

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138063.5 + phase: 0 /pseudo
(840 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
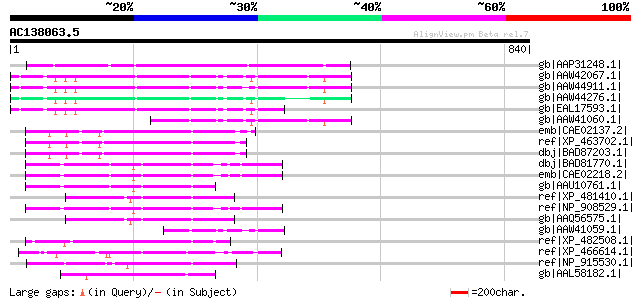
Score E
Sequences producing significant alignments: (bits) Value
gb|AAP31248.1| transposase [Fusarium oxysporum f. sp. melonis] 209 3e-52
gb|AAW42067.1| conserved hypothetical protein [Cryptococcus neof... 138 9e-31
gb|AAW44911.1| conserved hypothetical protein [Cryptococcus neof... 137 2e-30
gb|AAW44276.1| conserved hypothetical protein [Cryptococcus neof... 128 9e-28
gb|EAL17593.1| hypothetical protein CNBM0450 [Cryptococcus neofo... 126 3e-27
gb|AAW41060.1| conserved hypothetical protein [Cryptococcus neof... 125 4e-27
emb|CAE02137.2| OSJNBa0074L08.5 [Oryza sativa (japonica cultivar... 105 6e-21
ref|XP_463702.1| putative far-red impaired response protein [Ory... 104 1e-20
dbj|BAD87203.1| far-red impaired response-like [Oryza sativa (ja... 104 1e-20
dbj|BAD81770.1| putative far-red impaired response protein [Oryz... 92 5e-17
emb|CAE02218.2| OSJNBb0002N06.8 [Oryza sativa (japonica cultivar... 91 1e-16
gb|AAU10761.1| unknown protein [Oryza sativa (japonica cultivar-... 91 2e-16
ref|XP_481410.1| far-red impaired response protein -like [Oryza ... 91 2e-16
ref|NP_908529.1| P0702D12.15 [Oryza sativa (japonica cultivar-gr... 91 2e-16
gb|AAQ56575.1| putative transposase [Oryza sativa (japonica cult... 91 2e-16
gb|AAW41059.1| hypothetical protein CNA05530 [Cryptococcus neofo... 89 8e-16
ref|XP_482508.1| putative far-red impaired response protein [Ory... 87 3e-15
ref|XP_466614.1| putative far-red impaired response protein [Ory... 83 3e-14
ref|NP_915530.1| P0529E05.9 [Oryza sativa (japonica cultivar-gro... 83 3e-14
gb|AAL58182.1| putative far-red impaired response protein [Oryza... 82 7e-14
>gb|AAP31248.1| transposase [Fusarium oxysporum f. sp. melonis]
Length = 836
Score = 209 bits (532), Expect = 3e-52
Identities = 158/536 (29%), Positives = 259/536 (47%), Gaps = 24/536 (4%)
Query: 28 YEDREEMINWARGQAIEAGFTLIIDKSYSGSGRRKPKLVLAFERSGEYKGTKKSKREGTG 87
Y REE+ + A G+ +I +S S + + ++ +R G + S R T
Sbjct: 21 YSSREELRSAINAWAAPRGYAFVIKRS-SKTANGRTHVIFNCDR-GAGRIPSLSDRRQTT 78
Query: 88 SRKCGCPFRLRGYFSATK-LWSLNIVSGLH--NHKMEPKLE--GHMLAGRLTAKESKIVG 142
+R+ GC F + S K +WSL G H H EP H +L+ +E V
Sbjct: 79 TRRTGCLFSVLAKESLCKTIWSLRHRPGPHFSQHNHEPSFSEVAHPTLRQLSRQEEITVN 138
Query: 143 DMTRNLIKPKNILLDLKGRWKDNRTVAKQ--IYNARHRYKLSIRGSR*EMQELFQKFEEN 200
+T I PK I L+ + T+A Q IYN + + + + + L + E
Sbjct: 139 QLTNAGIAPKEIGSFLR---ITSNTLATQQDIYNCIAKGRRDLSKGQSNIHALADQLNEE 195
Query: 201 NYVFNYRTVSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQYKMPLFEIVGLTST 260
+ +N + S V + FAHP+S++ T+P VL++DSTYKTN++KMPL +IVG+ +
Sbjct: 196 GF-WNRICLDESSRVTAVLFAHPKSLEYLKTYPEVLILDSTYKTNRFKMPLLDIVGVDAC 254
Query: 261 EMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKK-QMPKVIVTDRDTGLMNAVANIFPKS 319
+ T+ + FAFL+ E+E +FTW LQ ++ + +P VI+TDR MNAV++ FP S
Sbjct: 255 QRTFCIAFAFLSGEEEGDFTWALQALRSVYEDHNIGLPSVILTDRCLACMNAVSSCFPGS 314
Query: 320 TALVCEFHILKNVRARC-ILDCKVKDSKGKHVKSSDLVNTIMNAWEVIVSSPSEELYADA 378
+C +HI K V++ C + KD+ S+ N W IV+S +E++Y +
Sbjct: 315 ALFLCLWHINKAVQSYCRPAFTRGKDNPQGLGGESEEWKEFFNFWHEIVASTTEDIYNER 374
Query: 379 VLQFRKVCEQFPKFLK---YVESTILEPLKKKIVKAWTNRVMHFGCTTTNGVESAHGTLK 435
+ +F+K P ++ Y+ T L+ KK VKAW N +HF T+ VE H +K
Sbjct: 375 LEKFKK--RYIPDYINEVGYILETWLDLYKKSFVKAWVNTHLHFEQYATSRVEGIHSLIK 432
Query: 436 GYLEDSKGDLAKGWKAINAMLTVQFTEIQATFGRNSTVLEHKFKGNKMYKSLEYNVSRTA 495
+L S+ DL + W+ I +L Q ++++A R + + +Y ++ +S A
Sbjct: 433 LHLNHSQVDLFEAWRVIKLVLMNQLSQLEANQARQH-ISNPIRESRVLYSNIRGWISHEA 491
Query: 496 MNYIYKQASRVKDCGSDSKKCGCVIRWTHALPCTCLIYKKLNLGSCIRLDEINIHW 551
+ + Q R+ + C V T LPC + L + L+ + HW
Sbjct: 492 LRKVETQRERLL---KEVPVCTGVFTRTLGLPCAHSLQPLLKQNQPLLLNHFHSHW 544
>gb|AAW42067.1| conserved hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21] gi|50260537|gb|EAL23192.1|
hypothetical protein CNBA5360 [Cryptococcus neoformans
var. neoformans B-3501A] gi|50259285|gb|EAL21958.1|
hypothetical protein CNBC0980 [Cryptococcus neoformans
var. neoformans B-3501A] gi|50256296|gb|EAL19021.1|
hypothetical protein CNBH1230 [Cryptococcus neoformans
var. neoformans B-3501A] gi|57229113|gb|AAW45547.1|
conserved hypothetical protein [Cryptococcus neoformans
var. neoformans JEC21] gi|58271396|ref|XP_572854.1|
conserved hypothetical protein [Cryptococcus neoformans
var. neoformans JEC21] gi|58264436|ref|XP_569374.1|
