

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139748.4 + phase: 0 /pseudo
(36 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
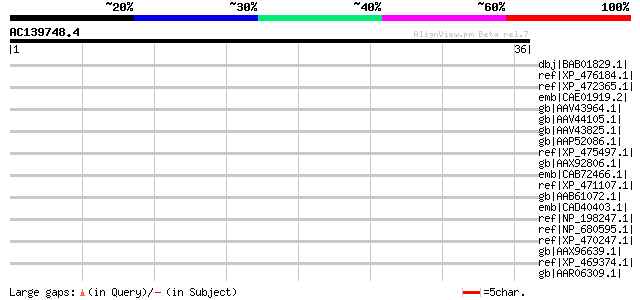
Score E
Sequences producing significant alignments: (bits) Value
dbj|BAB01829.1| unnamed protein product [Arabidopsis thaliana] 42 0.006
ref|XP_476184.1| unknown protein [Oryza sativa (japonica cultiva... 41 0.010
ref|XP_472365.1| OSJNBb0012E08.4 [Oryza sativa (japonica cultiva... 40 0.018
emb|CAE01919.2| OSJNBb0078D11.1 [Oryza sativa (japonica cultivar... 40 0.018
gb|AAV43964.1| putative polyprotein [Oryza sativa (japonica cult... 40 0.018
gb|AAV44105.1| unknown protein [Oryza sativa (japonica cultivar-... 40 0.018
gb|AAV43825.1| putative polyprotein [Oryza sativa (japonica cult... 40 0.018
gb|AAP52086.1| putative transposase [Oryza sativa (japonica cult... 39 0.030
ref|XP_475497.1| unknown protein [Oryza sativa (japonica cultiva... 39 0.030
gb|AAX92806.1| transposon protein, putative, CACTA, En/Spm sub-c... 38 0.087
emb|CAB72466.1| putative protein [Arabidopsis thaliana] gi|11358... 37 0.11
ref|XP_471107.1| OSJNBa0042N22.14 [Oryza sativa (japonica cultiv... 37 0.11
gb|AAB61072.1| contains similarity to a DNAJ-like domain [Arabid... 37 0.11
emb|CAD40403.1| OSJNBa0004L19.21 [Oryza sativa (japonica cultiva... 37 0.15
ref|NP_198247.1| hypothetical protein [Arabidopsis thaliana] 34 0.97
ref|NP_680595.1| hypothetical protein [Arabidopsis thaliana] 34 0.97
ref|XP_470247.1| Unknown protein [Oryza sativa (japonica cultiva... 34 1.3
gb|AAX96639.1| transposon protein, putative, CACTA, En/Spm sub-c... 34 1.3
ref|XP_469374.1| hypothetical protein [Oryza sativa (japonica cu... 33 2.2
gb|AAR06309.1| hypothetical protein [Oryza sativa (japonica cult... 32 3.7
>dbj|BAB01829.1| unnamed protein product [Arabidopsis thaliana]
Length = 194
Score = 41.6 bits (96), Expect = 0.006
Identities = 14/24 (58%), Positives = 22/24 (91%)
Query: 3 LPNRIKNNPKYYPWFKNCIGAIDG 26
+P +IK+N ++YP+FK+C+GAIDG
Sbjct: 68 VPTKIKDNTRFYPYFKDCVGAIDG 91
>ref|XP_476184.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|53982300|gb|AAV25279.1| unknow protein [Oryza sativa
(japonica cultivar-group)] gi|48843769|gb|AAT47028.1|
unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 251
Score = 40.8 bits (94), Expect = 0.010
Identities = 14/21 (66%), Positives = 20/21 (94%)
Query: 6 RIKNNPKYYPWFKNCIGAIDG 26
+I+ NP+++P+FKNCIGAIDG
Sbjct: 152 KIRTNPRFFPYFKNCIGAIDG 172
>ref|XP_472365.1| OSJNBb0012E08.4 [Oryza sativa (japonica cultivar-group)]
gi|38345208|emb|CAD40780.2| OSJNBb0012E08.4 [Oryza
sativa (japonica cultivar-group)]
Length = 261
Score = 40.0 bits (92), Expect = 0.018
Identities = 15/30 (50%), Positives = 23/30 (76%)
Query: 1 MELPNRIKNNPKYYPWFKNCIGAIDGLLLR 30
++ P +I NP++ P+FK+CIGAIDG +R
Sbjct: 110 LDTPTKIAGNPRWDPYFKDCIGAIDGTHIR 139
>emb|CAE01919.2| OSJNBb0078D11.1 [Oryza sativa (japonica cultivar-group)]
gi|38345857|emb|CAD41055.2| OSJNBa0084K11.21 [Oryza
sativa (japonica cultivar-group)]
gi|50927943|ref|XP_473499.1| OSJNBa0084K11.21 [Oryza
sativa (japonica cultivar-group)]
