNFKPPQLPTFDGRSDPSEHVTVFNTRMSVYGVADSL 93
H DLDE F+ ++ P FKP + +DG ++P +TV+ + G +
Sbjct: 411 HDDDLDEVAA--FTDDLRRVDWPAGFKPTGIEKYDGTTNPGSWLTVYGLAIRAAGGDNKA 468
Query: 94 KCKLLAGTLADAALRWYMSLPRFSIVGYQDLTKKFTQQFSGTRHRKVLSTSLFNVRQGPN 153
L LAD+A W LPR +I + +L F F GT R L+NV Q
Sbjct: 469 MANYLPVALADSARSWLHGLPRGTIGSWAELRDHFIANFQGTFERPGTQYDLYNVIQKSG 528
Query: 154 ESLREYLARFNDSTIKVSNPNQEVFVGAFQNGLRAGQFNESLAQKPADSMEEIIARAECY 213
ESLR+Y+ RF++ K+S+ +V + AF G+R + +KP +++ + +A Y
Sbjct: 529 ESLRDYIRRFSEQRNKISDITDDVIIAAFTKGIRHEELVGKFGRKPPRTVKLMFEKANEY 588
Query: 214 VKGEESNAEKRARDVKEKGSSGGERRNHYVPPNRDRGTFKKPYERNQPRYVPEHFTPLNT 273
K E++ K+ G S ++N P G+ + +P
Sbjct: 589 AKAEDAVTAS-----KQSGPSW--KQNKGTPATGGGGSNNHKDRKRKPA----------- 630
Query: 274 RPEKILKEVFESKIIPPPPFNRSKTMGQDKNAWCKYHLIEGHNTDDCVHLKREIEKLLLN 333
E+ + +R T + N+ C +H H DC K+ ++
Sbjct: 631 -------ELVATASHSSRQRSRVNTFDKIMNSQCPHHPNSNHVAKDCFVYKQFADQYTKT 683
Query: 334 GKLRGYAKEKHHSERREDKPNP-----EPKHTLHTISGGFAGGGESSNLRKKYVRQVMLL 388
+ + +E+ S +++D P K H G A ES +K R++ +
Sbjct: 684 AR-KNSDEEQSTSRKKDDGDAPAGFQDHRKELNHIFGGPLAY--ESKRKQKLTEREINAV 740
Query: 389 GDSPSSPKKYPHVTFS---SEDFGRVLPHDDDPLVISVQLFNWEIKRVLIDTGSSADVLY 445
++ + S+ RV+ PLV+ + N +++R LID GS+ ++L+
Sbjct: 741 QPDTPQYLRWSEIAIKFDRSDHPDRVVHPGRYPLVLDPVVRNVKLRRTLIDGGSALNILF 800
Query: 446 FDAFSKMGLSEEQLQPFNGTLSG-FTGEKVHVRGYVTLKTIFGTGDQQTSIKVRYLVINS 504
M + +L+P N G G G +TL FGT + + + + V +
Sbjct: 801 AKTLDDMQIPRSELKPSNAPFHGVIPGLSATPLGQITLPVTFGTRENFRTENISFEVADF 860
Query: 505 PSSYNVIIGRPSINLLDAFVSTKHLLMKYPLDSGRVGVVHGDQKIARECYHASLRLKKHK 564
++Y+ I+GRP++ A +++MK P G + + D K A C S +
Sbjct: 861 ETAYHAILGRPALAKFMAVPHYTYMMMKMPGPRGVLSL-RSDIKQAVTCDKESCDMAHTH 919
Query: 565 EFEVQVIDLDPREDFPQERIEPI--------EEVKDVVIGPLPHQITKIGTSLSGLEEEI 616
E D+ E P+ E K I P TK E
Sbjct: 920 EMASAREDIRLAAATASEGEIPVTKTSKSGESEAKTKKIPLDPSDPTKTA-------ESA 972
Query: 617 LVQLLRENVDLFAWAPSDMPGIDIGVACHHLAVRTSVKPVVQRKRKMGEEKRKAVDEEVK 676
L+ L+ N D+FAW PSDMPGI V H L V+ KP+ QR R+ ++++ A+ EE+
Sbjct: 973 LITFLQNNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAIKEELT 1032
Query: 677 KLQEAHFICEIKYPTWLANTVLVKKASGKWRMCVDYTDLNMACPKDPYPLPSIDHLIDNA 736
KL A FI E+ +P WLAN VLV+K +G+WRMCVDYTDLN +CPKDP+ LP ID ++D+
Sbjct: 1033 KLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPRIDQVVDST 1092
Query: 737 SGYKTLSFMDAYSGYNQIKMDPLDAPKTAFMTNQKNYHYRVMSFGLRNAGATFQRSMDTI 796
+G + LSF+D YSGY+QI++ D KT+F+T Y Y M FGL+NAGAT+QR +
Sbjct: 1093 AGCELLSFLDCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMPFGLKNAGATYQRMIQRC 1152
Query: 797 FSNQIGRNLEVYIDDLVVKTSEKQSHSVDLKEIFQQIRKFSMRLNPTKCTFGVQAGKFLG 856
FS QIGRN+E Y+DD+VVKT +K DL+E F IR F M+LNP KCTFGV +GK LG
Sbjct: 1153 FSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFVSIRAFRMKLNPEKCTFGVPSGKLLG 1212
Query: 857 FLLTKKGIEANPDKCQAIINMRNPCNIREVQQLTGRLAALSRFLSCAGDKAFAFFATIKK 916
F+++ +GI+ANP+K AI+NM+ P ++VQ+LTG +AALSRF+S G++ FF +KK
Sbjct: 1213 FMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKLLKK 1272
Query: 917 KEEFEWNQECDEAFGKIKQFLTTPPILHRPTKGAGLFLYLSVSENALSSVLVEESDER-- 974
+ F+W E +AF K+ LT PP+L P L LY+S + +S+VL E +E
Sbjct: 1273 TDNFQWGPEAQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTVLAVEREEEGH 1332
Query: 975 ----EKPIYFVSRVLKGAELRYQKIEKLALAVIITARKLRPYFQSHKVVIRTNYPVKQIL 1030
++PIYFVS VL ++ RY +++KL H V + T++P+ IL
Sbjct: 1333 VQKVQRPIYFVSEVLADSKTRYPQVQKL--------------LYGHSVTVVTSFPLGDIL 1378
Query: 1031 GKLDLAGRMLSWSVELSEYDIQFAPRNNIKSQVLADFVVEFTSPIQEAVP----HVWLLS 1086
+ GR+ W++EL DI F PR +IKSQ LADFV E+T QE P W +
Sbjct: 1379 HNREANGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTE-CQEDTPAENMEHWTMH 1437
Query: 1087 VDGSSNIKGSGAGIILEGPGDLLIEQSLKFDFKASNNQAEYEALIAGMLLAQEMGAKNLR 1146
DGS + G+GAG++L P + L F AS+N AEYEAL+ G+ +A +G K L
Sbjct: 1438 FDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIKRLI 1497
Query: 1147 ARSDSQLMTNQISGEYQTKDQQLSKYLARVRKLAGDFQFFEAIYVPRESNSRADLLAKLA 1206
R DSQL+ NQ+ E+ D + Y VRKL F E +V R +N AD LA
Sbjct: 1498 VRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLANFG 1557
Query: 1207 STKKPGNNRTVIQEVIS-----APSTDEKAVFELNQEPEGWMTPLLKFLTGSFVAKNDEY 1261
S ++ + ++ + + +T ++ W PL++FLT + ++ +
Sbjct: 1558 SKREVAPSDVFVEHLYTPTVPHKDTTQVAGTHDVAVVETDWREPLIRFLTSQELPQDKDE 1617
Query: 1262 AQLVRRRATKFVVIAGKLYKRGRASPLLRCLGEGETELVLLEVHEGVCGSHIGGRSLAAK 1321
A+ + RR+ +V+ +LYK+ + L RC+ E +L ++H G+CG+H R++ K
Sbjct: 1618 AERISRRSKLYVMHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNHAAARTIVGK 1677
Query: 1322 LLRAGYYWPRMAHDCCEFVKKCDKCQRFSDKKNAPANELTSVFSPWPFHKWGVDIVGPFP 1381
R G++WP D + V+ C+ CQ F+ + + PA EL ++ WPF WG+D+VGPF
Sbjct: 1678 AYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWGLDMVGPFK 1737
Query: 1382 QAPGQLKFLIVDVDYFTKWVEAEAVSKITAERVVKFYWKKIICHFGLPKYIVTDNGTQFA 1441
+A G L V +D F+KW+EA+ V ITA+ F+ I+ FG+P I+TDNGTQF
Sbjct: 1738 KAVGGYTHLFVAIDKFSKWIEAKPVVTITADNARDFF-INIVHRFGVPNRIITDNGTQFT 1796
Query: 1442 SSKVVNFCKQLGIETKFVSVIHPQANGQAESANKMIVNDIKKKLEDA----KGLWAEQLH 1497
+FC+ GI+ + SV HP +NGQ E AN MI+ IK ++ D G W +QL
Sbjct: 1797 GGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPYAGKWVQQLP 1856
Query: 1498 EVLWSYHTTPHSTTGETPFTMVYGADAMLPVEIDTPTWRREHFSEESNEVGIRCTMDMID 1557
VLWS TTP TG++PF +VYGA+AMLP E++ + R +F EE E + ++
Sbjct: 1857 SVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFREERYEEDRVDDLHRLE 1916
Query: 1558 EVREAAHIREFAAKQRAARRYNSKVIPRSMKEGDLVLKQVVAPTRIGKLLPSWKGPYRVK 1617
E REAA IR Q R +N V R+ GDLVL+++ KL P W+GP+ +
Sbjct: 1917 EAREAALIRSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTTRDRHKLSPLWEGPFIIS 1976
Query: 1618 EKLQHGAYKLEELSGNPV 1635
E + G+Y+L+ G V
Sbjct: 1977 EVTRPGSYRLKREDGTLV 1994
>ref|XP_475589.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|46485807|gb|AAS98432.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
gi|46391121|gb|AAS90648.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 2000
Score = 925 bits (2391), Expect = 0.0
Identities = 566/1649 (34%), Positives = 856/1649 (51%), Gaps = 96/1649 (5%)
Query: 25 VQQQEKTDSHHG----DLDEPEPQPFSTEIWNAPVPDNFKPPQLPTFDGRSDPSEHVTVF 80
+Q++ HH D D F+ ++ P FKP + +DG ++P +TV+
Sbjct: 396 IQRRAARGYHHSPDRRDDDLDGVAAFTDDLRRVDWPAGFKPTGIEKYDGTTNPESWLTVY 455
Query: 81 NTRMSVYGVADSLKCKLLAGTLADAALRWYMSLPRFSIVGYQDLTKKFTQQFSGTRHRKV 140
+ G + L LAD+A W LPR +I + +L F F GT R
Sbjct: 456 GLAIRAAGGDNKAMANYLPVALADSARSWLHGLPRGTIGSWAELRDHFIANFQGTFERPG 515
