

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137996.10 + phase: 0 /pseudo
(1594 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
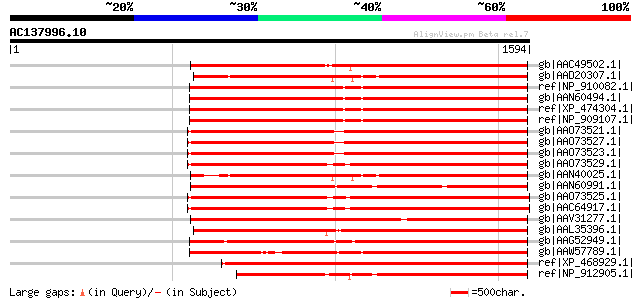
Sequences producing significant alignments: (bits) Value
gb|AAC49502.1| Pol [Zea mays] gi|7489803|pir||T04112 pol protein... 948 0.0
gb|AAD20307.1| copia-type pol polyprotein [Zea mays] 947 0.0
ref|NP_910082.1| putative gag-pol polyprotein [Oryza sativa (jap... 943 0.0
gb|AAN60494.1| Putative Zea mays retrotransposon Opie-2 [Oryza s... 939 0.0
ref|XP_474304.1| OSJNBb0004A17.2 [Oryza sativa (japonica cultiva... 939 0.0
ref|NP_909107.1| unnamed protein product [Oryza sativa (japonica... 939 0.0
gb|AAO73521.1| gag-pol polyprotein [Glycine max] 923 0.0
gb|AAO73527.1| gag-pol polyprotein [Glycine max] 921 0.0
gb|AAO73523.1| gag-pol polyprotein [Glycine max] 921 0.0
gb|AAO73529.1| gag-pol polyprotein [Glycine max] 920 0.0
gb|AAN40025.1| putative gag-pol polyprotein [Zea mays] 912 0.0
gb|AAN60991.1| Putative Zea mays retrotransposon Opie-2 [Oryza s... 911 0.0
gb|AAO73525.1| gag-pol polyprotein [Glycine max] 910 0.0
gb|AAC64917.1| gag-pol polyprotein [Glycine max] 908 0.0
gb|AAV31277.1| putative polyprotein [Oryza sativa (japonica cult... 907 0.0
gb|AAL35396.1| Opie2a pol [Zea mays] 907 0.0
gb|AAG52949.1| gag/pol polyprotein [Arabidopsis thaliana] 904 0.0
gb|AAW57789.1| putative polyprotein [Oryza sativa (japonica cult... 870 0.0
ref|XP_468929.1| putative polyprotein [Oryza sativa (japonica cu... 848 0.0
ref|NP_912905.1| unnamed protein product [Oryza sativa (japonica... 846 0.0
>gb|AAC49502.1| Pol [Zea mays] gi|7489803|pir||T04112 pol protein homolog - maize
retrotransposon Opie-2
Length = 1068
Score = 948 bits (2450), Expect = 0.0
Identities = 498/1061 (46%), Positives = 665/1061 (61%), Gaps = 34/1061 (3%)
Query: 554 SWYLDSGCSRHMTGEKALFLTLTMKDGGE--VKFGGNQTGKIIGTGTIGNSSI-SINNVW 610
SW +DSGC+ HMTGEK +F + + + FG GK+ G G I S+ SI+NV+
Sbjct: 10 SWIIDSGCTNHMTGEKKMFTSYVKNKDSQDSIIFGDGNQGKVKGLGKIAISNEHSISNVF 69
Query: 611 LVDGLKHNLLSISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKINFSDLADQ 670
LV+ L +NLLS+SQ C+ GY+ F+ + ++ + D S+ FKG +Y ++F+
Sbjct: 70 LVESLGYNLLSVSQLCNMGYNCLFTNIDVSVFRRCDGSLAFKGVLDGKLYLVDFAKEEAG 129
Query: 671 KVVCLLSMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGACQKGKIVK 730
CL++ W+WH+RL H + + K+ K + V GL N+ + D C ACQ GK V
Sbjct: 130 LDACLIAKTSMGWLWHRRLAHVGMKNLHKLLKGEHVIGLTNVQFEKDRPCAACQAGKQVG 189
Query: 731 SSFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWVKFIKSKDYAC 790
S +K++++TSRPLE+LH+DLFGPV S+ GSKYGLVIVDD+SR+TWV F++ K
Sbjct: 190 GSHHTKNVMTTSRPLEMLHMDLFGPVAYLSIGGSKYGLVIVDDFSRFTWVFFLQEKSETQ 249
Query: 791 EVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRTPQQNGVVE 850
F + Q+E ELK+ K+RSD G EF+N E F E+ GI HEFS+P TPQQNGVVE
Sbjct: 250 GTLKRFLRRAQNEFELKVKKIRSDNGSEFKNLQVEEFLEEEGIKHEFSAPYTPQQNGVVE 309
Query: 851 RKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYELFKGRRPNIS 910
RKNRTL +MARTM+ E + FW EAVNT+C+ NR+Y+ +L+ T+YEL G +PN+S
Sbjct: 310 RKNRTLIDMARTMLGEFKTPECFWTEAVNTACHAINRVYLHRILKNTSYELLTGNKPNVS 369
Query: 911 YFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVEESMHVKFDD 970
YF FG CYIL K KF KA G LGY +KAYRV+N + VE S V FD
Sbjct: 370 YFRVFGSKCYILVKKGRNSKFAPKAVEGFLLGYDSNTKAYRVFNKSSGLVEVSGDVVFD- 428
Query: 971 REPGSKTSEQSESNAGTTDSEDASESDQPSDSEKYTEVESSPEAEYNSEAEPSPKVQNES 1030
E EQ D +D E D P+ + + + E + +PSP
Sbjct: 429 -ETNGSPREQ------VVDCDDVDEEDIPTAAIRTMAIGEVRPQEQDEREQPSPSTMVHP 481
Query: 1031 ASEDFQDNTQQVI----------------------QPKFKHKSSHPEELIIGSKDSPRRT 1068
++D + QQ + Q + + HP + I+G T
Sbjct: 482 PTQDDEQVHQQEVCDQGGAQDDHVLEEEAQPAPPTQVRAMIQRDHPVDQILGDISKGVTT 541
Query: 1069 RSHFRQEESLIGLLSIIEPKTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTLL 1128
RS +S IEP VEEAL D W+LAMQEEL NFK P P ++ ++
Sbjct: 542 RSRLVNFCEHNSFVSSIEPFRVEEALLDPDWVLAMQEELNNFKRNEVWTLVPRP-KQNVV 600
Query: 1129 EQNGYSKTS*MNKEK*PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHG 1188
+ + K VA+GY+Q G+D+ ETFAPVARLE+IR+LL+YA +H
Sbjct: 601 GTKWVFRNKQDERGVVTRNKARLVAKGYAQVAGLDFEETFAPVARLESIRILLAYAAHHS 660
Query: 1189 IILYQMDVKSAFLNGVIEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLS 1248
LYQMDVKSAFLNG I+EEVYV+QPPGFED ++PDHV KL K+LYGLKQAPRAWY+ L
Sbjct: 661 FRLYQMDVKSAFLNGPIKEEVYVEQPPGFEDERYPDHVCKLSKALYGLKQAPRAWYECLR 720
Query: 1249 NFLIKNDFERGQVDTTLFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMS 1308
+FLI N F+ G+ D TLF +T D+ + QIYVDDIIFGSTN C+ FS++M +FEMS
Sbjct: 721 DFLIANAFKVGKADPTLFTKTCDGDLFVCQIYVDDIIFGSTNQKSCEEFSRVMTQKFEMS 780
Query: 1309 MMGELKFFLGIQINQSKEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGT 1368
MMGEL +FLG Q+ Q K+G ++ QTKYT++LLK+F ++D K TPM G
Sbjct: 781 MMGELNYFLGFQVKQLKDGTFISQTKYTQDLLKRFGMKDAKPAKTPMGTDGHTDLNKGGK 840
Query: 1369 VVDQKLYRGMIGSLLYLTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLG 1428
VDQK YR MIGSLLYL ASRP I+ SVC+CARFQSDP+E HL AVKRI RYL T G
Sbjct: 841 SVDQKAYRSMIGSLLYLCASRPDIMLSVCMCARFQSDPKECHLVAVKRILRYLVATPCFG 900
Query: 1429 LLYRKSLDYKLIGFCDADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEY 1488
L Y K + L+G+ D+DYAG +++RKSTSG CQFLG +L+SW SK+Q ++A+STAEAEY
Sbjct: 901 LWYPKGSTFDLVGYSDSDYAGCKVDRKSTSGTCQFLGRSLVSWNSKKQTSVALSTAEAEY 960
Query: 1489 ISAASCCTQLLWMKHQLEDYQINANSIPIYCDNTAAICLSKNPILHSRAKHIEIKHHFIR 1548
++A CC QLLWM+ L D+ N + +P+ CDN +AI +++NP+ HSR KHI+I+HHF+R
Sbjct: 961 VAAGQCCAQLLWMRQTLRDFGYNLSKVPLLCDNESAIRMAENPVEHSRTKHIDIRHHFLR 1020
Query: 1549 DYVQKGILDIQFIDIEHQWADIFTKPLSVERFDFIKKNLNM 1589
D+ QKG +++ + E+Q ADIFTKPL + F ++ LN+
Sbjct: 1021 DHQQKGDIEVFHVSTENQLADIFTKPLDEKTFCRLRSELNV 1061
>gb|AAD20307.1| copia-type pol polyprotein [Zea mays]
Length = 1063
Score = 947 bits (2449), Expect = 0.0
Identities = 512/1066 (48%), Positives = 671/1066 (62%), Gaps = 52/1066 (4%)
Query: 565 MTGEKALFLTLTMKDGGE--VKFGGNQTGKIIGTGTIGNS-SISINNVWLVDGLKHNLLS 621
MTGEK +F + + + FG G + G G I S SI+NV+LVD L +NLLS
Sbjct: 1 MTGEKRMFSSYEKNQDPQRAITFGDGNQGLVKGLGKIAISPDHSISNVFLVDSLDYNLLS 60
Query: 622 ISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKINFSDLADQKVVCLLSMNDK 681
+SQ C GY+ F+ T+ + D SI FKG +Y ++F D A+ CL++ +
Sbjct: 61 VSQLCQMGYNCLFTDIGVTVFRRSDDSIAFKGVLEGQLYLVDF-DRAELDT-CLIAKTNM 118
Query: 682 KWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGACQKGKIVKSSFKSKDIVST 741
W+WH+RL H + + K+ K + + GL N+ + D +C ACQ GK V + K+I++T
Sbjct: 119 GWLWHRRLAHVGMKNLHKLLKGEHILGLTNVHFEKDRICSACQAGKQVGTHHPHKNIMTT 178
Query: 742 SRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWVKFIKSKDYACEVFSSFCTQIQ 801
RPLELLH+DLFGP+ S+ GSKY LVIVDDYSR+TWV F++ K E F + Q
Sbjct: 179 DRPLELLHMDLFGPIAYISIGGSKYCLVIVDDYSRFTWVFFLQEKSQTQETLKGFLRRAQ 238
Query: 802 SEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRTPQQNGVVERKNRTLQEMAR 861
+E L+I K+RSD G EF+N E F E+ GI HEFSSP TPQQNGVVERKNRTL +MAR
Sbjct: 239 NEFGLRIKKIRSDNGTEFKNSQIESFLEEEGIKHEFSSPYTPQQNGVVERKNRTLLDMAR 298
Query: 862 TMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYELFKGRRPNISYFHQFGCTCYI 921
TM+ E FWAEAVNT+CY NR+Y+ +L+KT+YEL G++PNISYF FG C+I
Sbjct: 299 TMLDEYKTPDRFWAEAVNTACYAINRLYLHRILKKTSYELLTGKKPNISYFRVFGSKCFI 358
Query: 922 LNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVEESMHVKFDDREPGSKTSEQS 981
L + KF K G LGY ++AYRV+N T VE S V FD+ GS+ +
Sbjct: 359 LIKRGRKSKFAPKTVEGFLLGYDSNTRAYRVFNKSTGLVEVSCDVVFDETN-GSQVEQVD 417
Query: 982 ESNAGTTDS-------------------EDASESDQPSDSEKYT-----EVESSPEAEYN 1017
G + E S DQPS S + + E E+ + E N
Sbjct: 418 LDEIGEEQAPCIALRNMSIGDVCPKESEEPPSTQDQPSSSMQASPPTQNEDEAQNDEEQN 477
Query: 1018 SEAEPSPKVQNESASEDFQDNTQQVIQPKFKH-------KSSHPEELIIGSKDSPRRTRS 1070
E EP N+ + + +P+ H + HP + I+G TRS
Sbjct: 478 QEDEPPQDDSNDQGGDTNDQEKEDEEEPRPPHPRVHQAIQRDHPVDTILGDIHKGVTTRS 537
Query: 1071 ---HFRQEESLIGLLSIIEPKTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTL 1127
HF + S + S IEP VEEAL D W++AMQEEL NF P P + +
Sbjct: 538 RVAHFCEHYSFV---SSIEPHRVEEALQDSDWVVAMQEELNNFTRNEVWHLVPRPNQNVV 594
Query: 1128 LEQNGYSKTS*MNKEK*----PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSY 1183
+K NK+ K VA+GYSQ EG+D+ ET+APVARLE+IR+LL+Y
Sbjct: 595 -----GTKWVFRNKQDEHGVVTRNKARLVAKGYSQVEGLDFGETYAPVARLESIRILLAY 649
Query: 1184 AINHGIILYQMDVKSAFLNGVIEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAW 1243
A HG LYQMDVKSAFLNG I+EEVYV+QPPGFED ++P+HVY+L K+LYGLKQAPRAW
Sbjct: 650 ATYHGFKLYQMDVKSAFLNGPIKEEVYVEQPPGFEDSEYPNHVYRLSKALYGLKQAPRAW 709
Query: 1244 YDRLSNFLIKNDFERGQVDTTLFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQD 1303
Y+ L +FLI N F+ G+ D TLF +TL+ D+ + QIYVDDIIFGSTN S C+ FS++M
Sbjct: 710 YECLRDFLIANGFKVGKADPTLFTKTLENDLFVCQIYVDDIIFGSTNKSTCEEFSRIMTQ 769
Query: 1304 EFEMSMMGELKFFLGIQINQSKEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSK 1363
+FEMSMMGELK+FLG Q+ Q +EG ++ QTKYT+++L KF ++D K + TPM L
Sbjct: 770 KFEMSMMGELKYFLGFQVKQLQEGTFICQTKYTQDILTKFGMKDAKPIKTPMGTNGHLDL 829
Query: 1364 EDTGTVVDQKLYRGMIGSLLYLTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKG 1423
+ G VDQK+YR MIGSLLYL ASRP I+ SVC+CARFQSDP+ESHLTAVKRI RYL
Sbjct: 830 DTGGKSVDQKVYRSMIGSLLYLCASRPDIMLSVCMCARFQSDPKESHLTAVKRILRYLAY 889
Query: 1424 TTNLGLLYRKSLDYKLIGFCDADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMST 1483
