

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139743.4 - phase: 0 /pseudo
(105 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
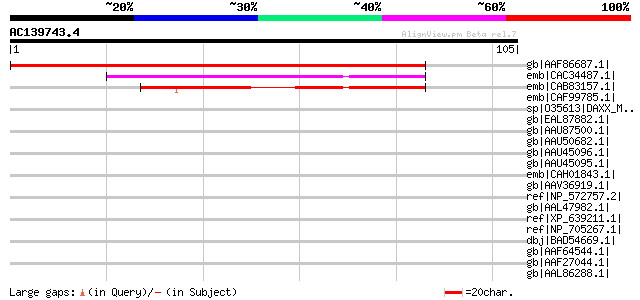
Score E
Sequences producing significant alignments: (bits) Value
gb|AAF86687.1| MTD1 [Medicago truncatula] 174 6e-43
emb|CAC34487.1| putative protein [Arabidopsis thaliana] gi|30023... 54 1e-06
emb|CAB83157.1| putative protein [Arabidopsis thaliana] gi|15229... 52 4e-06
emb|CAF99785.1| unnamed protein product [Tetraodon nigroviridis] 40 0.014
sp|O35613|DAXX_MOUSE Death domain-associated protein 6 (Daxx) gi... 40 0.018
gb|EAL87882.1| hypothetical protein Afu1g01700 [Aspergillus fumi... 39 0.040
gb|AAU87500.1| death-domain associated protein [Mus musculus] 39 0.040
gb|AAU50682.1| death domain-associated protein [Mus musculus] 39 0.040
gb|AAU45096.1| death domain-associated protein [Mus musculus] 39 0.040
gb|AAU45095.1| death domain-associated protein [Mus musculus] 39 0.040
emb|CAH01843.1| unnamed protein product [Kluyveromyces lactis NR... 37 0.12
gb|AAV36919.1| RE02581p [Drosophila melanogaster] 37 0.12
ref|NP_572757.2| CG2025-PA [Drosophila melanogaster] gi|22832115... 37 0.12
gb|AAL47982.1| GH13968p [Drosophila melanogaster] 37 0.12
ref|XP_639211.1| putative DNA polymerase epsilon subunit A [Dict... 37 0.12
ref|NP_705267.1| hypothetical protein [Plasmodium falciparum 3D7... 37 0.15
dbj|BAD54669.1| unknown protein [Oryza sativa (japonica cultivar... 37 0.15
gb|AAF64544.1| unknown protein [Arabidopsis thaliana] gi|1522997... 36 0.20
gb|AAF27044.1| unknown protein, 3' partial [Arabidopsis thaliana] 36 0.20
gb|AAL86288.1| unknown protein [Arabidopsis thaliana] 36 0.20
>gb|AAF86687.1| MTD1 [Medicago truncatula]
Length = 231
Score = 174 bits (440), Expect = 6e-43
Identities = 86/86 (100%), Positives = 86/86 (100%)
Query: 1 MELETVAFKRVGSRTSVLFDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMD 60
MELETVAFKRVGSRTSVLFDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMD
Sbjct: 1 MELETVAFKRVGSRTSVLFDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMD 60
Query: 61 ENEAESKYNGGALDCMEALEEVLPIR 86
ENEAESKYNGGALDCMEALEEVLPIR
Sbjct: 61 ENEAESKYNGGALDCMEALEEVLPIR 86
>emb|CAC34487.1| putative protein [Arabidopsis thaliana]
gi|30023750|gb|AAP13408.1| At5g21940 [Arabidopsis
thaliana] gi|22531283|gb|AAM97145.1| unknown protein
[Arabidopsis thaliana] gi|29294065|gb|AAO73902.1|
expressed protein [Arabidopsis thaliana]
gi|22326936|ref|NP_680182.1| expressed protein
[Arabidopsis thaliana]
Length = 264
Score = 53.5 bits (127), Expect = 1e-06
Identities = 29/66 (43%), Positives = 38/66 (56%), Gaps = 1/66 (1%)
Query: 21 APRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGGALDCMEALE 80
+P D S+++SSSIGRNSDD + ENE ES Y G L+ ME+LE
Sbjct: 24 SPSPSPSDSSSSPSSSASSSIGRNSDDGEKSSEDGGDDAGENEVESPYK-GPLEMMESLE 82
Query: 81 EVLPIR 86
+VLP+R
Sbjct: 83 QVLPVR 88
>emb|CAB83157.1| putative protein [Arabidopsis thaliana]
gi|15229793|ref|NP_189971.1| hypothetical protein
[Arabidopsis thaliana] gi|11290088|pir||T47421
hypothetical protein T28A8.140 - Arabidopsis thaliana
Length = 270
Score = 52.0 bits (123), Expect = 4e-06