Query: 141 LSTSLFNVRQGPNESLREYLARFNDSTIKVSNPNQEVFVGAFQNGLRAGQFNESLAQKPA 200
L+NV Q ESLR+Y+ RF++ K+S+ +V + AF G+R + +KP
Sbjct: 516 TQYDLYNVIQKSGESLRDYIRRFSEQRNKISDITDDVIIAAFTKGIRHEELVGKFGRKPP 575
Query: 201 DSMEEIIARAECYVKGEESNAEKRARDVKEKGSSGGERRNHYVPPNRDRGTFKKPYERNQ 260
+++ + +A Y K E++ K+ G S ++N P G+ + +
Sbjct: 576 RTVKLMFEKANEYAKAEDAVTAS-----KQSGPSW--KQNKGTPATGGGGSNNHKDRKRK 628
Query: 261 PRYVPEHFTPLNTRPEKILKEVFESKIIPPPPFNRSKTMGQDKNAWCKYHLIEGHNTDDC 320
P E+ + +R T + N+ C +H H DC
Sbjct: 629 PA------------------ELVATASHSSRQRSRVNTFDKIMNSQCPHHPNSNHVAKDC 670
Query: 321 VHLKREIEKLLLNGKLRGYAKEKHHSERREDKPNP-----EPKHTLHTISGGFAGGGESS 375
K+ ++ + + +E+ S +++D P K H G A ES
Sbjct: 671 FIYKQFADQYTKTAR-KNSDEEQSTSRKKDDGDTPAGFQDHRKELNHIFGGPLAY--ESK 727
Query: 376 NLRKKYVRQVMLLGDSPSSPKKYPHVTFS---SEDFGRVLPHDDDPLVISVQLFNWEIKR 432
+K R++ + ++ + S+ RV+ PLV+ + N +++R
Sbjct: 728 RKQKLTEREINAVQPDTPQYLRWSEIAIKFDRSDHPDRVVHPGRYPLVLDPVVRNVKLRR 787
Query: 433 VLIDTGSSADVLYFDAFSKMGLSEEQLQPFNGTLSG-FTGEKVHVRGYVTLKTIFGTGDQ 491
LID GS+ ++L+ M + +L+P N G G G +TL FGT +
Sbjct: 788 TLIDGGSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPLGQITLPVTFGTREN 847
Query: 492 QTSIKVRYLVINSPSSYNVIIGRPSINLLDAFVSTKHLLMKYPLDSGRVGV--VHGDQKI 549
+ + + V N ++Y+ I+GRP++ A +++MK SG GV + D K
Sbjct: 848 FRTENISFEVANFETAYHAILGRPALAKFMAVPHYTYMMMKM---SGPRGVLSLRSDIKQ 904
Query: 550 ARECYHASLRLKKHKEFEVQVIDLDPREDFPQERIEPIEEVKDVVIGPLPHQITKIGTSL 609
A C S + + +E RED E G +P TK TS
Sbjct: 905 AVTCDKESCDMAQTREM------ASAREDIRLAAATASE-------GEIP--ATK--TSK 947
Query: 610 SGLEEEILVQLLRENVDLFAWAPSDMPGIDI----GVACHHLAVRTSVKPVVQRKRKMGE 665
SG E ++ + PSD + V H L V+ KP+ QR R+ +
Sbjct: 948 SGESEAKTKKIPLD--------PSDPTKTAVIGAEEVIEHSLHVKEDAKPIKQRLRRFAQ 999
Query: 666 EKRKAVDEEVKKLQEAHFICEIKYPTWLANTVLVKKASGKWRMCVDYTDLNMACPKDPYP 725
+++ A+ EE+ KL A FI E+ +P WLAN VLV+K +G+WRMCVDYTDLN +CPKDP+
Sbjct: 1000 DRKDAIKEELTKLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFG 1059
Query: 726 LPSIDHLIDNASGYKTLSFMDAYSGYNQIKMDPLDAPKTAFMTNQKNYHYRVMSFGLRNA 785
LP ID ++D+ +G + LSF+D YSGY+QI++ D KT+F+T Y Y M FGL+NA
Sbjct: 1060 LPRIDQVVDSTAGCELLSFLDCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMPFGLKNA 1119
Query: 786 GATFQRSMDTIFSNQIGRNLEVYIDDLVVKTSEKQSHSVDLKEIFQQIRKFSMRLNPTKC 845
GAT+QR + FS QIGRN+E Y+DD+VVKT +K DL+E F IR F M+LNP KC
Sbjct: 1120 GATYQRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKC 1179
Query: 846 TFGVQAGKFLGFLLTKKGIEANPDKCQAIINMRNPCNIREVQQLTGRLAALSRFLSCAGD 905
TFGV +GK LGF+++ +GI+ANP+K AI+NM+ P ++VQ+LTG +AALSRF+S G+
Sbjct: 1180 TFGVPSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGE 1239
Query: 906 KAFAFFATIKKKEEFEWNQECDEAFGKIKQFLTTPPILHRPTKGAGLFLYLSVSENALSS 965
+ FF +KK ++F+W E +AF K+ LT PP+L P L LY+S + +S+
Sbjct: 1240 RGMPFFKLLKKTDDFQWGPEAQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVST 1299
Query: 966 VLVEESDER------EKPIYFVSRVLKGAELRYQKIEKLALAVIITARKLRPYFQSHKVV 1019
VLV E +E ++PIYFVS VL ++ RY +++KL ++IT RKL YFQ H V
Sbjct: 1300 VLVVEREEEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVT 1359
Query: 1020 IRTNYPVKQILGKLDLAGRMLSWSVELSEYDIQFAPRNNIKSQVLADFVVEFTSPIQEAV 1079
+ T++P+ IL GR+ W++EL DI F PR +IKSQ LADFV E+T QE
Sbjct: 1360 VVTSFPLGDILHNRKANGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTE-CQEDT 1418
Query: 1080 P----HVWLLSVDGSSNIKGSGAGIILEGPGDLLIEQSLKFDFKASNNQAEYEALIAGML 1135
P W + DGS + G+GAG++L P + L F AS+N AEYEAL+ G+
Sbjct: 1419 PAENMEHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLR 1478
Query: 1136 LAQEMGAKNLRARSDSQLMTNQISGEYQTKDQQLSKYLARVRKLAGDFQFFEAIYVPRES 1195
+A +G K L R DSQL+ NQ+ E+ D + Y VRKL F E +V R +
Sbjct: 1479 IAISLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHN 1538
Query: 1196 NSRADLLAKLASTKKPGNNRTVIQEVIS-----APSTDEKAVFELNQEPEGWMTPLLKFL 1250
N AD LA S ++ + ++ + + +T + ++ W PL++FL
Sbjct: 1539 NEAADRLANFGSKREAAPSDVFVEHLYTPTVPHKDTTQDADTHDVAMVEADWREPLIRFL 1598
Query: 1251 TGSFVAKNDEYAQLVRRRATKFVVIAGKLYKRGRASPLLRCLGEGETELVLLEVHEGVCG 1310
T + ++ + A+ + RR+ +V+ +LYK+ + L RC+ E +L ++H G+CG
Sbjct: 1599 TSQELPQDKDEAERISRRSKLYVLHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSGICG 1658
Query: 1311 SHIGGRSLAAKLLRAGYYWPRMAHDCCEFVKKCDKCQRFSDKKNAPANELTSVFSPWPFH 1370
+H R++ K R G++WP D + V+ C+ CQ F+ + + PA EL ++ WPF
Sbjct: 1659 NHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFA 1718
Query: 1371 KWGVDIVGPFPQAPGQLKFLIVDVDYFTKWVEAEAVSKITAERVVKFYWKKIICHFGLPK 1430
WG+D+VGPF +A G L V +D F+KW+EA+ V ITA+ F+ I+ FG+P
Sbjct: 1719 VWGLDMVGPFKKAVGGYTHLFVAIDKFSKWIEAKPVVTITADNARDFF-ISIVHRFGVPN 1777
Query: 1431 YIVTDNGTQFASSKVVNFCKQLGIETKFVSVIHPQANGQAESANKMIVNDIKKKLEDA-- 1488
I+TDNGTQF +FC+ GI+ + SV HP +NGQ E AN MI+ IK ++ D
Sbjct: 1778 RIITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLK 1837
Query: 1489 --KGLWAEQLHEVLWSYHTTPHSTTGETPFTMVYGADAMLPVEIDTPTWRREHFSEESNE 1546
G W +QL VLWS TTP TG++PF +VYGA+AMLP E++ + R +F EE E
Sbjct: 1838 PYAGKWVQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFREERYE 1897
Query: 1547 VGIRCTMDMIDEVREAAHIREFAAKQRAARRYNSKVIPRSMKEGDLVLKQVVAPTRIGKL 1606
+ ++E REAA I+ Q R +N V R+ GDLVL+++ KL
Sbjct: 1898 EDRVDDLHRLEEAREAALIQSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTTRDRHKL 1957
Query: 1607 LPSWKGPYRVKEKLQHGAYKLEELSGNPV 1635
P W+GP+ + E + G+Y+L+ G V
Sbjct: 1958 SPLWEGPFIISEVTRPGSYRLKREDGTLV 1986
>gb|AAP52687.1| putative gag-pol precursor [Oryza sativa (japonica cultivar-group)]
gi|37532196|ref|NP_920400.1| putative gag-pol precursor
[Oryza sativa (japonica cultivar-group)]
gi|20451040|gb|AAM22011.1| Putative gag-pol precursor
[Oryza sativa (japonica cultivar-group)]
Length = 1933
Score = 922 bits (2383), Expect = 0.0
Identities = 558/1577 (35%), Positives = 823/1577 (51%), Gaps = 99/1577 (6%)
Query: 32 DSHHGDLDEPEPQPFSTEIWNAPVPDNFKPPQLPTFDGRSDPSEHVTVFNTRMSVYGVAD 91
D H DLD F+ + P FKP + +DG ++P +TV+ + G
Sbjct: 413 DRHDDDLDGVAA--FTDNLRRVDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDS 470
Query: 92 SLKCKLLAGTLADAALRWYMSLPRFSIVGYQDLTKKFTQQFSGTRHRKVLSTSLFNVRQG 151
L LAD+A W LPR +I + +L F F GT R L+NV Q
Sbjct: 471 KAMANYLPVALADSARSWLHGLPRGTIGSWAELRDHFIANFQGTFERPGTQFDLYNVVQK 530
Query: 152 PNESLREYLARFNDSTIKVSNPNQEVFVGAFQNGLRAGQFNESLAQKPADSMEEIIARAE 211
ESLR+Y+ RF++ K+S+ +V + AF G+R + +KP +++ + +A
Sbjct: 531 SGESLRDYIRRFSEQRNKISDITDDVIIAAFTKGIRHEELVGKFGRKPPRTVKLMFEKAN 590