conserved hypothetical protein [Cryptococcus neoformans
var. neoformans JEC21]
Length = 932
Score = 138 bits (347), Expect = 9e-31
Identities = 140/595 (23%), Positives = 247/595 (40%), Gaps = 67/595 (11%)
Query: 2 DDLVEPDLEVYFIDVAEFFVTK---EKKKYEDREEMINWARGQAIEAGFTLIIDKSYSGS 58
DD+++ ++ + D E T E+ Y+ ++E + A GF +I +S
Sbjct: 70 DDIIQAPIDSFNDD--EMISTPGFDEQAYYQSKDECKGAVQAYAASQGFAAVIRRSK--- 124
Query: 59 GRRKPKLVLAFERS---------GEYKGTKKSKREGTGSR-----KCGCPFRLRGYFSAT 104
R+KP R EY+ T+ + R K GCPF + A
Sbjct: 125 -RKKPTKTHKEGRELIQYSCIFGDEYRCTRGDSFKDVRIRTIQTVKNGCPFAAVAWEMAE 183
Query: 105 ---------KLWSLNIVSGLHNHKMEPKLEGHMLAGRLTAKESK-IVGDMTRNLIKPKNI 154
K W+ +V+ HNH + +L E+K ++ + + +I
Sbjct: 184 AEVKEHGGDKCWTWKVVNASHNHPPVANIGIALLRRASRTPEAKELINTLIDSRTPFPSI 243
Query: 155 LLDLKGRWKDNRTVAKQIYNARHRYKLSIRGSR*EMQELFQKFEENNYVFNYRTVSGSRT 214
+ + R+ ++ ++ R + +R ++ N+ + +R +R
Sbjct: 244 ISTMSQRFPTVMLTSEDLHRLAFRRRADLRAGSSASAACIRQLVSNDEL--HRPFVNTRD 301
Query: 215 -VQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQYKMPLFEIVGLTSTEMTYNVGFAFLTM 273
+ L + + L + FP VL D TYKTN Y+MP+ IVG TST MTY G +
Sbjct: 302 ELIGLVYTKRTARALIHRFPEVLFFDCTYKTNLYRMPMLHIVGSTSTGMTYTAGVILMLR 361
Query: 274 EKEDNFTWGLQESANLLKCKKQMPKVIVTDRDTGLMNAVANIFPKSTALVCEFHILKNVR 333
E + +T L L+ K KV++TDRD L+NA+ ++ PK+ C +H+ +NV+
Sbjct: 362 ETTNWYTQALNSFFELV--GKPDVKVVITDRDPALINALMSVLPKAYRFSCFWHLQENVK 419
Query: 334 ARCILDCKVKDSKGKHVKSSDLVNTIMNAW-EVIVSSPSEELYADAVLQFRKVCEQFP-- 390
+ I C V + V+ AW V + S+E + +RK+ E +P
Sbjct: 420 SN-IRPCFVDEE-----NIVMAVSNFCKAWVRYCVHAKSDEQLEEG---YRKMEELYPGQ 470
Query: 391 ---KFLKYVESTILEPLKKKIVKAWTNRVMHFGCTTTNGVESAHGTLKGYLEDSKGDLAK 447
+ + Y+ L+ +K++ V A+ N+ +HFG T + +E H TLK ++ GDL
Sbjct: 471 KYARAISYIRG--LDEIKERFVHAYINKQLHFGQTGNSRLEGQHATLKKSIDTKYGDLLL 528
Query: 448 GWKAINAMLTVQFTEIQATFGRNSTVLEHKFKGNKMYKSLEYNVSRTAMNYIYKQASRVK 507
+++ Q+ +I T + L+ ++SR AM + +Q + K
Sbjct: 529 VIGSLSTYFDQQWLKILKRIELERTRCATHIPST--FVRLKGSISRAAMKLLAEQLTLAK 586
Query: 508 ----------DCGSDSKKCGCVIRWTHALPCTCLIYKKLNLGSCIRLDEINIHWK 552
D +S C +H LPC + + + D I+ HW+
Sbjct: 587 RYLGDYDEGVDDFEESHPCSGSFTKSHGLPCAHRLISFVRERRHLEKDHIHPHWR 641
>gb|AAW44911.1| conserved hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21] gi|58270124|ref|XP_572218.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21]
Length = 920
Score = 137 bits (344), Expect = 2e-30
Identities = 137/589 (23%), Positives = 241/589 (40%), Gaps = 67/589 (11%)
Query: 2 DDLVEPDLEVYFIDVAEFFVTK---EKKKYEDREEMINWARGQAIEAGFTLIIDKSYSGS 58
DD+++ ++ + D E T E+ Y+ ++E + A GF +I +S
Sbjct: 70 DDIIQAPIDSFNDD--EMISTPGFDEQAYYQSKDECKGAVQAYAASQGFAAVIRRSK--- 124
Query: 59 GRRKPKLVLAFERS---------GEYKGTKKSKREGTGSR-----KCGCPFRLRGYFSAT 104
R+KP R EY+ T+ + R K GCPF + A
Sbjct: 125 -RKKPTKTHKEGRELIQYSCIFGDEYRCTRGDSFKDVRIRTIQTVKNGCPFAAVAWEMAE 183
Query: 105 ---------KLWSLNIVSGLHNHKMEPKLEGHMLAGRLTAKESK-IVGDMTRNLIKPKNI 154
K W+ +V+ HNH + +L E+K ++ + + +I
Sbjct: 184 AEVKEHGGDKCWTWKVVNASHNHPPVANIGIALLRRASRTPEAKELINTLIDSRTPFPSI 243
Query: 155 LLDLKGRWKDNRTVAKQIYNARHRYKLSIRGSR*EMQELFQKFEENNYVFNYRTVSGSRT 214
+ + R+ ++ ++ R + +R ++ N+ + +R +R
Sbjct: 244 ISTMSQRFPTVMLTSEDLHRLAFRRRADLRAGSSASAACIRQLVSNDEL--HRPFVNTRD 301
Query: 215 -VQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQYKMPLFEIVGLTSTEMTYNVGFAFLTM 273
+ L + + L + FP VL D TYKTN Y+MP+ IVG TST MTY G +
Sbjct: 302 ELIGLVYTKRTARALIHRFPEVLFFDCTYKTNLYRMPMLHIVGSTSTGMTYTAGVILMLR 361
Query: 274 EKEDNFTWGLQESANLLKCKKQMPKVIVTDRDTGLMNAVANIFPKSTALVCEFHILKNVR 333
E + +T L L+ K KV++TDRD L+NA+ ++ PK+ C +H+ +NV+
Sbjct: 362 ETTNWYTQALNSFFELV--GKPDVKVVITDRDPALINALMSVLPKAYRFSCFWHLQENVK 419
Query: 334 ARCILDCKVKDSKGKHVKSSDLVNTIMNAWEVIVSSPSEELYADAVLQFRKVCEQFPKFL 393
+ I C V + V+ AW EELY +++ + +
Sbjct: 420 SN-IRPCFVDEE-----NIVMAVSNFCKAWLEEGYRKMEELYPG---------QKYARAI 464
Query: 394 KYVESTILEPLKKKIVKAWTNRVMHFGCTTTNGVESAHGTLKGYLEDSKGDLAKGWKAIN 453
Y+ L+ +K++ V A+ N+ +HFG T + +E H TLK ++ GDL +++
Sbjct: 465 SYIRG--LDEIKERFVHAYINKQLHFGQTGNSRLEGQHATLKKSIDTKYGDLLLVIGSLS 522
Query: 454 AMLTVQFTEIQATFGRNSTVLEHKFKGNKMYKSLEYNVSRTAMNYIYKQASRVK------ 507
Q+ +I T + L+ ++SR AM + +Q + K
Sbjct: 523 TYFDQQWLKILKRIELERTRCATHIPST--FVRLKGSISRAAMKLLAEQLTLAKRYLGDY 580
Query: 508 ----DCGSDSKKCGCVIRWTHALPCTCLIYKKLNLGSCIRLDEINIHWK 552
D +S C +H LPC + + + D I+ HW+
Sbjct: 581 DEGVDDFEESHPCSGSFTKSHGLPCAHRLISFVRERRHLEKDHIHPHWR 629
>gb|AAW44276.1| conserved hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21] gi|58268854|ref|XP_571583.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21]