Identities = 33/61 (54%), Positives = 37/61 (60%), Gaps = 12/61 (19%)
Query: 28 DQGKVC--STTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGGALDCMEALEEVLPI 85
DQ C S+TSS SIG NSDDD+ ENE ES YN G LD ME+LEE LPI
Sbjct: 15 DQEFACLSSSTSSDSIGENSDDDEG---------GENEIESSYN-GPLDMMESLEEALPI 64
Query: 86 R 86
+
Sbjct: 65 K 65
>emb|CAF99785.1| unnamed protein product [Tetraodon nigroviridis]
Length = 1679
Score = 40.0 bits (92), Expect = 0.014
Identities = 32/107 (29%), Positives = 52/107 (47%), Gaps = 16/107 (14%)
Query: 2 ELETVAFKRVGSRTSVLFDAPRDHDDDQGKVCSTTSSSS----IGRNSDDDDDDEVSSER 57
+L T+ KR G R +A +++Q VC+ +S++ +G +SDDDDDD+ S+
Sbjct: 692 KLRTIYAKRTGKR-----EANMGVEENQS-VCTPSSAAKRKRKMGGDSDDDDDDDDDSDD 745
Query: 58 SMDENEAESKYNGGALDCMEALEEVLPIRLFQSFMVMQEKYLKFLQW 104
D+ E E + + +E +EVL + F YLK W
Sbjct: 746 QPDDEEEEEEEETKKVKKVETNDEVLVAPSPRIF------YLKLCSW 786
>sp|O35613|DAXX_MOUSE Death domain-associated protein 6 (Daxx) gi|2253707|gb|AAC53284.1|
Daxx [Mus musculus] gi|6681135|ref|NP_031855.1| Fas
death domain-associated protein [Mus musculus]
Length = 739
Score = 39.7 bits (91), Expect = 0.018
Identities = 28/95 (29%), Positives = 42/95 (43%), Gaps = 13/95 (13%)
Query: 1 MELETVAFKRVGSRTSVLFDAPRDHDDDQG-------------KVCSTTSSSSIGRNSDD 47
M+ +T +R R +L AP+ D Q + C TTS + + DD
Sbjct: 388 MQDKTEEGERQKRRARLLGTAPQPSDPPQASSESGEGPSGMASQECPTTSKAETDDDDDD 447
Query: 48 DDDDEVSSERSMDENEAESKYNGGALDCMEALEEV 82
DDDD+ +E S +E E E + D E LE++
Sbjct: 448 DDDDDEDNEESEEEEEEEEEEKEATEDEDEDLEQL 482
>gb|EAL87882.1| hypothetical protein Afu1g01700 [Aspergillus fumigatus Af293]
Length = 1044
Score = 38.5 bits (88), Expect = 0.040
Identities = 21/56 (37%), Positives = 27/56 (47%)
Query: 18 LFDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGGAL 73
+F P D D D+ + S++SSSS SD D D E S S D + S G L
Sbjct: 983 IFSPPSDDDSDESESSSSSSSSSSESESDKDSDSENESHSSADHGDIMSSGTVGKL 1038
>gb|AAU87500.1| death-domain associated protein [Mus musculus]
Length = 651
Score = 38.5 bits (88), Expect = 0.040
Identities = 19/50 (38%), Positives = 27/50 (54%)
Query: 33 CSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGGALDCMEALEEV 82
C TTS + + DDDDDD+ +E S +E E E + D E LE++
Sbjct: 345 CPTTSKAETDDDDDDDDDDDEDNEESEEEEEEEEEEKEATEDEDEDLEQL 394
>gb|AAU50682.1| death domain-associated protein [Mus musculus]
Length = 90
Score = 38.5 bits (88), Expect = 0.040
Identities = 19/50 (38%), Positives = 27/50 (54%)
Query: 33 CSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGGALDCMEALEEV 82
C TTS + + DDDDDD+ +E S +E E E + D E LE++
Sbjct: 16 CPTTSKAETDDDDDDDDDDDEDNEESEEEEEEEEEEKEATEDEDEDLEQL 65
>gb|AAU45096.1| death domain-associated protein [Mus musculus]
Length = 715
Score = 38.5 bits (88), Expect = 0.040
Identities = 19/50 (38%), Positives = 27/50 (54%)
Query: 33 CSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGGALDCMEALEEV 82
C TTS + + DDDDDD+ +E S +E E E + D E LE++
Sbjct: 409 CPTTSKAETDDDDDDDDDDDEDNEESEEEEEEEEEEKEATEDEDEDLEQL 458
>gb|AAU45095.1| death domain-associated protein [Mus musculus]
Length = 621
Score = 38.5 bits (88), Expect = 0.040
Identities = 19/50 (38%), Positives = 27/50 (54%)
