

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138130.17 + phase: 0 /pseudo
(626 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
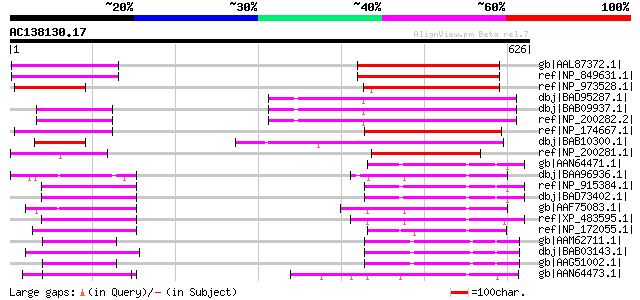
Score E
Sequences producing significant alignments: (bits) Value
gb|AAL87372.1| At1g10340/F14N23_22 [Arabidopsis thaliana] gi|183... 192 4e-47
ref|NP_849631.1| ankyrin repeat family protein [Arabidopsis thal... 192 4e-47
ref|NP_973528.1| ankyrin repeat family protein [Arabidopsis thal... 191 8e-47
dbj|BAD95287.1| putative protein [Arabidopsis thaliana] 179 2e-43
dbj|BAB09937.1| unnamed protein product [Arabidopsis thaliana] 179 2e-43
ref|NP_200282.2| ankyrin repeat family protein [Arabidopsis thal... 179 2e-43
ref|NP_174667.1| ankyrin repeat family protein [Arabidopsis thal... 179 2e-43
dbj|BAB10300.1| ankyrin-like protein [Arabidopsis thaliana] gi|1... 169 3e-40
ref|NP_200281.1| ankyrin repeat family protein [Arabidopsis thal... 147 1e-33
gb|AAN64471.1| hypothetical protein, 5'-partial [Oryza sativa (j... 102 4e-20
dbj|BAA96936.1| ankyrin-like protein [Arabidopsis thaliana] gi|6... 102 4e-20
ref|NP_915384.1| P0506B12.26 [Oryza sativa (japonica cultivar-gr... 98 7e-19
dbj|BAD73402.1| ankyrin-like protein [Oryza sativa (japonica cul... 98 7e-19
gb|AAF75083.1| It contains Ank repeat PF|00023. EST gb|AI996003... 97 1e-18
ref|XP_483595.1| ankyrin-like protein [Oryza sativa (japonica cu... 97 2e-18
ref|NP_172055.1| ankyrin repeat family protein [Arabidopsis thal... 96 3e-18
gb|AAM62711.1| ankyrin-like protein [Arabidopsis thaliana] 95 6e-18
dbj|BAB03143.1| ankyrin-like protein [Arabidopsis thaliana] 95 6e-18
gb|AAG51002.1| ankyrin-like protein; 93648-91299 [Arabidopsis th... 95 6e-18
gb|AAN64473.1| putative ankyrin repeat containing protein [Oryza... 94 1e-17
>gb|AAL87372.1| At1g10340/F14N23_22 [Arabidopsis thaliana]
gi|18391143|ref|NP_563867.1| ankyrin repeat family
protein [Arabidopsis thaliana]
gi|13937240|gb|AAK50112.1| At1g10340/F14N23_22
[Arabidopsis thaliana] gi|4914336|gb|AAD32884.1|
F14N23.22 [Arabidopsis thaliana]
Length = 578
Score = 192 bits (487), Expect = 4e-47
Identities = 92/171 (53%), Positives = 123/171 (71%)
Query: 420 KGKIENVNHTKRKHYHEMHKEALLNARNTIVLVAVLIATVTFAAGISPPGGVYQEGPKKG 479
+ K + V ++ ++MH EAL NARNTI +VAVLIA+V +A GI+PPGGVYQ+GP +G
Sbjct: 379 RSKEQEVERGRQNLEYQMHIEALQNARNTIAIVAVLIASVAYAGGINPPGGVYQDGPWRG 438
Query: 480 ISMAGETSAFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFM 539
S+ G+T+AFKVFAI N IALFTSL +VI+LVSIIP++RKP LL H++MWV+V FM
Sbjct: 439 KSLVGKTTAFKVFAICNNIALFTSLGIVILLVSIIPYKRKPLKRLLVATHRMMWVSVGFM 498
Query: 540 GTGYVAATWVILPHNQEMQWLSVVLLALGGGFLGTIFISLSVMLVEHWLRK 590
T Y+AA+WV +PH QWL ++A+ GG L +F L V + HW +K
Sbjct: 499 ATAYIAASWVTIPHYHGTQWLFPAIVAVAGGALTVLFFYLGVETIGHWFKK 549
Score = 69.3 bits (168), Expect = 3e-10
Identities = 38/131 (29%), Positives = 69/131 (52%), Gaps = 2/131 (1%)
Query: 3 QEFFDAIKKNDMITFSSIVKEREGILNQKTDDTF--SAPLHLASKYGCIEMVSEIVKLCP 60
Q F AI KND+ F +V++ E L ++ ++ + LH+A+K+G E+VS+I++L P
Sbjct: 2 QPIFHAILKNDLPAFLELVEDSESSLEERNEEEHLNNTVLHMAAKFGHRELVSKIIELRP 61
Query: 61 DMVSAENKNMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLDLV 120
+VS+ N TP+H A +V ++M +LE N + +AC ++
Sbjct: 62 SLVSSRNAYRNTPLHLAAILGDVNIVMQMLETGLEVCSARNINNHTPLHLACRSNSIEAA 121
Query: 121 NLLLNLSEIVG 131
L+ ++ +G
Sbjct: 122 RLIAEKTQSIG 132
Score = 37.0 bits (84), Expect = 1.9
Identities = 29/129 (22%), Positives = 55/129 (42%), Gaps = 5/129 (3%)
Query: 23 EREGILNQKTDDTFSAPLHLASKYGCIEMVSEIVKLCPDMVS-----AENKNMETPIHEA 77
E ++ +KT L LA G +V I++ PD+ E+ + T +H A
Sbjct: 119 EAARLIAEKTQSIGLGELILAISSGSTSIVGTILERFPDLAREEAWVVEDGSQSTLLHHA 178
Query: 78 CRQENVKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLDLVNLLLNLSEIVGQEVAGF 137
C + + ++ +LL ++ LNP S +A G + ++ L+ + +
Sbjct: 179 CDKGDFELTTILLGLDQGLEEALNPNGLSPLHLAVLRGSVVILEEFLDKVPLSFSSITPS 238
Query: 138 DQACFHVAA 146
+ FH+AA
Sbjct: 239 KETVFHLAA 247
Score = 35.8 bits (81), Expect = 4.3
Identities = 30/134 (22%), Positives = 60/134 (44%), Gaps = 7/134 (5%)
Query: 17 FSSIVKEREGILNQKTDDTFSAPLHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHE 76
F + +E ++ + T LH A G E+ + ++ L + A N N +P+H
Sbjct: 155 FPDLAREEAWVVEDGSQSTL---LHHACDKGDFELTTILLGLDQGLEEALNPNGLSPLHL 211
Query: 77 ACRQENVKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLD----LVNLLLNLSEIVGQ 132
A + +V +L L+ P + + P+ ++ F +A + ++D + L S+I+ Q
Sbjct: 212 AVLRGSVVILEEFLDKVPLSFSSITPSKETVFHLAARNKNMDAFVFMAESLGINSQILLQ 271
Query: 133 EVAGFDQACFHVAA 146
+ H+AA
Sbjct: 272 QTDESGNTVLHIAA 285
>ref|NP_849631.1| ankyrin repeat family protein [Arabidopsis thaliana]
Length = 574
Score = 192 bits (487), Expect = 4e-47
Identities = 92/171 (53%), Positives = 123/171 (71%)
Query: 420 KGKIENVNHTKRKHYHEMHKEALLNARNTIVLVAVLIATVTFAAGISPPGGVYQEGPKKG 479
+ K + V ++ ++MH EAL NARNTI +VAVLIA+V +A GI+PPGGVYQ+GP +G
Sbjct: 375 RSKEQEVERGRQNLEYQMHIEALQNARNTIAIVAVLIASVAYAGGINPPGGVYQDGPWRG 434
Query: 480 ISMAGETSAFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFM 539
S+ G+T+AFKVFAI N IALFTSL +VI+LVSIIP++RKP LL H++MWV+V FM
Sbjct: 435 KSLVGKTTAFKVFAICNNIALFTSLGIVILLVSIIPYKRKPLKRLLVATHRMMWVSVGFM 494
Query: 540 GTGYVAATWVILPHNQEMQWLSVVLLALGGGFLGTIFISLSVMLVEHWLRK 590
T Y+AA+WV +PH QWL ++A+ GG L +F L V + HW +K
Sbjct: 495 ATAYIAASWVTIPHYHGTQWLFPAIVAVAGGALTVLFFYLGVETIGHWFKK 545
Score = 69.3 bits (168), Expect = 3e-10
Identities = 38/131 (29%), Positives = 69/131 (52%), Gaps = 2/131 (1%)
Query: 3 QEFFDAIKKNDMITFSSIVKEREGILNQKTDDTF--SAPLHLASKYGCIEMVSEIVKLCP 60
Q F AI KND+ F +V++ E L ++ ++ + LH+A+K+G E+VS+I++L P
