

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138130.15 + phase: 0
(174 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
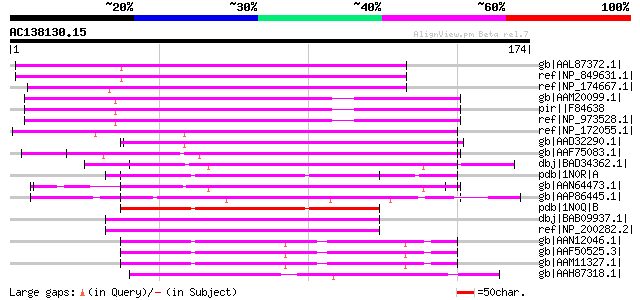
Score E
Sequences producing significant alignments: (bits) Value
gb|AAL87372.1| At1g10340/F14N23_22 [Arabidopsis thaliana] gi|183... 70 2e-11
ref|NP_849631.1| ankyrin repeat family protein [Arabidopsis thal... 70 2e-11
ref|NP_174667.1| ankyrin repeat family protein [Arabidopsis thal... 67 3e-10
gb|AAM20099.1| unknown protein [Arabidopsis thaliana] gi|1581033... 65 8e-10
pir||F84638 hypothetical protein At2g24600 [imported] - Arabidop... 65 8e-10
ref|NP_973528.1| ankyrin repeat family protein [Arabidopsis thal... 65 8e-10
ref|NP_172055.1| ankyrin repeat family protein [Arabidopsis thal... 61 1e-08
gb|AAD32290.1| ankyrin-like protein [Arabidopsis thaliana] gi|25... 59 6e-08
gb|AAF75083.1| It contains Ank repeat PF|00023. EST gb|AI996003... 58 1e-07
dbj|BAD34362.1| ankyrin-like protein [Oryza sativa (japonica cul... 58 1e-07
pdb|1N0R|A Chain A, 4ank: A Designed Ankyrin Repeat Protein With... 58 1e-07
gb|AAN64473.1| putative ankyrin repeat containing protein [Oryza... 57 2e-07
gb|AAP86445.1| ion channel NompC [Danio rerio] gi|34330186|ref|N... 57 3e-07
pdb|1N0Q|B Chain B, 3ank: A Designed Ankyrin Repeat Protein With... 55 8e-07
dbj|BAB09937.1| unnamed protein product [Arabidopsis thaliana] 55 1e-06
ref|NP_200282.2| ankyrin repeat family protein [Arabidopsis thal... 55 1e-06
gb|AAN12046.1| CG7462-PC, isoform C [Drosophila melanogaster] gi... 54 1e-06
gb|AAF50525.3| CG7462-PB, isoform B [Drosophila melanogaster] gi... 54 1e-06
gb|AAM11327.1| GH01626p [Drosophila melanogaster] 54 1e-06
gb|AAH87318.1| LOC495949 protein [Xenopus laevis] 54 1e-06
>gb|AAL87372.1| At1g10340/F14N23_22 [Arabidopsis thaliana]
gi|18391143|ref|NP_563867.1| ankyrin repeat family
protein [Arabidopsis thaliana]
gi|13937240|gb|AAK50112.1| At1g10340/F14N23_22
[Arabidopsis thaliana] gi|4914336|gb|AAD32884.1|
F14N23.22 [Arabidopsis thaliana]
Length = 578
Score = 70.5 bits (171), Expect = 2e-11
Identities = 40/133 (30%), Positives = 68/133 (51%), Gaps = 2/133 (1%)
Query: 3 QEFFNAIKNNDISTFSSIVKVREGILNQRTDDTF--NTPLHLASKYGCIEMVSEIVRLCP 60
Q F+AI ND+ F +V+ E L +R ++ NT LH+A+K+G E+VS+I+ L P
Sbjct: 2 QPIFHAILKNDLPAFLELVEDSESSLEERNEEEHLNNTVLHMAAKFGHRELVSKIIELRP 61
Query: 61 DMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLV 120
+VS+ N TP+H A +V ++M +LE N + +AC ++
Sbjct: 62 SLVSSRNAYRNTPLHLAAILGDVNIVMQMLETGLEVCSARNINNHTPLHLACRSNSIEAA 121
Query: 121 NLLLNLSEIVEPG 133
L+ ++ + G
Sbjct: 122 RLIAEKTQSIGLG 134
Score = 36.2 bits (82), Expect = 0.39
Identities = 31/124 (25%), Positives = 56/124 (45%), Gaps = 14/124 (11%)
Query: 33 DDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEV 92
D + +T LH A G E+ + ++ L + A N N +P+H A + +V +L L+
Sbjct: 168 DGSQSTLLHHACDKGDFELTTILLGLDQGLEEALNPNGLSPLHLAVLRGSVVILEEFLDK 227
Query: 93 NPTAACKLNPTCKSAFFVACSHGHLDL---------VNLLLNLSEIVEPGLAGFDQACFH 143
P + + P+ ++ F +A + ++D +N + L + E G H
Sbjct: 228 VPLSFSSITPSKETVFHLAARNKNMDAFVFMAESLGINSQILLQQTDESG-----NTVLH 282
Query: 144 IAAS 147
IAAS
Sbjct: 283 IAAS 286
Score = 34.3 bits (77), Expect = 1.5
Identities = 27/125 (21%), Positives = 53/125 (41%), Gaps = 5/125 (4%)
Query: 27 ILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVS-----AENENMETPIHEACRQE 81
++ ++T L LA G +V I+ PD+ E+ + T +H AC +
Sbjct: 123 LIAEKTQSIGLGELILAISSGSTSIVGTILERFPDLAREEAWVVEDGSQSTLLHHACDKG 182
Query: 82 NVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQAC 141
+ ++ +LL ++ LNP S +A G + ++ L+ + + +
Sbjct: 183 DFELTTILLGLDQGLEEALNPNGLSPLHLAVLRGSVVILEEFLDKVPLSFSSITPSKETV 242
Query: 142 FHIAA 146
FH+AA
Sbjct: 243 FHLAA 247
>ref|NP_849631.1| ankyrin repeat family protein [Arabidopsis thaliana]
Length = 574
Score = 70.5 bits (171), Expect = 2e-11
Identities = 40/133 (30%), Positives = 68/133 (51%), Gaps = 2/133 (1%)
Query: 3 QEFFNAIKNNDISTFSSIVKVREGILNQRTDDTF--NTPLHLASKYGCIEMVSEIVRLCP 60
Q F+AI ND+ F +V+ E L +R ++ NT LH+A+K+G E+VS+I+ L P
Sbjct: 2 QPIFHAILKNDLPAFLELVEDSESSLEERNEEEHLNNTVLHMAAKFGHRELVSKIIELRP 61
Query: 61 DMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLV 120
+VS+ N TP+H A +V ++M +LE N + +AC ++
Sbjct: 62 SLVSSRNAYRNTPLHLAAILGDVNIVMQMLETGLEVCSARNINNHTPLHLACRSNSIEAA 121
Query: 121 NLLLNLSEIVEPG 133
L+ ++ + G
Sbjct: 122 RLIAEKTQSIGLG 134
Score = 36.2 bits (82), Expect = 0.39
Identities = 31/124 (25%), Positives = 56/124 (45%), Gaps = 14/124 (11%)
Query: 33 DDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEV 92
D + +T LH A G E+ + ++ L + A N N +P+H A + +V +L L+
Sbjct: 164 DGSQSTLLHHACDKGDFELTTILLGLDQGLEEALNPNGLSPLHLAVLRGSVVILEEFLDK 223
Query: 93 NPTAACKLNPTCKSAFFVACSHGHLDL---------VNLLLNLSEIVEPGLAGFDQACFH 143
P + + P+ ++ F +A + ++D +N + L + E G H
Sbjct: 224 VPLSFSSITPSKETVFHLAARNKNMDAFVFMAESLGINSQILLQQTDESG-----NTVLH 278
Query: 144 IAAS 147
IAAS
Sbjct: 279 IAAS 282
Score = 32.7 bits (73), Expect = 4.3
Identities = 30/138 (21%), Positives = 54/138 (38%), Gaps = 28/138 (20%)
Query: 37 NTPLHLASKYGCIE-----------------------MVSEIVRLCPDMVS-----AENE 68
+TPLHLA + IE +V I+ PD+ E+
Sbjct: 106 HTPLHLACRSNSIEAARLIAEKTQSIGLGELILAISSIVGTILERFPDLAREEAWVVEDG 165
Query: 69 NMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLLNLSE 128
+ T +H AC + + ++ +LL ++ LNP S +A G + ++ L+
Sbjct: 166 SQSTLLHHACDKGDFELTTILLGLDQGLEEALNPNGLSPLHLAVLRGSVVILEEFLDKVP 225
Query: 129 IVEPGLAGFDQACFHIAA 146
+ + + FH+AA
Sbjct: 226 LSFSSITPSKETVFHLAA 243
>ref|NP_174667.1| ankyrin repeat family protein [Arabidopsis thaliana]
gi|25518163|pir||D86464 F12G12.13 protein - Arabidopsis
thaliana gi|10086472|gb|AAG12532.1| Hypothetical Protein
[Arabidopsis thaliana]
Length = 573
Score = 66.6 bits (161), Expect = 3e-10
