

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140034.3 + phase: 0
(199 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
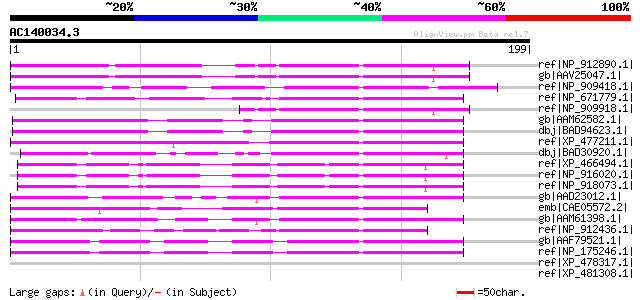
Sequences producing significant alignments: (bits) Value
ref|NP_912890.1| unnamed protein product [Oryza sativa (japonica... 66 6e-10
gb|AAV25047.1| putative polyprotein [Oryza sativa (japonica cult... 66 6e-10
ref|NP_909418.1| P0701D05.29 [Oryza sativa (japonica cultivar-gr... 63 6e-09
ref|NP_671779.1| expressed protein [Arabidopsis thaliana] 62 9e-09
ref|NP_909918.1| putative mutator-like transposase [Oryza sativa... 56 7e-07
gb|AAM62582.1| unknown [Arabidopsis thaliana] 56 7e-07
dbj|BAD94623.1| hypothetical protein [Arabidopsis thaliana] gi|1... 56 7e-07
ref|XP_477211.1| calcineurin-like phosphoesterase-like [Oryza sa... 54 3e-06
dbj|BAD30920.1| serine/threonine phosphatase PP7-related-like [O... 53 6e-06
ref|XP_466494.1| calcineurin-like phosphoesterase-like protein [... 52 1e-05
ref|NP_916020.1| OSJNBb0021A09.19 [Oryza sativa (japonica cultiv... 52 1e-05
ref|NP_918073.1| P0702H08.15 [Oryza sativa (japonica cultivar-gr... 52 1e-05
gb|AAD23012.1| expressed protein [Arabidopsis thaliana] gi|18400... 52 1e-05
emb|CAE05572.2| OSJNBb0013O03.13 [Oryza sativa (japonica cultiva... 50 4e-05
gb|AAM61398.1| unknown [Arabidopsis thaliana] 50 4e-05
ref|NP_912436.1| Putative transposable element [Oryza sativa (ja... 49 8e-05
gb|AAF79521.1| F21D18.16 [Arabidopsis thaliana] gi|25405918|pir|... 47 3e-04
ref|NP_175246.1| calcineurin-like phosphoesterase family protein... 47 3e-04
ref|XP_478317.1| serine/threonine phosphatase PP7 -related-like ... 43 0.005
ref|XP_481308.1| calcineurin-like phosphoesterase-like [Oryza sa... 41 0.017
>ref|NP_912890.1| unnamed protein product [Oryza sativa (japonica cultivar-group)]
Length = 1230
Score = 65.9 bits (159), Expect = 6e-10
Identities = 56/178 (31%), Positives = 81/178 (45%), Gaps = 19/178 (10%)
Query: 1 MPMGEMTVTLDDFACLTHLPIEGRMLDYEKKMPKHEGAALLMTYLGVSQHEAEKICNQEY 60
+P GE+TVTL+D A + LPI G+ + + + YLG+ A Q
Sbjct: 592 LPCGELTVTLEDVAMILGLPIRGQAVTGDTASGNWRER--VEEYLGLEPPVAPDGQRQTK 649
Query: 61 GGYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRWYLMYLVGCLLFDDRS 120
+ LR + + P E +E A V+ YC R Y++Y+ G +LF D
Sbjct: 650 TSGVPLSWLRANFG------------QCPAEADE-ATVQRYC-RAYVLYIFGSILFPDSG 695
Query: 121 NKRIELIYLTTMADGYAGMRNYSWGGMTLAYLYGELADACR--PGHRALGGSVTLLTV 176