T GL Y + + LIG+ DAD+AG +I RKSTSG CQFLG +L+SWASK+Q ++A+ST
Sbjct: 890 TPKFGLWYPRGSTFDLIGYSDADWAGCKINRKSTSGTCQFLGRSLVSWASKKQNSVALST 949
Query: 1484 AEAEYISAASCCTQLLWMKHQLEDYQINANSIPIYCDNTAAICLSKNPILHSRAKHIEIK 1543
AEAEYI+A CC QLLWM+ L DY +P+ CDN +AI ++ NP+ HSR KHI I+
Sbjct: 950 AEAEYIAAGHCCAQLLWMRQTLLDYGYKLTKVPLLCDNESAIKMADNPVEHSRTKHIAIR 1009
Query: 1544 HHFIRDYVQKGILDIQFIDIEHQWADIFTKPLSVERFDFIKKNLNM 1589
+HF+RD+ QKG ++I +I+ + Q ADIFTKPL + F ++ LN+
Sbjct: 1010 YHFLRDHQQKGDIEISYINTKDQLADIFTKPLDEQSFTRLRHELNI 1055
>ref|NP_910082.1| putative gag-pol polyprotein [Oryza sativa (japonica cultivar-group)]
gi|28269414|gb|AAO37957.1| putative gag-pol polyprotein
[Oryza sativa (japonica cultivar-group)]
Length = 1969
Score = 943 bits (2438), Expect = 0.0
Identities = 495/1047 (47%), Positives = 668/1047 (63%), Gaps = 16/1047 (1%)
Query: 551 KQRSWYLDSGCSRHMTGEKALFLTLTMKDGGE-VKFGGNQTGKIIGTGTIG-NSSISINN 608
+ SW +DSGCSRHMTGE F +LT G E + FG +G+++ GTI N + +
Sbjct: 750 RSNSWLVDSGCSRHMTGEAKWFTSLTRASGDETITFGDASSGRVMAKGTIKVNDKFMLKD 809
Query: 609 VWLVDGLKHNLLSISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKINFSDLA 668
V LV LK+NLLS+SQ CD +V F K +++ + + F RV V+ NF A
Sbjct: 810 VALVSKLKYNLLSVSQLCDENLEVRFKKDRSRVLDASESPV-FDISRVGRVFFANFDSSA 868
Query: 669 DQKVVCLL-SMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGACQKGK 727
CL+ S N + WH+RLGH + +S+IS + L++GLP + D +C C+ GK
Sbjct: 869 PGPSRCLIASENRDLFFWHRRLGHIGFDHLSRISGMDLIRGLPKLKVQKDLVCAPCRHGK 928
Query: 728 IVKSSFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWVKFIKSKD 787
+ SS K +V T P +LLH+D GP S+ G Y LV+VDD+SR++WV F++SK+
Sbjct: 929 MTSSSHKPVTMVMTDGPGQLLHMDTVGPARVQSVGGKWYVLVVVDDFSRYSWVYFLESKE 988
Query: 788 YACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRTPQQNG 847
F S + E + +RSD G EF+N FE FC+ G+ H+FSSP PQQNG
Sbjct: 989 ETFGFFQSLARSLALEFPGALRAIRSDNGSEFKNSAFESFCDSSGVEHQFSSPYVPQQNG 1048
Query: 848 VVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYELFKGRRP 907
VVERKNRTL EMARTM+ E + FW EA++ +C+I NR+++R +L KT YEL GRRP
Sbjct: 1049 VVERKNRTLVEMARTMLDEFTTPRKFWTEAISAACFISNRVFLRTILHKTPYELRFGRRP 1108
Query: 908 NISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVEESMHVK 967
+S+ FGC C++L + + L KF++++ GIFLGY+ S+AYRVY T + E+ V
Sbjct: 1109 KVSHLRVFGCKCFVLKSGN-LDKFESRSLDGIFLGYATHSRAYRVYVLSTNKIVETCEVT 1167
Query: 968 FDDREPGSKTSEQSESNAGTTDSEDASESDQ-----PSDSEKYTEVESSPEAEYNSEAEP 1022
FD+ PG++ + ED+ + D P DS + SP S P
Sbjct: 1168 FDEASPGARPEISGVPDESIFVDEDSDDDDDDSIPPPLDSTPPVQETGSPSTTSPSGDAP 1227
Query: 1023 SPKVQNESASEDFQDNTQQVIQPKFKHKSSHPEELIIGSKDSPRRTRSHFRQEESLIGLL 1082
+ + SA+E+ T P+ P+ +I G + R RS+ + +
Sbjct: 1228 TT---SSSAAEEIDGGTSGPTAPRHIQNRHPPDSMIGGLGERVTRNRSYELVNSAFV--- 1281
Query: 1083 SIIEPKTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTLLEQNGYSKTS*MNKE 1142
+ EPK V ALSD+ W+ AM EEL NF+ P ++ K
Sbjct: 1282 ASFEPKNVCHALSDENWVNAMHEELENFERNKVWSLVEPPLGFNVIGTKWVFKNKLGEDG 1341
Query: 1143 K*PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQMDVKSAFLN 1202
K VAQG++Q EG+D+ ETFAPVARLEAIR+LL++A + G L+QMDVKSAFLN
Sbjct: 1342 SIVRNKARLVAQGFTQVEGLDFEETFAPVARLEAIRILLAFAASKGFKLFQMDVKSAFLN 1401
Query: 1203 GVIEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDFERGQVD 1262
GVIEEEVYVKQPPGFE+ K P+HV+KL+K+LYGLKQAPRAWY+RL FL++N FE G VD
Sbjct: 1402 GVIEEEVYVKQPPGFENPKFPNHVFKLEKALYGLKQAPRAWYERLKTFLLQNGFEMGAVD 1461
Query: 1263 TTLFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELKFFLGIQIN 1322
TLF D L+VQIYVDDIIFG ++ +L FS +M EFEMSMMGEL FFLG+QI
Sbjct: 1462 KTLFTLHSGIDFLLVQIYVDDIIFGGSSHALVAQFSDVMSREFEMSMMGELTFFLGLQIK 1521
Query: 1323 QSKEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGTVVDQKLYRGMIGSL 1382
Q+KEG++VHQTKY+KELLKKF + DCK + TPM T +L ++ G VDQ+ YR MIGSL
Sbjct: 1522 QTKEGIFVHQTKYSKELLKKFDMADCKPIATPMATTSSLGPDEDGEEVDQREYRSMIGSL 1581
Query: 1383 LYLTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGLLYRKSLDYKLIGF 1442
LYLTASRP I FSVCLCARFQ+ PR SH AVKRIFRY+K T G+ Y S + F
Sbjct: 1582 LYLTASRPDIHFSVCLCARFQASPRTSHRQAVKRIFRYIKSTLEYGIWYSCSSALSVRAF 1641
Query: 1443 CDADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEYISAASCCTQLLWMK 1502
DAD+AG +I+RKSTSG C FLG +L+SW+S++Q+++A STAEAEY++AAS C+Q+LWM
Sbjct: 1642 SDADFAGCKIDRKSTSGTCHFLGTSLVSWSSRKQSSVAQSTAEAEYVAAASACSQVLWMI 1701
Query: 1503 HQLEDYQINANSIPIYCDNTAAICLSKNPILHSRAKHIEIKHHFIRDYVQKGILDIQFID 1562
L+DY ++ + +P+ CDNT+AI ++KNP+ HSR KHIEI++HF+RD V+KG + ++F++
Sbjct: 1702 STLKDYGLSFSGVPLLCDNTSAINIAKNPVQHSRTKHIEIRYHFLRDNVEKGTIVLEFVE 1761
Query: 1563 IEHQWADIFTKPLSVERFDFIKKNLNM 1589
E Q ADIFTKPL RF+F++ L +
Sbjct: 1762 SEKQLADIFTKPLDRSRFEFLRSELGV 1788
>gb|AAN60494.1| Putative Zea mays retrotransposon Opie-2 [Oryza sativa (japonica
cultivar-group)] gi|34902378|ref|NP_912535.1| Putative
Zea mays retrotransposon Opie-2 [Oryza sativa (japonica
cultivar-group)]
Length = 2145
Score = 939 bits (2427), Expect = 0.0
Identities = 493/1047 (47%), Positives = 667/1047 (63%), Gaps = 16/1047 (1%)
Query: 551 KQRSWYLDSGCSRHMTGEKALFLTLTMKDGGE-VKFGGNQTGKIIGTGTIG-NSSISINN 608
+ SW +DSGCSRHMTGE F +LT E + FG +G+++ GTI N + +
Sbjct: 750 RSNSWLVDSGCSRHMTGEAKWFTSLTRASSDETITFGDASSGRVMAKGTIKVNDKFMLKD 809
Query: 609 VWLVDGLKHNLLSISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKINFSDLA 668
V LV LK+NLLS+SQ CD +V F K +++ + + F RV V+ NF A
Sbjct: 810 VALVSKLKYNLLSVSQLCDENLEVRFKKDRSRVLDASESPV-FDISRVGRVFFANFDSSA 868
Query: 669 DQKVVCLL-SMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGACQKGK 727
CL+ S N + WH+RLGH + +S+IS + L++GLP + D +C C+ GK
Sbjct: 869 PGPSRCLIASENRDLFFWHRRLGHIGFDHLSRISGMDLIRGLPKLKVPKDLVCAPCRHGK 928
Query: 728 IVKSSFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWVKFIKSKD 787
+ SS K +V T P +LLH+D GP S+ G Y LV+VDD+SR++WV F++SK+
Sbjct: 929 MTSSSHKPVTMVMTDGPGQLLHMDTVGPARVQSVGGKWYVLVVVDDFSRYSWVYFLESKE 988
Query: 788 YACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRTPQQNG 847
F S + E + +RSD G EF+N FE FC+ G+ H+FSSP PQQNG
Sbjct: 989 ETFGFFQSLARSLALEFPGALRAIRSDNGSEFKNSAFESFCDSSGVEHQFSSPYVPQQNG 1048
Query: 848 VVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYELFKGRRP 907
VVERKNRTL EMARTM+ E + FW EA++ +C+I NR+++R +L KT YEL GRRP
Sbjct: 1049 VVERKNRTLVEMARTMLDEFTTPRKFWTEAISAACFISNRVFLRTILHKTPYELRFGRRP 1108
Query: 908 NISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVEESMHVK 967
+S+ FGC C++L + + L KF++++ GIFLGY+ S+AYRVY T + E+ V
Sbjct: 1109 KVSHLRVFGCKCFVLKSGN-LDKFESRSLDGIFLGYATHSRAYRVYVLSTNKIVETCEVT 1167
Query: 968 FDDREPGSKTSEQSESNAGTTDSEDASESDQ-----PSDSEKYTEVESSPEAEYNSEAEP 1022
FD+ PG++ + ED+ + D P DS + SP S P
Sbjct: 1168 FDEASPGARPEISGVPDESIFVDEDSDDDDDDSIPPPLDSTPPVQETGSPSTTSPSGDAP 1227
Query: 1023 SPKVQNESASEDFQDNTQQVIQPKFKHKSSHPEELIIGSKDSPRRTRSHFRQEESLIGLL 1082
+ + SA+E+ T P+ P+ +I G + R RS+ + +
Sbjct: 1228 TT---SSSAAEEIDGGTSGPTAPRHIQNRHPPDSMIGGLGERVTRNRSYELVNSAFV--- 1281
Query: 1083 SIIEPKTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTLLEQNGYSKTS*MNKE 1142
+ EPK V ALSD+ W+ AM EEL NF+ P ++ K
Sbjct: 1282 ASFEPKNVCHALSDENWVNAMHEELENFERNKVWSLVEPPLGFNVIGTKWVFKNKLGEDG 1341
Query: 1143 K*PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQMDVKSAFLN 1202
K VAQG++Q EG+D+ ETFAPVARLEAIR+LL++A + G L+QMDVKSAFLN
Sbjct: 1342 SIVRNKARLVAQGFTQVEGLDFEETFAPVARLEAIRILLAFAASKGFKLFQMDVKSAFLN 1401
Query: 1203 GVIEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDFERGQVD 1262
GVIEEEVYVKQPPGFE+ K P+HV+KL+K+LYGLKQAPRAWY+RL FL++N FE G VD
Sbjct: 1402 GVIEEEVYVKQPPGFENPKFPNHVFKLEKALYGLKQAPRAWYERLKTFLLQNGFEMGAVD 1461
Query: 1263 TTLFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELKFFLGIQIN 1322
TLF D L+VQIYVDDIIFG ++ +L FS +M EFEMSMMGEL FFLG+QI
Sbjct: 1462 KTLFTLHSGIDFLLVQIYVDDIIFGGSSHALVAQFSDVMSREFEMSMMGELTFFLGLQIK 1521
Query: 1323 QSKEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGTVVDQKLYRGMIGSL 1382
Q+KEG++VHQTKY+KELLKKF + DCK + TPM T +L ++ G VDQ+ YR MIGSL
Sbjct: 1522 QTKEGIFVHQTKYSKELLKKFDMADCKPIATPMATTSSLGPDEDGEEVDQREYRSMIGSL 1581
Query: 1383 LYLTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGLLYRKSLDYKLIGF 1442
LYLTASRP I FSVCLCARFQ+ PR SH AVKR+FRY+K T G+ Y S + F
Sbjct: 1582 LYLTASRPDIHFSVCLCARFQASPRTSHRQAVKRMFRYIKSTLEYGIWYSCSSALSVRAF 1641
Query: 1443 CDADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEYISAASCCTQLLWMK 1502
DAD+AG +I+RKSTSG C FLG +L+SW+S++Q+++A STAEAEY++AAS C+Q+LWM
Sbjct: 1642 SDADFAGCKIDRKSTSGTCHFLGTSLVSWSSRKQSSVAQSTAEAEYVAAASACSQVLWMI 1701
Query: 1503 HQLEDYQINANSIPIYCDNTAAICLSKNPILHSRAKHIEIKHHFIRDYVQKGILDIQFID 1562
L+DY ++ + +P+ CDNT+AI ++KNP+ HSR KHIEI++HF+RD V+KG + ++F++
Sbjct: 1702 STLKDYGLSFSGVPLLCDNTSAINIAKNPVQHSRTKHIEIRYHFLRDNVEKGTIVLEFVE 1761
Query: 1563 IEHQWADIFTKPLSVERFDFIKKNLNM 1589
E Q ADIFTKPL RF+F++ L +
Sbjct: 1762 SEKQLADIFTKPLDRSRFEFLRSELGV 1788
>ref|XP_474304.1| OSJNBb0004A17.2 [Oryza sativa (japonica cultivar-group)]
gi|32488723|emb|CAE03600.1| OSJNBb0004A17.2 [Oryza sativa
(japonica cultivar-group)]
Length = 1877
Score = 939 bits (2427), Expect = 0.0
Identities = 493/1047 (47%), Positives = 667/1047 (63%), Gaps = 16/1047 (1%)
Query: 551 KQRSWYLDSGCSRHMTGEKALFLTLTMKDGGE-VKFGGNQTGKIIGTGTIG-NSSISINN 608
+ SW +DSGCSRHMTGE F +LT E + FG +G+++ GTI N + +