Identities = 39/129 (30%), Positives = 69/129 (53%), Gaps = 2/129 (1%)
Query: 7 NAIKNNDISTFSSIVKVREGILNQRT--DDTFNTPLHLASKYGCIEMVSEIVRLCPDMVS 64
+AI ND+ST ++ + +L +R D T LHLA++ G E+V I++LCP +V
Sbjct: 23 DAILANDVSTLLALAEGNLSVLRERYHWDSLGGTVLHLATELGHKEIVEAIIKLCPSLVG 82
Query: 65 AENENMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLL 124
N + +TP+H A R + ++ +L +N ++AF VAC + + D+ +L+L
Sbjct: 83 VTNLDGDTPLHFAARWGHATIVAQILASGYAEFTPVNGRGETAFVVACRYTNPDVASLIL 142
Query: 125 NLSEIVEPG 133
+ + G
Sbjct: 143 EETSSITIG 151
Score = 34.7 bits (78), Expect = 1.1
Identities = 19/85 (22%), Positives = 39/85 (45%)
Query: 33 DDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEV 92
D +TPLH A +E+ ++ + + N++ TP+H A + ++ +L +
Sbjct: 181 DGELSTPLHHACNANNLEITKMLLEIDESLAERVNKDGFTPLHLAAMKCSIPILKEFSDK 240
Query: 93 NPTAACKLNPTCKSAFFVACSHGHL 117
P L P ++ F +A H ++
Sbjct: 241 APRYFDILTPAKETVFHLAAEHKNI 265
Score = 32.7 bits (73), Expect = 4.3
Identities = 12/42 (28%), Positives = 25/42 (58%)
Query: 60 PDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAACKLN 101
P + + + TP+H AC N+++ +LLE++ + A ++N
Sbjct: 174 PKLAWNADGELSTPLHHACNANNLEITKMLLEIDESLAERVN 215
>gb|AAM20099.1| unknown protein [Arabidopsis thaliana] gi|15810331|gb|AAL07053.1|
unknown protein [Arabidopsis thaliana]
gi|20197978|gb|AAD23887.2| expressed protein
[Arabidopsis thaliana] gi|18400588|ref|NP_565575.1|
ankyrin repeat family protein [Arabidopsis thaliana]
gi|42570312|ref|NP_850055.2| ankyrin repeat family
protein [Arabidopsis thaliana]
Length = 548
Score = 65.1 bits (157), Expect = 8e-10
Identities = 42/148 (28%), Positives = 75/148 (50%), Gaps = 9/148 (6%)
Query: 6 FNAIKNNDISTFSSIVKVREGILNQRTDD--TFNTPLHLASKYGCIEMVSEIVRLCPDMV 63
F+AI ND+ F +V+ RE L +R+++ T NT LH+A+K G E+V++I+ L P ++
Sbjct: 5 FDAILQNDLPAFLGLVEARESSLEERSEEQNTNNTVLHVAAKLGHRELVAKIIELRPSLL 64
Query: 64 SAENENMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLL 123
S+ N +TP+H A +V ++M +L+ N ++ HL V++
Sbjct: 65 SSRNAYGDTPLHLAALLGDVNIVMQMLDTGLELYSARNNKNQTPL-------HLAFVSIF 117
Query: 124 LNLSEIVEPGLAGFDQACFHIAASRGHT 151
+ ++ + D + A S G T
Sbjct: 118 MEAAKFIVEKTNSVDLDELNFALSSGST 145
Score = 38.9 bits (89), Expect = 0.061
Identities = 33/119 (27%), Positives = 57/119 (47%), Gaps = 4/119 (3%)
Query: 33 DDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEV 92
D + +T LH A G +E+ S ++ L + A N +P+H A ++ +V +L ++
Sbjct: 168 DGSRSTLLHYACDKGDLELTSILLGLNQGLEEALNSKGLSPLHLAVQRGSVIILEEFMDK 227
Query: 93 NPTAACKLNPTCKSAFFVACSHGHLD-LVNLLLNLSEIVEPGLAGFDQ---ACFHIAAS 147
+P + C P+ ++ F +A + + D V + NL L DQ HIAAS
Sbjct: 228 SPLSFCVRTPSKETVFHLAARNKNTDAFVFMAENLGTSSPILLKKKDQQGNTVLHIAAS 286
>pir||F84638 hypothetical protein At2g24600 [imported] - Arabidopsis thaliana
Length = 620
Score = 65.1 bits (157), Expect = 8e-10
Identities = 42/148 (28%), Positives = 75/148 (50%), Gaps = 9/148 (6%)
Query: 6 FNAIKNNDISTFSSIVKVREGILNQRTDD--TFNTPLHLASKYGCIEMVSEIVRLCPDMV 63
F+AI ND+ F +V+ RE L +R+++ T NT LH+A+K G E+V++I+ L P ++
Sbjct: 56 FDAILQNDLPAFLGLVEARESSLEERSEEQNTNNTVLHVAAKLGHRELVAKIIELRPSLL 115
Query: 64 SAENENMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLL 123
S+ N +TP+H A +V ++M +L+ N ++ HL V++
Sbjct: 116 SSRNAYGDTPLHLAALLGDVNIVMQMLDTGLELYSARNNKNQTPL-------HLAFVSIF 168
Query: 124 LNLSEIVEPGLAGFDQACFHIAASRGHT 151
+ ++ + D + A S G T
Sbjct: 169 MEAAKFIVEKTNSVDLDELNFALSSGST 196
Score = 38.9 bits (89), Expect = 0.061
Identities = 33/119 (27%), Positives = 57/119 (47%), Gaps = 4/119 (3%)
Query: 33 DDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEV 92
D + +T LH A G +E+ S ++ L + A N +P+H A ++ +V +L ++
Sbjct: 219 DGSRSTLLHYACDKGDLELTSILLGLNQGLEEALNSKGLSPLHLAVQRGSVIILEEFMDK 278
Query: 93 NPTAACKLNPTCKSAFFVACSHGHLD-LVNLLLNLSEIVEPGLAGFDQ---ACFHIAAS 147
+P + C P+ ++ F +A + + D V + NL L DQ HIAAS
Sbjct: 279 SPLSFCVRTPSKETVFHLAARNKNTDAFVFMAENLGTSSPILLKKKDQQGNTVLHIAAS 337
>ref|NP_973528.1| ankyrin repeat family protein [Arabidopsis thaliana]
Length = 601
Score = 65.1 bits (157), Expect = 8e-10
Identities = 42/148 (28%), Positives = 75/148 (50%), Gaps = 9/148 (6%)
Query: 6 FNAIKNNDISTFSSIVKVREGILNQRTDD--TFNTPLHLASKYGCIEMVSEIVRLCPDMV 63
F+AI ND+ F +V+ RE L +R+++ T NT LH+A+K G E+V++I+ L P ++
Sbjct: 5 FDAILQNDLPAFLGLVEARESSLEERSEEQNTNNTVLHVAAKLGHRELVAKIIELRPSLL 64
Query: 64 SAENENMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLL 123
S+ N +TP+H A +V ++M +L+ N ++ HL V++
Sbjct: 65 SSRNAYGDTPLHLAALLGDVNIVMQMLDTGLELYSARNNKNQTPL-------HLAFVSIF 117
Query: 124 LNLSEIVEPGLAGFDQACFHIAASRGHT 151
+ ++ + D + A S G T
Sbjct: 118 MEAAKFIVEKTNSVDLDELNFALSSGST 145
Score = 38.9 bits (89), Expect = 0.061
Identities = 33/119 (27%), Positives = 57/119 (47%), Gaps = 4/119 (3%)
Query: 33 DDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEV 92
D + +T LH A G +E+ S ++ L + A N +P+H A ++ +V +L ++
Sbjct: 168 DGSRSTLLHYACDKGDLELTSILLGLNQGLEEALNSKGLSPLHLAVQRGSVIILEEFMDK 227
Query: 93 NPTAACKLNPTCKSAFFVACSHGHLD-LVNLLLNLSEIVEPGLAGFDQ---ACFHIAAS 147
+P + C P+ ++ F +A + + D V + NL L DQ HIAAS
Sbjct: 228 SPLSFCVRTPSKETVFHLAARNKNTDAFVFMAENLGTSSPILLKKKDQQGNTVLHIAAS 286
>ref|NP_172055.1| ankyrin repeat family protein [Arabidopsis thaliana]
gi|25406852|pir||E86190 hypothetical protein [imported]
- Arabidopsis thaliana gi|4836914|gb|AAD30616.1|
Hypothetical protein [Arabidopsis thaliana]
Length = 627
Score = 60.8 bits (146), Expect = 1e-08
Identities = 42/155 (27%), Positives = 71/155 (45%), Gaps = 6/155 (3%)
Query: 2 DQEFFNAIKNNDISTFSSIVKVREGI-----LNQRTDDTFNTPLHLASKYGCIEMVSEIV 56
D A + ++ +++ GI L+ + + TPL+ A++ G +V E++
Sbjct: 114 DSPLHLAARTGNLGKVMELIRACNGIEELKELSSKQNLEGETPLYSAAENGHSLVVEEML 173
Query: 57 R-LCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHG 115
+ + D S + N P H A +Q +++ L LLE P A ++ +C +A A S G
