

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140913.9 - phase: 2 /pseudo
(273 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
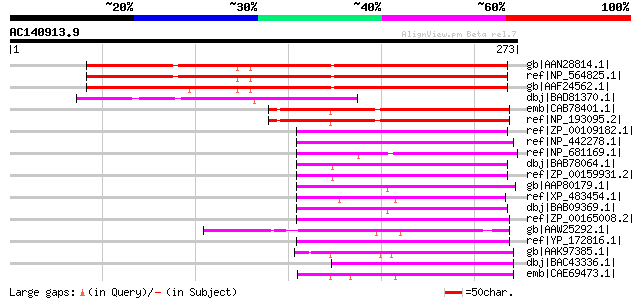
Score E
Sequences producing significant alignments: (bits) Value
gb|AAN28814.1| At1g64150/F22C12_10 [Arabidopsis thaliana] gi|150... 246 6e-64
ref|NP_564825.1| expressed protein [Arabidopsis thaliana] 246 6e-64
gb|AAF24562.1| F22C12.9 [Arabidopsis thaliana] 242 7e-63
dbj|BAD81370.1| hypothetical protein [Oryza sativa (japonica cul... 96 1e-18
emb|CAB78401.1| putative protein [Arabidopsis thaliana] gi|46783... 94 5e-18
ref|NP_193095.2| expressed protein [Arabidopsis thaliana] 94 5e-18
ref|ZP_00109182.1| COG2119: Predicted membrane protein [Nostoc p... 83 9e-15
ref|NP_442278.1| transmembrane protein FT27 [Synechocystis sp. P... 78 3e-13
ref|NP_681169.1| hypothetical protein tlr0379 [Thermosynechococc... 77 4e-13
dbj|BAB78064.1| alr1698 [Nostoc sp. PCC 7120] gi|25531406|pir||A... 76 9e-13
ref|ZP_00159931.2| COG2119: Predicted membrane protein [Anabaena... 76 9e-13
gb|AAP80179.1| At5g36290 [Arabidopsis thaliana] gi|21537321|gb|A... 72 2e-11
ref|XP_483454.1| putative transmembrane protein(TPA regulated lo... 72 2e-11
dbj|BAB09369.1| transmembrane protein FT27/PFT27-like [Arabidops... 71 3e-11
ref|ZP_00165008.2| COG2119: Predicted membrane protein [Synechoc... 69 2e-10
gb|AAW25292.1| unknown [Schistosoma japonicum] 68 3e-10
ref|YP_172816.1| hypothetical protein syc2106_d [Synechococcus e... 67 4e-10
gb|AAK97385.1| putative membrane protein [Crithidia fasciculata] 65 2e-09
dbj|BAC43336.1| putative transmembrane protein [Arabidopsis thal... 63 1e-08
emb|CAE69473.1| Hypothetical protein CBG15669 [Caenorhabditis br... 62 1e-08
>gb|AAN28814.1| At1g64150/F22C12_10 [Arabidopsis thaliana]
gi|15010676|gb|AAK73997.1| At1g64150/F22C12_10
[Arabidopsis thaliana]
Length = 370
Score = 246 bits (627), Expect = 6e-64
Identities = 148/237 (62%), Positives = 168/237 (70%), Gaps = 13/237 (5%)
Query: 42 VLKFMLFSAFFALQDAFPAVAASDFATGLNS-IPIFGDVGDLSTGFASYHRHFC*YFSLN 100
VL F+ S AL PA AAS S + FGD+GD+S+GFAS +FS
Sbjct: 114 VLMFLAVSGSVALLGTDPAFAASSIPNVTQSLVTSFGDLGDISSGFAS--AFLLIFFSEL 171
Query: 101 WETRLFSLQHC*QLEIQPVLF-----SLGHLAH----LRRTFHYVDELLPFRFGETDLPI 151
+ F +F +LG + L RTFHYVDE+LPFRFG TDLPI
Sbjct: 172 GDKTFFIAALLAARNSAATVFVGTFGALGIMTIISVVLGRTFHYVDEVLPFRFGGTDLPI 231
Query: 152 DDIAAVCLLVYFGVSTLLDASSSDSQKSDDEQKEAELAVSDFSGDGAGILAAASTIVSTF 211
DDIAAVCLLVYFGVSTLLDA S D K+D+EQKEAELAVS+ SG+GAGI+AAA+TI+STF
Sbjct: 232 DDIAAVCLLVYFGVSTLLDAVS-DEGKADEEQKEAELAVSELSGNGAGIVAAANTIISTF 290
