

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140031.2 + phase: 0 /pseudo
(484 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
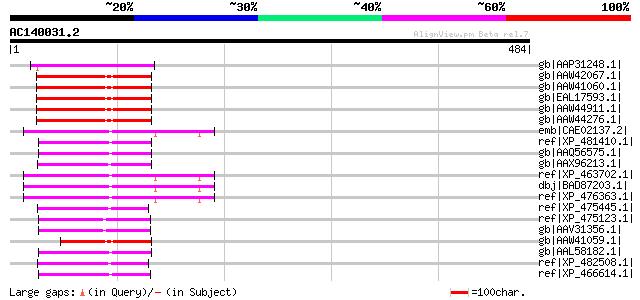
Score E
Sequences producing significant alignments: (bits) Value
gb|AAP31248.1| transposase [Fusarium oxysporum f. sp. melonis] 94 1e-17
gb|AAW42067.1| conserved hypothetical protein [Cryptococcus neof... 90 2e-16
gb|AAW41060.1| conserved hypothetical protein [Cryptococcus neof... 90 2e-16
gb|EAL17593.1| hypothetical protein CNBM0450 [Cryptococcus neofo... 90 2e-16
gb|AAW44911.1| conserved hypothetical protein [Cryptococcus neof... 90 2e-16
gb|AAW44276.1| conserved hypothetical protein [Cryptococcus neof... 90 2e-16
emb|CAE02137.2| OSJNBa0074L08.5 [Oryza sativa (japonica cultivar... 69 4e-10
ref|XP_481410.1| far-red impaired response protein -like [Oryza ... 68 6e-10
gb|AAQ56575.1| putative transposase [Oryza sativa (japonica cult... 68 6e-10
gb|AAX96213.1| transposon protein, putative, unclassified [Oryza... 68 6e-10
ref|XP_463702.1| putative far-red impaired response protein [Ory... 67 1e-09
dbj|BAD87203.1| far-red impaired response-like [Oryza sativa (ja... 67 1e-09
ref|XP_476363.1| far-red impaired response-like protein [Oryza s... 67 2e-09
ref|XP_475445.1| putative far-red impaired response protein [Ory... 67 2e-09
ref|XP_475123.1| putative Mutator-like transposase [Oryza sativa... 67 2e-09
gb|AAV31356.1| hypothetical protein [Oryza sativa (japonica cult... 67 2e-09
gb|AAW41059.1| hypothetical protein CNA05530 [Cryptococcus neofo... 66 2e-09
gb|AAL58182.1| putative far-red impaired response protein [Oryza... 66 3e-09
ref|XP_482508.1| putative far-red impaired response protein [Ory... 65 5e-09
ref|XP_466614.1| putative far-red impaired response protein [Ory... 65 5e-09
>gb|AAP31248.1| transposase [Fusarium oxysporum f. sp. melonis]
Length = 836
Score = 94.0 bits (232), Expect = 1e-17
Identities = 45/119 (37%), Positives = 71/119 (58%), Gaps = 3/119 (2%)
Query: 20 LIQHP--LSFFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEEN 77
L HP L + +P VL++DSTY+TN +KMP+ +IVGV + T+ + F F++ E+E +
Sbjct: 213 LFAHPKSLEYLKTYPEVLILDSTYKTNRFKMPLLDIVGVDACQRTFCIAFAFLSGEEEGD 272
Query: 78 FVWVLQMMCKLLTS-RMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVKVRC 135
F W LQ + + + +P V++TD+ MNAVS+ P S C +H+ K V+ C
Sbjct: 273 FTWALQALRSVYEDHNIGLPSVILTDRCLACMNAVSSCFPGSALFLCLWHINKAVQSYC 331
Score = 43.9 bits (102), Expect = 0.011
Identities = 22/70 (31%), Positives = 39/70 (55%), Gaps = 4/70 (5%)
Query: 284 YSWLGGYMSRYALGNIVLEEERCRGTLCMDKEICGCV*RKSYGLPCACFVATKIRDDKPI 343
YS + G++S AL + + ER L + +C V ++ GLPCA + ++ ++P+
Sbjct: 480 YSNIRGWISHEALRKVETQRER----LLKEVPVCTGVFTRTLGLPCAHSLQPLLKQNQPL 535
Query: 344 LLDEIHLHWN 353
LL+ H HW+
Sbjct: 536 LLNHFHSHWH 545
>gb|AAW42067.1| conserved hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21] gi|50260537|gb|EAL23192.1|