Length = 895
Score = 128 bits (321), Expect = 9e-28
Identities = 137/595 (23%), Positives = 238/595 (39%), Gaps = 104/595 (17%)
Query: 2 DDLVEPDLEVYFIDVAEFFVTK---EKKKYEDREEMINWARGQAIEAGFTLIIDKSYSGS 58
DD+++ ++ + D E T E+ Y+ ++E + A GF +I +S
Sbjct: 70 DDIIQAPIDSFNDD--EMISTPGFDEQAYYQSKDECKGAVQAYAASQGFAAVIRRSK--- 124
Query: 59 GRRKPKLVLAFERS---------GEYKGTKKSKREGTGSR-----KCGCPFRLRGYFSAT 104
R+KP R EY+ T+ + R K GCPF + A
Sbjct: 125 -RKKPTKTHKEGRELIQYSCIFGDEYRCTRGDSFKDVRIRTIQTVKNGCPFAAVAWEMAE 183
Query: 105 ---------KLWSLNIVSGLHNHKMEPKLEGHMLAGRLTAKESK-IVGDMTRNLIKPKNI 154
K W+ +V+ HNH + +L E+K ++ + + +I
Sbjct: 184 AEVKEHGGDKCWTWKVVNASHNHPPVANIGIALLRRASRTPEAKELINTLIDSRTPFPSI 243
Query: 155 LLDLKGRWKDNRTVAKQIYNARHRYKLSIRGSR*EMQELFQKFEENNYVFNYRTVSGSRT 214
+ + R+ ++ ++ R + +R ++ N+ + +R +R
Sbjct: 244 ISTMSQRFPTVMLTSEDLHRLAFRRRADLRAGSSASAACIRQLVSNDEL--HRPFVNTRD 301
Query: 215 -VQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQYKMPLFEIVGLTSTEMTYNVGFAFLTM 273
+ L + + L + FP VL D TYKTN Y+MP+ IVG TST MTY G +
Sbjct: 302 ELIGLVYTKRTARALIHRFPEVLFFDCTYKTNLYRMPMLHIVGSTSTGMTYTAGVILMLR 361
Query: 274 EKEDNFTWGLQESANLLKCKKQMPKVIVTDRDTGLMNAVANIFPKSTALVCEFHILKNVR 333
E + +T L L+ K KV++TDRD L+NA+ ++ PK+ C +H+ +NV+
Sbjct: 362 ETTNWYTQALNSFFELV--GKPDVKVVITDRDPALINALMSVLPKAYRFSCFWHLQENVK 419
Query: 334 ARCILDCKVKDSKGKHVKSSDLVNTIMNAW-EVIVSSPSEELYADAVLQFRKVCEQFP-- 390
+ I C V + V+ AW V + S+E + +RK+ E +P
Sbjct: 420 SN-IRPCFVDEE-----NIVMAVSNFCKAWVRYCVHAKSDEQLEEG---YRKMEELYPGQ 470
Query: 391 ---KFLKYVESTILEPLKKKIVKAWTNRVMHFGCTTTNGVESAHGTLKGYLEDSKGDLAK 447
+ + Y+ L+ +K++ V A+ N+ +HFG T + +E H TLK ++ GDL
Sbjct: 471 KYARAISYIRG--LDEIKERFVHAYINKQLHFGQTGNSRLEGQHATLKKSIDTKYGDLL- 527
Query: 448 GWKAINAMLTVQFTEIQATFGRNSTVLEHKFKGNKMYKSLEYNVSRTAMNYIYKQASRVK 507
L+ ++SR AM + +Q + K
Sbjct: 528 --------------------------------------LLKGSISRAAMKLLAEQLTLAK 549
Query: 508 ----------DCGSDSKKCGCVIRWTHALPCTCLIYKKLNLGSCIRLDEINIHWK 552
D +S C +H LPC + + + D I+ HW+
Sbjct: 550 RYLGDYDEGVDDFEESHPCSGSFTKSHGLPCAHRLISFVRERRHLEKDHIHPHWR 604
>gb|EAL17593.1| hypothetical protein CNBM0450 [Cryptococcus neoformans var.
neoformans B-3501A] gi|57226908|gb|AAW43368.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21] gi|58267038|ref|XP_570675.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21]
Length = 580
Score = 126 bits (316), Expect = 3e-27
Identities = 119/478 (24%), Positives = 207/478 (42%), Gaps = 55/478 (11%)
Query: 2 DDLVEPDLEVYFIDVAEFFVTK---EKKKYEDREEMINWARGQAIEAGFTLIIDKSYSGS 58
DD+++ ++ + D E T E+ Y+ ++E + A GF +I +S
Sbjct: 70 DDIIQAPIDSFNDD--EMISTPGFDEQAYYQSKDECKGAVQAYAASQGFAAVIRRSK--- 124
Query: 59 GRRKPKLVLAFERS---------GEYKGTKKSKREGTGSR-----KCGCPFRLRGYFSAT 104
R+KP R EY+ T+ + R K GCPF + A
Sbjct: 125 -RKKPTKTHKEGRELIQYSCIFGDEYRCTRGDSFKDVRIRTIQTVKNGCPFAAVAWEMAE 183
Query: 105 ---------KLWSLNIVSGLHNHKMEPKLEGHMLAGRLTAKESK-IVGDMTRNLIKPKNI 154
K W+ +V+ HNH + +L E+K ++ + + +I
Sbjct: 184 AEVKEHGGDKCWTWKVVNASHNHPPVANIGIALLRRASRTPEAKELINTLIDSRTPFPSI 243
Query: 155 LLDLKGRWKDNRTVAKQIYNARHRYKLSIRGSR*EMQELFQKFEENNYVFNYRTVSGSRT 214
+ + R+ ++ ++ R + +R ++ N+ + +R +R
Sbjct: 244 ISTMSQRFPTVMLTSEDLHRLAFRRRADLRAGSSASAACIRQLVSNDEL--HRPFVNTRD 301
Query: 215 -VQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQYKMPLFEIVGLTSTEMTYNVGFAFLTM 273
+ L + + L + FP VL D TYKTN Y+MP+ IVG TST MTY G +
Sbjct: 302 ELIGLVYTKRTARALIHRFPEVLFFDCTYKTNLYRMPMLHIVGSTSTGMTYTAGVILMLR 361
Query: 274 EKEDNFTWGLQESANLLKCKKQMPKVIVTDRDTGLMNAVANIFPKSTALVCEFHILKNVR 333
E + +T L L+ K KV++TDRD L+NA+ ++ PK+ C +H+ +NV+
Sbjct: 362 ETTNWYTQALNSFFELV--GKPDVKVVITDRDPALINALMSVLPKAYRFSCFWHLQENVK 419
Query: 334 ARCILDCKVKDSKGKHVKSSDLVNTIMNAW-EVIVSSPSEELYADAVLQFRKVCEQFP-- 390
+ I C V + V+ AW V + S+E + +RK+ E +P
Sbjct: 420 SN-IRPCFVDEE-----NIVMAVSNFCKAWVRYCVHAKSDEQLEEG---YRKMEELYPGQ 470
Query: 391 ---KFLKYVESTILEPLKKKIVKAWTNRVMHFGCTTTNGVESAHGTLKGYLEDSKGDL 445
+ + Y+ L+ +K++ V A+ N+ +HFG T + +E H TLK ++ GDL
Sbjct: 471 KYARAISYIRG--LDEIKERFVHAYINKQLHFGQTGNSRLEGQHATLKKSIDTKYGDL 526
>gb|AAW41060.1| conserved hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21] gi|58258933|ref|XP_566879.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21]
Length = 678
Score = 125 bits (315), Expect = 4e-27
Identities = 96/341 (28%), Positives = 156/341 (45%), Gaps = 31/341 (9%)
Query: 228 LFNTFPTVLVMDSTYKTNQYKMPLFEIVGLTSTEMTYNVGFAFLTMEKEDNFTWGLQESA 287
L + FP VL D TYKTN Y+MP+ IVG TST MTY G + E + +T L
Sbjct: 62 LIHRFPEVLFFDCTYKTNLYRMPMLHIVGSTSTGMTYTAGVILMLRETTNWYTQALNSFF 121