Sbjct: 832 RSNSWLVDSGCSRHMTGEAKWFTSLTRASSDETITFGDASSGRVMAKGTIKVNDKFMLKD 891
Query: 609 VWLVDGLKHNLLSISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKINFSDLA 668
V LV LK+NLLS+SQ CD +V F K +++ + + F RV V+ NF A
Sbjct: 892 VALVSKLKYNLLSVSQLCDENLEVRFKKDRSRVLDASESPV-FDISRVGRVFFANFDSSA 950
Query: 669 DQKVVCLL-SMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGACQKGK 727
CL+ S N + WH+RLGH + +S+IS + L++GLP + D +C C+ GK
Sbjct: 951 PGPSRCLIASENRDLFFWHRRLGHIGFDHLSRISGMDLIRGLPKLKVPKDLVCAPCRHGK 1010
Query: 728 IVKSSFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWVKFIKSKD 787
+ SS K +V T P +LLH+D GP S+ G Y LV+VDD+SR++WV F++SK+
Sbjct: 1011 MTSSSHKPVTMVMTDGPGQLLHMDTVGPARVQSVGGKWYVLVVVDDFSRYSWVYFLESKE 1070
Query: 788 YACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRTPQQNG 847
F S + E + +RSD G EF+N FE FC+ G+ H+FSSP PQQNG
Sbjct: 1071 ETFGFFQSLARSLALEFPGALRAIRSDNGSEFKNSAFESFCDSSGVEHQFSSPYVPQQNG 1130
Query: 848 VVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYELFKGRRP 907
VVERKNRTL EMARTM+ E + FW EA++ +C+I NR+++R +L KT YEL GRRP
Sbjct: 1131 VVERKNRTLVEMARTMLDEFTTPRKFWTEAISAACFISNRVFLRTILHKTPYELRFGRRP 1190
Query: 908 NISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVEESMHVK 967
+S+ FGC C++L + + L KF++++ GIFLGY+ S+AYRVY T + E+ V
Sbjct: 1191 KVSHLRVFGCKCFVLKSGN-LDKFESRSLDGIFLGYATHSRAYRVYVLSTNKIVETCEVT 1249
Query: 968 FDDREPGSKTSEQSESNAGTTDSEDASESDQ-----PSDSEKYTEVESSPEAEYNSEAEP 1022
FD+ PG++ + ED+ + D P DS + SP S P
Sbjct: 1250 FDEASPGARPEISGVPDESIFVDEDSDDDDDDSIPPPLDSTPPVQETGSPSTTSPSGDAP 1309
Query: 1023 SPKVQNESASEDFQDNTQQVIQPKFKHKSSHPEELIIGSKDSPRRTRSHFRQEESLIGLL 1082
+ + SA+E+ T P+ P+ +I G + R RS+ + +
Sbjct: 1310 TT---SSSAAEEIDGGTSGPTAPRHIQNRHPPDSMIGGLGERVTRNRSYELVNSAFV--- 1363
Query: 1083 SIIEPKTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTLLEQNGYSKTS*MNKE 1142
+ EPK V ALSD+ W+ AM EEL NF+ P ++ K
Sbjct: 1364 ASFEPKNVCHALSDENWVNAMHEELENFERNKVWSLVEPPLGFNVIGTKWVFKNKLGEDG 1423
Query: 1143 K*PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQMDVKSAFLN 1202
K VAQG++Q EG+D+ ETFAPVARLEAIR+LL++A + G L+QMDVKSAFLN
Sbjct: 1424 SIVRNKARLVAQGFTQVEGLDFEETFAPVARLEAIRILLAFAASKGFKLFQMDVKSAFLN 1483
Query: 1203 GVIEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDFERGQVD 1262
GVIEEEVYVKQPPGFE+ K P+HV+KL+K+LYGLKQAPRAWY+RL FL++N FE G VD
Sbjct: 1484 GVIEEEVYVKQPPGFENPKFPNHVFKLEKALYGLKQAPRAWYERLKTFLLQNGFEMGAVD 1543
Query: 1263 TTLFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELKFFLGIQIN 1322
TLF D L+VQIYVDDIIFG ++ +L FS +M EFEMSMMGEL FFLG+QI
Sbjct: 1544 KTLFTLHSGIDFLLVQIYVDDIIFGGSSHALVAQFSDVMSREFEMSMMGELTFFLGLQIK 1603
Query: 1323 QSKEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGTVVDQKLYRGMIGSL 1382
Q+KEG++VHQTKY+KELLKKF + DCK + TPM T +L ++ G VDQ+ YR MIGSL
Sbjct: 1604 QTKEGIFVHQTKYSKELLKKFDMADCKPIATPMATTSSLGPDEDGEEVDQREYRSMIGSL 1663
Query: 1383 LYLTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGLLYRKSLDYKLIGF 1442
LYLTASRP I FSVCLCARFQ+ PR SH AVKR+FRY+K T G+ Y S + F
Sbjct: 1664 LYLTASRPDIHFSVCLCARFQASPRTSHRQAVKRMFRYIKSTLEYGIWYSCSSALSVRAF 1723
Query: 1443 CDADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEYISAASCCTQLLWMK 1502
DAD+AG +I+RKSTSG C FLG +L+SW+S++Q+++A STAEAEY++AAS C+Q+LWM
Sbjct: 1724 SDADFAGCKIDRKSTSGTCHFLGTSLVSWSSRKQSSVAQSTAEAEYVAAASACSQVLWMI 1783
Query: 1503 HQLEDYQINANSIPIYCDNTAAICLSKNPILHSRAKHIEIKHHFIRDYVQKGILDIQFID 1562
L+DY ++ + +P+ CDNT+AI ++KNP+ HSR KHIEI++HF+RD V+KG + ++F++
Sbjct: 1784 STLKDYGLSFSGVPLLCDNTSAINIAKNPVQHSRTKHIEIRYHFLRDNVEKGTIVLEFVE 1843
Query: 1563 IEHQWADIFTKPLSVERFDFIKKNLNM 1589
E Q ADIFTKPL RF+F++ L +
Sbjct: 1844 SEKQLADIFTKPLDRSRFEFLRSELGV 1870
>ref|NP_909107.1| unnamed protein product [Oryza sativa (japonica cultivar-group)]
Length = 1410
Score = 939 bits (2426), Expect = 0.0
Identities = 494/1047 (47%), Positives = 667/1047 (63%), Gaps = 16/1047 (1%)
Query: 551 KQRSWYLDSGCSRHMTGEKALFLTLTMKDGGE-VKFGGNQTGKIIGTGTIG-NSSISINN 608
+ SW +DSGCSRHMTGE F +LT G E + FG +G+++ GTI N + +
Sbjct: 365 RSNSWLVDSGCSRHMTGEAKWFTSLTRASGDETITFGDASSGRVMAKGTIKVNDKFMLKD 424
Query: 609 VWLVDGLKHNLLSISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKINFSDLA 668
V LV LK+NLLS+SQ CD +V F K +++ + + F RV V+ NF A
Sbjct: 425 VALVSKLKYNLLSVSQLCDENLEVRFKKDRSRVLDASESPV-FDISRVGRVFFANFDSSA 483
Query: 669 DQKVVCLL-SMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGACQKGK 727
CL+ S N + WH+RLGH + +S+IS + L++GLP + D +C C+ GK
Sbjct: 484 PGPSRCLVASENRDLFFWHRRLGHIGFDHLSRISGMDLIRGLPKLKAPKDLVCAPCRHGK 543
Query: 728 IVKSSFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWVKFIKSKD 787
+ SS K +V T P +LLH++ GP S+ G Y LV+VDD+SR++WV F++SK+
Sbjct: 544 MTSSSHKPVTMVMTDGPGQLLHMNTVGPARVQSVGGKWYVLVVVDDFSRYSWVYFLESKE 603
Query: 788 YACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRTPQQNG 847
F S + E + +RSD G EF+N FE FC+ G+ H+FSSP PQQNG
Sbjct: 604 ETFGFFQSLARSLALEFPGALRAIRSDNGSEFKNSAFESFCDSSGVEHQFSSPYVPQQNG 663
Query: 848 VVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYELFKGRRP 907
VVERKNRTL EMARTM+ E + FW EA++ +C+I NR+++R +L KT YEL GRRP
Sbjct: 664 VVERKNRTLVEMARTMLDEFTTPRKFWTEAISAACFISNRVFLRTILHKTPYELRFGRRP 723
Query: 908 NISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVEESMHVK 967
+S+ FGC C++L + + L KF++++ GIFLGY+ S+AYRVY T + E+ V
Sbjct: 724 KVSHLRVFGCKCFVLKSGN-LDKFESRSLDGIFLGYATHSRAYRVYVLSTNKIVETCEVT 782
Query: 968 FDDREPGSKTSEQSESNAGTTDSEDASESDQ-----PSDSEKYTEVESSPEAEYNSEAEP 1022
FD+ PG++ + ED+ + D P DS + SP S P
Sbjct: 783 FDEASPGARPEISGVLDESIFVDEDSDDDDDDSIPPPLDSTPPVQETGSPSTTSPSGDAP 842
Query: 1023 SPKVQNESASEDFQDNTQQVIQPKFKHKSSHPEELIIGSKDSPRRTRSHFRQEESLIGLL 1082
+ + SA+E+ T P+ P+ +I G + R RS+ + +
Sbjct: 843 TT---SSSAAEEIDGGTSGPTAPRHIQNRHPPDSMIGGLGERVTRNRSYDLVNSAFV--- 896
Query: 1083 SIIEPKTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTLLEQNGYSKTS*MNKE 1142
+ EPK V ALSD+ W+ AM EEL NF+ P ++ K
Sbjct: 897 ASFEPKNVCHALSDENWVNAMHEELENFERNKVWSLVEPPLGFNVIGTKWVFKNKLGEDG 956
Query: 1143 K*PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQMDVKSAFLN 1202
K VAQG++Q EG+D+ ETFAPVARLEAIR+LL++A + G L+QMDVKSAFLN
Sbjct: 957 SIVRNKARLVAQGFTQVEGLDFEETFAPVARLEAIRILLAFAASKGFKLFQMDVKSAFLN 1016
Query: 1203 GVIEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDFERGQVD 1262
GVIEEEVYVKQPPGFE+ K P+HV+KL K+LYGLKQAPRAWY+RL FL++N FE G VD
Sbjct: 1017 GVIEEEVYVKQPPGFENPKFPNHVFKLDKALYGLKQAPRAWYERLKTFLLQNGFEMGAVD 1076
Query: 1263 TTLFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELKFFLGIQIN 1322
TLF D L+VQIYVDDIIFG ++ +L FS +M EFEMSMMGEL FFLG+QI
Sbjct: 1077 KTLFTLHSGIDFLLVQIYVDDIIFGGSSHALVAQFSDVMSREFEMSMMGELTFFLGLQIK 1136
Query: 1323 QSKEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGTVVDQKLYRGMIGSL 1382
Q+KEG++VHQTKY+KELLKKF + DCK + TPM T +L ++ G VDQ+ YR MIGSL
Sbjct: 1137 QTKEGIFVHQTKYSKELLKKFDMADCKPIATPMATTSSLGPDEDGEEVDQREYRSMIGSL 1196
Query: 1383 LYLTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGLLYRKSLDYKLIGF 1442
LYLTASRP I FSVCLCARFQ+ PR SH AVKRIFRY+K T G+ Y S + F
Sbjct: 1197 LYLTASRPDIHFSVCLCARFQASPRTSHRQAVKRIFRYIKSTLEYGIWYSCSSALSVRAF 1256
Query: 1443 CDADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEYISAASCCTQLLWMK 1502
DAD+AG +I+RKSTSG C FLG +L+SW+S++Q+++A STAEAEY++AAS C+Q+LWM
Sbjct: 1257 SDADFAGCKIDRKSTSGTCHFLGTSLVSWSSRKQSSVAQSTAEAEYVAAASACSQVLWMI 1316
Query: 1503 HQLEDYQINANSIPIYCDNTAAICLSKNPILHSRAKHIEIKHHFIRDYVQKGILDIQFID 1562
L+DY ++ + +P+ CDNT+AI ++KNP+ HSR KHIEI++HF+RD V+KG + ++F++
Sbjct: 1317 STLKDYGLSFSGVPLLCDNTSAINIAKNPVQHSRTKHIEIRYHFLRDNVEKGTIVLEFVE 1376
Query: 1563 IEHQWADIFTKPLSVERFDFIKKNLNM 1589
E Q ADIFTKPL RF+F++ L +
Sbjct: 1377 SEKQLADIFTKPLDRSRFEFLRSELGV 1403
>gb|AAO73521.1| gag-pol polyprotein [Glycine max]
Length = 1574
Score = 923 bits (2385), Expect = 0.0
Identities = 480/1045 (45%), Positives = 656/1045 (61%), Gaps = 29/1045 (2%)
Query: 546 LRAREKQRSWYLDSGCSRHMTGEKALFLTLTMKDGGEVKFGGNQTGKIIGTGTIGNSSI- 604
LRA K+ WYLDSGCSRHMTG K L + V FG GKIIG G + + +
Sbjct: 552 LRASAKE-DWYLDSGCSRHMTGVKEFLLNIEPCSTSYVTFGDGSKGKIIGMGKLVHDGLP 610
Query: 605 SINNVWLVDGLKHNLLSISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKINF 664
S+N V LV GL NL+SISQ CD G++V F+K+ C LV + + KG R ++ +
Sbjct: 611 SLNKVLLVKGLTANLISISQLCDEGFNVNFTKSEC-LVTNEKSEVLMKGSRSKDNCYLWT 669
Query: 665 SDLADQKVVCLLSMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGACQ 724
CL S D+ +WH+R GH + R + KI V+G+PN+ +CG CQ
Sbjct: 670 PQETSYSSTCLSSKEDEVRIWHQRFGHLHLRGMKKIIDKGAVRGIPNLKIEEGRICGECQ 729
Query: 725 KGKIVKSSFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWVKFIK 784
GK VK S + +TSR LELLH+DL GP+ SL G +Y V+VDD+SR+TWVKFI+
Sbjct: 730 IGKQVKMSHQKLQHQTTSRVLELLHMDLMGPMQVESLGGKRYAYVVVDDFSRFTWVKFIR 789
Query: 785 SKDYACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRTPQ 844
K EVF ++Q EK+ I ++RSD+G EFEN FC GI HEFS+ TPQ
Sbjct: 790 EKSETFEVFKELSLRLQREKDCVIKRIRSDHGREFENSRLTEFCTSEGITHEFSAAITPQ 849
Query: 845 QNGVVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYELFKG 904
QNG+VERKNRTLQE AR M+H L + WAEA+NT+CYI NR+ +R T YE++KG
Sbjct: 850 QNGIVERKNRTLQEAARVMLHAKELPYNLWAEAMNTACYIHNRVTLRRGTPTTLYEIWKG 909
Query: 905 RRPNISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVEESM 964