++L +AD + +SWG LA+LY +L DACR G L G V LL V
Sbjct: 696 GDMASWMWLPLLAD-WDEAGTFSWGSAALAWLYRQLCDACRRQGGDANLAGCVWLLQV 752
>gb|AAV25047.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1754
Score = 65.9 bits (159), Expect = 6e-10
Identities = 56/178 (31%), Positives = 81/178 (45%), Gaps = 19/178 (10%)
Query: 1 MPMGEMTVTLDDFACLTHLPIEGRMLDYEKKMPKHEGAALLMTYLGVSQHEAEKICNQEY 60
+P GE+TVTL+D A + LPI G+ + + + YLG+ A Q
Sbjct: 974 LPCGELTVTLEDVAMILGLPIRGQAVTGDTASGNWRER--VEEYLGLEPPVAPDGQRQTK 1031
Query: 61 GGYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRWYLMYLVGCLLFDDRS 120
+ LR + + P E +E A V+ YC R Y++Y+ G +LF D
Sbjct: 1032 TSGVPLSWLRANFG------------QCPAEADE-ATVQRYC-RAYVLYIFGSILFPDSG 1077
Query: 121 NKRIELIYLTTMADGYAGMRNYSWGGMTLAYLYGELADACR--PGHRALGGSVTLLTV 176
++L +AD + +SWG LA+LY +L DACR G L G V LL V
Sbjct: 1078 GDMASWMWLPLLAD-WDEAGTFSWGSAALAWLYRQLCDACRRQGGDANLAGCVWLLQV 1134
>ref|NP_909418.1| P0701D05.29 [Oryza sativa (japonica cultivar-group)]
gi|13486671|dbj|BAB39908.1| hypothetical protein~similar
to Oryza sativa chromosome 1, P0499C11.11 [Oryza sativa
(japonica cultivar-group)]
Length = 792
Score = 62.8 bits (151), Expect = 6e-09
Identities = 57/187 (30%), Positives = 82/187 (43%), Gaps = 27/187 (14%)
Query: 1 MPMGEMTVTLDDFACLTHLPIEGRMLDYEKKMPKHEGAALLMTYLGVSQHEAEKICNQEY 60
MP GE+T+TL D A + LPI G + P++E L+ YLG +A
Sbjct: 278 MPCGEITITLQDVAMILGLPIAGHAVTVNPTEPQNE---LVERYLG----KAPPPDRPRP 330
Query: 61 GGYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRWYLMYLVGCLLFDDRS 120
G +S+ R N D E +++ AR Y++ L+ LLF D S
Sbjct: 331 GLRVSWVR---------AEFNNCPEDADEETIKQHARA-------YILSLISGLLFPDAS 374
Query: 121 NKRIELIYLTTMADGYAGMRNYSWGGMTLAYLYGELADACRPGHRALGGSVTLLTVRKLK 180
+AD + +YSWG TLAYLY + DACR R GS + L+
Sbjct: 375 GDLYTFYPFPLIAD-LENIGSYSWGSATLAYLYRAMCDACR---RQSDGSNLTGCLLLLQ 430
Query: 181 FYIYVNF 187
F+ + +F
Sbjct: 431 FWSWEHF 437
>ref|NP_671779.1| expressed protein [Arabidopsis thaliana]
Length = 667
Score = 62.0 bits (149), Expect = 9e-09
Identities = 52/172 (30%), Positives = 79/172 (45%), Gaps = 23/172 (13%)
Query: 3 MGEMTVTLDDFACLTHLPIEGRMLDYEKKMPKHEGAALLMTYLGVSQHEAEKICNQEYGG 62
+GEMTVTL+D A L L I+G+ + + +A+ YLG S N GG
Sbjct: 96 VGEMTVTLEDIALLLGLGIDGKPV---IGLTYTTCSAVCERYLGKS-----PASNSASGG 147
Query: 63 YISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRWYLMYLVGCLLFDDRSNK 122
+ L+D ++ D EE+ R RT R YL+YLVG +F +
