

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139706.5 - phase: 0
(424 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
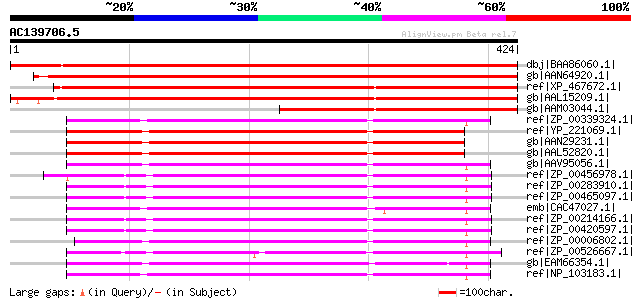
Score E
Sequences producing significant alignments: (bits) Value
dbj|BAA86060.1| senescencs-related protein [Pyrus pyrifolia] 726 0.0
gb|AAN64920.1| putative dehydrogenase [Lycopersicon esculentum] 702 0.0
ref|XP_467672.1| putative dihydropyrimidine dehydrogenase [Oryza... 676 0.0
gb|AAL15209.1| putative dehydrogenase [Arabidopsis thaliana] gi|... 674 0.0
gb|AAM03044.1| putative dihydroorotate dehydrogenase [Oryza sativa] 341 3e-92
ref|ZP_00339324.1| COG0167: Dihydroorotate dehydrogenase [Silici... 285 2e-75
ref|YP_221069.1| dihydroorotate dehydrogenase family protein [Br... 280 8e-74
gb|AAN29231.1| dihydroorotate dehydrogenase family protein [Bruc... 280 8e-74
gb|AAL52820.1| DIHYDROPYRIMIDINE DEHYDROGENASE (NADP+) [Brucella... 278 2e-73
gb|AAV95056.1| dihydroorotate dehydrogenase family protein [Sili... 277 5e-73
ref|ZP_00456978.1| Dihydroorotate dehydrogenase 1 [Burkholderia ... 275 3e-72
ref|ZP_00283910.1| COG0167: Dihydroorotate dehydrogenase [Burkho... 275 3e-72
ref|ZP_00465097.1| Dihydroorotate dehydrogenase 1 [Burkholderia ... 273 7e-72
emb|CAC47027.1| PUTATIVE OXIDOREDUCTASE IRON-SULFUR PROTEIN [Sin... 273 7e-72
ref|ZP_00214166.1| COG0167: Dihydroorotate dehydrogenase [Burkho... 273 1e-71
ref|ZP_00420597.1| Dihydroorotate dehydrogenase 1 [Burkholderia ... 270 6e-71
ref|ZP_00006802.1| COG0167: Dihydroorotate dehydrogenase [Rhodob... 267 4e-70
ref|ZP_00526667.1| Dihydroorotate dehydrogenase 1 [Solibacter us... 267 5e-70
gb|EAM66354.1| Dihydroorotate dehydrogenase 1 [Jannaschia sp. CCS1] 266 7e-70
ref|NP_103183.1| probable oxidoreductase [Mesorhizobium loti MAF... 266 9e-70
>dbj|BAA86060.1| senescencs-related protein [Pyrus pyrifolia]
Length = 424
Score = 726 bits (1875), Expect = 0.0
Identities = 361/425 (84%), Positives = 391/425 (91%), Gaps = 2/425 (0%)
Query: 1 MANLSMTQLKTRNSASRFSFNFSKKV-YPRPSRVDFKVFASEGQVSEPDLSVKVNGLHMP 59
MA++S TQ+ +N + FS +V RPSRV F+VFASE + +EPDLSV VNGLHMP
Sbjct: 1 MASISFTQIGGKNPVAEFSRRPRPEVRLTRPSRVGFRVFASESK-AEPDLSVTVNGLHMP 59
Query: 60 NPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYARLRANANGSAKG 119
NPFVIGSGPPGTNYTVMKRAFDEGWG VIAKTVSL+A KV NVTPRYARLR + NGSAKG
Sbjct: 60 NPFVIGSGPPGTNYTVMKRAFDEGWGAVIAKTVSLEADKVKNVTPRYARLRVDGNGSAKG 119
Query: 120 EIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAWEELIDRVEQTGI 179
+IIGW+NIELISDRPL+ MLKEFKQLK+EYPDRILIASIMEEYNKA WEELIDRVEQTG+
Sbjct: 120 QIIGWENIELISDRPLDIMLKEFKQLKQEYPDRILIASIMEEYNKAGWEELIDRVEQTGV 179
Query: 180 DAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKMTPNITDISQPAR 239
DA+EINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKMTPNITDI+QPAR
Sbjct: 180 DALEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKMTPNITDITQPAR 239
Query: 240 VALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAVHPIALGKVMSIAK 299
VALSSGCEGV+AINTIMSVMGINL TLRPEPCVEGYSTPGGYSAKAVHPIAL KVM+IAK
Sbjct: 240 VALSSGCEGVSAINTIMSVMGINLTTLRPEPCVEGYSTPGGYSAKAVHPIALAKVMNIAK 299
Query: 300 MMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYGLVKKLSLELKDF 359
+MKSEF +YSLS IGGVE+GGDAAEFILLGANTVQVCTGVMMHGYG+VKKL EL+DF
Sbjct: 300 LMKSEFGDTDYSLSGIGGVESGGDAAEFILLGANTVQVCTGVMMHGYGVVKKLCSELQDF 359
Query: 360 MKKHNFTSIEDFRGASLQYFTTHTDLVHRQQEAIKQRKAIKKGLQSDKDWTGDGFVKESE 419
MK+HNF+SIEDFRGASL+YFTTHTDLV RQQEAIKQRKAI+KGLQSDKDWTGDGFVKE+E
Sbjct: 360 MKQHNFSSIEDFRGASLEYFTTHTDLVRRQQEAIKQRKAIRKGLQSDKDWTGDGFVKETE 419
Query: 420 SMVSN 424
SMVSN
Sbjct: 420 SMVSN 424
>gb|AAN64920.1| putative dehydrogenase [Lycopersicon esculentum]
Length = 429
Score = 702 bits (1813), Expect = 0.0
Identities = 347/404 (85%), Positives = 374/404 (91%), Gaps = 6/404 (1%)
Query: 21 NFSKKVYPRPSRVDFKVFASEGQVSEPDLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAF 80
NF +K RV F++ A EGQ EPDLSV VNGL MPNPFVIGSGPPGTNYTVMKRAF
Sbjct: 32 NFGRK------RVGFRIMALEGQSVEPDLSVTVNGLKMPNPFVIGSGPPGTNYTVMKRAF 85
Query: 81 DEGWGGVIAKTVSLDAAKVINVTPRYARLRANANGSAKGEIIGWQNIELISDRPLETMLK 140
DEGWGGVIAKTVSL+A KV NVTPRYA+LRA+ANGSAKG+IIGWQNIELISDRPLETMLK