hypothetical protein CNBA5360 [Cryptococcus neoformans
var. neoformans B-3501A] gi|50259285|gb|EAL21958.1|
hypothetical protein CNBC0980 [Cryptococcus neoformans
var. neoformans B-3501A] gi|50256296|gb|EAL19021.1|
hypothetical protein CNBH1230 [Cryptococcus neoformans
var. neoformans B-3501A] gi|57229113|gb|AAW45547.1|
conserved hypothetical protein [Cryptococcus neoformans
var. neoformans JEC21] gi|58271396|ref|XP_572854.1|
conserved hypothetical protein [Cryptococcus neoformans
var. neoformans JEC21] gi|58264436|ref|XP_569374.1|
conserved hypothetical protein [Cryptococcus neoformans
var. neoformans JEC21]
Length = 932
Score = 89.7 bits (221), Expect = 2e-16
Identities = 44/107 (41%), Positives = 70/107 (65%), Gaps = 2/107 (1%)
Query: 26 SFFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMM 85
+ + FP VL D TY+TN+Y+MPM IVG TST +TY+ G + E + L
Sbjct: 315 ALIHRFPEVLFFDCTYKTNLYRMPMLHIVGSTSTGMTYTAGVILMLRETTNWYTQALNSF 374
Query: 86 CKLLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVK 132
+L+ + +V KVV+TD+D L+NA+ +VLP+++ +C++H+Q+NVK
Sbjct: 375 FELV-GKPDV-KVVITDRDPALINALMSVLPKAYRFSCFWHLQENVK 419
>gb|AAW41060.1| conserved hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21] gi|58258933|ref|XP_566879.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21]
Length = 678
Score = 89.7 bits (221), Expect = 2e-16
Identities = 44/107 (41%), Positives = 70/107 (65%), Gaps = 2/107 (1%)
Query: 26 SFFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMM 85
+ + FP VL D TY+TN+Y+MPM IVG TST +TY+ G + E + L
Sbjct: 61 ALIHRFPEVLFFDCTYKTNLYRMPMLHIVGSTSTGMTYTAGVILMLRETTNWYTQALNSF 120
Query: 86 CKLLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVK 132
+L+ + +V KVV+TD+D L+NA+ +VLP+++ +C++H+Q+NVK
Sbjct: 121 FELV-GKPDV-KVVITDRDPALINALMSVLPKAYRFSCFWHLQENVK 165
>gb|EAL17593.1| hypothetical protein CNBM0450 [Cryptococcus neoformans var.
neoformans B-3501A] gi|57226908|gb|AAW43368.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21] gi|58267038|ref|XP_570675.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21]
Length = 580
Score = 89.7 bits (221), Expect = 2e-16
Identities = 44/107 (41%), Positives = 70/107 (65%), Gaps = 2/107 (1%)
Query: 26 SFFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMM 85
+ + FP VL D TY+TN+Y+MPM IVG TST +TY+ G + E + L
Sbjct: 315 ALIHRFPEVLFFDCTYKTNLYRMPMLHIVGSTSTGMTYTAGVILMLRETTNWYTQALNSF 374
Query: 86 CKLLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVK 132
+L+ + +V KVV+TD+D L+NA+ +VLP+++ +C++H+Q+NVK
Sbjct: 375 FELV-GKPDV-KVVITDRDPALINALMSVLPKAYRFSCFWHLQENVK 419
>gb|AAW44911.1| conserved hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21] gi|58270124|ref|XP_572218.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21]
Length = 920
Score = 89.7 bits (221), Expect = 2e-16
Identities = 44/107 (41%), Positives = 70/107 (65%), Gaps = 2/107 (1%)
Query: 26 SFFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMM 85