Sbjct: 148 MVKLSWLKDNFSE----------CPDDASFEEVER-RT---RAYLLYLVGSTIFSTTTGN 193
Query: 123 RIELIYLTTMADGYAGMRNYSWGGMTLAYLYGELADACRPGHRALGGSVTLL 174
++ ++YL D + ++WG LA+LY L +A + G +TLL
Sbjct: 194 KVPVMYLPLFED-FDDAGTFAWGAAALAFLYRALGNASVKSQSTICGCLTLL 244
>ref|NP_909918.1| putative mutator-like transposase [Oryza sativa (japonica
cultivar-group)] gi|29150382|gb|AAO72391.1| mutator-like
transposase [Oryza sativa (japonica cultivar-group)]
gi|28209524|gb|AAO37542.1| putative mutator-like
transposase [Oryza sativa (japonica cultivar-group)]
Length = 1381
Score = 55.8 bits (133), Expect = 7e-07
Identities = 36/90 (40%), Positives = 49/90 (54%), Gaps = 5/90 (5%)
Query: 89 PEELEELARVRTYCVRWYLMYLVGCLLFDDRSNKRIELIYLTTMADGYAGMRNYSWGGMT 148
P E +E A V+ YC R Y++Y+ G +LF D ++L +AD + +SWG
Sbjct: 804 PAEADE-ATVQRYC-RAYVLYIFGSILFPDSGGDMASWMWLPLLAD-WDEAGTFSWGSAA 860
Query: 149 LAYLYGELADACR--PGHRALGGSVTLLTV 176
LA+LY +L DACR G L G V LL V
Sbjct: 861 LAWLYRQLCDACRRQGGDANLAGCVWLLQV 890
>gb|AAM62582.1| unknown [Arabidopsis thaliana]
Length = 478
Score = 55.8 bits (133), Expect = 7e-07
Identities = 43/173 (24%), Positives = 77/173 (43%), Gaps = 22/173 (12%)
Query: 2 PMGEMTVTLDDFACLTHLPIEGRMLDYEKKMPKHEGAALLMTYLGVSQHEAEKICNQEYG 61
P GEMT+TLD+ + + L ++G+ + K+ + L + + E G
Sbjct: 84 PCGEMTITLDEVSLILGLAVDGKPVVGVKEKDEDPSQVCLKLLGKLPKGELS-------G 136
Query: 62 GYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRWYLMYLVGCLLFDDRSN 121
++ L++ + A + +E+E Y R YL+Y+VG +F
Sbjct: 137 NRVTAKWLKESFAECPKGATM-------KEIE-------YHTRAYLIYIVGSTIFATTDP 182
Query: 122 KRIELIYLTTMADGYAGMRNYSWGGMTLAYLYGELADACRPGHRALGGSVTLL 174
+I + YL D + Y+WG LA+LY ++ +A + +GG +TLL
Sbjct: 183 SKISVDYLILFED-FEKAGEYAWGAAALAFLYRQIGNASQRSQSIIGGCLTLL 234
>dbj|BAD94623.1| hypothetical protein [Arabidopsis thaliana]
gi|18394553|ref|NP_564042.1| expressed protein
[Arabidopsis thaliana] gi|25371349|pir||E86314 F2H15.15
protein - Arabidopsis thaliana gi|9665070|gb|AAF97272.1|
Contains similarity to a hypothetical protein At2g25010
gi|4559351 from Arabidopsis thaliana chromosome II
gb|AC006585
Length = 478
Score = 55.8 bits (133), Expect = 7e-07
Identities = 43/173 (24%), Positives = 77/173 (43%), Gaps = 22/173 (12%)
Query: 2 PMGEMTVTLDDFACLTHLPIEGRMLDYEKKMPKHEGAALLMTYLGVSQHEAEKICNQEYG 61
P GEMT+TLD+ + + L ++G+ + K+ + L + + E G
Sbjct: 84 PCGEMTITLDEVSLILGLAVDGKPVVGVKEKDEDPSQVCLRLLGKLPKGELS-------G 136