Sbjct: 2 QPIFHAILKNDLPAFLELVEDSESSLEERNEEEHLNNTVLHMAAKFGHRELVSKIIELRP 61
Query: 61 DMVSAENKNMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLDLV 120
+VS+ N TP+H A +V ++M +LE N + +AC ++
Sbjct: 62 SLVSSRNAYRNTPLHLAAILGDVNIVMQMLETGLEVCSARNINNHTPLHLACRSNSIEAA 121
Query: 121 NLLLNLSEIVG 131
L+ ++ +G
Sbjct: 122 RLIAEKTQSIG 132
Score = 35.8 bits (81), Expect = 4.3
Identities = 30/134 (22%), Positives = 60/134 (44%), Gaps = 7/134 (5%)
Query: 17 FSSIVKEREGILNQKTDDTFSAPLHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHE 76
F + +E ++ + T LH A G E+ + ++ L + A N N +P+H
Sbjct: 151 FPDLAREEAWVVEDGSQSTL---LHHACDKGDFELTTILLGLDQGLEEALNPNGLSPLHL 207
Query: 77 ACRQENVKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLD----LVNLLLNLSEIVGQ 132
A + +V +L L+ P + + P+ ++ F +A + ++D + L S+I+ Q
Sbjct: 208 AVLRGSVVILEEFLDKVPLSFSSITPSKETVFHLAARNKNMDAFVFMAESLGINSQILLQ 267
Query: 133 EVAGFDQACFHVAA 146
+ H+AA
Sbjct: 268 QTDESGNTVLHIAA 281
>ref|NP_973528.1| ankyrin repeat family protein [Arabidopsis thaliana]
Length = 601
Score = 191 bits (484), Expect = 8e-47
Identities = 96/167 (57%), Positives = 122/167 (72%), Gaps = 3/167 (1%)
Query: 427 NHTKRKHY---HEMHKEALLNARNTIVLVAVLIATVTFAAGISPPGGVYQEGPKKGISMA 483
+H KR H HEMH EAL NARNTI +VAVLIA+V++A GI+PPGGVYQ+GP KG S+
Sbjct: 383 HHVKRGHKSLEHEMHIEALQNARNTIAIVAVLIASVSYAGGINPPGGVYQDGPWKGKSLV 442
Query: 484 GETSAFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGY 543
G T+AFKVFAI N IALFTSL +VI+LVSIIP++RKP LL H++MWV+V FM T Y
Sbjct: 443 GNTAAFKVFAICNNIALFTSLCIVILLVSIIPYQRKPLKKLLVATHRMMWVSVGFMATAY 502
Query: 544 VAATWVILPHNQEMQWLSVVLLALGGGFLGTIFISLSVMLVEHWLRK 590
VAA+ V +PH +WL V++++ GG L +F L V + HW +K
Sbjct: 503 VAASLVTIPHFPGTRWLFPVIISVAGGSLTVLFSYLGVETISHWFKK 549
Score = 61.2 bits (147), Expect = 9e-08
Identities = 31/88 (35%), Positives = 58/88 (65%), Gaps = 2/88 (2%)
Query: 6 FDAIKKNDMITFSSIVKEREGILNQKTDD--TFSAPLHLASKYGCIEMVSEIVKLCPDMV 63
FDAI +ND+ F +V+ RE L +++++ T + LH+A+K G E+V++I++L P ++
Sbjct: 5 FDAILQNDLPAFLGLVEARESSLEERSEEQNTNNTVLHVAAKLGHRELVAKIIELRPSLL 64
Query: 64 SAENKNMETPIHEACRQENVKVLMLLLE 91
S+ N +TP+H A +V ++M +L+
Sbjct: 65 SSRNAYGDTPLHLAALLGDVNIVMQMLD 92
Score = 36.6 bits (83), Expect = 2.5
Identities = 22/86 (25%), Positives = 44/86 (50%)
Query: 33 DDTFSAPLHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACRQENVKVLMLLLEV 92
D + S LH A G +E+ S ++ L + A N +P+H A ++ +V +L ++
Sbjct: 168 DGSRSTLLHYACDKGDLELTSILLGLNQGLEEALNSKGLSPLHLAVQRGSVIILEEFMDK 227
Query: 93 NPTAACKLNPTCKSAFLVACSHGHLD 118
+P + C P+ ++ F +A + + D
Sbjct: 228 SPLSFCVRTPSKETVFHLAARNKNTD 253
>dbj|BAD95287.1| putative protein [Arabidopsis thaliana]
Length = 422
Score = 179 bits (455), Expect = 2e-43
Identities = 106/306 (34%), Positives = 164/306 (52%), Gaps = 11/306 (3%)
Query: 313 AENRQLQAIFIRDGGKRSTPSSFSLELDNTSSPSPTSRHSLSRRYISKEMEVLTEMVSYD 372
A++ ++ + + +T + +D+TS RH LS I ++ + D
Sbjct: 112 AQSANIRQLLYSLDAEDNTVLHVAASVDSTS----LVRHILSETTIDVTLKNKKGFAAVD 167
Query: 373 CISPPPVSESTESISPQPQVSERFENGTYNPYYFSPTNLVKQKHHHNKGKIEN------- 425
I V S+ + + + Y + P L++ ++ K E+
Sbjct: 168 LIDKEGVDFPLLSLWFRDEAEKIQRPARYVKFAHEPVELIRNTNNGEKLSSESRAMDLLR 227
Query: 426 VNHTKRKHYHEMHKEALLNARNTIVLVAVLIATVTFAAGISPPGGVYQEGPKKGISMAGE 485
R EMH E+L NARNTI +VAVLIA+V F GI+PPGGV+Q+GP G + AG
Sbjct: 228 EGRDPRNKEREMHSESLQNARNTITIVAVLIASVAFTCGINPPGGVHQDGPFIGKATAGR 287
Query: 486 TSAFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGYVA 545
T AFK+F+++N IALFTSLS+V +LVSII +R K + + IAHK+MW+AVA M T Y A
Sbjct: 288 TLAFKIFSVANNIALFTSLSIVTLLVSIISYRTKALKMCVVIAHKMMWLAVASMATAYAA 347
Query: 546 ATWVILPHNQEMQWLSVVLLALGGGFLGTIFISLSVMLVEHWLRKSSWRKKRKESGDGTA 605
+ W+ +PHN+ +WL A+ LG++F+ +S M+V+H L+K R+ + +
Sbjct: 348 SAWITVPHNEGSKWLVYTTSAIASVALGSMFVYVSFMMVKHILKKDKLRRNQSHGKTPST 407
Query: 606 ESDKES 611
D E+
Sbjct: 408 TLDMEA 413
>dbj|BAB09937.1| unnamed protein product [Arabidopsis thaliana]
Length = 652
Score = 179 bits (455), Expect = 2e-43
Identities = 106/306 (34%), Positives = 164/306 (52%), Gaps = 11/306 (3%)
Query: 313 AENRQLQAIFIRDGGKRSTPSSFSLELDNTSSPSPTSRHSLSRRYISKEMEVLTEMVSYD 372
A++ ++ + + +T + +D+TS RH LS I ++ + D
Sbjct: 288 AQSANIRQLLYSLDAEDNTVLHVAASVDSTS----LVRHILSETTIDVTLKNKKGFAAVD 343
Query: 373 CISPPPVSESTESISPQPQVSERFENGTYNPYYFSPTNLVKQKHHHNKGKIEN------- 425
I V S+ + + + Y + P L++ ++ K E+
Sbjct: 344 LIDKEGVDFPLLSLWFRDEAEKIQRPARYVKFAHEPVELIRNTNNGEKLSSESRAMDLLR 403
Query: 426 VNHTKRKHYHEMHKEALLNARNTIVLVAVLIATVTFAAGISPPGGVYQEGPKKGISMAGE 485
R EMH E+L NARNTI +VAVLIA+V F GI+PPGGV+Q+GP G + AG
Sbjct: 404 EGRDPRNKEREMHSESLQNARNTITIVAVLIASVAFTCGINPPGGVHQDGPFIGKATAGR 463
Query: 486 TSAFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGYVA 545
T AFK+F+++N IALFTSLS+V +LVSII +R K + + IAHK+MW+AVA M T Y A
Sbjct: 464 TLAFKIFSVANNIALFTSLSIVTLLVSIISYRTKALKMCVVIAHKMMWLAVASMATAYAA 523
Query: 546 ATWVILPHNQEMQWLSVVLLALGGGFLGTIFISLSVMLVEHWLRKSSWRKKRKESGDGTA 605
+ W+ +PHN+ +WL A+ LG++F+ +S M+V+H L+K R+ + +
Sbjct: 524 SAWITVPHNEGSKWLVYTTSAIASVALGSMFVYVSFMMVKHILKKDKLRRNQSHGKTPST 583
Query: 606 ESDKES 611
D E+
Sbjct: 584 TLDMEA 589
Score = 55.1 bits (131), Expect = 7e-06
Identities = 27/92 (29%), Positives = 49/92 (52%)
Query: 33 DDTFSAPLHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACRQENVKVLMLLLEV 92
D+ S LH A E ++++ LCP +VS N + TP+H A N+ +L +LE
Sbjct: 65 DEEQSTLLHKAVTQRNEEYATKVIDLCPSLVSVTNVDGNTPLHLAAEIGNINILWKMLET 124
Query: 93 NPTAACKLNPTCKSAFLVACSHGHLDLVNLLL 124
K+N ++AF++AC + +++ +L+
Sbjct: 125 GEAECMKINKQGQTAFILACLNNNVNSARILV 156
Score = 39.3 bits (90), Expect = 0.39
Identities = 20/46 (43%), Positives = 29/46 (62%), Gaps = 5/46 (10%)