Sbjct: 86 DEGWGGVIAKTVSLEADKVKNVTPRYAKLRADANGSAKGQIIGWQNIELISDRPLETMLK 145
Query: 141 EFKQLKEEYPDRILIASIMEEYNKAAWEELIDRVEQTGIDAIEINFSCPHGMPERKMGAA 200
EFKQLKEEYPDRILIASIMEEYNKAAWEELI R E+TGIDA EINFSCPHGMPER+MGAA
Sbjct: 146 EFKQLKEEYPDRILIASIMEEYNKAAWEELIYRCEETGIDAFEINFSCPHGMPERRMGAA 205
Query: 201 VGQDCALLEEVCGWINAKATVPVWAKMTPNITDISQPARVALSSGCEGVAAINTIMSVMG 260
VGQDC LLEEVCGWINA ATVPVWAKMTPNITDI++PARVAL+ GCEGV+AINTIMSVMG
Sbjct: 206 VGQDCDLLEEVCGWINAVATVPVWAKMTPNITDITKPARVALNQGCEGVSAINTIMSVMG 265
Query: 261 INLNTLRPEPCVEGYSTPGGYSAKAVHPIALGKVMSIAKMMKSEFDSENYSLSAIGGVET 320
INL+TLRPEPCVEGYSTPGGYS+KAVHPIAL KVM+IA+MMKSEF ++YSLSAIGGVET
Sbjct: 266 INLDTLRPEPCVEGYSTPGGYSSKAVHPIALAKVMNIARMMKSEFGDKDYSLSAIGGVET 325
Query: 321 GGDAAEFILLGANTVQVCTGVMMHGYGLVKKLSLELKDFMKKHNFTSIEDFRGASLQYFT 380
GGDAAEFILLGA+TVQVCTGVMMHGYGLVK L ELKDFM+KHNF+SIEDFRG SL+YFT
Sbjct: 326 GGDAAEFILLGADTVQVCTGVMMHGYGLVKTLCSELKDFMRKHNFSSIEDFRGTSLEYFT 385
Query: 381 THTDLVHRQQEAIKQRKAIKKGLQSDKDWTGDGFVKESESMVSN 424
THTDLV RQQEAI+QRKA+KKGLQSDKDWTGDGFV+E+ESMVSN
Sbjct: 386 THTDLVRRQQEAIRQRKAVKKGLQSDKDWTGDGFVQETESMVSN 429
>ref|XP_467672.1| putative dihydropyrimidine dehydrogenase [Oryza sativa (japonica
cultivar-group)] gi|46390439|dbj|BAD15901.1| putative
dihydropyrimidine dehydrogenase [Oryza sativa (japonica
cultivar-group)]
Length = 414
Score = 676 bits (1744), Expect = 0.0
Identities = 333/388 (85%), Positives = 359/388 (91%), Gaps = 2/388 (0%)
Query: 37 VFASEGQVSEPDLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDA 96
V AS G EPDLSV+VNGL MPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDA
Sbjct: 29 VRASAG-AGEPDLSVRVNGLKMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDA 87
Query: 97 AKVINVTPRYARLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIA 156
KVINVTPRYARLRA+ NGS K IIGWQNIELISDRPLETML EFKQLK+EYPDRILI
Sbjct: 88 EKVINVTPRYARLRADPNGSTKSPIIGWQNIELISDRPLETMLNEFKQLKKEYPDRILIG 147
Query: 157 SIMEEYNKAAWEELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWIN 216
SIMEEYNKAAW ELI+RVE++G+DA+EINFSCPHGMPERKMGAAVGQDC LLEEVCGWIN
Sbjct: 148 SIMEEYNKAAWHELIERVEESGVDALEINFSCPHGMPERKMGAAVGQDCDLLEEVCGWIN 207
Query: 217 AKATVPVWAKMTPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYS 276
KATVPVWAKMTPNITDI++PAR++L SGCEGV+AINTIMSVMGINL TLRPEPCVEGYS
Sbjct: 208 EKATVPVWAKMTPNITDITKPARISLKSGCEGVSAINTIMSVMGINLKTLRPEPCVEGYS 267
Query: 277 TPGGYSAKAVHPIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQ 336
TPGGYSA+AVHPIAL KVM IA+MMK EF ++ SLSAIGGVETG DAAEFILLGA+TVQ
Sbjct: 268 TPGGYSARAVHPIALAKVMQIARMMKEEF-ADGQSLSAIGGVETGNDAAEFILLGADTVQ 326
Query: 337 VCTGVMMHGYGLVKKLSLELKDFMKKHNFTSIEDFRGASLQYFTTHTDLVHRQQEAIKQR 396
VCTGVMMHGYGLVKKL EL+DFM++HNF+SIEDFRGASL YFTTHTDLVHRQ+EAI QR
Sbjct: 327 VCTGVMMHGYGLVKKLCAELQDFMRQHNFSSIEDFRGASLPYFTTHTDLVHRQREAINQR 386
Query: 397 KAIKKGLQSDKDWTGDGFVKESESMVSN 424
KAI+KGL+SDKDWTGDGFVKE+ESMVSN
Sbjct: 387 KAIRKGLESDKDWTGDGFVKETESMVSN 414
>gb|AAL15209.1| putative dehydrogenase [Arabidopsis thaliana]
gi|14334712|gb|AAK59534.1| putative dehydrogenase
[Arabidopsis thaliana] gi|9294485|dbj|BAB02704.1|
senescencs-related protein; dihydroorotate
dehydrogenase-like protein [Arabidopsis thaliana]
gi|15229529|ref|NP_188408.1| dihydroorotate
dehydrogenase family protein / dihydroorotate oxidase
family protein [Arabidopsis thaliana]
gi|24850451|gb|AAN64919.1| putative dehydrogenase
[Arabidopsis thaliana]
Length = 426
Score = 674 bits (1739), Expect = 0.0
Identities = 338/429 (78%), Positives = 381/429 (88%), Gaps = 8/429 (1%)
Query: 1 MANLS--MTQLKTRNSASRFSFNF---SKKVYPRPSRVDFKVFASEGQVSEPDLSVKVNG 55
MA++S + + +S + S +F S++ + P+RV K+ S SEPDLSV VNG
Sbjct: 1 MASMSFALNRFSGLSSKTTLSADFDPSSRRSFLPPTRVGLKI--SSAAESEPDLSVTVNG 58
Query: 56 LHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYARLRANANG 115
L MPNPFVIGSGPPGTNYTVMKRAFDEGWG VIAKTVSLDA+KVINVTPRYARLR +NG
Sbjct: 59 LKMPNPFVIGSGPPGTNYTVMKRAFDEGWGAVIAKTVSLDASKVINVTPRYARLRTGSNG 118
Query: 116 SAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAWEELIDRVE 175
SAK ++IGWQNIELISDRPLETMLKEF++LK+EYPDRILIAS+MEEYNK AWEELIDRVE
Sbjct: 119 SAKTDVIGWQNIELISDRPLETMLKEFERLKKEYPDRILIASVMEEYNKTAWEELIDRVE 178