+ + FP VL D TY+TN+Y+MPM IVG TST +TY+ G + E + L
Sbjct: 315 ALIHRFPEVLFFDCTYKTNLYRMPMLHIVGSTSTGMTYTAGVILMLRETTNWYTQALNSF 374
Query: 86 CKLLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVK 132
+L+ + +V KVV+TD+D L+NA+ +VLP+++ +C++H+Q+NVK
Sbjct: 375 FELV-GKPDV-KVVITDRDPALINALMSVLPKAYRFSCFWHLQENVK 419
>gb|AAW44276.1| conserved hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21] gi|58268854|ref|XP_571583.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21]
Length = 895
Score = 89.7 bits (221), Expect = 2e-16
Identities = 44/107 (41%), Positives = 70/107 (65%), Gaps = 2/107 (1%)
Query: 26 SFFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMM 85
+ + FP VL D TY+TN+Y+MPM IVG TST +TY+ G + E + L
Sbjct: 315 ALIHRFPEVLFFDCTYKTNLYRMPMLHIVGSTSTGMTYTAGVILMLRETTNWYTQALNSF 374
Query: 86 CKLLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVK 132
+L+ + +V KVV+TD+D L+NA+ +VLP+++ +C++H+Q+NVK
Sbjct: 375 FELV-GKPDV-KVVITDRDPALINALMSVLPKAYRFSCFWHLQENVK 419
>emb|CAE02137.2| OSJNBa0074L08.5 [Oryza sativa (japonica cultivar-group)]
gi|50926620|ref|XP_473257.1| OSJNBa0074L08.5 [Oryza
sativa (japonica cultivar-group)]
Length = 1061
Score = 68.6 bits (166), Expect = 4e-10
Identities = 46/184 (25%), Positives = 81/184 (44%), Gaps = 8/184 (4%)
Query: 14 LKTYFALIQHPLSFFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHE 73
+K+ F + + + D+TY+TN Y MP IVGVT F+ E
Sbjct: 507 VKSMFWTDGRSTQLYEQYGEFVSFDTTYKTNRYNMPFAPIVGVTGHGNICIFACAFLGDE 566
Query: 74 KEENFVWVLQMMCKLLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVKV 133
E F WV + + + P+ ++TD+D + A+ V P S NC FH+ K +
Sbjct: 567 TTETFKWVFETFLTAMGGKH--PETIITDQDLAMRAAIRQVFPNSKHRNCLFHILKKCRE 624
Query: 134 R---CILDCRYPMAKKDGKEVKHGDVVK-KIMRVW*VIIESHPLKN--YIQMH*WSSKMF 187
R D R + ++ H + + + +W +I + LKN Y+++ + K F
Sbjct: 625 RSGNTFSDKRRKDLYAEFNDIVHNSLTRAEFESLWLQMIAQYNLKNIKYLEIMWRTRKNF 684
Query: 188 VMIF 191
+ ++
Sbjct: 685 IPVY 688
>ref|XP_481410.1| far-red impaired response protein -like [Oryza sativa (japonica
cultivar-group)] gi|35215248|dbj|BAC92598.1| putative
far-red impaired response protein [Oryza sativa
(japonica cultivar-group)] gi|35215057|dbj|BAC92415.1|
putative far-red impaired response protein [Oryza sativa
(japonica cultivar-group)]
Length = 1132
Score = 68.2 bits (165), Expect = 6e-10
Identities = 32/105 (30%), Positives = 52/105 (49%), Gaps = 2/105 (1%)
Query: 28 FNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCK 87
+ + + D+TY+TN Y MP VGV T G F+ E E F WV +
Sbjct: 624 YEEYGDCISFDTTYRTNRYNMPFAPFVGVIGHGTTCLFGCAFLGDETAETFKWVFETFAT 683
Query: 88 LLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVK 132
+ + PK ++TD+D + +A++ V + NC FH++KN +
Sbjct: 684 AMGGKH--PKTIITDQDNAMRSAIAQVFQNTKHRNCLFHIKKNCR 726
>gb|AAQ56575.1| putative transposase [Oryza sativa (japonica cultivar-group)]
Length = 1037
Score = 68.2 bits (165), Expect = 6e-10
Identities = 32/105 (30%), Positives = 52/105 (49%), Gaps = 2/105 (1%)
