

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139600.10 + phase: 0
(81 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
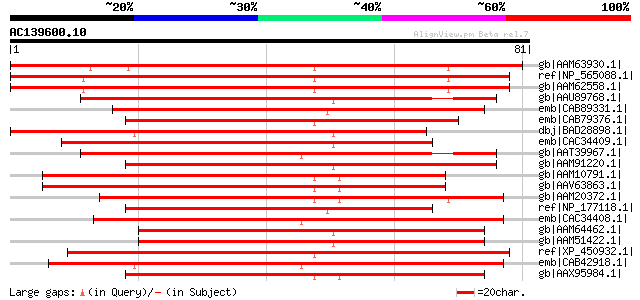
Score E
Sequences producing significant alignments: (bits) Value
gb|AAM63930.1| unknown [Arabidopsis thaliana] gi|62319539|dbj|BA... 109 2e-23
ref|NP_565088.1| zinc finger homeobox family protein / ZF-HD hom... 94 7e-19
gb|AAM62558.1| unknown [Arabidopsis thaliana] 94 7e-19
gb|AAU89768.1| ZF-HD homeobox protein-like [Solanum tuberosum] 86 2e-16
emb|CAB89331.1| putative protein [Arabidopsis thaliana] gi|15242... 82 5e-15
emb|CAB79376.1| putative protein [Arabidopsis thaliana] gi|42205... 82 5e-15
dbj|BAD28898.1| unknown protein [Oryza sativa (japonica cultivar... 81 8e-15
emb|CAC34409.1| ZF-HD homeobox protein [Flaveria bidentis] 80 1e-14
gb|AAT39967.1| putative ZF-HD homeobox protein [Solanum demissum] 79 2e-14
gb|AAM91220.1| unknown protein [Arabidopsis thaliana] gi|9294358... 79 4e-14
gb|AAM10791.1| hypothetical protein At2g02540/T822.16 [Arabidops... 78 5e-14
gb|AAV63863.1| hypothetical protein At2g02540 [Arabidopsis thali... 78 5e-14
gb|AAM20372.1| unknown protein [Arabidopsis thaliana] gi|1837762... 78 5e-14
ref|NP_177118.1| zinc finger homeobox family protein / ZF-HD hom... 77 9e-14
emb|CAC34408.1| ZF-HD homeobox protein [Flaveria bidentis] 77 1e-13
gb|AAM64462.1| unknown [Arabidopsis thaliana] 76 2e-13
gb|AAM51422.1| unknown protein [Arabidopsis thaliana] gi|2025947... 76 2e-13
ref|XP_450932.1| putative ZF-HD homeobox protein [Oryza sativa (... 76 2e-13
emb|CAB42918.1| putative protein [Arabidopsis thaliana] gi|15230... 75 3e-13
gb|AAX95984.1| ZF-HD protein dimerisation region, putative [Oryz... 75 4e-13
>gb|AAM63930.1| unknown [Arabidopsis thaliana] gi|62319539|dbj|BAD94968.1|
hypothetical protein [Arabidopsis thaliana]
gi|9294233|dbj|BAB02135.1| unnamed protein product
[Arabidopsis thaliana] gi|42572555|ref|NP_974373.1|
zinc finger homeobox family protein / ZF-HD homeobox
family protein [Arabidopsis thaliana]
Length = 100
Score = 109 bits (273), Expect = 2e-23
Identities = 60/99 (60%), Positives = 70/99 (70%), Gaps = 19/99 (19%)
Query: 1 MRKRQVVVRRS--EDPQRS----------VVKYGECQKNHAANVGGYAVDGCREFMPS-- 46
MRKRQVV+RR+ E+P RS V+YGECQKNHAA VGGYAVDGCREFM S
Sbjct: 1 MRKRQVVLRRASPEEPSRSSSTASSLTVRTVRYGECQKNHAAAVGGYAVDGCREFMASRG 60
Query: 47 ---TNGSLTCAACGCHRNFHKREV--EVVSESSSPHSNG 80