Query: 62 GYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRWYLMYLVGCLLFDDRSN 121
++ L++ + A + +E+E Y R YL+Y+VG +F
Sbjct: 137 NRVTAKWLKESFAECPKGATM-------KEIE-------YHTRAYLIYIVGSTIFATTDP 182
Query: 122 KRIELIYLTTMADGYAGMRNYSWGGMTLAYLYGELADACRPGHRALGGSVTLL 174
+I + YL D + Y+WG LA+LY ++ +A + +GG +TLL
Sbjct: 183 SKISVDYLILFED-FEKAGEYAWGAAALAFLYRQIGNASQRSQSIIGGCLTLL 234
>ref|XP_477211.1| calcineurin-like phosphoesterase-like [Oryza sativa (japonica
cultivar-group)] gi|50510004|dbj|BAD30617.1|
calcineurin-like phosphoesterase-like [Oryza sativa
(japonica cultivar-group)] gi|33146683|dbj|BAC80078.1|
calcineurin-like phosphoesterase-like [Oryza sativa
(japonica cultivar-group)]
Length = 327
Score = 53.9 bits (128), Expect = 3e-06
Identities = 44/175 (25%), Positives = 75/175 (42%), Gaps = 9/175 (5%)
Query: 1 MPMGEMTVTLDDFACLTHLPIEGRMLDYEKKMPKHEGAALLMTYLGVSQHEAEKICNQEY 60
+P+GEMT+TL D +CL LPI GR + + + L+ QH +K +E
Sbjct: 87 LPVGEMTITLQDVSCLWGLPIHGRPITGQADGSWVDMIERLLGIPMEEQHMKQKKRKKED 146
Query: 61 G-GYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRWYLMYLVGCLLFDDR 119
+SY R + R V+ E+ + R ++ ++G ++F D
Sbjct: 147 DMTMVSYSRYSISLSKLRDRFRVMPKNATEREI-------NWYTRALVLDIIGSMVFTDT 199
Query: 120 SNKRIELIYLTTMADGYAGMRNYSWGGMTLAYLYGELADACRPGHRALGGSVTLL 174
S + +YL M + + Y+WG L+ LY +L+ A + + LL
Sbjct: 200 SGDGVPAMYLQFMMN-LSEQTEYNWGAAALSMLYRQLSIASEKERAEISRPLLLL 253
>dbj|BAD30920.1| serine/threonine phosphatase PP7-related-like [Oryza sativa
(japonica cultivar-group)]
Length = 619
Score = 52.8 bits (125), Expect = 6e-06
Identities = 52/171 (30%), Positives = 77/171 (44%), Gaps = 22/171 (12%)
Query: 5 EMTVTLDDFACLTHLPIEGRMLDYEKKMPKHEGAALLMTYLGVSQHEAEKICNQEYGGYI 64
EM +L D + + +P+ G ++ E + + + +LG S E++C G I
Sbjct: 96 EMAPSLRDVSYILGIPVTGHVVTAEP-IGDEAVRRMCLHFLGESPGNGEQLC-----GLI 149
Query: 65 SYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRWYLMYLVGCLLFDDRSNKRI 124
RL Y + E+P + E+A Y R YL+YLVG LF D +
Sbjct: 150 ---RLTWLYRKFHQLP------ENPT-INEIA----YSTRAYLLYLVGSTLFPDTMRGFV 195
Query: 125 ELIYLTTMADGYAGMRNYSWGGMTLAYLYGELADACRPGHRA-LGGSVTLL 174
YL +AD + +R Y+WG LA+LY L+ A P GS TLL
Sbjct: 196 SPRYLPLLAD-FRKIREYAWGSAALAHLYRGLSVAVTPNATTQFLGSATLL 245
>ref|XP_466494.1| calcineurin-like phosphoesterase-like protein [Oryza sativa
(japonica cultivar-group)] gi|50726478|dbj|BAD34087.1|
calcineurin-like phosphoesterase-like protein [Oryza
