

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139356.2 + phase: 0
(180 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
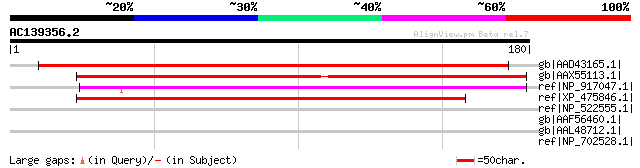
Score E
Sequences producing significant alignments: (bits) Value
gb|AAD43165.1| Hypothetical Protein [Arabidopsis thaliana] gi|15... 172 4e-42
gb|AAX55113.1| hypothetical protein At2g16190 [Arabidopsis thali... 163 2e-39
ref|NP_917047.1| P0034C09.31 [Oryza sativa (japonica cultivar-gr... 152 5e-36
ref|XP_475846.1| unknown protein [Oryza sativa (japonica cultiva... 142 4e-33
ref|NP_522555.1| PROBABLE EXCINUCLEASE ABC SUBUNIT A (DNA REPAIR... 35 0.73
gb|AAF56460.1| CG11902-PA [Drosophila melanogaster] gi|24650050|... 34 2.1
gb|AAL48712.1| RE15630p [Drosophila melanogaster] 34 2.1
ref|NP_702528.1| DNA-3-methyladenine glycosylase, putative [Plas... 33 2.8
>gb|AAD43165.1| Hypothetical Protein [Arabidopsis thaliana]
gi|15222106|ref|NP_175358.1| hydroxyproline-rich
glycoprotein family protein [Arabidopsis thaliana]
gi|25372894|pir||E96529 hypothetical protein F13F21.24
[imported] - Arabidopsis thaliana
Length = 331
Score = 172 bits (436), Expect = 4e-42
Identities = 83/163 (50%), Positives = 110/163 (66%)
Query: 11 RRRSAPRSGPIPIPFIWATNHRAKIHNLNHLLQNKIFSITGDVQCKSCQTKFQMPFDLRT 70
R + +S I PF WATN R +I +L +L N+I +ITG+VQC+ C+ +Q+ ++LR
Sbjct: 157 RSTVSKKSDTISPPFPWATNRRGEIQSLEYLESNQITTITGEVQCRHCEKVYQVSYNLRE 216
Query: 71 KFDVISQYLVTNMNTMHDRAPKILMNPRLPRCAHCNQENSVKPVIAEKKKNINWLFLLLG 130
+F + ++ +T M DRA K P RC C +E +VKPVIAE+K INWLFLLLG
Sbjct: 217 RFAEVVKFYLTEKRKMRDRAHKDWAYPEQRRCELCGREKAVKPVIAERKSQINWLFLLLG 276
Query: 131 QMLGCCKLEQLKYFCKYNNHHRTGAKNRVLYLTYFELCQQLDP 173
Q LG C LEQLK FCK++ +HRTGAK+RVLYLTY LC+ L P
Sbjct: 277 QTLGFCTLEQLKNFCKHSKNHRTGAKDRVLYLTYMGLCKMLQP 319
>gb|AAX55113.1| hypothetical protein At2g16190 [Arabidopsis thaliana]
gi|52354251|gb|AAU44446.1| hypothetical protein
AT2G16190 [Arabidopsis thaliana]
gi|4678214|gb|AAD26960.1| hypothetical protein
[Arabidopsis thaliana] gi|15226704|ref|NP_179215.1|
hypothetical protein [Arabidopsis thaliana]
gi|25372895|pir||F84537 hypothetical protein At2g16190
[imported] - Arabidopsis thaliana
Length = 303
Score = 163 bits (413), Expect = 2e-39
Identities = 76/156 (48%), Positives = 101/156 (64%), Gaps = 2/156 (1%)
Query: 24 PFIWATNHRAKIHNLNHLLQNKIFSITGDVQCKSCQTKFQMPFDLRTKFDVISQYLVTNM 83