Query: 33 CSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGGALDCMEALEEV 82
C TTS + + DDDDDD+ +E S +E E E + D E LE++
Sbjct: 315 CPTTSKAETDDDDDDDDDDDEDNEESEEEEEEEEEEKEATEDEDEDLEQL 364
>emb|CAH01843.1| unnamed protein product [Kluyveromyces lactis NRRL Y-1140]
gi|50306061|ref|XP_452992.1| unnamed protein product
[Kluyveromyces lactis]
Length = 1194
Score = 37.0 bits (84), Expect = 0.12
Identities = 21/65 (32%), Positives = 34/65 (52%), Gaps = 1/65 (1%)
Query: 3 LETVAFKRVGSRTSV-LFDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDE 61
+ETV R R +V L ++ D+DDD G + + S + + DDD +DE + DE
Sbjct: 97 VETVTSLRGRKRKNVSLRESDDDNDDDDGDISMSKRSRNRSKIEDDDSEDEYKDDGDDDE 156
Query: 62 NEAES 66
E ++
Sbjct: 157 EEEDN 161
>gb|AAV36919.1| RE02581p [Drosophila melanogaster]
Length = 1147
Score = 37.0 bits (84), Expect = 0.12
Identities = 17/44 (38%), Positives = 22/44 (49%)
Query: 27 DDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNG 70
+D K S + S N DDDD+DE E S +E E E + G
Sbjct: 1034 EDTAKCASKSDDESESENDDDDDEDEEEEESSEEEEEEEKEKEG 1077
Score = 32.7 bits (73), Expect = 2.2
Identities = 14/39 (35%), Positives = 20/39 (50%)
Query: 33 CSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGG 71
C++ S +DDDDD++ E S +E E E K G
Sbjct: 1039 CASKSDDESESENDDDDDEDEEEEESSEEEEEEEKEKEG 1077
>ref|NP_572757.2| CG2025-PA [Drosophila melanogaster] gi|22832115|gb|AAF48105.2|
CG2025-PA [Drosophila melanogaster]
Length = 1147
Score = 37.0 bits (84), Expect = 0.12
Identities = 17/44 (38%), Positives = 22/44 (49%)
Query: 27 DDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNG 70
+D K S + S N DDDD+DE E S +E E E + G
Sbjct: 1034 EDTAKCASKSDDESESENDDDDDEDEEEEESSEEEEEEEKEKEG 1077
Score = 32.7 bits (73), Expect = 2.2
Identities = 14/39 (35%), Positives = 20/39 (50%)
Query: 33 CSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGG 71
C++ S +DDDDD++ E S +E E E K G
Sbjct: 1039 CASKSDDESESENDDDDDEDEEEEESSEEEEEEEKEKEG 1077
>gb|AAL47982.1| GH13968p [Drosophila melanogaster]
Length = 343
Score = 37.0 bits (84), Expect = 0.12
Identities = 17/44 (38%), Positives = 22/44 (49%)
Query: 27 DDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNG 70
+D K S + S N DDDD+DE E S +E E E + G
Sbjct: 230 EDTAKCASKSDDESESENDDDDDEDEEEEESSEEEEEEEKEKEG 273
Score = 32.7 bits (73), Expect = 2.2
Identities = 14/39 (35%), Positives = 20/39 (50%)
Query: 33 CSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGG 71
C++ S +DDDDD++ E S +E E E K G
Sbjct: 235 CASKSDDESESENDDDDDEDEEEEESSEEEEEEEKEKEG 273
>ref|XP_639211.1| putative DNA polymerase epsilon subunit A [Dictyostelium discoideum]
gi|60467818|gb|EAL65833.1| putative DNA polymerase
epsilon subunit A [Dictyostelium discoideum]
Length = 2380
Score = 37.0 bits (84), Expect = 0.12
Identities = 21/65 (32%), Positives = 35/65 (53%), Gaps = 3/65 (4%)
Query: 44 NSDDDDDDEVS---SERSMDENEAESKYNGGALDCMEALEEVLPIRLFQSFMVMQEKYLK 100
N+DDDDDD+VS E+ ++N+ K G + + E LP ++ SF+++ Y+
Sbjct: 1975 NADDDDDDDVSENEEEQQQNKNKKLKKIKNGKIITNWNIAEFLPPQIQTSFIIIISDYIY 2034
Query: 101 FLQWE 105
L E
Sbjct: 2035 KLHTE 2039
>ref|NP_705267.1| hypothetical protein [Plasmodium falciparum 3D7]
gi|23615512|emb|CAD52504.1| hypothetical protein