Query: 522 TILLTIAHKVMWVAVAFMGTGYVAATWVILPHNQEM-----QWLSV 562
T LL I H++MWV VA M + Y+AA+ + LPH ++ +WL V
Sbjct: 598 TKLLVITHRLMWVDVAAMESAYLAASSITLPHFGDVMLLCYRWLDV 643
>ref|NP_200282.2| ankyrin repeat family protein [Arabidopsis thaliana]
Length = 598
Score = 179 bits (455), Expect = 2e-43
Identities = 106/306 (34%), Positives = 164/306 (52%), Gaps = 11/306 (3%)
Query: 313 AENRQLQAIFIRDGGKRSTPSSFSLELDNTSSPSPTSRHSLSRRYISKEMEVLTEMVSYD 372
A++ ++ + + +T + +D+TS RH LS I ++ + D
Sbjct: 288 AQSANIRQLLYSLDAEDNTVLHVAASVDSTS----LVRHILSETTIDVTLKNKKGFAAVD 343
Query: 373 CISPPPVSESTESISPQPQVSERFENGTYNPYYFSPTNLVKQKHHHNKGKIEN------- 425
I V S+ + + + Y + P L++ ++ K E+
Sbjct: 344 LIDKEGVDFPLLSLWFRDEAEKIQRPARYVKFAHEPVELIRNTNNGEKLSSESRAMDLLR 403
Query: 426 VNHTKRKHYHEMHKEALLNARNTIVLVAVLIATVTFAAGISPPGGVYQEGPKKGISMAGE 485
R EMH E+L NARNTI +VAVLIA+V F GI+PPGGV+Q+GP G + AG
Sbjct: 404 EGRDPRNKEREMHSESLQNARNTITIVAVLIASVAFTCGINPPGGVHQDGPFIGKATAGR 463
Query: 486 TSAFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGYVA 545
T AFK+F+++N IALFTSLS+V +LVSII +R K + + IAHK+MW+AVA M T Y A
Sbjct: 464 TLAFKIFSVANNIALFTSLSIVTLLVSIISYRTKALKMCVVIAHKMMWLAVASMATAYAA 523
Query: 546 ATWVILPHNQEMQWLSVVLLALGGGFLGTIFISLSVMLVEHWLRKSSWRKKRKESGDGTA 605
+ W+ +PHN+ +WL A+ LG++F+ +S M+V+H L+K R+ + +
Sbjct: 524 SAWITVPHNEGSKWLVYTTSAIASVALGSMFVYVSFMMVKHILKKDKLRRNQSHGKTPST 583
Query: 606 ESDKES 611
D E+
Sbjct: 584 TLDMEA 589
Score = 55.1 bits (131), Expect = 7e-06
Identities = 27/92 (29%), Positives = 49/92 (52%)
Query: 33 DDTFSAPLHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACRQENVKVLMLLLEV 92
D+ S LH A E ++++ LCP +VS N + TP+H A N+ +L +LE
Sbjct: 65 DEEQSTLLHKAVTQRNEEYATKVIDLCPSLVSVTNVDGNTPLHLAAEIGNINILWKMLET 124
Query: 93 NPTAACKLNPTCKSAFLVACSHGHLDLVNLLL 124
K+N ++AF++AC + +++ +L+
Sbjct: 125 GEAECMKINKQGQTAFILACLNNNVNSARILV 156
>ref|NP_174667.1| ankyrin repeat family protein [Arabidopsis thaliana]
gi|25518163|pir||D86464 F12G12.13 protein - Arabidopsis
thaliana gi|10086472|gb|AAG12532.1| Hypothetical Protein
[Arabidopsis thaliana]
Length = 573
Score = 179 bits (454), Expect = 2e-43
Identities = 88/165 (53%), Positives = 118/165 (71%)
Query: 429 TKRKHYHEMHKEALLNARNTIVLVAVLIATVTFAAGISPPGGVYQEGPKKGISMAGETSA 488
T K +MH EALLNARNTI +VAVLIA+V F GI+PPGGVYQEGP KG S AG T A
Sbjct: 379 TPSKRESKMHAEALLNARNTITIVAVLIASVAFTCGINPPGGVYQEGPYKGKSTAGRTLA 438
Query: 489 FKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGYVAATW 548
F+VF+ISN IALFTSL +VI+LVSIIP+R +P L + H+++WVAVA M YV+A
Sbjct: 439 FQVFSISNNIALFTSLCIVILLVSIIPYRTRPLKNFLKLTHRILWVAVASMALAYVSAAS 498
Query: 549 VILPHNQEMQWLSVVLLALGGGFLGTIFISLSVMLVEHWLRKSSW 593
+I+PH + +WL +L++ LG +F ++ ++ HWL+K ++
Sbjct: 499 IIIPHVEGKRWLFTTVLSISTLMLGGLFAFMTYKVIRHWLKKMTF 543
Score = 65.1 bits (157), Expect = 7e-09
Identities = 37/120 (30%), Positives = 65/120 (53%), Gaps = 2/120 (1%)
Query: 7 DAIKKNDMITFSSIVKEREGILNQKT--DDTFSAPLHLASKYGCIEMVSEIVKLCPDMVS 64
DAI ND+ T ++ + +L ++ D LHLA++ G E+V I+KLCP +V
Sbjct: 23 DAILANDVSTLLALAEGNLSVLRERYHWDSLGGTVLHLATELGHKEIVEAIIKLCPSLVG 82
Query: 65 AENKNMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLDLVNLLL 124
N + +TP+H A R + ++ +L +N ++AF+VAC + + D+ +L+L
Sbjct: 83 VTNLDGDTPLHFAARWGHATIVAQILASGYAEFTPVNGRGETAFVVACRYTNPDVASLIL 142
Score = 37.0 bits (84), Expect = 1.9
Identities = 25/114 (21%), Positives = 48/114 (41%)
Query: 4 EFFDAIKKNDMITFSSIVKEREGILNQKTDDTFSAPLHLASKYGCIEMVSEIVKLCPDMV 63
EF+ + + + ER L D S PLH A +E+ ++++ +
Sbjct: 152 EFYATFVLGEYTDIARRMLERFPKLAWNADGELSTPLHHACNANNLEITKMLLEIDESLA 211
Query: 64 SAENKNMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHL 117
NK+ TP+H A + ++ +L + P L P ++ F +A H ++
Sbjct: 212 ERVNKDGFTPLHLAAMKCSIPILKEFSDKAPRYFDILTPAKETVFHLAAEHKNI 265
>dbj|BAB10300.1| ankyrin-like protein [Arabidopsis thaliana]
gi|15240620|ref|NP_199825.1| ankyrin repeat family
protein [Arabidopsis thaliana]
Length = 535
Score = 169 bits (427), Expect = 3e-40
Identities = 111/329 (33%), Positives = 168/329 (50%), Gaps = 9/329 (2%)
Query: 273 ATLSYISQFQAVAIRTKVDINTRNNEGLTALDILDQAMDNAENRQLQAIFIRDGGKRSTP 332
A++ ++S + K+++ T+N++G A+D+L++ D+ + + + I D
Sbjct: 188 ASVGFLSLVSYIVHEIKIEVTTQNDKGFEAVDLLNK--DDEDFKMMSMILGHDSEIVQRA 245
Query: 333 SSFSLELDNTSSPSPTSRHSLSRRYISKEMEVLTEMVS------YDCISPPPVSESTESI 386
+S + S+ + + E+ E V+ ++ I P + +
Sbjct: 246 ASSPRDAYTPSTQTEVENSEIHHEQGLVAPEIKEENVTNENNKVFEAIDLPTKEDGDLKM 305
Query: 387 SPQPQVSERFENGTYNPYYFSPTNLVKQKHHHNKGKIENVNHTKRKHYHEMHKEALLNAR 446
SE F+ + +P + + I + R EM EAL NAR
Sbjct: 306 LAGTD-SETFQLPSSRTGILTPETETEMVISNTLHGIRHGLRESRIKEKEMQSEALQNAR 364
Query: 447 NTIVLVAVLIATVTFAAGISPPGGVYQEGPKKGISMAGETSAFKVFAISNIIALFTSLSV 506
NTI +VAVLIA+VTF G++PPGGVYQ+G G + AG T AFKVF++SN IALFTSL +
Sbjct: 365 NTITVVAVLIASVTFTCGLNPPGGVYQDGHFIGKATAGGTVAFKVFSVSNSIALFTSLCI 424
Query: 507 VIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGYVAATWVILPHNQEMQWLSVVLLA 566
VI+L+SIIPFR K L I HK++W+AV M + YVA T V LPH++ +W+ L
Sbjct: 425 VILLLSIIPFRTKSLKTFLIITHKMIWLAVIAMASAYVAGTCVTLPHSRGNKWVLKATLV 484
Query: 567 LGGGFLGTIFISLSVMLVEHWLRKSSWRK 595
+ LG +FI L L H RK RK
Sbjct: 485 IACVMLGGMFIFLWFKLANHMSRKKKMRK 513
Score = 50.1 bits (118), Expect = 2e-04
Identities = 25/63 (39%), Positives = 41/63 (64%), Gaps = 1/63 (1%)
Query: 30 QKTDDTFSAP-LHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACRQENVKVLML 88
++ D++F LHLA K G E+V +IV++ P +VS+ N +TP+H A R + +L+L
Sbjct: 20 EEQDESFGGTFLHLAVKLGNEELVKKIVEIHPSLVSSTNTKSDTPLHLAARLGHTSILLL 79
Query: 89 LLE 91
+LE
Sbjct: 80 MLE 82
>ref|NP_200281.1| ankyrin repeat family protein [Arabidopsis thaliana]
Length = 480