Query: 288 NLLKCKKQMPKVIVTDRDTGLMNAVANIFPKSTALVCEFHILKNVRARCILDCKVKDSKG 347
L+ K KV++TDRD L+NA+ ++ PK+ C +H+ +NV++ I C V +
Sbjct: 122 ELV--GKPDVKVVITDRDPALINALMSVLPKAYRFSCFWHLQENVKSN-IRPCFVDEE-- 176
Query: 348 KHVKSSDLVNTIMNAW-EVIVSSPSEELYADAVLQFRKVCEQFP-----KFLKYVESTIL 401
V+ AW V + S+E + +RK+ E +P + + Y+ L
Sbjct: 177 ---NIVMAVSNFCKAWVRYCVHAKSDEQLEEG---YRKMEELYPGQKYARAISYIRG--L 228
Query: 402 EPLKKKIVKAWTNRVMHFGCTTTNGVESAHGTLKGYLEDSKGDLAKGWKAINAMLTVQFT 461
+ +K++ V A+ N+ +HFG T + +E H TLK ++ GDL +++ Q+
Sbjct: 229 DEIKERFVHAYINKQLHFGQTGNSRLEGQHATLKKSIDTKYGDLLLVIGSLSTYFDQQWL 288
Query: 462 EIQATFGRNSTVLEHKFKGNKMYKSLEYNVSRTAMNYIYKQASRVK----------DCGS 511
+I T + L+ ++SR AM + +Q + K D
Sbjct: 289 KILKRIELERTRCATHIPST--FVRLKGSISRAAMKLLAEQLTLAKRYLGDYDEGVDDFE 346
Query: 512 DSKKCGCVIRWTHALPCTCLIYKKLNLGSCIRLDEINIHWK 552
+S C +H LPC + + + D I+ HW+
Sbjct: 347 ESHPCSGSFTKSHGLPCAHRLISFVRERRHLEKDHIHPHWR 387
>emb|CAE02137.2| OSJNBa0074L08.5 [Oryza sativa (japonica cultivar-group)]
gi|50926620|ref|XP_473257.1| OSJNBa0074L08.5 [Oryza
sativa (japonica cultivar-group)]
Length = 1061
Score = 105 bits (262), Expect = 6e-21
Identities = 98/395 (24%), Positives = 167/395 (41%), Gaps = 43/395 (10%)
Query: 26 KKYEDREEMINWARGQAIEAGFTLIIDKSYSGSGRRK----PKLVLAFERSGEYKGTKKS 81
KK+ EE ++ A + GF+++I Y + +++ K R G+ +K
Sbjct: 301 KKFRTHEEARSFFNFYAYQVGFSVVITHHYKSTSKKRFGQITKYTYQCYRYGKNDDGQKK 360
Query: 82 KREGTGSRK---------CGCPFRLRGYFSATKLWSLNIVSGLHNHKMEPKLEGHMLAGR 132
K++ T ++ C C +R ++ S+N+ HNH + P E L
Sbjct: 361 KKKKTQQQRNTQVIDRTDCKCMLTVRLDGDVWRILSMNLN---HNHTLTPNREAKFLRSH 417
Query: 133 --LTAKESKIVGDM------TRNLIKPKNILLDLKGRWKDNRTVAKQIYNARHRYKLSIR 184
+T +E K++ + TRN+I +IL L+G K + N R
Sbjct: 418 KHMTDEEKKLIRTLKQCNIPTRNMI---SILSFLRGGIAALPYTGKDVSNVGTAINSETR 474
Query: 185 GSR*E--MQELFQKFEENNYVFNYRTVSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTY 242
+ M+ L QK ++ + T+ + V+ +F+ S +L+ + + D+TY
Sbjct: 475 NTDMNQVMEYLRQKEAKDPGYYYKLTLDENNKVKSMFWTDGRSTQLYEQYGEFVSFDTTY 534
Query: 243 KTNQYKMPLFEIVGLTSTEMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKKQMPKVIVT 302
KTN+Y MP IVG+T AFL E + F W + + K P+ I+T
Sbjct: 535 KTNRYNMPFAPIVGVTGHGNICIFACAFLGDETTETFKWVFETFLTAMGGKH--PETIIT 592
Query: 303 DRDTGLMNAVANIFPKSTALVCEFHILKNVRARCILDCKVKDSKGKHVKSSDLVNTIMNA 362
D+D + A+ +FP S C FHILK R R K K + + +D+V+ +
Sbjct: 593 DQDLAMRAAIRQVFPNSKHRNCLFHILKKCRERSGNTFSDKRRKDLYAEFNDIVHNSLTR 652
Query: 363 WEVIVSSPSEELYADAVLQFRKVCEQFPKFLKYVE 397
E E L+ + Q+ K +KY+E
Sbjct: 653 AEF------ESLWLQMIAQYNL------KNIKYLE 675
>ref|XP_463702.1| putative far-red impaired response protein [Oryza sativa (japonica
cultivar-group)]
Length = 1114
Score = 104 bits (259), Expect = 1e-20
Identities = 94/380 (24%), Positives = 161/380 (41%), Gaps = 37/380 (9%)
Query: 26 KKYEDREEMINWARGQAIEAGFTLIIDKSYSGSGRRK----PKLVLAFERSGEYKGTKKS 81
KK+ EE ++ A + GF+++I Y + +++ K R G+ +K
Sbjct: 354 KKFRTHEEARSFFNFYAYQVGFSVVITHHYKSTSKKRFGQITKYTYQCYRYGKNDDGQKK 413
Query: 82 KREGTGSRK---------CGCPFRLRGYFSATKLWSLNIVSGLHNHKMEPKLEGHMLAGR 132
K++ T ++ C C +R ++ S+N+ HNH + P E L
Sbjct: 414 KKKKTQQQRNTQVIDRTDCKCMLTVRLEGDVWRILSMNLN---HNHTLTPNREAKFLRSH 470
Query: 133 --LTAKESKIVGDM------TRNLIKPKNILLDLKGRWKDNRTVAKQIYNARHRYKLSIR 184
+T +E K++ + TRN+I +IL L+G K + N R
Sbjct: 471 KHMTDEEKKLIRTLKQCNIPTRNMI---SILSFLRGGIAALPYTRKDVSNVGTAINSETR 527
Query: 185 GSR*E--MQELFQKFEENNYVFNYRTVSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTY 242
+ M+ L QK ++ + T+ + V+ +F+ S +L+ + + D+TY
Sbjct: 528 NTDMNQVMEYLRQKEAKDPGYYYKLTLDENNKVKSMFWTDGRSTQLYEQYGEFVSFDTTY 587
Query: 243 KTNQYKMPLFEIVGLTSTEMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKKQMPKVIVT 302
KTN+Y MP IVG+T AFL E + F W + + K P+ I+T
Sbjct: 588 KTNRYNMPFAPIVGVTGHGNICIFACAFLGDETTETFKWVFETFLTAMGGKH--PETIIT 645
Query: 303 DRDTGLMNAVANIFPKSTALVCEFHILKNVRARCILDCKVKDSKGKHVKSSDLVNTIMNA 362
D+D + A+ +FP S C FHILK R R K K + + +D+V+ +
Sbjct: 646 DQDLAMRAAIRQVFPNSKHRNCLFHILKKCRERSGNTFSDKRRKDLYAEFNDIVHNSLTR 705
Query: 363 WEVIVSSPSEELYADAVLQF 382
E E L+ + Q+
Sbjct: 706 AEF------ESLWLQMIAQY 719
>dbj|BAD87203.1| far-red impaired response-like [Oryza sativa (japonica
cultivar-group)]
Length = 1130
Score = 104 bits (259), Expect = 1e-20
Identities = 94/380 (24%), Positives = 161/380 (41%), Gaps = 37/380 (9%)
Query: 26 KKYEDREEMINWARGQAIEAGFTLIIDKSYSGSGRRK----PKLVLAFERSGEYKGTKKS 81
KK+ EE ++ A + GF+++I Y + +++ K R G+ +K
Sbjct: 370 KKFRTHEEARSFFNFYAYQVGFSVVITHHYKSTSKKRFGQITKYTYQCYRYGKNDDGQKK 429