Sbjct: 174 KHMDLDTASVKARNGFDPFHVAAKQGHIEALKKLLETFPNLAMTVDLSCTTALHTAASQG 233
Query: 116 HLDLVNLLLNLSEIVEPGLAGFDQACFHIAASRGH 150
H D+VNLLL + + H AA GH
Sbjct: 234 HTDVVNLLLKTDSHLAKIAKNNGKTALHSAARMGH 268
Score = 44.3 bits (103), Expect = 0.001
Identities = 32/116 (27%), Positives = 54/116 (45%), Gaps = 12/116 (10%)
Query: 9 IKNNDISTFSSIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENE 68
+K+ D+ T S VK R G P H+A+K G IE + +++ P++ +
Sbjct: 173 LKHMDLDTAS--VKARNGF----------DPFHVAAKQGHIEALKKLLETFPNLAMTVDL 220
Query: 69 NMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLL 124
+ T +H A Q + V+ LLL+ + A K+A A GH ++V L+
Sbjct: 221 SCTTALHTAASQGHTDVVNLLLKTDSHLAKIAKNNGKTALHSAARMGHREVVKSLI 276
Score = 43.1 bits (100), Expect = 0.003
Identities = 27/93 (29%), Positives = 48/93 (51%)
Query: 31 RTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLL 90
RTD T LH+A K +V E+V+ P ++S E+ TP+H A + +K++ L+
Sbjct: 285 RTDKKGQTALHMAVKGQNEGIVLELVKPDPAILSVEDSKGNTPLHTATNKGRIKIVRCLV 344
Query: 91 EVNPTAACKLNPTCKSAFFVACSHGHLDLVNLL 123
+ +N +A +A G+ +LV++L
Sbjct: 345 SFDGINLNAMNKAGDTALDIAEKIGNPELVSVL 377
>gb|AAD32290.1| ankyrin-like protein [Arabidopsis thaliana] gi|25408188|pir||E84725
ankyrin-like protein [imported] - Arabidopsis thaliana
gi|15225141|ref|NP_180741.1| ankyrin repeat family
protein [Arabidopsis thaliana]
Length = 662
Score = 58.9 bits (141), Expect = 6e-08
Identities = 36/114 (31%), Positives = 58/114 (50%), Gaps = 1/114 (0%)
Query: 38 TPLHLASKYGCIEMVSEIVR-LCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTA 96
TPL+ A++ G +V E+++ + + S N P H A +Q +++VL +LLE P
Sbjct: 191 TPLYTAAENGHSIVVEEMLKHMDLETASIAARNGFDPFHVAAKQGHLEVLKILLETFPNL 250
Query: 97 ACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQACFHIAASRGH 150
A + +C +A A + GH+D+VNLLL + + H AA GH
Sbjct: 251 AMTTDLSCTTALHTAATQGHIDVVNLLLETDSNLAKIAKNNGKTALHSAARMGH 304
Score = 45.8 bits (107), Expect = 5e-04
Identities = 28/114 (24%), Positives = 52/114 (45%)
Query: 39 PLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAAC 98
P H+A+K G +E++ ++ P++ + + T +H A Q ++ V+ LLLE + A
Sbjct: 227 PFHVAAKQGHLEVLKILLETFPNLAMTTDLSCTTALHTAATQGHIDVVNLLLETDSNLAK 286
Query: 99 KLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQACFHIAASRGHTG 152
K+A A GH+++V L+ + Q H+A + G
Sbjct: 287 IAKNNGKTALHSAARMGHVEVVKSLIGKDPSIGFRTDKKGQTALHMAVKGQNDG 340
Score = 38.1 bits (87), Expect = 0.10
Identities = 31/128 (24%), Positives = 60/128 (46%), Gaps = 12/128 (9%)
Query: 28 LNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLM 87
L TD + T LH A+ G I++V+ ++ ++ N +T +H A R +V+V+
Sbjct: 250 LAMTTDLSCTTALHTAATQGHIDVVNLLLETDSNLAKIAKNNGKTALHSAARMGHVEVVK 309
Query: 88 LLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFD------QAC 141
L+ +P+ + + ++A +A G D + + E+V+P +A
Sbjct: 310 SLIGKDPSIGFRTDKKGQTALHMAVK-GQNDGI-----VVELVKPDVAVLSVEDNKGNTP 363
Query: 142 FHIAASRG 149
HIA ++G
Sbjct: 364 LHIATNKG 371
Score = 36.6 bits (83), Expect = 0.30
Identities = 25/93 (26%), Positives = 45/93 (47%)
Query: 31 RTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLL 90
RTD T LH+A K +V E+V+ ++S E+ TP+H A + +K++ L+
Sbjct: 321 RTDKKGQTALHMAVKGQNDGIVVELVKPDVAVLSVEDNKGNTPLHIATNKGRIKIVRCLV 380
Query: 91 EVNPTAACKLNPTCKSAFFVACSHGHLDLVNLL 123
+N + V+ G+ +LV++L
Sbjct: 381 SFEGINLNPINKAGDTPLDVSEKIGNAELVSVL 413
Score = 32.0 bits (71), Expect = 7.4
Identities = 23/90 (25%), Positives = 45/90 (49%), Gaps = 1/90 (1%)
Query: 8 AIKNNDISTFSSIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAEN 67
A+K + +VK +L+ D+ NTPLH+A+ G I++V +V ++ N
Sbjct: 333 AVKGQNDGIVVELVKPDVAVLSVE-DNKGNTPLHIATNKGRIKIVRCLVSFEGINLNPIN 391
Query: 68 ENMETPIHEACRQENVKVLMLLLEVNPTAA 97
+ +TP+ + + N +++ +L E A
Sbjct: 392 KAGDTPLDVSEKIGNAELVSVLKEAGAATA 421
>gb|AAF75083.1| It contains Ank repeat PF|00023. EST gb|AI996003 comes from this
gene. [Arabidopsis thaliana]
gi|15222993|ref|NP_172250.1| ankyrin repeat family
protein [Arabidopsis thaliana] gi|25406998|pir||C86212
hypothetical protein [imported] - Arabidopsis thaliana
Length = 543
Score = 57.8 bits (138), Expect = 1e-07
Identities = 39/136 (28%), Positives = 68/136 (49%), Gaps = 6/136 (4%)
Query: 20 IVKVREGILNQ---RTDDTFNTPLHLASKYGCIEMVSEIVRLCPDM--VSAENENMETPI 74
+ K RE LNQ + + + T L++A++YG +E+V E++ C D+ V + N
Sbjct: 47 LTKTRESELNQLLGKQNQSGETALYVAAEYGDVEIVKEMIN-CYDLALVEIKARNGFDAF 105
Query: 75 HEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGL 134
H A +Q ++ VL +L E + A ++ + +A A + GH ++VN LL L +
Sbjct: 106 HIAAKQGDLDVLKVLAEAHSELAMTVDLSNTTALHTAATQGHTEVVNFLLELGSSLAGIA 165
Query: 135 AGFDQACFHIAASRGH 150
+ H A+ GH
Sbjct: 166 KSNGKTALHSASRNGH 181
Score = 47.8 bits (112), Expect = 1e-04
Identities = 35/147 (23%), Positives = 66/147 (44%), Gaps = 12/147 (8%)
Query: 5 FFNAIKNNDISTFSSIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVS 64
F A K D+ + + L D + T LH A+ G E+V+ ++ L +
Sbjct: 105 FHIAAKQGDLDVLKVLAEAHSE-LAMTVDLSNTTALHTAATQGHTEVVNFLLELGSSLAG 163
Query: 65 AENENMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLL 124
N +T +H A R +VKV+ LL P A +++ ++A +A ++++V L+
Sbjct: 164 IAKSNGKTALHSASRNGHVKVIKALLASEPAIAIRMDKKGQTALHMAVKGTNVEVVEELI 223
Query: 125 NLSEIVEPGLAGFDQACFHIAASRGHT 151
D++ +IA ++G+T
Sbjct: 224 KA-----------DRSSINIADTKGNT 239
>dbj|BAD34362.1| ankyrin-like protein [Oryza sativa (japonica cultivar-group)]
gi|50725332|dbj|BAD34405.1| ankyrin-like protein [Oryza
sativa (japonica cultivar-group)]
Length = 562
Score = 57.8 bits (138), Expect = 1e-07
Identities = 41/127 (32%), Positives = 62/127 (48%), Gaps = 3/127 (2%)
Query: 26 GILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSA--ENENMETPIHEACRQENV 83
G L R + T L+++++ G E+VSEI++ C D+ SA + N H A +Q ++
Sbjct: 78 GELAARQNQDGETALYVSAEKGHTEVVSEILKFC-DLQSAGLKATNSFDAFHIAAKQGHL 136
