

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138087.1 + phase: 0
(1399 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
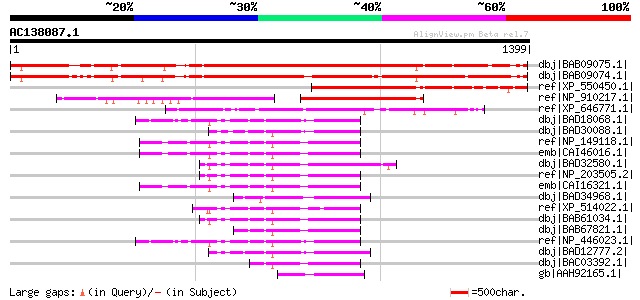
Score E
Sequences producing significant alignments: (bits) Value
dbj|BAB09075.1| unnamed protein product [Arabidopsis thaliana] g... 1173 0.0
dbj|BAB09074.1| unnamed protein product [Arabidopsis thaliana] g... 1138 0.0
ref|XP_550450.1| unknown protein [Oryza sativa (japonica cultiva... 520 e-145
ref|NP_910217.1| EST AU031036(E60617) corresponds to a region of... 372 e-101
ref|XP_646771.1| hypothetical protein DDB0191321 [Dictyostelium ... 247 2e-63
dbj|BAD18068.1| RGPR-p117 [Rattus norvegicus] 149 7e-34
dbj|BAD30088.1| RGPR-p117 [Oryctolagus cuniculus] 143 5e-32
ref|NP_149118.1| regucalcin gene promotor region related protein... 142 1e-31
emb|CAI46016.1| hypothetical protein [Homo sapiens] 140 4e-31
dbj|BAD32580.1| mKIAA1928 protein [Mus musculus] 138 1e-30
ref|NP_203505.2| regucalcin gene promotor region related protein... 137 2e-30
emb|CAI16321.1| regucalcin gene promotor region related protein ... 137 3e-30
dbj|BAD34968.1| regucalcin gene promotor region related protein ... 136 6e-30
ref|XP_514022.1| PREDICTED: hypothetical protein XP_514022 [Pan ... 135 7e-30
dbj|BAB61034.1| RGPR-p117 [Mus musculus] 135 1e-29
dbj|BAB67821.1| KIAA1928 protein [Homo sapiens] 132 6e-29
ref|NP_446023.1| regucalcin gene promotor region related protein... 131 1e-28
dbj|BAD12777.2| RGPR-p117 [Bos taurus] gi|46559376|ref|NP_996854... 131 1e-28
dbj|BAC03392.1| FLJ00305 protein [Homo sapiens] 127 3e-27
gb|AAH92165.1| LOC553356 protein [Danio rerio] 127 3e-27
>dbj|BAB09075.1| unnamed protein product [Arabidopsis thaliana]
gi|15238109|ref|NP_199560.1| expressed protein
[Arabidopsis thaliana]
Length = 1361
Score = 1173 bits (3035), Expect = 0.0
Identities = 691/1464 (47%), Positives = 887/1464 (60%), Gaps = 172/1464 (11%)
Query: 1 MASNPPFHVEDQDDEDFFDKLVEDDVGNVN------------DEANDSDDVKAFSNLSIG 48
MAS F +EDQ DEDFFDKLV+D D+ +DSDD++AFSNLSIG
Sbjct: 1 MASASQFLLEDQTDEDFFDKLVDDAYSPTEAQASSSVTELKFDDESDSDDIRAFSNLSIG 60
Query: 49 -----GDDADVNASAFEN--SSGGGSGGEGKERKEEGDVKLDGGNVQEGSSSGCDGMMDR 101
G D +N + N ++ G SG G++ E + + + R
Sbjct: 61 KDPLGGGDGTLNEAILGNDVANEGASGSVGED--EPSSIAPEAVQFPHSDARELRDDEMR 118
Query: 102 SDHGMESRNSSGSSADKSNRRSSLDVKEKDWNAFNVDS--NGGAGSESYSDFFSEFGDQN 159
S+ + + N VKE DW +F+ D N G G SYSDFF+E
Sbjct: 119 SEVADMPLSETAKECTIVNEPGIPGVKELDWGSFDADLSVNDGRGFGSYSDFFTELDATA 178
Query: 160 GKGYDHDLNTEVKHANEIPGDQYAQTYNRDSNTEVKL---GNEIPSDGMNASVDYVQYQE 216
G N +V + GN + +D N SV + Q+Q
Sbjct: 179 G--------------------------NLQGKADVAVATGGNLVANDTNNTSVGFEQHQG 212
Query: 217 GQSYDASARNSTSGEDVNSSQYWESLYPGWKYDYNTGQWYQVDEHNATAATQGSSE---- 272
+D S SG+ V++SQ WE+LYPGWKYD +TGQW+QVD H+A+ +Q S E
Sbjct: 213 QLHHD-----SASGQYVDNSQSWENLYPGWKYDASTGQWFQVDGHDASMNSQESYENSTS 267
Query: 273 ------VNTAEVSYMQQTAQSAVAGTLAESAATETVPSWNQVSQGNNGYPEHMIFDPQYP 326
N ++V+Y +Q+ SAVAGT+ E V +WNQVSQ +NGYPEHM+FD QYP
Sbjct: 268 NWENVAANNSDVAYQRQSTASAVAGTV------ENVSTWNQVSQVSNGYPEHMVFDSQYP 321
Query: 327 GWYYDTIAQEWRSLETYHSSIQYAVQGHG----NGHASSGTFSHNDNSLYRDYGQVGYYE 382
GWYYDTIAQEWRSL++Y+ + Q Q + NG++ + +++++ Y +
Sbjct: 322 GWYYDTIAQEWRSLDSYNQAFQTTGQANDQQVQNGNSFTAVDHSRESNVHDVYDKNQILR 381
Query: 383 SQGVGSQAANNNWSGSYGINHQQDLDRHTTDTA-------TKSGGSAYGGNQQFDHSFGS 435
+Q Q+ + +W SY +QQ + + A T + S GGNQQ ++ + +
Sbjct: 382 TQKFDIQSQHGSWDQSYYDKNQQATNMWQPENAGAAEAAVTPASLSNSGGNQQVNNLYST 441
Query: 436 SNSVNKNQQNASSSFGSVPLYNKVNHGHGLVNGTVEVQRFAPSGNFGQHYNYSNTQFDEQ 495
+ + S VQ F P QH N +N +
Sbjct: 442 GPVAEQFKPYESG-----------------------VQSFIP-----QHMNVANVTQNGP 473
Query: 496 KNISNDYAESHQPFGYSNQSYQSGHQQSYAPNVGRSSAGRPPHALVTFGFGGKLIILKDS 555
+ SN + + + QS+QS Q ++P+ GRSS GRPPHALV FGFGGKLI++KD
Sbjct: 474 MSFSNGFYSRQESVDDAPQSFQSS--QLFSPSAGRSSDGRPPHALVNFGFGGKLILMKDD 531
Query: 556 S--LSSSTYGSQ-GAAQGSVSVLNLMEAVSGSIGSSSIGNGAGDYFRALGQQSIPGPLVG 612
S L +S++GSQ G S+SVLNL E +SGS SS+G + YF L QQS+PGPLVG
Sbjct: 532 SGSLQNSSFGSQKGTGGSSISVLNLAEVISGSASYSSLGENSLSYFSCLDQQSLPGPLVG 591
Query: 613 GSVGSKELNKWIDERIAHCGSPDMDYKKSERMRLLLSLLKIACQYYGKLRSPFGTDTILK 672
G+VGSK+L+KW+DERI +C S MD+ + + +++LLSLL+I+CQYYGKLRSPFG+D + K
Sbjct: 592 GNVGSKDLHKWLDERILNCESSYMDFSRGKLLKMLLSLLRISCQYYGKLRSPFGSDALQK 651
Query: 673 DNDTPGSAVAKLFASAKMSGKE--YGVLSHCLQNLPSEAQMRATASEVQNLLVSGKKKEA 730
+ D+ +AVAKLFA AK G + Y +S CLQ+LP E+QM+ TASEVQNLL SG+K EA
Sbjct: 652 ETDSAEAAVAKLFAIAKEDGVQNGYAPISQCLQHLPPESQMQVTASEVQNLLASGRKMEA 711
Query: 731 LQYAQEGQLWGPALVLASQLGEKFYVDTVKQMALRQLVAGSPLRTLCLLIAGQPAEVFSS 790
LQ AQEG LWGPALV+A+QLG++FYVDTVKQMALRQLV GSPLRTLCLL+AGQPAEVFS+
Sbjct: 712 LQCAQEGHLWGPALVIAAQLGQQFYVDTVKQMALRQLVPGSPLRTLCLLVAGQPAEVFST 771
Query: 791 DS-SNSGDPSAFNMPQNPAQFGSSGMLDDWEENLAVITSNRTKDDELVIIHLGDCLWKER 849
S S+ P + N+P QFG S MLD WEENL +IT+NRT DDELVI HLGDC+WKER
Sbjct: 772 GSTSDISFPGSVNLPPQQPQFGCSSMLDSWEENLGIITANRTTDDELVITHLGDCMWKER 831
Query: 850 SEITAAHICYLIAEANFESYSDSARLCLIGADHWKFPRTYASPEAIQRTELYEYSKVLGN 909
EI AAHICYLIA+ NF++YSD+ARLCL+GADHWK+PRTYASPEAIQRTELYEYSK LGN
Sbjct: 832 GEIIAAHICYLIADKNFDTYSDTARLCLVGADHWKYPRTYASPEAIQRTELYEYSKTLGN 891
Query: 910 SQFILLPFQPYKLIYAYMLAEVGKVSDSLKYCQAVLKSLKTGRAPEVETWKQRLSSLEER 969
SQ+ LL FQPYK++YA+MLAEVGK+S + KYCQAVLK LKTGR+PEVE WKQ +SSLEER
Sbjct: 892 SQYTLLTFQPYKVMYAHMLAEVGKLSTAQKYCQAVLKCLKTGRSPEVEMWKQFVSSLEER 951
Query: 970 IRTHQQGGYAANLAPGKLVGKLLNFFDSTAHRVVGGLPPPAP-SSQGNVHGNE-QNYQSG 1027
IR HQQGGY ANL P KLVG LLNFF S HR VGG+PPPAP S++GN+ GNE Q+ Q
Sbjct: 952 IRIHQQGGYTANLHPEKLVGVLLNFFGSKTHRPVGGMPPPAPHSTKGNLQGNEYQHQQQE 1011
Query: 1028 AHRVSNSQSTMAMSSLVPSGSMEPNGEWTADNNRMTKSNRSVSEPDFGRSPRQETSHDAQ 1087
A +++ SQS MSSL+P S+EP E RM RSVSEPDFGR+P QE + ++
Sbjct: 1012 ATKLAYSQSVNTMSSLMPPASVEPTHESGGSGRRMAVHTRSVSEPDFGRTPIQEMADSSK 1071
Query: 1088 GKASEGT-----------SRFSRFSFGSQLLQKTMGLVLKPRPGKQAKLGEKNKFYYDEN 1136
KA +G SRFSRF FG + + T+G VL R K+AKLG +N+FYYD+
Sbjct: 1072 EKAVDGVTKLKSSGSVAGSRFSRFGFG--IFKDTVGRVL-ARSSKEAKLGAENQFYYDDK 1128
Query: 1137 LKRWVEEGAAPPAEETALPPPPTTAAFQNGLTEYNLQSALKTEGPPSKEGSDLKTSNPEL 1196
LKRWVE G PPAEE ALPPPPT AFQN Y +S + G + N
Sbjct: 1129 LKRWVERGVEPPAEEAALPPPPTIGAFQNNSLGYENKSDMIPSNGNWSSGGPTPSEN--- 1185