Query: 212 LLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEK 268
LVFVAEWGDKSFFSTIALAAASSPLGVIAG+LAGHG ATL+AVLGGSLLG FLSEK
Sbjct: 291 ALVFVAEWGDKSFFSTIALAAASSPLGVIAGALAGHGAATLLAVLGGSLLGNFLSEK 347
>ref|NP_564825.1| expressed protein [Arabidopsis thaliana]
Length = 370
Score = 246 bits (627), Expect = 6e-64
Identities = 148/237 (62%), Positives = 168/237 (70%), Gaps = 13/237 (5%)
Query: 42 VLKFMLFSAFFALQDAFPAVAASDFATGLNS-IPIFGDVGDLSTGFASYHRHFC*YFSLN 100
VL F+ S AL PA AAS S + FGD+GD+S+GFAS +FS
Sbjct: 114 VLMFLAVSGSVALLGTDPAFAASSIPNVTQSLVTSFGDLGDISSGFAS--AFLLIFFSEL 171
Query: 101 WETRLFSLQHC*QLEIQPVLF-----SLGHLAH----LRRTFHYVDELLPFRFGETDLPI 151
+ F +F +LG + L RTFHYVDE+LPFRFG TDLPI
Sbjct: 172 GDKTFFIAALLAARNSAATVFVGTFGALGIMTIISVVLGRTFHYVDEVLPFRFGGTDLPI 231
Query: 152 DDIAAVCLLVYFGVSTLLDASSSDSQKSDDEQKEAELAVSDFSGDGAGILAAASTIVSTF 211
DDIAAVCLLVYFGVSTLLDA S D K+D+EQKEAELAVS+ SG+GAGI+AAA+TI+STF
Sbjct: 232 DDIAAVCLLVYFGVSTLLDAVS-DEGKADEEQKEAELAVSELSGNGAGIVAAANTIISTF 290
Query: 212 LLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEK 268
LVFVAEWGDKSFFSTIALAAASSPLGVIAG+LAGHG ATL+AVLGGSLLG FLSEK
Sbjct: 291 ALVFVAEWGDKSFFSTIALAAASSPLGVIAGALAGHGAATLLAVLGGSLLGNFLSEK 347
>gb|AAF24562.1| F22C12.9 [Arabidopsis thaliana]
Length = 388
Score = 242 bits (618), Expect = 7e-63
Identities = 150/253 (59%), Positives = 170/253 (66%), Gaps = 27/253 (10%)
Query: 42 VLKFMLFSAFFALQDAFPAVAASDFATGLNS-IPIFGDVGDLSTGFASYHRH-FC*---- 95
VL F+ S AL PA AAS S + FGD+GD+S+GFAS FC
Sbjct: 114 VLMFLAVSGSVALLGTDPAFAASSIPNVTQSLVTSFGDLGDISSGFASVRESSFCTEPGT 173
Query: 96 -----------YFSLNWETRLFSLQHC*QLEIQPVLF-----SLGHLAH----LRRTFHY 135
+FS + F +F +LG + L RTFHY
Sbjct: 174 ISSSIPAFLLIFFSELGDKTFFIAALLAARNSAATVFVGTFGALGIMTIISVVLGRTFHY 233
Query: 136 VDELLPFRFGETDLPIDDIAAVCLLVYFGVSTLLDASSSDSQKSDDEQKEAELAVSDFSG 195
VDE+LPFRFG TDLPIDDIAAVCLLVYFGVSTLLDA S D K+D+EQKEAELAVS+ SG
Sbjct: 234 VDEVLPFRFGGTDLPIDDIAAVCLLVYFGVSTLLDAVS-DEGKADEEQKEAELAVSELSG 292
Query: 196 DGAGILAAASTIVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAV 255
+GAGI+AAA+TI+STF LVFVAEWGDKSFFSTIALAAASSPLGVIAG+LAGHG ATL+AV
Sbjct: 293 NGAGIVAAANTIISTFALVFVAEWGDKSFFSTIALAAASSPLGVIAGALAGHGAATLLAV 352
Query: 256 LGGSLLGTFLSEK 268
LGGSLLG FLSEK
Sbjct: 353 LGGSLLGNFLSEK 365
>dbj|BAD81370.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|56784056|dbj|BAD81293.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 266
Score = 95.5 bits (236), Expect = 1e-18
Identities = 60/161 (37%), Positives = 82/161 (50%), Gaps = 16/161 (9%)