Query: 84 KVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQACFH 143
VL LL+ P A N +A A + GH+D+VNLLL + + H
Sbjct: 137 DVLKELLQAFPALAMTTNSVNATALDTAATQGHIDIVNLLLETDASLARIARNNGKTVLH 196
Query: 144 IAASRGH 150
AA GH
Sbjct: 197 SAARMGH 203
Score = 47.4 bits (111), Expect = 2e-04
Identities = 36/134 (26%), Positives = 65/134 (47%), Gaps = 14/134 (10%)
Query: 41 HLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAACKL 100
H+A+K G ++++ E+++ P + N T + A Q ++ ++ LLLE + + A
Sbjct: 128 HIAAKQGHLDVLKELLQAFPALAMTTNSVNATALDTAATQGHIDIVNLLLETDASLARIA 187
Query: 101 NPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGF-----DQACFHIAASRGHTGENK 155
K+ A GH+++V LLN +PG+ GF Q H+A+ G+N
Sbjct: 188 RNNGKTVLHSAARMGHVEVVTALLN----KDPGI-GFRTDKKGQTALHMASK----GQNA 238
Query: 156 EFSLLCLHVFLTLL 169
E L L L+++
Sbjct: 239 EILLELLKPDLSVI 252
Score = 43.5 bits (101), Expect = 0.002
Identities = 30/109 (27%), Positives = 54/109 (49%), Gaps = 6/109 (5%)
Query: 21 VKVREGILNQ------RTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPI 74
V+V +LN+ RTD T LH+ASK E++ E+++ ++ E+ +
Sbjct: 204 VEVVTALLNKDPGIGFRTDKKGQTALHMASKGQNAEILLELLKPDLSVIHVEDNKGNRAL 263
Query: 75 HEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLL 123
H A R+ N ++ L+ V +N ++AF +A G+ +L N+L
Sbjct: 264 HVATRKGNTVIVQTLISVKEIVINAVNRAGETAFAIAEKLGNEELSNIL 312
Score = 42.4 bits (98), Expect = 0.005
Identities = 29/120 (24%), Positives = 57/120 (47%), Gaps = 6/120 (5%)
Query: 37 NTPLHLASKYGCIEMVSEIV-----RLCPDMVSAENENMETPIHEACRQENVKVLMLLLE 91
+T LHLA++ G + V +I L ++ + +N++ ET ++ + + + +V+ +L+
Sbjct: 50 DTELHLAARAGSVPHVQKIFAASDPELVGELAARQNQDGETALYVSAEKGHTEVVSEILK 109
Query: 92 VNPTAACKLNPTCK-SAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQACFHIAASRGH 150
+ L T AF +A GHLD++ LL + + AA++GH
Sbjct: 110 FCDLQSAGLKATNSFDAFHIAAKQGHLDVLKELLQAFPALAMTTNSVNATALDTAATQGH 169
Score = 37.0 bits (84), Expect = 0.23
Identities = 25/92 (27%), Positives = 44/92 (47%), Gaps = 6/92 (6%)
Query: 71 ETPIHEACRQENVKVLMLLL-----EVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLLN 125
+T +H A R +V + + E+ A + N ++A +V+ GH ++V+ +L
Sbjct: 50 DTELHLAARAGSVPHVQKIFAASDPELVGELAARQNQDGETALYVSAEKGHTEVVSEILK 109
Query: 126 LSEIVEPGLAGFDQA-CFHIAASRGHTGENKE 156
++ GL + FHIAA +GH KE
Sbjct: 110 FCDLQSAGLKATNSFDAFHIAAKQGHLDVLKE 141
>pdb|1N0R|A Chain A, 4ank: A Designed Ankyrin Repeat Protein With Four
Identical Consensus Repeats
Length = 126
Score = 57.8 bits (138), Expect = 1e-07
Identities = 38/113 (33%), Positives = 62/113 (54%), Gaps = 3/113 (2%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAA 97
TPLHLA++ G +E+V ++ D V+A+++N TP+H A R +++V+ LLLE
Sbjct: 4 TPLHLAARNGHLEVVKLLLEAGAD-VNAKDKNGRTPLHLAARNGHLEVVKLLLEAGADVN 62
Query: 98 CKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQACFHIAASRGH 150
K + ++ +A +GHL++V LLL V + H+AA GH
Sbjct: 63 AK-DKNGRTPLHLAARNGHLEVVKLLLEAGADVNAKDKN-GRTPLHLAARNGH 113
Score = 55.8 bits (133), Expect = 5e-07
Identities = 33/92 (35%), Positives = 55/92 (58%), Gaps = 2/92 (2%)
Query: 33 DDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEV 92
D TPLHLA++ G +E+V ++ D V+A+++N TP+H A R +++V+ LLLE
Sbjct: 32 DKNGRTPLHLAARNGHLEVVKLLLEAGAD-VNAKDKNGRTPLHLAARNGHLEVVKLLLEA 90
Query: 93 NPTAACKLNPTCKSAFFVACSHGHLDLVNLLL 124
K + ++ +A +GHL++V LLL
Sbjct: 91 GADVNAK-DKNGRTPLHLAARNGHLEVVKLLL 121
Score = 44.7 bits (104), Expect = 0.001
Identities = 24/59 (40%), Positives = 39/59 (65%), Gaps = 1/59 (1%)
Query: 33 DDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLE 91
D TPLHLA++ G +E+V ++ D V+A+++N TP+H A R +++V+ LLLE
Sbjct: 65 DKNGRTPLHLAARNGHLEVVKLLLEAGAD-VNAKDKNGRTPLHLAARNGHLEVVKLLLE 122
Score = 38.9 bits (89), Expect = 0.061
Identities = 26/82 (31%), Positives = 40/82 (48%), Gaps = 2/82 (2%)
Query: 69 NMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLLNLSE 128
N TP+H A R +++V+ LLLE K + ++ +A +GHL++V LLL
Sbjct: 1 NGRTPLHLAARNGHLEVVKLLLEAGADVNAK-DKNGRTPLHLAARNGHLEVVKLLLEAGA 59
Query: 129 IVEPGLAGFDQACFHIAASRGH 150
V + H+AA GH
Sbjct: 60 DVNAKDKN-GRTPLHLAARNGH 80
>gb|AAN64473.1| putative ankyrin repeat containing protein [Oryza sativa (japonica
cultivar-group)]
Length = 565
Score = 57.0 bits (136), Expect = 2e-07
Identities = 36/138 (26%), Positives = 70/138 (50%), Gaps = 12/138 (8%)
Query: 9 IKNNDISTFSSIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENE 68
IK +D++T + +K R G LH+A+K G +E+V+E+++ P++ +
Sbjct: 106 IKYHDVAT--ACIKARSGY----------DALHIAAKQGDVEVVNELLKALPELSMTVDA 153
Query: 69 NMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLLNLSE 128
+ T ++ A Q +++V+ LLLE + + A K+A A +GH+++V L+
Sbjct: 154 SNTTALNTAATQGHMEVVRLLLEADASLAVIARSNGKTALHSAARNGHVEVVRALMEAEP 213
Query: 129 IVEPGLAGFDQACFHIAA 146
+ + Q H+AA
Sbjct: 214 SIAARVDKKGQTALHMAA 231
Score = 50.8 bits (120), Expect = 2e-05
Identities = 38/148 (25%), Positives = 67/148 (44%), Gaps = 9/148 (6%)
Query: 8 AIKNNDISTFSSIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAEN 67
A K D+ + ++K L+ D + T L+ A+ G +E+V ++ +
Sbjct: 128 AAKQGDVEVVNELLKALPE-LSMTVDASNTTALNTAATQGHMEVVRLLLEADASLAVIAR 186
Query: 68 ENMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLLNLS 127
N +T +H A R +V+V+ L+E P+ A +++ ++A +A LD+V+ LL
Sbjct: 187 SNGKTALHSAARNGHVEVVRALMEAEPSIAARVDKKGQTALHMAAKGTRLDIVDALL--- 243
Query: 128 EIVEPGLAGF----DQACFHIAASRGHT 151
EP L HIAA + T
Sbjct: 244 -AGEPTLLNLADSKGNTALHIAARKART 270
Score = 48.5 bits (114), Expect = 8e-05
Identities = 31/115 (26%), Positives = 61/115 (52%), Gaps = 3/115 (2%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSA--ENENMETPIHEACRQENVKVLMLLLEVNPT 95
TPL +A++YG + +V+E+++ D+ +A + + +H A +Q +V+V+ LL+ P
Sbjct: 88 TPLFVAAEYGYVALVAEMIKY-HDVATACIKARSGYDALHIAAKQGDVEVVNELLKALPE 146