Query: 212 CYVKGEESNAEKRARDVKEK------GSSGGERRNHYVPPNRDRGTFKKPYERNQPRYVP 265
Y K E++ + K + GG NH +DR
Sbjct: 591 EYAKAEDAVTASKQSGPSWKPNKGTPATGGGGSNNH-----KDR---------------- 629
Query: 266 EHFTPLNTRPEKILKEVFESKIIPPPPFNRSKTMGQDKNAWCKYHLIEGHNTDDCVHLKR 325
+PE+++ S +R T + N C +H H DC K+
Sbjct: 630 ------KRKPEELVATATHSS----RQRSRVNTFDKIMNPQCPHHPNSNHVAKDCFVYKQ 679
Query: 326 EIEKLLLNGKLRGYAKEKHHSERREDKPNPEPKHTLHTISGGFAGGG----ESSNLRKKY 381
E+ R + E+H + R++D + H GG ES RK
Sbjct: 680 FAEQYTKT--TRKNSDEEHSTSRKKDDGDTPAGFQDHRKELNHIFGGPLAYESKRKRKLT 737
Query: 382 VRQVMLLGDSPSSPKKYPHVTFS---SEDFGRVLPHDDDPLVISVQLFNWEIKRVLIDTG 438
R++ + ++ + S+ RV+ PLV+ + N +++R LID G
Sbjct: 738 EREINAVQPDTPQYLRWSEIAIKFDRSDHPDRVVHPGRYPLVLDPVVRNVKLRRTLIDGG 797
Query: 439 SSADVLYFDAFSKMGLSEEQLQPFNGTLSG-FTGEKVHVRGYVTLKTIFGTGDQQTSIKV 497
S+ ++L+ M + +L+P N G G +TL FGT + + +
Sbjct: 798 SALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPLCQITLPVTFGTRENFRTENI 857
Query: 498 RYLVINSPSSYNVIIGRPSINLLDAFVSTKHLLMKYPLDSGRVGVVHGDQKIARECYHAS 557
+ V + ++Y+ I+GRP++ A +++MK P G + + D K A C S
Sbjct: 858 SFEVADFETTYHAILGRPALAKFMAVPHYTYMMMKMPGPRGVLSL-RSDIKQAVTCDKES 916
Query: 558 LRLKKHKEFEVQVIDLDPREDFPQERIEPIEEVKDVVIGPLPHQITKIGTSLSGLEEEIL 617
+ + +E RED E G +P +TK TS SG E
Sbjct: 917 CDMAQTREMA------SAREDIQLAAATASE-------GEVP--VTK--TSKSGESEAKT 959
Query: 618 VQL---------LRENVDLFAWAPSDMPGIDIGVACHHLAVRTSVKPVVQRKRKMGEEKR 668
++ N D+FAW PSDMPGI V H L V+ KP+ QR R+ ++++
Sbjct: 960 KKIPLDPSDPTKTANNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRK 1019
Query: 669 KAVDEEVKKLQEAHFICEIKYPTWLANTVLVKKASGKWRMCVDYTDLNMACPKDPYPLPS 728
A+ EE+ KL A FI E+ +P WLAN VLV+K +G+WRMCVDYTDLN +CPKDP+ LP
Sbjct: 1020 DAIKEELTKLLAAGFIKEVHHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPR 1079
Query: 729 IDHLIDNASGYKTLSFMDAYSGYNQIKMDPLDAPKTAFMTNQKNYHYRVMSFGLRNAGAT 788
ID ++D+ +G + LSF+D YSGY+QI++ D KT+F+T Y Y MSFGL+NAGAT
Sbjct: 1080 IDQVVDSTAGCELLSFLDCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMSFGLKNAGAT 1139
Query: 789 FQRSMDTIFSNQIGRNLEVYIDDLVVKTSEKQSHSVDLKEIFQQIRKFSMRLNPTKCTFG 848
+QR + FS QIGRN+E Y+DD+VVKT +K DL+E F I F M+LNP KCTFG
Sbjct: 1140 YQRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIHAFRMKLNPEKCTFG 1199
Query: 849 VQAGKFLGFLLTKKGIEANPDKCQAIINMRNPCNIREVQQLTGRLAALSRFLSCAGDKAF 908
V +GK LGF+++ +GI+ANP+K AI+NM+ P ++VQ+LTG +AALSRF+S G++
Sbjct: 1200 VPSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGM 1259
Query: 909 AFFATIKKKEEFEWNQECDEAFGKIKQFLTTPPILHRPTKGAGLFLYLSVSENALSSVLV 968
FF +KK ++F+W E +AF K+ LT PP+L P L LY+S + +S+VLV
Sbjct: 1260 PFFKLLKKTDDFQWGPEAQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTVLV 1319
Query: 969 EESDER------EKPIYFVSRVLKGAELRYQKIEKLALAVIITARKLRPYFQSHKVVIRT 1022
E ++ ++PIYFVS VL ++ RY +++KL ++IT RKL YFQ H V + T
Sbjct: 1320 VEREKEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVT 1379
Query: 1023 NYPVKQILGKLDLAGRMLSWSVELSEYDIQFAPRNNIKSQVLADFVVEFTSPIQEAVP-- 1080
++P+ IL + GR+ W++EL DI F PR +IKSQ LADFV E+T QE P
Sbjct: 1380 SFPLGDILHNREANGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTE-CQEDTPAE 1438
Query: 1081 --HVWLLSVDGSSNIKGSGAGIILEGPGDLLIEQSLKFDFKASNNQAEYEALIAGMLLAQ 1138
W + DGS + G+GAG++L P + L F AS+N AEYEAL+ G+ +A
Sbjct: 1439 KMEHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAI 1498
Query: 1139 EMGAKNLRARSDSQLMTNQISGEYQTKDQQLSKYLARVRKLAGDFQFFEAIYVPRESNSR 1198
+G K L R DSQL+ NQ+ E+ D ++ Y VRKL F E +V R +N
Sbjct: 1499 SLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMTAYRQEVRKLEDKFDGLELSHVLRHNNEA 1558
Query: 1199 ADLLAKLASTKKPGNNRTVIQEVISAPSTDEK------AVFELNQEPEGWMTPLLKFLTG 1252
AD LA S K+ G V E + P+ K ++ W PL++FLT
Sbjct: 1559 ADRLANFGS-KREGAPSDVFVEHLYTPTVPHKDTTQVAGTHDVAMVEADWREPLIRFLTS 1617
Query: 1253 SFVAKNDEYAQLVRRRATKFVVIAGKLYKRGRASPLLRCLGEGETELVLLEVHEGVCGSH 1312
+ ++ + A+ + RR+ +V+ +LYK+ + L RC+ E +L ++H G+CG+H
Sbjct: 1618 QELPQDKDEAERISRRSKLYVMHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNH 1677
Query: 1313 IGGRSLAAKLLRAGYYWPRMAHDCCEFVKKCDKCQRFSDKKNAPANELTSVFSPWPFHKW 1372
R++ K R G++WP D + V+ C+ CQ F+ + + PA EL ++ WPF W
Sbjct: 1678 AAARTIIGKAYRQGFFWPAAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVW 1737
Query: 1373 GVDIVGPFPQAPGQLKFLIVDVDYFTKWVEAEAVSKITAERVVKFYWKKIICHFGLPKYI 1432
G+D+VGPF +A G L V +D F+KW+EA+ V ITA+ F+ I+ FG+P I
Sbjct: 1738 GLDMVGPFKKAVGGFTHLFVAIDKFSKWIEAKPVVTITADNARDFF-INIVHRFGVPNRI 1796
Query: 1433 VTDNGTQFASSKVVNFCKQLGIETKFVSVIHPQANGQAESANKMIVNDIKKKLEDA---- 1488
+TDNG QF +FC+ GI+ + SV HP +NGQ E AN MI+ IK ++ D
Sbjct: 1797 ITDNGRQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPY 1856
Query: 1489 KGLWAEQLHEVLWSYHTTPHSTTGETPFTMVYGADAMLPVEIDTPTWRREHFSEESNEVG 1548
G W EQL VLWS TTP TG++PF +VYGA+AMLP E++ + R +F EE E
Sbjct: 1857 AGKWVEQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFREERYEED 1916
Query: 1549 IRCTMDMIDEVREAAHI 1565
+ ++EVREAA I
Sbjct: 1917 RVDDLHQLEEVREAALI 1933
>gb|AAP54065.1| putative gypsy-type retrotransposon [Oryza sativa (japonica
cultivar-group)] gi|37534952|ref|NP_921778.1| putative
gypsy-type retrotransposon [Oryza sativa (japonica
cultivar-group)]
Length = 1995
Score = 919 bits (2375), Expect = 0.0
Identities = 564/1641 (34%), Positives = 843/1641 (51%), Gaps = 109/1641 (6%)
Query: 32 DSHHGDLDEPEPQPFSTEIWNAPVPDNFKPPQLPTFDGRSDPSEHVTVFNTRMSVYGVAD 91
D H DLD F+ ++ A P FKP + +DG ++P +TV+ + G
Sbjct: 413 DRHDDDLDGVAA--FTDDLRRADWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDS 470
Query: 92 SLKCKLLAGTLADAALRWYMSLPRFSIVGYQDLTKKFTQQFSGTRHRKVLSTSLFNVRQG 151
L LAD+A W LPR +I + +L F F GT R L+NV Q
Sbjct: 471 KAMANYLPVALADSARSWLHGLPRGTIGSWAELRDHFIANFQGTFERPGTQYDLYNVIQK 530
Query: 152 PNESLREYLARFNDSTIKVSNPNQEVFVGAFQNGLRAGQFNESLAQKPADSMEEIIARAE 211
ESLR+Y+ RF++ K+S+ +V + AF G+R + +KP +++ + +
Sbjct: 531 SGESLRDYIRRFSEQRNKISDITDDVIIAAFTKGIRHEELVGKFGRKPPRTVKLMFEKVN 590
Query: 212 CYVKGEESNAEKRARDVKEKGSSGGERRNHYVPPNRDRGTFKKPYERNQPRYVPEHFTPL 271
Y K E++ K S G + N P G+ +
Sbjct: 591 EYAKAEDAVTAS-------KQSGPGWKPNKGTPATGGGGS--------------NNHKDR 629
Query: 272 NTRPEKILKEVFESKIIPPPPFNRSKTMGQDKNAWCKYHLIEGHNTDDCVHLKREIEKLL 331
+PE+++ S P R T + N+ C ++ H DC K+ E+
Sbjct: 630 KRKPEELVATATHSSRQRP----RVNTFDKIMNSQCPHYPNSNHVAKDCFVYKQFAEQYT 685
Query: 332 LNGKLRGYAKEKHHSERREDKPNP-----EPKHTLHTISGGFAGGGESSNLRKKYVRQVM 386
+ + + +E+ S +++D P K H G A ES +K R++