Query: 82 KREGTGSRK---------CGCPFRLRGYFSATKLWSLNIVSGLHNHKMEPKLEGHMLAGR 132
K++ T ++ C C +R ++ S+N+ HNH + P E L
Sbjct: 430 KKKKTQQQRNTQVIDRTDCKCMLTVRLEGDVWRILSMNLN---HNHTLTPNREAKFLRSH 486
Query: 133 --LTAKESKIVGDM------TRNLIKPKNILLDLKGRWKDNRTVAKQIYNARHRYKLSIR 184
+T +E K++ + TRN+I +IL L+G K + N R
Sbjct: 487 KHMTDEEKKLIRTLKQCNIPTRNMI---SILSFLRGGIAALPYTRKDVSNVGTAINSETR 543
Query: 185 GSR*E--MQELFQKFEENNYVFNYRTVSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTY 242
+ M+ L QK ++ + T+ + V+ +F+ S +L+ + + D+TY
Sbjct: 544 NTDMNQVMEYLRQKEAKDPGYYYKLTLDENNKVKSMFWTDGRSTQLYEQYGEFVSFDTTY 603
Query: 243 KTNQYKMPLFEIVGLTSTEMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKKQMPKVIVT 302
KTN+Y MP IVG+T AFL E + F W + + K P+ I+T
Sbjct: 604 KTNRYNMPFAPIVGVTGHGNICIFACAFLGDETTETFKWVFETFLTAMGGKH--PETIIT 661
Query: 303 DRDTGLMNAVANIFPKSTALVCEFHILKNVRARCILDCKVKDSKGKHVKSSDLVNTIMNA 362
D+D + A+ +FP S C FHILK R R K K + + +D+V+ +
Sbjct: 662 DQDLAMRAAIRQVFPNSKHRNCLFHILKKCRERSGNTFSDKRRKDLYAEFNDIVHNSLTR 721
Query: 363 WEVIVSSPSEELYADAVLQF 382
E E L+ + Q+
Sbjct: 722 AEF------ESLWLQMIAQY 735
>dbj|BAD81770.1| putative far-red impaired response protein [Oryza sativa (japonica
cultivar-group)]
Length = 883
Score = 92.4 bits (228), Expect = 5e-17
Identities = 96/427 (22%), Positives = 178/427 (41%), Gaps = 37/427 (8%)
Query: 26 KKYEDREEMINWARGQAIEAGFTLIIDKSYSGSGRRKPKLVLAFERSGEYKGTKKSKREG 85
++++ + N+ A+ GF + +DK + + +++ + G TKK
Sbjct: 126 QRFKTERDAYNFYNVYAVSKGFGIRLDKDHMNTKKQRTMRQICCSHQGRNPKTKKP---- 181
Query: 86 TGSRKCGCPFRLRGYFS-ATKLWSLNIVSGLHNHKMEPKL---EGHMLAGRLTAKESKIV 141
S + GCP ++ S A WS+ V HNH M+ + + + ++ I+
Sbjct: 182 --SVRIGCPAMMKINMSGAGSGWSVTKVVSAHNHPMKKSVGVTKNYQSHNQIDEGTRGII 239
Query: 142 GDMTRNLIKPKNILLDLKGRWKDNRTVAKQIYNARHRYKLSIRG--SR*EMQELFQKFEE 199
+M + + N+ L G ++ A R +IR S +MQ+ ++
Sbjct: 240 EEMVDSSMSLTNMYGMLSGM-HGGPSMVPFTRKAMDRVAYAIRRDESSDDMQKTLDVLKD 298
Query: 200 -----NNYVFNYRTVSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQYKMPLFEI 254
N+ ++ + R V+++F++H S F F V+ D+TYKTN+Y MP
Sbjct: 299 LQKRSKNFFYSIQVDEACR-VKNIFWSHAVSRLNFEHFGDVITFDTTYKTNKYNMPFAPF 357
Query: 255 VGLTSTEMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKKQMPKVIVTDRDTGLMNAVAN 314
VG+ + + G A L E E++FTW + K +P I+TD + A+
Sbjct: 358 VGVNNHFQSTFFGCALLREETEESFTWLFNTFKECMNGK--VPIGILTDNCPSMAAAIRT 415
Query: 315 IFPKSTALVCEFHILKNVRARCILDCKVKDSKGKHVKSSDLVNTIMNAWEVIVSSPSEEL 374
+FP + VC+ H+LK K K+ G +T A+ +++ E
Sbjct: 416 VFPNTIHRVCKSHVLK----------KAKEFMGNIYSK---CHTFKKAFHKVLTQTLTE- 461
Query: 375 YADAVLQFRKVCEQFPKFLKYVESTILEPLKKKIVKAWTNRVMHFGCTTTNGVESAHGTL 434
+ V + K+ + K V + +++K + + G TTT ESA+
Sbjct: 462 -EEFVAAWHKLIRDY-NLEKSVYLRHIWDIRRKWAFVYFSHRFFAGMTTTQRSESANHVF 519
Query: 435 KGYLEDS 441
K ++ S
Sbjct: 520 KMFVSPS 526
>emb|CAE02218.2| OSJNBb0002N06.8 [Oryza sativa (japonica cultivar-group)]
gi|50923349|ref|XP_472035.1| OSJNBb0002N06.8 [Oryza
sativa (japonica cultivar-group)]
Length = 885
Score = 91.3 bits (225), Expect = 1e-16
Identities = 95/427 (22%), Positives = 177/427 (41%), Gaps = 37/427 (8%)
Query: 26 KKYEDREEMINWARGQAIEAGFTLIIDKSYSGSGRRKPKLVLAFERSGEYKGTKKSKREG 85
++++ + N+ A+ GF + +DK + +++ + G TKK
Sbjct: 128 QRFKTERDAYNFYNVYAVSKGFGIRLDKDRMNTEKQRTMRQICCSHQGRNPKTKKP---- 183
Query: 86 TGSRKCGCPFRLR-GYFSATKLWSLNIVSGLHNHKMEPKL---EGHMLAGRLTAKESKIV 141
S + GCP ++ A WS+ V HNH M+ + + + ++ I+
Sbjct: 184 --SVRIGCPAMMKINRSGAGSGWSVTKVVSTHNHPMKKSVGVTKNYQSHNQIDEGTRGII 241
Query: 142 GDMTRNLIKPKNILLDLKGRWKDNRTVAKQIYNARHRYKLSIRG--SR*EMQELFQKFEE 199
+M + + N+ L G ++ A R +IR S +MQ+ ++
Sbjct: 242 EEMVDSSMSLTNMYGMLSGM-HGGPSMVPFTRKAMDRLAYAIRRDESSDDMQKTLDVLKD 300
Query: 200 -----NNYVFNYRTVSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQYKMPLFEI 254
N+ ++ + R V+++F++H S F F V+ D+TYKTN+Y MP
Sbjct: 301 LQKRSKNFFYSIQVDEACR-VKNIFWSHAVSRLNFEHFGDVITFDTTYKTNKYNMPFAPF 359
Query: 255 VGLTSTEMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKKQMPKVIVTDRDTGLMNAVAN 314
VG+ + + G A L E E++FTW + K +P I+TD + A+
Sbjct: 360 VGVNNHFQSTFFGCALLREETEESFTWLFNTFKECMNGK--VPIGILTDNCPSMAAAIRT 417
Query: 315 IFPKSTALVCEFHILKNVRARCILDCKVKDSKGKHVKSSDLVNTIMNAWEVIVSSPSEEL 374
+FP + VC++H+LK K K+ G +T A+ +++ E
Sbjct: 418 VFPNTIHRVCKWHVLK----------KAKEFMGNIYSKR---HTFKKAFHKVLTQTLTE- 463
Query: 375 YADAVLQFRKVCEQFPKFLKYVESTILEPLKKKIVKAWTNRVMHFGCTTTNGVESAHGTL 434
+ V + K+ + K V + +++K + + G TTT ESA+
Sbjct: 464 -EEFVAAWHKLIRDY-NLEKSVYLRHIWDIRRKWAFVYFSHRFFAGMTTTQRSESANHVF 521
Query: 435 KGYLEDS 441
K ++ S
Sbjct: 522 KMFVSPS 528