sativa (japonica cultivar-group)]
Length = 766
Score = 52.0 bits (123), Expect = 1e-05
Identities = 54/172 (31%), Positives = 83/172 (47%), Gaps = 24/172 (13%)
Query: 4 GEMTVTLDDFACLTHLPIEGRMLDYEKKMPKHEGAALLMTYLGVSQHEAEKICNQEYGGY 63
GEMTVTL D + L LPI+GR + H+ A +++ LG H+ + + +
Sbjct: 153 GEMTVTLQDVSMLLALPIDGRPV---CSTTDHDYAQMVIDCLG---HD-PRGPSMPGKSF 205
Query: 64 ISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRWYLMYLVGCLLFDDRSNKR 123
+ Y L+ + E PE ++ R VR Y++ L+ +LF D + R
Sbjct: 206 LHYKWLKKHF------------YELPEGTDDQTVERH--VRAYILSLLCGVLFPDGTG-R 250
Query: 124 IELIYLTTMADGYAGMRNYSWGGMTLAYLYGELAD-ACRPGHRALGGSVTLL 174
+ LIYL +AD + + YSWG LA+LY L A + +GGS+ LL
Sbjct: 251 MSLIYLPLIAD-LSLVGTYSWGSAVLAFLYRSLCSVASSHNIKNIGGSLLLL 301
>ref|NP_916020.1| OSJNBb0021A09.19 [Oryza sativa (japonica cultivar-group)]
Length = 761
Score = 52.0 bits (123), Expect = 1e-05
Identities = 54/172 (31%), Positives = 82/172 (47%), Gaps = 24/172 (13%)
Query: 4 GEMTVTLDDFACLTHLPIEGRMLDYEKKMPKHEGAALLMTYLGVSQHEAEKICNQEYGGY 63
GEM VTL D A L LPI+GR + H+ A +++ LG H+ + + +
Sbjct: 170 GEMAVTLQDVAMLLALPIDGRPV---CSTTDHDYAQMVIDCLG---HD-PRGPSMPGKSF 222
Query: 64 ISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRWYLMYLVGCLLFDDRSNKR 123
+ Y L+ + E PE ++ R VR Y++ L+ +LF D + R
Sbjct: 223 LHYKWLKKHF------------YELPEGADDQTVERH--VRAYILSLLCGVLFPDGTG-R 267
Query: 124 IELIYLTTMADGYAGMRNYSWGGMTLAYLYGELAD-ACRPGHRALGGSVTLL 174
+ LIYL +AD + + YSWG LA+LY L A + +GGS+ LL
Sbjct: 268 MSLIYLPLIAD-LSRVGTYSWGSAVLAFLYRSLCSVASSHNIKNIGGSLLLL 318
>ref|NP_918073.1| P0702H08.15 [Oryza sativa (japonica cultivar-group)]
Length = 718
Score = 52.0 bits (123), Expect = 1e-05
Identities = 54/172 (31%), Positives = 82/172 (47%), Gaps = 24/172 (13%)
Query: 4 GEMTVTLDDFACLTHLPIEGRMLDYEKKMPKHEGAALLMTYLGVSQHEAEKICNQEYGGY 63
GEM VTL D A L LPI+GR + H+ A +++ LG H+ + + +
Sbjct: 184 GEMAVTLQDVAMLLALPIDGRPV---CSTTDHDYAQMVIDCLG---HD-PRGPSMPGKSF 236
Query: 64 ISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRWYLMYLVGCLLFDDRSNKR 123
+ Y L+ + E PE ++ R VR Y++ L+ +LF D + R
Sbjct: 237 LHYKWLKKHF------------YELPEGADDQTVERH--VRAYILSLLCGVLFPDGTG-R 281
Query: 124 IELIYLTTMADGYAGMRNYSWGGMTLAYLYGELAD-ACRPGHRALGGSVTLL 174
+ LIYL +AD + + YSWG LA+LY L A + +GGS+ LL
Sbjct: 282 MSLIYLPLIAD-LSRVGTYSWGSAVLAFLYRSLCSVASSHNIKNIGGSLLLL 332
>gb|AAD23012.1| expressed protein [Arabidopsis thaliana]
gi|18400697|ref|NP_565582.1| expressed protein