Length = 377
Score = 40.0 bits (92), Expect = 0.018
Identities = 15/30 (50%), Positives = 23/30 (76%)
Query: 1 MELPNRIKNNPKYYPWFKNCIGAIDGLLLR 30
++ P +I NP++ P+FK+CIGAIDG +R
Sbjct: 133 LDTPTKIAGNPRWDPYFKDCIGAIDGTHIR 162
>gb|AAV43964.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 561
Score = 40.0 bits (92), Expect = 0.018
Identities = 15/30 (50%), Positives = 23/30 (76%)
Query: 1 MELPNRIKNNPKYYPWFKNCIGAIDGLLLR 30
++ P +I NP++ P+FK+CIGAIDG +R
Sbjct: 110 LDTPTKIAGNPRWDPYFKDCIGAIDGTHIR 139
>gb|AAV44105.1| unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 1220
Score = 40.0 bits (92), Expect = 0.018
Identities = 15/30 (50%), Positives = 23/30 (76%)
Query: 1 MELPNRIKNNPKYYPWFKNCIGAIDGLLLR 30
++ P +I NP++ P+FK+CIGAIDG +R
Sbjct: 215 LDTPTKIAGNPRWDPYFKDCIGAIDGTHIR 244
>gb|AAV43825.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1067
Score = 40.0 bits (92), Expect = 0.018
Identities = 15/30 (50%), Positives = 23/30 (76%)
Query: 1 MELPNRIKNNPKYYPWFKNCIGAIDGLLLR 30
++ P +I NP++ P+FK+CIGAIDG +R
Sbjct: 62 LDTPTKIAGNPRWDPYFKDCIGAIDGTHIR 91
>gb|AAP52086.1| putative transposase [Oryza sativa (japonica cultivar-group)]
gi|37530994|ref|NP_919799.1| putative transposase [Oryza
sativa (japonica cultivar-group)]
gi|16519473|gb|AAL25182.1| Putative transposase [Oryza
sativa]
Length = 435
Score = 39.3 bits (90), Expect = 0.030
Identities = 13/21 (61%), Positives = 20/21 (94%)
Query: 6 RIKNNPKYYPWFKNCIGAIDG 26
+I+ +P++YP+FKNC+GAIDG
Sbjct: 157 KIEKDPRFYPYFKNCLGAIDG 177
>ref|XP_475497.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|51038051|gb|AAT93855.1| unknown protein [Oryza sativa
(japonica cultivar-group)] gi|48475221|gb|AAT44290.1|
unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 200
Score = 39.3 bits (90), Expect = 0.030
Identities = 13/21 (61%), Positives = 20/21 (94%)
Query: 6 RIKNNPKYYPWFKNCIGAIDG 26
+I+ +P++YP+FKNC+GAIDG
Sbjct: 157 KIEKDPRFYPYFKNCLGAIDG 177
>gb|AAX92806.1| transposon protein, putative, CACTA, En/Spm sub-class [Oryza sativa
(japonica cultivar-group)]
Length = 297
Score = 37.7 bits (86), Expect = 0.087
Identities = 14/23 (60%), Positives = 19/23 (81%)
Query: 4 PNRIKNNPKYYPWFKNCIGAIDG 26
P+ I P++YP+F+NCIGAIDG
Sbjct: 147 PHPILRKPQFYPFFQNCIGAIDG 169
>emb|CAB72466.1| putative protein [Arabidopsis thaliana] gi|11358076|pir||T47439
hypothetical protein T18B22.40 - Arabidopsis thaliana
Length = 377
Score = 37.4 bits (85), Expect = 0.11
Identities = 13/26 (50%), Positives = 21/26 (80%)
Query: 1 MELPNRIKNNPKYYPWFKNCIGAIDG 26
+ +P +I+ + +YP+FK+CIGAIDG
Sbjct: 127 VRVPAKIRESTSFYPYFKDCIGAIDG 152
>ref|XP_471107.1| OSJNBa0042N22.14 [Oryza sativa (japonica cultivar-group)]
gi|38569160|emb|CAE03671.3| OSJNBa0042N22.14 [Oryza
sativa (japonica cultivar-group)]
Length = 401
Score = 37.4 bits (85), Expect = 0.11
Identities = 15/27 (55%), Positives = 21/27 (77%)
Query: 4 PNRIKNNPKYYPWFKNCIGAIDGLLLR 30
P++I NP + P+FK+CIGAIDG +R
Sbjct: 101 PSKILGNPMWDPYFKDCIGAIDGTHVR 127
>gb|AAB61072.1| contains similarity to a DNAJ-like domain [Arabidopsis thaliana]
gi|7485307|pir||T01797 hypothetical protein A_TM021B04.9