>gb|AAU10761.1| unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 822
Score = 90.9 bits (224), Expect = 2e-16
Identities = 76/319 (23%), Positives = 140/319 (43%), Gaps = 21/319 (6%)
Query: 26 KKYEDREEMINWARGQAIEAGFTLIIDKSYSGSGRRKPKLVLAFERSGEYKGTKKSKREG 85
++++ + N+ A+ GF + +DK + +++ + G TKK
Sbjct: 128 QRFKTERDAYNFYNVYAVSKGFGIRLDKDRMNTKKQRTMRQICCSHQGRNPKTKKP---- 183
Query: 86 TGSRKCGCPFRLRGYFS-ATKLWSLNIVSGLHNHKMEPKL---EGHMLAGRLTAKESKIV 141
S + GCP ++ S A WS+ V HNH M+ + + + ++ I+
Sbjct: 184 --SVRIGCPAMMKINMSGAGSGWSVTKVVSAHNHPMKKSVGVTKNYQSHNQIDEGTRGII 241
Query: 142 GDMTRNLIKPKNILLDLKGRWKDNRTVAKQIYNARHRYKLSIRG--SR*EMQELFQKFEE 199
+M + + N+ L G ++ A R +IR S +MQ+ ++
Sbjct: 242 EEMVDSSMSLTNMYGMLSGM-HGGPSMVPFTRKAMDRVAYAIRRDESSDDMQKTLDVLKD 300
Query: 200 -----NNYVFNYRTVSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQYKMPLFEI 254
N+ ++ + R V+++F++H S F F V+ D+TYKTN+Y MP
Sbjct: 301 LQKRSKNFFYSIQVDEACR-VKNIFWSHAVSRLNFEHFGDVITFDTTYKTNKYNMPFAPF 359
Query: 255 VGLTSTEMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKKQMPKVIVTDRDTGLMNAVAN 314
VG+ + + G A L E E++FTW + K +P I+TD + A+
Sbjct: 360 VGVNNHFQSTFFGCALLREETEESFTWLFNTFKECMNGK--VPIGILTDNCPSMAAAIRT 417
Query: 315 IFPKSTALVCEFHILKNVR 333
+FP + VC++H+LK +
Sbjct: 418 VFPNTIHRVCKWHVLKKAK 436
>ref|XP_481410.1| far-red impaired response protein -like [Oryza sativa (japonica
cultivar-group)] gi|35215248|dbj|BAC92598.1| putative
far-red impaired response protein [Oryza sativa
(japonica cultivar-group)] gi|35215057|dbj|BAC92415.1|
putative far-red impaired response protein [Oryza sativa
(japonica cultivar-group)]
Length = 1132
Score = 90.5 bits (223), Expect = 2e-16
Identities = 67/284 (23%), Positives = 124/284 (43%), Gaps = 15/284 (5%)
Query: 91 CGCPFRLRGYFSATKLWSLNIVSGLHNHKMEPKLEGHMLAGR--LTAKESKIVGDMTRNL 148
C C + F+ +W + + HNH + P+ E L +T KE +++ +
Sbjct: 479 CKCMMMVEKMFALGDVWQIATLDLKHNHALCPRREAKFLKSHKNMTIKEKRLIRTLKECN 538
Query: 149 IKPKN---ILLDLKGRWKDNRTVAKQIYNARHRYKLSIRGSR*EMQELF-----QKFEEN 200
I ++ IL L+G K + N R + +M+++ ++ E+
Sbjct: 539 IPTRSMIVILSFLRGGLPALPYTKKDVSNVGTAINSETRNN--DMKQVLAYLRKKEIEDP 596
Query: 201 NYVFNYRTVSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQYKMPLFEIVGLTST 260
+ ++ ++ V +F+ S +L+ + + D+TY+TN+Y MP VG+
Sbjct: 597 GISYKFKLDENNK-VTSMFWTDGRSTQLYEEYGDCISFDTTYRTNRYNMPFAPFVGVIGH 655
Query: 261 EMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKKQMPKVIVTDRDTGLMNAVANIFPKST 320
T G AFL E + F W + A + K PK I+TD+D + +A+A +F +
Sbjct: 656 GTTCLFGCAFLGDETAETFKWVFETFATAMGGKH--PKTIITDQDNAMRSAIAQVFQNTK 713
Query: 321 ALVCEFHILKNVRARCILDCKVKDSKGKHVKSSDLVNTIMNAWE 364
C FHI KN R + K +K + + D+++ + E
Sbjct: 714 HRNCLFHIKKNCREKTGSMFSQKSNKNLYDEYDDILSNCLTEAE 757
>ref|NP_908529.1| P0702D12.15 [Oryza sativa (japonica cultivar-group)]
Length = 832
Score = 90.5 bits (223), Expect = 2e-16
Identities = 96/427 (22%), Positives = 177/427 (40%), Gaps = 37/427 (8%)
Query: 26 KKYEDREEMINWARGQAIEAGFTLIIDKSYSGSGRRKPKLVLAFERSGEYKGTKKSKREG 85
++++ + N+ A+ GF + +DK + ++ + G TKK
Sbjct: 75 QRFKTERDAYNFYNVYAVSKGFGIRLDKDRKNTKNQRTMRQICCSHQGRNPKTKKP---- 130
Query: 86 TGSRKCGCPFRLRGYFS-ATKLWSLNIVSGLHNHKMEPKL---EGHMLAGRLTAKESKIV 141
S + GCP ++ S A WS+ V HNH M+ + + + ++ I+
Sbjct: 131 --SVRIGCPAMMKINRSRAGSGWSVTKVVSAHNHPMKKSVGVTKNYQSHNQIDEGTRGII 188
Query: 142 GDMTRNLIKPKNILLDLKGRWKDNRTVAKQIYNARHRYKLSIRG--SR*EMQELFQKFEE 199
+M + + N+ L G ++ A R +IR S +MQ+ ++
Sbjct: 189 EEMVDSSMSLTNMYGMLSGM-HGGPSMVPFTRKAMDRLAYAIRRDESSDDMQKTLDVLKD 247
Query: 200 -----NNYVFNYRTVSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQYKMPLFEI 254
N+ ++ + R V+++F++H S F F V+ D+TYKTN+Y MP
Sbjct: 248 LQKRSKNFFYSIQVDEACR-VKNIFWSHAVSRLNFEHFGDVITFDTTYKTNKYNMPFAPF 306
Query: 255 VGLTSTEMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKKQMPKVIVTDRDTGLMNAVAN 314
VG+ + + G A L E E++FTW + K +P I+TD + A+
Sbjct: 307 VGVNNHFQSTFFGCALLREETEESFTWLFNTFKECMNGK--VPIGILTDNCPSMAAAIRT 364
Query: 315 IFPKSTALVCEFHILKNVRARCILDCKVKDSKGKHVKSSDLVNTIMNAWEVIVSSPSEEL 374
+FP + VC++H+LK K K+ G +T A+ +++ E
Sbjct: 365 VFPNTIHRVCKWHVLK----------KAKEFMGNIYSKR---HTFKKAFHKVLTQTLTE- 410
Query: 375 YADAVLQFRKVCEQFPKFLKYVESTILEPLKKKIVKAWTNRVMHFGCTTTNGVESAHGTL 434
+ V + K+ + K V + +++K + + G TTT ESA+
Sbjct: 411 -EEFVAAWHKLIRDY-NLEKSVYLRHIWDIRRKWAFVYFSHRFFAGMTTTQRSESANHVF 468
Query: 435 KGYLEDS 441
K ++ S
Sbjct: 469 KMFVSPS 475
>gb|AAQ56575.1| putative transposase [Oryza sativa (japonica cultivar-group)]
Length = 1037
Score = 90.5 bits (223), Expect = 2e-16
Identities = 67/284 (23%), Positives = 124/284 (43%), Gaps = 15/284 (5%)