Query: 1197 TPGIPPIPPGTNHFSARGRVGIRSRYVDTFN-QGGGNSANLFQSPSVPSAKPVVAANAKF 1255
+ GIPPI G+N FSARGR G+R+RYVDT+N G GNS + SPSV +AKP + A AKF
Sbjct: 1186 SSGIPPISHGSNQFSARGRTGVRARYVDTYNPPGRGNSHTMIPSPSVQTAKPPIPAKAKF 1245
Query: 1256 FIP-TPAPSSNEQTME-AIEENNQEDDLAYENPSTSYRNDWSFQSPKHASASTWQRCPSM 1313
F+P PA SN+Q ME A E QE+ A E ++S QR PSM
Sbjct: 1246 FVPAAPASFSNDQAMEPAAAETRQEEISADEVVASS----------GAPPPMMMQRYPSM 1295
Query: 1314 GNFANHEAVVS--GSNSRSPHSRRTVSWGGSTDVTYSPTKMREIMPLGEALGMPPSTYMS 1371
N + +S G N + P SRRT SW G+ + +++P PST+
Sbjct: 1296 DNIQRNGLGISVNGDNHQPPTSRRTASWSGNFNTSFTPP-------------TSPSTFKP 1342
Query: 1372 DDISSMRTSMKSGNFGEDLHEVDL 1395
++S S + GE+L EV+L
Sbjct: 1343 VLLNS-----SSSSLGEELQEVEL 1361
>dbj|BAB09074.1| unnamed protein product [Arabidopsis thaliana]
gi|15238106|ref|NP_199559.1| expressed protein
[Arabidopsis thaliana]
Length = 1350
Score = 1138 bits (2943), Expect = 0.0
Identities = 678/1464 (46%), Positives = 886/1464 (60%), Gaps = 183/1464 (12%)
Query: 1 MASNPPFHVEDQDDEDFFDKLVEDDVGNVN---------DEANDSDDVKAFSNLSI---- 47
MAS F ++DQ DEDFFDKLV+D D+ +DSDD KAF+NLS+
Sbjct: 1 MASTADFLLDDQTDEDFFDKLVDDSYTPTASSSAKELKFDDGSDSDDAKAFANLSVVDDV 60
Query: 48 -GGDDADVNASAFEN--SSGGGSGGEGKERKEEGDVKLDGGNVQEGSSSGCDGMMDRSDH 104
G D +N + N ++ G SG GKE V ++ D + RS+
Sbjct: 61 LGDGDVALNEAGLGNDVANEGTSGSVGKEEPSSSIAPEAVQFVNSDANRLRDVDVVRSEV 120
Query: 105 GMESRNSSGSSADKSNRRSSLDVKEKDWNAFNVDS--NGGAGSESYSDFFSEFGDQNGKG 162
+ +G ++ + S VKE DW +F DS N G G SYSDFF+E G
Sbjct: 121 DDMALTETGKESNIVDGSGSPGVKEVDWGSFYADSSVNDGGGFGSYSDFFTELDATAG-- 178
Query: 163 YDHDLNTEVKHANEIPGDQYAQTYNRDSNTEVKLGNEIPSDGMNASVDYVQYQEGQSYDA 222
++ + + A G+ A N NT V L N S + Q+Q +D
Sbjct: 179 ---NVQGQAEVAVATGGNLVA---NDTINTSVGLDN---------SAGFEQHQGQVQHD- 222
Query: 223 SARNSTSGEDVNSSQYWESLYPGWKYDYNTGQWYQVDEHNATAATQGSSEVNT------- 275
S SG+ V++SQ WE+LYPGWKYD +TGQWYQVD +AT +Q S +T
Sbjct: 223 ----SGSGQYVDNSQSWENLYPGWKYDASTGQWYQVDGQDATVNSQESYINSTGNWESVA 278
Query: 276 ---AEVSYMQQTAQSAVAGTLAESAATETVPSWNQVSQGNNGYPEHMIFDPQYPGWYYDT 332
++V+Y++Q+ SA+AGT E+V +WNQVSQ NGYPEHM+FD QYPGWYYDT
Sbjct: 279 ADNSDVAYLKQSTTSAMAGT------AESVSTWNQVSQVGNGYPEHMVFDAQYPGWYYDT 332
Query: 333 IAQEWRSLETYHSSIQYAVQGHG------NGHASSGTFSHNDNSLYRDYGQVGY-YESQG 385
IAQEWRSL++Y+ + Q V G NGHA + T+ +N S D +++Q
Sbjct: 333 IAQEWRSLDSYNQASQTTVTGQAHDQQVQNGHARTTTYHNNSQSSVYDVNNKNQTFKAQD 392
Query: 386 VGSQAANNNWSGSYGINHQQDLDR-------HTTDTATKSGGSAYGGNQQFDHSFGSSNS 438
Q + +W SY N+QQ + T S +GGNQQ ++ + + +
Sbjct: 393 FAIQGQHGSWDESYYANNQQAGNTWQPVNVGKAEPAVTSDSLSRFGGNQQVNNLYSTESV 452
Query: 439 VNKNQQNASSSFGSVPLYNKVNHGHGLVNGTVEVQRFAPSGNFGQHYNYSNTQFDEQKNI 498
+ + N T+ Q F P QH N ++ + +
Sbjct: 453 AEQFKPN-----------------------TIGAQSFIP-----QHMNVASATQNGPLSF 484
Query: 499 SNDYAESHQPFGYSNQSYQSGHQQSYAPNVGRSSAGRPPHALVTFGFGGKLIILKDS--S 556
SND Q ++ +S+Q+ Q ++P+VGRSS RPPHALV+FGFGGKLI++KD+ S
Sbjct: 485 SNDLYNRQQSVDHAQKSFQNN--QLFSPSVGRSSDRRPPHALVSFGFGGKLIVMKDNNGS 542
Query: 557 LSSSTYGSQGAAQGSVSVLNLMEAVSGSIGSSSIGNGAGDYFRALGQQSIPGPLVGGSVG 616
L ++++GSQG S++VLNL E +SGS SS G + YFR L QQS+PGPLVGG+VG
Sbjct: 543 LQNTSFGSQGIGGSSITVLNLAEVISGSASYSSPGEDSLSYFRCLHQQSLPGPLVGGNVG 602
Query: 617 SKELNKWIDERIAHCGSPDMDYKKSERMRLLLSLLKIACQYYGKLRSPFGTDTILKDNDT 676
SKEL+KWIDER+ HC S +MD+ + + +++LLSLL+I+CQYYGKLRSPFG+D K+ DT
Sbjct: 603 SKELHKWIDERLLHCESSNMDFSRGKLLKMLLSLLRISCQYYGKLRSPFGSDASQKETDT 662
Query: 677 PGSAVAKLFASAKMSGKE--YGVLSHCLQNLPSEAQMRATASEVQNLLVSGKKKEALQYA 734
P +AVAKLFA AK G + Y +S CLQ+LP E+QM+ TASEVQNLL SG+K EALQ A
Sbjct: 663 PEAAVAKLFAFAKKDGIQNGYAPISQCLQHLPPESQMQVTASEVQNLLASGRKMEALQCA 722
Query: 735 QEGQLWGPALVLASQLGEKFYVDTVKQMALRQLVAGSPLRTLCLLIAGQPAEVFSSDSSN 794
QEG LWGPALV+A+QLG++FYVDTVKQMALRQL+ GSPLRTLCLL+AGQPAEV + SS+
Sbjct: 723 QEGHLWGPALVIAAQLGDQFYVDTVKQMALRQLIPGSPLRTLCLLVAGQPAEVCPTGSSS 782
Query: 795 SGDPSAFNMPQNPAQFGSSGMLDDWEENLAVITSNRTKDDELVIIHLGDCLWKERSEITA 854
S MLD+WEENL +IT+NRT DD+LVIIHLGD +WKER EI A
Sbjct: 783 S-------------------MLDNWEENLGIITANRTTDDDLVIIHLGDSMWKERGEIIA 823
Query: 855 AHICYLIAEANFESYSDSARLCLIGADHWKFPRTYASPEAIQRTELYEYSKVLGNSQFIL 914
AHICYLIA+ NF+ YS+SARLCL+GADHWK PRTYASP+AIQRTELYEYSK LGNSQ+IL
Sbjct: 824 AHICYLIADKNFDPYSESARLCLVGADHWKCPRTYASPDAIQRTELYEYSKTLGNSQYIL 883
Query: 915 LPFQPYKLIYAYMLAEVGKVSDSLKYCQAVLKSLKTGRAPEVETWKQRLSSLEERIRTHQ 974
LPFQPYK+IYA+MLAEVGK+S + KYCQAV++ LKT R+ EVE WKQ SSLEERIR+HQ
Sbjct: 884 LPFQPYKIIYAHMLAEVGKLSTAQKYCQAVIRCLKTSRSSEVEMWKQFASSLEERIRSHQ 943
Query: 975 QGGYAANLAPGKLVGKLLNFFDSTAHRVVGGLPPPAP-SSQGNVHGNE-QNYQSGAHRVS 1032
+GG NLAP KLVGKLLN + G+PPPAP S+ GN NE Q+ Q A ++S
Sbjct: 944 EGG---NLAPAKLVGKLLN--------SLWGMPPPAPHSTTGNPQVNEYQHQQQEAAKLS 992
Query: 1033 NSQSTMAMSSLVPSGSMEPNGEWTADNNRMTKSNRSVSEPDFGRSPRQETSHDAQGKASE 1092
SQS MSSL+P S+EP EW + M +RSVSEPDF R+P Q+ + ++ KA +
Sbjct: 993 YSQSANTMSSLMPPASIEPVHEWGGNGRTMAAHSRSVSEPDFSRTPIQDQTDSSKDKAPD 1052
Query: 1093 G-----------TSRFSRFSFGSQLLQKTMGLVLKPRPGKQAKLGEKNKFYYDENLKRWV 1141
G +SRFSRF G +L+ T+G V R +AKLG +N+FYYD+NLKRWV
Sbjct: 1053 GVTQVKSTRKVPSSRFSRFGIG--ILKNTVGKVFPSRSSNEAKLGNENQFYYDDNLKRWV 1110
Query: 1142 EEGAAPPAEETALPPPPTTAAFQNGLTEYNLQSALKTEGPPSKEGSDLKTSNP-ELTPGI 1200
E G PPAEE ALPPPPT+ F++ + +S +K E PS + P E +PGI
Sbjct: 1111 ERGVEPPAEEAALPPPPTSVPFRSNSLGHENKSEIKNEMSPSSGSWSSGSPTPSENSPGI 1170
Query: 1201 PPIPPGTNHFSARGRVGIRSRYVDTFNQGGGNSANLFQSPSVPSAKPVVAANAKFFIP-T 1259
PP+ G+N FSARGR+G+R+RYVDT+NQG S++++QSP V S+KP + A AKFF+P
Sbjct: 1171 PPVSQGSNQFSARGRMGVRARYVDTYNQG---SSSMYQSPPVQSSKPPIPAKAKFFVPAA 1227
Query: 1260 PAPSSNEQTMEAI----EENNQEDDLAYENPSTSYRNDWSFQSPKHASASTWQRCPSMGN 1315
PA +N+Q ME++ + N D+ + + SFQSP S QR PS+ N
Sbjct: 1228 PASFANDQVMESVSAETRQENSGDEAVVGSAGAPGPSQASFQSPT-PSPIAMQRFPSVDN 1286
Query: 1316 FANHEAVVSGSNSRSPH--SRRTVSWGGSTDVT--YSPTKMREIMPLGEALGMPPSTYMS 1371
+ S N P SRRT SW GS + + SPT ST+
Sbjct: 1287 IRRSGSGTS-LNGDLPQSVSRRTASWSGSVNSSSFMSPTS--------------ASTFRP 1331
Query: 1372 DDISSMRTSMKSGNFGEDLHEVDL 1395
++S S + GE+L EV+L
Sbjct: 1332 SPLNS-----SSSSLGEELQEVEL 1350
>ref|XP_550450.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|55295836|dbj|BAD67704.1| unknown protein [Oryza sativa
(japonica cultivar-group)]
Length = 577
Score = 520 bits (1340), Expect = e-145
Identities = 310/603 (51%), Positives = 380/603 (62%), Gaps = 48/603 (7%)