P+ WAT KI + L N I I+G V CK+C + ++L KF + Y+ N
Sbjct: 150 PYPWATKKPGKIQSFRDLSSNNINVISGQVHCKTCDRTDTVEYNLEEKFSELYGYIKVNK 209
Query: 84 NTMHDRAPKILMNPRLPRCAHCNQENSVKPVIAEKKKNINWLFLLLGQMLGCCKLEQLKY 143
M RAP P+L C C E +KPV++E+K+ INWLFLLLGQMLGCC L+QL+Y
Sbjct: 210 EEMRHRAPGSWSTPKLIPCRTCKSE--MKPVMSERKEEINWLFLLLGQMLGCCTLDQLRY 267
Query: 144 FCKYNNHHRTGAKNRVLYLTYFELCQQLDPSGPFNV 179
FC+ N+ HRTG+K+RV+Y+TY LC+QLDP GPFN+
Sbjct: 268 FCQLNSKHRTGSKDRVVYITYLSLCKQLDPEGPFNL 303
>ref|NP_917047.1| P0034C09.31 [Oryza sativa (japonica cultivar-group)]
gi|18565431|dbj|BAB84618.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 316
Score = 152 bits (383), Expect = 5e-36
Identities = 72/157 (45%), Positives = 95/157 (59%), Gaps = 2/157 (1%)
Query: 25 FIWATNHRAKIHN--LNHLLQNKIFSITGDVQCKSCQTKFQMPFDLRTKFDVISQYLVTN 82
F W T+ + + L +L I S+ G C C + + +DL KF + Y+ N
Sbjct: 160 FPWVTSADLPVLHCTLESMLLKGITSVEGKATCNRCSAEVPIAYDLDAKFREVRDYVAAN 219
Query: 83 MNTMHDRAPKILMNPRLPRCAHCNQENSVKPVIAEKKKNINWLFLLLGQMLGCCKLEQLK 142
++ M DRAP+ M PRLP C C ++ + P I +K+ INWLFL LGQMLGCC LE LK
Sbjct: 220 IHIMDDRAPEHWMCPRLPDCGSCGKKACMWPQIPNEKREINWLFLFLGQMLGCCTLEGLK 279
Query: 143 YFCKYNNHHRTGAKNRVLYLTYFELCQQLDPSGPFNV 179
+FCK +H TGAK RVLY Y E+C+QLDP GPFN+
Sbjct: 280 FFCKNTKNHCTGAKTRVLYYAYIEMCRQLDPQGPFNI 316
>ref|XP_475846.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|47900492|gb|AAT39249.1| unknown protein [Oryza sativa
(japonica cultivar-group)]
Length = 315
Score = 142 bits (358), Expect = 4e-33
Identities = 65/135 (48%), Positives = 88/135 (65%)
Query: 24 PFIWATNHRAKIHNLNHLLQNKIFSITGDVQCKSCQTKFQMPFDLRTKFDVISQYLVTNM 83
P+ WATN AK H+L L + I +I G+ +C+ C T+ + +++ TKF +S Y N
Sbjct: 172 PYPWATNEVAKHHSLVELARRDIININGEARCRRCDTRKMIVYNIATKFREVSDYFRQNY 231
Query: 84 NTMHDRAPKILMNPRLPRCAHCNQENSVKPVIAEKKKNINWLFLLLGQMLGCCKLEQLKY 143
M+DRA MNP +P C C E ++PVIA +K+ INWLFLLLG+ LG C L+QLKY
Sbjct: 232 QHMNDRAQARWMNPVVPNCDSCGHERCMRPVIAAEKERINWLFLLLGETLGLCTLDQLKY 291
Query: 144 FCKYNNHHRTGAKNR 158
FC + N HRTGAK+R
Sbjct: 292 FCAHTNRHRTGAKDR 306
>ref|NP_522555.1| PROBABLE EXCINUCLEASE ABC SUBUNIT A (DNA REPAIR ATP-BINDING) ABC
TRANSPORTER PROTEIN [Ralstonia solanacearum GMI1000]
gi|17431467|emb|CAD18145.1| PROBABLE EXCINUCLEASE ABC
SUBUNIT A (DNA REPAIR ATP-BINDING) ABC TRANSPORTER
PROTEIN [Ralstonia solanacearum]