Sbjct: 686 KSTR-KNSDEEQSTSRKKDDGDTPAGFQDHRKELNHIFGGPLAY--ESKRKQKLTEREIN 742
Query: 387 LLGDSPSSPKKYPHVTFS---SEDFGRVLPHDDDPLVISVQLFNWEIKRVLIDTGSSADV 443
+ ++ + S+ RV+ PLV+ + N +++R LID GS+ ++
Sbjct: 743 AVQPDTPQYLRWSEIAIKFDRSDHPDRVVHPGRYPLVLDPVVRNVKLRRTLIDGGSALNI 802
Query: 444 LYFDAFSKMGLSEEQLQPFNGTLSG-FTGEKVHVRGYVTLKTIFGTGDQQTSIKVRYLVI 502
L+ M + +L+P N G G G +TL FGT + + + + V
Sbjct: 803 LFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPLGQITLPVTFGTRENFRTENISFEVA 862
Query: 503 NSPSSYNVIIGRPSINLLDAFVSTKHLLMKYPLDSGRVGVVHGDQKIARECYHASLRLKK 562
+ ++Y+ I+GRP++ A +++MK P G + + D K A C S + +
Sbjct: 863 DFETAYHAILGRPALAKFMAVPHYTYMMMKMPGPRGVLSL-RSDIKQAVTCDKESCDMAQ 921
Query: 563 HKEFEVQVIDLDPREDFPQERIEPIEEVKDVVIGPLPHQITKIGTSLSGLEEEILVQL-- 620
+E RED E G +P TK TS SG E ++
Sbjct: 922 TREMA------SAREDIRLAAATASE-------GEVP--ATK--TSKSGESEAKTKKIPL 964
Query: 621 -------LRENVDLFAWAPSDMPGIDIGVACHHLAVRTSVKPVVQRKRKMGEEKRKAVDE 673
N D+FAW PSDMPGI V H L V+ KP+ QR R+ ++++ A+ E
Sbjct: 965 DPSDPTKTANNKDIFAWKPSDMPGIPREVIEHSLHVQEDAKPIKQRLRRFAQDRKDAIKE 1024
Query: 674 EVKKLQEAHFICEIKYPTWLANTVLVKKASGKWRMCVDYTDLNMACPKDPYPLPSIDHLI 733
E+ KL A FI E+ +P WLAN VLV+K +G+WRMCVDYTDLN +CPKDP+ LP ID
Sbjct: 1025 ELTKLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPRID--- 1081
Query: 734 DNASGYKTLSFMDAYSGYNQIKMDPLDAPKTAFMTNQKNYHYRVMSFGLRNAGATFQRSM 793
QI++ D KT+F+T Y Y M FGL+NAGAT+QR +
Sbjct: 1082 -------------------QIRLKESDCLKTSFITPFGAYCYVTMPFGLKNAGATYQRMI 1122
Query: 794 DTIFSNQIGRNLEVYIDDLVVKTSEKQSHSVDLKEIFQQIRKFSMRLNPTKCTFGVQAGK 853
FS QIGRN+E Y+DD+VVKT +K DL+E F IR F M+LNP KCTFGV +GK
Sbjct: 1123 QRCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFTSIRAFRMKLNPEKCTFGVPSGK 1182
Query: 854 FLGFLLTKKGIEANPDKCQAIINMRNPCNIREVQQLTGRLAALSRFLSCAGDKAFAFFAT 913
LGF+++ +GI+ANP+K AI+NM+ P ++VQ+LTG +AALSRF+S G++ FF
Sbjct: 1183 LLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKL 1242
Query: 914 IKKKEEFEWNQECDEAFGKIKQFLTTPPILHRPTKGAGLFLYLSVSENALSSVLVEESDE 973
+KK ++F+W E +AF K+ LT PP+L P L LY+S + +S+VLV E +E
Sbjct: 1243 LKKTDDFQWGPEAQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTVLVVEREE 1302
Query: 974 R------EKPIYFVSRVLKGAELRYQKIEKLALAVIITARKLRPYFQSHKVVIRTNYPVK 1027
++PIYFVS VL ++ RY +++KL ++IT RKL YFQ H V + T++P+
Sbjct: 1303 EGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSFPLG 1362
Query: 1028 QILGKLDLAGRMLSWSVELSEYDIQFAPRNNIKSQVLADFVVEFTSPIQEAVP----HVW 1083
IL + GR+ W++EL DI F PR +IKSQ LADFV E+T QE P W
Sbjct: 1363 DILHNREANGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTE-CQEDTPAEKMEHW 1421
Query: 1084 LLSVDGSSNIKGSGAGIILEGPGDLLIEQSLKFDFKASNNQAEYEALIAGMLLAQEMGAK 1143
+ DGS + G+GAG++L P + L F AS+N AEYEAL+ G+ +A +G K
Sbjct: 1422 TMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIK 1481
Query: 1144 NLRARSDSQLMTNQISGEYQTKDQQLSKYLARVRKLAGDFQFFEAIYVPRESNSRADLLA 1203
L R DSQL+ NQ+ E+ D + Y VRKL F E +V R +N AD L
Sbjct: 1482 RLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLV 1541
Query: 1204 KLASTKKPGNNRTVIQEVIS-----APSTDEKAVFELNQEPEGWMTPLLKFLTGSFVAKN 1258
S ++ + ++ + + +T ++ W PL++FLT + ++
Sbjct: 1542 NFGSKREVAPSDVFVEHLYTPTVPHKDTTQAAGTHDVAMVEADWREPLIRFLTSQELPQD 1601
Query: 1259 DEYAQLVRRRATKFVVIAGKLYKRGRASPLLRCLGEGETELVLLEVHEGVCGSHIGGRSL 1318
+ A+ + RR+ +V+ +LYK+ + L RC+ E +L ++H G+CG+H R++
Sbjct: 1602 KDEAERISRRSKLYVMHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNHAAARTI 1661
Query: 1319 AAKLLRAGYYWPRMAHDCCEFVKKCDKCQRFSDKKNAPANELTSVFSPWPFHKWGVDIVG 1378
K R G++WP D + V+ C+ CQ F+ + + PA EL ++ WPF WG+D+VG
Sbjct: 1662 VGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWGLDMVG 1721
Query: 1379 PFPQAPGQLKFLIVDVDYFTKWVEAEAVSKITAERVVKFYWKKIICHFGLPKYIVTDNGT 1438
PF +A G L V +D F+KW+EA+ V ITA+ F+ I+ FG+P I+TDNGT
Sbjct: 1722 PFKKAVGGYTHLFVAIDKFSKWIEAKPVVTITADNARDFF-INIVHRFGVPNRIITDNGT 1780
Query: 1439 QFASSKVVNFCKQLGIETKFVSVIHPQANGQAESANKMIVNDIKKKLEDA----KGLWAE 1494
QF +FC+ GI+ + SV HP +NGQ E AN MI+ IK ++ D G W +
Sbjct: 1781 QFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPYAGKWVQ 1840
Query: 1495 QLHEVLWSYHTTPHSTTGETPFTMVYGADAMLPVEIDTPTWRREHFSEESNEVGIRCTMD 1554
QL VLWS TTP TG++PF +VYGA+AMLP E++ + R +F E+ E +
Sbjct: 1841 QLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFHEDRYEEDRVDDLH 1900
Query: 1555 MIDEVREAAHIREFAAKQRAARRYNSKVIPRSMKEGDLVLKQVVAPTRIGKLLPSWKGPY 1614
++EVREAA IR Q R +N V R+ GDLVL+++ KL P W+GP+
Sbjct: 1901 RLEEVREAALIRSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTTRDRHKLSPLWEGPF 1960
Query: 1615 RVKEKLQHGAYKLEELSGNPV 1635
+ E + G+Y+L+ G V
Sbjct: 1961 IISEVTRPGSYRLKREDGTLV 1981
>gb|AAR96234.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 2004
Score = 917 bits (2369), Expect = 0.0
Identities = 565/1646 (34%), Positives = 848/1646 (51%), Gaps = 119/1646 (7%)
Query: 32 DSHHGDLDEPEPQPFSTEIWNAPVPDNFKPPQLPTFDGRSDPSEHVTVFNTRMSVYGVAD 91
D H DLD F+ ++ P FKP + +DG ++P +TV+ + G
Sbjct: 413 DRHDDDLDGVAA--FTDDLRRVDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDS 470
Query: 92 SLKCKLLAGTLADAALRWYMSLPRFSIVGYQDLTKKFTQQFSGTRHRKVLSTSLFNVRQG 151
L LAD+A W LPR +I + +L F F GT R L+NV Q
Sbjct: 471 KAMANYLPVALADSARSWLHGLPRGTIGSWAELRDHFIANFQGTFERPGTQFDLYNVVQK 530
Query: 152 PNESLREYLARFNDSTIKVSNPNQEVFVGAFQNGLRAGQFNESLAQKPADSMEEIIARAE 211
ESLR+Y+ RF++ K+S+ +V + AF G+R + +KP +++ + +A
Sbjct: 531 SGESLRDYIRRFSEQRNKISDITDDVIIAAFTKGIRHEELVGKFGRKPPRTVKLMFEKAN 590
Query: 212 CYVKGEES-NAEKRA----RDVKEKGSSGGERRNHYVPPNRDRGTFKKPYERNQPRYVPE 266
Y K E++ A K++ + K+ ++GG N++ +DR
Sbjct: 591 EYAKAEDAVTASKQSGPSWKPKKDTPATGGGGSNNH----KDR----------------- 629
Query: 267 HFTPLNTRPEKILKEVFESKIIPPPPFNRSKTMGQDKNAWCKYHLIEGHNTDDCVHLKRE 326
+PE+++ S +R T + N+ C +H H DC K+
Sbjct: 630 -----KRKPEELVATAIHSS----RQRSRVNTFDKIMNSQCPHHPNSNHVAKDCFVYKQF 680
Query: 327 IEKLLLNGKLRGYAKEKHHSERREDKPNP-----EPKHTLHTISGGFAGGGESSNLRKKY 381
E+ + + +E+ S +++D P K H G A ES +K
Sbjct: 681 AEQYTKTTR-KNSDEEQSTSRKKDDGDTPVGFQDHRKELNHIFGGPLAY--ESKRKQKLT 737
Query: 382 VRQVMLLGDSPSSPKKYPHVTFS---SEDFGRVLPHDDDPLVISVQLFNWEIKRVLIDTG 438
R++ + ++ + S+ RV+ PLV+ + N +++R LID G
Sbjct: 738 EREINAVQPDTPQYLRWSEIAIKFDRSDHPDRVVHPGRYPLVLDPVVRNVKLRRTLIDGG 797
Query: 439 SSADVLYFDAFSKMGLSEEQLQPFNGTLSG-FTGEKVHVRGYVTLKTIFGTGDQQTSIKV 497
S+ ++L+ M + +L+P N G G G +TL FGT + + +
Sbjct: 798 SALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPLGQITLPVTFGTRENFRTENI 857