- Arabidopsis thaliana
Length = 1609
Score = 37.4 bits (85), Expect = 0.11
Identities = 12/24 (50%), Positives = 20/24 (83%)
Query: 3 LPNRIKNNPKYYPWFKNCIGAIDG 26
+P++I ++YP+FK+C+GAIDG
Sbjct: 1204 VPSKISKTTRFYPYFKDCVGAIDG 1227
>emb|CAD40403.1| OSJNBa0004L19.21 [Oryza sativa (japonica cultivar-group)]
Length = 639
Score = 37.0 bits (84), Expect = 0.15
Identities = 13/17 (76%), Positives = 16/17 (93%)
Query: 10 NPKYYPWFKNCIGAIDG 26
NP+YYP+F +CIGAIDG
Sbjct: 500 NPRYYPYFNDCIGAIDG 516
>ref|NP_198247.1| hypothetical protein [Arabidopsis thaliana]
Length = 148
Score = 34.3 bits (77), Expect = 0.97
Identities = 11/23 (47%), Positives = 19/23 (81%)
Query: 3 LPNRIKNNPKYYPWFKNCIGAID 25
+P +I+ + + YP+FK+C+GAID
Sbjct: 8 VPRKIRESTRLYPYFKDCVGAID 30
>ref|NP_680595.1| hypothetical protein [Arabidopsis thaliana]
Length = 284
Score = 34.3 bits (77), Expect = 0.97
Identities = 10/20 (50%), Positives = 19/20 (95%)
Query: 7 IKNNPKYYPWFKNCIGAIDG 26
+KN+ +Y+P+F++C+GA+DG
Sbjct: 90 LKNDDRYWPFFRDCVGALDG 109
>ref|XP_470247.1| Unknown protein [Oryza sativa (japonica cultivar-group)]
gi|21397269|gb|AAM51833.1| Unknown protein [Oryza sativa
(japonica cultivar-group)]
Length = 1202
Score = 33.9 bits (76), Expect = 1.3
Identities = 12/19 (63%), Positives = 17/19 (89%)
Query: 12 KYYPWFKNCIGAIDGLLLR 30
K+YP+FK+CIGA+DG +R
Sbjct: 691 KFYPYFKDCIGALDGTHIR 709
>gb|AAX96639.1| transposon protein, putative, CACTA, En/Spm sub-class [Oryza sativa
(japonica cultivar-group)]
Length = 689
Score = 33.9 bits (76), Expect = 1.3
Identities = 11/21 (52%), Positives = 18/21 (85%)
Query: 6 RIKNNPKYYPWFKNCIGAIDG 26
+I NP+++P+FK+C+GAI G
Sbjct: 117 KIATNPRFFPFFKDCLGAIGG 137
>ref|XP_469374.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|27552543|gb|AAO19366.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 515
Score = 33.1 bits (74), Expect = 2.2
Identities = 11/17 (64%), Positives = 15/17 (87%)
Query: 10 NPKYYPWFKNCIGAIDG 26
NP++YP+F +CI AIDG
Sbjct: 267 NPRFYPYFNDCIAAIDG 283
>gb|AAR06309.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|50916453|ref|XP_468623.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 767
Score = 32.3 bits (72), Expect = 3.7
Identities = 11/17 (64%), Positives = 16/17 (93%)
Query: 10 NPKYYPWFKNCIGAIDG 26
N +++P+FK+CIGAIDG
Sbjct: 533 NRRFFPYFKDCIGAIDG 549
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.324 0.146 0.491
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 70,243,283
Number of Sequences: 2540612
Number of extensions: 1427524
Number of successful extensions: 2904
Number of sequences better than 10.0: 29
Number of HSP's better than 10.0 without gapping: 29
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 2875
Number of HSP's gapped (non-prelim): 29
length of query: 36
length of database: 863,360,394
effective HSP length: 12
effective length of query: 24
effective length of database: 832,873,050
effective search space: 19988953200
effective search space used: 19988953200
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 69 (31.2 bits)
Medicago: description of AC139748.4