Score = 147 bits (371), Expect = 1e-33
Identities = 76/132 (57%), Positives = 97/132 (72%), Gaps = 1/132 (0%)
Query: 437 MHKEALLNARNTIVLVAVLIATVTFAAGISPPGGVYQEGPK-KGISMAGETSAFKVFAIS 495
MH EAL NARNTI +VA+LIA+VTFA G++PPGG+YQE KG S+A +T AFK+F +S
Sbjct: 205 MHSEALQNARNTITVVAILIASVTFAVGMNPPGGIYQESTSSKGKSVAAKTVAFKIFYVS 264
Query: 496 NIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGYVAATWVILPHNQ 555
N IALFTSL +VI+LVSIIPF+ K +L I HK+M V+VA + T YVA W+ILPH +
Sbjct: 265 NSIALFTSLWIVILLVSIIPFKPKSLKNVLVITHKMMSVSVAALATSYVAVGWIILPHFE 324
Query: 556 EMQWLSVVLLAL 567
+WL L +
Sbjct: 325 GTKWLLYTTLGI 336
Score = 51.2 bits (121), Expect = 1e-04
Identities = 36/135 (26%), Positives = 66/135 (48%), Gaps = 18/135 (13%)
Query: 1 MDQEFFDAIKKNDMITFSSIVKEREGILNQK-TDDTFSAPLHLASKYGCIEMVSEIVKLC 59
M F+AI+KND TF+ +++E+ ++ ++ ++ + LHL +K G E I+ +C
Sbjct: 1 MTPPIFNAIRKNDEATFNQLIQEKPSVIEERDKENNGESVLHLVTKIGHQEFAKTIIGIC 60
Query: 60 P--------------DMVSAE--NKNMETPIHEACRQENVKVLMLLLEVNPTAACKL-NP 102
P D+ AE N + TP+H A ++K+L + P++ L P
Sbjct: 61 PSLSTPLDDISEVENDLKLAELVNNDGLTPLHCAAVSNSIKILKVFSHKTPSSFDILTQP 120
Query: 103 TCKSAFLVACSHGHL 117
++ F +A H +L
Sbjct: 121 HNETVFHLAVRHKNL 135
>gb|AAN64471.1| hypothetical protein, 5'-partial [Oryza sativa (japonica
cultivar-group)]
Length = 284
Score = 102 bits (254), Expect = 4e-20
Identities = 59/192 (30%), Positives = 108/192 (55%), Gaps = 10/192 (5%)
Query: 432 KHYHEMHKEALLNARNTIVLVAVLIATVTFAAGISPPGGVYQEGPKKGISMAGETSAFKV 491
K ++H+E + NA N++ +VAVL ATV FAA + PGG G+++A +FK+
Sbjct: 98 KELRKLHREGINNATNSVTVVAVLFATVAFAAIFTVPGG----NDNNGVAIAVHAVSFKI 153
Query: 492 FAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGYVAATWVIL 551
F I N IALFTSL+VV+V ++++ K + ++ I +K+MW+A ++++ ++++
Sbjct: 154 FFIFNAIALFTSLAVVVVQITLVRGETKAERRVVEIINKLMWLASVCTTVAFISSAYIVV 213
Query: 552 PHNQEMQWLSVVLLALGGGFLGTIFISLSVMLVEHWLRKSSWRKKRK---ESGDGTAESD 608
+ QW ++++ +GG + + +++ +V R S RKK K SG + + +
Sbjct: 214 --GKHFQWAALLVTLIGGVIMAGVLGTMTYYVVRS-KRTRSIRKKVKSTRRSGSNSWQQN 270
Query: 609 KESEDSDFQSSY 620
E DS+ Y
Sbjct: 271 SEFSDSEIDRIY 282
>dbj|BAA96936.1| ankyrin-like protein [Arabidopsis thaliana]
gi|67633894|gb|AAY78871.1| ankyrin repeat family protein
[Arabidopsis thaliana] gi|15238604|ref|NP_200815.1|
ankyrin repeat family protein [Arabidopsis thaliana]
Length = 548
Score = 102 bits (254), Expect = 4e-20
Identities = 64/201 (31%), Positives = 108/201 (52%), Gaps = 17/201 (8%)
Query: 412 VKQKHHHNKGKIENVNHTKR------KHYHEMHKEALLNARNTIVLVAVLIATVTFAAGI 465
+K + HH ++E+ T++ K ++MH E L NA N+ +VAVLIATV FAA
Sbjct: 330 IKHEVHH---QLEHARETRKRVQGIAKRINKMHVEGLDNAINSTTVVAVLIATVAFAAIF 386
Query: 466 SPPGGVYQE------GPKKGISMAGETSAFKVFAISNIIALFTSLSVVIVLVSIIPFRRK 519
+ PG E G G + + AF +F I + IALF SL+VV+V S++ K
Sbjct: 387 TVPGQYADELSSLLPGQSLGEANIADRPAFAIFFIFDSIALFISLAVVVVQTSVVAIEHK 446
Query: 520 PQTILLTIAHKVMWVAVAFMGTGYVAATWVILPHNQEMQWLSVVLLALGGGFLGTIFISL 579
+ ++ + +K+MW+A + ++A +V++ +E +WL+V + G + T ++
Sbjct: 447 AKKNMMAVINKLMWLACVLISVAFLALAFVVV--GEEERWLAVGVTVFGATIMLTTLGTM 504
Query: 580 SVMLVEHWLRKSSWRKKRKES 600
++ H + S+ RK RKES
Sbjct: 505 CYWVIMHRIEASNVRKSRKES 525
Score = 57.0 bits (136), Expect = 2e-06
Identities = 45/163 (27%), Positives = 78/163 (47%), Gaps = 16/163 (9%)
Query: 2 DQEFFDAIKKNDMITFSSIVK---EREGILN---QKTDDTFSAPLHLASKYGCIEMVSEI 55
D + A+++ D I+ E E L +K + L++A++YG ++V+E+
Sbjct: 33 DSQLLSAVRRGDFSAVKEILSNHMESEDELRDLLRKQNQCGETALYVAAEYGDADVVAEL 92
Query: 56 VKLCPDMVSAENK--NMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFLVACS 113
+K D+ AE K N P H A +Q + VL +L+E +P + ++ + +A A +
Sbjct: 93 IKYY-DLEDAETKARNGFDPFHIAAKQGELDVLRVLMEEHPELSMTVDLSNTTALHTAAA 151
Query: 114 HGHLDLVNLLLNLSEIVGQEVAGF----DQACFHVAAVRGHTE 152
GH+++V LL E G +A + H AA GH E
Sbjct: 152 QGHVEVVEYLL---EAAGSSLAAIAKSNGKTALHSAARNGHAE 191
Score = 46.2 bits (108), Expect = 0.003
Identities = 26/89 (29%), Positives = 51/89 (57%)
Query: 40 LHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACRQENVKVLMLLLEVNPTAACK 99
LH A++ G E+V IV + PD + +K +TP+H A + +++ V++ L++ + ++
Sbjct: 181 LHSAARNGHAEVVKAIVAVEPDTATRTDKKGQTPLHMAVKGQSIDVVVELMKGHRSSLNM 240
Query: 100 LNPTCKSAFLVACSHGHLDLVNLLLNLSE 128
+ +A VA G + +V LLL+ +E
Sbjct: 241 ADSKGNTALHVATRKGRIKIVELLLDNNE 269
Score = 39.7 bits (91), Expect = 0.30
Identities = 21/66 (31%), Positives = 37/66 (55%)
Query: 31 KTDDTFSAPLHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACRQENVKVLMLLL 90
+TD PLH+A K I++V E++K ++ + T +H A R+ +K++ LLL
Sbjct: 206 RTDKKGQTPLHMAVKGQSIDVVVELMKGHRSSLNMADSKGNTALHVATRKGRIKIVELLL 265
Query: 91 EVNPTA 96
+ N T+
Sbjct: 266 DNNETS 271
Score = 36.2 bits (82), Expect = 3.3
Identities = 25/116 (21%), Positives = 53/116 (45%), Gaps = 1/116 (0%)
Query: 6 FDAIKKNDMITFSSIVKEREGILNQKTDDTFSAPLHLASKYGCIEMVSEIVKLCPDMVSA 65
F K + ++ E L+ D + + LH A+ G +E+V +++ ++A
Sbjct: 112 FHIAAKQGELDVLRVLMEEHPELSMTVDLSNTTALHTAAAQGHVEVVEYLLEAAGSSLAA 171
Query: 66 ENK-NMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLDLV 120
K N +T +H A R + +V+ ++ V P A + + ++ +A +D+V
Sbjct: 172 IAKSNGKTALHSAARNGHAEVVKAIVAVEPDTATRTDKKGQTPLHMAVKGQSIDVV 227
>ref|NP_915384.1| P0506B12.26 [Oryza sativa (japonica cultivar-group)]
Length = 583
Score = 98.2 bits (243), Expect = 7e-19
Identities = 56/196 (28%), Positives = 109/196 (55%), Gaps = 10/196 (5%)
Query: 428 HTKRKHYHEMHKEALLNARNTIVLVAVLIATVTFAAGISPPGGVYQEGPKKGISMAGETS 487
H K ++H+E + NA N++ +VAVL ATV FAA + PGG G+++ + +
Sbjct: 393 HGIAKELRKLHREGINNATNSVTVVAVLFATVAFAAIFTVPGG----NANNGVAVVVQAA 448
Query: 488 AFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGYVAAT 547