Query: 815 MLDDWEENLAVITSNRTKDDELVIIHLGDCLWKERSEITAAHICYLIAEANFESYSDSAR 874
MLDDW+ENLA+IT+NRTK D+LVI HLGDCLWKE++E+ AAH CYL AE N ESYS+S+R
Sbjct: 1 MLDDWQENLAIITANRTKGDDLVITHLGDCLWKEKNEVAAAHSCYLAAELNIESYSESSR 60
Query: 875 LCLIGADHWKFPRTYASPEAIQRTELYEYSKVLGNSQFILLPFQPYKLIYAYMLAEVGKV 934
+CLIGADH + PRT+ SPEAIQRTE+YEY+KVLGNSQ+ILLPFQPYKLIYAYML EVGKV
Sbjct: 61 MCLIGADHLRCPRTFTSPEAIQRTEVYEYAKVLGNSQYILLPFQPYKLIYAYMLVEVGKV 120
Query: 935 SDSLKYCQAVLKSLK-TGRAPEVETWKQRLSSLEERIRTHQQGGYAANLAPGKLVGKLLN 993
SDSL+YCQA LK LK +GRAPE+E WKQ SSLEERIRT+QQGGY NLAP KLVGKL
Sbjct: 121 SDSLRYCQACLKVLKASGRAPELEAWKQLFSSLEERIRTYQQGGYGTNLAPAKLVGKLFT 180
Query: 994 FFDSTAHRVVGGLPPP-APSSQGNVHGNEQNYQSGAHRVSNSQSTMAMSSLVPSGSMEPN 1052
D + R++G P +P QG++ + A N+Q MAMSSL+ S S +
Sbjct: 181 SLDKSLSRMMGTQPSALSPVPQGSLTERDSYSAPAATNFVNNQPVMAMSSLMSSVSEQSM 240
Query: 1053 GEWTADN--NRMTKSNRSVSEPDFGRSPRQETSHD-AQGKASEGTSRFSRFSFGSQLLQK 1109
E + + +R NRSVSEPDFGR+P Q D AQ + G+SRF LLQK
Sbjct: 241 SEMSGNTGPDRKVTHNRSVSEPDFGRTPNQGAGLDNAQSTSGSGSSRF------GWLLQK 294
Query: 1110 TMGLVLKPRPGKQAKLGEKNKFYYDENLKRWVEEGAAPPAEETALPPPPTTAAFQNGLTE 1169
TMGLV R QAKLGE+NKFYYDE LKRWVEEGA PAEE LPPPPT A FQNG+ +
Sbjct: 295 TMGLV--SRSHHQAKLGEQNKFYYDEKLKRWVEEGADIPAEEPPLPPPPTKALFQNGIPD 352
Query: 1170 Y--NLQSALKTEGPPSKEGSDLKTSNPELTPGIPPIPPGTNHFSARGRVGIRSRYVDTFN 1227
N ++ E L S P + G+PP+PP N FSARGR+G+RSRYVDTFN
Sbjct: 353 QSSNGPGSVSYTANGFSEARPLNPSGP--SSGMPPMPPSQNQFSARGRMGVRSRYVDTFN 410
Query: 1228 QGGGNSAN-LFQSPSVPSAKPVVAANAKFFIPTPAPSSNEQTMEAIEENNQEDDLAYENP 1286
+GG ++ + P+ PS P+ + A FF+PTPA ++EQ + +Q + P
Sbjct: 411 KGGASATGPSYNKPATPSMNPL--SGATFFVPTPATVASEQIPDPTVNVHQ------DQP 462
Query: 1287 STSYRNDWSFQSPKHA-----SASTWQRCPSMGNFANHEAVVSGSNSRSPHSR-RTVSWG 1340
S++ S SP + S QR PSM N SGS + S SR R SW
Sbjct: 463 SSTIALRESSASPPPSVQSIPVQSNIQRYPSMDNI----MTPSGSGNGSSFSRSRAASWS 518
Query: 1341 GS-------TDVTYSPTKMREIMPLGEALGMPPSTYMSDDISSMRTSMKSGN-FGEDLHE 1392
G+ V+ SP R +M + P S SS +S++ N GEDLHE
Sbjct: 519 GAYSEQLSGNAVSRSPDGQRTMM----QSPLIPGQKQSHSRSSSNSSLQFNNGLGEDLHE 574
Query: 1393 VDL 1395
V+L
Sbjct: 575 VEL 577
>ref|NP_910217.1| EST AU031036(E60617) corresponds to a region of the predicted
gene.~Similar to Human mRNA for KIAA0310 gene. (AB002308)
[Oryza sativa (japonica cultivar-group)]
Length = 1172
Score = 372 bits (954), Expect = e-101
Identities = 198/338 (58%), Positives = 242/338 (71%), Gaps = 12/338 (3%)
Query: 783 QPAEVFSSDSSNSGDPSAFNMPQNPAQ-FGSSGMLDDWEENLAVITSNRTKDDELVIIHL 841
QPA+VF++++ G+ ++PQ P + S MLDDW+ENLA+IT+NRTK D+LVI HL
Sbjct: 803 QPADVFNAENPVDGNYGNLHIPQRPVEAVNSKSMLDDWQENLAIITANRTKGDDLVITHL 862
Query: 842 GDCLWKERSEITAAHICYLIAEANFESYSDSARLCLIGADHWKFPRTYASPEAIQRTELY 901
GDCLWKE++E+ AAH CYL AE N ESYS+S+R+CLIGADH + PRT+ SPEAIQRTE+Y
Sbjct: 863 GDCLWKEKNEVAAAHSCYLAAELNIESYSESSRMCLIGADHLRCPRTFTSPEAIQRTEVY 922
Query: 902 EYSKVLGNSQFILLPFQPYKLIYAYMLAEVGKVSDSLKYCQAVLKSLK-TGRAPEVETWK 960
EY+KVLGNSQ+ILLPFQPYKLIYAYML EVGKVSDSL+YCQA LK LK +GRAPE+E WK
Sbjct: 923 EYAKVLGNSQYILLPFQPYKLIYAYMLVEVGKVSDSLRYCQACLKVLKASGRAPELEAWK 982
Query: 961 QRLSSLEERIRTHQQGGYAANLAPGKLVGKLLNFFDSTAHRVVGGLPPP-APSSQGNVHG 1019
Q SSLEERIRT+QQGGY NLAP KLVGKL D + R++G P +P QG++
Sbjct: 983 QLFSSLEERIRTYQQGGYGTNLAPAKLVGKLFTSLDKSLSRMMGTQPSALSPVPQGSLTE 1042
Query: 1020 NEQNYQSGAHRVSNSQSTMAMSSLVPSGSMEPNGEWTADN--NRMTKSNRSVSEPDFGRS 1077
+ A N+Q MAMSSL+ S S + E + + +R NRSVSEPDFGR+
Sbjct: 1043 RDSYSAPAATNFVNNQPVMAMSSLMSSVSEQSMSEMSGNTGPDRKVTHNRSVSEPDFGRT 1102
Query: 1078 PRQETSHD-AQGKASEGTSRFSRFSFGSQLLQKTMGLV 1114
P Q D AQ + G+SRF LLQKTMGLV
Sbjct: 1103 PNQGAGLDNAQSTSGSGSSRF------GWLLQKTMGLV 1134
Score = 278 bits (710), Expect = 1e-72
Identities = 217/710 (30%), Positives = 323/710 (44%), Gaps = 133/710 (18%)
Query: 127 VKEKDWNAFNVDSNGGAGSESYSDFFSEFGDQNGKGYDHDLNTEVKHANEIPG--DQYAQ 184
VK+ W+AF +S G G + + +F + G + A+ +P
Sbjct: 103 VKQVQWSAFATNSGVG-GDDPFGEFMGDDAFFGGNNTHTMTGDQPIQASLVPPTTSTIGS 161
Query: 185 TYNRDSNTEVKLGNEIPSDGMNAS--VDYVQYQEGQSYDASARNSTSGEDVNSSQYWESL 242
++ SN + + P A+ +D+ + S A+A +STS + +Y+ESL
Sbjct: 162 MDHQFSNQVDNIADSQPGWPATAAEFMDHNTSVQSDSTTAAAVDSTSTDP----KYFESL 217
Query: 243 YPGWKYDYNTGQWYQVD---------EHNATAATQGSSEVNTAE--------VSYMQQTA 285
YPGWKYD T QWYQVD ++ G V+ + VSY+ ++
Sbjct: 218 YPGWKYDEATQQWYQVDSSYTAQGNADNLGALPVVGGDNVHQQQQQQQQQFTVSYLHDSS 277
Query: 286 QSAVAGTLAESAATETVPSW--NQVSQGNNGYPEHMIFDPQYPGWYYDTIAQEWRSLETY 343
Q+ + T+AE +T SW N+ + G YP +M+F +YPGWY+DT Q+W SLE+Y
Sbjct: 278 QAGLE-TIAEEGSTMAT-SWTPNESNTGAVEYPSNMVFYAEYPGWYFDTNTQQWLSLESY 335
Query: 344 H-----------SSIQYAVQGHGNGHAS---SGTFSHN---------DNSLYRD-YGQVG 379
+S YA GH S +G +SH DNSL YG
Sbjct: 336 QQGGVQAETTAAASAGYAGTGHNVAQTSDSYTGDYSHQGQQQHGSLGDNSLSDSFYGSNQ 395
Query: 380 YYESQ-----------------------------------------GVGSQAANNNWSGS 398
+ E+Q G GS ++ +
Sbjct: 396 HTENQTAQQANVESLESSKYYHADINTYAHSTSQYTSSEDHQASYKGFGSSTSHQSVYKG 455
Query: 399 Y--GINHQQDLDRH----------TTDTATKSGGSAYGGNQQFD-------HSFGSSNSV 439
+ + HQ H T A G GNQ + H G
Sbjct: 456 FEPSVGHQSTFTSHHSGYNGSESSTVQQAAHQGFKPSTGNQNYKGFEPYSGHQLGYKGYD 515
Query: 440 NKNQQNASSSFGSVP-------LYNKVNHGHGLVNGTVEVQRFAPSGNFGQHYNYSNTQF 492
Q FG Y ++ + NG + Q P+ N Q +++
Sbjct: 516 YSTGQTGHQEFGPSTDSQANHVAYQQLPSHYSSFNGAAKPQDSVPTANMPQMQTRADS-- 573
Query: 493 DEQKNISNDYAESHQPFGYSNQSYQSGH----QQSYAPNVGRSSAGRPPHALVTFGFGGK 548
D N+ N+Y + ++ Q + S + Q Y+ + RSSAGRPPHALVTFGFGGK
Sbjct: 574 DGCMNLPNNYLSTGSSVNFAQQQFISSNALPQQFGYSSHEQRSSAGRPPHALVTFGFGGK 633
Query: 549 LIILKDSSLSSSTY--GSQGAAQGSVSVLNLMEAVSGSIGSSSIGNGAG-DYFRALGQQS 605
L++++++ S+ + G+QG G+VS+LN+ E VS + SI +G+ YF AL +Q
Sbjct: 634 LVVVRETISMSTNFDSGNQGNPCGTVSILNVSEIVSDRVDHPSIPSGSALSYFHALCRQP 693
Query: 606 IPGPLVGGSVGSKELNKWIDERIAHCGSPDMDYKKSERMRLLLSLLKIACQYYGKLRSPF 665
IPGPLVGGS +K++NKW+DE S +++ + +LL+SLLKI CQ+YGKLRSPF
Sbjct: 694 IPGPLVGGSAAAKDVNKWLDEITGGYDSSIREFQGGDDQKLLISLLKILCQHYGKLRSPF 753
Query: 666 GTDTILKDNDTPGSAVAKLFASAKMSGK---EYGVLSHCLQNLPSEAQMR 712
G+D + D P AV KLF+S K SG EYG + HC++N+PSE Q++
Sbjct: 754 GSDPSQEGIDGPEMAVTKLFSSCKSSGAHKGEYGAIVHCMKNIPSENQIQ 803
>ref|XP_646771.1| hypothetical protein DDB0191321 [Dictyostelium discoideum]
gi|60474012|gb|EAL71949.1| hypothetical protein
DDB0191321 [Dictyostelium discoideum]
Length = 1992
Score = 247 bits (631), Expect = 2e-63