Query: 498 RYLVINSPSSYNVIIGRPSINLLDAFVSTKHLLMKYPLDSGRVGVVHGDQKIARECYHAS 557
+ V + ++Y+ I+GRP++ A +++MK P G + + D K A C S
Sbjct: 858 SFEVADFETAYHAILGRPALAKFMAVPHYTYMMMKMPGPRGVLSL-RSDIKQAVTCDKES 916
Query: 558 LRLKKHKEFEVQVIDLDPREDFPQERIEPIEEVKDVVIGPLPHQITKIGTSLSGLEEEIL 617
+ + +E RED E G +P TK TS SG E
Sbjct: 917 CDMAQTREMA------SAREDIRLAAATASE-------GEVP--ATK--TSKSGESEAKT 959
Query: 618 VQL---------LRENVDLFAWAPSDMPGIDIGVACHHLAVRTSVKPVVQRKRKMGEEKR 668
++ N D+FAW PSDMPGI V H L V+ KP+ QR R+ ++++
Sbjct: 960 KKIPLDPSNPTKTANNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRK 1019
Query: 669 KAVDEEVKKLQEAHFICEIKYPTWLANTVLVKKASGKWRMCVDYTDLNMACPKDPYPLPS 728
A+ EE+ KL A FI E+ +P WLAN VLV+K +G+WRMCVDYTDLN +CPKDP+ LP
Sbjct: 1020 DAIKEELTKLLAAGFIKEVHHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPR 1079
Query: 729 IDHLIDNASGYKTLSFMDAYSGYNQIKMDPLDAPKTAFMTNQKNYHYRVMSFGLRNAGAT 788
ID QI++ D KT+F+T Y Y M FGL+NAGAT
Sbjct: 1080 ID----------------------QIRLKESDCLKTSFITPFGAYCYVTMPFGLKNAGAT 1117
Query: 789 FQRSMDTIFSNQIGRNLEVYIDDLVVKTSEKQSHSVDLKEIFQQIRKFSMRLNPTKCTFG 848
+QR + FS QIGRN+E Y+DD+VVKT +K DL+E F IR F M+LNP KCTFG
Sbjct: 1118 YQRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFG 1177
Query: 849 VQAGKFLGFLLTKKGIEANPDKCQAIINMRNPCNIREVQQLTGRLAALSRFLSCAGDKAF 908
V +GK LGF+++ +GI+ANP+K AI+NM+ P ++VQ+LTG +AALSRF+S G++
Sbjct: 1178 VPSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGM 1237
Query: 909 AFFATIKKKEEFEWNQECDEAFGKIKQFLTTPPILHRPTKGAGLFLYLSVSENALSSVLV 968
FF +KK ++F+W E +AF K+ LT PP+L P L LY+S + +S+VLV
Sbjct: 1238 PFFKLLKKTDDFQWGPEAQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTVLV 1297
Query: 969 EESDER------EKPIYFVSRVLKGAELRYQKIEKLALAVIITARKLRPYFQSHKVVIRT 1022
E +E ++PIYFVS VL ++ RY +++KL ++IT RKL YFQ H V + T
Sbjct: 1298 VEREEEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVT 1357
Query: 1023 NYPVKQILGKLDLAGRMLSWSVELSEYDIQFAPRNNIKSQVLADFVVEFTSPIQEAVP-- 1080
++P+ IL + GR+ W++EL DI F PR +IKSQ LADFV E+T QE P
Sbjct: 1358 SFPLGDILHNREANGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTE-CQEDTPAE 1416
Query: 1081 --HVWLLSVDGSSNIKGSGAGIILEGPGDLLIEQSLKFDFKASNNQAEYEALIAGMLLAQ 1138
W + DGS + G+GAG++L P + L F AS+N AEYEAL+ G+ +A
Sbjct: 1417 KMEHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAI 1476
Query: 1139 EMGAKNLRARSDSQLMTNQISGEYQTKDQQLSKYLARVRKLAGDFQFFEAIYVPRESNSR 1198
+G K L R DSQL+ NQ+ E+ D + Y VRKL F E +V R +N
Sbjct: 1477 SLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEA 1536
Query: 1199 ADLLAKLASTKKPGNNRTVIQEVIS-----APSTDEKAVFELNQEPEGWMTPLLKFLTGS 1253
AD LA S ++ + ++ + + +T ++ W PL++FLT
Sbjct: 1537 ADRLANFGSKREVAPSDVFVEHLYTPTVPHKDTTQVAGTHDVAMVEADWREPLIRFLTSQ 1596
Query: 1254 FVAKNDEYAQLVRRRATKFVVIAGKLYKRGRASPLLRCLGEGETELVLLEVHEGVCGSHI 1313
+ ++ + A+ + RR+ +V+ +LYK+ + L RC+ E +L ++H G+CG+H
Sbjct: 1597 ELPQDKDEAERISRRSKLYVMHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNHA 1656
Query: 1314 GGRSLAAKLLRAGYYWPRMAHDCCEFVKKCDKCQRFSDKKNAPANELTSVFSPWPFHKWG 1373
R++ K R G++WP D + V+ C+ CQ F+ + + PA EL ++ WPF WG
Sbjct: 1657 AARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWG 1716
Query: 1374 VDIVGPFPQAPGQLKFLIVDVDYFTKWVEAEAVSKITAERVVKFYWKKIICHFGLPKYIV 1433
+D+VGPF +A G L V +D F+KW+EA+ V ITA+ F+ I+ FG+P I+
Sbjct: 1717 LDMVGPFKKAVGGYTHLFVAIDKFSKWIEAKPVVTITADNAPDFF-INIVHRFGVPNRII 1775
Query: 1434 TDNGTQFASSKVVNFCKQLGIETKFVSVIHPQANGQAESANKMIVNDIKKKLEDA----K 1489
TDNG QF + C+ GI+ + SV HP +NGQ E AN MI+ IK ++ D
Sbjct: 1776 TDNGRQFTGGVFKDCCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPYA 1835
Query: 1490 GLWAEQLHEVLWSYHTTPHSTTGETPFTMVYGADAMLPVEIDTPTWRREHFSEESNEVGI 1549
G W EQL VLWS TTP TG++PF +VYGA+AMLP E++ + R +F EE E
Sbjct: 1836 GKWVEQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFREERYEEDR 1895
Query: 1550 RCTMDMIDEVREAAHIREFAAKQRAARRYNSKVIPRSMKEGDLVLKQVVAPTRIGKLLPS 1609
+ ++EVREAA I+ Q R +N V R+ GDLVL+++ KL P
Sbjct: 1896 VDDLHRLEEVREAALIQSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTTRDRHKLSPL 1955
Query: 1610 WKGPYRVKEKLQHGAYKLEELSGNPV 1635
W+GP+ + + G+Y+L+ G V
Sbjct: 1956 WEGPFIISAVTRPGSYRLKREDGTLV 1981
>ref|XP_474973.1| OSJNBb0015G09.7 [Oryza sativa (japonica cultivar-group)]
gi|38346888|emb|CAE03913.2| OSJNBb0015G09.7 [Oryza sativa
(japonica cultivar-group)]
Length = 1991
Score = 915 bits (2364), Expect = 0.0
Identities = 563/1652 (34%), Positives = 845/1652 (51%), Gaps = 111/1652 (6%)
Query: 25 VQQQEKTDSHHG----DLDEPEPQPFSTEIWNAPVPDNFKPPQLPTFDGRSDPSEHVTVF 80
+Q++ HH D D F+ ++ P FKP + +DG ++P +TV+
Sbjct: 396 IQRRAARGYHHSPDRRDDDLDGVAAFTDDLRRVDWPAGFKPTGIEKYDGTTNPESWLTVY 455
Query: 81 NTRMSVYGVADSLKCKLLAGTLADAALRWYMSLPRFSIVGYQDLTKKFTQQFSGTRHRKV 140
+ G + L LAD+A W LPR +I + +L F F GT R
Sbjct: 456 GLAIRAAGGDNKAMANYLPVALADSARSWLHGLPRGTIGSWAELRDHFIANFQGTFERPG 515
Query: 141 LSTSLFNVRQGPNESLREYLARFNDSTIKVSNPNQEVFVGAFQNGLRAGQFNESLAQKPA 200
L+NV Q ESLR+Y+ RF++ K+S+ +V + AF G+R + +KP
Sbjct: 516 TQYDLYNVIQKSGESLRDYIRRFSEQRNKISDITDDVIIAAFTKGIRHEELVGKFGRKPP 575
Query: 201 DSMEEIIARAECYVKGEESNAEKRARDVKEKGSSGGERRNHYVPPNRDRGTFKKPYERNQ 260
+++ + +A Y K E++ K+ G S ++N P G+ + +
Sbjct: 576 RTVKLMFEKANEYAKAEDAVTAS-----KQSGPSW--KQNKGTPATGGGGSNNHKDRKRK 628
Query: 261 PRYVPEHFTPLNTRPEKILKEVFESKIIPPPPFNRSKTMGQDKNAWCKYHLIEGHNTDDC 320
P E+ + +R T + N+ C +H H DC
Sbjct: 629 PA------------------ELVATASHSSRQRSRVNTFDKIMNSQCPHHPNSNHVAKDC 670
Query: 321 VHLKREIEKLLLNGKLRGYAKEKHHSERREDKPNP-----EPKHTLHTISGGFAGGGESS 375
K+ ++ + + +E+ S +++D P K H G A ES
Sbjct: 671 FIYKQFADQYTKTAR-KNSDEEQSTSRKKDDGDTPAGFQDHRKELNHIFGGPLAY--ESK 727
Query: 376 NLRKKYVRQVMLLGDSPSSPKKYPHVTFS---SEDFGRVLPHDDDPLVISVQLFNWEIKR 432
+K R++ + ++ + S+ RV+ PLV+ + N +++R
Sbjct: 728 RKQKLTEREINAVQPDTPQYLRWSEIAIKFDRSDHPDRVVHPGRYPLVLDPVVRNVKLRR 787
Query: 433 VLIDTGSSADVLYFDAFSKMGLSEEQLQPFNGTLSG-FTGEKVHVRGYVTLKTIFGTGDQ 491
LID GS+ ++L+ M + +L+P N G G G +TL FGT +
Sbjct: 788 TLIDGGSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPLGQITLPVTFGTREN 847
Query: 492 QTSIKVRYLVINSPSSYNVIIGRPSINLLDAFVSTKHLLMKYPLDSGRVGVVHGDQKIAR 551
+ + + V + ++Y+ I+GRP++ A +++MK P G + + D K A
Sbjct: 848 FRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMKMPGPRGVLSL-RSDIKQAV 906
Query: 552 ECYHASLRLKKHKEFEVQVIDLDPREDFPQERIEPIEEVKDVVIGPLPHQITKIGTSLSG 611
C S + + +E RED E G +P TK TS SG
Sbjct: 907 TCDKESCDMAQTREMA------SAREDIRLAAATASE-------GEIP--ATK--TSKSG 949