Query: 37 DASMTVLKFMLFSAFFALQDAFPAVAASDFATGLNSIPIFGDVGDLSTGFASYHRHFC*Y 96
DAS L + LQ + A+A ++F + + GD+GD+STGFAS F
Sbjct: 82 DASSCGLALAAAAGVLMLQGSQQALAGTEF---MGMQDVVGDLGDISTGFASA---FLLI 135
Query: 97 FSLNWETRLFSLQHC*QLEIQPVLFSLGHLAHLR----------RTFHYVDELLPFRFGE 146
F R F + + LG L R FHYVD ++PF FG
Sbjct: 136 FFSELGDRTFFIAALLAARNSGAIIFLGTFGALAVMTIISVVLGRAFHYVDGIIPFSFGG 195
Query: 147 TDLPIDDIAAVCLLVYFGVSTLLDASSSDSQKSDDEQKEAE 187
TD P+DD A CLLVY+G++TLLDA+S D +K ++EQ+E E
Sbjct: 196 TDFPVDDFLAACLLVYYGITTLLDAASGDEEKMNEEQEEVE 236
Score = 43.1 bits (100), Expect = 0.008
Identities = 29/74 (39%), Positives = 41/74 (55%), Gaps = 3/74 (4%)
Query: 188 LAVSDFSG--DGAGILAAAST-IVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSL 244
LA ++F G D G L ST S FLL+F +E GD++FF LAA +S + G+
Sbjct: 106 LAGTEFMGMQDVVGDLGDISTGFASAFLLIFFSELGDRTFFIAALLAARNSGAIIFLGTF 165
Query: 245 AGHGVATLIAVLGG 258
V T+I+V+ G
Sbjct: 166 GALAVMTIISVVLG 179
>emb|CAB78401.1| putative protein [Arabidopsis thaliana] gi|4678385|emb|CAB41117.1|
putative protein [Arabidopsis thaliana]
gi|7487743|pir||T06661 hypothetical protein T6G15.140 -
Arabidopsis thaliana
Length = 273
Score = 93.6 bits (231), Expect = 5e-18
Identities = 53/139 (38%), Positives = 85/139 (61%), Gaps = 12/139 (8%)
Query: 140 LPFRFGETDLPIDDIAAVCLLVYFGVSTLLDA---------SSSDSQKSDDEQKEAELAV 190
+P +F +T LPI + AA+ LL++FG+ ++ DA + ++ E EAE V
Sbjct: 137 VPAQF-QTTLPIGEYAAIALLMFFGLKSIKDAWDLPPVEAKNGEETGIELGEYSEAEELV 195
Query: 191 SDFSGDGAGILAAASTIVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVA 250
+ + + + +F LVF AEWGD+S +T+AL AA SPLGV +G++AGH VA
Sbjct: 196 KEKASKK--LTNPLEILWKSFSLVFFAEWGDRSMLATVALGAAQSPLGVASGAIAGHLVA 253
Query: 251 TLIAVLGGSLLGTFLSEKV 269
T++A++GG+ L ++SEK+
Sbjct: 254 TVLAIMGGAFLANYISEKL 272
Score = 39.3 bits (90), Expect = 0.12
Identities = 23/68 (33%), Positives = 37/68 (53%), Gaps = 2/68 (2%)
Query: 195 GDGAGILAAA--STIVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATL 252
G + +LAA S + F L+FV+E GDK+FF LA V+ GS+ + T+
Sbjct: 66 GGPSSVLAAVAKSGFTAAFSLIFVSEIGDKTFFIAALLAMQYEKTLVLLGSMGALSLMTI 125
Query: 253 IAVLGGSL 260
++V+ G +
Sbjct: 126 LSVVIGKI 133
>ref|NP_193095.2| expressed protein [Arabidopsis thaliana]
Length = 359
Score = 93.6 bits (231), Expect = 5e-18
Identities = 53/139 (38%), Positives = 85/139 (61%), Gaps = 12/139 (8%)
Query: 140 LPFRFGETDLPIDDIAAVCLLVYFGVSTLLDA---------SSSDSQKSDDEQKEAELAV 190
+P +F +T LPI + AA+ LL++FG+ ++ DA + ++ E EAE V
Sbjct: 203 VPAQF-QTTLPIGEYAAIALLMFFGLKSIKDAWDLPPVEAKNGEETGIELGEYSEAEELV 261
Query: 191 SDFSGDGAGILAAASTIVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVA 250