Query: 96 AACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQACFHIAASRGH 150
+ ++ + +A A + GH+++V LLL + + H AA GH
Sbjct: 147 LSMTVDASNTTALNTAATQGHMEVVRLLLEADASLAVIARSNGKTALHSAARNGH 201
Score = 40.4 bits (93), Expect = 0.021
Identities = 27/117 (23%), Positives = 57/117 (48%), Gaps = 1/117 (0%)
Query: 7 NAIKNNDISTFSSIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAE 66
+A +N + ++++ I R D T LH+A+K +++V ++ P +++
Sbjct: 195 SAARNGHVEVVRALMEAEPSIA-ARVDKKGQTALHMAAKGTRLDIVDALLAGEPTLLNLA 253
Query: 67 NENMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLL 123
+ T +H A R+ ++ LLE+ T +N + ++AF A G+ + V +L
Sbjct: 254 DSKGNTALHIAARKARTPIVKRLLELPDTDLKAINRSRETAFDTAEKMGNTESVAVL 310
>gb|AAP86445.1| ion channel NompC [Danio rerio] gi|34330186|ref|NP_899192.1|
transient receptor potential cation channel, subfamily
N, member 1 [Danio rerio]
Length = 1614
Score = 56.6 bits (135), Expect = 3e-07
Identities = 44/147 (29%), Positives = 68/147 (45%), Gaps = 7/147 (4%)
Query: 8 AIKNNDISTFSSIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAEN 67
A + + SS++ +R I TD TPLHLA++ E+V +RL P++ + N
Sbjct: 686 AAMSGQLDVCSSLLNLRADIT--ATDSRGQTPLHLAAESDHSEVVKLFLRLRPELSTLAN 743
Query: 68 ENMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKS--AFFVACSHGHLDLVNLLLN 125
E+ T H A + +V V+ LL N LN +A + GH ++V +LL
Sbjct: 744 EDGSTCTHIAAAKGSVSVIRELLMFNQGGVGTLNHKAHGLCPLHLAAAGGHAEVVKVLLE 803
Query: 126 L-SEIVEPGLAGFDQACFHIAASRGHT 151
+ + E G H+AA GHT
Sbjct: 804 AGASVTEEDAEG--MTAVHLAAKHGHT 828
Score = 53.5 bits (127), Expect = 2e-06
Identities = 42/137 (30%), Positives = 70/137 (50%), Gaps = 17/137 (12%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENME--TPIHEACRQENVKVLMLLLEVNPT 95
TPLHLAS+ G E V ++ CP + + N++ +P+H A + + V+ LLL + +
Sbjct: 900 TPLHLASQSGH-ESVVRLLLNCPGVQADAETNIQGSSPLHLAAQSGHTAVVGLLLSRSSS 958
Query: 96 AACKLNPTCKSAFFVACSHGHLDLVNLLLNL-SEIVEPGLAGFDQACFHIAASRGHTGEN 154
+ + +SA +A +HGH+D+V +LL +EI ++G+ H AA G
Sbjct: 959 LLHQADRRGRSALHLAAAHGHVDMVRVLLGQGAEINHTDMSGW--TALHYAAEAG----- 1011
Query: 155 KEFSLLCLHVFLTLLST 171
CL V L L+ +
Sbjct: 1012 ------CLEVLLFLVES 1022
Score = 42.7 bits (99), Expect = 0.004
Identities = 34/123 (27%), Positives = 64/123 (51%), Gaps = 20/123 (16%)
Query: 39 PLHLASKYGCIEMVSEIV--RLCPDMVSAENENMETPIHEACRQENVKVLMLLLE--VNP 94
PL LA++ G + +V E++ + P + +A+ N +T +H CR+ +V++ +L+E NP
Sbjct: 154 PLLLAAEAGNVGIVRELLSSQSEPQIRAAKTANGDTALHICCRRRDVEMAKILVEFGANP 213
Query: 95 TAACKLNPTCKSAFFVACSHGHLDLVNLLL------NLSEIVEPGLAGFDQACFHIAASR 148
+ N ++ +A G +++ L N+S+ + D++ HIAA R
Sbjct: 214 DSQ---NDEGQTPLHIAAHEGDENMLKFLYLCKANANISDKM-------DRSPLHIAAER 263
Query: 149 GHT 151
GHT
Sbjct: 264 GHT 266
Score = 42.0 bits (97), Expect = 0.007
Identities = 30/120 (25%), Positives = 58/120 (48%), Gaps = 6/120 (5%)
Query: 16 TFSSIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCP-----DMVSAENENM 70
T I+ + + T T TPLH +++ G ++ E++R P ++ ++N
Sbjct: 520 TIIQILMEHQADITAVTRQTGETPLHYSARVGNTAVLQEMLRNVPTNQIQTAINKHSKNG 579
Query: 71 ETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIV 130
+P+ A Q + +V+ +LL+ N + K+A +A GH D+V++LL+ V
Sbjct: 580 WSPLLLAADQGHTEVVKILLQ-NNARVDVFDEEGKAAIHLAAQRGHQDIVDVLLSQKAFV 638
Score = 40.0 bits (92), Expect = 0.027
Identities = 32/126 (25%), Positives = 64/126 (50%), Gaps = 4/126 (3%)
Query: 33 DDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEV 92
D+ +HLA++ G ++V +++ V+A+ + TP+H + + + +++ LL+E
Sbjct: 609 DEEGKAAIHLAAQRGHQDIV-DVLLSQKAFVNAKTKQGLTPLHLSAQNGSARLVRLLVEN 667
Query: 93 NPTAACKLNPTCKSAFFVACSHGHLDLVNLLLNL-SEIVEPGLAGFDQACFHIAASRGHT 151
+ + L+ ++ +A G LD+ + LLNL ++I G Q H+AA H+
Sbjct: 668 HQASVDALSLRKQTPLHLAAMSGQLDVCSSLLNLRADITATDSRG--QTPLHLAAESDHS 725
Query: 152 GENKEF 157
K F
Sbjct: 726 EVVKLF 731
Score = 38.9 bits (89), Expect = 0.061
Identities = 33/133 (24%), Positives = 55/133 (40%), Gaps = 4/133 (3%)
Query: 33 DDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEV 92
+D TPLH+A+ G M+ + + LC + ++ +P+H A + + V+ +L E
Sbjct: 217 NDEGQTPLHIAAHEGDENML-KFLYLCKANANISDKMDRSPLHIAAERGHTNVVEILTEK 275
Query: 93 NPTAACKLNPTCKSAFFVACSHGH-LDLVNLLLNLSEIVEPGLAGFDQACFHIAASRGHT 151
+ + +A GH ++ L + P +G C H AA RGHT
Sbjct: 276 FRSCVLARTKDGNTLLHIASQCGHPTTALSFLRKGVPLHMPNKSG--AVCLHAAAKRGHT 333
Query: 152 GENKEFSLLCLHV 164
K HV
Sbjct: 334 AVVKALLQKGAHV 346
Score = 38.5 bits (88), Expect = 0.079
Identities = 32/115 (27%), Positives = 55/115 (47%), Gaps = 3/115 (2%)
Query: 39 PLHLASKYGCIEM-VSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAA 97
PL L ++ +E + + R PD++ + E+ + + A R+ + VL LLE+ +
Sbjct: 16 PLALRGEWMALEQKIKTLERGDPDILQCDEESGMSLLMLAVRESRLSVLDRLLELGANPS 75
Query: 98 CKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQACFHIAASRGHTG 152
K +SA +A +H ++V LL+ ++ P DQ H AASR G
Sbjct: 76 DKTKDG-RSALHIAAAHSKDEIVKLLVRKTDPNSPA-GPNDQLPLHYAASRSTGG 128
Score = 36.6 bits (83), Expect = 0.30
Identities = 24/87 (27%), Positives = 45/87 (51%), Gaps = 1/87 (1%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAA 97
TPLH++++ E +E++ V+AE EN ET +H A R +++++ L++
Sbjct: 389 TPLHISARVKEGERAAEMLLKSGAEVNAEQENGETALHVAARHGSLQMIRALIQEGGDPR 448
Query: 98 CKLNPTCKSAFFVACSHGHLDLVNLLL 124
+ + +S VA H H +V +L
Sbjct: 449 WR-SRVGESPLHVAVRHCHAHVVQEIL 474
>pdb|1N0Q|B Chain B, 3ank: A Designed Ankyrin Repeat Protein With Three
Identical Consensus Repeats gi|28373835|pdb|1N0Q|A Chain
A, 3ank: A Designed Ankyrin Repeat Protein With Three
Identical Consensus Repeats
Length = 93
Score = 55.1 bits (131), Expect = 8e-07
Identities = 32/87 (36%), Positives = 54/87 (61%), Gaps = 2/87 (2%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAA 97
TPLHLA++ G +E+V ++ D V+A+++N TP+H A R +++V+ LLLE
Sbjct: 4 TPLHLAARNGHLEVVKLLLEAGAD-VNAKDKNGRTPLHLAARNGHLEVVKLLLEAGADVN 62
Query: 98 CKLNPTCKSAFFVACSHGHLDLVNLLL 124
K + ++ +A +GHL++V LLL