R+P++ +FH FG CYIL ++ +K D K+ GIFLGYS S+AYRV+NS T+ V ES+
Sbjct: 910 RKPSVKHFHIFGSPCYILADREQRRKMDPKSDAGIFLGYSTNSRAYRVFNSRTRTVMESI 969
Query: 965 HVKFDDREPGSKTSEQSESNAGTTDSEDASESDQPSDSEKYTEVESSPEAEYNSEAEPSP 1024
+V DD P K + + + DA++S +
Sbjct: 970 NVVVDDLSPARKKDVEEDVRTSGDNVADAAKSGE-------------------------- 1003
Query: 1025 KVQNESASEDFQDNTQQVIQPKFKHKSSHPEELIIGSKDSPRRTRSHFRQEESLIGLLSI 1084
+N ++ D + Q + + + HP+ELIIG + TRS + S +S
Sbjct: 1004 NAENSDSATDESNINQPDKRSSTRIQKMHPKELIIGDPNRGVTTRSREVEIVSNSCFVSK 1063
Query: 1085 IEPKTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTLLEQNGYSKTS*MNKEK* 1144
IEPK V+EAL+D+ WI AMQEEL FK P P ++ K +
Sbjct: 1064 IEPKNVKEALTDEFWINAMQEELEQFKRNEVWELVPRPEGTNVIGTKWIFKNKTNEEGVI 1123
Query: 1145 PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQMDVKSAFLNGV 1204
K VAQGY+Q EG+D+ ETFAPVARLE+IRLLL A LYQMDVKSAFLNG
Sbjct: 1124 TRNKARLVAQGYTQIEGVDFDETFAPVARLESIRLLLGVACILKFKLYQMDVKSAFLNGY 1183
Query: 1205 IEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDFERGQVDTT 1264
+ EEVYV+QP GF D HPDHVY+LKK+LYGLKQAPRAWY+RL+ FL + + +G +D T
Sbjct: 1184 LNEEVYVEQPKGFADPTHPDHVYRLKKALYGLKQAPRAWYERLTEFLTQQGYRKGGIDKT 1243
Query: 1265 LFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELKFFLGIQINQS 1324
LF + ++++I QIYVDDI+FG + + + F + MQ EFEMS++GEL +FLG+Q+ Q
Sbjct: 1244 LFVKQDAENLMIAQIYVDDIVFGGMSNEMLRHFVQQMQSEFEMSLVGELTYFLGLQVKQM 1303
Query: 1325 KEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGTVVDQKLYRGMIGSLLY 1384
++ +++ Q++Y K ++KKF +E+ TP LSK++ GT VDQ LYR MIGSLLY
Sbjct: 1304 EDSIFLSQSRYAKNIVKKFGMENASHKRTPAPTHLKLSKDEAGTSVDQSLYRSMIGSLLY 1363
Query: 1385 LTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGLLYRKSLDYKLIGFCD 1444
LTASRP I ++V +CAR+Q++P+ SHLT VKRI +Y+ GT++ G++Y + L+G+CD
Sbjct: 1364 LTASRPDITYAVGVCARYQANPKISHLTQVKRILKYVNGTSDYGIMYCHCSNPMLVGYCD 1423
Query: 1445 ADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEYISAASCCTQLLWMKHQ 1504
AD+AG +RKSTSG C +LG NLISW SK+Q +++STAEAEYI+A S C+QL+WMK
Sbjct: 1424 ADWAGSADDRKSTSGGCFYLGNNLISWFSKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQM 1483
Query: 1505 LEDYQINANSIPIYCDNTAAICLSKNPILHSRAKHIEIKHHFIRDYVQKGILDIQFIDIE 1564
L++Y + + + +YCDN +AI +SKNP+ HSR KHI+I+HH+IRD V ++ ++ +D E
Sbjct: 1484 LKEYNVEQDVMTLYCDNMSAINISKNPVQHSRTKHIDIRHHYIRDLVDDKVITLKHVDTE 1543
Query: 1565 HQWADIFTKPLSVERFDFIKKNLNM 1589
Q ADIFTK L +F+ ++ L +
Sbjct: 1544 EQIADIFTKALDANQFEKLRGKLGI 1568
>gb|AAO73527.1| gag-pol polyprotein [Glycine max]
Length = 1576
Score = 921 bits (2380), Expect = 0.0
Identities = 480/1045 (45%), Positives = 656/1045 (61%), Gaps = 29/1045 (2%)
Query: 546 LRAREKQRSWYLDSGCSRHMTGEKALFLTLTMKDGGEVKFGGNQTGKIIGTGTIGNSSI- 604
LRA K+ WYLDSGCSRHMTG K L + V FG GKIIG G + + +
Sbjct: 554 LRASAKE-DWYLDSGCSRHMTGVKEFLLNIEPCSTSYVTFGDGSKGKIIGMGKLVHDGLP 612
Query: 605 SINNVWLVDGLKHNLLSISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKINF 664
S+N V LV GL NL+SISQ CD G++V F+K+ C LV + + KG R ++ +
Sbjct: 613 SLNKVLLVKGLTANLISISQLCDEGFNVNFTKSEC-LVTNEKSEVLMKGSRSKDNCYLWT 671
Query: 665 SDLADQKVVCLLSMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGACQ 724
CL S D+ +WH+R GH + R + KI V+G+PN+ +CG CQ
Sbjct: 672 PQETSYSSTCLSSKEDEVRIWHQRFGHLHLRGMKKIIDKGAVRGIPNLKIEEGRICGECQ 731
Query: 725 KGKIVKSSFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWVKFIK 784
GK VK S + +TSR LELLH+DL GP+ SL G +Y V+VDD+SR+TWV FI+
Sbjct: 732 IGKQVKMSHQKLRHQTTSRVLELLHMDLMGPMQVESLGGKRYAYVVVDDFSRFTWVNFIR 791
Query: 785 SKDYACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRTPQ 844
K EVF ++Q EK+ I ++RSD+G EFEN F FC GI HEFS+ TPQ
Sbjct: 792 EKSETFEVFKELSLRLQREKDCVIKRIRSDHGREFENSRFTEFCTSEGITHEFSAAITPQ 851
Query: 845 QNGVVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYELFKG 904
QNG+VERKNRTLQE AR M+H L + WAEA+NT+CYI NR+ +R T YE++KG
Sbjct: 852 QNGIVERKNRTLQEAARVMLHAKELPYNLWAEAMNTACYIHNRVTLRRGTPTTLYEIWKG 911
Query: 905 RRPNISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVEESM 964
R+P++ +FH FG CYIL ++ +K D K+ GIFLGYS S+AYRV+NS T+ V ES+
Sbjct: 912 RKPSVKHFHIFGSPCYILADREQRRKMDPKSDAGIFLGYSTNSRAYRVFNSRTRTVMESI 971
Query: 965 HVKFDDREPGSKTSEQSESNAGTTDSEDASESDQPSDSEKYTEVESSPEAEYNSEAEPSP 1024
+V DD P K + + + DA++S +
Sbjct: 972 NVVVDDLSPARKKDVEEDVRTLGDNVADAAKSGE-------------------------- 1005
Query: 1025 KVQNESASEDFQDNTQQVIQPKFKHKSSHPEELIIGSKDSPRRTRSHFRQEESLIGLLSI 1084
+N ++ D + Q + + + HP+ELIIG + TRS + S +S
Sbjct: 1006 NAENSDSATDESNINQPDKRSSTRIQKMHPKELIIGDPNRGVTTRSREVEIVSNSCFVSK 1065
Query: 1085 IEPKTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTLLEQNGYSKTS*MNKEK* 1144
IEPK V+EAL+D+ WI AMQEEL FK P P ++ K +
Sbjct: 1066 IEPKNVKEALTDEFWINAMQEELEQFKRNEVWELVPRPEGTNVIGTKWIFKNKTNEEGVI 1125
Query: 1145 PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQMDVKSAFLNGV 1204
K VAQGY+Q EG+D+ ETFAPVARLE+IRLLL A LYQMDVKSAFLNG
Sbjct: 1126 TRNKARLVAQGYTQIEGVDFDETFAPVARLESIRLLLGVACILKFKLYQMDVKSAFLNGY 1185
Query: 1205 IEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDFERGQVDTT 1264
+ EEVYV+QP GF D HPDHVY+LKK+LYGLKQAPRAWY+RL+ FL + + +G +D T
Sbjct: 1186 LNEEVYVEQPKGFADPTHPDHVYRLKKALYGLKQAPRAWYERLTEFLTQQGYRKGGIDKT 1245
Query: 1265 LFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELKFFLGIQINQS 1324
LF + ++++I QIYVDDI+FG + + + F + MQ EFEMS++GEL +FLG+Q+ Q
Sbjct: 1246 LFVKQDAENLMIAQIYVDDIVFGGMSNEMLRHFVQQMQSEFEMSLVGELTYFLGLQVKQM 1305
Query: 1325 KEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGTVVDQKLYRGMIGSLLY 1384
++ +++ Q++Y K ++KKF +E+ TP LSK++ GT VDQ LYR MIGSLLY
Sbjct: 1306 EDSIFLSQSRYAKNIVKKFGMENASHKRTPAPTHLKLSKDEAGTSVDQSLYRSMIGSLLY 1365
Query: 1385 LTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGLLYRKSLDYKLIGFCD 1444
LTASRP I ++V +CAR+Q++P+ SHLT VKRI +Y+ GT++ G++Y + L+G+CD
Sbjct: 1366 LTASRPDITYAVGVCARYQANPKISHLTQVKRILKYVNGTSDYGIMYCHCSNPMLVGYCD 1425
Query: 1445 ADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEYISAASCCTQLLWMKHQ 1504
AD+AG +RKSTSG C +LG NLISW SK+Q +++STAEAEYI+A S C+QL+WMK
Sbjct: 1426 ADWAGSADDRKSTSGGCFYLGNNLISWFSKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQM 1485
Query: 1505 LEDYQINANSIPIYCDNTAAICLSKNPILHSRAKHIEIKHHFIRDYVQKGILDIQFIDIE 1564
L++Y + + + +YCDN +AI +SKNP+ HSR KHI+I+HH+IRD V ++ ++ +D E
Sbjct: 1486 LKEYNVEQDVMTLYCDNMSAINISKNPVQHSRTKHIDIRHHYIRDLVDDKVITLKHVDTE 1545
Query: 1565 HQWADIFTKPLSVERFDFIKKNLNM 1589
Q ADIFTK L +F+ ++ L +
Sbjct: 1546 EQIADIFTKALDANQFEKLRGKLGI 1570
>gb|AAO73523.1| gag-pol polyprotein [Glycine max]
Length = 1576
Score = 921 bits (2380), Expect = 0.0
Identities = 479/1045 (45%), Positives = 654/1045 (61%), Gaps = 29/1045 (2%)
Query: 546 LRAREKQRSWYLDSGCSRHMTGEKALFLTLTMKDGGEVKFGGNQTGKIIGTGTIGNSSI- 604
LRA K+ WYLDSGCSRHMTG K L + V FG GKIIG G + + +
Sbjct: 554 LRASAKE-DWYLDSGCSRHMTGVKEFLLNIEPCSTSYVTFGDGSKGKIIGMGKLVHDGLP 612
Query: 605 SINNVWLVDGLKHNLLSISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKINF 664
S+N V LV GL NL+SISQ CD G++V F+K+ C LV + + KG R ++ +
Sbjct: 613 SLNKVLLVKGLTANLISISQLCDEGFNVNFTKSEC-LVTNEKSEVLMKGSRSKDNCYLWT 671
Query: 665 SDLADQKVVCLLSMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGACQ 724
CL S D+ +WH+R GH + R + KI V+G+PN+ +CG CQ
Sbjct: 672 PQETSYSSTCLSSKEDEVRIWHQRFGHLHLRGMKKILDKSAVRGIPNLKIEEGRICGECQ 731
Query: 725 KGKIVKSSFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWVKFIK 784
GK VK S + +TSR LELLH+DL GP+ SL G +Y V+VDD+SR+TWV FI+
Sbjct: 732 IGKQVKMSHQKLQHQTTSRVLELLHMDLMGPMQVESLGGKRYAYVVVDDFSRFTWVNFIR 791
Query: 785 SKDYACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRTPQ 844
K EVF ++Q EK+ I ++RSD+G EFEN F FC GI HEFS+ TPQ
Sbjct: 792 EKSGTFEVFKKLSLRLQREKDCVIKRIRSDHGREFENSRFTEFCTSEGITHEFSAAITPQ 851
Query: 845 QNGVVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYELFKG 904
QNG+VERKNRTLQE AR M+H L + WAEA+NT+CYI NR+ +R T YE++KG
Sbjct: 852 QNGIVERKNRTLQEAARVMLHAKELPYNLWAEAMNTACYIHNRVTLRRGTPTTLYEIWKG 911
Query: 905 RRPNISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVEESM 964
R+P++ +FH FG CYIL ++ +K D K+ GIFLGYS S+AYRV+NS T+ V ES+
Sbjct: 912 RKPSVKHFHIFGSPCYILADREQRRKMDPKSDAGIFLGYSTNSRAYRVFNSRTRTVMESI 971
Query: 965 HVKFDDREPGSKTSEQSESNAGTTDSEDASESDQPSDSEKYTEVESSPEAEYNSEAEPSP 1024
+V DD P K + + + DA++S +
Sbjct: 972 NVVVDDLSPARKKDVEEDVRTSGDNVADAAKSGE-------------------------- 1005
Query: 1025 KVQNESASEDFQDNTQQVIQPKFKHKSSHPEELIIGSKDSPRRTRSHFRQEESLIGLLSI 1084
+N ++ D + Q + + + HP+ELIIG + TRS + S +S
Sbjct: 1006 NAENSDSATDESNINQPDKRSSTRIQKMHPKELIIGDPNRGVTTRSREVEIVSNSCFVSK 1065
Query: 1085 IEPKTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTLLEQNGYSKTS*MNKEK* 1144
IEPK V+EAL+D+ WI AMQEEL FK P P ++ K +
Sbjct: 1066 IEPKNVKEALTDEFWINAMQEELEQFKRNEVWELVPRPEGTNVIGTKWIFKNKTNEEGVI 1125
Query: 1145 PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQMDVKSAFLNGV 1204
K VAQGY+Q EG+D+ ETFAPVARLE+IRLLL A LYQMDVKSAFLNG
Sbjct: 1126 TRNKARLVAQGYTQIEGVDFDETFAPVARLESIRLLLGVACILKFKLYQMDVKSAFLNGY 1185
Query: 1205 IEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDFERGQVDTT 1264
+ EEVYV+QP GF D HPDHVY+LKK+LYGLKQAPRAWY+RL+ FL + + +G +D T
Sbjct: 1186 LNEEVYVEQPKGFADPTHPDHVYRLKKALYGLKQAPRAWYERLTEFLTQQGYRKGGIDKT 1245
Query: 1265 LFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELKFFLGIQINQS 1324