+ + + + +F LVF AEWGD+S +T+AL AA SPLGV +G++AGH VA
Sbjct: 262 KEKASKK--LTNPLEILWKSFSLVFFAEWGDRSMLATVALGAAQSPLGVASGAIAGHLVA 319
Query: 251 TLIAVLGGSLLGTFLSEKV 269
T++A++GG+ L ++SEK+
Sbjct: 320 TVLAIMGGAFLANYISEKL 338
Score = 39.3 bits (90), Expect = 0.12
Identities = 23/68 (33%), Positives = 37/68 (53%), Gaps = 2/68 (2%)
Query: 195 GDGAGILAAA--STIVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATL 252
G + +LAA S + F L+FV+E GDK+FF LA V+ GS+ + T+
Sbjct: 132 GGPSSVLAAVAKSGFTAAFSLIFVSEIGDKTFFIAALLAMQYEKTLVLLGSMGALSLMTI 191
Query: 253 IAVLGGSL 260
++V+ G +
Sbjct: 192 LSVVIGKI 199
>ref|ZP_00109182.1| COG2119: Predicted membrane protein [Nostoc punctiforme PCC 73102]
Length = 206
Score = 82.8 bits (203), Expect = 9e-15
Identities = 43/115 (37%), Positives = 68/115 (58%), Gaps = 1/115 (0%)
Query: 155 AAVCLLVYFGVSTLLDASSSDSQKSDDEQ-KEAELAVSDFSGDGAGILAAASTIVSTFLL 213
A + L + FG+ L DAS S D E +EAE AV + + + ++ F+L
Sbjct: 70 AEIVLFLAFGIKLLYDASKMSSAACDTEVIEEAEAAVKKADLELPKKKTSLAIVIEAFIL 129
Query: 214 VFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEK 268
F+AEWGD++ +TIALAA ++P+GV G++ GH + IAV+GG ++ +SE+
Sbjct: 130 TFMAEWGDRTQIATIALAAGNNPIGVTVGAILGHTICAAIAVIGGKMIAGRISER 184
>ref|NP_442278.1| transmembrane protein FT27 [Synechocystis sp. PCC 6803]
gi|1001617|dbj|BAA10348.1| transmembrane protein FT27
[Synechocystis sp. PCC 6803]
gi|1723176|sp|P52876|Y615_SYNY3 Hypothetical UPF0016
protein sll0615 gi|1256592|gb|AAA96398.1| similar to Mus
musculus transmembrane protein (clone pFT27); Method:
conceptual translation supplied by author; ORF206
Length = 206
Score = 77.8 bits (190), Expect = 3e-13
Identities = 42/117 (35%), Positives = 67/117 (56%)
Query: 155 AAVCLLVYFGVSTLLDASSSDSQKSDDEQKEAELAVSDFSGDGAGILAAASTIVSTFLLV 214
A V L + FG L DA + + +E ++AE A++ + +V +F L
Sbjct: 70 AEVALFLIFGTKLLWDARRIKATANLEEMEDAEKAIASGEKKLKIVPRGWGIVVESFALT 129
Query: 215 FVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEKVTS 271
FVAEWGD++ +TIALAA+++ GV AG++ GH + +IAV+GG + +SEK +
Sbjct: 130 FVAEWGDRTQIATIALAASNNAWGVSAGAILGHTICAVIAVMGGKFVAGRISEKTVT 186
Score = 35.8 bits (81), Expect = 1.3
Identities = 21/54 (38%), Positives = 31/54 (56%), Gaps = 1/54 (1%)
Query: 212 LLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFL 265
LL+ V+E GDK+FF + LA V+ G + G T+++VL G + TFL
Sbjct: 10 LLITVSELGDKTFFIAMILAMRYPRRWVLVGVVGGLAAMTILSVLMGQIF-TFL 62
>ref|NP_681169.1| hypothetical protein tlr0379 [Thermosynechococcus elongatus BP-1]
gi|22294100|dbj|BAC07931.1| tlr0379 [Thermosynechococcus
elongatus BP-1]
Length = 211