[Arabidopsis thaliana] gi|25371347|pir||B84643
hypothetical protein At2g25010 [imported] - Arabidopsis
thaliana
Length = 509
Score = 52.0 bits (123), Expect = 1e-05
Identities = 48/175 (27%), Positives = 80/175 (45%), Gaps = 21/175 (12%)
Query: 1 MPMGEMTVTLDDFACLTHLPIEGRMLDYEKKMPKHEGAALLMTYLGVSQHEAEKICNQEY 60
+P+GEMT+TLD+ A + L I+G + K G + M G + N+E
Sbjct: 92 LPLGEMTITLDEVALVLGLEIDGDPIVGSK-----VGDEVAMDMCGRLLGKLPSAANKE- 145
Query: 61 GGYISYPRLRDFYTSYLGRANVLAGTEDPEELE-ELARVRTYCVRWYLMYLVGCLLFDDR 119
++ R++ ++L R +E PE+ ++ + T R YL+YL+G +F
Sbjct: 146 ---VNCSRVK---LNWLKR----TFSECPEDASFDVVKCHT---RAYLLYLIGSTIFATT 192
Query: 120 SNKRIELIYLTTMADGYAGMRNYSWGGMTLAYLYGELADACRPGHRALGGSVTLL 174
++ + YL D + Y+WG LA LY L +A + G +TLL
Sbjct: 193 DGDKVSVKYLPLFED-FDQAGRYAWGAAALACLYRALGNASLKSQSNICGCLTLL 246
>emb|CAE05572.2| OSJNBb0013O03.13 [Oryza sativa (japonica cultivar-group)]
Length = 1611
Score = 50.1 bits (118), Expect = 4e-05
Identities = 41/162 (25%), Positives = 72/162 (44%), Gaps = 21/162 (12%)
Query: 1 MPMGEMTVTLDDFACLTHLPIEGRMLDYEKKMP--KHEGAALLMTYLGVSQHEAEKICNQ 58
+P GEMT+TL D A + LP++G + + P + + L L + Q + K
Sbjct: 994 LPSGEMTITLQDVAMILALPLQGHAVTGRTETPGWRAQVEQLFGIQLNIEQGQGGK--KM 1051
Query: 59 EYGGYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRWYLMYLVGCLLFDD 118
+ G +S+ + ++ E RV Y R Y+++L+G +LF D
Sbjct: 1052 QNGIPLSW---------------LSQNFSHLDDDAEPWRVECY-ARAYILHLLGGVLFPD 1095
Query: 119 RSNKRIELIYLTTMADGYAGMRNYSWGGMTLAYLYGELADAC 160
I++ +A+ + +SWG LA+ Y +L +AC
Sbjct: 1096 AGGDIASAIWIPLVAN-LGDLGRFSWGSAVLAWTYRQLCEAC 1136
>gb|AAM61398.1| unknown [Arabidopsis thaliana]
Length = 509
Score = 50.1 bits (118), Expect = 4e-05
Identities = 47/175 (26%), Positives = 75/175 (42%), Gaps = 21/175 (12%)
Query: 1 MPMGEMTVTLDDFACLTHLPIEGRMLDYEKKMPKHEGAALLMTYLGVSQHEAEKICNQEY 60
+P+GEMT+TLD+ A + L I+G + K+ + LG A K N
Sbjct: 92 LPLGEMTITLDEVALVLGLEIDGDPI-VGSKVDDEVAMDMCGRLLGKLPSAANKEVN--- 147
Query: 61 GGYISYPRLRDFYTSYLGRANVLAGTEDPEELE-ELARVRTYCVRWYLMYLVGCLLFDDR 119
S +L ++ +E PE+ ++ + T R YL+YL+G +F
Sbjct: 148 ---CSRVKLNWLKRTF---------SECPEDASFDVVKCHT---RAYLLYLIGSTIFATT 192
Query: 120 SNKRIELIYLTTMADGYAGMRNYSWGGMTLAYLYGELADACRPGHRALGGSVTLL 174
++ + YL D + Y+WG LA LY L +A + G +TLL
Sbjct: 193 DGDKVSVKYLPLFED-FDQAGRYAWGAAALACLYRALGNASLKSQSNICGCLTLL 246
>ref|NP_912436.1| Putative transposable element [Oryza sativa (japonica