LF + ++++I QIYVDDI+FG + + + F + MQ EFEMS++GEL +FLG+Q+ Q
Sbjct: 1246 LFVKQDAENLMIAQIYVDDIVFGGMSNEMLRHFVQQMQSEFEMSLVGELTYFLGLQVKQM 1305
Query: 1325 KEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGTVVDQKLYRGMIGSLLY 1384
++ +++ Q++Y K ++KKF +E+ TP LSK++ GT VDQK YR MIGSLLY
Sbjct: 1306 EDSIFLSQSRYAKNIVKKFGMENASHKRTPAPTHLKLSKDEAGTSVDQKPYRSMIGSLLY 1365
Query: 1385 LTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGLLYRKSLDYKLIGFCD 1444
LTASRP I ++V +CAR+Q++P+ SHL VKRI +Y+ GT++ G++Y L+G+CD
Sbjct: 1366 LTASRPDITYAVGVCARYQANPKISHLNQVKRILKYVNGTSDYGIMYCHCSSSMLVGYCD 1425
Query: 1445 ADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEYISAASCCTQLLWMKHQ 1504
AD+AG +RKSTSG C +LG NLISW SK+Q +++STAEAEYI+A S C+QL+WMK
Sbjct: 1426 ADWAGSADDRKSTSGGCFYLGNNLISWFSKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQM 1485
Query: 1505 LEDYQINANSIPIYCDNTAAICLSKNPILHSRAKHIEIKHHFIRDYVQKGILDIQFIDIE 1564
L++Y + + + +YCDN +AI +SKNP+ HSR KHI+I+HH+IRD V ++ ++ +D E
Sbjct: 1486 LKEYNVEQDVMTLYCDNMSAINISKNPVQHSRTKHIDIRHHYIRDLVDDKVITLKHVDTE 1545
Query: 1565 HQWADIFTKPLSVERFDFIKKNLNM 1589
Q ADIFTK L +F+ ++ L +
Sbjct: 1546 EQIADIFTKALDANQFEKLRGKLGI 1570
>gb|AAO73529.1| gag-pol polyprotein [Glycine max]
Length = 1577
Score = 920 bits (2378), Expect = 0.0
Identities = 479/1045 (45%), Positives = 652/1045 (61%), Gaps = 29/1045 (2%)
Query: 546 LRAREKQRSWYLDSGCSRHMTGEKALFLTLTMKDGGEVKFGGNQTGKIIGTGTIGNSSI- 604
LRA K+ WYLDSGCSRHMTG K + + V FG GKI G G + + +
Sbjct: 555 LRASAKE-DWYLDSGCSRHMTGVKEFLVNIEPCSTSYVTFGDGSKGKITGMGKLVHEGLP 613
Query: 605 SINNVWLVDGLKHNLLSISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKINF 664
S+N V LV GL NL+SISQ CD G++V F+K+ C LV + + KG R ++ +
Sbjct: 614 SLNKVLLVKGLTVNLISISQLCDEGFNVNFTKSEC-LVTNEKSEVLMKGSRSKDNCYLWT 672
Query: 665 SDLADQKVVCLLSMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGACQ 724
+ CL S D+ +WH+R GH + R + KI V+G+PN+ +CG CQ
Sbjct: 673 PQESSHSSTCLFSKEDEVKIWHQRFGHLHLRGMKKIIDKGAVRGIPNLKIEEGRICGECQ 732
Query: 725 KGKIVKSSFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWVKFIK 784
GK VK S + +TSR LELLH+DL GP+ SL G +Y V+VDD+SR+TWV FI+
Sbjct: 733 IGKQVKMSHQKLQHQTTSRVLELLHMDLMGPMQVESLGGKRYAYVVVDDFSRFTWVNFIR 792
Query: 785 SKDYACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRTPQ 844
K EVF ++Q EK+ I ++RSD+G EFEN F FC GI HEFS+ TPQ
Sbjct: 793 EKSDTFEVFKELSLRLQREKDCVIKRIRSDHGREFENSKFTEFCTSEGITHEFSAAITPQ 852
Query: 845 QNGVVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYELFKG 904
QNG+VERKNRTLQE AR M+H L + WAEA+NT+CYI NR+ +R T YE++KG
Sbjct: 853 QNGIVERKNRTLQEAARVMLHAKELPYNLWAEAMNTACYIHNRVTLRRGTPTTLYEIWKG 912
Query: 905 RRPNISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVEESM 964
R+P + +FH FG CYIL ++ +K D K+ GIFLGYS S+AYRV+NS T+ V ES+
Sbjct: 913 RKPTVKHFHIFGSPCYILADREQRRKMDPKSDAGIFLGYSTNSRAYRVFNSRTRTVMESI 972
Query: 965 HVKFDDREPGSKTSEQSESNAGTTDSEDASESDQPSDSEKYTEVESSPEAEYNSEAEPSP 1024
+V DD P K +D E + S +S+ AE + A P
Sbjct: 973 NVVVDDLTPARK--------------KDVEEDVRTSGDNVADTAKSAENAENSDSATDEP 1018
Query: 1025 KVQNESASEDFQDNTQQVIQPKFKHKSSHPEELIIGSKDSPRRTRSHFRQEESLIGLLSI 1084
+ +P + + HP+ELIIG + TRS + S +S
Sbjct: 1019 NINQPDK------------RPSIRIQKMHPKELIIGDPNRGVTTRSREIEIVSNSCFVSK 1066
Query: 1085 IEPKTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTLLEQNGYSKTS*MNKEK* 1144
IEPK V+EAL+D+ WI AMQEEL FK P P ++ K +
Sbjct: 1067 IEPKNVKEALTDEFWINAMQEELEQFKRNEVWELVPRPEGTNVIGTKWIFKNKTNEEGVI 1126
Query: 1145 PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQMDVKSAFLNGV 1204
K VAQGY+Q EG+D+ ETFAPVARLE+IRLLL A LYQMDVKSAFLNG
Sbjct: 1127 TRNKARLVAQGYTQIEGVDFDETFAPVARLESIRLLLGVACILKFKLYQMDVKSAFLNGY 1186
Query: 1205 IEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDFERGQVDTT 1264
+ EE YV+QP GF D HPDHVY+LKK+LYGLKQAPRAWY+RL+ FL + + +G +D T
Sbjct: 1187 LNEEAYVEQPKGFVDPTHPDHVYRLKKALYGLKQAPRAWYERLTEFLTQQGYRKGGIDKT 1246
Query: 1265 LFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELKFFLGIQINQS 1324
LF + ++++I QIYVDDI+FG + + + F + MQ EFEMS++GEL +FLG+Q+ Q
Sbjct: 1247 LFVKQDAENLMIAQIYVDDIVFGGMSNEMLRHFVQQMQSEFEMSLVGELTYFLGLQVKQM 1306
Query: 1325 KEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGTVVDQKLYRGMIGSLLY 1384
++ +++ Q+KY K ++KKF +E+ TP LSK++ GT VDQ LYR MIGSLLY
Sbjct: 1307 EDSIFLSQSKYAKNIVKKFGMENASHKRTPAPTHLKLSKDEAGTSVDQSLYRSMIGSLLY 1366
Query: 1385 LTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGLLYRKSLDYKLIGFCD 1444
LTASRP I ++V +CAR+Q++P+ SHL VKRI +Y+ GT++ G++Y L+G+CD
Sbjct: 1367 LTASRPDITYAVGVCARYQANPKISHLNQVKRILKYVNGTSDYGIMYCHCSGSMLVGYCD 1426
Query: 1445 ADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEYISAASCCTQLLWMKHQ 1504
AD+AG +RKSTSG C +LG NLISW SK+Q +++STAEAEYI+A S C+QL+WMK
Sbjct: 1427 ADWAGSADDRKSTSGGCFYLGNNLISWFSKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQM 1486
Query: 1505 LEDYQINANSIPIYCDNTAAICLSKNPILHSRAKHIEIKHHFIRDYVQKGILDIQFIDIE 1564
L++Y + + + +YCDN +AI +SKNP+ HSR KHI+I+HH+IR+ V ++ ++ +D E
Sbjct: 1487 LKEYNVEQDVMTLYCDNMSAINISKNPVQHSRTKHIDIRHHYIRELVDDKVITLEHVDTE 1546
Query: 1565 HQWADIFTKPLSVERFDFIKKNLNM 1589
Q ADIFTK L ++F+ ++ L +
Sbjct: 1547 EQIADIFTKALDAKQFEKLRGKLGI 1571
>gb|AAN40025.1| putative gag-pol polyprotein [Zea mays]
Length = 1892
Score = 912 bits (2357), Expect = 0.0
Identities = 497/1076 (46%), Positives = 651/1076 (60%), Gaps = 94/1076 (8%)
Query: 554 SWYLDSGCSRHMTGEKALFLTLTMKDGGE--VKFGGNQTGKIIGTGTIGNSSISINNVWL 611
SW LDSGC+ HMTGEK +F + + + FG G + G
Sbjct: 863 SWILDSGCTNHMTGEKKMFSSYEKNKDPQRAITFGDGNQGLVKGV--------------- 907
Query: 612 VDGLKHNLLSISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKINFSDLADQK 671
T+ + D SI FKG +Y + F D A+
Sbjct: 908 ----------------------------TVFRRSDDSIAFKGVLEGQLYLVVF-DRAELD 938
Query: 672 VVCLLSMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGACQKGKIVKS 731
CL++ + W+WH+RL H + + K+ K + + GL N+ + D +C ACQ GK V +
Sbjct: 939 T-CLIAKTNMGWLWHRRLAHVGMKNLHKLLKGEHILGLTNVHFEKDRICSACQAGKQVGT 997
Query: 732 SFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWVKFIKSKDYACE 791
K+I++T RPLELLH+DLFGP+ S+ GSKY LVIVDDYSR+TWV F++ K E
Sbjct: 998 HHPHKNIMTTDRPLELLHMDLFGPIAYISIGGSKYCLVIVDDYSRFTWVFFLQEKSQTQE 1057
Query: 792 VFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRTPQQNGVVER 851
F + Q+E L+I K+RSD G EF+N E F E+ GI HEFSSP TPQQNGVVER
Sbjct: 1058 TLKGFLRRAQNEFGLRIKKIRSDNGTEFKNSQIESFLEEEGIKHEFSSPYTPQQNGVVER 1117
Query: 852 KNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYELFKGRRPNISY 911
KNRTL +MARTM+ E FWAEAVNT+CY NR+Y+ +L+KT+YEL G++PNISY
Sbjct: 1118 KNRTLLDMARTMLDEYKTPDRFWAEAVNTACYAINRLYLHRILKKTSYELLTGKKPNISY 1177
Query: 912 FHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVEESMHVKFDDR 971
F FG C+IL + KF K G LGY ++AYRV+N T VE S V FD+
Sbjct: 1178 FRVFGSKCFILIKRGRKSKFAPKTVEGFLLGYDSNTRAYRVFNKSTGLVEVSCDVVFDET 1237
Query: 972 EPGSKTSEQSESNAGTTDS-------------------EDASESDQPSDSEKYT-----E 1007
GS+ + G + E S DQP S + + E
Sbjct: 1238 N-GSQVEQVDLDEIGEEQAPCIALRNMSIGDVCPKESEEPPSTQDQPPSSMQASPPTQNE 1296
Query: 1008 VESSPEAEYNSEAEPSPKVQNESASEDFQDNTQQVIQPKFKH-------KSSHPEELIIG 1060
E+ + E N E +P N+ + + +P+ H + HP + I+G
Sbjct: 1297 DEAQNDEEQNQEVKPPQDKSNDQGGDTNDQEKEDEEEPRPPHPRVHQAIQRDHPVDTILG 1356
Query: 1061 SKDSPRRTRS---HFRQEESLIGLLSIIEPKTVEEALSDDGWILAMQEELINFKGMMCGI 1117
TRS HF + S + S IEP VEEAL D W++AMQEEL NF
Sbjct: 1357 DIHKGVTTRSRVAHFCEHYSFV---SSIEPHRVEEALQDSDWVVAMQEELNNFTRNEVWH 1413
Query: 1118 WYPYPFRRTLLEQNGYSKTS*MNKEK*----PETKPDFVAQGYSQQEGIDYTETFAPVAR 1173
P P + + +K NK+ K VA+GYSQ EG+D+ ET+APVAR
Sbjct: 1414 LVPRPNQNVV-----GTKWVFRNKQDEHGVVTRNKARLVAKGYSQVEGLDFDETYAPVAR 1468
Query: 1174 LEAIRLLLSYAINHGIILYQMDVKSAFLNGVIEEEVYVKQPPGFEDLKHPDHVYKLKKSL 1233
LE+IR+LL+YA HG LYQMDVKSAFLNG I+EEVYV+QPPGFED ++P+HVY+L K+L
Sbjct: 1469 LESIRILLAYATYHGFKLYQMDVKSAFLNGPIKEEVYVEQPPGFEDSEYPNHVYRLSKAL 1528
Query: 1234 YGLKQAPRAWYDRLSNFLIKNDFERGQVDTTLFRRTLKKDILIVQIYVDDIIFGSTNASL 1293
YGLKQAPRAWY+ L +FLI N F+ G+ D TLF +TL+ D+ + QIYVDDIIFGSTN S
Sbjct: 1529 YGLKQAPRAWYECLRDFLIANGFKVGKADPTLFTKTLENDLFVCQIYVDDIIFGSTNEST 1588
Query: 1294 CK*FSKLMQDEFEMSMMGELKFFLGIQINQSKEGVYVHQTKYTKELLKKFKLEDCKVMNT 1353
C+ FS++M +FEMSMMGELK+FLG Q+ Q +EG ++ QTKYT+++L KF ++D K + T
Sbjct: 1589 CEEFSRIMTQKFEMSMMGELKYFLGFQVKQLREGTFISQTKYTQDILAKFGMKDAKPIKT 1648
Query: 1354 PMHPTCTLSKEDTGTVVDQKLYRGMIGSLLYLTASRPAILFSVCLCARFQSDPRESHLTA 1413
PM L + G VDQK+YR MIGSLLYL ASRP I+ SVC+CARFQSDP+ESHLTA
Sbjct: 1649 PMGTNGHLDLDTGGKSVDQKVYRSMIGSLLYLCASRPDIMLSVCMCARFQSDPKESHLTA 1708
Query: 1414 VKRIFRYLKGTTNLGLLYRKSLDYKLIGFCDADYAGDRIERKSTSGNCQFLGENLISWAS 1473
VKRI RYL T GL Y + + LIG+ DAD+AG +I RKSTSG CQFLG +L+SWAS
Sbjct: 1709 VKRILRYLAYTPKFGLWYPRGSTFDLIGYSDADWAGCKINRKSTSGTCQFLGRSLVSWAS 1768
Query: 1474 KRQATIAMSTAEAEYISAASCCTQLLWMKHQLEDYQINANSIPIYCDNTAAICLSKNPIL 1533
K+Q ++A+STAEAEYI+A CC QLLWM+ L DY +P+ CDN +AI ++ NP+
Sbjct: 1769 KKQNSVALSTAEAEYIAAGHCCAQLLWMRQTLRDYGYKLTKVPLLCDNESAIKMADNPVE 1828
Query: 1534 HSRAKHIEIKHHFIRDYVQKGILDIQFIDIEHQWADIFTKPLSVERFDFIKKNLNM 1589
HSR KHI I++HF+RD+ QKG ++I +I+ + Q ADIFTKPL + F ++ LN+
Sbjct: 1829 HSRTKHIAIRYHFLRDHQQKGDIEISYINTKDQLADIFTKPLDEQSFTRLRHELNI 1884
>gb|AAN60991.1| Putative Zea mays retrotransposon Opie-2 [Oryza sativa (japonica