Score = 77.4 bits (189), Expect = 4e-13
Identities = 50/124 (40%), Positives = 71/124 (56%), Gaps = 7/124 (5%)
Query: 155 AAVCLLVYFGVSTLLDASSSDSQKSDDEQKEA----ELAVSDFSGDGAGILAAASTIVST 210
AA+ L FG+ L+ ++ +DE + A E A ++ S G+ AA V
Sbjct: 70 AAILLFTIFGLRMLIQGWRMGNKPCEDECEAAVETVEKAEANLSRWGSNPAWAA--FVEA 127
Query: 211 FLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEK-V 269
F L +AEWGD++ +TI LAAAS GV G++AGHG+ T IAVLGG L+ +SE+ +
Sbjct: 128 FSLTLMAEWGDRTQIATITLAAASQAFGVALGAIAGHGICTAIAVLGGGLIAGRISERTL 187
Query: 270 TSSG 273
T SG
Sbjct: 188 TLSG 191
>dbj|BAB78064.1| alr1698 [Nostoc sp. PCC 7120] gi|25531406|pir||AD2018 hypothetical
protein alr1698 [imported] - Nostoc sp. (strain PCC
7120) gi|17229190|ref|NP_485738.1| hypothetical protein
alr1698 [Nostoc sp. PCC 7120]
Length = 209
Score = 76.3 bits (186), Expect = 9e-13
Identities = 42/118 (35%), Positives = 68/118 (57%), Gaps = 4/118 (3%)
Query: 155 AAVCLLVYFGVSTLLDAS----SSDSQKSDDEQKEAELAVSDFSGDGAGILAAASTIVST 210
A + L + FG+ L DAS ++D + +E +EA+ AV S I+
Sbjct: 70 AEITLFIAFGLKLLYDASKMSAAADKAEVMEEMEEAKAAVEKADLQLPKQKTPLSIILEA 129
Query: 211 FLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEK 268
F+L F+AEWGD++ +TIALAA ++ +GV G++ GH + IAV+GG ++ +SE+
Sbjct: 130 FVLTFMAEWGDRTQIATIALAAGNNIIGVTIGAILGHAICAAIAVIGGKMIAGKISER 187
>ref|ZP_00159931.2| COG2119: Predicted membrane protein [Anabaena variabilis ATCC
29413]
Length = 233
Score = 76.3 bits (186), Expect = 9e-13
Identities = 42/118 (35%), Positives = 68/118 (57%), Gaps = 4/118 (3%)
Query: 155 AAVCLLVYFGVSTLLDAS----SSDSQKSDDEQKEAELAVSDFSGDGAGILAAASTIVST 210
A + L + FG+ L DAS ++D + +E +EA+ AV S I+
Sbjct: 94 AEITLFIAFGLKLLYDASKMSAAADKAEVMEEMEEAKAAVEKADLQLPKQKTPLSIILEA 153
Query: 211 FLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEK 268
F+L F+AEWGD++ +TIALAA ++ +GV G++ GH + IAV+GG ++ +SE+
Sbjct: 154 FVLTFMAEWGDRTQIATIALAAGNNIIGVTIGAILGHAICAAIAVIGGKMIAGKISER 211
>gb|AAP80179.1| At5g36290 [Arabidopsis thaliana] gi|21537321|gb|AAM61662.1|
transmembrane protein FT27/PFT27-like [Arabidopsis
thaliana] gi|18421551|ref|NP_568535.1| expressed protein
[Arabidopsis thaliana] gi|30692937|ref|NP_851098.1|
expressed protein [Arabidopsis thaliana]
gi|15450794|gb|AAK96668.1| transmembrane protein
FT27/PFT27-like [Arabidopsis thaliana]
Length = 293
Score = 72.0 bits (175), Expect = 2e-11
Identities = 42/124 (33%), Positives = 67/124 (53%), Gaps = 6/124 (4%)
Query: 155 AAVCLLVYFGVSTLLDASSSDSQKSDDEQKEAELAVSDFSGDGAGILA------AASTIV 208
AA L +FG+ L A S KS+ +++ E+ SG G +
Sbjct: 151 AATVLYAFFGLRLLYIAWRSTDSKSNQKKEMEEVEEKLESGQGKTPFRRLFSRFCTPIFL 210