Query: 91 CGCPFRLRGYFSATKLWSLNIVSGLHNHKMEPKLEGHMLAGR--LTAKESKIVGDMTRNL 148
C C + F+ +W + + HNH + P+ E L +T KE +++ +
Sbjct: 384 CKCMMMVEKMFALGDVWQIATLDLKHNHALCPRREAKFLKSHKNMTIKEKRLIRTLKECN 443
Query: 149 IKPKN---ILLDLKGRWKDNRTVAKQIYNARHRYKLSIRGSR*EMQELF-----QKFEEN 200
I ++ IL L+G K + N R + +M+++ ++ E+
Sbjct: 444 IPTRSMIVILSFLRGGLPALPYTKKDVSNVGTAINSETRNN--DMKQVLAYLRKKEIEDP 501
Query: 201 NYVFNYRTVSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQYKMPLFEIVGLTST 260
+ ++ ++ V +F+ S +L+ + + D+TY+TN+Y MP VG+
Sbjct: 502 GISYKFKLDENNK-VTSMFWTDGRSTQLYEEYGDCISFDTTYRTNRYNMPFAPFVGVIGH 560
Query: 261 EMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKKQMPKVIVTDRDTGLMNAVANIFPKST 320
T G AFL E + F W + A + K PK I+TD+D + +A+A +F +
Sbjct: 561 GTTCLFGCAFLGDETAETFKWVFETFATAMGGKH--PKTIITDQDNAMRSAIAQVFQNTK 618
Query: 321 ALVCEFHILKNVRARCILDCKVKDSKGKHVKSSDLVNTIMNAWE 364
C FHI KN R + K +K + + D+++ + E
Sbjct: 619 HRNCLFHIKKNCREKTGSMFSQKSNKNLYDEYDDILSNCLTEAE 662
>gb|AAW41059.1| hypothetical protein CNA05530 [Cryptococcus neoformans var.
neoformans JEC21] gi|58258931|ref|XP_566878.1|
hypothetical protein CNA05530 [Cryptococcus neoformans
var. neoformans JEC21]
Length = 206
Score = 88.6 bits (218), Expect = 8e-16
Identities = 60/197 (30%), Positives = 96/197 (48%), Gaps = 19/197 (9%)
Query: 249 MPLFEIVGLTSTEMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKKQMPKVIVTDRDTGL 308
MP+ IVG TST MTY G + E + +T L L+ K KV++TDRD L
Sbjct: 1 MPMLHIVGSTSTGMTYTAGVILMLRETTNWYTQALNSFFELVG--KPDVKVVITDRDPAL 58
Query: 309 MNAVANIFPKSTALVCEFHILKNVRARCILDCKVKDSKGKHVKSSDLVNTIMNAWEVIVS 368
+NA+ ++ PK+ C +H+ +NV++ I C V + V+ AW
Sbjct: 59 INALMSVLPKAYRFSCFWHLQENVKSN-IRPCFVDEE-----NIVMAVSNFCKAWLEEGY 112
Query: 369 SPSEELYADAVLQFRKVCEQFPKFLKYVESTILEPLKKKIVKAWTNRVMHFGCTTTNGVE 428
EELY +++ + + Y+ L+ +K++ V A+ N+ +HFG T + +E
Sbjct: 113 RKMEELYPG---------QKYARAISYIRG--LDEIKERFVHAYINKQLHFGQTGNSRLE 161
Query: 429 SAHGTLKGYLEDSKGDL 445
H TLK ++ GDL
Sbjct: 162 GQHATLKKSIDTKYGDL 178
>ref|XP_482508.1| putative far-red impaired response protein [Oryza sativa (japonica
cultivar-group)] gi|25553698|dbj|BAC24942.1| putative
far-red impaired response protein [Oryza sativa
(japonica cultivar-group)]
Length = 1148
Score = 86.7 bits (213), Expect = 3e-15
Identities = 84/354 (23%), Positives = 156/354 (43%), Gaps = 32/354 (9%)
Query: 26 KKYEDREEMINWARGQAIEAGFTLIIDKSYSGSGRRKPKLV--LAFERSGEYKGTKKSKR 83
K + E N+ A+ AGF+++ +Y + +++ V + F + + K T +SK
Sbjct: 369 KNITEAHEFFNF---YALLAGFSIVRAHNYHTTSKKRNGEVTRVTFRCNRQGKPTSQSKS 425
Query: 84 EGT-------------GSRKCGCPFRLRGYFSATKLWSLNIVSGLHNHKMEPKLEGHMLA 130
+ C C + ++W+++ V HNH++ P+ E L
Sbjct: 426 VSAEETVVSERNTNENDATDCKCALVIS---ERNEIWNISRVQLDHNHQLSPRDEVRFLK 482
Query: 131 GR--LTAKESKIVGDMTRNLIKPKN---ILLDLKGRWKDNRTVAKQIYNARHRY-KLSIR 184
+T +E ++ + I ++ IL L+G K I N R K +
Sbjct: 483 SHKHMTTEEKMLIRTLKECNIPTRHMIVILSVLRGGLTSLPYTKKDISNVRTTINKETSS 542
Query: 185 GSR*EMQELFQKFEENNYVFNYR-TVSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTYK 243
+ E F++ +E + F Y + S+ V++LF+ S + + + V+ D+TY
Sbjct: 543 NDIMKTLEFFRRKKEKDPHFFYEFDLDESKKVKNLFWTDGRSREWYEKYGDVVSFDTTYF 602
Query: 244 TNQYKMPLFEIVGLTSTEMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKKQMPKVIVTD 303
TN+Y +P VG+ T G AFL E + F W + + K+ PK I+TD
Sbjct: 603 TNKYNLPFAPFVGIYGHGNTIVFGCAFLHDETSETFKWLFRTFLKAMSQKE--PKTIITD 660
Query: 304 RDTGLMNAVANIFPKSTALVCEFHILKNVRARCILDCKVKDSKGKHVKSSDLVN 357
+D + +A+A +F + C FHI+K +A + +K +G + + D++N
Sbjct: 661 QDGAMRSAIAQVFQNAKHRNCFFHIVK--KAFNLSGNLLKAKEGLYDEYEDIIN 712
>ref|XP_466614.1| putative far-red impaired response protein [Oryza sativa (japonica
cultivar-group)] gi|47497275|dbj|BAD19318.1| putative
far-red impaired response protein [Oryza sativa
(japonica cultivar-group)]
Length = 880
Score = 83.2 bits (204), Expect = 3e-14
Identities = 102/456 (22%), Positives = 186/456 (40%), Gaps = 69/456 (15%)
Query: 14 IDVAEFFVTKEKKKYEDREEMINWARGQAIEAGFTLIIDKSYSGSGRRKPKLVLAFERSG 73
++ E + T +++ +E + A E GF+ + K+Y RR P V R
Sbjct: 49 LEEQEEYRTMTAMRFKTEKEGFLFYNRYAKEKGFS--VRKNYI---RRDPITVAVTHRQY 103
Query: 74 E----------YKGTKKSKREGTGSRKCGCPFRLRGYFSATKL-WSLNIVSGLHNHKMEP 122
+ Y RE +CGC K W + HNH +
Sbjct: 104 QCSREGHRKEVYMEAANRSREPKALTRCGCNALFEIKLDKKKGDWFVVKYVAKHNHPLAK 163
Query: 123 KLEGHMLAGRLTAKESKIVGDMTRNLIKPKNI------LLDLK----GRWKDNRTVAKQI 172
E L T ++ N+++ K + ++D+ G + V++ +
Sbjct: 164 SDEVAFLRSHRTISNAQ-----KANILELKEVGLRQHQVMDVMERHHGGFDATGFVSRDL 218