+F++F I N IALFTSL+VV+V ++++ K + ++ + +K+MW+A ++A+
Sbjct: 449 SFRIFFIFNAIALFTSLAVVVVQITVVRGETKSERKVVEVINKLMWLASVCTTISFIASC 508
Query: 548 WVILPHNQEMQWLSVVLLALGGGFLGTIFISLSVMLVEHWLRKSSWRKKRK---ESGDGT 604
+++L + QW ++++ +GG + + +++ +V+ R RKK K SG +
Sbjct: 509 YIVL--GRHFQWAALLVSLIGGITMAGVLGTMTYYVVKS-KRMRKIRKKEKMSRRSGSSS 565
Query: 605 AESDKESEDSDFQSSY 620
+ E +++ Y
Sbjct: 566 WYDNTELSETELNQVY 581
Score = 48.9 bits (115), Expect = 5e-04
Identities = 36/116 (31%), Positives = 61/116 (52%), Gaps = 2/116 (1%)
Query: 39 PLHLASKYGCIEMVSEIVK-LCPDMVSAENKNMETPIHEACRQENVKVLMLLLEVNPTAA 97
PL A++ G +E+V E+++ L + V+A+N++ +H A R+ V+ +L N A
Sbjct: 123 PLVAAAERGHLEVVRELLRHLDAEGVAAKNRSGYDALHVAAREGRHAVVQEMLLHNRLLA 182
Query: 98 CKLNPTCKSAFLVACSHGHLDLVNLLLNLSEIVGQEVAGFD-QACFHVAAVRGHTE 152
P S + A + GH ++V LLL L + E+A + + H AA +GH E
Sbjct: 183 KTFGPANTSPLISAATRGHTEVVKLLLELDDFGLVEMAKDNGKNSLHFAARQGHVE 238
Score = 37.7 bits (86), Expect = 1.1
Identities = 23/89 (25%), Positives = 45/89 (49%)
Query: 23 EREGILNQKTDDTFSAPLHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACRQEN 82
E++ L ++ D LH+A K +++ +V P +V +KN T +H A R++
Sbjct: 245 EKDPQLARRNDKKGQTALHMAVKGTNCDVLRALVDADPAIVMLPDKNGNTALHVATRKKR 304
Query: 83 VKVLMLLLEVNPTAACKLNPTCKSAFLVA 111
+++ +LL + T L K+A+ +A
Sbjct: 305 AEIVAVLLRLPDTHVNALTRDHKTAYDIA 333
>dbj|BAD73402.1| ankyrin-like protein [Oryza sativa (japonica cultivar-group)]
Length = 556
Score = 98.2 bits (243), Expect = 7e-19
Identities = 56/196 (28%), Positives = 109/196 (55%), Gaps = 10/196 (5%)
Query: 428 HTKRKHYHEMHKEALLNARNTIVLVAVLIATVTFAAGISPPGGVYQEGPKKGISMAGETS 487
H K ++H+E + NA N++ +VAVL ATV FAA + PGG G+++ + +
Sbjct: 366 HGIAKELRKLHREGINNATNSVTVVAVLFATVAFAAIFTVPGG----NANNGVAVVVQAA 421
Query: 488 AFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGYVAAT 547
+F++F I N IALFTSL+VV+V ++++ K + ++ + +K+MW+A ++A+
Sbjct: 422 SFRIFFIFNAIALFTSLAVVVVQITVVRGETKSERKVVEVINKLMWLASVCTTISFIASC 481
Query: 548 WVILPHNQEMQWLSVVLLALGGGFLGTIFISLSVMLVEHWLRKSSWRKKRK---ESGDGT 604
+++L + QW ++++ +GG + + +++ +V+ R RKK K SG +
Sbjct: 482 YIVL--GRHFQWAALLVSLIGGITMAGVLGTMTYYVVKS-KRMRKIRKKEKMSRRSGSSS 538
Query: 605 AESDKESEDSDFQSSY 620
+ E +++ Y
Sbjct: 539 WYDNTELSETELNQVY 554
Score = 48.9 bits (115), Expect = 5e-04
Identities = 36/116 (31%), Positives = 61/116 (52%), Gaps = 2/116 (1%)
Query: 39 PLHLASKYGCIEMVSEIVK-LCPDMVSAENKNMETPIHEACRQENVKVLMLLLEVNPTAA 97
PL A++ G +E+V E+++ L + V+A+N++ +H A R+ V+ +L N A
Sbjct: 96 PLVAAAERGHLEVVRELLRHLDAEGVAAKNRSGYDALHVAAREGRHAVVQEMLLHNRLLA 155
Query: 98 CKLNPTCKSAFLVACSHGHLDLVNLLLNLSEIVGQEVAGFD-QACFHVAAVRGHTE 152
P S + A + GH ++V LLL L + E+A + + H AA +GH E
Sbjct: 156 KTFGPANTSPLISAATRGHTEVVKLLLELDDFGLVEMAKDNGKNSLHFAARQGHVE 211
Score = 37.7 bits (86), Expect = 1.1
Identities = 23/89 (25%), Positives = 45/89 (49%)
Query: 23 EREGILNQKTDDTFSAPLHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACRQEN 82
E++ L ++ D LH+A K +++ +V P +V +KN T +H A R++
Sbjct: 218 EKDPQLARRNDKKGQTALHMAVKGTNCDVLRALVDADPAIVMLPDKNGNTALHVATRKKR 277
Query: 83 VKVLMLLLEVNPTAACKLNPTCKSAFLVA 111
+++ +LL + T L K+A+ +A
Sbjct: 278 AEIVAVLLRLPDTHVNALTRDHKTAYDIA 306
>gb|AAF75083.1| It contains Ank repeat PF|00023. EST gb|AI996003 comes from this
gene. [Arabidopsis thaliana]
gi|15222993|ref|NP_172250.1| ankyrin repeat family
protein [Arabidopsis thaliana] gi|25406998|pir||C86212
hypothetical protein [imported] - Arabidopsis thaliana
Length = 543
Score = 97.4 bits (241), Expect = 1e-18
Identities = 63/217 (29%), Positives = 114/217 (52%), Gaps = 18/217 (8%)
Query: 400 TYNPYYFSPTNLVKQKHHHNKGKIEN-VNHTK---------RKHYHEMHKEALLNARNTI 449
T P +P +KQ K ++ N + HT+ K ++MH E L NA N+
Sbjct: 300 TIKPSGPNPARELKQTVSDIKHEVHNQLEHTRLTRKRVQGIAKQLNKMHTEGLNNAINST 359
Query: 450 VLVAVLIATVTFAAGISPPGGVYQE------GPKKGISMAGETSAFKVFAISNIIALFTS 503
+VAVLIATV FAA + PG ++ G G + T+ F +F I + IALF S
Sbjct: 360 TVVAVLIATVAFAAIFTVPGQYVEDTSKIPDGHSLGEANIASTTPFIIFFIFDSIALFIS 419
Query: 504 LSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGYVAATWVILPHNQEMQWLSVV 563
L+VV+V S++ K + ++ + +K+MW+A + ++A ++V++ +E +WL++
Sbjct: 420 LAVVVVQTSVVVIESKAKKQMMAVINKLMWLACVLISVAFLALSFVVV--GEEEKWLAIW 477
Query: 564 LLALGGGFLGTIFISLSVMLVEHWLRKSSWRKKRKES 600
+ A+G + T ++ +++H + ++ R R+ S
Sbjct: 478 VTAIGATIMITTLGTMCYWIIQHKIEAANLRNIRRSS 514
Score = 57.8 bits (138), Expect = 1e-06
Identities = 40/138 (28%), Positives = 68/138 (48%), Gaps = 6/138 (4%)
Query: 20 IVKEREGILNQ---KTDDTFSAPLHLASKYGCIEMVSEIVKLCPDMVSAENK--NMETPI 74
+ K RE LNQ K + + L++A++YG +E+V E++ C D+ E K N
Sbjct: 47 LTKTRESELNQLLGKQNQSGETALYVAAEYGDVEIVKEMIN-CYDLALVEIKARNGFDAF 105
Query: 75 HEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLDLVNLLLNLSEIVGQEV 134
H A +Q ++ VL +L E + A ++ + +A A + GH ++VN LL L +
Sbjct: 106 HIAAKQGDLDVLKVLAEAHSELAMTVDLSNTTALHTAATQGHTEVVNFLLELGSSLAGIA 165
Query: 135 AGFDQACFHVAAVRGHTE 152
+ H A+ GH +
Sbjct: 166 KSNGKTALHSASRNGHVK 183
Score = 47.4 bits (111), Expect = 0.001
Identities = 34/148 (22%), Positives = 66/148 (43%), Gaps = 1/148 (0%)
Query: 5 FFDAIKKNDMITFSSIVKEREGILNQKTDDTFSAPLHLASKYGCIEMVSEIVKLCPDMVS 64
F A K+ D+ ++ E L D + + LH A+ G E+V+ +++L +
Sbjct: 105 FHIAAKQGDLDVLK-VLAEAHSELAMTVDLSNTTALHTAATQGHTEVVNFLLELGSSLAG 163
Query: 65 AENKNMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLDLVNLLL 124
N +T +H A R +VKV+ LL P A +++ ++A +A ++++V L+
Sbjct: 164 IAKSNGKTALHSASRNGHVKVIKALLASEPAIAIRMDKKGQTALHMAVKGTNVEVVEELI 223
Query: 125 NLSEIVGQEVAGFDQACFHVAAVRGHTE 152
H+AA +G ++
Sbjct: 224 KADRSSINIADTKGNTALHIAARKGRSQ 251
>ref|XP_483595.1| ankyrin-like protein [Oryza sativa (japonica cultivar-group)]