Query: 176 QTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKMTPNITDIS 235
QTG+DA+EINFSCPHGMPER+MGAAVGQDCALL+EVCGWINAKATVPVWAKMTPNITDI+
Sbjct: 179 QTGVDALEINFSCPHGMPERRMGAAVGQDCALLDEVCGWINAKATVPVWAKMTPNITDIT 238
Query: 236 QPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAVHPIALGKVM 295
+PARV+L SGCEG+AAINTIMSVMGI++ TLRPEPCVEGYSTPGGYS KAV PIAL KVM
Sbjct: 239 EPARVSLKSGCEGIAAINTIMSVMGIDMKTLRPEPCVEGYSTPGGYSYKAVRPIALAKVM 298
Query: 296 SIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYGLVKKLSLE 355
+IAKMMKSEF SE+ SLS IGGVETG DAAEFILLG+NTVQVCTGVMMHGYG VK L E
Sbjct: 299 NIAKMMKSEF-SEDRSLSGIGGVETGYDAAEFILLGSNTVQVCTGVMMHGYGHVKTLCAE 357
Query: 356 LKDFMKKHNFTSIEDFRGASLQYFTTHTDLVHRQQEAIKQRKAIKKGLQSDKDWTGDGFV 415
LKDFMK+HNF++IE+FRG SLQYFTTHTDLV RQ+EA++QRKA K+GL+SDKDWTGDGFV
Sbjct: 358 LKDFMKQHNFSTIEEFRGHSLQYFTTHTDLVKRQKEAVEQRKAEKRGLKSDKDWTGDGFV 417
Query: 416 KESESMVSN 424
KE+ESMVSN
Sbjct: 418 KETESMVSN 426
>gb|AAM03044.1| putative dihydroorotate dehydrogenase [Oryza sativa]
Length = 201
Score = 341 bits (874), Expect = 3e-92
Identities = 167/199 (83%), Positives = 186/199 (92%), Gaps = 1/199 (0%)
Query: 226 KMTPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKA 285
+MTPNITDI++PAR++L SGCEGV+AINTIMSVMGINL TLRPEPCVEGYSTPGGYSA+A
Sbjct: 4 QMTPNITDITRPARISLKSGCEGVSAINTIMSVMGINLKTLRPEPCVEGYSTPGGYSARA 63
Query: 286 VHPIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHG 345
VHPIAL KVM IA+MMK EF ++ SLSAIGGVETG DAAEFILLGA+TVQVCTGVMMHG
Sbjct: 64 VHPIALAKVMQIARMMKEEF-ADGQSLSAIGGVETGNDAAEFILLGADTVQVCTGVMMHG 122
Query: 346 YGLVKKLSLELKDFMKKHNFTSIEDFRGASLQYFTTHTDLVHRQQEAIKQRKAIKKGLQS 405
YGLVKKL EL+DFM++HNF+SIEDFRGASL YFTTHTDLVHRQ+EAI QRKAI+KGL+S
Sbjct: 123 YGLVKKLCAELQDFMRQHNFSSIEDFRGASLPYFTTHTDLVHRQREAINQRKAIRKGLES 182
Query: 406 DKDWTGDGFVKESESMVSN 424
DKDWTGDGFVKE+ESMVSN
Sbjct: 183 DKDWTGDGFVKETESMVSN 201
>ref|ZP_00339324.1| COG0167: Dihydroorotate dehydrogenase [Silicibacter sp. TM1040]
Length = 434
Score = 285 bits (728), Expect = 2e-75
Identities = 156/358 (43%), Positives = 216/358 (59%), Gaps = 12/358 (3%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVT-PRY 106
DL+ + G+ PNPF + S PP ++RAF+ GWGGV+ KT+ + V+NV PRY
Sbjct: 3 DLTTEFLGIKSPNPFWLASAPPTDKEYNVRRAFEAGWGGVVWKTLGAEGPPVVNVNGPRY 62
Query: 107 ARLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAA 166
+ A ++G NIELI+DRPL+ L+E ++K++YPDR LIASIM +AA
Sbjct: 63 GAI-----WGADRRLLGLNNIELITDRPLDVNLEEMTRVKKDYPDRALIASIMVPCEEAA 117
Query: 167 WEELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAK 226
W+ ++ RV +TG D IE+NF CPHGM ER MG+AVGQ ++ V W PV K
Sbjct: 118 WKAILPRVAETGCDGIELNFGCPHGMAERGMGSAVGQVPEYIQMVTEWCKQYYDKPVIVK 177
Query: 227 MTPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAV 286
+TPNITDI PAR A + + V+ INTI S+ +NL+ + PEP + G T GGY AV
Sbjct: 178 LTPNITDIRHPARAAKAGNADAVSLINTINSITSVNLDAMSPEPMIGGKGTHGGYCGPAV 237
Query: 287 HPIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGY 346
PIA+ V I++ + + +SAIGGV T DAAEFI LGA VQVCT M +G+
Sbjct: 238 KPIAMNMVAEISR----DPQTAGLPISAIGGVTTWRDAAEFIALGAGNVQVCTAAMTYGF 293
Query: 347 GLVKKLSLELKDFMKKHNFTSIEDFRGASLQYFT--THTDLVHRQQEAIKQRKAIKKG 402
+V+++ L D+M + ++SIEDFRG ++ T + DL + + I Q IK G
Sbjct: 294 KVVEEMISGLSDWMDEKGYSSIEDFRGMAVPNVTDWQYLDLNYVTKAKISQDDCIKCG 351
>ref|YP_221069.1| dihydroorotate dehydrogenase family protein [Brucella abortus
biovar 1 str. 9-941] gi|62195408|gb|AAX73708.1|
dihydroorotate dehydrogenase family protein [Brucella
abortus biovar 1 str. 9-941]
Length = 436
Score = 280 bits (715), Expect = 8e-74
Identities = 146/334 (43%), Positives = 207/334 (61%), Gaps = 10/334 (2%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVT-PRY 106
DLS G+ PNPF + S PP ++RAF GWGGV+ KT+ + V+NV PRY
Sbjct: 3 DLSTNFLGIKSPNPFWLASAPPTDKAYNVERAFKAGWGGVVWKTLGAEGPPVVNVNGPRY 62
Query: 107 ARLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAA 166
+ A ++G NIELI+DRPLE L+E K++K +PDR LIASIM + A
Sbjct: 63 GAIHG-----ADRRLLGLNNIELITDRPLEINLREMKEVKMRWPDRALIASIMVPCEEEA 117
Query: 167 WEELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAK 226
W+ ++ VE+TG D IE+NF CPHGM ER MGAAVGQ + V W + +PV K
Sbjct: 118 WKAILPLVEETGADGIELNFGCPHGMSERGMGAAVGQVPEYVGMVVKWCKQYSRMPVITK 177