cultivar-group)] gi|34902324|ref|NP_912508.1| Putative
Zea mays retrotransposon Opie-2 [Oryza sativa (japonica
cultivar-group)]
Length = 2011
Score = 911 bits (2355), Expect = 0.0
Identities = 495/1051 (47%), Positives = 657/1051 (62%), Gaps = 43/1051 (4%)
Query: 555 WYLDSGCSRHMTGEKALFLTLTM--KDGGEVKFGGNQTGKIIGTGTIGNSS-ISINNVWL 611
W LDSGC++HMTG++A+F T + + +V FG N GK+IG G I S+ +SI+NV L
Sbjct: 558 WVLDSGCTQHMTGDRAMFTTFEVGGNEQEKVTFGDNSKGKVIGLGKIAISNDLSIDNVSL 617
Query: 612 VDGLKHNLLSISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKINFSDLADQK 671
V L NLLS++Q CD F + + DKS FKG R N+Y +F+
Sbjct: 618 VKSLNFNLLSVAQICDLVLSCAFFPQEVIVSSLLDKSCVFKGFRYGNLYLGDFNSSEANL 677
Query: 672 VVCLLSMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGACQKGKIVKS 731
CL++ W+WH+RL H +SK+SK LV GL ++ + D LC ACQ GK V
Sbjct: 678 KTCLVAKTSLGWLWHRRLAHVGMNQLSKLSKRDLVVGLKDVKFEKDKLCSACQAGKQVAC 737
Query: 732 SFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWVKFIKSKDYACE 791
S +K I+STSRPLELLH++ FGP S+ G+ + LVIVDDYSR+TW+ F+ K E
Sbjct: 738 SHPTKSIMSTSRPLELLHMEFFGPTTYKSIGGNSHCLVIVDDYSRYTWMFFLHDKSIVAE 797
Query: 792 VFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRTPQQNGVVER 851
+F F + Q+E ++K+RSD G +F+N E +C+ GI HE S+ +PQQNGVVE
Sbjct: 798 LFKKFAKRGQNEFNCTLVKIRSDNGSKFKNTNIEDYCDDLGIKHELSATYSPQQNGVVEM 857
Query: 852 KNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYELFKGRRPNISY 911
KNRTL EMARTM+ E ++ FWAEA+NT+C+ NR+Y+ +L+KT+YEL GR+PN++Y
Sbjct: 858 KNRTLIEMARTMLDEYGVSDSFWAEAINTACHATNRLYLHRLLKKTSYELIVGRKPNVAY 917
Query: 912 FHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVEESMHVKFDDR 971
F FGC CYI L KF+++ G LGY+ SKAYRVYN VEE+ V+FD+
Sbjct: 918 FRVFGCKCYIYRKGVRLTKFESRCDEGFLLGYASNSKAYRVYNKNKGIVEETADVQFDET 977
Query: 972 EPGSKTSEQSESNAGTTDSEDASESDQPSDSEKYTEVESSPEAEYNSEAEPSPKVQNESA 1031
GS+ ++ + G A ++ D K EVE P +++ EPS A
Sbjct: 978 N-GSQEGHENLDDVGDEGLMRAMKNMSIGD-VKPIEVEDKPST--STQDEPSTSATPSQA 1033
Query: 1032 S-EDFQDNTQQVIQPKFKH---KSSHPEELIIGSKDSPRRTRSHFRQEESLIGLLSIIEP 1087
E ++ Q + P H HP + ++G +TRS +S +EP
Sbjct: 1034 QVEVEEEKAQDLPMPPRIHTALSKDHPIDQVLGDISKGVQTRSRVASICEHYSFVSCLEP 1093
Query: 1088 KTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTLLEQ---NGYSKTS*MNKEK* 1144
K V+EAL D WI AM +EL NF +W TL+E+ + T + + K
Sbjct: 1094 KHVDEALCDPDWINAMHKELNNFARNK--VW-------TLVERLRDHNVIGTKWVFRNKQ 1144
Query: 1145 PE------TKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQMDVKS 1198
E K FVAQG++Q EG+D+ ETFAPV RLEAI +LL++A I L+QMDVKS
Sbjct: 1145 DENGLVVRNKARFVAQGFTQVEGLDFGETFAPVTRLEAICILLAFASCFNIKLFQMDVKS 1204
Query: 1199 AFLNGVIEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDFER 1258
AFLNG I E V+V+QPPGFED K+P+HVYKL K+LYGLKQAPRAWY+RL +FL+ DF+
Sbjct: 1205 AFLNGEIAELVFVEQPPGFEDPKYPNHVYKLSKALYGLKQAPRAWYERLRDFLLSKDFKI 1264
Query: 1259 GQVDTTLFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELKFFLG 1318
G+VDTTLF + + D + QIYVDDIIFG TN CK F +M EFEMSM+GEL FF G
Sbjct: 1265 GKVDTTLFTKIIGDDFFVCQIYVDDIIFGCTNEVFCKEFGDMMSREFEMSMIGELSFFHG 1324
Query: 1319 IQINQSKEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGTVVDQKLYRGM 1378
+QI Q K+G F LED K + TPM L ++ G VD KLYR M
Sbjct: 1325 LQIKQLKDGT--------------FGLEDAKPIKTPMATNGHLDLDEGGKPVDLKLYRSM 1370
Query: 1379 IGSLLYLTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGLLYRKSLDYK 1438
IGSLLYLTASRP I+FSVC+CARFQ+ P+E HL AVKRI RYLK ++ +GL Y K +K
Sbjct: 1371 IGSLLYLTASRPDIMFSVCMCARFQAAPKECHLVAVKRILRYLKHSSTIGLWYPKGAKFK 1430
Query: 1439 LIGFCDADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEYISAASCCTQL 1498
L+G+ D+DYAG +++RKSTSG+CQ LG +L+SW+SK+Q +A+ AEAEY+SA SCC QL
Sbjct: 1431 LVGYSDSDYAGCKVDRKSTSGSCQMLGRSLVSWSSKKQNFVALFIAEAEYVSAGSCCAQL 1490
Query: 1499 LWMKHQLEDYQINANSIPIYCDNTAAICLSKNPILHSRAKHIEIKHHFIRDYVQKGILDI 1558
LWMK L DY I+ P+ C+N +AI ++ NP+ HSR KHI+I+HHF+RD+V K + I
Sbjct: 1491 LWMKQILLDYGISFTKTPLLCENDSAIKIANNPVQHSRTKHIDIRHHFLRDHVAKCDIVI 1550
Query: 1559 QFIDIEHQWADIFTKPLSVERFDFIKKNLNM 1589
I E Q ADIFTKPL RF ++ LN+
Sbjct: 1551 SHIRTEDQLADIFTKPLDETRFCKLRNELNL 1581
>gb|AAO73525.1| gag-pol polyprotein [Glycine max]
Length = 1576
Score = 910 bits (2351), Expect = 0.0
Identities = 478/1050 (45%), Positives = 646/1050 (61%), Gaps = 29/1050 (2%)
Query: 546 LRAREKQRSWYLDSGCSRHMTGEKALFLTLTMKDGGEVKFGGNQTGKIIGTGTIGNSSI- 604
LRA K+ WYLDSGCSRHMTG K + + V FG GKI G G + + +
Sbjct: 554 LRASAKE-DWYLDSGCSRHMTGVKEFLVNIEPCSTSYVTFGDGSKGKITGMGKLVHDGLP 612
Query: 605 SINNVWLVDGLKHNLLSISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKINF 664
S+N V LV GL NL+SISQ CD G++V F+K+ C LV + + KG R ++ +
Sbjct: 613 SLNKVLLVKGLTANLISISQLCDEGFNVNFTKSEC-LVTNEKSEVLMKGSRSKDNCYLWT 671
Query: 665 SDLADQKVVCLLSMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGACQ 724
CL S D+ +WH+R GH + R + KI V+G+PN+ +CG CQ
Sbjct: 672 PQETSYSSTCLSSKEDEVKIWHQRFGHLHLRGMKKIIDKGAVRGIPNLKIEEGRICGECQ 731
Query: 725 KGKIVKSSFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWVKFIK 784
GK VK S + +TS LELLH+DL GP+ SL G +Y V+VDD+SR+TWV FI+
Sbjct: 732 IGKQVKMSHQKLQHQTTSMVLELLHMDLMGPMQVESLGGKRYAYVVVDDFSRFTWVNFIR 791
Query: 785 SKDYACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRTPQ 844
K EVF ++Q EK+ I ++RSD+G EFEN F FC GI HEFS+ TPQ
Sbjct: 792 EKSDTFEVFKELSLRLQREKDCVIKRIRSDHGREFENSKFTEFCTSEGITHEFSAAITPQ 851
Query: 845 QNGVVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYELFKG 904
QNG+VERKNRTLQE R M+H L + WAEA+NT+CYI NR+ +R T YE++KG
Sbjct: 852 QNGIVERKNRTLQEATRVMLHAKELPYNLWAEAMNTACYIHNRVTLRRGTPTTLYEIWKG 911
Query: 905 RRPNISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVEESM 964
R+P + +FH FG CYIL ++ +K D K+ GIFLGYS S+AYRV+NS T+ V ES+
Sbjct: 912 RKPTVKHFHIFGSPCYILADREQRRKMDPKSDAGIFLGYSTNSRAYRVFNSRTRTVMESI 971
Query: 965 HVKFDDREPGSKTSEQSESNAGTTDSEDASESDQPSDSEKYTEVESSPEAEYNSEAEPSP 1024
+V DD P K +D E + S+ +S+ AE + P
Sbjct: 972 NVVVDDLTPARK--------------KDVEEDVRTSEDNVADTAKSAENAEKSDSTTDEP 1017
Query: 1025 KVQNESASEDFQDNTQQVIQPKFKHKSSHPEELIIGSKDSPRRTRSHFRQEESLIGLLSI 1084
+ S P + + P+ELIIG + TRS + S +S
Sbjct: 1018 NINQPDKS------------PFIRIQKMQPKELIIGDPNRGVTTRSREIEIVSNSCFVSK 1065
Query: 1085 IEPKTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTLLEQNGYSKTS*MNKEK* 1144
IEPK V+EAL+D+ WI AMQEEL FK P P ++ K +
Sbjct: 1066 IEPKNVKEALTDEFWINAMQEELEQFKRNEVWELVPRPEGTNVIGTKWIFKNKTNEEGVI 1125
Query: 1145 PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQMDVKSAFLNGV 1204
K VAQGY+Q EG+D+ ETFAPVARLE+IRLLL A LYQMDVKSAFLNG
Sbjct: 1126 TRNKARLVAQGYTQIEGVDFDETFAPVARLESIRLLLGVACILKFKLYQMDVKSAFLNGY 1185
Query: 1205 IEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDFERGQVDTT 1264
+ EE YV+QP GF D H DHVY+LKK+LYGLKQAPRAWY+RL+ FL + + +G +D T
Sbjct: 1186 LNEEAYVEQPKGFVDPTHLDHVYRLKKALYGLKQAPRAWYERLTEFLTQQGYRKGGIDKT 1245
Query: 1265 LFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELKFFLGIQINQS 1324
LF + ++++I QIYVDDI+FG + + + F MQ EFEMS++GEL +FLG+Q+ Q
Sbjct: 1246 LFVKQDAENLMIAQIYVDDIVFGGMSNEMLRHFVPQMQSEFEMSLVGELHYFLGLQVKQM 1305
Query: 1325 KEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGTVVDQKLYRGMIGSLLY 1384
++ +++ Q+KY K ++KKF +E+ TP LSK++ GT VDQ LYR MIGSLLY
Sbjct: 1306 EDSIFLSQSKYAKNIVKKFGMENASHKRTPAPTHLKLSKDEAGTSVDQNLYRSMIGSLLY 1365
Query: 1385 LTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGLLYRKSLDYKLIGFCD 1444
LTASRP I F+V +CAR+Q++P+ SHL VKRI +Y+ GT++ G++Y D L+G+CD
Sbjct: 1366 LTASRPDITFAVGVCARYQANPKISHLNQVKRILKYVNGTSDYGIMYCHCSDSMLVGYCD 1425
Query: 1445 ADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEYISAASCCTQLLWMKHQ 1504
AD+AG +RK TSG C +LG NLISW SK+Q +++STAEAEYI+A S C+QL+WMK
Sbjct: 1426 ADWAGSADDRKCTSGGCFYLGTNLISWFSKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQM 1485
Query: 1505 LEDYQINANSIPIYCDNTAAICLSKNPILHSRAKHIEIKHHFIRDYVQKGILDIQFIDIE 1564
L++Y + + + +YCDN +AI +SKNP+ H+R KHI+I+HH+IRD V I+ ++ +D E
Sbjct: 1486 LKEYNVEQDVMTLYCDNMSAINISKNPVQHNRTKHIDIRHHYIRDLVDDKIITLEHVDTE 1545
Query: 1565 HQWADIFTKPLSVERFDFIKKNLNMHFVSD 1594
Q ADIFTK L +F+ ++ L + D
Sbjct: 1546 EQVADIFTKALDANQFEKLRGKLGTCLLED 1575
>gb|AAC64917.1| gag-pol polyprotein [Glycine max]
Length = 1550
Score = 908 bits (2347), Expect = 0.0
Identities = 477/1050 (45%), Positives = 647/1050 (61%), Gaps = 29/1050 (2%)
Query: 546 LRAREKQRSWYLDSGCSRHMTGEKALFLTLTMKDGGEVKFGGNQTGKIIGTGTIGNSSI- 604
LRA K+ WYLDSGCSRHMTG K + + V FG GKI G G + + +
Sbjct: 528 LRASAKE-DWYLDSGCSRHMTGVKEFLVNIEPCSTSYVTFGDGSKGKITGMGKLVHDGLP 586
Query: 605 SINNVWLVDGLKHNLLSISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKINF 664
S+N V LV GL NL+SISQ CD G++V F+K+ C LV + + KG R ++ +
Sbjct: 587 SLNKVLLVKGLTANLISISQLCDEGFNVNFTKSEC-LVTNEKSEVLMKGSRSKDNCYLWT 645
Query: 665 SDLADQKVVCLLSMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGACQ 724
CL S D+ +WH+R GH + R + KI V+G+PN+ +CG CQ
Sbjct: 646 PQETSYSSTCLFSKEDEVKIWHQRFGHLHLRGMKKIIDKGAVRGIPNLKIEEGRICGECQ 705
Query: 725 KGKIVKSSFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWVKFIK 784
GK VK S + +TSR LELLH+DL GP+ SL +Y V+VDD+SR+TWV FI+
Sbjct: 706 IGKQVKMSNQKLQHQTTSRVLELLHMDLMGPMQVESLGRKRYAYVVVDDFSRFTWVNFIR 765
Query: 785 SKDYACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRTPQ 844
K EVF ++Q EK+ I ++RSD+G EFEN F FC GI HEFS+ TPQ