Identities = 248/923 (26%), Positives = 381/923 (40%), Gaps = 125/923 (13%)
Query: 421 SAYGGNQQF-DHS---FGSSNSVNKNQQNASSSFGSVPLYNKVNHGHGLVNGTVEVQRFA 476
S +GGNQ +HS F + ++ N Q +++ + Y N G+ + NG Q+
Sbjct: 1082 SQFGGNQVIRNHSLPDFSNHFPISPNDQFNNNNNNN---YQGYNGGNSMFNGPPPQQQQQ 1138
Query: 477 PSGNFGQHYNYSNTQFDEQKNISNDYAESHQPFGYSNQSYQSG----HQQSYAPNVGRS- 531
Q Q +Q+ + P +N + + H +S + N +
Sbjct: 1139 QQQQQQQQQQQQQQQQQQQQQQQQQQQQGMMPINNNNNNNNNNNFFHHARSTSWNGNDAH 1198
Query: 532 SAGRP----PHALVTFGFGGKLIILKDSSLSSSTYGSQGAAQGSVSVLNLMEAVSGSIGS 587
G P PH + FGFGGK++ +K +S T S+ + + IG
Sbjct: 1199 QRGIPFYSRPHPVFAFGFGGKVVSIKSNSGIQVTQPDGTVKTSKYSIQ--ISPIKSLIGH 1256
Query: 588 SSIGNGAGDYFRALGQQSIPGPLVGGSVGSKE-LNKWIDERIAHCGSPDMDY---KKSER 643
++ N + PGPL S SK+ +NKWI ER+ S +D K SE
Sbjct: 1257 NTFYNNLSTF---------PGPLTSSS--SKDTINKWISERVHDIASGKIDQLCSKDSEA 1305
Query: 644 MRLLLSLLKIACQYYGKLRSPFGTDTILKDNDTPGSAVAKLFASAKMSGKEYGVLSHCLQ 703
+L LLKI C YG L + +D +VAK S K L Q
Sbjct: 1306 STILWELLKILCNNYGNL---------MNSSDKDDKSVAKSIQSLLSDNKTTPTLFQPTQ 1356
Query: 704 NL--PSEAQMRATASEVQNLLVSGKKKEALQYAQEGQLWGPALVLASQLGEKFYVDTVKQ 761
S+A + VQ+L+ G A Q+A + +LW ALVLA QLG + Y T+
Sbjct: 1357 KNIGRSDADSSLLLNRVQSLMTQGDIIAAHQFALDNELWSHALVLAHQLGAEVYGRTMSI 1416
Query: 762 MALRQLVAGSPLRTLCLLIAGQPAEVFSSDSSNSGDPSAFNMPQNPAQFGSSGMLDDWEE 821
L GSPL+TL +L +G ++F P N + LD W+E
Sbjct: 1417 FTNSSLPTGSPLKTLYILYSGNTRDLFKDI------PQGITNNNNNNNNNNGAFLDSWKE 1470
Query: 822 NLAVITSNRTKDDELVIIHLGDCLWKERSEITAAHICYLIAEANFESYSD-SARLCLIGA 880
N+A + N + VI +GDC+W+ + I AAHICYL+A+ F ++ +R+ L+G
Sbjct: 1471 NIATLLFNGKANGCKVIGEIGDCIWQVQKRIEAAHICYLLADYQFGPINNPGSRIVLLGG 1530
Query: 881 DHWKFPRTYASPEAIQRTELYEYSKVLGNSQFILLPFQPYKLIYAYMLAEVGKVSDSLKY 940
DH K + + + E++QR+E+YE+SK+LGNSQ+ L YK IYA L ++G V SLKY
Sbjct: 1531 DH-KIDKHFITSESLQRSEIYEFSKILGNSQYSLPIIFVYKFIYATYLCDLGLVDLSLKY 1589
Query: 941 CQAVLKSLKTGRAP------EVETWKQRLSSLEERIRTHQQGGYAANLAPGKLVGKLLNF 994
+ LK + +++T+ +R++S + G + ++
Sbjct: 1590 LNHIKHLLKNHQVSNPCFIYQLDTFAERITSFSSNNKLSASGSWIPSIF----------- 1638
Query: 995 FDSTAHRVVGGLPPPAPSSQGNVHGNEQNYQSGAHRVSNSQSTMAMSSLVPSGSMEPNGE 1054
+ +++GG P+ N N N + ST + P+
Sbjct: 1639 --TKGFKILGGSNRQVPTVGAN---NNNNEMISPPSIVTGSSTPISTPTKSQQQQPPSQP 1693
Query: 1055 WTADNNRMTK-SNRSVSEPDFGRSPR-QETSHDAQGK-------ASEGTSRFSRFSFGSQ 1105
+A K + S F P+ T++++ K S+ T+ S S
Sbjct: 1694 ISASKLPFKKDEDDEDSYSSFSTKPKASSTNNNSSNKVDSDLSSTSDNTASPSSPSKDEN 1753
Query: 1106 LLQKTMGLVL------KPRPGKQAKLGEKNKFYYDENLKRWVEEGAAPPAEETALPPPPT 1159
+ GL+ KP+ K+ L N FYY+E L +WVE G A + PPPP
Sbjct: 1754 SNETKSGLLAGLFWRKKPKT-KEINLDNNNDFYYNEELGKWVERGKEAEAIAASQPPPPP 1812
Query: 1160 -----TAAFQNGLTEYNLQSALKTEGPPSKEGSDLKTSNPELTPG-----------IPPI 1203
+ + +G++ PPS S T P G P
Sbjct: 1813 PTESYSPSHTDGMSTPPPFGGGGNASPPSVLSSSPSTRLPSGGGGGGSGSGGAPSSDPSQ 1872
Query: 1204 PPGTNHFSARGRVG----IRSRYVDTFNQ----GGGNSANLFQSPSVPSAKPVVAANAKF 1255
PPG N +S G + RYVDT NQ GG N+++ S S+P P
Sbjct: 1873 PPGVNKYSLVSPGGPNTRRKPRYVDTLNQTVYTGGVNNSS---SQSLPPKPPTSTFTP-- 1927
Query: 1256 FIPTPAPSSNEQTMEAIEENNQE 1278
F+P PS+ AI E Q+
Sbjct: 1928 FMPAFDPSA------AIVEQQQQ 1944
>dbj|BAD18068.1| RGPR-p117 [Rattus norvegicus]
Length = 1057
Score = 149 bits (376), Expect = 7e-34
Identities = 164/630 (26%), Positives = 270/630 (42%), Gaps = 92/630 (14%)
Query: 340 LETYHSSIQYAVQGHGNGHASSGTFSHNDNSLYRDYGQVGYYESQGVGSQAANNNWSGSY 399
L+ + S Y+ G+ + + S T + D Y +Y YY Q GS
Sbjct: 83 LKGSYPSHPYSRSGYEDPYQSYHTPTPRDEYAYGNY----YYHGHPQLPQGERVARQGSP 138
Query: 400 GINHQQDLD-RHTTDTATKSGGSAYGGNQQFDHSFGSSNSVNKNQQNASSSFGSVPLYNK 458
I H+ D RH ++ + S + N F S N N N +S+FG
Sbjct: 139 YIWHEDHRDQRHFSEHHWEKHNSTFEANSDTQFQFTSKNPYN---DNPASAFGLE----- 190
Query: 459 VNHGHGLVNGTVEVQRFAPSGNFGQHYNYSNTQFDEQKNISNDYAESHQPFGYSNQSYQS 518
G + Q+ + + SN + +S+ E Q + + Y+
Sbjct: 191 -QPGEFFPESGAQKQKPSLTSK-------SNLLQQHESGLSSSSYELSQYMTEAPEQYEP 242
Query: 519 GHQQSYAP-NVGRSSAGRP--------PHALVTFGFGGKLIILKDSSLSSSTYGSQGAAQ 569
++ P +SA P PH V+FG GG+LI + +S + Q
Sbjct: 243 MVSATWRPIQADDTSATVPKAPMRFYVPHVSVSFGPGGQLICVSPNSPADG--------Q 294
Query: 570 GSVSVLNLMEAVSGSIGSSSIGNGAGDYFRALGQQSIPGPLVGGSVGSKELNKWIDERIA 629
++ ++ ME + D+ ++ PGPL+ + ++ + ++ A
Sbjct: 295 TALVEVHSMEVI------------LNDFEDQEEMRAFPGPLIREDIHKVDIMTFCQKKAA 342
Query: 630 HCGSPDMDYKKSERMRLLLSLLKIACQYYGKLRSPFGTDTILKD-----NDTPGSAVAKL 684
C + + LL LL + C+ G + + +++D VA L
Sbjct: 343 QCLKSETPGSRDSA--LLWQLLVLLCRQNGSMVGSDIAELLMQDCKKLEKYKRQPPVANL 400
Query: 685 FASAKMSGKEYGVLSHCLQNL-----PSEAQMRATASE-VQNLLVSGKKKEALQYAQEGQ 738
++ +++ VLS +NL P A E NLL G+KKEAL++A +
Sbjct: 401 I---NLTDEDWPVLSSGTRNLLTGEIPLNVDTPAQIVEKFTNLLYYGRKKEALEWAMKNH 457
Query: 739 LWGPALVLASQLGEKFYVDTVKQMALRQLVAGSPLRTLCLLIAGQPAEVFSSDSSNSGDP 798
LWG AL LAS++ + Y + V L PL+TL L++G+ + ++ GD
Sbjct: 458 LWGHALFLASKMDPRTY-NWVMSGFTTTLALNDPLQTLFQLMSGRIPQA----ATCCGDK 512
Query: 799 SAFNMPQNPAQFGSSGMLDDWEENLAVITSNRTKDDEL---VIIHLGDCLWKERSEITAA 855
Q+G DW +LAVI SN+ D EL I+ +GD L + + A+
Sbjct: 513 ----------QWG------DWRPHLAVILSNQAGDAELYQRAIVSMGDTL-AGKGLVEAS 555
Query: 856 HICYLIAEANFESYS-DSARLCLIGADHWKFPRTYASPEAIQRTELYEYSKVLGNSQFIL 914
H CYL+A F Y+ + L L+G++H + +A+ EAIQRTE++EY ++LG + +
Sbjct: 556 HFCYLMAHVPFGHYTVKTDHLALVGSNHSQEFLKFATIEAIQRTEIFEYCQMLGRPKSFI 615
Query: 915 LPFQPYKLIYAYMLAEVGKVSDSLKYCQAV 944
FQ YKL+YA LA+ G S +L YC+A+
Sbjct: 616 PSFQVYKLLYASRLADYGLASQALHYCEAI 645
>dbj|BAD30088.1| RGPR-p117 [Oryctolagus cuniculus]
Length = 1045
Score = 143 bits (360), Expect = 5e-32
Identities = 129/427 (30%), Positives = 198/427 (46%), Gaps = 70/427 (16%)
Query: 537 PHALVTFGFGGKLIILKDSSLSSSTYGSQGAAQGSVSVLNLMEAVSGSIGSSSIGNGAGD 596
PH V FG G+L+ + SS Q ++ ++ ME + +
Sbjct: 274 PHVSVGFGPRGQLVCVSPSSPMDG--------QTALVEVHSMEVILNDLEEQE------- 318
Query: 597 YFRALGQQSIPGPLVGGSVGSKELNKWIDERIAHC---GSPDMDYKKSERMRLLLSLLKI 653
+S PGPL+ V ++ + ++ A G+P LL LL +
Sbjct: 319 -----EMRSFPGPLIREDVHKVDVMTFCQQKAAQSRKSGTPG-----GRDSALLWQLLVL 368
Query: 654 ACQYYGKLRSPFGTDTILKD-----NDTPGSAVAKLFASAKMSGKEYGVLSHCLQNL--- 705
C+ G + + +L+D VA L ++ +++ VLS +NL
Sbjct: 369 LCRQNGSMAGSDIAELLLQDCKKLEKYKKQPPVANLI---NLADEDWPVLSSGPRNLLTG 425
Query: 706 ---PSEAQMRATASEVQNLLVSGKKKEALQYAQEGQLWGPALVLASQLGEKFYVDTVKQM 762
PS + LL G+KKEAL++A + LWG AL L+S++ + Y V
Sbjct: 426 ETPPSVETPAQVVGKFTQLLYYGRKKEALEWAMKNHLWGHALFLSSKMEPRTY-KWVMSG 484