Sbjct: 63 AK-DKNGRTPLHLAARNGHLEVVKLLL 88
Score = 44.7 bits (104), Expect = 0.001
Identities = 24/59 (40%), Positives = 39/59 (65%), Gaps = 1/59 (1%)
Query: 33 DDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLE 91
D TPLHLA++ G +E+V ++ D V+A+++N TP+H A R +++V+ LLLE
Sbjct: 32 DKNGRTPLHLAARNGHLEVVKLLLEAGAD-VNAKDKNGRTPLHLAARNGHLEVVKLLLE 89
Score = 38.9 bits (89), Expect = 0.061
Identities = 26/82 (31%), Positives = 40/82 (48%), Gaps = 2/82 (2%)
Query: 69 NMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLLNLSE 128
N TP+H A R +++V+ LLLE K + ++ +A +GHL++V LLL
Sbjct: 1 NGRTPLHLAARNGHLEVVKLLLEAGADVNAK-DKNGRTPLHLAARNGHLEVVKLLLEAGA 59
Query: 129 IVEPGLAGFDQACFHIAASRGH 150
V + H+AA GH
Sbjct: 60 DVNAKDKN-GRTPLHLAARNGH 80
>dbj|BAB09937.1| unnamed protein product [Arabidopsis thaliana]
Length = 652
Score = 54.7 bits (130), Expect = 1e-06
Identities = 27/92 (29%), Positives = 49/92 (52%)
Query: 33 DDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEV 92
D+ +T LH A E ++++ LCP +VS N + TP+H A N+ +L +LE
Sbjct: 65 DEEQSTLLHKAVTQRNEEYATKVIDLCPSLVSVTNVDGNTPLHLAAEIGNINILWKMLET 124
Query: 93 NPTAACKLNPTCKSAFFVACSHGHLDLVNLLL 124
K+N ++AF +AC + +++ +L+
Sbjct: 125 GEAECMKINKQGQTAFILACLNNNVNSARILV 156
>ref|NP_200282.2| ankyrin repeat family protein [Arabidopsis thaliana]
Length = 598
Score = 54.7 bits (130), Expect = 1e-06
Identities = 27/92 (29%), Positives = 49/92 (52%)
Query: 33 DDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEV 92
D+ +T LH A E ++++ LCP +VS N + TP+H A N+ +L +LE
Sbjct: 65 DEEQSTLLHKAVTQRNEEYATKVIDLCPSLVSVTNVDGNTPLHLAAEIGNINILWKMLET 124
Query: 93 NPTAACKLNPTCKSAFFVACSHGHLDLVNLLL 124
K+N ++AF +AC + +++ +L+
Sbjct: 125 GEAECMKINKQGQTAFILACLNNNVNSARILV 156
>gb|AAN12046.1| CG7462-PC, isoform C [Drosophila melanogaster]
gi|8132557|gb|AAF73309.1| ankyrin 2 [Drosophila
melanogaster] gi|21356447|ref|NP_648148.1| CG7462-PC,
isoform C [Drosophila melanogaster]
Length = 1159
Score = 54.3 bits (129), Expect = 1e-06
Identities = 38/116 (32%), Positives = 57/116 (48%), Gaps = 9/116 (7%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLE--VNPT 95
TPLHL +KYG I++ +++ D V A+ +N TP+H AC N +V +LLLE +P
Sbjct: 537 TPLHLTAKYGHIKVAQLLLQKEAD-VDAQGKNGVTPLHVACHYNNQQVALLLLEKGASPH 595
Query: 96 AACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVE-PGLAGFDQACFHIAASRGH 150
A K T +A +D+ LL + AGF H+++ GH
Sbjct: 596 ATAKNGHT---PLHIAARKNQMDIATTLLEYGALANAESKAGFTP--LHLSSQEGH 646
Score = 43.5 bits (101), Expect = 0.002
Identities = 35/129 (27%), Positives = 59/129 (45%), Gaps = 7/129 (5%)
Query: 23 VREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVR--LCPDMVSAENENMETPIHEACRQ 80
+R G T ++ TPLH+A+ GC+ +V +++ PD+ + ETP+H A R
Sbjct: 390 LRHGASISATTESGLTPLHVAAFMGCMNIVIYLLQHDASPDVPTVRG---ETPLHLAARA 446
Query: 81 ENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQA 140
++ +LL + ++ +A G++D+V LLL V+ A
Sbjct: 447 NQTDIIRILLRNGAQVDARAREQ-QTPLHIASRLGNVDIVMLLLQHGAQVDATTKDMYTA 505
Query: 141 CFHIAASRG 149
HIAA G
Sbjct: 506 -LHIAAKEG 513
Score = 39.3 bits (90), Expect = 0.046
Identities = 32/113 (28%), Positives = 55/113 (48%), Gaps = 3/113 (2%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAA 97
+PLH+A+K+G MVS ++ + + A+ + TP+H A R + +V+ +LLE +
Sbjct: 240 SPLHVAAKWGKTNMVSLLLEKGGN-IEAKTRDGLTPLHCAARSGHEQVVDMLLERGAPIS 298
Query: 98 CKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQACFHIAASRGH 150
K + +A H+D +LL V+ + A H+AA GH
Sbjct: 299 AK-TKNGLAPLHMAAQGEHVDAARILLYHRAPVDEVTVDYLTA-LHVAAHCGH 349
Score = 39.3 bits (90), Expect = 0.046
Identities = 32/117 (27%), Positives = 56/117 (47%), Gaps = 11/117 (9%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAA 97
TPLHLA++ +++ ++R V A +TP+H A R NV ++MLLL+
Sbjct: 438 TPLHLAARANQTDIIRILLRNGAQ-VDARAREQQTPLHIASRLGNVDIVMLLLQ----HG 492
Query: 98 CKLNPTCK---SAFFVACSHGHLDLVNLLLNLSEIVEPGL-AGFDQACFHIAASRGH 150
+++ T K +A +A G ++ +L+ ++ GF H+ A GH
Sbjct: 493 AQVDATTKDMYTALHIAAKEGQDEVAAVLIENGAALDAATKKGFTP--LHLTAKYGH 547
Score = 37.4 bits (85), Expect = 0.18
Identities = 32/117 (27%), Positives = 54/117 (45%), Gaps = 8/117 (6%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAA 97
TPLH+AS YG + + +++ D+ + N+ +P+H A + ++ LLLE
Sbjct: 207 TPLHIASHYGNQNIANLLIQKGADVNYSAKHNI-SPLHVAAKWGKTNMVSLLLEKGGNIE 265
Query: 98 CKLNPTCKSAFFVACSHGHLDLVNLLLN----LSEIVEPGLAGFDQAC--FHIAASR 148
K + A GH +V++LL +S + GLA A H+ A+R
Sbjct: 266 AKTRDGL-TPLHCAARSGHEQVVDMLLERGAPISAKTKNGLAPLHMAAQGEHVDAAR 321
Score = 36.6 bits (83), Expect = 0.30
Identities = 35/114 (30%), Positives = 56/114 (48%), Gaps = 12/114 (10%)
Query: 19 SIVKVREGILNQRTDDTFNT----PLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPI 74
++ +V E + N +T N LHLASK G I +VSE++R +V + + T +
Sbjct: 23 NLERVLEHLKNNIDINTSNANGLNALHLASKDGHIHVVSELLRR-GAIVDSATKKGNTAL 81
Query: 75 HEACRQENVKVLMLLLEVNPTAACKLNPTCKSAF---FVACSHGHLDLVNLLLN 125
H A +V+ LLLE N +N ++ F ++A H +V LLL+
Sbjct: 82 HIASLAGQEEVVKLLLEHN----ASVNVQSQNGFTPLYMAAQENHDAVVRLLLS 131
Score = 33.1 bits (74), Expect = 3.3
Identities = 24/98 (24%), Positives = 51/98 (51%), Gaps = 2/98 (2%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAA 97
TPLH+AS +G MV +++ ++ +A + TP+H+ +Q + ++ LLLE A
Sbjct: 702 TPLHVASHFGQANMVRFLLQNGANVDAATSIGY-TPLHQTAQQGHCHIVNLLLEHKANAN 760
Query: 98 CKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLA 135
+ ++ +A G++ +++ L +++ E A
Sbjct: 761 AQ-TVNGQTPLHIARKLGYISVLDSLKTITKEDETAAA 797
Score = 32.7 bits (73), Expect = 4.3
Identities = 29/126 (23%), Positives = 60/126 (47%), Gaps = 17/126 (13%)
Query: 19 SIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEAC 78
+++ + +G T +TPLH+A++ +++ + ++ + +AE++ TP+H +
Sbjct: 584 ALLLLEKGASPHATAKNGHTPLHIAARKNQMDIATTLLEYGA-LANAESKAGFTPLHLSS 642