Query: 227 MTPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAV 286
+TPNITDI +PAR A ++G + V+ INTI S++ ++L+T+ P P ++G + GGY AV
Sbjct: 178 LTPNITDIRKPARAAFANGTDAVSLINTINSIVSVDLDTMSPVPSIDGRGSHGGYCGPAV 237
Query: 287 HPIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGY 346
PIAL V IA+ + ++ +S IGG+ T DAAEFI LG TVQVCT M +G+
Sbjct: 238 KPIALNMVAEIAR----DAETAGLPISGIGGITTWRDAAEFIALGCGTVQVCTAAMTYGF 293
Query: 347 GLVKKLSLELKDFMKKHNFTSIEDFRGASLQYFT 380
+V+++ L D+M + +++DF+G ++ + T
Sbjct: 294 KVVQEMIDGLSDWMDSKGYQTLDDFQGCAVSHVT 327
>gb|AAN29231.1| dihydroorotate dehydrogenase family protein [Brucella suis 1330]
gi|23501189|ref|NP_697316.1| dihydroorotate
dehydrogenase family protein [Brucella suis 1330]
Length = 436
Score = 280 bits (715), Expect = 8e-74
Identities = 146/334 (43%), Positives = 207/334 (61%), Gaps = 10/334 (2%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVT-PRY 106
DLS G+ PNPF + S PP ++RAF GWGGV+ KT+ + V+NV PRY
Sbjct: 3 DLSTNFLGIKSPNPFWLASAPPTDKAYNVERAFKAGWGGVVWKTLGAEGPPVVNVNDPRY 62
Query: 107 ARLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAA 166
+ A ++G NIELI+DRPLE L+E K++K +PDR LIASIM + A
Sbjct: 63 GAIHG-----ADRRLLGLNNIELITDRPLEINLREMKEVKMRWPDRALIASIMVPCEEEA 117
Query: 167 WEELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAK 226
W+ ++ VE+TG D IE+NF CPHGM ER MGAAVGQ + V W + +PV K
Sbjct: 118 WKAILPLVEETGADGIELNFGCPHGMSERGMGAAVGQVPEYVGMVVKWCKQYSRMPVITK 177
Query: 227 MTPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAV 286
+TPNITDI +PAR A ++G + V+ INTI S++ ++L+T+ P P ++G + GGY AV
Sbjct: 178 LTPNITDIRKPARAAFANGTDAVSLINTINSIVSVDLDTMSPVPSIDGRGSHGGYCGPAV 237
Query: 287 HPIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGY 346
PIAL V IA+ + ++ +S IGG+ T DAAEFI LG TVQVCT M +G+
Sbjct: 238 KPIALNMVAEIAR----DAETAGLPISGIGGITTWRDAAEFIALGCGTVQVCTAAMTYGF 293
Query: 347 GLVKKLSLELKDFMKKHNFTSIEDFRGASLQYFT 380
+V+++ L D+M + +++DF+G ++ + T
Sbjct: 294 KVVQEMIDGLSDWMDSKGYQTLDDFQGCAVSHVT 327
>gb|AAL52820.1| DIHYDROPYRIMIDINE DEHYDROGENASE (NADP+) [Brucella melitensis 16M]
gi|17987922|ref|NP_540556.1| DIHYDROPYRIMIDINE
DEHYDROGENASE (NADP+) [Brucella melitensis 16M]
gi|25529497|pir||AI3456 dihydropyrimidine dehydrogenase
(NADP) (EC 1.3.1.2) [imported] - Brucella melitensis
(strain 16M)
Length = 436
Score = 278 bits (712), Expect = 2e-73
Identities = 145/334 (43%), Positives = 207/334 (61%), Gaps = 10/334 (2%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVT-PRY 106
DLS G+ PNPF + S PP ++RAF GWGGV+ KT+ + V+NV PRY
Sbjct: 3 DLSTNFLGIKSPNPFWLASAPPTDKAYNVERAFKAGWGGVVWKTLGAEGPPVVNVNGPRY 62
Query: 107 ARLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAA 166
+ A ++G NIELI+DRPLE L+E K++K +PDR LIASIM + A
Sbjct: 63 GAIHG-----ADRRLLGLNNIELITDRPLEINLREMKEVKMRWPDRALIASIMVPCEEEA 117
Query: 167 WEELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAK 226
W+ ++ VE+TG D IE+NF CPHGM ER MGAAVGQ + V W + +PV K
Sbjct: 118 WKAILPLVEETGADGIELNFGCPHGMSERGMGAAVGQVPEYVGMVVKWCKQYSRMPVITK 177
Query: 227 MTPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAV 286
+TPNITDI +PAR A ++G + ++ INTI S++ ++L+T+ P P ++G + GGY AV
Sbjct: 178 LTPNITDIRKPARAAFANGTDALSLINTINSIVSVDLDTMSPVPSIDGRGSHGGYCGPAV 237
Query: 287 HPIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGY 346
PIAL V IA+ + ++ +S IGG+ T DAAEFI LG TVQVCT M +G+
Sbjct: 238 KPIALNMVAEIAR----DAETAGLPISGIGGITTWQDAAEFIALGCGTVQVCTAAMTYGF 293
Query: 347 GLVKKLSLELKDFMKKHNFTSIEDFRGASLQYFT 380
+V+++ L D+M + +++DF+G ++ + T
Sbjct: 294 KVVQEMIDGLSDWMDSKGYQTLDDFQGCAVSHVT 327
>gb|AAV95056.1| dihydroorotate dehydrogenase family protein [Silicibacter pomeroyi
DSS-3] gi|56696653|ref|YP_167014.1| dihydroorotate
dehydrogenase family protein [Silicibacter pomeroyi
DSS-3]
Length = 434
Score = 277 bits (708), Expect = 5e-73
Identities = 151/358 (42%), Positives = 211/358 (58%), Gaps = 12/358 (3%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVT-PRY 106
DL G+ PNPF + S PP ++RAF+ GWGGV+ KT+ + V+NV PRY
Sbjct: 3 DLKTNFLGIESPNPFWLASAPPTDKEYNVRRAFEAGWGGVVWKTLGSEGPPVVNVNGPRY 62
Query: 107 ARLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAA 166