[Plasmodium falciparum 3D7]
Length = 158
Score = 36.6 bits (83), Expect = 0.15
Identities = 16/50 (32%), Positives = 27/50 (54%)
Query: 20 DAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYN 69
D D DDD + TT++ + + DDDDDD+ + D+++ E + N
Sbjct: 90 DDDDDDDDDDDESIDTTNNDNDDNDDDDDDDDDDDDDDDDDDDDDEDEIN 139
>dbj|BAD54669.1| unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 228
Score = 36.6 bits (83), Expect = 0.15
Identities = 24/61 (39%), Positives = 34/61 (55%), Gaps = 13/61 (21%)
Query: 39 SSIGRNSDDDDDD---EVSSERSMDENEAESKYN----------GGALDCMEALEEVLPI 85
SSIG +S+D+ DD EV S+ S AE+ GGAL C++AL++ LPI
Sbjct: 85 SSIGADSEDEQDDGEEEVESKASSAAVAAETCRRKTKTKCGGGGGGALACLDALDDALPI 144
Query: 86 R 86
+
Sbjct: 145 K 145
>gb|AAF64544.1| unknown protein [Arabidopsis thaliana] gi|15229976|ref|NP_187189.1|
myb family transcription factor [Arabidopsis thaliana]
Length = 1055
Score = 36.2 bits (82), Expect = 0.20
Identities = 30/96 (31%), Positives = 45/96 (46%), Gaps = 9/96 (9%)
Query: 9 KRVGSRTSVLFDAPRDHDDDQGKVCSTTS---SSSIGRNSDDDDDDEVSSERSMDENEAE 65
KRV + + +A + DD G+ CS T S S R + + E S RS + +
Sbjct: 315 KRVYKKRVKVEEAECNDSDDNGEACSATQGLRSKSQRRKAAIEASREKYSPRS--PKKRD 372
Query: 66 SKYNGGALDCMEALEE----VLPIRLFQSFMVMQEK 97
K+ GA D ++AL E +LP L +S + Q K
Sbjct: 373 DKHTSGAFDALQALAELSASMLPANLMESELSAQLK 408
>gb|AAF27044.1| unknown protein, 3' partial [Arabidopsis thaliana]
Length = 618
Score = 36.2 bits (82), Expect = 0.20
Identities = 30/96 (31%), Positives = 45/96 (46%), Gaps = 9/96 (9%)
Query: 9 KRVGSRTSVLFDAPRDHDDDQGKVCSTTS---SSSIGRNSDDDDDDEVSSERSMDENEAE 65
KRV + + +A + DD G+ CS T S S R + + E S RS + +
Sbjct: 315 KRVYKKRVKVEEAECNDSDDNGEACSATQGLRSKSQRRKAAIEASREKYSPRS--PKKRD 372
Query: 66 SKYNGGALDCMEALEE----VLPIRLFQSFMVMQEK 97
K+ GA D ++AL E +LP L +S + Q K
Sbjct: 373 DKHTSGAFDALQALAELSASMLPANLMESELSAQLK 408
>gb|AAL86288.1| unknown protein [Arabidopsis thaliana]
Length = 820
Score = 36.2 bits (82), Expect = 0.20
Identities = 30/96 (31%), Positives = 45/96 (46%), Gaps = 9/96 (9%)
Query: 9 KRVGSRTSVLFDAPRDHDDDQGKVCSTTS---SSSIGRNSDDDDDDEVSSERSMDENEAE 65
KRV + + +A + DD G+ CS T S S R + + E S RS + +
Sbjct: 80 KRVYKKRVKVEEAECNDSDDNGEACSATQGLRSKSQRRKAAIEASREKYSPRS--PKKRD 137
Query: 66 SKYNGGALDCMEALEE----VLPIRLFQSFMVMQEK 97
K+ GA D ++AL E +LP L +S + Q K
Sbjct: 138 DKHTSGAFDALQALAELSASMLPANLMESELSAQLK 173
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.312 0.129 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 173,879,433
Number of Sequences: 2540612
Number of extensions: 6577398
Number of successful extensions: 76668
Number of sequences better than 10.0: 432
Number of HSP's better than 10.0 without gapping: 218
Number of HSP's successfully gapped in prelim test: 234
Number of HSP's that attempted gapping in prelim test: 73310
Number of HSP's gapped (non-prelim): 2024
length of query: 105
length of database: 863,360,394
effective HSP length: 81
effective length of query: 24
effective length of database: 657,570,822
effective search space: 15781699728
effective search space used: 15781699728
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 68 (30.8 bits)
Medicago: description of AC139743.4