Query: 79 RQENVKVLMLLLE----VNPTAACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGL 134
++ + ++ LL+E VN A L P HL +N++EI+E
Sbjct: 643 QEGHAEISNLLIEHKAAVNHPAKNGLTPM------------HLCAQEDNVNVAEILEKNG 690
Query: 135 AGFDQA 140
A D A
Sbjct: 691 ANIDMA 696
Score = 32.7 bits (73), Expect = 4.3
Identities = 23/75 (30%), Positives = 38/75 (50%), Gaps = 5/75 (6%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLL----EVN 93
T LH+A+ G + + ++ D +A N TP+H AC++ +KV+ LLL ++
Sbjct: 339 TALHVAAHCGHVRVAKLLLDRNAD-ANARALNGFTPLHIACKKNRLKVVELLLRHGASIS 397
Query: 94 PTAACKLNPTCKSAF 108
T L P +AF
Sbjct: 398 ATTESGLTPLHVAAF 412
>gb|AAF50525.3| CG7462-PB, isoform B [Drosophila melanogaster]
gi|45551532|ref|NP_729285.2| CG7462-PB, isoform B
[Drosophila melanogaster]
Length = 1571
Score = 54.3 bits (129), Expect = 1e-06
Identities = 38/116 (32%), Positives = 57/116 (48%), Gaps = 9/116 (7%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLE--VNPT 95
TPLHL +KYG I++ +++ D V A+ +N TP+H AC N +V +LLLE +P
Sbjct: 537 TPLHLTAKYGHIKVAQLLLQKEAD-VDAQGKNGVTPLHVACHYNNQQVALLLLEKGASPH 595
Query: 96 AACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVE-PGLAGFDQACFHIAASRGH 150
A K T +A +D+ LL + AGF H+++ GH
Sbjct: 596 ATAKNGHT---PLHIAARKNQMDIATTLLEYGALANAESKAGFTP--LHLSSQEGH 646
Score = 43.5 bits (101), Expect = 0.002
Identities = 35/129 (27%), Positives = 59/129 (45%), Gaps = 7/129 (5%)
Query: 23 VREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVR--LCPDMVSAENENMETPIHEACRQ 80
+R G T ++ TPLH+A+ GC+ +V +++ PD+ + ETP+H A R
Sbjct: 390 LRHGASISATTESGLTPLHVAAFMGCMNIVIYLLQHDASPDVPTVRG---ETPLHLAARA 446
Query: 81 ENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQA 140
++ +LL + ++ +A G++D+V LLL V+ A
Sbjct: 447 NQTDIIRILLRNGAQVDARAREQ-QTPLHIASRLGNVDIVMLLLQHGAQVDATTKDMYTA 505
Query: 141 CFHIAASRG 149
HIAA G
Sbjct: 506 -LHIAAKEG 513
Score = 39.3 bits (90), Expect = 0.046
Identities = 32/113 (28%), Positives = 55/113 (48%), Gaps = 3/113 (2%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAA 97
+PLH+A+K+G MVS ++ + + A+ + TP+H A R + +V+ +LLE +
Sbjct: 240 SPLHVAAKWGKTNMVSLLLEKGGN-IEAKTRDGLTPLHCAARSGHEQVVDMLLERGAPIS 298
Query: 98 CKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQACFHIAASRGH 150
K + +A H+D +LL V+ + A H+AA GH
Sbjct: 299 AK-TKNGLAPLHMAAQGEHVDAARILLYHRAPVDEVTVDYLTA-LHVAAHCGH 349
Score = 39.3 bits (90), Expect = 0.046
Identities = 32/117 (27%), Positives = 56/117 (47%), Gaps = 11/117 (9%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAA 97
TPLHLA++ +++ ++R V A +TP+H A R NV ++MLLL+
Sbjct: 438 TPLHLAARANQTDIIRILLRNGAQ-VDARAREQQTPLHIASRLGNVDIVMLLLQ----HG 492
Query: 98 CKLNPTCK---SAFFVACSHGHLDLVNLLLNLSEIVEPGL-AGFDQACFHIAASRGH 150
+++ T K +A +A G ++ +L+ ++ GF H+ A GH
Sbjct: 493 AQVDATTKDMYTALHIAAKEGQDEVAAVLIENGAALDAATKKGFTP--LHLTAKYGH 547
Score = 37.4 bits (85), Expect = 0.18
Identities = 32/117 (27%), Positives = 54/117 (45%), Gaps = 8/117 (6%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAA 97
TPLH+AS YG + + +++ D+ + N+ +P+H A + ++ LLLE
Sbjct: 207 TPLHIASHYGNQNIANLLIQKGADVNYSAKHNI-SPLHVAAKWGKTNMVSLLLEKGGNIE 265
Query: 98 CKLNPTCKSAFFVACSHGHLDLVNLLLN----LSEIVEPGLAGFDQAC--FHIAASR 148
K + A GH +V++LL +S + GLA A H+ A+R
Sbjct: 266 AKTRDGL-TPLHCAARSGHEQVVDMLLERGAPISAKTKNGLAPLHMAAQGEHVDAAR 321
Score = 36.6 bits (83), Expect = 0.30
Identities = 35/114 (30%), Positives = 56/114 (48%), Gaps = 12/114 (10%)
Query: 19 SIVKVREGILNQRTDDTFNT----PLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPI 74
++ +V E + N +T N LHLASK G I +VSE++R +V + + T +
Sbjct: 23 NLERVLEHLKNNIDINTSNANGLNALHLASKDGHIHVVSELLRR-GAIVDSATKKGNTAL 81
Query: 75 HEACRQENVKVLMLLLEVNPTAACKLNPTCKSAF---FVACSHGHLDLVNLLLN 125
H A +V+ LLLE N +N ++ F ++A H +V LLL+
Sbjct: 82 HIASLAGQEEVVKLLLEHN----ASVNVQSQNGFTPLYMAAQENHDAVVRLLLS 131
Score = 33.1 bits (74), Expect = 3.3
Identities = 24/98 (24%), Positives = 51/98 (51%), Gaps = 2/98 (2%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAA 97
TPLH+AS +G MV +++ ++ +A + TP+H+ +Q + ++ LLLE A
Sbjct: 702 TPLHVASHFGQANMVRFLLQNGANVDAATSIGY-TPLHQTAQQGHCHIVNLLLEHKANAN 760
Query: 98 CKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLA 135
+ ++ +A G++ +++ L +++ E A
Sbjct: 761 AQ-TVNGQTPLHIARKLGYISVLDSLKTITKEDETAAA 797
Score = 32.7 bits (73), Expect = 4.3
Identities = 29/126 (23%), Positives = 60/126 (47%), Gaps = 17/126 (13%)
Query: 19 SIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEAC 78
+++ + +G T +TPLH+A++ +++ + ++ + +AE++ TP+H +
Sbjct: 584 ALLLLEKGASPHATAKNGHTPLHIAARKNQMDIATTLLEYGA-LANAESKAGFTPLHLSS 642
Query: 79 RQENVKVLMLLLE----VNPTAACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGL 134
++ + ++ LL+E VN A L P HL +N++EI+E
Sbjct: 643 QEGHAEISNLLIEHKAAVNHPAKNGLTPM------------HLCAQEDNVNVAEILEKNG 690
Query: 135 AGFDQA 140
A D A
Sbjct: 691 ANIDMA 696
Score = 32.7 bits (73), Expect = 4.3
Identities = 23/75 (30%), Positives = 38/75 (50%), Gaps = 5/75 (6%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLL----EVN 93
T LH+A+ G + + ++ D +A N TP+H AC++ +KV+ LLL ++
Sbjct: 339 TALHVAAHCGHVRVAKLLLDRNAD-ANARALNGFTPLHIACKKNRLKVVELLLRHGASIS 397
Query: 94 PTAACKLNPTCKSAF 108
T L P +AF
Sbjct: 398 ATTESGLTPLHVAAF 412
>gb|AAM11327.1| GH01626p [Drosophila melanogaster]
Length = 1009
Score = 54.3 bits (129), Expect = 1e-06
Identities = 38/116 (32%), Positives = 57/116 (48%), Gaps = 9/116 (7%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLE--VNPT 95
TPLHL +KYG I++ +++ D V A+ +N TP+H AC N +V +LLLE +P
Sbjct: 387 TPLHLTAKYGHIKVAQLLLQKEAD-VDAQGKNGVTPLHVACHYNNQQVALLLLEKGASPH 445
Query: 96 AACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVE-PGLAGFDQACFHIAASRGH 150
A K T +A +D+ LL + AGF H+++ GH
Sbjct: 446 ATAKNGHT---PLHIAARKNQMDIATTLLEYGALANAESKAGFTP--LHLSSQEGH 496
Score = 43.5 bits (101), Expect = 0.002