Query: 612 LEEEILVQL---------LRENVDLFAWAPSDMPGIDIGVACHHLAVRTSVKPVVQRKRK 662
E ++ N D+FAW PSDMPGI V H L V+ KP+ QR R+
Sbjct: 950 ESEAKTKKIPLDPSDPAKTANNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRR 1009
Query: 663 MGEEKRKAVDEEVKKLQEAHFICEIKYPTWLANTVLVKKASGKWRMCVDYTDLNMACPKD 722
++++ A+ EE+ KL A FI E+ +P WLAN VLV+K +G+WRMCVDYTDLN +CPKD
Sbjct: 1010 FAQDRKDAIKEELTKLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKD 1069
Query: 723 PYPLPSIDHLIDNASGYKTLSFMDAYSGYNQIKMDPLDAPKTAFMTNQKNYHYRVMSFGL 782
P+ LP ID QI++ D KT+F+T Y Y M FGL
Sbjct: 1070 PFGLPRID----------------------QIRLKESDCLKTSFITPFGAYCYVTMPFGL 1107
Query: 783 RNAGATFQRSMDTIFSNQIGRNLEVYIDDLVVKTSEKQSHSVDLKEIFQQIRKFSMRLNP 842
+NAGAT+QR + FS QIGRN+E Y+DD+VVKT +K DL+E F IR F M+LNP
Sbjct: 1108 KNAGATYQRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNP 1167
Query: 843 TKCTFGVQAGKFLGFLLTKKGIEANPDKCQAIINMRNPCNIREVQQLTGRLAALSRFLSC 902
KCTFGV +GK LGF+++ +GI+ANP+K AI+NM+ P ++VQ+LTG +AALSRF+S
Sbjct: 1168 EKCTFGVPSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSR 1227
Query: 903 AGDKAFAFFATIKKKEEFEWNQECDEAFGKIKQFLTTPPILHRPTKGAGLFLYLSVSENA 962
G++ FF +KK + F+W E +AF K+ LT PP+L P L LY+S +
Sbjct: 1228 LGERGMPFFKLLKKTDNFQWGPEAQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQV 1287
Query: 963 LSSVLVEESDER------EKPIYFVSRVLKGAELRYQKIEKLALAVIITARKLRPYFQSH 1016
+S+VLV E +E ++PIYFVS VL ++ RY +++KL ++IT RKL YFQ H
Sbjct: 1288 VSTVLVVEREEEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGH 1347
Query: 1017 KVVIRTNYPVKQILGKLDLAGRMLSWSVELSEYDIQFAPRNNIKSQVLADFVVEFTSPIQ 1076
V + T++P+ IL + GR+ W++EL DI F PR +IKSQ LADFV E+T Q
Sbjct: 1348 SVTVVTSFPLGDILHNREANGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTE-CQ 1406
Query: 1077 EAVP----HVWLLSVDGSSNIKGSGAGIILEGPGDLLIEQSLKFDFKASNNQAEYEALIA 1132
E P W + DGS + G+GAG++L P + L F AS+N AEYEAL+
Sbjct: 1407 EDTPAENMEHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLH 1466
Query: 1133 GMLLAQEMGAKNLRARSDSQLMTNQISGEYQTKDQQLSKYLARVRKLAGDFQFFEAIYVP 1192
G+ +A +G K+L R DSQL+ NQ+ E+ D + Y VRKL F E +V
Sbjct: 1467 GLRIAISLGIKHLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVL 1526
Query: 1193 RESNSRADLLAKLASTKKPGNNRTVIQEVIS-----APSTDEKAVFELNQEPEGWMTPLL 1247
R +N AD LA S ++ + ++ + S +T ++ W L+
Sbjct: 1527 RHNNEAADRLANFGSKREAAPSDVFVEHLYSPTVPHKDATQVAGTHDVAMVEADWRESLI 1586
Query: 1248 KFLTGSFVAKNDEYAQLVRRRATKFVVIAGKLYKRGRASPLLRCLGEGETELVLLEVHEG 1307
+FLT + ++ + A+ + RR+ +V+ +LYK+ + L RC+ E +L ++H G
Sbjct: 1587 RFLTSQELPQDKDEAERISRRSKLYVLHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSG 1646
Query: 1308 VCGSHIGGRSLAAKLLRAGYYWPRMAHDCCEFVKKCDKCQRFSDKKNAPANELTSVFSPW 1367
+CG+H R++ K R G++WP D + V+ C+ CQ F+ + + PA EL ++ W
Sbjct: 1647 ICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSW 1706
Query: 1368 PFHKWGVDIVGPFPQAPGQLKFLIVDVDYFTKWVEAEAVSKITAERVVKFYWKKIICHFG 1427
PF WG+D+VGPF +A G L V +D F+KW+EA+ V ITA+ F+ I+ FG
Sbjct: 1707 PFAVWGLDMVGPFKKAVGGYTHLFVAIDKFSKWIEAKPVVTITADNARDFF-ISIVHRFG 1765
Query: 1428 LPKYIVTDNGTQFASSKVVNFCKQLGIETKFVSVIHPQANGQAESANKMIVNDIKKKLED 1487
+P I+TDNGTQF +FC+ GI+ + SV HP +NGQ E AN MI+ IK ++ D
Sbjct: 1766 VPNRIITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFD 1825
Query: 1488 A----KGLWAEQLHEVLWSYHTTPHSTTGETPFTMVYGADAMLPVEIDTPTWRREHFSEE 1543
G W +QL VLWS TTP TG++PF +VYGA+AMLP E++ + R +F EE
Sbjct: 1826 RLKPYAGKWVQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFREE 1885
Query: 1544 SNEVGIRCTMDMIDEVREAAHIREFAAKQRAARRYNSKVIPRSMKEGDLVLKQVVAPTRI 1603
E + ++E REAA IR Q R +N V R+ GDLVL+++
Sbjct: 1886 RYEEDRVDDLHRLEEAREAALIRSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTTRDR 1945
Query: 1604 GKLLPSWKGPYRVKEKLQHGAYKLEELSGNPV 1635
KL P W+GP+ + E + G+Y+L+ G V
Sbjct: 1946 HKLSPLWEGPFIISEVTRPGSYRLKREDGTLV 1977
>gb|AAL76007.1| prpol [Zea mays]
Length = 1317
Score = 912 bits (2357), Expect = 0.0
Identities = 494/1308 (37%), Positives = 785/1308 (59%), Gaps = 39/1308 (2%)
Query: 356 EPKHTLHTISGGFAGGGESSNLRKKYVRQVMLLG-DSPSSPKKYPHV--TFSSEDFG-RV 411
+PK L I+GG + + +K+ R+V +G P ++ H+ TFS ED +
Sbjct: 13 DPKLVL-PITGGSSSEPANKKQKKEAQRRVQHVGVQGPFIKSRWSHIPITFSQEDLQLKD 71
Query: 412 LPHDDDPLVISVQLFNWEIKRVLIDTGSSADVLYFDAFSKMGLSEEQLQPFNGTLSGFTG 471
PH+D +VIS + + + VL+DTGS+AD+++ AF +M E+++ L GF G
Sbjct: 72 YPHND-AMVISCVIKGFLVHNVLVDTGSAADIIFAKAFRQMQEPEDKIHDATHPLCGFGG 130
Query: 472 EKVHVRGYVTLKTIFGTGDQQTSIKVRYLVINSPSSYNVIIGRPSINLLDAFVSTKHLLM 531
++ G +T+ FG + + +V + +++ YN IIGR ++N +A + +L M
Sbjct: 131 RQIVALGKITMSVTFGFINNTRTEQVVFDIVDMEYPYNAIIGRGTLNAFEAILHPAYLCM 190
Query: 532 KYPLDSGRVGVVHGDQKIARECYHASLRLKKHKEFEVQVIDLDPRE-----DFPQERIEP 586
K P D G + + HG Q+ AR R + + + ++D E F +E+
Sbjct: 191 KIPSDQGPIAI-HGSQEAAR-------RAEGNWTDSKAIHNIDGAEACEQYKFRREKAAS 242
Query: 587 IEEVKDVVI-GPLPHQITKIGTSLSGLEEEILVQLLRENVDLFAWAPSDMPGIDIGVACH 645
++ K +++ + Q +G+ LS +E+ L++ L N D+FAW+ +D+ G++ V H
Sbjct: 243 ADQPKPMLLCEDIAEQKVLLGSQLSEEQEKTLIRFLFNNKDVFAWSANDLCGVNRDVIEH 302
Query: 646 HLAVRTSVKPVVQRKRKMGEEKRKAVDEEVKKLQEAHFICEIKYPTWLANTVLVKKASGK 705
L V S +P QR RKM ++K + EVK+L A I E+KYP WLANTV+VKKA+GK
Sbjct: 303 SLNVDPSFRPRKQRLRKMSDDKAEGARNEVKRLLSAGVIREVKYPEWLANTVMVKKANGK 362
Query: 706 WRMCVDYTDLNMACPKDPYPLPSIDHLIDNASGYKTLSFMDAYSGYNQIKMDPLDAPKTA 765
WRMC+D+TDLN ACPKD +PLP ID L+D A+ + +S +D YSGY+QI M D PKT+
Sbjct: 363 WRMCIDFTDLNKACPKDEFPLPRIDSLVDAAASSELMSLLDCYSGYHQIWMKKEDEPKTS 422
Query: 766 FMTNQKNYHYRVMSFGLRNAGATFQRSMDTIFSNQIGRNLEVYIDDLVVKTSEKQSHSVD 825
F+T Y Y M GL+NAG +F R + +QIGRN+ Y+DD++VK++++++H D
Sbjct: 423 FITPSGTYCYLRMPEGLKNAGGSFSRMTAKVLQSQIGRNVLTYVDDIIVKSTKQENHIAD 482
Query: 826 LKEIFQQIRKFSMRLNPTKCTFGVQAGKFLGFLLTKKGIEANPDKCQAIINMRNPCNIRE 885
L+E F R+ ++LNP KC FGV+ GKFLG L++ KGIEANP K +AI+ M P +
Sbjct: 483 LQETFASFRQAGLKLNPEKCVFGVKKGKFLGCLVSTKGIEANPSKIEAILRMEPPTTKKG 542
Query: 886 VQQLTGRLAALSRFLSCAGDKAFAFFATIKKKEEFEWNQECDEAFGKIKQFLTTPPILHR 945
Q+LTGRLA+L+RF+S + ++ FF +K E F+W +AF ++KQ+L L
Sbjct: 543 AQRLTGRLASLNRFISRSAERNLPFFEVLKSAEVFQWGPIQQKAFEELKQYLIDLTTLTP 602
Query: 946 PTKGAGLFLYLSVSENALSSVLVEE----SDEREKPIYFVSRVLKGAELRYQKIEKLALA 1001
P GA L LY++ S +A+S+ LV+E +R+ PIYFVS VL ++ Y ++EK+ A
Sbjct: 603 PMSGAPLLLYVAASHSAVSAALVQEKLDGQVKRQAPIYFVSEVLSLSKKNYTELEKVLYA 662
Query: 1002 VIITARKLRPYFQSHKVVIRTNYPVKQILGKLDLAGRMLSWSVELSEYDIQFAPRNNIKS 1061
V++ +RKLR YFQ++ +++ ++ P+K I+ + GR+ W+ EL+E+ I++ R++I+S