Sbjct: 766 EKSDTFEVFKELSLRLQREKDCVIKRIRSDHGREFENSKFTEFCTSEGITHEFSAAITPQ 825
Query: 845 QNGVVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYELFKG 904
QNG+VERKNRTLQE AR M+H L + WAEA+NT+CYI NR+ +R T YE++KG
Sbjct: 826 QNGIVERKNRTLQEAARVMLHAKELPYNLWAEAMNTACYIHNRVTLRRGTPTTLYEIWKG 885
Query: 905 RRPNISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVEESM 964
R+P + +FH G CYIL ++ +K D K+ GIFLGYS S+AYRV+NS T+ V ES+
Sbjct: 886 RKPTVKHFHICGSPCYILADREQRRKMDPKSDAGIFLGYSTNSRAYRVFNSRTRTVMESI 945
Query: 965 HVKFDDREPGSKTSEQSESNAGTTDSEDASESDQPSDSEKYTEVESSPEAEYNSEAEPSP 1024
+V DD P K +D E + S +S+ AE + A P
Sbjct: 946 NVVVDDLTPARK--------------KDVEEDVRTSGDNVADTAKSAENAENSDSATDEP 991
Query: 1025 KVQNESASEDFQDNTQQVIQPKFKHKSSHPEELIIGSKDSPRRTRSHFRQEESLIGLLSI 1084
+ +P + + HP+ELIIG + TRS + S +S
Sbjct: 992 NINQPDK------------RPSIRIQKMHPKELIIGDPNRGVTTRSREIEIISNSCFVSK 1039
Query: 1085 IEPKTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTLLEQNGYSKTS*MNKEK* 1144
IEPK V+EAL+D+ WI AMQEEL FK P P ++ K +
Sbjct: 1040 IEPKNVKEALTDEFWINAMQEELEQFKRNEVWELVPRPEGTNVIGTKWIFKNKTNEEGVI 1099
Query: 1145 PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQMDVKSAFLNGV 1204
K VAQGY+Q EG+D+ ETFAP ARLE+IRLLL A LYQMDVKSAFLNG
Sbjct: 1100 TRNKARLVAQGYTQIEGVDFDETFAPGARLESIRLLLGVACILKFKLYQMDVKSAFLNGY 1159
Query: 1205 IEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDFERGQVDTT 1264
+ EE YV+QP GF D HPDHVY+LKK+LYGLKQAPRAWY+RL+ FL + + +G +D T
Sbjct: 1160 LNEEAYVEQPKGFVDPTHPDHVYRLKKALYGLKQAPRAWYERLTEFLTQQGYRKGGIDKT 1219
Query: 1265 LFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELKFFLGIQINQS 1324
LF + ++++I QIYVDDI+FG + + + F + MQ EFEMS++GEL +FLG+Q+ Q
Sbjct: 1220 LFVKQDAENLMIAQIYVDDIVFGGMSNEMLRHFVQQMQSEFEMSLVGELTYFLGLQVKQM 1279
Query: 1325 KEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGTVVDQKLYRGMIGSLLY 1384
++ +++ Q+KY K ++KKF +E+ TP LSK++ GT VDQ LYR MIGSLLY
Sbjct: 1280 EDSIFLSQSKYAKNIVKKFGMENASHKRTPAPTHLKLSKDEAGTSVDQSLYRSMIGSLLY 1339
Query: 1385 LTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGLLYRKSLDYKLIGFCD 1444
LTASRP I ++V CAR+Q++P+ SHL VKRI +Y+ GT++ G++Y D L+G+CD
Sbjct: 1340 LTASRPDITYAVGGCARYQANPKISHLNQVKRILKYVNGTSDYGIMYCHCSDSMLVGYCD 1399
Query: 1445 ADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEYISAASCCTQLLWMKHQ 1504
AD+AG +RKST G C +LG N ISW SK+Q +++STAEAEYI+A S C+QL+WMK
Sbjct: 1400 ADWAGSVDDRKSTFGGCFYLGTNFISWFSKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQM 1459
Query: 1505 LEDYQINANSIPIYCDNTAAICLSKNPILHSRAKHIEIKHHFIRDYVQKGILDIQFIDIE 1564
L++Y + + + +YCDN +AI +SKNP+ HSR KHI+I+HH+IRD V ++ ++ +D E
Sbjct: 1460 LKEYNVEQDVMTLYCDNLSAINISKNPVQHSRTKHIDIRHHYIRDLVDDKVITLEHVDTE 1519
Query: 1565 HQWADIFTKPLSVERFDFIKKNLNMHFVSD 1594
Q ADIFTK L +F+ ++ L + + D
Sbjct: 1520 EQIADIFTKALDANQFEKLRGKLGICLLED 1549
>gb|AAV31277.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1577
Score = 907 bits (2345), Expect = 0.0
Identities = 484/1042 (46%), Positives = 645/1042 (61%), Gaps = 27/1042 (2%)
Query: 555 WYLDSGCSRHMTGEKALFLTLTM--KDGGEVKFGGNQTGKIIGTGTIGNSS-ISINNVWL 611
W LDSGC++HMTG++A+F T + + +V FG N GK+IG G I S+ +SI+NV L
Sbjct: 549 WVLDSGCTQHMTGDRAMFTTFEVGRNEQEKVTFGDNSKGKVIGLGKIAISNDLSIDNVSL 608
Query: 612 VDGLKHNLLSISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKINFSDLADQK 671
V L NLLS++Q CD F + + DKS FKG R N+Y ++F+
Sbjct: 609 VKSLNFNLLSVAQICDLSLSCAFFPQEVIVSSLLDKSCVFKGFRYGNLYLVDFNSSEANL 668
Query: 672 VVCLLSMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGACQKGKIVKS 731
CL++ W+WH+RL H +SK SK LV GL ++ + D LC ACQ GK V
Sbjct: 669 KTCLVAKTSLGWLWHRRLAHVGMNQLSKFSKRDLVMGLKDVKFEKDKLCSACQAGKQVAC 728
Query: 732 SFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWVKFIKSKDYACE 791
S +K I+STS+PLELLH+DLF P S+ G+ + LVIVDDYSR+TWV F+ K +
Sbjct: 729 SHPTKSIMSTSKPLELLHMDLFDPTTYKSIGGNSHCLVIVDDYSRYTWVFFLHDKSIVAD 788
Query: 792 VFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRTPQQNGVVER 851
+F F + Q+E ++K+RS+ G EF+N E +C+ GI HE + +PQQNGVVER
Sbjct: 789 LFKKFAKRAQNEFSCTLVKIRSNIGSEFKNTNIEDYCDDLGIKHELFATYSPQQNGVVER 848
Query: 852 KNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYELFKGRRPNISY 911
KNRTL EMARTM+ E ++ FWAEA+NT+C+ NR+Y+ +L+KT+YE+ GR+PNI+Y
Sbjct: 849 KNRTLIEMARTMLDEYGVSDSFWAEAINTACHATNRLYLHRVLKKTSYEVIVGRKPNIAY 908
Query: 912 FHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVEESMHVKFDDR 971
F FGC CYI L KF+++ G LGY+ +SKAYRVYN VEE+ V+FD+
Sbjct: 909 FRVFGCKCYIHRKGVRLTKFESRCDEGFLLGYASKSKAYRVYNKNKGIVEETADVQFDET 968
Query: 972 EPGSKTSEQSESNAGTTDSEDASESDQPSDSEKYTEVESSPEAEYNSEAEPSPKVQNESA 1031
GS+ ++ + G ++ D K EVE P +++ EPS A
Sbjct: 969 N-GSQEGHENLDDVGDEGLMRVMKNMSIGDV-KPIEVEDKPST--STQDEPSTSAMPSQA 1024
Query: 1032 SEDFQDN-TQQVIQPKFKHKS---SHPEELIIGSKDSPRRTRSHFRQEESLIGLLSIIEP 1087
+ ++ Q+ P H + HP + ++G +TRS +S +E
Sbjct: 1025 QVEVEEEKAQEPPMPPRIHTALSKDHPIDQVLGDISKGVQTRSRVTSICEHYSFVSCLER 1084
Query: 1088 KTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTLLEQNGYSKTS*MNKEK*PET 1147
K V+EAL D W+ AM EEL NF P ++ +
Sbjct: 1085 KHVDEALCDPDWMNAMHEELKNFARNKVWTLVERPRDHNVIGTKWVFRNKQDENGLVVRN 1144
Query: 1148 KPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQMDVKSAFLNGVIEE 1207
K VAQG++Q EG+D+ ETFAPVARLEAI +LL++A I L+QMDVKSAFLN
Sbjct: 1145 KARLVAQGFTQVEGLDFGETFAPVARLEAICILLAFASCFDIKLFQMDVKSAFLN----- 1199
Query: 1208 EVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDFERGQVDTTLFR 1267
D K+P+HVYKL K+LYGL+QAPRAWY+RL +FL+ DF+ G+VD TLF
Sbjct: 1200 -----------DTKYPNHVYKLSKALYGLRQAPRAWYERLRDFLLSKDFKIGKVDITLFT 1248
Query: 1268 RTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELKFFLGIQINQSKEG 1327
+ + D + QIYVDDIIFGSTN CK F +M EFEMSM+GEL FFLG+QI Q K G
Sbjct: 1249 KIIGDDFFVYQIYVDDIIFGSTNEVFCKEFGDMMSREFEMSMIGELSFFLGLQIKQLKNG 1308
Query: 1328 VYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGTVVDQKLYRGMIGSLLYLTA 1387
+V QTKY K+LLK+F LED K + TPM L ++ G VD KLYR MIGSLLYLT
Sbjct: 1309 TFVSQTKYIKDLLKRFGLEDAKPIKTPMATNGHLDLDEGGKPVDLKLYRSMIGSLLYLTV 1368
Query: 1388 SRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGLLYRKSLDYKLIGFCDADY 1447
SRP I+FSVC+CARFQ+ P+E HL AVKRI RYLK ++ +GL Y K +KL+G+ D DY
Sbjct: 1369 SRPDIMFSVCMCARFQAAPKECHLVAVKRILRYLKHSSTIGLWYPKGAKFKLVGYSDPDY 1428
Query: 1448 AGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEYISAASCCTQLLWMKHQLED 1507
AG +++RKSTS +CQ LG +L+SW+SK+Q ++A+STAE EY+SA SCC QLLWMK L D
Sbjct: 1429 AGCKVDRKSTSSSCQMLGRSLVSWSSKKQNSVALSTAETEYVSAGSCCAQLLWMKQTLLD 1488
Query: 1508 YQINANSIPIYCDNTAAICLSKNPILHSRAKHIEIKHHFIRDYVQKGILDIQFIDIEHQW 1567
Y I+ P+ CDN AI ++ NP+ HSR KHI+I+HHF+RD+V K + I I E Q
Sbjct: 1489 YGISFTKTPLLCDNDGAIKIANNPVQHSRTKHIDIRHHFLRDHVAKCDIVISHIRTEDQL 1548
Query: 1568 ADIFTKPLSVERFDFIKKNLNM 1589
ADIFTKPL RF ++ LN+
Sbjct: 1549 ADIFTKPLDETRFCKLRNELNI 1570
>gb|AAL35396.1| Opie2a pol [Zea mays]
Length = 1048
Score = 907 bits (2344), Expect = 0.0
Identities = 480/1048 (45%), Positives = 649/1048 (61%), Gaps = 30/1048 (2%)
Query: 565 MTGEKALFLTLTMKDGGE--VKFGGNQTGKIIGTGTIGNSSI-SINNVWLVDGLKHNLLS 621
MTGEK +F + + + FG GK+ G G I SS SI+NV+LV+ L +N LS
Sbjct: 1 MTGEKKMFTSYVKNKDSQDSIIFGDGNQGKVKGLGKIAISSEHSISNVFLVESLGYNFLS 60
Query: 622 ISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKINFSDLADQKVVCLLSMNDK 681
+SQ C+ GY+ F+ + ++ + D S+ FKG +Y ++F+ CL++
Sbjct: 61 VSQLCNMGYNCLFTNVDVSVFRRSDGSLAFKGVLDGKLYLVDFAKEEAGLDACLIAKTSM 120
Query: 682 KWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGACQKGKIVKSSFKSKDIVST 741
W+WH+RL H + + K+ K + V GL N+ + D C ACQ GK V+ S +K++++T
Sbjct: 121 GWLWHRRLAHVGMKNLHKLLKGEHVIGLTNVQFKKDRPCVACQAGKQVRGSHHTKNVMTT 180
Query: 742 SRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWVKFIKSKDYACEVFSSFCTQIQ 801
SRPLE+LH+DLFGPV S+ GSKYGLVIVDD+SR+TWV F++ K + + Q
Sbjct: 181 SRPLEMLHMDLFGPVAYLSIGGSKYGLVIVDDFSRFTWVFFLQDKSETQGTLKRYLRRAQ 240
Query: 802 SEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRTPQQNGVVERKNRTLQEMAR 861
+E ELK+ K+RSD G EF+N E F + GI HEFS+P TPQQNGVVERKNRTL +MAR
Sbjct: 241 NEFELKVKKIRSDNGSEFKNLQVEEFLVEEGIKHEFSAPYTPQQNGVVERKNRTLMDMAR 300
Query: 862 TMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYELFKGRRPNISYFHQFGCTCYI 921
TM+ E + FW+EAVNT+C+ NR+Y+ +L+ T+YEL G +PN+SYF FG CYI
Sbjct: 301 TMLGEFKTPERFWSEAVNTACHSINRVYLHRLLKNTSYELLTGNKPNVSYFRVFGSKCYI 360
Query: 922 LNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVEESMHVKFDDR---------- 971
L K KF KA G LGY +KAYRV+N + VE S V FD+
Sbjct: 361 LVKKGRTSKFAPKAVEGFLLGYDSNTKAYRVFNKSSGLVEVSSDVVFDETNGSLREQVVN 420
Query: 972 -----EPGSKTSEQSESNAGTTDSEDASESDQPSDSEKYTEVESSPEAEYNSEAEPSPKV 1026
E T+ G ++ E DQPS + T V + + E P
Sbjct: 421 LDDVDEEDVPTAAMRTMAIGDVRPQEHLEQDQPSST---TMVHPPTQ---DDEQAPQVGA 474
Query: 1027 QNESASEDFQDNTQQV-----IQPKFKHKSSHPEELIIGSKDSPRRTRSHFRQEESLIGL 1081
++ ++D Q ++ Q + + +HP I+G TRS
Sbjct: 475 HDQGGAQDVQVEEEEAPQAPPTQVRATIQRNHPVNQILGDISKGVTTRSRLVNFCEHYSF 534
Query: 1082 LSIIEPKTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTLLEQNGYSKTS*MNK 1141
+S IEP VEE L D W+LAMQEEL NFK P P ++ ++ +
Sbjct: 535 VSSIEPFRVEEVLLDPDWVLAMQEELNNFKRNEVWTLVPRP-KQNVVGTKWVFRNKQDEH 593
Query: 1142 EK*PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQMDVKSAFL 1201
K VA+GY+Q G+D+ ETFAPVARL++IR+ L+YA +H LYQMDVKSAFL