Identities = 35/129 (27%), Positives = 59/129 (45%), Gaps = 7/129 (5%)
Query: 23 VREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVR--LCPDMVSAENENMETPIHEACRQ 80
+R G T ++ TPLH+A+ GC+ +V +++ PD+ + ETP+H A R
Sbjct: 240 LRHGASISATTESGLTPLHVAAFMGCMNIVIYLLQHDASPDVPTVRG---ETPLHLAARA 296
Query: 81 ENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQA 140
++ +LL + ++ +A G++D+V LLL V+ A
Sbjct: 297 NQTDIIRILLRNGAQVDARAREQ-QTPLHIASRLGNVDIVMLLLQHGAQVDATTKDMYTA 355
Query: 141 CFHIAASRG 149
HIAA G
Sbjct: 356 -LHIAAKEG 363
Score = 39.3 bits (90), Expect = 0.046
Identities = 32/113 (28%), Positives = 55/113 (48%), Gaps = 3/113 (2%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAA 97
+PLH+A+K+G MVS ++ + + A+ + TP+H A R + +V+ +LLE +
Sbjct: 90 SPLHVAAKWGKTNMVSLLLEKGGN-IEAKTRDGLTPLHCAARSGHEQVVDMLLERGAPIS 148
Query: 98 CKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQACFHIAASRGH 150
K + +A H+D +LL V+ + A H+AA GH
Sbjct: 149 AK-TKNGLAPLHMAAQGEHVDAARILLYHRAPVDEVTVDYLTA-LHVAAHCGH 199
Score = 39.3 bits (90), Expect = 0.046
Identities = 32/117 (27%), Positives = 56/117 (47%), Gaps = 11/117 (9%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAA 97
TPLHLA++ +++ ++R V A +TP+H A R NV ++MLLL+
Sbjct: 288 TPLHLAARANQTDIIRILLRNGAQ-VDARAREQQTPLHIASRLGNVDIVMLLLQ----HG 342
Query: 98 CKLNPTCK---SAFFVACSHGHLDLVNLLLNLSEIVEPGL-AGFDQACFHIAASRGH 150
+++ T K +A +A G ++ +L+ ++ GF H+ A GH
Sbjct: 343 AQVDATTKDMYTALHIAAKEGQDEVAAVLIENGAALDAATKKGFTP--LHLTAKYGH 397
Score = 37.4 bits (85), Expect = 0.18
Identities = 32/117 (27%), Positives = 54/117 (45%), Gaps = 8/117 (6%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAA 97
TPLH+AS YG + + +++ D+ + N+ +P+H A + ++ LLLE
Sbjct: 57 TPLHIASHYGNQNIANLLIQKGADVNYSAKHNI-SPLHVAAKWGKTNMVSLLLEKGGNIE 115
Query: 98 CKLNPTCKSAFFVACSHGHLDLVNLLLN----LSEIVEPGLAGFDQAC--FHIAASR 148
K + A GH +V++LL +S + GLA A H+ A+R
Sbjct: 116 AKTRDGL-TPLHCAARSGHEQVVDMLLERGAPISAKTKNGLAPLHMAAQGEHVDAAR 171
Score = 33.1 bits (74), Expect = 3.3
Identities = 25/98 (25%), Positives = 49/98 (49%), Gaps = 2/98 (2%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAA 97
TPLH+AS +G MV +++ + V A TP+H+ +Q + ++ LLLE A
Sbjct: 552 TPLHVASHFGQANMVRFLLQNGAN-VDAATSIGYTPLHQTAQQGHCHIVNLLLEHKANAN 610
Query: 98 CKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLA 135
+ ++ +A G++ +++ L +++ E A
Sbjct: 611 AQ-TVNGQTPLHIARKLGYISVLDSLKTITKEDETAAA 647
Score = 32.7 bits (73), Expect = 4.3
Identities = 23/75 (30%), Positives = 38/75 (50%), Gaps = 5/75 (6%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLL----EVN 93
T LH+A+ G + + ++ D +A N TP+H AC++ +KV+ LLL ++
Sbjct: 189 TALHVAAHCGHVRVAKLLLDRNAD-ANARALNGFTPLHIACKKNRLKVVELLLRHGASIS 247
Query: 94 PTAACKLNPTCKSAF 108
T L P +AF
Sbjct: 248 ATTESGLTPLHVAAF 262
Score = 32.7 bits (73), Expect = 4.3
Identities = 29/126 (23%), Positives = 60/126 (47%), Gaps = 17/126 (13%)
Query: 19 SIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEAC 78
+++ + +G T +TPLH+A++ +++ + ++ + +AE++ TP+H +
Sbjct: 434 ALLLLEKGASPHATAKNGHTPLHIAARKNQMDIATTLLEYGA-LANAESKAGFTPLHLSS 492
Query: 79 RQENVKVLMLLLE----VNPTAACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGL 134
++ + ++ LL+E VN A L P HL +N++EI+E
Sbjct: 493 QEGHAEISNLLIEHKAAVNHPAKNGLTPM------------HLCAQEDNVNVAEILEKNG 540
Query: 135 AGFDQA 140
A D A
Sbjct: 541 ANIDMA 546
>gb|AAH87318.1| LOC495949 protein [Xenopus laevis]
Length = 293
Score = 54.3 bits (129), Expect = 1e-06
Identities = 33/129 (25%), Positives = 60/129 (45%), Gaps = 13/129 (10%)
Query: 41 HLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAACKL 100
H+A++ G + ++ ++ + PD+ E++ TP+H A +V+ +LLE C
Sbjct: 150 HIATREGDVAIIQYLLDVFPDIWKTESKIKRTPLHTAAMHGCFEVIEVLLE-----RCNY 204
Query: 101 NPTCKSA-----FFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQACFHIAASRGHTGENK 155
+P CK + F A +GHL + LL+ ++ H+AA TG+N+
Sbjct: 205 DPDCKDSCGVTPFMDAVQNGHLSIAQLLIEKKKVCYSAFDRMGAQALHLAAV---TGQNE 261
Query: 156 EFSLLCLHV 164
L H+
Sbjct: 262 SLQYLVSHL 270
Score = 40.4 bits (93), Expect = 0.021
Identities = 31/114 (27%), Positives = 51/114 (44%), Gaps = 4/114 (3%)
Query: 37 NTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTA 96
+T LH A+++G + +++ +V V N + + P+HEA + L+ LL
Sbjct: 46 DTLLHYAARHGHLSILTYLVEAVGMDVEVLNNDYKRPLHEAASMGHRDCLLYLLSRGTEV 105
Query: 97 ACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFD-QACFHIAASRG 149
C L + +AC+ LD++ L + P L D CFHIA G
Sbjct: 106 DC-LKKADWTPLMMACTKTKLDIIKDL--IEHRANPMLKNKDGWNCFHIATREG 156
Score = 35.0 bits (79), Expect = 0.87
Identities = 25/110 (22%), Positives = 45/110 (40%), Gaps = 1/110 (0%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAA 97
TPLH A+ +GC E++ ++ C ++ TP +A + ++ + LL+E
Sbjct: 181 TPLHTAAMHGCFEVIEVLLERCNYDPDCKDSCGVTPFMDAVQNGHLSIAQLLIEKKKVCY 240
Query: 98 CKLNPTCKSAFFVACSHGHLD-LVNLLLNLSEIVEPGLAGFDQACFHIAA 146
+ A +A G + L L+ +L V + H AA
Sbjct: 241 SAFDRMGAQALHLAAVTGQNESLQYLVSHLGVNVNEKATSMQLSALHYAA 290
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.323 0.137 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 284,654,000
Number of Sequences: 2540612
Number of extensions: 10341257
Number of successful extensions: 38501
Number of sequences better than 10.0: 1958
Number of HSP's better than 10.0 without gapping: 369
Number of HSP's successfully gapped in prelim test: 1619
Number of HSP's that attempted gapping in prelim test: 31545
Number of HSP's gapped (non-prelim): 6278
length of query: 174
length of database: 863,360,394
effective HSP length: 119
effective length of query: 55
effective length of database: 561,027,566
effective search space: 30856516130
effective search space used: 30856516130
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 70 (31.6 bits)
Medicago: description of AC138130.15