Query: 209 STFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEK 268
+F+L F+AEWGD+S +TIALA + +GV G+ GH V T +AV+GGS+L + +S++
Sbjct: 211 ESFILTFLAEWGDRSQIATIALATHKNAIGVAIGASIGHTVCTSLAVVGGSMLASRISQR 270
Query: 269 VTSS 272
++
Sbjct: 271 TVAT 274
Score = 34.3 bits (77), Expect = 3.9
Identities = 18/66 (27%), Positives = 36/66 (54%)
Query: 207 IVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLS 266
+ S+F ++ V E GD++F +A V++G+L+ V T+++ G ++ +S
Sbjct: 85 LFSSFSMILVTEIGDETFIIAALMAMRHPKATVLSGALSALFVMTILSTGLGRIVPNLIS 144
Query: 267 EKVTSS 272
K T+S
Sbjct: 145 RKHTNS 150
>ref|XP_483454.1| putative transmembrane protein(TPA regulated locus protein) [Oryza
sativa (japonica cultivar-group)]
gi|42407963|dbj|BAD09101.1| putative transmembrane
protein(TPA regulated locus protein) [Oryza sativa
(japonica cultivar-group)]
Length = 282
Score = 71.6 bits (174), Expect = 2e-11
Identities = 44/118 (37%), Positives = 65/118 (54%), Gaps = 5/118 (4%)
Query: 155 AAVCLLVYFGVSTLLDASSSDS---QKSDDEQKEAELAVSDFSGDGAGILAAAST--IVS 209
AA L +FG+ L A SDS QK + E+ E +L I + T +
Sbjct: 141 AATVLYAFFGLRLLYIAWRSDSKASQKKEIEEVEEKLEAGQGKSTFRRIFSRFCTPIFLE 200
Query: 210 TFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSE 267
+F+L F+AEWGD+S +TIALA + +GV G+ GH + T AV+GGS+L + +S+
Sbjct: 201 SFVLTFLAEWGDRSQIATIALATHKNAVGVAVGATLGHTICTSFAVVGGSMLASKISQ 258
>dbj|BAB09369.1| transmembrane protein FT27/PFT27-like [Arabidopsis thaliana]
Length = 325
Score = 71.2 bits (173), Expect = 3e-11
Identities = 42/120 (35%), Positives = 65/120 (54%), Gaps = 6/120 (5%)
Query: 155 AAVCLLVYFGVSTLLDASSSDSQKSDDEQKEAELAVSDFSGDGAGILA------AASTIV 208
AA L +FG+ L A S KS+ +++ E+ SG G +
Sbjct: 151 AATVLYAFFGLRLLYIAWRSTDSKSNQKKEMEEVEEKLESGQGKTPFRRLFSRFCTPIFL 210
Query: 209 STFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEK 268
+F+L F+AEWGD+S +TIALA + +GV G+ GH V T +AV+GGS+L + +S++
Sbjct: 211 ESFILTFLAEWGDRSQIATIALATHKNAIGVAIGASIGHTVCTSLAVVGGSMLASRISQR 270
Score = 34.3 bits (77), Expect = 3.9
Identities = 18/66 (27%), Positives = 36/66 (54%)
Query: 207 IVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLS 266
+ S+F ++ V E GD++F +A V++G+L+ V T+++ G ++ +S
Sbjct: 85 LFSSFSMILVTEIGDETFIIAALMAMRHPKATVLSGALSALFVMTILSTGLGRIVPNLIS 144
Query: 267 EKVTSS 272
K T+S
Sbjct: 145 RKHTNS 150
>ref|ZP_00165008.2| COG2119: Predicted membrane protein [Synechococcus elongatus PCC
7942]
Length = 220
Score = 68.6 bits (166), Expect = 2e-10
Identities = 39/115 (33%), Positives = 59/115 (50%)
Query: 155 AAVCLLVYFGVSTLLDASSSDSQKSDDEQKEAELAVSDFSGDGAGILAAASTIVSTFLLV 214