T +LTCAACGCHR+FH+RE+ EVV + +SP S G
Sbjct: 61 EEGTVAALTCAACGCHRSFHRREIETEVVCDCNSPPSTG 99
>ref|NP_565088.1| zinc finger homeobox family protein / ZF-HD homeobox family protein
[Arabidopsis thaliana] gi|12324813|gb|AAG52375.1|
hypothetical protein; 104370-104062 [Arabidopsis
thaliana] gi|25406359|pir||G96775 hypothetical protein
F1M20.34 [imported] - Arabidopsis thaliana
Length = 102
Score = 94.4 bits (233), Expect = 7e-19
Identities = 51/101 (50%), Positives = 65/101 (63%), Gaps = 23/101 (22%)
Query: 1 MRKRQVVVRR-----------------SEDPQRSVVKYGECQKNHAANVGGYAVDGCREF 43
M+KRQ+V+++ S + S V+Y ECQKNHAAN+GGYAVDGCREF
Sbjct: 2 MKKRQMVIKQRSRNSNTSSSWTTTSSSSSSSEISNVRYVECQKNHAANIGGYAVDGCREF 61
Query: 44 MPS----TNGSLTCAACGCHRNFHKREV--EVVSESSSPHS 78
M + T +L CAACGCHRNFH++EV EVV E S P++
Sbjct: 62 MAAGVEGTVDALRCAACGCHRNFHRKEVDTEVVCEYSPPNA 102
>gb|AAM62558.1| unknown [Arabidopsis thaliana]
Length = 101
Score = 94.4 bits (233), Expect = 7e-19
Identities = 51/101 (50%), Positives = 65/101 (63%), Gaps = 23/101 (22%)
Query: 1 MRKRQVVVRR-----------------SEDPQRSVVKYGECQKNHAANVGGYAVDGCREF 43
M+KRQ+V+++ S + S V+Y ECQKNHAAN+GGYAVDGCREF
Sbjct: 1 MKKRQMVIKQRSRNSNTSSSWTTTSSSSSSSEISNVRYVECQKNHAANIGGYAVDGCREF 60
Query: 44 MPS----TNGSLTCAACGCHRNFHKREV--EVVSESSSPHS 78
M + T +L CAACGCHRNFH++EV EVV E S P++
Sbjct: 61 MAAGVEGTVDALRCAACGCHRNFHRKEVDTEVVCEYSPPNA 101
>gb|AAU89768.1| ZF-HD homeobox protein-like [Solanum tuberosum]
Length = 290
Score = 85.9 bits (211), Expect = 2e-16
Identities = 43/71 (60%), Positives = 51/71 (71%), Gaps = 9/71 (12%)
Query: 12 EDPQRSVVKYGECQKNHAANVGGYAVDGCREFMPSTNG------SLTCAACGCHRNFHKR 65
+ P +VV Y EC KNHAA++GG+AVDGC EFMPST SL CAACGCHRNFH+R
Sbjct: 40 QPPPCTVVIYKECLKNHAASLGGHAVDGCGEFMPSTESTPSDPISLKCAACGCHRNFHRR 99
Query: 66 EVEVVSESSSP 76
E S++SSP
Sbjct: 100 E---PSDNSSP 107
>emb|CAB89331.1| putative protein [Arabidopsis thaliana]
gi|15242243|ref|NP_197025.1| zinc finger homeobox family
protein / ZF-HD homeobox family protein [Arabidopsis
thaliana] gi|45773756|gb|AAS76682.1| At5g15210
[Arabidopsis thaliana] gi|11288375|pir||T49956
hypothetical protein F8M21.100 - Arabidopsis thaliana
Length = 271
Score = 81.6 bits (200), Expect = 5e-15
Identities = 38/64 (59%), Positives = 46/64 (71%), Gaps = 6/64 (9%)
Query: 17 SVVKYGECQKNHAANVGGYAVDGCREFMPSTN------GSLTCAACGCHRNFHKREVEVV 70
+V Y EC KNHAA +GG+A+DGC EFMPS + SLTCAACGCHRNFH+RE +