Query: 173 YN--ARHRYKLSIRGSR*EMQELFQKFEENN--YVFNYRTVSGSRTVQDLFFAHPESVKL 228
YN R R K + G + + FQ ++++ + F Y T R ++ LF++ P+S
Sbjct: 219 YNYFTRLRKKHILGGDAERVIKYFQWRQKHDMEFFFEYETDEAGR-LKRLFWSDPQSRID 277
Query: 229 FNTFPTVLVMDSTYKTNQYKMPLFEIVGLTSTEMTYNVGFAFLTMEKEDNFTWGLQESAN 288
++ F V+V DSTY+ N+Y +P VG+ T G A + EK + W L++
Sbjct: 278 YDAFGDVVVFDSTYRVNRYNLPFIPFVGVNHHGSTIIFGCAIVADEKVATYEWILKQ--- 334
Query: 289 LLKCKKQM-PKVIVTDRDTGLMNAVANIFPKSTALVCEFHILKNVRARCILDCKVKDSKG 347
L C Q P+ ++TD D + A+A + P S +C +HI +N+
Sbjct: 335 FLDCMYQKHPRALITDGDNAMRRAIAAVMPDSEHWLCTWHIEQNM--------------A 380
Query: 348 KHVKSSDLVNTIMNAWEVIVSSP-SEELYADAVLQFR---KVCEQFPKFLKYVESTILEP 403
+H++ +++ + +V +P E + ++F+ K CE ++ +
Sbjct: 381 RHLRPD-----MLSDFRTLVHAPYDHEEFERKWVEFKVKHKGCEDNQWLVR------MYN 429
Query: 404 LKKKIVKAWTNRVMHFGCTTTNGVESAHGTLKGYLE 439
L+KK A+T + G + ES + L YL+
Sbjct: 430 LRKKWATAYTKGLFFLGMKSNQRSESLNSKLHRYLD 465
>ref|NP_915530.1| P0529E05.9 [Oryza sativa (japonica cultivar-group)]
Length = 773
Score = 83.2 bits (204), Expect = 3e-14
Identities = 85/360 (23%), Positives = 155/360 (42%), Gaps = 27/360 (7%)
Query: 27 KYEDREEMINWARGQAIEAGFTLII---DKSYSGSGRRKPKLVLAFERSGEYKGTKKSKR 83
+++D + + A + GF++ DKS + R K K V + E + + +R
Sbjct: 156 EFDDEDIAYEFYNRYAGDVGFSIRKFWHDKSSTNVIRTK-KFVCSREGFNKRNTSDACQR 214
Query: 84 EGTGSRKCGCPFRLRGYFSATKLWSLNIVSGLHNHKMEPKLEGHMLAG--RLTAKESKIV 141
+ +R GC + + T +++ S HNH++ + H+L R+T + +
Sbjct: 215 KRADTR-VGCMAEMTIKITPTGKYAIASFSNTHNHELITPSKAHLLRSQRRMTEAQKAQI 273
Query: 142 GDMTRNLIKPKNILLDLKGRWKDNRTVAKQIYNARHRYKLSIRGSR*E----------MQ 191
+ + ++ K ++ R R + + R YK +R R + +Q
Sbjct: 274 DILNDSGVRSKEGH-EVMSRQAGGR---QSLTFTRKDYKNYLRSKRMKSIQEGDTGAILQ 329
Query: 192 ELFQKFEENNYVFNYRTVSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQYKMPL 251
L K EN F V + ++F+A SV F+ F V+ D+TY+TN Y P
Sbjct: 330 YLQDKQMENPSFFYAIQVDEDEMMTNIFWADARSVLDFDYFGDVICFDTTYRTNNYGRPF 389
Query: 252 FEIVGLTSTEMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKKQMPKVIVTDRDTGLMNA 311
VG+ + T G A L E F W + + K+ P+ I+TD+ ++NA
Sbjct: 390 ALFVGVNHHKQTVVFGAALLYDETTSTFEWLFETFKRAMSGKE--PRTILTDQCAAIINA 447
Query: 312 VANIFPKSTALVCEFHILKNVR---ARCILDCKV-KDSKGKHVKSSDLVNTIMNAWEVIV 367
+ +FP ST +C +H+ +N + K K+ GK V + V+ ++AW ++
Sbjct: 448 IGTVFPNSTHRLCVWHMYQNAAVHLSHIFQGSKTFKNDFGKCVFDFEEVDEFISAWNKMI 507
>gb|AAL58182.1| putative far-red impaired response protein [Oryza sativa (japonica
cultivar-group)] gi|31433683|gb|AAP55167.1| putative
far-red impaired response protein [Oryza sativa
(japonica cultivar-group)] gi|37537156|ref|NP_922881.1|
putative far-red impaired response protein [Oryza sativa
(japonica cultivar-group)]
Length = 721
Score = 82.0 bits (201), Expect = 7e-14
Identities = 63/263 (23%), Positives = 116/263 (43%), Gaps = 15/263 (5%)
Query: 82 KREGTGSRKCGCPFRLRGYFSA-TKLWSLNIVSGLHNHKMEPK-----LEGHMLAGRLTA 135
KRE +CGC L + T +W ++ H+H++ L H
Sbjct: 81 KREPRALTRCGCKAMLEIELNGETGMWFVSGFEARHSHRLANLDLVAFLRSHREVNDAQK 140
Query: 136 KESKIVGDMTRNLIKPKNILLDLKGRWKDNRTVAKQIYNARHRYKLS-IRGSR*EM---Q 191
E+ +G + ++ + G + V + +YN RYK I G ++
Sbjct: 141 AEAVELGVGGLRTCQIMEVMENNNGGYDKVGFVTRDLYNFFARYKKKRIEGRDADLVVNH 200
Query: 192 ELFQKFEENNYVFNYRTVSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQYKMPL 251
+ Q+ ++ ++ F Y ++ +++LF+A +S + F V++ DSTY+ N+Y +P
Sbjct: 201 LMAQQEQDPDFFFRY-SIDEKGRLRNLFWADSQSQLDYEAFSDVVIFDSTYRVNRYNLPF 259
Query: 252 FEIVGLTSTEMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKKQM-PKVIVTDRDTGLMN 310
VG T G L+ E ++ W LQ LL+ Q PK ++TD D +
Sbjct: 260 VPFVGANHHRSTVIFGCGILSNESVSSYVWLLQ---TLLEAMHQKHPKSLITDGDASMAK 316
Query: 311 AVANIFPKSTALVCEFHILKNVR 333
A+ + P + +C +HI +N++
Sbjct: 317 AIRKVMPNTDHRLCSWHIEENMK 339
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.319 0.136 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,442,936,467
Number of Sequences: 2540612
Number of extensions: 62332069
Number of successful extensions: 143824
Number of sequences better than 10.0: 386
Number of HSP's better than 10.0 without gapping: 146
Number of HSP's successfully gapped in prelim test: 240
Number of HSP's that attempted gapping in prelim test: 143303
Number of HSP's gapped (non-prelim): 571
length of query: 840
length of database: 863,360,394
effective HSP length: 137
effective length of query: 703
effective length of database: 515,296,550
effective search space: 362253474650
effective search space used: 362253474650
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 80 (35.4 bits)
Medicago: description of AC138063.5