Query: 763 ALRQLVAGSPLRTLCLLIAGQPAEVFSSDSSNSGDPSAFNMPQNPAQFGSSGMLDDWEEN 822
L PL+TL L++G+ + ++ GD Q+G DW +
Sbjct: 485 FTSTLALNDPLQTLFQLLSGRIPQA----ATCCGD----------TQWG------DWRPH 524
Query: 823 LAVITSNRTKDDEL---VIIHLGDCLWKERSEITAAHICYLIAEANFESYSDSA-RLCLI 878
LAVI SN+ D EL VI+ +GD L + + AAH CYL+A+ F Y+ +A RL L+
Sbjct: 525 LAVILSNQAGDPELRRRVIVSMGDTL-ASKGLVEAAHFCYLMAQVPFGHYTVNADRLALL 583
Query: 879 GADH-WKFPRTYASPEAIQRTELYEYSKVLGNSQFILLPFQPYKLIYAYMLAEVGKVSDS 937
G+ H +FPR +AS EAIQRTE+ EY + LG ++ FQ YKL+YA LA+ G S +
Sbjct: 584 GSSHSQEFPR-FASAEAIQRTEVLEYCRTLGGRPGLIASFQVYKLLYASRLADYGLASQA 642
Query: 938 LKYCQAV 944
YC+A+
Sbjct: 643 FHYCEAI 649
>ref|NP_149118.1| regucalcin gene promotor region related protein [Homo sapiens]
gi|14517637|dbj|BAB61035.1| RGPR-p117 [Homo sapiens]
Length = 1060
Score = 142 bits (357), Expect = 1e-31
Identities = 163/626 (26%), Positives = 261/626 (41%), Gaps = 100/626 (15%)
Query: 349 YAVQGHGNGHASSGTFSHNDNSLYRDY---GQVGYYESQGVGSQAANNNWSGSYGINHQQ 405
Y+ G+ N + S + + + Y Y G + + + V Q + W Y Q+
Sbjct: 98 YSRPGYENSYQSYQSPTMREEYAYGSYYYHGHPQWLQEERVPRQRSPYIWHEDY--REQK 155
Query: 406 DLDRHTTDTATKSGGSAYGGNQQFDHSFGSSNSVNKNQQNASSSFGSVPLYNKVNHGHG- 464
LD H + + HS +NS Q N+ + P N G
Sbjct: 156 YLDEH---------------HYENQHSPFGTNSETHFQSNSRNPCKDSPASNSGQEWPGE 200
Query: 465 LVNGTV--EVQRFAPSGNFGQHYNYSNTQFDEQKNISNDYAESHQPFGYSNQSYQSGHQQ 522
L G++ E Q+ PS + SN + +S+ E Q + +
Sbjct: 201 LFPGSLLAEAQKNKPS-----LASESNLLQQRESGLSSSSYELSQYIRDAPERDDPPASA 255
Query: 523 SYAPNVGRSSAGRP--------PHALVTFGFGGKLIILKDSSLSSSTYGSQGAAQGSVSV 574
+++P S+ P PH V+FG GG+L+ + SS + Q ++
Sbjct: 256 AWSPVQADVSSAGPKAPMKFYIPHVPVSFGPGGQLVHVGPSSPTDG--------QAALVE 307
Query: 575 LNLMEAVSGSIGSSSIGNGAGDYFRALGQQSIPGPLVGGSVGSKELNKWIDERIAH-CGS 633
L+ ME + D +S GPL+ V ++ + ++ A C S
Sbjct: 308 LHSMEVI------------LNDSEEQEEMRSFSGPLIREDVHKVDIMTFCQQKAAQSCKS 355
Query: 634 PDMDYKKSERMRLLLSLLKIACQYYGKLRSPFGTDTILKD-----NDTPGSAVAKLFASA 688
+ + S LL LL + C+ G + + +++D VA L
Sbjct: 356 ETLGSRDSA---LLWQLLVLLCRQNGSMVGSDIAELLMQDCKKLEKYKRQPPVANLI--- 409
Query: 689 KMSGKEYGVLSHCLQNL------PSEAQMRATASEVQNLLVSGKKKEALQYAQEGQLWGP 742
++ +++ VLS NL PS + LL G+KKEAL++A + LWG
Sbjct: 410 NLTDEDWPVLSSGTPNLLTGEIPPSVETPAQIVEKFTRLLYYGRKKEALEWAMKNHLWGH 469
Query: 743 ALVLASQLGEKFYVDTVKQMALRQLVAGSPLRTLCLLIAGQPAEVFSSDSSNSGDPSAFN 802
AL L+S++ + Y V L PL+TL L++G+
Sbjct: 470 ALFLSSKMDPQTY-SWVMSGFTSTLALNDPLQTLFQLMSGR------------------- 509
Query: 803 MPQNPAQFGSSGMLDDWEENLAVITSNRTKDDEL---VIIHLGDCLWKERSEITAAHICY 859
+PQ G DW +LAVI SN+ D EL I+ +GD L + + AAH CY
Sbjct: 510 IPQAATCCGEK-QWGDWRPHLAVILSNQAGDPELYQRAIVAIGDTL-AGKGLVEAAHFCY 567
Query: 860 LIAEANFESYS-DSARLCLIGADHWKFPRTYASPEAIQRTELYEYSKVLGNSQFILLPFQ 918
L+A F Y+ + L L+G+ H + +A+ EAIQRTE++EY ++LG + + FQ
Sbjct: 568 LMAHVPFGHYTVKTDHLVLLGSSHSQEFLKFATTEAIQRTEIFEYCQMLGRPKSFIPSFQ 627
Query: 919 PYKLIYAYMLAEVGKVSDSLKYCQAV 944
YKL+YA LA+ G VS +L YC+A+
Sbjct: 628 VYKLLYASRLADYGLVSQALHYCEAI 653
>emb|CAI46016.1| hypothetical protein [Homo sapiens]
Length = 1061
Score = 140 bits (352), Expect = 4e-31
Identities = 165/627 (26%), Positives = 261/627 (41%), Gaps = 101/627 (16%)
Query: 349 YAVQGHGNGHASSGTFSHNDNSLYRDY---GQVGYYESQGVGSQAANNNWSGSYGINHQQ 405
Y+ G+ N + S + + + Y Y G + + V Q + W Y Q+
Sbjct: 98 YSRPGYENSYQSYQSPTMREEYAYGSYYYHGHPQWLREERVPRQRSPYIWHEDY--REQK 155
Query: 406 DLDRHTTDTATKSGGSAYGGNQQFDHSFGSSNSVNKNQQNASSSFGSVPLYNKVNHGHG- 464
LD H + + HS +NS Q N+ + P N G
Sbjct: 156 YLDEH---------------HYENQHSPFGTNSETHFQSNSRNPCKDSPASNSGQEWPGE 200
Query: 465 LVNGTV--EVQRFAPSGNFGQHYNYSNTQFDEQKNISNDYAESHQPFGYSNQSYQSGHQQ 522
L G++ E Q+ PS + SN + +S+ E Q + +
Sbjct: 201 LFPGSLLAEAQKNKPS-----LASESNLLQQRESGLSSSSYELSQYIRDAPERDDPPASA 255
Query: 523 SYAPNVGR--SSAGRP-------PHALVTFGFGGKLIILKDSSLSSSTYGSQGAAQGSVS 573
+++P SSAG PH V+FG GG+L+ + SS + Q ++
Sbjct: 256 AWSPVQAEDVSSAGPKAPMKFYIPHVPVSFGPGGQLVHVGPSSPTDG--------QAALV 307
Query: 574 VLNLMEAVSGSIGSSSIGNGAGDYFRALGQQSIPGPLVGGSVGSKELNKWIDERIAH-CG 632
L+ ME + D +S GPL+ V ++ + ++ A C
Sbjct: 308 ELHSMEVI------------LNDSEEQEEMRSFSGPLIREDVHKVDIMTFCQQKAAQSCK 355
Query: 633 SPDMDYKKSERMRLLLSLLKIACQYYGKLRSPFGTDTILKD-----NDTPGSAVAKLFAS 687
S + + S LL LL + C+ G + + +++D VA L
Sbjct: 356 SETLGSRDSA---LLWQLLVLLCRQNGSMVGSDIAELLMQDCKKLEKYKRQPPVANLI-- 410
Query: 688 AKMSGKEYGVLSHCLQNL------PSEAQMRATASEVQNLLVSGKKKEALQYAQEGQLWG 741
++ +++ VLS NL PS + LL G+KKEAL++A + LWG
Sbjct: 411 -NLTDEDWPVLSSGTPNLLTGEIPPSVETPAQIVEKFTRLLYYGRKKEALEWAMKNHLWG 469
Query: 742 PALVLASQLGEKFYVDTVKQMALRQLVAGSPLRTLCLLIAGQPAEVFSSDSSNSGDPSAF 801
AL L+S++ + Y V L PL+TL L++G+
Sbjct: 470 HALFLSSKMDPQTY-SWVMSGFTSTLALNDPLQTLFQLMSGR------------------ 510
Query: 802 NMPQNPAQFGSSGMLDDWEENLAVITSNRTKDDEL---VIIHLGDCLWKERSEITAAHIC 858
+PQ G DW +LAVI SN+ D EL I+ +GD L + + AAH C
Sbjct: 511 -IPQATTCCGEK-QWGDWRPHLAVILSNQAGDPELYQRAIVAIGDTL-AGKGLVEAAHFC 567
Query: 859 YLIAEANFESYS-DSARLCLIGADHWKFPRTYASPEAIQRTELYEYSKVLGNSQFILLPF 917
YL+A F Y+ + L L+G+ H + +A+ EAIQRTE++EY ++LG + + F
Sbjct: 568 YLMAHVPFGHYTVKTDHLVLLGSSHSQEFLKFATTEAIQRTEIFEYCQMLGRPKSFIPSF 627
Query: 918 QPYKLIYAYMLAEVGKVSDSLKYCQAV 944
Q YKL+YA LA+ G VS +L YC+A+
Sbjct: 628 QVYKLLYASRLADYGLVSQALHYCEAI 654
>dbj|BAD32580.1| mKIAA1928 protein [Mus musculus]
Length = 1064
Score = 138 bits (348), Expect = 1e-30
Identities = 146/585 (24%), Positives = 243/585 (40%), Gaps = 103/585 (17%)
Query: 511 YSNQSYQSGHQQSYAPNVGRSSAGRP-------------------PHALVTFGFGGKLII 551
Y Y + + Y P V S+A RP PH V+FG GG+L+
Sbjct: 240 YELSQYMTAAPEEYEPMV--SAAWRPIQADDTSATVPKAPMRFYVPHVSVSFGPGGQLVC 297
Query: 552 LKDSSLSSSTYGSQGAAQGSVSVLNLMEAVSGSIGSSSIGNGAGDYFRALGQQSIPGPLV 611
+ +S + Q ++ ++ ME + D+ ++ PGPL+
Sbjct: 298 VPPNSPADG--------QTALVEVHSMEVL------------LNDFEDQEEMRAFPGPLI 337
Query: 612 GGSVGSKELNKWIDERIAHCGSPDMDYKKSERMRLLLSLLKIACQYYGKLRSPFGTDTIL 671
+ ++ + ++ C + + LL LL + C+ G + + ++
Sbjct: 338 REDIHKVDIMTFCQQKATQCLKSETPGSRDSA--LLWQLLVLLCRQNGSMVGSDIAELLM 395
Query: 672 KD-----NDTPGSAVAKLFASAKMSGKEYGVLSHCLQNL------PSEAQMRATASEVQN 720
+D VA L ++ +++ VLS ++L P+ +
Sbjct: 396 QDCKKLEKYKRQPPVANLI---NLTDEDWPVLSSGTRDLLTGEIPPNVDTPAQIVEKFTK 452
Query: 721 LLVSGKKKEALQYAQEGQLWGPALVLASQLGEKFYVDTVKQMALRQLVAGSPLRTLCLLI 780
LL G+KKEAL++A + LWG AL LAS++ + Y + V L PL+TL L+