+ A ++G NIELI+DRPLET L+E ++K +YPDR +I SIM +A
Sbjct: 63 GAIYG-----ADRRLLGLNNIELITDRPLETNLEEIARVKADYPDRAVIVSIMVPCEEAE 117
Query: 167 WEELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAK 226
W+ ++ RV +TG D IE+NF CPHGM ER MGAAVGQ +E V W PV K
Sbjct: 118 WKAILPRVAETGADGIELNFGCPHGMSERGMGAAVGQVPEYIEMVTRWCKTYYDKPVIVK 177
Query: 227 MTPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAV 286
+TPNITDI PAR A + G + V+ INTI S+ +NL+ PEP ++G + GGY AV
Sbjct: 178 LTPNITDIRYPARAARNGGADAVSLINTISSITSVNLDNFSPEPSIDGKGSHGGYCGPAV 237
Query: 287 HPIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGY 346
PIA+ V I++ + ++ +S IGGV T DAAEF+ LG TVQVCT M +G+
Sbjct: 238 KPIAMNMVAEISR----DPATQGLPISGIGGVTTWRDAAEFMSLGCGTVQVCTAAMTYGF 293
Query: 347 GLVKKLSLELKDFMKKHNFTSIEDFRGASLQYFT--THTDLVHRQQEAIKQRKAIKKG 402
+V+++ L +M + +TS+ DF G ++ T + +L + + I Q + IK G
Sbjct: 294 RVVEEMKTGLSQWMDEKGYTSVNDFIGRAVPNVTDWQYLNLNYVAKAKIDQDQCIKCG 351
>ref|ZP_00456978.1| Dihydroorotate dehydrogenase 1 [Burkholderia cenocepacia AU 1054]
gi|67092837|gb|EAM10386.1| Dihydroorotate dehydrogenase
1 [Burkholderia cenocepacia AU 1054]
Length = 508
Score = 275 bits (702), Expect = 3e-72
Identities = 153/381 (40%), Positives = 224/381 (58%), Gaps = 16/381 (4%)
Query: 29 RPSRVDFKVFASEGQV-SEP---DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGW 84
RP R + + +EP DL + G+ PNPF + S PP + RAF+ GW
Sbjct: 51 RPDRATHRTHTEHDPILTEPNMADLRCTIAGITSPNPFWLASAPPTDKAYNVNRAFEAGW 110
Query: 85 GGVIAKTVSLDAAKVINVTPRYARLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQ 144
GGV+ KT+ LD V+NV+ RY ++ N I G NIELI+DRPL+ L+E Q
Sbjct: 111 GGVVWKTLGLDP-HVVNVSSRYGAVQWNGQ-----RIAGLNNIELITDRPLDVNLREIAQ 164
Query: 145 LKEEYPDRILIASIMEEYNKAAWEELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQD 204
+K ++PDR LI S+M N+ W+ ++ VE TG DA+E+NF CPHGM ER MGAAVGQ
Sbjct: 165 VKRDWPDRALIVSLMVPCNERDWKWILPLVEDTGADAVELNFGCPHGMSERGMGAAVGQV 224
Query: 205 CALLEEVCGWINAKATVPVWAKMTPNITDISQPARVALSSGCEGVAAINTIMSVMGINLN 264
+E V W+ +P K+TPNI+DI +R A G +GV+ INTI S++ ++L+
Sbjct: 225 PEYVEMVTRWVKEGTKLPCLVKLTPNISDIRMGSRAAYKGGADGVSLINTINSIVAVDLD 284
Query: 265 TLRPEPCVEGYSTPGGYSAKAVHPIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDA 324
+ P P V+G T GGY AV PIAL V IA+ + ++ N +S IGG+ + DA
Sbjct: 285 QMAPMPTVDGKGTHGGYCGPAVKPIALNMVAEIAR----DPETPNLPISGIGGISSWRDA 340
Query: 325 AEFILLGANTVQVCTGVMMHGYGLVKKLSLELKDFMKKHNFTSIEDFRGASLQYFT--TH 382
AEF++LGA +VQVCT M +G+ +V L+ L ++M + + +++D RG ++ T +
Sbjct: 341 AEFMVLGAGSVQVCTAAMHYGFRIVSDLADGLSNWMDEKGYATLDDIRGRAVPNVTDWKY 400
Query: 383 TDLVHRQQEAIKQRKAIKKGL 403
+L + + I Q + I+ GL
Sbjct: 401 LNLKYDIKARIDQDRCIQCGL 421
>ref|ZP_00283910.1| COG0167: Dihydroorotate dehydrogenase [Burkholderia fungorum LB400]
Length = 444
Score = 275 bits (702), Expect = 3e-72
Identities = 149/358 (41%), Positives = 216/358 (59%), Gaps = 12/358 (3%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYA 107
DL + G+ PNPF + S PP + RAF+ GWGGV+ KT+ LD V+NV+ RY
Sbjct: 3 DLRCTIAGITSPNPFWLASAPPTDKAYNVNRAFEAGWGGVVWKTLGLDP-HVVNVSSRYG 61
Query: 108 RLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAW 167
++ N I G NIELI+DRPL+ LKE Q+K ++PDR +I S+M N+ W
Sbjct: 62 AVQWNGQ-----RIAGLNNIELITDRPLDVNLKEIAQVKRDWPDRAMIVSLMVPCNERDW 116
Query: 168 EELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKM 227
+ ++ VE TG DA+E+NF CPHGM ER MGAAVGQ +E V W+ +P K+
Sbjct: 117 KWILPLVEDTGADAVELNFGCPHGMSERGMGAAVGQVPEYVEMVTRWVKEGTKLPCLVKL 176
Query: 228 TPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAVH 287
TPNI+DI +R A G +GV+ INTI S++ ++L+ + P P V+G T GGY AV
Sbjct: 177 TPNISDIRLGSRAAYKGGADGVSLINTINSIVAVDLDAMSPLPMVDGKGTHGGYCGPAVK 236
Query: 288 PIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYG 347
PIAL V IA+ + ++ N +S IGG+ + DAAEF++LGA +VQVCT M +G+
Sbjct: 237 PIALNMVAEIAR----DAETPNLPISGIGGISSWRDAAEFMVLGAGSVQVCTAAMHYGFR 292
Query: 348 LVKKLSLELKDFMKKHNFTSIEDFRGASLQYFT--THTDLVHRQQEAIKQRKAIKKGL 403
+V L+ L ++M + + +++D RG ++ T + +L + + I Q K I+ GL
Sbjct: 293 IVSDLADGLSNWMDEKGYATLDDIRGRAVPNVTDWKYLNLKYDIKARIDQDKCIQCGL 350
>ref|ZP_00465097.1| Dihydroorotate dehydrogenase 1 [Burkholderia cenocepacia HI2424]
gi|67098550|gb|EAM15754.1| Dihydroorotate dehydrogenase