Query: 28 FNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCK 87
+ + + D+TY+TN Y MP VGV T G F+ E E F WV +
Sbjct: 529 YEEYGDCISFDTTYRTNRYNMPFAPFVGVIGHGTTCLFGCAFLGDETAETFKWVFETFAT 588
Query: 88 LLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVK 132
+ + PK ++TD+D + +A++ V + NC FH++KN +
Sbjct: 589 AMGGKH--PKTIITDQDNAMRSAIAQVFQNTKHRNCLFHIKKNCR 631
>gb|AAX96213.1| transposon protein, putative, unclassified [Oryza sativa (japonica
cultivar-group)]
Length = 1318
Score = 68.2 bits (165), Expect = 6e-10
Identities = 31/106 (29%), Positives = 54/106 (50%), Gaps = 2/106 (1%)
Query: 27 FFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMC 86
F+ F ++ D+ Y+TN Y +P VG+TS G+ F+ E E F W+ +
Sbjct: 777 FYEMFDDIVSFDTMYKTNRYDLPFAPFVGITSHGDNCLFGYAFLQDETSETFQWLFKTFL 836
Query: 87 KLLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVK 132
+ + VP ++TD+D + A++ V P++ NC FH+ N +
Sbjct: 837 DCMGGK--VPATIITDQDLAMKAAIAIVFPDTVHRNCLFHMLSNAR 880
>ref|XP_463702.1| putative far-red impaired response protein [Oryza sativa (japonica
cultivar-group)]
Length = 1114
Score = 67.0 bits (162), Expect = 1e-09
Identities = 45/184 (24%), Positives = 81/184 (43%), Gaps = 8/184 (4%)
Query: 14 LKTYFALIQHPLSFFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHE 73
+K+ F + + + D+TY+TN Y MP IVGVT F+ E
Sbjct: 560 VKSMFWTDGRSTQLYEQYGEFVSFDTTYKTNRYNMPFAPIVGVTGHGNICIFACAFLGDE 619
Query: 74 KEENFVWVLQMMCKLLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVKV 133
E F WV + + + P+ ++TD+D + A+ V P S NC FH+ K +
Sbjct: 620 TTETFKWVFETFLTAMGGKH--PETIITDQDLAMRAAIRQVFPNSKHRNCLFHILKKCRE 677
Query: 134 R---CILDCRYPMAKKDGKEVKHGDVVK-KIMRVW*VIIESHPLKN--YIQMH*WSSKMF 187
R D R + ++ H + + + +W +I + L+N Y+++ + K F
Sbjct: 678 RSGNTFSDKRRKDLYAEFNDIVHNSLTRAEFESLWLQMIAQYNLENIKYLEIMWRTRKNF 737
Query: 188 VMIF 191
+ ++
Sbjct: 738 IPVY 741
>dbj|BAD87203.1| far-red impaired response-like [Oryza sativa (japonica
cultivar-group)]
Length = 1130
Score = 67.0 bits (162), Expect = 1e-09
Identities = 45/184 (24%), Positives = 81/184 (43%), Gaps = 8/184 (4%)
Query: 14 LKTYFALIQHPLSFFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHE 73
+K+ F + + + D+TY+TN Y MP IVGVT F+ E
Sbjct: 576 VKSMFWTDGRSTQLYEQYGEFVSFDTTYKTNRYNMPFAPIVGVTGHGNICIFACAFLGDE 635
Query: 74 KEENFVWVLQMMCKLLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVKV 133
E F WV + + + P+ ++TD+D + A+ V P S NC FH+ K +
Sbjct: 636 TTETFKWVFETFLTAMGGKH--PETIITDQDLAMRAAIRQVFPNSKHRNCLFHILKKCRE 693
Query: 134 R---CILDCRYPMAKKDGKEVKHGDVVK-KIMRVW*VIIESHPLKN--YIQMH*WSSKMF 187
R D R + ++ H + + + +W +I + L+N Y+++ + K F
Sbjct: 694 RSGNTFSDKRRKDLYAEFNDIVHNSLTRAEFESLWLQMIAQYNLENIKYLEIMWRTRKNF 753
Query: 188 VMIF 191
+ ++
Sbjct: 754 IPVY 757
>ref|XP_476363.1| far-red impaired response-like protein [Oryza sativa (japonica
cultivar-group)] gi|22324476|dbj|BAC10390.1| far-red
impaired response-like protein [Oryza sativa (japonica