A V FG+ D+ E++EAE V + + ++ F LV
Sbjct: 83 AEVAFFAIFGLKLWRDSLGMPQVGDSAEEEEAEELVLGAEAKLGKQVTVFTVVLEAFSLV 142
Query: 215 FVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEKV 269
FVAEWGD++ F+T+ALAAA + GV G++ GH + +IAV G + +SE+V
Sbjct: 143 FVAEWGDRTQFTTMALAAAGNAWGVALGAILGHAIVAVIAVNVGRWVSRHISERV 197
>gb|AAW25292.1| unknown [Schistosoma japonicum]
Length = 261
Score = 67.8 bits (164), Expect = 3e-10
Identities = 54/185 (29%), Positives = 87/185 (46%), Gaps = 31/185 (16%)
Query: 105 LFSLQHC*QLEIQPVLFSLGHLAHLRRTFHYVDELLPFRFGETDLPIDDIAAVCLLVYFG 164
+ S+QH L +F+L + L Y ++P RF L + L + FG
Sbjct: 59 IMSMQHPRALVYCGAMFALITMTMLSALLGYATTIVP-RFVTLYL------SGVLFLIFG 111
Query: 165 VSTLLDASSSDSQKSDDEQKEAELAVSDF-SGD---GAGILAAASTIVS----------- 209
+ L +A + S + DE E + ++ SGD G + +++S
Sbjct: 112 IKMLYEAYTMSSSSAKDEFDEVHMQITQSKSGDIETGTSVPETPRSLISKPILIIKKILT 171
Query: 210 -----TFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTF 264
F+L F+AEWGD+S +TI LAA S LGVI G + GH + T +AV L+G F
Sbjct: 172 PIFVEAFVLTFLAEWGDRSQITTIVLAATKSALGVIVGGVLGHALCTGLAV----LMGRF 227
Query: 265 LSEKV 269
+++++
Sbjct: 228 VAQRI 232
>ref|YP_172816.1| hypothetical protein syc2106_d [Synechococcus elongatus PCC 6301]
gi|56687074|dbj|BAD80296.1| hypothetical protein
[Synechococcus elongatus PCC 6301]
Length = 220
Score = 67.4 bits (163), Expect = 4e-10
Identities = 39/115 (33%), Positives = 58/115 (49%)
Query: 155 AAVCLLVYFGVSTLLDASSSDSQKSDDEQKEAELAVSDFSGDGAGILAAASTIVSTFLLV 214
A V FG+ D+ E++EAE V + + + F LV
Sbjct: 83 AEVAFFAIFGLKLWRDSLGMPQVGDSAEEEEAEELVLGAEAKLGKQVTVFTVVSEAFSLV 142
Query: 215 FVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEKV 269
FVAEWGD++ F+T+ALAAA + GV G++ GH + +IAV G + +SE+V
Sbjct: 143 FVAEWGDRTQFTTMALAAAGNAWGVALGAILGHAIVAVIAVNVGRWVSRHISERV 197
>gb|AAK97385.1| putative membrane protein [Crithidia fasciculata]
Length = 259
Score = 65.1 bits (157), Expect = 2e-09
Identities = 49/139 (35%), Positives = 71/139 (50%), Gaps = 22/139 (15%)
Query: 154 IAAVCLLVYFGVSTLLDA---SSSDSQKSDDEQKEAELAVSDFSGDGA---GILAAA--- 204
+AAV LV FG L D + ++ ++S+DE EA A+ + A G +A++
Sbjct: 79 LAAVLFLV-FGGKILFDELVRNKAEDEESEDEMAEAAAALRRRDPNDAVETGSVASSVYT 137
Query: 205 ------------STIVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATL 252
+V F L FVAEWGD+S +TIALAAA +P V G + GH + T
Sbjct: 138 SAPARRWRRLLNPVMVEAFTLTFVAEWGDRSQLATIALAAAKNPYAVTVGGVLGHALCTG 197
Query: 253 IAVLGGSLLGTFLSEKVTS 271
AVL G+L+ +S K +
Sbjct: 198 GAVLCGNLIAQRVSMKTVN 216
Score = 34.7 bits (78), Expect = 3.0
Identities = 18/63 (28%), Positives = 35/63 (54%)