1 [Burkholderia cenocepacia HI2424]
Length = 437
Score = 273 bits (698), Expect = 7e-72
Identities = 148/358 (41%), Positives = 216/358 (59%), Gaps = 12/358 (3%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYA 107
DL + G+ PNPF + S PP + RAF+ GWGGV+ KT+ LD V+NV+ RY
Sbjct: 3 DLRCTIAGITSPNPFWLASAPPTDKAYNVNRAFEAGWGGVVWKTLGLDP-HVVNVSSRYG 61
Query: 108 RLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAW 167
++ N I G NIELI+DRPL+ L+E Q+K ++PDR LI S+M N+ W
Sbjct: 62 AVQWNGQ-----RIAGLNNIELITDRPLDVNLREIAQVKRDWPDRALIVSLMVPCNERDW 116
Query: 168 EELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKM 227
+ ++ VE TG DA+E+NF CPHGM ER MGAAVGQ +E V W+ +P K+
Sbjct: 117 KWILPLVEDTGADAVELNFGCPHGMSERGMGAAVGQVPEYVEMVTRWVKEGTKLPCLVKL 176
Query: 228 TPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAVH 287
TPNI+DI +R A G +GV+ INTI S++ ++L+ + P P V+G T GGY AV
Sbjct: 177 TPNISDIRMGSRAAYKGGADGVSLINTINSIVAVDLDQMAPMPTVDGKGTHGGYCGPAVK 236
Query: 288 PIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYG 347
PIAL V IA+ + ++ N +S IGG+ + DAAEF++LGA +VQVCT M +G+
Sbjct: 237 PIALNMVAEIAR----DPETPNLPISGIGGISSWRDAAEFMVLGAGSVQVCTAAMHYGFR 292
Query: 348 LVKKLSLELKDFMKKHNFTSIEDFRGASLQYFT--THTDLVHRQQEAIKQRKAIKKGL 403
+V L+ L ++M + + +++D RG ++ T + +L + + I Q + I+ GL
Sbjct: 293 IVSDLADGLSNWMDEKGYATLDDIRGRAVPNVTDWKYLNLKYDIKARIDQDRCIQCGL 350
>emb|CAC47027.1| PUTATIVE OXIDOREDUCTASE IRON-SULFUR PROTEIN [Sinorhizobium
meliloti] gi|15966201|ref|NP_386554.1| PUTATIVE
OXIDOREDUCTASE IRON-SULFUR PROTEIN [Sinorhizobium
meliloti 1021]
Length = 437
Score = 273 bits (698), Expect = 7e-72
Identities = 156/360 (43%), Positives = 210/360 (58%), Gaps = 16/360 (4%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVT-PRY 106
DL G+ PNPF + S PP ++RAF GWGGV+ KT+ + V+NV PRY
Sbjct: 3 DLRNNFVGIKSPNPFWLASAPPTDKAYNVERAFKAGWGGVVWKTLGEEGPPVVNVNGPRY 62
Query: 107 ARLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAA 166
+ A ++G+ NIELI+DR L L+E KQ+K +PDR LIASIM + A
Sbjct: 63 GAI-----WGADRRLLGFNNIELITDRDLYVNLREMKQVKMNWPDRALIASIMVPCEENA 117
Query: 167 WEELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAK 226
W+ ++ VE+TG D IE+NF CPHGM ER MG+AVGQ +E V W +PV K
Sbjct: 118 WKAILPLVEETGADGIELNFGCPHGMSERGMGSAVGQVPEYIEMVVRWCKQYTRMPVITK 177
Query: 227 MTPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAV 286
+TPNITDI +PAR A + G + V+ INTI S+ G+NL+T PEP ++G + GGY AV
Sbjct: 178 LTPNITDIRKPARAAKAGGTDAVSLINTINSITGVNLDTFSPEPSIDGRGSHGGYCGPAV 237
Query: 287 HPIALGKVMSIAKMMKSEFDSENYSL--SAIGGVETGGDAAEFILLGANTVQVCTGVMMH 344
PIAL V IA+ D E Y L S IGGV T DAAEF+ LGA VQVCT M +
Sbjct: 238 KPIALNMVAEIAR------DPETYGLPISGIGGVTTWRDAAEFMALGAGNVQVCTAAMTY 291
Query: 345 GYGLVKKLSLELKDFMKKHNFTSIEDFRGASLQYFT--THTDLVHRQQEAIKQRKAIKKG 402
G+ +V+++ L D+M +++D G ++ T + +L + + I Q IK G
Sbjct: 292 GFKIVQEMITGLSDWMDAKGHRTLDDISGRAVPNVTDWQYLNLNYIAKAKIDQDACIKCG 351
>ref|ZP_00214166.1| COG0167: Dihydroorotate dehydrogenase [Burkholderia cepacia R18194]
Length = 438
Score = 273 bits (697), Expect = 1e-71
Identities = 148/358 (41%), Positives = 216/358 (59%), Gaps = 12/358 (3%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYA 107
DL + G+ PNPF + S PP + RAF+ GWGGV+ KT+ LD V+NV+ RY
Sbjct: 3 DLRCTIAGITSPNPFWLASAPPTDKAYNVNRAFEAGWGGVVWKTLGLDP-HVVNVSSRYG 61
Query: 108 RLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAW 167
++ N I G NIELI+DRPL+ L+E Q+K ++PDR LI S+M N+ W
Sbjct: 62 AVQWNGQ-----RIAGLNNIELITDRPLDVNLREIAQVKRDWPDRALIVSLMVPCNERDW 116
Query: 168 EELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKM 227
+ ++ VE TG DA+E+NF CPHGM ER MGAAVGQ +E V W+ +P K+
Sbjct: 117 KWILPLVEDTGADAVELNFGCPHGMSERGMGAAVGQVPEYVEMVTRWVKEGTKLPCLVKL 176
Query: 228 TPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAVH 287
TPNI+DI +R A G +GV+ INTI S++ ++L+ + P P V+G T GGY AV
Sbjct: 177 TPNISDIRMGSRAAYKGGADGVSLINTINSIVAVDLDHMAPMPMVDGKGTHGGYCGPAVK 236
Query: 288 PIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYG 347
PIAL V IA+ + ++ N +S IGG+ + DAAEF++LGA +VQVCT M +G+
Sbjct: 237 PIALNMVAEIAR----DPETPNLPISGIGGISSWRDAAEFMVLGAGSVQVCTAAMHYGFR 292
Query: 348 LVKKLSLELKDFMKKHNFTSIEDFRGASLQYFT--THTDLVHRQQEAIKQRKAIKKGL 403