cultivar-group)] gi|50508936|dbj|BAD31841.1| far-red
impaired response-like protein [Oryza sativa (japonica
cultivar-group)]
Length = 772
Score = 66.6 bits (161), Expect = 2e-09
Identities = 45/184 (24%), Positives = 81/184 (43%), Gaps = 8/184 (4%)
Query: 14 LKTYFALIQHPLSFFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHE 73
+K+ F + + + D+TY+TN Y MP IVGVT F+ E
Sbjct: 245 VKSMFWTDGRSTQLYEQYGEFVSFDTTYKTNRYNMPFAPIVGVTGHGNICIFACAFLGDE 304
Query: 74 KEENFVWVLQMMCKLLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVKV 133
E F WV + + + P+ ++TD+D + A+ V P S NC FH+ K +
Sbjct: 305 TTETFKWVFETFLTAMGRKH--PETIITDQDLAMRAAIRQVFPNSKHRNCLFHILKKYRE 362
Query: 134 R---CILDCRYPMAKKDGKEVKHGDVVK-KIMRVW*VIIESHPLKN--YIQMH*WSSKMF 187
R D R + ++ H + + + +W +I + L+N Y+++ + K F
Sbjct: 363 RSGNTFSDKRRKDLYAEFNDIVHNSLTRAEFESLWLQMIAQYNLENIKYLEIMWRTRKNF 422
Query: 188 VMIF 191
+ ++
Sbjct: 423 IPVY 426
>ref|XP_475445.1| putative far-red impaired response protein [Oryza sativa (japonica
cultivar-group)] gi|46576038|gb|AAT01399.1| putative
far-red impaired response protein [Oryza sativa
(japonica cultivar-group)]
Length = 888
Score = 66.6 bits (161), Expect = 2e-09
Identities = 30/103 (29%), Positives = 55/103 (53%), Gaps = 2/103 (1%)
Query: 27 FFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMC 86
++ + V+ D+TY TN Y +P VG++ T G F+ E E F W+ +
Sbjct: 528 WYEKYGDVVSFDTTYFTNRYNLPFAPFVGISGHGNTIVFGCAFLHDETSETFKWMFRTFL 587
Query: 87 KLLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQK 129
K ++ + PK+++TD+D + +A++ V + NC+FH+ K
Sbjct: 588 KAMSQK--EPKIIITDQDGAMRSAIAQVFQNAKHGNCFFHIVK 628
>ref|XP_475123.1| putative Mutator-like transposase [Oryza sativa (japonica
cultivar-group)] gi|46063440|gb|AAS79743.1| putative
Mutator-like transposase [Oryza sativa (japonica
cultivar-group)]
Length = 1510
Score = 66.6 bits (161), Expect = 2e-09
Identities = 33/104 (31%), Positives = 56/104 (53%), Gaps = 2/104 (1%)
Query: 28 FNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCK 87
++ F VL +DSTY TN Y M GV + +G F+ EK E+FVW+ Q K
Sbjct: 930 YSVFGDVLSVDSTYSTNQYNMKFVPFTGVNHHLQSVFLGASFLADEKIESFVWLFQTFLK 989
Query: 88 LLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNV 131
+ P++++TD+D ++ A++ +LP + C +H+ + V
Sbjct: 990 --ATGGVAPRLIITDEDASMKAAIAQILPNTVHRLCMWHIMEKV 1031
>gb|AAV31356.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
Length = 896
Score = 66.6 bits (161), Expect = 2e-09
Identities = 33/104 (31%), Positives = 56/104 (53%), Gaps = 2/104 (1%)
Query: 28 FNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCK 87
++ F VL +DSTY TN Y M GV + +G F+ EK E+FVW+ Q K
Sbjct: 321 YSVFGDVLSVDSTYSTNQYNMKFVPFTGVNHHLQSVFLGASFLADEKIESFVWLFQTFLK 380
Query: 88 LLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNV 131
+ P++++TD+D ++ A++ +LP + C +H+ + V
Sbjct: 381 --ATGGVAPRLIITDEDASMKAAIAQILPNTVHRLCMWHIMEKV 422
>gb|AAW41059.1| hypothetical protein CNA05530 [Cryptococcus neoformans var.