Sbjct: 52 AVATYKECLKNHAAGIGGHALDGCGEFMPSPSFNSNDPASLTCAACGCHRNFHRREEDPS 111
Query: 71 SESS 74
S S+
Sbjct: 112 SLSA 115
>emb|CAB79376.1| putative protein [Arabidopsis thaliana] gi|4220524|emb|CAA22997.1|
putative protein [Arabidopsis thaliana]
gi|21928089|gb|AAM78073.1| AT4g24660/F22K18_140
[Arabidopsis thaliana] gi|15233925|ref|NP_194197.1| zinc
finger homeobox family protein / ZF-HD homeobox family
protein [Arabidopsis thaliana]
gi|16612295|gb|AAL27510.1| AT4g24660/F22K18_140
[Arabidopsis thaliana] gi|7485966|pir||T05568
hypothetical protein F22K18.140 - Arabidopsis thaliana
Length = 220
Score = 81.6 bits (200), Expect = 5e-15
Identities = 35/56 (62%), Positives = 43/56 (76%), Gaps = 4/56 (7%)
Query: 19 VKYGECQKNHAANVGGYAVDGCREFMPS----TNGSLTCAACGCHRNFHKREVEVV 70
++Y EC KNHA N+GG+AVDGC EFMPS T +L CAACGCHRNFH++E E +
Sbjct: 47 IRYRECLKNHAVNIGGHAVDGCCEFMPSGEDGTLDALKCAACGCHRNFHRKETESI 102
>dbj|BAD28898.1| unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 113
Score = 80.9 bits (198), Expect = 8e-15
Identities = 41/82 (50%), Positives = 53/82 (64%), Gaps = 17/82 (20%)
Query: 1 MRKRQVVVRRSEDPQRSV---VKYGECQKNHAANVGGYAVDGCREFMPSTNG-------- 49
M KR VV+RR E R V+YGEC++NHAA+ GG+AVDGCREF+ + +G
Sbjct: 1 MMKRLVVLRRREPAVRFSCCGVRYGECRRNHAASTGGHAVDGCREFIAAEDGGGGNSTSA 60
Query: 50 ------SLTCAACGCHRNFHKR 65
+L CAACGCHR+FH+R
Sbjct: 61 VGVAAAALKCAACGCHRSFHRR 82
>emb|CAC34409.1| ZF-HD homeobox protein [Flaveria bidentis]
Length = 339
Score = 80.1 bits (196), Expect = 1e-14
Identities = 37/64 (57%), Positives = 45/64 (69%), Gaps = 6/64 (9%)
Query: 9 RRSEDPQRSVVKYGECQKNHAANVGGYAVDGCREFMPSTNG------SLTCAACGCHRNF 62
++ + Q VV Y EC KNHAA +GG+AVDGC EFMPS + SL CAACGCHRNF
Sbjct: 57 QQQQQQQPVVVSYRECLKNHAAAMGGHAVDGCGEFMPSPSSSPTDPTSLKCAACGCHRNF 116
Query: 63 HKRE 66
H+R+
Sbjct: 117 HRRD 120
>gb|AAT39967.1| putative ZF-HD homeobox protein [Solanum demissum]
Length = 291
Score = 79.3 bits (194), Expect = 2e-14
Identities = 41/71 (57%), Positives = 48/71 (66%), Gaps = 9/71 (12%)
Query: 12 EDPQRSVVKYGECQKNHAANVGGYAVDGCREFM------PSTNGSLTCAACGCHRNFHKR 65
+ P + V Y EC KNHAA++GG+AVDGC EFM PS SL CAACGCHRNFH+R
Sbjct: 44 QPPPCTAVIYKECLKNHAASLGGHAVDGCGEFMLSPESTPSDPISLKCAACGCHRNFHRR 103
Query: 66 EVEVVSESSSP 76
E S+ SSP
Sbjct: 104 E---PSDDSSP 111
>gb|AAM91220.1| unknown protein [Arabidopsis thaliana] gi|9294358|dbj|BAB02255.1|
unnamed protein product [Arabidopsis thaliana]