Length = 1945
Score = 35.4 bits (80), Expect = 0.73
Identities = 19/51 (37%), Positives = 26/51 (50%), Gaps = 4/51 (7%)
Query: 5 HTHVPPRRRSAPRSGPIPIPFIWATNHRAKIHNLNHLLQN----KIFSITG 51
H P R +APRSG P W T H A +HNL +L ++ ++TG
Sbjct: 1545 HPLQPRREVAAPRSGAAASPERWLTVHGASLHNLRNLTATLPLARLVAVTG 1595
>gb|AAF56460.1| CG11902-PA [Drosophila melanogaster] gi|24650050|ref|NP_651389.2|
CG11902-PA [Drosophila melanogaster]
Length = 1266
Score = 33.9 bits (76), Expect = 2.1
Identities = 18/51 (35%), Positives = 26/51 (50%), Gaps = 4/51 (7%)
Query: 22 PIPFIWATN---HRAKIHNLNHL-LQNKIFSITGDVQCKSCQTKFQMPFDL 68
P+ F W N H H L+ L +Q+ + + GD+QC C F+M DL
Sbjct: 770 PLRFRWRENLKLHMEVFHMLDELSIQDDLPKVDGDLQCGKCNRSFRMQKDL 820
>gb|AAL48712.1| RE15630p [Drosophila melanogaster]
Length = 1426
Score = 33.9 bits (76), Expect = 2.1
Identities = 18/51 (35%), Positives = 26/51 (50%), Gaps = 4/51 (7%)
Query: 22 PIPFIWATN---HRAKIHNLNHL-LQNKIFSITGDVQCKSCQTKFQMPFDL 68
P+ F W N H H L+ L +Q+ + + GD+QC C F+M DL
Sbjct: 930 PLRFRWRENLKLHMEVFHMLDELSIQDDLPKVDGDLQCGKCNRSFRMQKDL 980
>ref|NP_702528.1| DNA-3-methyladenine glycosylase, putative [Plasmodium falciparum
3D7] gi|23497713|gb|AAN37252.1| DNA-3-methyladenine
glycosylase, putative [Plasmodium falciparum 3D7]
Length = 501
Score = 33.5 bits (75), Expect = 2.8
Identities = 25/88 (28%), Positives = 42/88 (47%), Gaps = 6/88 (6%)
Query: 35 IHNLNHLLQNKIFSITGDVQCKSCQTKFQMPFDLRTKFDVISQYLVTNMNTMHDRAPKIL 94
IHN +++ N I +I + +S + + + + L KF+ ISQYL++N +H +
Sbjct: 172 IHNCLNIVTN-IQNIPDAILVRSTEPLYNIEYFLSNKFEKISQYLLSNSIFIHKNIKEQQ 230
Query: 95 MNPRLPRCAHCNQENSVKPVIAEKKKNI 122
N P+ Q N I+ K NI
Sbjct: 231 KNNSTPKNKRNQQGN-----ISNKDGNI 253
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.325 0.138 0.449
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 321,399,449
Number of Sequences: 2540612
Number of extensions: 12303692
Number of successful extensions: 32239
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 32233
Number of HSP's gapped (non-prelim): 8
length of query: 180
length of database: 863,360,394
effective HSP length: 120
effective length of query: 60
effective length of database: 558,486,954
effective search space: 33509217240
effective search space used: 33509217240
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 71 (32.0 bits)
Medicago: description of AC139356.2