Sbjct: 663 VLMASRKLRHYFQAYNIIVPSSQPLKDIMRNREATGRIGKWAAELNEFCIEYVHRSSIQS 722
Query: 1062 QVLADFVVEFTSPIQEAVPH----VWLLSVDGSSNIKGSGAGIILEGPGDLLIEQSLKFD 1117
Q LADF+ ++T QE + W + DGS G+GA +L P + I + K D
Sbjct: 723 QALADFIADWTPGAQEEETNKDNEAWTVFCDGSWGTFGAGAAAVLVSPSKVKICYAAKLD 782
Query: 1118 FKASNNQAEYEALIAGMLLAQEMGAKNLRARSDSQLMTNQISGEYQTKDQQLSKYLARVR 1177
F +NN AEYEAL+ G+ + MG + ++DSQ+++ I + KD +L KYL VR
Sbjct: 783 FNCTNNIAEYEALVLGLRKLKAMGIRRAILKTDSQVVSGHIDKSCKAKDPKLEKYLDMVR 842
Query: 1178 KLAGDFQFFEAIYVPRESNSRADLLAKLASTKKPGNNRTVIQEVISAPSTD--EKAVFEL 1235
++ F+ F +PR N ADLLAK A+ P + V E I APS + E+AV +
Sbjct: 843 RIEASFEGFSVKNIPRGQNEHADLLAKSAAQGLPLPS-DVFFETIKAPSVELLERAVLNI 901
Query: 1236 NQE-PEGWMTPLLKFLTGSFVAKNDEYAQLVRRRATKFVVIAGKLYKRGRASPLLRCLGE 1294
+ E W T ++ +L G F++ ++ Y + + RA +V+I G+LYK G +PLL+CL
Sbjct: 902 SPVFSEDWRTEIISYLQGKFLSDDEAYNKRIEARARPYVMIEGELYKHGVCAPLLKCLSR 961
Query: 1295 GETELVLLEVHEGVCGSHIGGRSLAAKLLRAGYYWPRMAHDCCEFVKKCDKCQRFSDKKN 1354
E ++ E+H G+CGSHIG R L K+ R G+YWP+ A D E V+KC+ CQ+ + +
Sbjct: 962 TEGIELMKEIHAGLCGSHIGSRPLLGKVFRQGFYWPKAASDAAELVQKCEGCQKCARDQK 1021
Query: 1355 APANELTSVFSP-WPFHKWGVDIVGPFPQAPGQLKFLIVDVDYFTKWVEAEAVSKITAER 1413
P++ LT + P WP +WG+D++GP P A G L++++V V+YF+KW+EA+ ++ IT+
Sbjct: 1022 QPSS-LTQLIQPTWPLQRWGLDLLGPLPPAQGNLRYVVVAVEYFSKWIEAKPLATITSAT 1080
Query: 1414 VVKFYWKKIICHFGLPKYIVTDNGTQFASSKVVNFCKQLGIETKFVSVIHPQANGQAESA 1473
V KF+W+ I+C FG+PK I DNGTQF S +FC Q+G + F SV HP++NG E A
Sbjct: 1081 VQKFFWQNIVCRFGVPKAITVDNGTQFDSEAFRDFCDQIGTKIHFASVRHPESNGLVERA 1140
Query: 1474 NKMIVNDIKKKL-EDAKGLWAEQLHEVLWSYHTTPHSTTGETPFTMVYGADAMLPVEIDT 1532
N +I+ I K + +G W +QL +V+WS++TT +TG TPF +++G +A+ P +
Sbjct: 1141 NGIIMTGIMKLIFNQPRGKWPDQLIKVVWSHNTTTSRSTGFTPFKLLFGDEAITPEKAKA 1200
Query: 1533 PTWRREHFSEESNEVGIRCTMDMIDEVREAA--HIREFAAKQRAARRYNSKVIPRSMKEG 1590
+ R +E +E D ++ +R A +I ++ A+ + + KV ++++ G
Sbjct: 1201 GSIRIVASAESDSEAAYSIEKDALEGIRLQAVENINKYQAE--TVKWRDRKVRLKNIEPG 1258
Query: 1591 DLVLKQVVAPTRIGKLLPSWKGPYRVKEKLQHGAYKLEELSGNPVPRT 1638
LVL++V P +GKL W GP+ V + G+Y+L++++G+ +PR+
Sbjct: 1259 HLVLRRVANPETVGKLQLKWDGPFLVASSSRPGSYRLKDMNGSDIPRS 1306
>emb|CAE02180.2| OSJNBa0080E14.11 [Oryza sativa (japonica cultivar-group)]
gi|50929995|ref|XP_474525.1| OSJNBa0080E14.11 [Oryza
sativa (japonica cultivar-group)]
Length = 2001
Score = 902 bits (2332), Expect = 0.0
Identities = 565/1646 (34%), Positives = 854/1646 (51%), Gaps = 113/1646 (6%)
Query: 32 DSHHGDLDEPEPQPFSTEIWNAPVPDNFKPPQLPTFDGRSDPSEHVTVFNTRMSVYGVAD 91
D H DLD F+ ++ P FKP + +DG ++P +TV+ + G
Sbjct: 413 DRHDDDLDGVAA--FTDDLRRVDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDS 470
Query: 92 SLKCKLLAGTLADAALRWYMSLPRFSIVGYQDLTKKFTQQFSGTRHRKVLSTSLFNVRQG 151
L LAD+A W LPR +I + +L F F GT R L+NV Q
Sbjct: 471 KAMVNYLPVALADSARSWLHGLPRGTIGSWAELRDHFIANFQGTFERPGTQFDLYNVVQK 530
Query: 152 PNESLREYLARFNDSTIKVSNPNQEVFVGAFQNGLRAGQFNESLAQKPADSMEEIIARAE 211
ESLR+Y+ RF++ K+S+ +V + AF G+R + +KP +++ + +A
Sbjct: 531 SGESLRDYIRRFSEQRNKISDITDDVIIAAFTKGIRHEELVGKFGRKPPRTVKLMFEKAN 590
Query: 212 CYVKGEES-NAEKRA----RDVKEKGSSGGERRNHYVPPNRDRGTFKKPYERNQPRYVPE 266
Y K E++ A K++ + K+ ++GG N++ +DR
Sbjct: 591 EYAKAEDAVTASKQSGPSWKPKKDTPATGGGGSNNH----KDR----------------- 629
Query: 267 HFTPLNTRPEKILKEVFESKIIPPPPFNRSKTMGQDKNAWCKYHLIEGHNTDDCVHLKRE 326
+PE+++ S +R T + N+ C ++ H DC K+
Sbjct: 630 -----KRKPEELVATAIHSS----RQRSRVNTFDKIMNSQCPHYPNSNHVAKDCFVYKQF 680
Query: 327 IEKLLLNGKLRGYAKEKHHSERREDKPNP-----EPKHTLHTISGGFAGGGESSNLRKKY 381
E+ + + +E+ S +++D P K H G A ES +K
Sbjct: 681 AEQYAKTTR-KNSDEEQSTSRKKDDGDTPVEFQDHRKELNHIFGGPLAY--ESKRKQKLT 737
Query: 382 VRQVMLLGDSPSSPKKYPHVTFS---SEDFGRVLPHDDDPLVISVQLFNWEIKRVLIDTG 438
R++ + ++ + S+ RV+ PLV+ + N +++R LID G
Sbjct: 738 EREINAVQPDTPQYLRWSEIAIKFDRSDHPDRVVHPGRYPLVLDPVVRNVKLRRTLIDGG 797
Query: 439 SSADVLYFDAFSKMGLSEEQLQPFNGTLSG-FTGEKVHVRGYVTLKTIFGTGDQQTSIKV 497
S+ ++L+ M + +L+P N G G G +TL FGT + + +
Sbjct: 798 SALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPLGQITLPVTFGTRENFRTENI 857
Query: 498 RYLVINSPSSYNVIIGRPSINLLDAFVSTKHLLMKYPLDSGRVGVVHGDQKIARECYHAS 557
+ V + ++Y+ I+GRP++ A +++MK P G + + D K A C S
Sbjct: 858 SFEVADFETAYHAILGRPALAKFMAVPHYTYMMMKMPGPRGVLSL-RSDIKQAVTCDKES 916
Query: 558 LRLKKHKEFEVQVIDLDPREDFPQERIEPIEEVKDVVIGPLPHQITKIGTSLSGLEEEIL 617
+ + +E RED E G +P TK TS SG E
Sbjct: 917 CDMAQTREMA------SAREDIRLAAATASE-------GEVP--ATK--TSKSGESEAKT 959
Query: 618 VQL---------LRENVDLFAWAPSDMPGIDIGVACHHLAVRTSVKPVVQRKRKMGEEKR 668
++ N D+FAW PSDMPGI V H L V+ KP+ QR R+ ++++
Sbjct: 960 KKIPLDPSNPTKTANNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRK 1019
Query: 669 KAVDEEVKKLQEAHFICEIKYPTWLANTVLVKKASGKWRMCVDYTDLNMACPKDPYPLPS 728
A+ EE+ KL A FI E+ +P WLAN VLV+K +G+WRMCVDYTDLN +CPKDP+ LP
Sbjct: 1020 DAIKEELTKLLAAGFIKEVHHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPR 1079
Query: 729 IDHLIDNASGYKTLSFMDAYSGYNQIKMDPLDAPKTAFMTNQKNYHYRVMSFGLRNAGAT 788
ID ++D+ +G + LSF+D YSGY+QI++ D KT+F+T Y Y M FGL+NAGAT
Sbjct: 1080 IDQVVDSTAGCELLSFLDCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMPFGLKNAGAT 1139
Query: 789 FQRSMDTIFSNQIGRNLEVYIDDLVVKTSEKQSHSVDLKEIFQQIRKFSMRLNPTKCTFG 848
+QR + FS QIGRN+E Y+DD+VVKT +K DL+E F IR F M+LNP K TFG
Sbjct: 1140 YQRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKYTFG 1199
Query: 849 VQAGKFLGFLLTKKGIEANPDKCQAIINMRNPCNIREVQQLTGRLAALSRFLSCAGDKAF 908
V +GK LGF+++ +GI+ANP+K AI+NM+ P ++VQ+LTG +AALSRF+S G++
Sbjct: 1200 VPSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGM 1259
Query: 909 AFFATIKKKEEFEWNQECDEAFGKIKQFLTTPPILHRPTKGAGLFLYLSVSENALSSVLV 968
FF +KK ++F+W E +AF K+ LT PP+L P L LY+S + +S+VLV
Sbjct: 1260 PFFKLLKKTDDFQWGPEAQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTVLV 1319
Query: 969 EESDER------EKPIYFVSRVLKGAELRYQKIEKLALAVIITARKLRPYFQSHKVVIRT 1022
E +E ++PIYFVS VL ++ RY +++KL ++IT RKL YFQ H V + T
Sbjct: 1320 VEREEEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVT 1379
Query: 1023 NYPVKQILGKLDLAGRMLSWSVELSEYDIQFAPRNNIKSQVLADFVVEFTSPIQEAVP-- 1080
++P+ IL + GR+ W++EL DI F PR +IKSQ LADF+ E+T QE P
Sbjct: 1380 SFPLGDILHNREANGRIAKWALELMSLDISFKPRISIKSQALADFIAEWTE-CQEDTPAE 1438
Query: 1081 --HVWLLSVDGSSNIKGSGAGIILEGPGDLLIEQSLKFDFKASNNQAEYEALIAGMLLAQ 1138
W + DGS + G+GAG++L P + L F AS+N AEYEAL+ G+ +A
Sbjct: 1439 KMEHWTMHFDGSKRLSGTGAGVVLISPAGERLSYVLWIHFSASHNVAEYEALLHGLRIAI 1498