gi|20260544|gb|AAM13170.1| unknown protein [Arabidopsis
thaliana] gi|15228530|ref|NP_189534.1| zinc finger
homeobox family protein / ZF-HD homeobox family protein
[Arabidopsis thaliana]
Length = 312
Score = 78.6 bits (192), Expect = 4e-14
Identities = 37/64 (57%), Positives = 43/64 (66%), Gaps = 6/64 (9%)
Query: 19 VKYGECQKNHAANVGGYAVDGCREFMPSTNG------SLTCAACGCHRNFHKREVEVVSE 72
V Y EC KNHAA +GG+A+DGC EFMPS + SL CAACGCHRNFH+RE + S
Sbjct: 50 VTYKECLKNHAAAIGGHALDGCGEFMPSPSSTPSDPTSLKCAACGCHRNFHRRETDDSSA 109
Query: 73 SSSP 76
P
Sbjct: 110 VPPP 113
>gb|AAM10791.1| hypothetical protein At2g02540/T822.16 [Arabidopsis thaliana]
Length = 310
Score = 78.2 bits (191), Expect = 5e-14
Identities = 37/67 (55%), Positives = 46/67 (68%), Gaps = 4/67 (5%)
Query: 6 VVVRRSEDPQRSVVKYGECQKNHAANVGGYAVDGCREFMPS-TNGS---LTCAACGCHRN 61
++V + + V+KY EC KNHAA +GG A+DGC EFMPS GS LTC+ C CHRN
Sbjct: 72 IMVTNIKKEKPVVIKYKECLKNHAATMGGNAIDGCGEFMPSGEEGSIEALTCSVCNCHRN 131
Query: 62 FHKREVE 68
FH+RE E
Sbjct: 132 FHRRETE 138
>gb|AAV63863.1| hypothetical protein At2g02540 [Arabidopsis thaliana]
gi|3184285|gb|AAC18932.1| hypothetical protein
[Arabidopsis thaliana] gi|7487811|pir||T00609
hypothetical protein At2g02540 [imported] - Arabidopsis
thaliana gi|15226993|ref|NP_178358.1| zinc finger
homeobox family protein / ZF-HD homeobox family protein
[Arabidopsis thaliana]
Length = 310
Score = 78.2 bits (191), Expect = 5e-14
Identities = 37/67 (55%), Positives = 46/67 (68%), Gaps = 4/67 (5%)
Query: 6 VVVRRSEDPQRSVVKYGECQKNHAANVGGYAVDGCREFMPS-TNGS---LTCAACGCHRN 61
++V + + V+KY EC KNHAA +GG A+DGC EFMPS GS LTC+ C CHRN
Sbjct: 72 IMVTNIKKEKPVVIKYKECLKNHAATMGGNAIDGCGEFMPSGEEGSIEALTCSVCNCHRN 131
Query: 62 FHKREVE 68
FH+RE E
Sbjct: 132 FHRRETE 138
>gb|AAM20372.1| unknown protein [Arabidopsis thaliana] gi|18377626|gb|AAL66963.1|
unknown protein [Arabidopsis thaliana]
gi|42571471|ref|NP_973826.1| zinc finger homeobox family
protein / ZF-HD homeobox family protein [Arabidopsis
thaliana] gi|15223757|ref|NP_172896.1| zinc finger
homeobox family protein / ZF-HD homeobox family protein
[Arabidopsis thaliana] gi|7262686|gb|AAF43944.1|
Contains similarity to a hypothetical protein from
Arabidopsis thaliana gb|AC004136.2
gi|25518026|pir||A86279 F14L17.21 protein - Arabidopsis
thaliana
Length = 312
Score = 78.2 bits (191), Expect = 5e-14
Identities = 38/69 (55%), Positives = 51/69 (73%), Gaps = 6/69 (8%)