+V L+ L ++M + + +++D RG ++ T + +L + + I Q + I+ GL
Sbjct: 293 IVSDLADGLSNWMDEKGYATLDDIRGRAVPNVTDWKYLNLKYDIKARIDQDRCIQCGL 350
>ref|ZP_00420597.1| Dihydroorotate dehydrogenase 1 [Burkholderia vietnamiensis G4]
gi|67535875|gb|EAM32599.1| Dihydroorotate dehydrogenase
1 [Burkholderia vietnamiensis G4]
Length = 435
Score = 270 bits (690), Expect = 6e-71
Identities = 149/358 (41%), Positives = 212/358 (58%), Gaps = 12/358 (3%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYA 107
DL + G+ PNPF + S PP + RAF+ GWGGV+ KT+ LD V+NV+ RY
Sbjct: 3 DLRCTIAGITSPNPFWLASAPPTDKAYNVNRAFEAGWGGVVWKTLGLDP-HVVNVSSRYG 61
Query: 108 RLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAW 167
++ N I G NIELI+DRPL+ L E Q+K ++PDR LI S+M N+ W
Sbjct: 62 AVQWNGQ-----RIAGLNNIELITDRPLDVNLAEIAQVKRDWPDRALIVSLMVPCNERDW 116
Query: 168 EELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKM 227
+ ++ VE TG DA+E+NF CPHGM ER MGAAVGQ +E V W+ +P K+
Sbjct: 117 KWILPLVEDTGADAVELNFGCPHGMSERGMGAAVGQVPEYVEMVTRWVKEGTKLPCLVKL 176
Query: 228 TPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAVH 287
TPNI+DI +R A G +GV+ INTI S++ ++L+ + P P V+G T GGY AV
Sbjct: 177 TPNISDIRMGSRAAYKGGADGVSLINTINSIVAVDLDQMAPMPTVDGKGTHGGYCGPAVK 236
Query: 288 PIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYG 347
PIAL V IA+ + + N +S IGG+ + DAAEFI+LGA +VQVCT M +G+
Sbjct: 237 PIALNMVAEIAR----DPQTPNLPISGIGGISSWRDAAEFIVLGAGSVQVCTAAMHYGFR 292
Query: 348 LVKKLSLELKDFMKKHNFTSIEDFRGASLQYFT--THTDLVHRQQEAIKQRKAIKKGL 403
+V L L ++M + +++D RG ++ T + +L + + I Q + I+ GL
Sbjct: 293 IVSDLVDGLSNWMDDKGYATLDDVRGRAVPNVTDWKYLNLKYDIKARIDQDRCIQCGL 350
>ref|ZP_00006802.1| COG0167: Dihydroorotate dehydrogenase [Rhodobacter sphaeroides
2.4.1]
Length = 434
Score = 267 bits (683), Expect = 4e-70
Identities = 143/351 (40%), Positives = 205/351 (57%), Gaps = 12/351 (3%)
Query: 55 GLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVT-PRYARLRANA 113
G+ PNPF + S PP ++RAF+ GWGGV+ KT+ + V+NV PRY +
Sbjct: 10 GIKSPNPFWLASAPPTDKEYNVRRAFEAGWGGVVWKTLGSEGPPVVNVNGPRYGAIYG-- 67
Query: 114 NGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAWEELIDR 173
A ++G NIELI+DRPLE L+E K +K +YPDR L+ S+M ++ +W+ ++
Sbjct: 68 ---ADRRLLGLNNIELITDRPLEVNLREIKSVKRDYPDRALVVSLMVPCDEESWKAILAH 124
Query: 174 VEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKMTPNITD 233
VE TG D +E+NF CPHGM ER MG+AVGQ +E V W+ + +P K+TPN+TD
Sbjct: 125 VEDTGADGVELNFGCPHGMAERGMGSAVGQVPEYIEMVTRWVKQHSQMPCIVKLTPNVTD 184
Query: 234 ISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAVHPIALGK 293
I +PA A G + V+ INTI S+ G+++++ P P ++G T GGY AV PIAL
Sbjct: 185 IRKPAEAARRGGADAVSLINTINSITGVDIDSFAPMPTIDGKGTHGGYCGPAVKPIALNM 244
Query: 294 VMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYGLVKKLS 353
V IA+ ++ +S IGGV T DA EF+LLGA VQVCT M +G+ +V+++
Sbjct: 245 VAEIAR----NPETHGLPISGIGGVTTWRDAVEFMLLGAGNVQVCTAAMTYGFRVVQEMI 300
Query: 354 LELKDFMKKHNFTSIEDFRGASLQYFT--THTDLVHRQQEAIKQRKAIKKG 402
L D+M F S D G ++ T + +L + + I Q IK G
Sbjct: 301 SGLSDYMDAKGFASTADLVGRAVPNVTDWQYLNLNYVTKAEIDQDLCIKCG 351
>ref|ZP_00526667.1| Dihydroorotate dehydrogenase 1 [Solibacter usitatus Ellin6076]
gi|67859205|gb|EAM54286.1| Dihydroorotate dehydrogenase
1 [Solibacter usitatus Ellin6076]
Length = 446
Score = 267 bits (682), Expect = 5e-70
Identities = 156/370 (42%), Positives = 209/370 (56%), Gaps = 23/370 (6%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYA 107
DLS G+ PNPF + SGPP + RAFD GWGG + KT+ ++NV+ RY+
Sbjct: 3 DLSTNFAGIRCPNPFWLASGPPTNCGEQIMRAFDAGWGGAVWKTIG---EPIVNVSSRYS 59
Query: 108 RLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAW 167
+ N ++G NIELISDRP+ET L+E ++K+ YP +IAS+M E + AW
Sbjct: 60 SVDWNGQ-----RMMGLNNIELISDRPIETNLREITEVKKRYPKHAVIASLMVESTREAW 114
Query: 168 EELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQ----DCALLEEVCGWINAKATVPV 223
+++ R E G D +E+NF CPHGM ER MGAAVGQ C + E W+ A PV
Sbjct: 115 HDIVKRSEDAGADGLELNFGCPHGMSERGMGAAVGQVPDYSCMITE----WVKEVARTPV 170
Query: 224 WAKMTPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSA 283
K+TPNITDI AR A + ++AINTI S+ GI+L+T P P V G ++ GGY
Sbjct: 171 LVKLTPNITDIRAVARAAKRGRADALSAINTINSITGIDLDTFTPRPNVGGKTSHGGYCG 230