cultivar-group)] gi|27476096|gb|AAO17027.1| Putative
transposable element [Oryza sativa (japonica
cultivar-group)]
Length = 663
Score = 48.9 bits (115), Expect = 8e-05
Identities = 44/160 (27%), Positives = 71/160 (43%), Gaps = 17/160 (10%)
Query: 1 MPMGEMTVTLDDFACLTHLPIEGRMLDYEKKMPKHEGAALLMTYLGVSQHEAEKICNQEY 60
+P GEMT+TL D A + LP+ G + + P A + G+ + Q
Sbjct: 92 LPSGEMTITLQDVAMILALPLRGHAVTGRTETPGWH--AQVQQLFGIPLN-----IEQGQ 144
Query: 61 GGYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRWYLMYLVGCLLFDDRS 120
GG + S+L + N +D E RV Y R Y+++L+G +LF D
Sbjct: 145 GGK---KKQNGIPLSWLSQ-NFSHLDDDAEPW----RVECYA-RAYILHLLGGVLFPDAG 195
Query: 121 NKRIELIYLTTMADGYAGMRNYSWGGMTLAYLYGELADAC 160
I++ +A+ + +SWG LA+ Y +L +AC
Sbjct: 196 GDIASAIWIPLVAN-LGDLGRFSWGSAVLAWTYRQLCEAC 234
>gb|AAF79521.1| F21D18.16 [Arabidopsis thaliana] gi|25405918|pir||D96521 protein
F21D18.16 [imported] - Arabidopsis thaliana
Length = 1340
Score = 47.0 bits (110), Expect = 3e-04
Identities = 45/174 (25%), Positives = 74/174 (41%), Gaps = 23/174 (13%)
Query: 1 MPMGEMTVTLDDFACLTHLPIEGRMLDYEKKMPKHEGAALLMTYLGVSQHEAEKICNQEY 60
+P GE+TVTL D L L ++G + K + A L LG + +
Sbjct: 101 LPAGEITVTLQDVNILLGLRVDGPAVTGSTK---YNWADLCEDLLGHRPGPKDL-----H 152
Query: 61 GGYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRWYLMYLVGCLLFDDRS 120
G ++S LR+ + + DP+E+ R + ++ L+ L+ D+S
Sbjct: 153 GSHVSLAWLRENFRNL---------PADPDEVTLKCHTRAF-----VLALMSGFLYGDKS 198
Query: 121 NKRIELIYLTTMADGYAGMRNYSWGGMTLAYLYGELADACRPGHRALGGSVTLL 174
+ L +L + D + + SWG TLA LY EL A + + G + LL
Sbjct: 199 KHDVALTFLPLLRD-FDEVAKLSWGSATLALLYRELCRASKRTVSTICGPLVLL 251
>ref|NP_175246.1| calcineurin-like phosphoesterase family protein [Arabidopsis
thaliana]
Length = 1338
Score = 47.0 bits (110), Expect = 3e-04
Identities = 45/174 (25%), Positives = 74/174 (41%), Gaps = 23/174 (13%)
Query: 1 MPMGEMTVTLDDFACLTHLPIEGRMLDYEKKMPKHEGAALLMTYLGVSQHEAEKICNQEY 60
+P GE+TVTL D L L ++G + K + A L LG + +
Sbjct: 101 LPAGEITVTLQDVNILLGLRVDGPAVTGSTK---YNWADLCEDLLGHRPGPKDL-----H 152
Query: 61 GGYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRWYLMYLVGCLLFDDRS 120
G ++S LR+ + + DP+E+ R + ++ L+ L+ D+S
Sbjct: 153 GSHVSLAWLRENFRNL---------PADPDEVTLKCHTRAF-----VLALMSGFLYGDKS 198
Query: 121 NKRIELIYLTTMADGYAGMRNYSWGGMTLAYLYGELADACRPGHRALGGSVTLL 174
+ L +L + D + + SWG TLA LY EL A + + G + LL
Sbjct: 199 KHDVALTFLPLLRD-FDEVAKLSWGSATLALLYRELCRASKRTVSTICGPLVLL 251