Query: 15 QRSVVKYGECQKNHAANVGGYAVDGCREFMPS-TNGS---LTCAACGCHRNFHKREV--E 68
++ ++KY EC KNHAA +GG A DGC EFMPS +GS LTC+AC CHRNFH++EV E
Sbjct: 84 KKPMIKYKECLKNHAAAMGGNATDGCGEFMPSGEDGSIEALTCSACNCHRNFHRKEVEGE 143
Query: 69 VVSESSSPH 77
+ + + SP+
Sbjct: 144 LAATAMSPY 152
>ref|NP_177118.1| zinc finger homeobox family protein / ZF-HD homeobox family
protein [Arabidopsis thaliana] gi|25363267|pir||F96717
hypothetical protein F24J1.29 [imported] - Arabidopsis
thaliana gi|6692255|gb|AAF24606.1| hypothetical
protein; 18366-17638 [Arabidopsis thaliana]
Length = 242
Score = 77.4 bits (189), Expect = 9e-14
Identities = 35/54 (64%), Positives = 40/54 (73%), Gaps = 6/54 (11%)
Query: 19 VKYGECQKNHAANVGGYAVDGCREFMPSTN------GSLTCAACGCHRNFHKRE 66
V Y EC KNHAAN+GG+A+DGC EFMPS SL CAACGCHRNFH+R+
Sbjct: 29 VCYKECLKNHAANLGGHALDGCGEFMPSPTATSTDPSSLRCAACGCHRNFHRRD 82
>emb|CAC34408.1| ZF-HD homeobox protein [Flaveria bidentis]
Length = 241
Score = 76.6 bits (187), Expect = 1e-13
Identities = 35/70 (50%), Positives = 43/70 (61%), Gaps = 6/70 (8%)
Query: 14 PQRSVVKYGECQKNHAANVGGYAVDGCREFM------PSTNGSLTCAACGCHRNFHKREV 67
P + V Y +C KNHA +GG+AVDGC EFM PS S CAACGCHRNFH+RE
Sbjct: 10 PPHTGVAYKQCLKNHAVGIGGHAVDGCGEFMPAPTYNPSQPSSFKCAACGCHRNFHRREP 69
Query: 68 EVVSESSSPH 77
+ ++ H
Sbjct: 70 TTTTIATRTH 79
>gb|AAM64462.1| unknown [Arabidopsis thaliana]
Length = 333
Score = 76.3 bits (186), Expect = 2e-13
Identities = 35/60 (58%), Positives = 44/60 (73%), Gaps = 6/60 (10%)
Query: 21 YGECQKNHAANVGGYAVDGCREFMPSTNG------SLTCAACGCHRNFHKREVEVVSESS 74
Y EC KNHAA +GG+A+DGC EFMPS + SL CAACGCHRNFH+R+ + ++SS
Sbjct: 55 YKECLKNHAAALGGHALDGCGEFMPSPSSISSDPTSLKCAACGCHRNFHRRDPDNNNDSS 114
>gb|AAM51422.1| unknown protein [Arabidopsis thaliana] gi|20259470|gb|AAM13855.1|
unknown protein [Arabidopsis thaliana]
gi|10177976|dbj|BAB11382.1| unnamed protein product
[Arabidopsis thaliana] gi|18421904|ref|NP_568570.1| zinc
finger homeobox protein-related / ZF-HD homeobox
protein-related [Arabidopsis thaliana]
Length = 334
Score = 76.3 bits (186), Expect = 2e-13
Identities = 35/60 (58%), Positives = 44/60 (73%), Gaps = 6/60 (10%)
Query: 21 YGECQKNHAANVGGYAVDGCREFMPSTNG------SLTCAACGCHRNFHKREVEVVSESS 74
Y EC KNHAA +GG+A+DGC EFMPS + SL CAACGCHRNFH+R+ + ++SS
Sbjct: 56 YKECLKNHAAALGGHALDGCGEFMPSPSSISSDPTSLKCAACGCHRNFHRRDPDNNNDSS 115
>ref|XP_450932.1| putative ZF-HD homeobox protein [Oryza sativa (japonica