Sbjct: 453 LLYYGRKKEALEWAMKNHLWGHALFLASKMDPRTY-NWVMSGFTSTLALNDPLQTLFQLM 511
Query: 781 AGQPAEVFSSDSSNSGDPSAFNMPQNPAQFGSSGMLDDWEENLAVITSNRTKDDEL---V 837
+G+ +PQ G DW +LAVI SN+ D EL
Sbjct: 512 SGR-------------------IPQAATVCGDK-QWGDWRPHLAVILSNQAGDTELYQRA 551
Query: 838 IIHLGDCLWKERSEITAAHICYLIAEANFESYS-DSARLCLIGADHWKFPRTYASPEAIQ 896
I+ +GD L + + A+H CYL+A F Y+ + L L+G+ H + +A+ EAIQ
Sbjct: 552 IVSMGDTL-AGKGLVEASHFCYLMAHVPFGHYTVKTDHLALVGSSHSQEFMKFATIEAIQ 610
Query: 897 RTELYEYSKVLGNSQFILLPFQPYKLIYAYMLAEVGKVSDSLKYCQAVLKSLKTGRAPEV 956
RTE++EY ++LG + + FQ YKL+YA LA+ G S +L YC+A+ ++ +
Sbjct: 611 RTEIFEYCQMLGRPKSFIPSFQVYKLLYASRLADYGLASQALHYCEAIGAAVLSQEGSSH 670
Query: 957 ETWKQRLSSLEERIRTH-----QQGGYAANLAPGKLVGKLLNFFDSTAHRVVGGLPPPAP 1011
L L E+++ ++ +L P LV D +R G P
Sbjct: 671 PVLLAELIKLAEKLKLSDPLVLERRRGDRDLEPDWLVQLRRKHKDLEQNRT--GAPRDPD 728
Query: 1012 SSQGNVHG-------------NEQNYQSGAHRVSNSQSTMAMSSL 1043
S+ +++G QNY + S ST +SL
Sbjct: 729 STPSDIYGAGGTTDTPYPDLSGHQNYSEDSEYSSTLWSTAEQTSL 773
>ref|NP_203505.2| regucalcin gene promotor region related protein [Mus musculus]
gi|37589164|gb|AAH59194.1| Regucalcin gene promotor
region related protein [Mus musculus]
Length = 1051
Score = 137 bits (346), Expect = 2e-30
Identities = 125/468 (26%), Positives = 206/468 (43%), Gaps = 83/468 (17%)
Query: 511 YSNQSYQSGHQQSYAPNVGRSSAGRP-------------------PHALVTFGFGGKLII 551
Y Y + + Y P V S+A RP PH V+FG GG+L+
Sbjct: 227 YELSQYMTAAPEEYEPMV--SAAWRPIQADDTSATVPKAPMRFYVPHVSVSFGPGGQLVC 284
Query: 552 LKDSSLSSSTYGSQGAAQGSVSVLNLMEAVSGSIGSSSIGNGAGDYFRALGQQSIPGPLV 611
+ +S + Q ++ ++ ME + D+ ++ PGPL+
Sbjct: 285 VPPNSPADG--------QTALVEVHSMEVL------------LNDFEDQEEMRAFPGPLI 324
Query: 612 GGSVGSKELNKWIDERIAHCGSPDMDYKKSERMRLLLSLLKIACQYYGKLRSPFGTDTIL 671
+ ++ + ++ C + + LL LL + C+ G + + ++
Sbjct: 325 REDIHKVDIMTFCQQKATQCLKSETPGSRDSA--LLWQLLVLLCRQNGSMVGSDIAELLM 382
Query: 672 KD-----NDTPGSAVAKLFASAKMSGKEYGVLSHCLQNL------PSEAQMRATASEVQN 720
+D VA L ++ +++ VLS ++L P+ +
Sbjct: 383 QDCKKLEKYKRQPPVANLI---NLTDEDWPVLSSGTRDLLTGEIPPNVDTPAQIVEKFTK 439
Query: 721 LLVSGKKKEALQYAQEGQLWGPALVLASQLGEKFYVDTVKQMALRQLVAGSPLRTLCLLI 780
LL G+KKEAL++A + LWG AL LAS++ + Y + V L PL+TL L+
Sbjct: 440 LLYYGRKKEALEWAMKNHLWGHALFLASKMDPRTY-NWVMSGFTSTLALNDPLQTLFQLM 498
Query: 781 AGQPAEVFSSDSSNSGDPSAFNMPQNPAQFGSSGMLDDWEENLAVITSNRTKDDEL---V 837
+G+ +PQ G DW +LAVI SN+ D EL
Sbjct: 499 SGR-------------------IPQAATVCGDK-QWGDWRPHLAVILSNQAGDTELYQRA 538
Query: 838 IIHLGDCLWKERSEITAAHICYLIAEANFESYS-DSARLCLIGADHWKFPRTYASPEAIQ 896
I+ +GD L + + A+H CYL+A F Y+ + L L+G+ H + +A+ EAIQ
Sbjct: 539 IVSMGDTL-AGKGLVEASHFCYLMAHVPFGHYTVKTDHLALVGSSHSQEFMKFATIEAIQ 597
Query: 897 RTELYEYSKVLGNSQFILLPFQPYKLIYAYMLAEVGKVSDSLKYCQAV 944
RTE++EY ++LG + + FQ YKL+YA LA+ G S +L YC+A+
Sbjct: 598 RTEIFEYCQMLGRPKSFIPSFQVYKLLYASRLADYGLASQALHYCEAI 645
>emb|CAI16321.1| regucalcin gene promotor region related protein (RGPR) [Homo
sapiens] gi|56204433|emb|CAI22848.1| regucalcin gene
promotor region related protein (RGPR) [Homo sapiens]
Length = 1061
Score = 137 bits (344), Expect = 3e-30
Identities = 163/627 (25%), Positives = 260/627 (40%), Gaps = 101/627 (16%)
Query: 349 YAVQGHGNGHASSGTFSHNDNSLYRDY---GQVGYYESQGVGSQAANNNWSGSYGINHQQ 405
Y+ G+ N + S + + + Y Y G + + + V Q + W Y Q+
Sbjct: 98 YSRPGYENSYQSYQSPTMREEYAYGSYYYHGHPQWLQEERVPRQRSPYIWHEDY--REQK 155
Query: 406 DLDRHTTDTATKSGGSAYGGNQQFDHSFGSSNSVNKNQQNASSSFGSVPLYNKVNHGHG- 464
LD H + + HS +NS Q N+ + P N G
Sbjct: 156 YLDEH---------------HYENQHSPFGTNSETHFQSNSRNPCKDSPASNSGQEWPGE 200
Query: 465 LVNGTV--EVQRFAPSGNFGQHYNYSNTQFDEQKNISNDYAESHQPFGYSNQSYQSGHQQ 522
L G++ E Q+ PS + SN + +S+ E Q + +
Sbjct: 201 LFPGSLLAEAQKNKPS-----LASESNLLQQRESGLSSSSYELSQYIRDAPERDDPPASA 255
Query: 523 SYAPNVGR--SSAGRP-------PHALVTFGFGGKLIILKDSSLSSSTYGSQGAAQGSVS 573
+++P SSAG PH V+FG GG+L+ + SS + Q ++
Sbjct: 256 AWSPVQAEDVSSAGPKAPMKFYIPHVPVSFGPGGQLVHVGPSSPTDG--------QAALV 307
Query: 574 VLNLMEAVSGSIGSSSIGNGAGDYFRALGQQSIPGPLVGGSVGSKELNKWIDERIAH-CG 632
L+ ME + D +S GPL+ V ++ + ++ A C
Sbjct: 308 ELHSMEVI------------LNDSEEQEEMRSFSGPLIREDVHKVDIMTFCQQKAAQSCK 355
Query: 633 SPDMDYKKSERMRLLLSLLKIACQYYGKLRSPFGTDTILKD-----NDTPGSAVAKLFAS 687
S + + S LL LL + C+ G + + +++D VA L
Sbjct: 356 SETLGSRDSA---LLWQLLVLLCRQNGSMVGSDIAELLMQDCKKLEKYKRQPPVANLI-- 410
Query: 688 AKMSGKEYGVLSHCLQNL------PSEAQMRATASEVQNLLVSGKKKEALQYAQEGQLWG 741
++ +++ VLS NL PS + LL G+KK L++A + LWG
Sbjct: 411 -NLTDEDWPVLSSGTPNLLTGEIPPSVETPAQIVEKFTRLLYYGRKKATLEWAMKNHLWG 469
Query: 742 PALVLASQLGEKFYVDTVKQMALRQLVAGSPLRTLCLLIAGQPAEVFSSDSSNSGDPSAF 801
AL L+S++ + Y V L PL+TL L++G+
Sbjct: 470 HALFLSSKMDPQTY-SWVMSGFTSTLALNDPLQTLFQLMSGR------------------ 510
Query: 802 NMPQNPAQFGSSGMLDDWEENLAVITSNRTKDDEL---VIIHLGDCLWKERSEITAAHIC 858
+PQ G DW +LAVI SN+ D EL I+ +GD L + + AAH C
Sbjct: 511 -IPQAATCCGEK-QWGDWRPHLAVILSNQAGDPELYQRAIVAIGDTL-AGKGLVEAAHFC 567
Query: 859 YLIAEANFESYS-DSARLCLIGADHWKFPRTYASPEAIQRTELYEYSKVLGNSQFILLPF 917
YL+A F Y+ + L L+G+ H + +A+ EAIQRTE++EY ++LG + + F
Sbjct: 568 YLMAHVPFGHYTVKTDHLVLLGSSHSQEFLKFATTEAIQRTEIFEYCQMLGRPKSFIPSF 627
Query: 918 QPYKLIYAYMLAEVGKVSDSLKYCQAV 944
Q YKL+YA LA+ G VS +L YC+A+
Sbjct: 628 QVYKLLYASRLADYGLVSQALHYCEAI 654
>dbj|BAD34968.1| regucalcin gene promotor region related protein [Gallus gallus]
Length = 929
Score = 136 bits (342), Expect = 6e-30
Identities = 112/381 (29%), Positives = 178/381 (46%), Gaps = 40/381 (10%)
Query: 604 QSIPGPLVGGSVGSKELNKWIDERIAHCGSPDMDYKKSERMRLLLSLLKIACQYYGKLRS 663
Q+ PGPL + ++ + ++IA S D+ ++ LL LL + C+ G +
Sbjct: 282 QAFPGPLAREDLHKVDVMTFCQQKIA--SSCDLSTQRGRDSALLWKLLVLLCRQNGSMVG 339
Query: 664 PFGTDTILKD--------NDTPGSAVAKLFASAKMSGKEYGVLSHCLQNLPSEAQMRATA 715
+ +++D P L A ++ G L +P + +A
Sbjct: 340 SDVAELLMQDCRQQERYKRQEPAVGPVSL---ADEEWRQLGTLDLITGEVPPVVETQAQI 396
Query: 716 SE-VQNLLVSGKKKEALQYAQEGQLWGPALVLASQLGEKFYVDTVKQMALRQLVAGSPLR 774
E LL G+KK+AL +A QLWG AL L+S++ + Y V L PL+
Sbjct: 397 VEKFTKLLYYGRKKDALVWAMRNQLWGHALFLSSKMDPRTY-SWVLSGFTSTLATNDPLQ 455
Query: 775 TLCLLIAGQPAEVFSSDSSNSGDPSAFNMPQNPAQFGSSGMLDDWEENLAVITSNRTKDD 834
T L++G+ + S GD A++G DW +LAV+ SN+ D
Sbjct: 456 TFFQLMSGRIPQAAQS----CGD----------AKWG------DWRPHLAVLLSNKVGDM 495
Query: 835 EL---VIIHLGDCLWKERSEITAAHICYLIAEANFESYSDSA-RLCLIGADHWKFPRTYA 890