>ref|XP_478317.1| serine/threonine phosphatase PP7 -related-like protein [Oryza
sativa (japonica cultivar-group)]
gi|23307580|dbj|BAC16715.1| serine/threonine phosphatase
PP7-related-like protein [Oryza sativa (japonica
cultivar-group)]
Length = 589
Score = 43.1 bits (100), Expect = 0.005
Identities = 46/175 (26%), Positives = 73/175 (41%), Gaps = 30/175 (17%)
Query: 5 EMTVTLDDFACLTHLPIEGRMLDYEKKMPKHEGAALLMTYLGVSQHEAEKICNQEYGGYI 64
EM +L D A + +P+ GR++ GA L E +C Q G
Sbjct: 83 EMAPSLRDAAYILGIPVTGRVVT--------TGAVL--------NKSVEDLCFQYLG--- 123
Query: 65 SYPRLRDFYTSYLG----RANVLAGTEDPEELEELARVRTYCVRWYLMYLVGCLLFDDRS 120
P RD S++ ++ E P + + Y R YL++L+G L +R
Sbjct: 124 QVPDCRDCRGSHVKLSWLQSKFSRIPERPTNDQTM-----YGTRAYLLFLIGSALLPERD 178
Query: 121 NKRIELIYLTTMADGYAGMRNYSWGGMTLAYLYGELADA-CRPGHRALGGSVTLL 174
+ YL ++D + ++ Y+WG LA+LY L+ A + L GS LL
Sbjct: 179 RGYVSPKYLPLLSD-FDKVQEYAWGAAALAHLYKALSIAVAHSARKRLFGSAALL 232
>ref|XP_481308.1| calcineurin-like phosphoesterase-like [Oryza sativa (japonica
cultivar-group)] gi|38175641|dbj|BAD01348.1|
calcineurin-like phosphoesterase-like [Oryza sativa
(japonica cultivar-group)] gi|38175657|dbj|BAD01362.1|
calcineurin-like phosphoesterase-like [Oryza sativa
(japonica cultivar-group)]
Length = 432
Score = 41.2 bits (95), Expect = 0.017
Identities = 22/71 (30%), Positives = 35/71 (48%), Gaps = 1/71 (1%)
Query: 104 RWYLMYLVGCLLFDDRSNKRIELIYLTTMADGYAGMRNYSWGGMTLAYLYGELADACRPG 163
R ++ ++G ++F D S + +YL M D A Y+WGG LA LY +L +
Sbjct: 15 RALVLEILGSIIFTDSSGDSVPAMYLQFM-DNLATRTEYNWGGAVLAMLYRQLNNGAEKA 73
Query: 164 HRALGGSVTLL 174
+ G + LL
Sbjct: 74 RSEIFGPLVLL 84
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.325 0.142 0.440
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 351,439,674
Number of Sequences: 2540612
Number of extensions: 14345572
Number of successful extensions: 32800
Number of sequences better than 10.0: 57
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 49
Number of HSP's that attempted gapping in prelim test: 32746
Number of HSP's gapped (non-prelim): 87
length of query: 199
length of database: 863,360,394
effective HSP length: 121
effective length of query: 78
effective length of database: 555,946,342
effective search space: 43363814676
effective search space used: 43363814676
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 72 (32.3 bits)
Medicago: description of AC140034.3