gi|42407837|dbj|BAD08980.1| ankyrin-like protein [Oryza
sativa (japonica cultivar-group)]
Length = 528
Score = 96.7 bits (239), Expect = 2e-18
Identities = 64/224 (28%), Positives = 114/224 (50%), Gaps = 17/224 (7%)
Query: 412 VKQKHHHNKGKIENVNHTK------RKHYHEMHKEALLNARNTIVLVAVLIATVTFAAGI 465
V H + +I+ TK +K ++H L NA N+ +VAVLIATV FAA
Sbjct: 305 VSDIRHDVQSQIKQTKQTKMQVQKIKKRLEKLHIGGLNNAINSNTVVAVLIATVAFAAIF 364
Query: 466 SPPGGVYQE------GPKKGISMAGETSAFKVFAISNIIALFTSLSVVIVLVSIIPFRRK 519
+ PG ++ G G + AF VF + + +ALF SL+VV+V S+I +K
Sbjct: 365 TVPGNFVEDITQAPPGMSLGQAYVASNPAFLVFLVFDALALFISLAVVVVQTSLIVVEQK 424
Query: 520 PQTILLTIAHKVMWVAVAFMGTGYVAATWVILPHNQEMQWLSVVLLALGGGFLGTIFISL 579
+ ++ + +K+MW+A F+ ++A T+V++ ++ WL+ +A+G + T S+
Sbjct: 425 AKRRMVFVMNKLMWLACLFISVAFIALTYVVV--GRDDWWLAWCTMAIGAVIMLTTLGSM 482
Query: 580 SVMLVEHWLRKSSWRK---KRKESGDGTAESDKESEDSDFQSSY 620
++ H + + RK + S T +SD + +S+++ Y
Sbjct: 483 CYCIIAHRMDERKIRKASTSQSRSWSQTVDSDPDLLNSEYKKMY 526
Score = 49.7 bits (117), Expect = 3e-04
Identities = 34/115 (29%), Positives = 54/115 (46%), Gaps = 1/115 (0%)
Query: 39 PLHLASKYGCIEMVSEIVKLCP-DMVSAENKNMETPIHEACRQENVKVLMLLLEVNPTAA 97
PL++A++ G ++V EI+K+ + N H A +Q +++VL LL+ P A
Sbjct: 59 PLYVAAERGHTDVVREILKVSDVQTAGVKANNSFDAFHIAAKQGHLEVLKELLQAFPALA 118
Query: 98 CKLNPTCKSAFLVACSHGHLDLVNLLLNLSEIVGQEVAGFDQACFHVAAVRGHTE 152
N +A A GH ++VNLLL + + + H AA GH E
Sbjct: 119 MTTNSVNATALDTAAILGHTEIVNLLLESDANLARIARNNGKTVLHSAARLGHVE 173
Score = 46.6 bits (109), Expect = 0.002
Identities = 29/110 (26%), Positives = 53/110 (47%)
Query: 24 REGILNQKTDDTFSAPLHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACRQENV 83
R+ + +TD LH+ASK E+V E++K ++ E+ P+H A R+ N+
Sbjct: 181 RDPGIGLRTDKKGQTALHMASKGQNAEIVIELLKPDISVIHLEDNKGNRPLHVATRKANI 240
Query: 84 KVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLDLVNLLLNLSEIVGQE 133
++ LL V +N + +A +A + +LVN+L + +E
Sbjct: 241 VIVQTLLSVEGIEVNAVNRSGHTALAIAEQLNNEELVNILREAGGVTAKE 290
Score = 43.5 bits (101), Expect = 0.020
Identities = 29/116 (25%), Positives = 55/116 (47%), Gaps = 1/116 (0%)
Query: 31 KTDDTFSAPLHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACRQENVKVLMLLL 90
K +++F A H+A+K G +E++ E+++ P + N T + A + +++ LLL
Sbjct: 87 KANNSFDA-FHIAAKQGHLEVLKELLQAFPALAMTTNSVNATALDTAAILGHTEIVNLLL 145
Query: 91 EVNPTAACKLNPTCKSAFLVACSHGHLDLVNLLLNLSEIVGQEVAGFDQACFHVAA 146
E + A K+ A GH+++V LL+ +G Q H+A+
Sbjct: 146 ESDANLARIARNNGKTVLHSAARLGHVEIVRSLLSRDPGIGLRTDKKGQTALHMAS 201
Score = 40.4 bits (93), Expect = 0.17
Identities = 28/91 (30%), Positives = 42/91 (45%), Gaps = 6/91 (6%)
Query: 68 KNMETPIHEACRQENVKVLMLLL-----EVNPTAACKLNPTCKSAFLVACSHGHLDLVNL 122
K +TP+H A R N ++ EV A + N ++ VA GH D+V
Sbjct: 15 KRGDTPLHLAARSGNAAGAQRIIAEFDPEVAAERAAQANHDGETPLYVAAERGHTDVVRE 74
Query: 123 LLNLSEIVGQEV-AGFDQACFHVAAVRGHTE 152
+L +S++ V A FH+AA +GH E
Sbjct: 75 ILKVSDVQTAGVKANNSFDAFHIAAKQGHLE 105
>ref|NP_172055.1| ankyrin repeat family protein [Arabidopsis thaliana]
gi|25406852|pir||E86190 hypothetical protein [imported]
- Arabidopsis thaliana gi|4836914|gb|AAD30616.1|
Hypothetical protein [Arabidopsis thaliana]
Length = 627
Score = 95.9 bits (237), Expect = 3e-18
Identities = 58/173 (33%), Positives = 99/173 (56%), Gaps = 8/173 (4%)
Query: 432 KHYHEMHKEALLNARNTIVLVAVLIATVTFAAGISPPGGVYQEGPKKGISMAGET----- 486
K ++H L NA N+ +VAVLIATV FAA + PG Y+E KG+ + GE
Sbjct: 428 KRLKKLHINGLNNAINSATVVAVLIATVAFAAIFTIPGQ-YEEDRTKGLLLLGEARIAGK 486
Query: 487 SAFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGYVAA 546
+ F VF I + +ALF SL+VV+V S++ +K + L+ + +K+MW+A F+ +V+
Sbjct: 487 APFLVFFIFDSLALFISLAVVVVQTSVVVIEQKAKKNLVFVINKLMWLACLFISVAFVSL 546
Query: 547 TWVILPHNQEMQWLSVVLLALGGGFLGTIFISLSVMLVEHWLRKSSWRKKRKE 599
+++++ +E WL++ +GG + T ++ +V H + +S + RKE
Sbjct: 547 SFIVV--GKEDIWLAICATIIGGTIMLTTIGAMCYCVVMHRIEESKLKSLRKE 597
Score = 62.4 bits (150), Expect = 4e-08
Identities = 40/126 (31%), Positives = 63/126 (49%), Gaps = 1/126 (0%)
Query: 28 LNQKTDDTFSAPLHLASKYGCIEMVSEIVK-LCPDMVSAENKNMETPIHEACRQENVKVL 86
L+ K + PL+ A++ G +V E++K + D S + +N P H A +Q +++ L
Sbjct: 145 LSSKQNLEGETPLYSAAENGHSLVVEEMLKHMDLDTASVKARNGFDPFHVAAKQGHIEAL 204
Query: 87 MLLLEVNPTAACKLNPTCKSAFLVACSHGHLDLVNLLLNLSEIVGQEVAGFDQACFHVAA 146
LLE P A ++ +C +A A S GH D+VNLLL + + + H AA
Sbjct: 205 KKLLETFPNLAMTVDLSCTTALHTAASQGHTDVVNLLLKTDSHLAKIAKNNGKTALHSAA 264
Query: 147 VRGHTE 152
GH E
Sbjct: 265 RMGHRE 270
Score = 47.8 bits (112), Expect = 0.001
Identities = 31/114 (27%), Positives = 54/114 (47%), Gaps = 1/114 (0%)
Query: 39 PLHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACRQENVKVLMLLLEVNPTAAC 98
P H+A+K G IE + ++++ P++ + + T +H A Q + V+ LLL+ + A
Sbjct: 191 PFHVAAKQGHIEALKKLLETFPNLAMTVDLSCTTALHTAASQGHTDVVNLLLKTDSHLAK 250
Query: 99 KLNPTCKSAFLVACSHGHLDLVNLLLNLSEIVGQEVAGFDQACFHVAAVRGHTE 152
K+A A GH ++V L+ +G Q H+ AV+G E
Sbjct: 251 IAKNNGKTALHSAARMGHREVVKSLIGNDASIGFRTDKKGQTALHM-AVKGQNE 303
Score = 40.8 bits (94), Expect = 0.13
Identities = 26/93 (27%), Positives = 47/93 (49%)
Query: 31 KTDDTFSAPLHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACRQENVKVLMLLL 90
+TD LH+A K +V E+VK P ++S E+ TP+H A + +K++ L+
Sbjct: 285 RTDKKGQTALHMAVKGQNEGIVLELVKPDPAILSVEDSKGNTPLHTATNKGRIKIVRCLV 344
Query: 91 EVNPTAACKLNPTCKSAFLVACSHGHLDLVNLL 123
+ +N +A +A G+ +LV++L
Sbjct: 345 SFDGINLNAMNKAGDTALDIAEKIGNPELVSVL 377
>gb|AAM62711.1| ankyrin-like protein [Arabidopsis thaliana]
Length = 534
Score = 95.1 bits (235), Expect = 6e-18
Identities = 56/188 (29%), Positives = 105/188 (55%), Gaps = 7/188 (3%)
Query: 428 HTKRKHYHEMHKEALLNARNTIVLVAVLIATVTFAAGISPPGGVYQEGPKKGISMAGETS 487