neoformans JEC21] gi|58258931|ref|XP_566878.1|
hypothetical protein CNA05530 [Cryptococcus neoformans
var. neoformans JEC21]
Length = 206
Score = 66.2 bits (160), Expect = 2e-09
Identities = 34/85 (40%), Positives = 55/85 (64%), Gaps = 2/85 (2%)
Query: 48 MPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCKLLTSRMNVPKVVVTDKDTTL 107
MPM IVG TST +TY+ G + E + L +L+ + +V KVV+TD+D L
Sbjct: 1 MPMLHIVGSTSTGMTYTAGVILMLRETTNWYTQALNSFFELV-GKPDV-KVVITDRDPAL 58
Query: 108 MNAVSNVLPESFAMNCYFHVQKNVK 132
+NA+ +VLP+++ +C++H+Q+NVK
Sbjct: 59 INALMSVLPKAYRFSCFWHLQENVK 83
>gb|AAL58182.1| putative far-red impaired response protein [Oryza sativa (japonica
cultivar-group)] gi|31433683|gb|AAP55167.1| putative
far-red impaired response protein [Oryza sativa
(japonica cultivar-group)] gi|37537156|ref|NP_922881.1|
putative far-red impaired response protein [Oryza sativa
(japonica cultivar-group)]
Length = 721
Score = 65.9 bits (159), Expect = 3e-09
Identities = 30/105 (28%), Positives = 57/105 (53%), Gaps = 2/105 (1%)
Query: 28 FNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCK 87
+ F V++ DSTY+ N Y +P VG T G +++E ++VW+LQ + +
Sbjct: 237 YEAFSDVVIFDSTYRVNRYNLPFVPFVGANHHRSTVIFGCGILSNESVSSYVWLLQTLLE 296
Query: 88 LLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVK 132
+ + PK ++TD D ++ A+ V+P + C +H+++N+K
Sbjct: 297 AMHQKH--PKSLITDGDASMAKAIRKVMPNTDHRLCSWHIEENMK 339
>ref|XP_482508.1| putative far-red impaired response protein [Oryza sativa (japonica
cultivar-group)] gi|25553698|dbj|BAC24942.1| putative
far-red impaired response protein [Oryza sativa
(japonica cultivar-group)]
Length = 1148
Score = 65.1 bits (157), Expect = 5e-09
Identities = 30/103 (29%), Positives = 53/103 (51%), Gaps = 2/103 (1%)
Query: 27 FFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMC 86
++ + V+ D+TY TN Y +P VG+ T G F+ E E F W+ +
Sbjct: 587 WYEKYGDVVSFDTTYFTNKYNLPFAPFVGIYGHGNTIVFGCAFLHDETSETFKWLFRTFL 646
Query: 87 KLLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQK 129
K ++ + PK ++TD+D + +A++ V + NC+FH+ K
Sbjct: 647 KAMSQK--EPKTIITDQDGAMRSAIAQVFQNAKHRNCFFHIVK 687
>ref|XP_466614.1| putative far-red impaired response protein [Oryza sativa (japonica
cultivar-group)] gi|47497275|dbj|BAD19318.1| putative
far-red impaired response protein [Oryza sativa
(japonica cultivar-group)]
Length = 880
Score = 65.1 bits (157), Expect = 5e-09
Identities = 31/104 (29%), Positives = 54/104 (51%), Gaps = 2/104 (1%)
Query: 28 FNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCK 87
++ F V+V DSTY+ N Y +P VGV T G V EK + W+L+
Sbjct: 278 YDAFGDVVVFDSTYRVNRYNLPFIPFVGVNHHGSTIIFGCAIVADEKVATYEWILKQFLD 337
Query: 88 LLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNV 131
+ + P+ ++TD D + A++ V+P+S C +H+++N+
Sbjct: 338 CMYQKH--PRALITDGDNAMRRAIAAVMPDSEHWLCTWHIEQNM 379
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.342 0.150 0.517
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 774,338,011
Number of Sequences: 2540612
Number of extensions: 30445816
Number of successful extensions: 103140
Number of sequences better than 10.0: 326
Number of HSP's better than 10.0 without gapping: 114
Number of HSP's successfully gapped in prelim test: 212
Number of HSP's that attempted gapping in prelim test: 102863
Number of HSP's gapped (non-prelim): 382
length of query: 484
length of database: 863,360,394
effective HSP length: 132
effective length of query: 352
effective length of database: 527,999,610
effective search space: 185855862720
effective search space used: 185855862720
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (22.0 bits)
S2: 77 (34.3 bits)
Medicago: description of AC140031.2