cultivar-group)] gi|46806323|dbj|BAD17515.1| putative
ZF-HD homeobox protein [Oryza sativa (japonica
cultivar-group)]
Length = 279
Score = 75.9 bits (185), Expect = 2e-13
Identities = 36/73 (49%), Positives = 46/73 (62%), Gaps = 4/73 (5%)
Query: 10 RSEDPQRSVVKYGECQKNHAANVGGYAVDGCREFMPS----TNGSLTCAACGCHRNFHKR 65
R + P +Y EC KNHA +GG+AVDGC EFM + T +L CAAC CHRNFH++
Sbjct: 46 RVKGPSCGGGRYRECLKNHAVGIGGHAVDGCGEFMAAGEEGTIDALRCAACNCHRNFHRK 105
Query: 66 EVEVVSESSSPHS 78
E E ++ SP S
Sbjct: 106 ESESLAGEGSPFS 118
>emb|CAB42918.1| putative protein [Arabidopsis thaliana]
gi|15230335|ref|NP_190658.1| zinc finger homeobox family
protein / ZF-HD homeobox family protein [Arabidopsis
thaliana] gi|51969440|dbj|BAD43412.1| unknown protein
[Arabidopsis thaliana] gi|7485700|pir||T08410
hypothetical protein F18B3.170 - Arabidopsis thaliana
Length = 249
Score = 75.5 bits (184), Expect = 3e-13
Identities = 37/78 (47%), Positives = 48/78 (61%), Gaps = 7/78 (8%)
Query: 7 VVRRSEDPQRSV---VKYGECQKNHAANVGGYAVDGCREFM----PSTNGSLTCAACGCH 59
+ + ++ P+ V KY ECQKNHAA+ GG+ VDGC EFM T G+L CAAC CH
Sbjct: 43 IKKENQKPKTRVDQGAKYRECQKNHAASTGGHVVDGCCEFMAGGEEGTLGALKCAACNCH 102
Query: 60 RNFHKREVEVVSESSSPH 77
R+FH++EV S H
Sbjct: 103 RSFHRKEVYGHRNSKQDH 120
>gb|AAX95984.1| ZF-HD protein dimerisation region, putative [Oryza sativa (japonica
cultivar-group)]
Length = 383
Score = 75.1 bits (183), Expect = 4e-13
Identities = 36/60 (60%), Positives = 42/60 (70%), Gaps = 4/60 (6%)
Query: 19 VKYGECQKNHAANVGGYAVDGCREFMPS-TNGS---LTCAACGCHRNFHKREVEVVSESS 74
VKY EC KNHAA +GG A DGC EFMPS GS L C+ACGCHRNFH++E + + S
Sbjct: 143 VKYRECLKNHAAAIGGNATDGCGEFMPSGEEGSLEALKCSACGCHRNFHRKEADDLDADS 202
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.316 0.129 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 137,109,076
Number of Sequences: 2540612
Number of extensions: 4497916
Number of successful extensions: 8967
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 44
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 8881
Number of HSP's gapped (non-prelim): 45
length of query: 81
length of database: 863,360,394
effective HSP length: 57
effective length of query: 24
effective length of database: 718,545,510
effective search space: 17245092240
effective search space used: 17245092240
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 68 (30.8 bits)
Medicago: description of AC139600.10