H K ++H+E + NA N++ +VAVL ATV FAA + PGG +G + A
Sbjct: 345 HNISKELRKLHREGINNATNSVTVVAVLFATVAFAAIFTVPGGDNNDGSAVVVGRA---- 400
Query: 488 AFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGYVAAT 547
+FK+F I N +ALFTSL+VV+V ++++ K + ++ + +K+MW+A ++A++
Sbjct: 401 SFKIFFIFNALALFTSLAVVVVQITLVRGETKAEKRVVEVINKLMWLASMCTSVAFLASS 460
Query: 548 WVILPHNQEMQWLSVVLLALGGGFLGTIFISLSVMLVEHWLRKSSWRKKRKESGDGTAES 607
++++ E W + ++ +GG + + +++ +V+ R S RKK K + + S
Sbjct: 461 YIVVGRKNE--WAAELVTVVGGVIMAGVLGTMTYYVVKS-KRTRSMRKKVKSARRSGSNS 517
Query: 608 DKESEDSD 615
S+ S+
Sbjct: 518 WHHSDFSN 525
Score = 51.6 bits (122), Expect = 8e-05
Identities = 25/89 (28%), Positives = 51/89 (57%)
Query: 40 LHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACRQENVKVLMLLLEVNPTAACK 99
LHLA++ G +E++ ++ P + +K +T +H A + ++ +V+ LLL+ +P +
Sbjct: 180 LHLAARQGHVEVIKALLSKDPQLARRIDKKGQTALHMAVKGQSSEVVKLLLDADPAIVMQ 239
Query: 100 LNPTCKSAFLVACSHGHLDLVNLLLNLSE 128
+ +C +A VA ++V LLL+L +
Sbjct: 240 PDKSCNTALHVATRKKRAEIVELLLSLPD 268
Score = 44.7 bits (104), Expect = 0.009
Identities = 29/130 (22%), Positives = 61/130 (46%)
Query: 23 EREGILNQKTDDTFSAPLHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACRQEN 82
+ + L+Q + + PL A+ G E+V++++ +++ N + +H A RQ +
Sbjct: 129 DHDATLSQTFGPSNATPLVSAAMRGHTEVVNQLLSKAGNLLEISRSNNKNALHLAARQGH 188
Query: 83 VKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLDLVNLLLNLSEIVGQEVAGFDQACF 142
V+V+ LL +P A +++ ++A +A ++V LLL+ + +
Sbjct: 189 VEVIKALLSKDPQLARRIDKKGQTALHMAVKGQSSEVVKLLLDADPAIVMQPDKSCNTAL 248
Query: 143 HVAAVRGHTE 152
HVA + E
Sbjct: 249 HVATRKKRAE 258
Score = 44.7 bits (104), Expect = 0.009
Identities = 30/136 (22%), Positives = 70/136 (51%), Gaps = 2/136 (1%)
Query: 18 SSIVKEREGILNQKTDDTFSAPLHLASKYGCIEMVSEIVKLCP-DMVSAENKNMETPIHE 76
+ + + R I+N+ ++ L A+ G +++V E++K + ++ +N++ P+H
Sbjct: 56 AEVAEIRASIVNE-VNELGETALFTAADKGHLDVVKELLKYSSRESIAKKNRSGYDPLHI 114
Query: 77 ACRQENVKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLDLVNLLLNLSEIVGQEVAG 136
A Q + ++ + L+ + T + P+ + + A GH ++VN LL+ + + +
Sbjct: 115 AAIQGHHAIVEVSLDHDATLSQTFGPSNATPLVSAAMRGHTEVVNQLLSKAGNLLEISRS 174
Query: 137 FDQACFHVAAVRGHTE 152
++ H+AA +GH E
Sbjct: 175 NNKNALHLAARQGHVE 190
>dbj|BAB03143.1| ankyrin-like protein [Arabidopsis thaliana]
Length = 1100
Score = 95.1 bits (235), Expect = 6e-18
Identities = 56/188 (29%), Positives = 105/188 (55%), Gaps = 7/188 (3%)
Query: 428 HTKRKHYHEMHKEALLNARNTIVLVAVLIATVTFAAGISPPGGVYQEGPKKGISMAGETS 487
H K ++H+E + NA N++ +VAVL ATV FAA + PGG +G + A
Sbjct: 911 HNISKELRKLHREGINNATNSVTVVAVLFATVAFAAIFTVPGGDNNDGSAVVVGRA---- 966
Query: 488 AFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGYVAAT 547
+FK+F I N +ALFTSL+VV+V ++++ K + ++ + +K+MW+A ++A++
Sbjct: 967 SFKIFFIFNALALFTSLAVVVVQITLVRGETKAEKRVVEVINKLMWLASMCTSVAFLASS 1026
Query: 548 WVILPHNQEMQWLSVVLLALGGGFLGTIFISLSVMLVEHWLRKSSWRKKRKESGDGTAES 607
++++ E W + ++ +GG + + +++ +V+ R S RKK K + + S
Sbjct: 1027 YIVVGRKNE--WAAELVTVVGGVIMAGVLGTMTYYVVKS-KRTRSMRKKVKSARRSGSNS 1083
Query: 608 DKESEDSD 615
S+ S+
Sbjct: 1084 WHHSDFSN 1091
Score = 48.5 bits (114), Expect = 6e-04
Identities = 31/137 (22%), Positives = 65/137 (46%)
Query: 20 IVKEREGILNQKTDDTFSAPLHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACR 79
++ + + L+Q + + PL A+ G E+V++++ +++ N + +H A R
Sbjct: 667 VLLDHDATLSQTFGPSNATPLVSAAMRGHTEVVNQLLSKAGNLLEISRSNNKNALHLAAR 726
Query: 80 QENVKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLDLVNLLLNLSEIVGQEVAGFDQ 139
Q +V+V+ LL +P A +++ ++A +A ++V LLL+ + +
Sbjct: 727 QGHVEVIKALLSKDPQLARRIDKKGQTALHMAVKGQSSEVVKLLLDADPAIVMQPDKSCN 786
Query: 140 ACFHVAAVRGHTESC*T 156
HVA + E C T
Sbjct: 787 TALHVATRKKRAEVCIT 803
Score = 47.0 bits (110), Expect = 0.002
Identities = 31/136 (22%), Positives = 71/136 (51%), Gaps = 2/136 (1%)
Query: 18 SSIVKEREGILNQKTDDTFSAPLHLASKYGCIEMVSEIVKLCP-DMVSAENKNMETPIHE 76
+ + + R I+N+ ++ L A+ G +++V E++K + ++ +N++ P+H
Sbjct: 597 AEVAEIRASIVNE-VNELGETALFTAADKGHLDVVKELLKYSSRESIAKKNRSGYDPLHI 655
Query: 77 ACRQENVKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLDLVNLLLNLSEIVGQEVAG 136
A Q + ++ +LL+ + T + P+ + + A GH ++VN LL+ + + +
Sbjct: 656 AAIQGHHAIVEVLLDHDATLSQTFGPSNATPLVSAAMRGHTEVVNQLLSKAGNLLEISRS 715
Query: 137 FDQACFHVAAVRGHTE 152
++ H+AA +GH E
Sbjct: 716 NNKNALHLAARQGHVE 731
>gb|AAG51002.1| ankyrin-like protein; 93648-91299 [Arabidopsis thaliana]
gi|15230470|ref|NP_187842.1| ankyrin repeat family
protein [Arabidopsis thaliana]
Length = 590
Score = 95.1 bits (235), Expect = 6e-18
Identities = 56/188 (29%), Positives = 105/188 (55%), Gaps = 7/188 (3%)
Query: 428 HTKRKHYHEMHKEALLNARNTIVLVAVLIATVTFAAGISPPGGVYQEGPKKGISMAGETS 487
H K ++H+E + NA N++ +VAVL ATV FAA + PGG +G + A
Sbjct: 401 HNISKELRKLHREGINNATNSVTVVAVLFATVAFAAIFTVPGGDNNDGSAVVVGRA---- 456
Query: 488 AFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGYVAAT 547
+FK+F I N +ALFTSL+VV+V ++++ K + ++ + +K+MW+A ++A++
Sbjct: 457 SFKIFFIFNALALFTSLAVVVVQITLVRGETKAEKRVVEVINKLMWLASMCTSVAFLASS 516
Query: 548 WVILPHNQEMQWLSVVLLALGGGFLGTIFISLSVMLVEHWLRKSSWRKKRKESGDGTAES 607
++++ E W + ++ +GG + + +++ +V+ R S RKK K + + S
Sbjct: 517 YIVVGRKNE--WAAELVTVVGGVIMAGVLGTMTYYVVKS-KRTRSMRKKVKSARRSGSNS 573
Query: 608 DKESEDSD 615
S+ S+
Sbjct: 574 WHHSDFSN 581
Score = 51.6 bits (122), Expect = 8e-05
Identities = 25/89 (28%), Positives = 51/89 (57%)
Query: 40 LHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACRQENVKVLMLLLEVNPTAACK 99
LHLA++ G +E++ ++ P + +K +T +H A + ++ +V+ LLL+ +P +
Sbjct: 236 LHLAARQGHVEVIKALLSKDPQLARRIDKKGQTALHMAVKGQSSEVVKLLLDADPAIVMQ 295