Query: 1139 EMGAKNLRARSDSQLMTNQISGEYQTKDQQLSKYLARVRKLAGDFQFFEAIYVPRESNSR 1198
+G K L R DSQL+ NQ+ E+ D + Y VRKL F E +V R +N
Sbjct: 1499 SLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEA 1558
Query: 1199 ADLLAKLASTKKPGNNRTVIQEVIS-----APSTDEKAVFELNQEPEGWMTPLLKFLTGS 1253
AD LA S ++ + ++ + + +T ++ W PL++FLT
Sbjct: 1559 ADRLANFGSKREVAPSDVFVEHLYTPTVPHKDTTQVAGTHDVAMVEADWREPLIRFLTSQ 1618
Query: 1254 FVAKNDEYAQLVRRRATKFVVIAGKLYKRGRASPLLRCLGEGETELVLLEVHEGVCGSHI 1313
+ ++ + A+ + RR+ +V+ +L GR + + E T ++ + C +H
Sbjct: 1619 ELPQDKDEAERISRRSKLYVMHEAEL---GRET-----IAERHT---FWDMRKPCCRTHH 1667
Query: 1314 GGRSLAAKLLRAGYYWPRMAHDCCEFVKKCDKCQRFSDKKNAPANELTSVFSPWPFHKWG 1373
+SL AG++ P D + V+ C+ CQ F+ + + PA EL ++ WPF WG
Sbjct: 1668 CRQSLP-----AGFFLPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWG 1722
Query: 1374 VDIVGPFPQAPGQLKFLIVDVDYFTKWVEAEAVSKITAERVVKFYWKKIICHFGLPKYIV 1433
+D+VGPF +A G L V +D F+KW+EA+ V ITA+ F+ + FG+P I+
Sbjct: 1723 LDMVGPFKKAVGGYTHLFVAIDKFSKWIEAKPVVTITADNARDFFINS-VHRFGVPNRII 1781
Query: 1434 TDNGTQFASSKVVNFCKQLGIETKFVSVIHPQANGQAESANKMIVNDIKKKLEDA----K 1489
TDNG QF +FC+ GI+ + SV HP +NGQ E AN MI+ IK ++ D
Sbjct: 1782 TDNGRQFTGGVFQDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPYA 1841
Query: 1490 GLWAEQLHEVLWSYHTTPHSTTGETPFTMVYGADAMLPVEIDTPTWRREHFSEESNEVGI 1549
G W EQL VLWS TTP TG++PF +VYGA+AMLP E++ + R +F EE E
Sbjct: 1842 GKWVEQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFREERYEEDR 1901
Query: 1550 RCTMDMIDEVREAAHIREFAAKQRAARRYNSKVIPRSMKEGDLVLKQVVAPTRIGKLLPS 1609
+ ++EVREAA I+ Q R +N V R+ GDLVL+++ KL P
Sbjct: 1902 VDDLHRLEEVREAALIQSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTTRDRHKLSPL 1961
Query: 1610 WKGPYRVKEKLQHGAYKLEELSGNPV 1635
W+GP+ + + G+Y+L+ G V
Sbjct: 1962 WEGPFIISAVTRPGSYRLKREDGTLV 1987
>dbj|BAD18986.1| GAG-POL precursor [Vitis vinifera]
Length = 1027
Score = 900 bits (2326), Expect = 0.0
Identities = 456/1015 (44%), Positives = 643/1015 (62%), Gaps = 12/1015 (1%)
Query: 635 MPGIDIGVACHHLAVRTSVKPVVQRKRKMGEEKRKAVDEEVKKLQEAHFICEIKYPTWLA 694
M GI + H L V ++ +PV QR R+ ++++ + E+ KL EA FI E+ YP WLA
Sbjct: 1 MKGIHPSITSHRLNVVSTARPVRQRIRRFHPDRQRVIRNEIDKLLEAGFIREVSYPDWLA 60
Query: 695 NTVLVKKASGKWRMCVDYTDLNMACPKDPYPLPSIDHLIDNASGYKTLSFMDAYSGYNQI 754
N V+V K GKWR+CVDYT+LN ACPKD +PLP ID ++D+ SG LSF+DA+SGY+QI
Sbjct: 61 NVVVVPKKEGKWRVCVDYTNLNNACPKDSFPLPRIDQIVDSTSGQGMLSFLDAFSGYHQI 120
Query: 755 KMDPLDAPKTAFMTNQKNYHYRVMSFGLRNAGATFQRSMDTIFSNQIGRNLEVYIDDLVV 814
M P D K AF+T Y Y+VM FGL+NAGAT+QR M IF IG ++EVYIDD+VV
Sbjct: 121 PMSPDDEEKIAFITPHDLYCYKVMPFGLKNAGATYQRLMTKIFKPLIGHSVEVYIDDIVV 180
Query: 815 KTSEKQSHSVDLKEIFQQIRKFSMRLNPTKCTFGVQAGKFLGFLLTKKGIEANPDKCQAI 874
K+ ++ H + L+E+F +R++ M+LNP+KC FGV A KFLGF+++++GIE +PD+ +A+
Sbjct: 181 KSKTREQHILHLQEVFYLLRRYGMKLNPSKCAFGVSARKFLGFMVSQRGIEVSPDQVKAV 240
Query: 875 INMRNPCNIREVQQLTGRLAALSRFLSCAGDKAFAFFATIKKKEEFEWNQECDEAFGKIK 934
+ P N +E+Q+LTG+L AL RF++ D+ FF I+K W C A +IK
Sbjct: 241 METPPPRNKKELQRLTGKLVALGRFIARFIDELRPFFLAIRKAGTHGWTDNCQNALERIK 300
Query: 935 QFLTTPPILHRPTKGAGLFLYLSVSENALSSVLVE-ESDEREKPIYFVSRVLKGAELRYQ 993
+L PPIL P L++YL+VSE A+S+VL S + +KPIY+VSR L E RY
Sbjct: 301 HYLMQPPILSSPIPKEKLYMYLAVSEWAISAVLFRCPSPKEQKPIYYVSRALADVETRYS 360
Query: 994 KIEKLALAVIITARKLRPYFQSHKVVIRTNYPVKQILGKLDLAGRMLSWSVELSEYDIQF 1053
K+E ++LA+ A+KLRPYFQ+H V++ T+ P++ IL K DL GRML W++ELSE+ I+F
Sbjct: 361 KMELISLALRSAAQKLRPYFQAHPVIVLTDQPLRNILHKPDLTGRMLQWAIELSEFGIEF 420
Query: 1054 APRNNIKSQVLADFVVEFT-SPIQEAVPHV---WLLSVDGSSNIKGSGAGIILEGPGDLL 1109
PR ++K QV+ADFV+E++ P Q W L VDG+S GSG G++L+ P
Sbjct: 421 QPRLSMKGQVMADFVLEYSRKPGQHEGSRKKEWWTLRVDGASRSSGSGVGLLLQSPTGEH 480
Query: 1110 IEQSLKFDFKASNNQAEYEALIAGMLLAQEMGAKNLRARSDSQLMTNQISGEYQTKDQQL 1169
+EQ+++ F ASNN+AEYEA+++G+ LA + LR SDSQL+ + EY+ KD ++
Sbjct: 481 LEQAIRLGFSASNNEAEYEAILSGLDLALALSVSKLRIFSDSQLVVKHVQEEYEAKDARM 540
Query: 1170 SKYLARVRKLAGDFQFFEAIYVPRESNSRADLLAKLASTKKPGNNRTVIQEVISAPSTDE 1229
++YLA+VR F + + R N RAD LA +A++ + V + PS E
Sbjct: 541 ARYLAKVRNTLQQFTEWTIEKIKRADNRRADALAGIAASLSIKEAILLPIHVQTNPSVSE 600
Query: 1230 KAVFELNQEPEG----WMTPLLKFLTGSFVAKNDEYAQLVRRRATKFVVIAGKLYKRGRA 1285
++ + P+ WM + +++ + + + A VR +A +F +I G LYKR
Sbjct: 601 ISICSTTEAPQADDQEWMNDITEYIRTGTLPGDPKQAHKVRVQAARFTLIGGHLYKRSFT 660
Query: 1286 SPLLRCLGEGETELVLLEVHEGVCGSHIGGRSLAAKLLRAGYYWPRMAHDCCEFVKKCDK 1345
P LRCLG E + VL E+HEG+ G+H GGRSLA + GYYWP M + +VK+CDK
Sbjct: 661 GPYLRCLGHSEAQYVLAELHEGIYGNHSGGRSLAHRAHSQGYYWPTMKKEAAAYVKRCDK 720
Query: 1346 CQRFSDKKNAPANELTSVFSPWPFHKWGVDIVGPFPQAPGQLKFLIVDVDYFTKWVEAEA 1405
CQR++ + P+ L S+ PWPF +WG+DIV P P AP Q KFL+V DYF+KWVEAEA
Sbjct: 721 CQRYAPIPHMPSTTLKSISGPWPFAQWGMDIVRPLPTAPAQKKFLLVATDYFSKWVEAEA 780
Query: 1406 VSKITAERVVKFYWKKIICHFGLPKYIVTDNGTQFASSKVVNFCKQLGIETKFVSVIHPQ 1465
+ + V KF WK IIC FG+P+ I+ DNG QF S NFC +L I + + +PQ
Sbjct: 781 YASTKDKDVTKFVWKNIICRFGIPQTIIADNGPQFDSIAFRNFCSELNIRNSYSTPRYPQ 840
Query: 1466 ANGQAESANKMIVNDIKKKLEDAKGLWAEQLHEVLWSYHTTPHSTTGETPFTMVYGADAM 1525
+NGQAE+ NK ++ +KK+LE AKG W E+L VLW+Y TTP TG TPF + YG DA+
Sbjct: 841 SNGQAEATNKTLITALKKRLEQAKGKWVEELPGVLWAYRTTPGRPTGNTPFALAYGMDAV 900
Query: 1526 LPVEIDTPTWRREHFSEESNEVGIRCTMDMIDEVREAAHIREFAAKQRAARRYNSKVIPR 1585
+P+EI PT + + + +D DEVRE+A IR +QRA+ YN KV PR
Sbjct: 901 IPIEIGLPTIWTNAAKQSDANMQLGRNLDWTDEVRESASIRMADYQQRASAHYNRKVRPR 960
Query: 1586 SMKEGDLVLKQVVAPTR---IGKLLPSWKGPYRVKEKLQHGAYKLEELSGNPVPR 1637
S+K G LVL++ T GK +W+GPY V + +GAY L++L G P+ R
Sbjct: 961 SLKNGTLVLRKFFENTTEVGAGKFQANWEGPYIVSKASDNGAYHLQKLDGTPLLR 1015
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.318 0.135 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,843,419,497
Number of Sequences: 2540612
Number of extensions: 126855210
Number of successful extensions: 374506
Number of sequences better than 10.0: 36669
Number of HSP's better than 10.0 without gapping: 4374
Number of HSP's successfully gapped in prelim test: 32296
Number of HSP's that attempted gapping in prelim test: 346958
Number of HSP's gapped (non-prelim): 43916
length of query: 1638
length of database: 863,360,394
effective HSP length: 142
effective length of query: 1496
effective length of database: 502,593,490
effective search space: 751879861040
effective search space used: 751879861040
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 82 (36.2 bits)
Medicago: description of AC141323.18