EL I+ +GD L + + AAH CYL+A+ F + A R+ L+G+ H + +A
Sbjct: 496 ELNHRAIVTMGDTL-AGKGAVEAAHFCYLMADIPFGYFGVKADRMALLGSSHRQAFTQFA 554
Query: 891 SPEAIQRTELYEYSKVLGNSQFILLPFQPYKLIYAYMLAEVGKVSDSLKYCQAVLKSLKT 950
+ EAIQR E++EY + L + LLPFQ YKL+YA LA+ G + +L YC+ + L
Sbjct: 555 TKEAIQRMEIFEYCQQLRHPTSFLLPFQVYKLLYASRLADHGLPAQALLYCEQIATVLLQ 614
Query: 951 GRAPEVETWKQRLSSLEERIR 971
Q+L+ L ER++
Sbjct: 615 QDPTSHPVLAQQLTKLAERLK 635
>ref|XP_514022.1| PREDICTED: hypothetical protein XP_514022 [Pan troglodytes]
Length = 1073
Score = 135 bits (341), Expect = 7e-30
Identities = 138/500 (27%), Positives = 215/500 (42%), Gaps = 102/500 (20%)
Query: 492 FDEQKNIS-NDYAESHQPFGYSNQSYQSGHQQ-----SYAPNVGR--------------- 530
+ EQK + + Y H PFG ++++Y + + S A N G+
Sbjct: 150 YREQKYLDEHHYENQHSPFGTNSETYFQSNSRNPCKDSPASNSGQEWPGELFPGSLLAEA 209
Query: 531 ------SSAGRP-------PHALVTFGFGGKLIILKDSSLSSSTYGSQGAAQGSVSVLNL 577
SSAG PH V+FG GG+L+ + SS + Q ++ L+
Sbjct: 210 QKNKDVSSAGPKAPMKFYIPHVPVSFGPGGQLVRVGPSSPTDG--------QAALVELHS 261
Query: 578 MEAVSGSIGSSSIGNGAGDYFRALGQQSIPGPLVGGSVGSKELNKWIDERIAH-CGSPDM 636
ME + D +S GPL+ V ++ + ++ A C S +
Sbjct: 262 MEVI------------LNDSEEQEEMRSFSGPLIREDVHKVDIMTFCQQKAAQSCKSETL 309
Query: 637 DYKKSERMRLLLSLLKIACQYYGKLRSPFGTDTILKD-----NDTPGSAVAKLFASAKMS 691
+ S LL LL + C+ G + + +++D VA L + ++
Sbjct: 310 GSRDSA---LLWQLLVLLCRQNGSMVGSDIAELLMQDCKKLEKYKRQPPVANLIS---LT 363
Query: 692 GKEYGVLSHCLQNL------PSEAQMRATASEVQNLLVSGKKKEALQYAQEGQLWGPALV 745
+++ VLS NL PS + LL G+KKEAL++A + LWG AL
Sbjct: 364 DEDWPVLSSGTPNLLTGEIPPSVETPAQIVEKFTRLLYYGRKKEALEWAMKNHLWGHALF 423
Query: 746 LASQLGEKFYVDTVKQMALRQLVAGSPLRTLCLLIAGQPAEVFSSDSSNSGDPSAFNMPQ 805
L+S++ Y V L PL+TL L++G+ +PQ
Sbjct: 424 LSSKMDPWTY-SWVMSGFTSTLALNDPLQTLFQLMSGR-------------------IPQ 463
Query: 806 NPAQFGSSGMLDDWEENLAVITSNRTKDDELVIIHLGDCLWKERSEITAAHICYLIAEAN 865
G DW +LAVI SN+ D EL + + AAH CYL+A
Sbjct: 464 AATCCGEK-QWGDWRPHLAVILSNQAGDPELY--------QPGKGLVEAAHFCYLVAHVP 514
Query: 866 FESYS-DSARLCLIGADHWKFPRTYASPEAIQRTELYEYSKVLGNSQFILLPFQPYKLIY 924
F Y+ + L L+G+ H + +A+ EAIQRTE++EY ++LG + + FQ YKL+Y
Sbjct: 515 FGHYTVKTDHLVLLGSSHSQEFLKFATTEAIQRTEIFEYCQMLGRPKSFIPSFQVYKLLY 574
Query: 925 AYMLAEVGKVSDSLKYCQAV 944
A LA+ G VS +L YC+A+
Sbjct: 575 ASRLADYGLVSQALHYCEAI 594
>dbj|BAB61034.1| RGPR-p117 [Mus musculus]
Length = 1051
Score = 135 bits (339), Expect = 1e-29
Identities = 124/468 (26%), Positives = 205/468 (43%), Gaps = 83/468 (17%)
Query: 511 YSNQSYQSGHQQSYAPNVGRSSAGRP-------------------PHALVTFGFGGKLII 551
Y Y + + Y P V S+A RP PH V+FG GG+L+
Sbjct: 227 YELSQYMTAAPEEYEPMV--SAAWRPIQADDTSATVPKAPMRFYVPHVSVSFGPGGQLVC 284
Query: 552 LKDSSLSSSTYGSQGAAQGSVSVLNLMEAVSGSIGSSSIGNGAGDYFRALGQQSIPGPLV 611
+ +S + Q ++ ++ ME + D+ ++ PG L+
Sbjct: 285 VSPNSPADG--------QTALVEVHSMEVL------------LNDFEDQEEMRAFPGALI 324
Query: 612 GGSVGSKELNKWIDERIAHCGSPDMDYKKSERMRLLLSLLKIACQYYGKLRSPFGTDTIL 671
+ ++ + ++ C + + LL LL + C+ G + + ++
Sbjct: 325 REDIPKVDIMTFCQQKATQCLKSETPGSRDSA--LLWQLLVLLCRQNGSMVGSDIAELLM 382
Query: 672 KD-----NDTPGSAVAKLFASAKMSGKEYGVLSHCLQNL------PSEAQMRATASEVQN 720
+D VA L ++ +++ VLS ++L P+ +
Sbjct: 383 QDCKKLEKYKRQPPVANLI---NLTDEDWPVLSSGTRDLLTGEIPPNVDTPAQIVEKFTK 439
Query: 721 LLVSGKKKEALQYAQEGQLWGPALVLASQLGEKFYVDTVKQMALRQLVAGSPLRTLCLLI 780
LL G+KKEAL++A + LWG AL LAS++ + Y + V L PL+TL L+
Sbjct: 440 LLYYGRKKEALEWAMKNHLWGHALFLASKMDPRTY-NWVMSGFTSTLALNDPLQTLFQLM 498
Query: 781 AGQPAEVFSSDSSNSGDPSAFNMPQNPAQFGSSGMLDDWEENLAVITSNRTKDDEL---V 837
+G+ +PQ G DW +LAVI SN+ D EL
Sbjct: 499 SGR-------------------IPQAATVCGDK-QWGDWRPHLAVILSNQAGDTELYQRA 538
Query: 838 IIHLGDCLWKERSEITAAHICYLIAEANFESYS-DSARLCLIGADHWKFPRTYASPEAIQ 896
I+ +GD L + + A+H CYL+A F Y+ + L L+G+ H + +A+ EAIQ
Sbjct: 539 IVSMGDTL-AGKGLVEASHFCYLMAHVPFGHYTVKTDHLALVGSSHSQEFMKFATIEAIQ 597
Query: 897 RTELYEYSKVLGNSQFILLPFQPYKLIYAYMLAEVGKVSDSLKYCQAV 944
RTE++EY ++LG + + FQ YKL+YA LA+ G S +L YC+A+
Sbjct: 598 RTEIFEYCQMLGRPKSFIPSFQVYKLLYASRLADYGLASQALHYCEAI 645
>dbj|BAB67821.1| KIAA1928 protein [Homo sapiens]
Length = 757
Score = 132 bits (333), Expect = 6e-29
Identities = 110/357 (30%), Positives = 170/357 (46%), Gaps = 44/357 (12%)
Query: 604 QSIPGPLVGGSVGSKELNKWIDERIAH-CGSPDMDYKKSERMRLLLSLLKIACQYYGKLR 662
+S GPL+ V ++ + ++ A C S + + S LL LL + C+ G +
Sbjct: 22 RSFSGPLIREDVHKVDIMTFCQQKAAQSCKSETLGSRDSA---LLWQLLVLLCRQNGSMV 78
Query: 663 SPFGTDTILKD-----NDTPGSAVAKLFASAKMSGKEYGVLSHCLQNL------PSEAQM 711
+ +++D VA L ++ +++ VLS NL PS
Sbjct: 79 GSDIAELLMQDCKKLEKYKRQPPVANLI---NLTDEDWPVLSSGTPNLLTGEIPPSVETP 135
Query: 712 RATASEVQNLLVSGKKKEALQYAQEGQLWGPALVLASQLGEKFYVDTVKQMALRQLVAGS 771
+ LL G+KKEAL++A + LWG AL L+S++ + Y V L
Sbjct: 136 AQIVEKFTRLLYYGRKKEALEWAMKNHLWGHALFLSSKMDPQTY-SWVMSGFTSTLALND 194
Query: 772 PLRTLCLLIAGQPAEVFSSDSSNSGDPSAFNMPQNPAQFGSSGMLDDWEENLAVITSNRT 831
PL+TL L++G+ +PQ G DW +LAVI SN+
Sbjct: 195 PLQTLFQLMSGR-------------------IPQAATCCGEK-QWGDWRPHLAVILSNQA 234
Query: 832 KDDEL---VIIHLGDCLWKERSEITAAHICYLIAEANFESYS-DSARLCLIGADHWKFPR 887
D EL I+ +GD L + + AAH CYL+A F Y+ + L L+G+ H +
Sbjct: 235 GDPELYQRAIVAIGDTL-AGKGLVEAAHFCYLMAHVPFGHYTVKTDHLVLLGSSHSQEFL 293
Query: 888 TYASPEAIQRTELYEYSKVLGNSQFILLPFQPYKLIYAYMLAEVGKVSDSLKYCQAV 944
+A+ EAIQRTE++EY ++LG + + FQ YKL+YA LA+ G VS +L YC+A+
Sbjct: 294 KFATTEAIQRTEIFEYCQMLGRPKSFIPSFQVYKLLYASRLADYGLVSQALHYCEAI 350
>ref|NP_446023.1| regucalcin gene promotor region related protein [Rattus norvegicus]
gi|13785516|dbj|BAB43905.1| RGPR-p117 [Rattus
norvegicus]
Length = 1058
Score = 131 bits (330), Expect = 1e-28
Identities = 158/631 (25%), Positives = 263/631 (41%), Gaps = 93/631 (14%)
Query: 340 LETYHSSIQYAVQGHGNGHASSGTFSHNDNSLYRDYGQVGYYESQGVGSQAANNNWSGSY 399
L+ + S Y+ G+ + + S T + D Y +Y YY Q GS
Sbjct: 83 LKGSYPSHPYSRSGYEDPYQSYHTPTPRDEYAYGNY----YYHGHPQLPQGERVARQGSP 138
Query: 400 GINHQQDLD-RHTTDTATKSGGSAYGGNQQFDHSFGSSNSVNKNQQNASSSFGSVPLYNK 458
I H+ D RH ++ + S + N F S N N N +S+FG
Sbjct: 139 YIWHEDHRDQRHFSEHHWEKHNSTFEANSDTQFQFTSKNPYN---DNPASAFGLE----- 190
Query: 459 VNHGHGLVNGTVEVQRFAPSGNFGQHYNYSNTQFDEQKNISNDYAESHQPFGYSNQSYQS 518
G + Q+ + + SN + +S+ E Q + + Y+
Sbjct: 191 -QPGEFFPESGAQKQKPSLTSK-------SNLLRQHESGLSSSSYELSQYMTEAPEQYEP 242
Query: 519 GHQQSYAP-NVGRSSAGRP--------PHALVTFGFGGKLIILKDSSLSSSTYGSQGAAQ 569