Query: 100 LNPTCKSAFLVACSHGHLDLVNLLLNLSE 128
+ +C +A VA ++V LLL+L +
Sbjct: 296 PDKSCNTALHVATRKKRAEIVELLLSLPD 324
Score = 47.0 bits (110), Expect = 0.002
Identities = 31/136 (22%), Positives = 71/136 (51%), Gaps = 2/136 (1%)
Query: 18 SSIVKEREGILNQKTDDTFSAPLHLASKYGCIEMVSEIVKLCP-DMVSAENKNMETPIHE 76
+ + + R I+N+ ++ L A+ G +++V E++K + ++ +N++ P+H
Sbjct: 112 AEVAEIRASIVNE-VNELGETALFTAADKGHLDVVKELLKYSSRESIAKKNRSGYDPLHI 170
Query: 77 ACRQENVKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLDLVNLLLNLSEIVGQEVAG 136
A Q + ++ +LL+ + T + P+ + + A GH ++VN LL+ + + +
Sbjct: 171 AAIQGHHAIVEVLLDHDATLSQTFGPSNATPLVSAAMRGHTEVVNQLLSKAGNLLEISRS 230
Query: 137 FDQACFHVAAVRGHTE 152
++ H+AA +GH E
Sbjct: 231 NNKNALHLAARQGHVE 246
Score = 45.4 bits (106), Expect = 0.005
Identities = 29/133 (21%), Positives = 63/133 (46%)
Query: 20 IVKEREGILNQKTDDTFSAPLHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACR 79
++ + + L+Q + + PL A+ G E+V++++ +++ N + +H A R
Sbjct: 182 VLLDHDATLSQTFGPSNATPLVSAAMRGHTEVVNQLLSKAGNLLEISRSNNKNALHLAAR 241
Query: 80 QENVKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLDLVNLLLNLSEIVGQEVAGFDQ 139
Q +V+V+ LL +P A +++ ++A +A ++V LLL+ + +
Sbjct: 242 QGHVEVIKALLSKDPQLARRIDKKGQTALHMAVKGQSSEVVKLLLDADPAIVMQPDKSCN 301
Query: 140 ACFHVAAVRGHTE 152
HVA + E
Sbjct: 302 TALHVATRKKRAE 314
>gb|AAN64473.1| putative ankyrin repeat containing protein [Oryza sativa (japonica
cultivar-group)]
Length = 565
Score = 94.4 bits (233), Expect = 1e-17
Identities = 78/311 (25%), Positives = 141/311 (45%), Gaps = 38/311 (12%)
Query: 339 LDNTSSPSPTSRHSLSRRYISKEMEVLTEMVSYDCI----SPPPVSESTESISPQPQVSE 394
L+ S T+ H +R+ + ++ L E+ D S ++ E + V+
Sbjct: 250 LNLADSKGNTALHIAARKARTPIVKRLLELPDTDLKAINRSRETAFDTAEKMGNTESVAV 309
Query: 395 RFENGTYNPYYFSPTN------------LVKQKHHHNKGKIENVNHTK------RKHYHE 436
E+G + SPT V H ++E T+ K ++
Sbjct: 310 LAEHGVPSARAMSPTGGGGGNPGRELKQQVSDIKHEVHSQLEQTRQTRVRMQGIAKQINK 369
Query: 437 MHKEALLNARNTIVLVAVLIATVTFAAGISPPG------GVYQEGPKKGISMAGETSAFK 490
+H E L NA N+ +VAVLIATV FAA + PG G G G + +AF
Sbjct: 370 LHDEGLNNAINSTTVVAVLIATVAFAAIFTVPGEYVDDAGSLTPGQALGEANISHQTAFL 429
Query: 491 VFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGYVAATWVI 550
+F + + +ALF SL+VV+V S++ RK + ++ + +K+MWVA + ++A ++V+
Sbjct: 430 IFFVFDSVALFISLAVVVVQTSVVVIERKAKKQMMAVINKLMWVACVLVSVAFLALSFVV 489
Query: 551 LPHNQEMQWLSVVLLALGGGFLGTIFISLSVMLVEH--------WLRKSSWRKKRKESGD 602
+ + +WL+V + +G L T ++ ++ H +++SS + R S
Sbjct: 490 V--GKAERWLAVGVTIMGATILVTTIGTMLYWVIAHRIEAKRMRSIKRSSLSRSRSFSAS 547
Query: 603 GTAESDKESED 613
G +E++ E+
Sbjct: 548 GMSEAEWVEEE 558
Score = 57.8 bits (138), Expect = 1e-06
Identities = 31/107 (28%), Positives = 58/107 (53%)
Query: 40 LHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACRQENVKVLMLLLEVNPTAACK 99
LH+A+K G +E+V+E++K P++ + + T ++ A Q +++V+ LLLE + + A
Sbjct: 125 LHIAAKQGDVEVVNELLKALPELSMTVDASNTTALNTAATQGHMEVVRLLLEADASLAVI 184
Query: 100 LNPTCKSAFLVACSHGHLDLVNLLLNLSEIVGQEVAGFDQACFHVAA 146
K+A A +GH+++V L+ + V Q H+AA
Sbjct: 185 ARSNGKTALHSAARNGHVEVVRALMEAEPSIAARVDKKGQTALHMAA 231
Score = 52.4 bits (124), Expect = 4e-05
Identities = 36/139 (25%), Positives = 68/139 (48%), Gaps = 3/139 (2%)
Query: 16 TFSSIVKEREGILNQKTDDTFSAPLHLASKYGCIEMVSEIVKLCPDMVSA--ENKNMETP 73
T S + L K + PL +A++YG + +V+E++K D+ +A + ++
Sbjct: 66 TLSGAPPDELRALLSKQNQAGETPLFVAAEYGYVALVAEMIKY-HDVATACIKARSGYDA 124
Query: 74 IHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLDLVNLLLNLSEIVGQE 133
+H A +Q +V+V+ LL+ P + ++ + +A A + GH+++V LLL +
Sbjct: 125 LHIAAKQGDVEVVNELLKALPELSMTVDASNTTALNTAATQGHMEVVRLLLEADASLAVI 184
Query: 134 VAGFDQACFHVAAVRGHTE 152
+ H AA GH E
Sbjct: 185 ARSNGKTALHSAARNGHVE 203
Score = 46.6 bits (109), Expect = 0.002
Identities = 33/144 (22%), Positives = 67/144 (45%), Gaps = 1/144 (0%)
Query: 8 AIKKNDMITFSSIVKEREGILNQKTDDTFSAPLHLASKYGCIEMVSEIVKLCPDMVSAEN 67
A K+ D+ + ++K L+ D + + L+ A+ G +E+V +++ +
Sbjct: 128 AAKQGDVEVVNELLKALPE-LSMTVDASNTTALNTAATQGHMEVVRLLLEADASLAVIAR 186
Query: 68 KNMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLDLVNLLLNLS 127
N +T +H A R +V+V+ L+E P+ A +++ ++A +A LD+V+ LL
Sbjct: 187 SNGKTALHSAARNGHVEVVRALMEAEPSIAARVDKKGQTALHMAAKGTRLDIVDALLAGE 246
Query: 128 EIVGQEVAGFDQACFHVAAVRGHT 151
+ H+AA + T
Sbjct: 247 PTLLNLADSKGNTALHIAARKART 270
Score = 37.0 bits (84), Expect = 1.9
Identities = 25/113 (22%), Positives = 53/113 (46%)
Query: 11 KNDMITFSSIVKEREGILNQKTDDTFSAPLHLASKYGCIEMVSEIVKLCPDMVSAENKNM 70
+N + + E E + + D LH+A+K +++V ++ P +++ +
Sbjct: 198 RNGHVEVVRALMEAEPSIAARVDKKGQTALHMAAKGTRLDIVDALLAGEPTLLNLADSKG 257
Query: 71 ETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLDLVNLL 123
T +H A R+ ++ LLE+ T +N + ++AF A G+ + V +L
Sbjct: 258 NTALHIAARKARTPIVKRLLELPDTDLKAINRSRETAFDTAEKMGNTESVAVL 310
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.330 0.140 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 985,497,008
Number of Sequences: 2540612
Number of extensions: 38997405
Number of successful extensions: 166850
Number of sequences better than 10.0: 1259
Number of HSP's better than 10.0 without gapping: 220
Number of HSP's successfully gapped in prelim test: 1058
Number of HSP's that attempted gapping in prelim test: 160933
Number of HSP's gapped (non-prelim): 5186
length of query: 626
length of database: 863,360,394
effective HSP length: 134
effective length of query: 492
effective length of database: 522,918,386
effective search space: 257275845912
effective search space used: 257275845912
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 78 (34.7 bits)
Medicago: description of AC138130.17