Query: 284 KAVHPIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMM 343
AV PIAL V IA + +S IGG+ +AAEFILLG TVQVCT M
Sbjct: 231 PAVKPIALHLVQQIA-----SDPAVGLPISGIGGIGGWREAAEFILLGCGTVQVCTAAMH 285
Query: 344 HGYGLVKKLSLELKDFMKKHNFTSIEDFRGASLQYFT--THTDLVHRQQEAIKQRKAIKK 401
+G+ +V+ + L ++M + F +IEDFRG S+ T H DL ++ I + K I
Sbjct: 286 YGFRIVEDMIDGLSNWMDEKGFRTIEDFRGLSVPRVTDWKHLDLNYKIVARINRDKCIGC 345
Query: 402 GLQSDKDWTG 411
L W G
Sbjct: 346 DLCYVACWDG 355
>gb|EAM66354.1| Dihydroorotate dehydrogenase 1 [Jannaschia sp. CCS1]
Length = 434
Score = 266 bits (681), Expect = 7e-70
Identities = 150/358 (41%), Positives = 210/358 (57%), Gaps = 13/358 (3%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVT-PRY 106
DL+ G+ PNPF + S PP ++RAF+ GWGGV+ KT+ + V+NV PRY
Sbjct: 3 DLTSNFIGITSPNPFWLASAPPTDKEYNVRRAFEAGWGGVVWKTLGSEGPPVVNVNGPRY 62
Query: 107 ARLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAA 166
+ A ++G NIELI+DR L T L+E K++K +YPDR +I SIM +AA
Sbjct: 63 GAIYG-----ADRRLLGLNNIELITDRDLYTNLEEIKRVKADYPDRAIIVSIMVPCEEAA 117
Query: 167 WEELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAK 226
W+ ++ VE TG D IE+NF CPHGM ER MGAAVGQ +E V W + PV K
Sbjct: 118 WKAILPLVEDTGADGIELNFGCPHGMSERGMGAAVGQVPEYIEMVTRWCKQYYSKPVIVK 177
Query: 227 MTPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAV 286
+TPNITDI++PA A G + V+ INTI S+ +NL+T+ PEP ++G + GGY AV
Sbjct: 178 LTPNITDITKPAAAAKRGGADAVSLINTINSITSVNLDTMSPEPSIDGKGSHGGYCGPAV 237
Query: 287 HPIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGY 346
PIA+ V IA+ ++ +S IGGV T DAAEF+ LGA VQVCT M +G+
Sbjct: 238 KPIAMSMVSEIARSP----ETHGLPISGIGGVTTWRDAAEFMTLGAGNVQVCTAAMTYGF 293
Query: 347 GLVKKLSLELKDFMKKHNFTSIEDFRGASLQYFT--THTDLVHRQQEAIKQRKAIKKG 402
+V+++ L F+ + T ++D G ++ T + +L + + I Q IK G
Sbjct: 294 KIVEEMISGLSQFLDEKGMT-LDDLVGRAVPNVTDWQYLNLNYVAKAQINQDDCIKCG 350
>ref|NP_103183.1| probable oxidoreductase [Mesorhizobium loti MAFF303099]
gi|14022359|dbj|BAB48969.1| probable oxidoreductase
[Mesorhizobium loti MAFF303099]
Length = 437
Score = 266 bits (680), Expect = 9e-70
Identities = 149/358 (41%), Positives = 212/358 (58%), Gaps = 12/358 (3%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVT-PRY 106
DL G+ PNPF + S PP + RAF GWGGV+ KT+ + V+NV PRY
Sbjct: 3 DLRNNFVGIKSPNPFWLASAPPTDKAYNVIRAFKAGWGGVVWKTLGEEGPPVVNVNGPRY 62
Query: 107 ARLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAA 166
+ A ++G NIELI+DR L+T L+E KQ+K ++PDR LIASIM +A+
Sbjct: 63 GAI-----WGADRRLLGLNNIELITDRDLQTNLREMKQVKMDWPDRALIASIMVPCEEAS 117
Query: 167 WEELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAK 226
W+ ++ VE+TG D IE+NF CPHGM ER MGAAVGQ +E V W +PV K
Sbjct: 118 WKAILPLVEETGADGIELNFGCPHGMSERGMGAAVGQVPEYIEMVVRWCKQYTRMPVITK 177
Query: 227 MTPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAV 286
+TPNITDI +PAR A + G + V+ INTI S+ G++L++ P P ++G + GGY AV
Sbjct: 178 LTPNITDIRKPARAAHAGGTDAVSLINTINSITGVDLDSFAPMPTIDGKGSHGGYCGPAV 237
Query: 287 HPIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGY 346
PIA+ V IA+ + ++ +S IGG+ T DAAEF+ LGA VQVCT M +G+
Sbjct: 238 KPIAMNMVAEIAR----DPETRGLPISGIGGITTWRDAAEFLALGAGNVQVCTAAMTYGF 293
Query: 347 GLVKKLSLELKDFMKKHNFTSIEDFRGASLQYFT--THTDLVHRQQEAIKQRKAIKKG 402
+V+++ L+++M + S++D G + T + +L + + I Q IK G
Sbjct: 294 KIVQEMIAGLENWMDEKGHRSLDDIIGRATPNVTDWQYLNLNYVAKAHIDQDACIKCG 351
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.316 0.133 0.386
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 703,638,160
Number of Sequences: 2540612
Number of extensions: 28885219
Number of successful extensions: 80022
Number of sequences better than 10.0: 486
Number of HSP's better than 10.0 without gapping: 317
Number of HSP's successfully gapped in prelim test: 169
Number of HSP's that attempted gapping in prelim test: 79144
Number of HSP's gapped (non-prelim): 553
length of query: 424
length of database: 863,360,394
effective HSP length: 131
effective length of query: 293
effective length of database: 530,540,222
effective search space: 155448285046
effective search space used: 155448285046
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 76 (33.9 bits)
Medicago: description of AC139706.5