++ P +SA P PH V+FG GG+LI + +S + Q
Sbjct: 243 MVSATWRPIQADDTSATVPKAPMRFYVPHVSVSFGPGGQLICVSPNSPADG--------Q 294
Query: 570 GSVSVLNLMEAVSGSIGSSSIGNGAGDYFRALGQQSIPGPLVGGSVGSKELNKWIDERIA 629
++ ++ ME + D+ ++ PGPL+ + ++ + ++ A
Sbjct: 295 TALVEVHSMEVI------------LNDFEDQEEMRAFPGPLIREDIHKVDIMTFCQKKAA 342
Query: 630 HCGSPDMDYKKSERMRLLLSLLKIACQYYGKLRSPFGTDTILKD-----NDTPGSAVAKL 684
C + + LL LL + C+ G + + +++D VA L
Sbjct: 343 QCLKSETPGSRDSA--LLWQLLVLLCRQNGSMVGSDIAELLMQDCKKLEKYKRQPPVANL 400
Query: 685 FASAKMSGKEYGVLSHCLQNL------PSEAQMRATASEVQNLLVSGKKKEALQYAQEGQ 738
++ +++ VLS +NL P+ + LL G+KKE L E
Sbjct: 401 I---NLTDEDWPVLSSGTRNLLTGEIPPNVDTPAQIVEKFTKLLYYGRKKEGLGMGYEEP 457
Query: 739 LWGPA-LVLASQLGEKFYVDTVKQMALRQLVAGSPLRTLCLLIAGQPAEVFSSDSSNSGD 797
L GP LAS++ + Y + V L PL+TL L++G+ + ++ GD
Sbjct: 458 LVGPMPCSLASKMDPRTY-NWVMSGFTTTLALNDPLQTLFQLMSGRIPQA----ATCCGD 512
Query: 798 PSAFNMPQNPAQFGSSGMLDDWEENLAVITSNRTKDDEL---VIIHLGDCLWKERSEITA 854
Q+G DW +LAVI SN+ D EL I+ +GD L + + A
Sbjct: 513 K----------QWG------DWRPHLAVILSNQAGDAELYQRAIVSMGDTL-AGKGLVEA 555
Query: 855 AHICYLIAEANFESYS-DSARLCLIGADHWKFPRTYASPEAIQRTELYEYSKVLGNSQFI 913
+H CYL+A F Y+ + L L+G++H + +A+ EAIQRTE++EY ++LG +
Sbjct: 556 SHFCYLMAHVPFGHYTVKTDHLALVGSNHSQEFLKFATIEAIQRTEIFEYCQMLGRPKSF 615
Query: 914 LLPFQPYKLIYAYMLAEVGKVSDSLKYCQAV 944
+ FQ YKL+YA LA+ G S +L YC+A+
Sbjct: 616 IPSFQVYKLLYASRLADYGLASQALHYCEAI 646
>dbj|BAD12777.2| RGPR-p117 [Bos taurus] gi|46559376|ref|NP_996854.2| regucalcin gene
promotor region related protein [Bos taurus]
Length = 1052
Score = 131 bits (330), Expect = 1e-28
Identities = 128/450 (28%), Positives = 203/450 (44%), Gaps = 62/450 (13%)
Query: 537 PHALVTFGFGGKLIILKDSSLSSSTYGSQGAAQGSVSVLNLMEAVSGSIGSSSIGNGAGD 596
PH V+FG GG+L+ + SS S Q ++ L+ ME + D
Sbjct: 278 PHMPVSFGPGGQLVCVSPSSPSDG--------QTALVELHSMEVI------------LND 317
Query: 597 YFRALGQQSIPGPLVGGSVGSKELNKWIDERIAHCGSPDMDYKKSERMRLLLSLLKIACQ 656
++ GPL+ V ++ + ++ A S + S LL LL + C+
Sbjct: 318 SEEQEEMRTFSGPLIREDVHKVDIMTFCQQKAAQ--SHKSETPGSRDSALLWQLLVLLCR 375
Query: 657 YYGKLRSPFGTDTILKD-----NDTPGSAVAKLFASAKMSGKEYGVLSHCLQNL------ 705
G + + +++D VA L ++ +++ VLS ++L
Sbjct: 376 QNGSMVGSDIAELLMQDWKKLEKYKRQPPVANLI---NLTDEDWPVLSSGTRDLLTGEIL 432
Query: 706 PSEAQMRATASEVQNLLVSGKKKEALQYAQEGQLWGPALVLASQLGEKFYVDTVKQMALR 765
PS + LL G+KKEAL++A + LWG AL LAS++ + + V
Sbjct: 433 PSVETPAQILEKFTKLLYYGRKKEALEWAMKNHLWGHALFLASKMDPRTH-SRVMSGFTS 491
Query: 766 QLVAGSPLRTLCLLIAGQPAEVFSSDSSNSGDPSAFNMPQNPAQFGSSGMLDDWEENLAV 825
L PL+TL L++G+ + ++ GD Q+G DW +LAV
Sbjct: 492 TLALNDPLQTLFQLMSGRIPQA----ATCCGDK----------QWG------DWRPHLAV 531
Query: 826 ITSNRTKDDEL---VIIHLGDCLWKERSEITAAHICYLIAEANFESYS-DSARLCLIGAD 881
I SN+ D EL I+ +GD L + + AAH CYL+A F Y+ + L L+G+
Sbjct: 532 ILSNQGGDPELYQRTIVAMGDTL-AGKGLVEAAHFCYLMAHVPFGYYTVKTDYLALLGSS 590
Query: 882 HWKFPRTYASPEAIQRTELYEYSKVLGNSQFILLPFQPYKLIYAYMLAEVGKVSDSLKYC 941
H + A+ EAIQRTE++EY ++LG + + FQ YKL+YA LA+ G S +L YC
Sbjct: 591 HSQEFLKSATTEAIQRTEIFEYCQMLGRPKSFIPSFQVYKLLYASRLADYGLTSQALHYC 650
Query: 942 QAVLKSLKTGRAPEVETWKQRLSSLEERIR 971
+ V +L + L L ER++
Sbjct: 651 EPVGTALLSHGESSHPVLLVELIKLAERLK 680
>dbj|BAC03392.1| FLJ00305 protein [Homo sapiens]
Length = 789
Score = 127 bits (319), Expect = 3e-27
Identities = 101/314 (32%), Positives = 152/314 (48%), Gaps = 40/314 (12%)
Query: 646 LLLSLLKIACQYYGKLRSPFGTDTILKD-----NDTPGSAVAKLFASAKMSGKEYGVLSH 700
LL LL + C+ G + + +++D VA L ++ +++ VLS
Sbjct: 80 LLWQLLVLLCRQNGSMVGSDIAELLMQDCKKLEKYKRQPPVANLI---NLTDEDWPVLSS 136
Query: 701 CLQNL------PSEAQMRATASEVQNLLVSGKKKEALQYAQEGQLWGPALVLASQLGEKF 754
NL PS + LL G+KKEAL++A + LWG AL L+S++ +
Sbjct: 137 GTPNLLTGEIPPSVETPAQIVEKFTRLLYYGRKKEALEWAMKNHLWGHALFLSSKMDPQT 196
Query: 755 YVDTVKQMALRQLVAGSPLRTLCLLIAGQPAEVFSSDSSNSGDPSAFNMPQNPAQFGSSG 814
Y V L PL+TL L++G+ +PQ G
Sbjct: 197 Y-SWVMSGFTSTLALNDPLQTLFQLMSGR-------------------IPQAATCCGEK- 235
Query: 815 MLDDWEENLAVITSNRTKDDEL---VIIHLGDCLWKERSEITAAHICYLIAEANFESYS- 870
DW +LAVI SN+ D EL I+ +GD L + + AAH CYL+A F Y+
Sbjct: 236 QWGDWRPHLAVILSNQAGDPELYQRAIVAIGDTL-AGKGLVEAAHFCYLMAHVPFGHYTV 294
Query: 871 DSARLCLIGADHWKFPRTYASPEAIQRTELYEYSKVLGNSQFILLPFQPYKLIYAYMLAE 930
+ L L+G+ H + +A+ EAIQRTE++EY ++LG + + FQ YKL+YA LA+
Sbjct: 295 KTDHLVLLGSSHSQEFLKFATTEAIQRTEIFEYCQMLGRPKSFIPSFQVYKLLYASRLAD 354
Query: 931 VGKVSDSLKYCQAV 944
G VS +L YC+A+
Sbjct: 355 YGLVSQALHYCEAI 368
>gb|AAH92165.1| LOC553356 protein [Danio rerio]
Length = 501
Score = 127 bits (318), Expect = 3e-27
Identities = 90/241 (37%), Positives = 125/241 (51%), Gaps = 28/241 (11%)
Query: 721 LLVSGKKKEALQYAQEGQLWGPALVLASQLGEKFYVDTVKQMALRQLVAGSPLRTLCLLI 780
LL+SG+KKEAL+ A + LWG AL LAS++ + Y +TV L PL+TL L+
Sbjct: 54 LLLSGRKKEALESAMQSGLWGHALFLASKMDSRSY-NTVLSRFTGSLTPSDPLQTLFQLL 112
Query: 781 AGQ-PAEVFSSDSSNSGDPSAFNMPQNPAQFGSSGMLDDWEENLAVITSNRTKDDEL--- 836
+G+ PA S S GD W +LAV+ SN T D +
Sbjct: 113 SGRIPAVSMSWSSEKWGD---------------------WRSHLAVMLSNETTDPVIHHR 151
Query: 837 VIIHLGDCLWKERSEITAAHICYLIAEANFESYSD-SARLCLIGADHWKFPRTYASPEAI 895
II +GD L R + A+HICYL A+ F Y + S RL L+G+ + AI
Sbjct: 152 SIIAMGDTL-ASRDMLFASHICYLTAQCPFGWYRNRSERLVLLGSKQSVMFSKFCHNSAI 210
Query: 896 QRTELYEYSKVLGNSQFILLPFQPYKLIYAYMLAEVGKVSDSLKYCQAVLKSLKTGRAPE 955
+ TEL+EY + LGN++F++ FQ YK +YA L + G VS + YC+ V K+L P
Sbjct: 211 RCTELFEYCQRLGNAEFVIPAFQVYKFLYACRLLDCGLVSQAFHYCEVVGKALLRIDQPH 270
Query: 956 V 956
V
Sbjct: 271 V 271
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.309 0.128 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,614,034,579
Number of Sequences: 2540612
Number of extensions: 122685456
Number of successful extensions: 385028
Number of sequences better than 10.0: 1628
Number of HSP's better than 10.0 without gapping: 175
Number of HSP's successfully gapped in prelim test: 1553
Number of HSP's that attempted gapping in prelim test: 358731
Number of HSP's gapped (non-prelim): 11463
length of query: 1399
length of database: 863,360,394
effective HSP length: 141
effective length of query: 1258
effective length of database: 505,134,102
effective search space: 635458700316
effective search space used: 635458700316
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 82 (36.2 bits)
Medicago: description of AC138087.1