Query: 208 VSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSE 267
+S+ ++ V+E GDK+FF +A L V G+++ T+++ L G ++ + LS
Sbjct: 14 LSSLSMILVSEIGDKTFFIACLMAMRHPRLTVYLGAISALAAMTVLSALMGVIVPSLLSV 73
Query: 268 KVT 270
+T
Sbjct: 74 YLT 76
>dbj|BAC43336.1| putative transmembrane protein [Arabidopsis thaliana]
gi|15221462|ref|NP_177032.1| expressed protein
[Arabidopsis thaliana] gi|5734713|gb|AAD49978.1| Is a
member of PF|01169 Uncharacterized (transmembrane
domain) protein family. [Arabidopsis thaliana]
gi|25353311|pir||A96711 hypothetical protein F24J5.11
[imported] - Arabidopsis thaliana
Length = 228
Score = 62.8 bits (151), Expect = 1e-08
Identities = 35/99 (35%), Positives = 55/99 (55%), Gaps = 1/99 (1%)
Query: 174 SDSQKSDDEQKEAELAVSDFSGDGAGILAAASTI-VSTFLLVFVAEWGDKSFFSTIALAA 232
SD +K++D+ K +++ + A S I + F + F EWGDKS +TI LAA
Sbjct: 109 SDLKKTNDQSKNSKIEDEQKKQKRPFLTAFFSPIFLKAFSINFFGEWGDKSQLATIGLAA 168
Query: 233 ASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEKVTS 271
+PLGV+ G + + T AVLGG L + +SE++ +
Sbjct: 169 DENPLGVVLGGIVAQTLCTTAAVLGGKSLASQISERIVA 207
>emb|CAE69473.1| Hypothetical protein CBG15669 [Caenorhabditis briggsae]
Length = 268
Score = 62.4 bits (150), Expect = 1e-08
Identities = 42/136 (30%), Positives = 66/136 (47%), Gaps = 20/136 (14%)
Query: 156 AVCLLVYFGVSTLLDA---SSSDSQKSDDE------QKEAELAVSDFSG-DGAGILAAAS 205
+ L FG+ L + S ++ Q+ +E ++E EL S F +G G+ +
Sbjct: 107 STALFALFGLKMLHEGWTMSPNEGQEGFEEAQAEVAKREGELDASKFEMLEGGGVAPQSE 166
Query: 206 T----------IVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAV 255
T + F L FVAEWGD+S +TI L A + GVI G + GH + T IAV
Sbjct: 167 TKKIFLFTSRIFIEAFTLTFVAEWGDRSQLTTIILGARENIAGVIGGGVLGHALCTGIAV 226
Query: 256 LGGSLLGTFLSEKVTS 271
+GG ++ +S + +
Sbjct: 227 IGGKIVAQRISVRTVT 242
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.325 0.138 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 417,430,186
Number of Sequences: 2540612
Number of extensions: 16082223
Number of successful extensions: 77429
Number of sequences better than 10.0: 159
Number of HSP's better than 10.0 without gapping: 86
Number of HSP's successfully gapped in prelim test: 73
Number of HSP's that attempted gapping in prelim test: 77193
Number of HSP's gapped (non-prelim): 264
length of query: 273
length of database: 863,360,394
effective HSP length: 126
effective length of query: 147
effective length of database: 543,243,282
effective search space: 79856762454
effective search space used: 79856762454
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 74 (33.1 bits)
Medicago: description of AC140913.9