Sbjct: 594 GVVTRNKARLVAKGYAQVAGLDFEETFAPVARLKSIRIWLAYAAHHSFRLYQMDVKSAFL 653
Query: 1202 NGVIEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDFERGQV 1261
NG I+EEVYV+QPPGFED + PDHV KL K+LYGLKQAPRAWY+ L +FLI N F+ G+
Sbjct: 654 NGPIKEEVYVEQPPGFEDERFPDHVCKLSKALYGLKQAPRAWYECLRDFLIANAFKVGKA 713
Query: 1262 DTTLFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELKFFLGIQI 1321
D TLF +T D+ + QIYVDDIIFGSTN + C+ FS++M +FEMSMMGEL +FLG Q+
Sbjct: 714 DPTLFTKTCDGDLFVCQIYVDDIIFGSTNQNSCEEFSRVMTQKFEMSMMGELSYFLGFQV 773
Query: 1322 NQSKEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGTVVDQKLYRGMIGS 1381
Q K+G ++ QTKYT++L+K+F ++D K TPM G VDQK YR IGS
Sbjct: 774 RQLKDGTFISQTKYTQDLIKRFGMKDAKPAKTPMGTDGHTDLNKGGKSVDQKAYRSTIGS 833
Query: 1382 LLYLTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGLLYRKSLDYKLIG 1441
LLYL ASRP I+ SVC+CARFQSDPRE HL AVKRI RYL T G+ Y K + LIG
Sbjct: 834 LLYLCASRPDIMLSVCMCARFQSDPRECHLVAVKRILRYLVATPCFGIWYPKGSTFDLIG 893
Query: 1442 FCDADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEYISAASCCTQLLWM 1501
+ D+DYA +++RKSTS CQFLG +L+SW SK+Q ++A+STAEAEY++ CC QLLWM
Sbjct: 894 YSDSDYARCKVDRKSTSRMCQFLGRSLVSWNSKKQTSVALSTAEAEYVAVGQCCAQLLWM 953
Query: 1502 KHQLEDYQINANSIPIYCDNTAAICLSKNPILHSRAKHIEIKHHFIRDYVQKGILDIQFI 1561
+ L D+ N + +P+ CDN +AI +++NP+ HSR KHI+I+HHF+RD+ QKG++++ +
Sbjct: 954 RQTLRDFGYNLSEVPLLCDNESAIRIAENPVEHSRTKHIDIRHHFLRDHQQKGVIEVFHV 1013
Query: 1562 DIEHQWADIFTKPLSVERFDFIKKNLNM 1589
E+ ADIFTKPL + F + LN+
Sbjct: 1014 SSENHLADIFTKPLDEQTFCKLHSELNV 1041
>gb|AAG52949.1| gag/pol polyprotein [Arabidopsis thaliana]
Length = 1643
Score = 904 bits (2337), Expect = 0.0
Identities = 469/1043 (44%), Positives = 662/1043 (62%), Gaps = 18/1043 (1%)
Query: 552 QRSWYLDSGCSRHMTGEKALFLTLTMKDGGEVKFGGNQTGKIIGTGTIGNSSIS-INNVW 610
++ WY DSG SRHMTG +A + V FGG G+I G G + + + NV+
Sbjct: 611 KKPWYFDSGASRHMTGSQANLNNYSSVKESNVMFGGGAKGRIKGKGDLTETEKPHLTNVY 670
Query: 611 LVDGLKHNLLSISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKINFSDLADQ 670
V+GL NL+S+SQ CD G V+F+K C N+ +++ + N Y + ++
Sbjct: 671 FVEGLTANLISVSQLCDEGLTVSFNKVKCWATNERNQNTLTGVRTGNNCY------MWEE 724
Query: 671 KVVCLLSMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGACQKGKIVK 730
+CL + + +WH+RLGH N R +SK+ ++V+G+P + + +CGAC +GK ++
Sbjct: 725 PKICLRAEKEDPVLWHQRLGHMNARSMSKLVNKEMVRGVPELKHIEKIVCGACNQGKQIR 784
Query: 731 SSFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWVKFIKSKDYAC 790
K + + T++ L+L+H+DL GP+ T S+ G +Y V+VDD+SR+ WV+FI+ K
Sbjct: 785 VQHKRVEGIQTTQVLDLIHMDLMGPMQTESIAGKRYVFVLVDDFSRYAWVRFIREKSETA 844
Query: 791 EVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRTPQQNGVVE 850
F Q+++EK++ I ++RSD GGEF NE F FCE GI H++S+PRTPQ NGVVE
Sbjct: 845 NSFKILALQLKNEKKMGIKQIRSDRGGEFMNEAFNSFCESQGIFHQYSAPRTPQSNGVVE 904
Query: 851 RKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYELFKGRRPNIS 910
RKNRTLQEMAR MIH N + + FWAEA++T+CY+ NR+Y+R +KT YE++KG++PN+S
Sbjct: 905 RKNRTLQEMARAMIHGNGVPEKFWAEAISTACYVINRVYVRLGSDKTPYEIWKGKKPNLS 964
Query: 911 YFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVEESMHVKFDD 970
YF FGC CYI+N KD L KFD++++ G FLGY+ S AYRV+N + +EESM+V FDD
Sbjct: 965 YFRVFGCVCYIMNDKDQLGKFDSRSEEGFFLGYATNSLAYRVWNKQRGKIEESMNVVFDD 1024
Query: 971 RE-PGSKTSEQSESNAGTTDSEDASESDQPSDSEKYTEVESSPEAEYNSEAEPSPKVQNE 1029
P + ++ + T+ S + E + ++ + + E + E P +V +
Sbjct: 1025 GSMPELQIIVRNRNEPQTSISNNHGEE---RNDNQFDNGDINKSGEESDEEVPPAQVHRD 1081
Query: 1030 SASEDF-QDNTQQVIQPKFKHKSSHPEELIIGSKDSPRRTRSHFRQEESLIG-LLSIIEP 1087
AS+D D + + + K + G K R S EE + +SI+EP
Sbjct: 1082 HASKDIIGDPSGERVTRGVKQDYRQ----LAGIKQKHRVMASFACFEEIMFSCFVSIVEP 1137
Query: 1088 KTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTLLEQNGYSKTS*MNKEK*PET 1147
K V+EAL D WILAM+EEL F P P + ++ K
Sbjct: 1138 KNVKEALEDHFWILAMEEELEEFSRHQVWDLVPRPPQVNVIGTKWIFKNKFDEVGNITRN 1197
Query: 1148 KPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQMDVKSAFLNGVIEE 1207
K VAQGY+Q EG+D+ ETFAPVARLE IR LL A G L+QMDVK AFLNG+IEE
Sbjct: 1198 KARLVAQGYTQVEGLDFDETFAPVARLECIRFLLGTACGMGFKLHQMDVKCAFLNGIIEE 1257
Query: 1208 EVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDFERGQVDTTLFR 1267
EVYV+QP GFE+L+ P++VYKLKK+LYGLKQAPRAWY+RL+ FLI + RG VD TLF
Sbjct: 1258 EVYVEQPKGFENLEFPEYVYKLKKALYGLKQAPRAWYERLTTFLIVQGYTRGSVDKTLFV 1317
Query: 1268 RTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELKFFLGIQINQSKEG 1327
+ I+I+QIYVDDI+FG T+ L K F K M EF MSM+GELK+FLG+QINQ+ EG
Sbjct: 1318 KNDVHGIIIIQIYVDDIVFGGTSDKLVKTFVKTMTTEFRMSMVGELKYFLGLQINQTDEG 1377
Query: 1328 VYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGTVVDQKLYRGMIGSLLYLTA 1387
+ + Q+ Y + L+K+F + K TPM T L K++ G VD+KLYRGMIGSLLYLTA
Sbjct: 1378 ITISQSTYAQNLVKRFGMCSSKPAPTPMSTTTKLFKDEKGVKVDEKLYRGMIGSLLYLTA 1437
Query: 1388 SRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGLLYRKSLDYKLIGFCDADY 1447
+RP + SV LCAR+QS+P+ SHL AVKRI +Y+ GT N GL Y + L+G+CDAD+
Sbjct: 1438 TRPDLCLSVGLCARYQSNPKASHLLAVKRIIKYVSGTINYGLNYTRDTSLVLVGYCDADW 1497
Query: 1448 AGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEYISAASCCTQLLWMKHQLED 1507
G+ +R+ST+G FLG NLISW SK+Q +++S+ ++EYI+ SCCTQLLWM+ D
Sbjct: 1498 GGNLDDRRSTTGGVFFLGSNLISWHSKKQNCVSLSSTQSEYIALGSCCTQLLWMRQMGLD 1557
Query: 1508 YQIN-ANSIPIYCDNTAAICLSKNPILHSRAKHIEIKHHFIRDYVQKGILDIQFIDIEHQ 1566
Y + + + + CDN +AI +SKNP+ HS KHI I+HHF+R+ V++ + ++ + E Q
Sbjct: 1558 YGMTFPDPLLVKCDNESAIAISKNPVQHSVTKHIAIRHHFVRELVEEKQITVEHVPTEIQ 1617
Query: 1567 WADIFTKPLSVERFDFIKKNLNM 1589
DIFTKPL + F ++K+L +
Sbjct: 1618 LVDIFTKPLDLNTFVNLQKSLGI 1640
>gb|AAW57789.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1799
Score = 870 bits (2247), Expect = 0.0
Identities = 475/1052 (45%), Positives = 648/1052 (61%), Gaps = 47/1052 (4%)
Query: 551 KQRSWYLDSGCSRHMTGEKALFLTLTMKDGGE-VKFGGNQTGKIIGTGTIG-NSSISINN 608
+ SW +DSGCSRHMTGE F +LT G E + FG +G+++ GTI N + +
Sbjct: 775 RSNSWLVDSGCSRHMTGEAKWFTSLTRASGDETITFGDASSGRVMAKGTIKVNDKFMLKD 834
Query: 609 VWLVDGLKHNLLSISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKINFSDLA 668
V LV LK+NLLS+SQ CD +V F K +++ + + F RV V+ NF A
Sbjct: 835 VALVSKLKYNLLSVSQLCDENLEVRFKKDRSRVLDASESPV-FDISRVGRVFFANFDSSA 893
Query: 669 DQKVVCLL-SMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGACQKGK 727
CL+ S N + WH+RLGH + +S+IS + L++GLP + D +C C+ GK
Sbjct: 894 PGPSRCLIASENRDLFFWHRRLGHIGFDHLSRISGMDLIRGLPKLKVPKDLVCAPCRHGK 953
Query: 728 IVKSSFKSKDIVSTSRPLELLHIDLFGPVNTASLYG------SKYGLVIVDDYSRWTWVK 781
+ SS K +V T P +LLH+D GP S+ G + + L++V W W
Sbjct: 954 MTSSSHKPVTMVMTDGPGQLLHMDTVGPARVQSVGGKWRKHLASFSLLLV----VWLWSF 1009
Query: 782 FIKSKDYACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPR 841
+ + +A V + ++I L ++ + S+ H+FSSP
Sbjct: 1010 LVLFEPFA--VIMALNSKI-LHLNLSVILLESN--------------------HQFSSPY 1046
Query: 842 TPQQNGVVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYEL 901
PQQNGVVERKNRTL EMARTM+ E + FW EA++ +C+I NR+++R +L KT YEL
Sbjct: 1047 VPQQNGVVERKNRTLVEMARTMLDEFTTPRKFWTEAISAACFISNRVFLRTILHKTPYEL 1106
Query: 902 FKGRRPNISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVE 961
GRRP +S+ FGC C++L + + L KF++++ GIFLGY+ S+AYRVY T +
Sbjct: 1107 RFGRRPKVSHLRVFGCKCFVLKSGN-LDKFESRSLDGIFLGYATHSRAYRVYVLSTNKIV 1165
Query: 962 ESMHVKFDDREPGSKTSEQSESNAGTTDSEDASESDQPSDSEKYTEVESSPEAEYNSEAE 1021
E+ V FD+ PG++ + ED+ + D S S + P E S
Sbjct: 1166 ETCEVTFDEASPGARPEISGVPDESIFVDEDSDDDDDDSISPPLD--STPPVQETGSPLT 1223
Query: 1022 PSPK----VQNESASEDFQDNTQQVIQPKFKHKSSHPEELIIGSKDSPRRTRSHFRQEES 1077
SP + SA+E+ T P+ P+ +I G + R RS+ +
Sbjct: 1224 TSPSGDAPTTSSSAAEEIDGGTSGPTAPRHIQNRHPPDSMIGGLGERVTRNRSYELVNSA 1283
Query: 1078 LIGLLSIIEPKTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTLLEQNGYSKTS 1137
+ + EPK V ALSD+ W+ AM EEL NF+ P +++ K
Sbjct: 1284 FV---ASFEPKNVCHALSDENWVNAMHEELENFERNKVWSLVEPPLGFSVIGTKWVFKNK 1340
Query: 1138 *MNKEK*PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQMDVK 1197
K VAQG++Q EG+D+ ETFAPVARLEAIR+LL++A + G L+QMDVK
Sbjct: 1341 LGEDGSIVRNKARLVAQGFTQVEGLDFEETFAPVARLEAIRILLAFAASKGFKLFQMDVK 1400
Query: 1198 SAFLNGVIEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDFE 1257
SAFLNGVIEEEVYVKQPPGFE+ K P+HV+KL K+LYGLKQAPRAWY+RL FL++N FE
Sbjct: 1401 SAFLNGVIEEEVYVKQPPGFENPKFPNHVFKLDKALYGLKQAPRAWYERLKTFLLQNGFE 1460
Query: 1258 RGQVDTTLFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELKFFL 1317
G VD TLF D L+VQIYVDDIIFG ++ +L F +M EFEMSMMGEL FFL
Sbjct: 1461 MGAVDKTLFTLHSGIDFLLVQIYVDDIIFGGSSHALVAQFFDVMSREFEMSMMGELTFFL 1520
Query: 1318 GIQINQSKEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGTVVDQKLYRG 1377
G+QI Q+KEG++VHQTKY+KELLKKF + DCK + TPM T +L ++ G VDQ+ YR
Sbjct: 1521 GLQIKQTKEGIFVHQTKYSKELLKKFDMADCKPIATPMATTSSLGPDEDGEEVDQREYRS 1580
Query: 1378 MIGSLLYLTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGLLYRKSLDY 1437
MIGSLLYLTASRP I FSVCLCARFQ+ PR SH AVKRIFRY+K T G+ Y S
Sbjct: 1581 MIGSLLYLTASRPDIHFSVCLCARFQASPRTSHRQAVKRIFRYIKSTLEYGIWYSCSSAL 1640
Query: 1438 KLIGFCDADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEYISAASCCTQ 1497
+ F DAD+AG +I+RKSTSG C FLG +L+SW+S++Q+++A STAEAEY++AAS C+Q
Sbjct: 1641 SVRAFSDADFAGCKIDRKSTSGTCHFLGTSLVSWSSRKQSSVAQSTAEAEYVAAASACSQ 1700
Query: 1498 LLWMKHQLEDYQINANSIPIYCDNTAAICLSKNPILHSRAKHIEIKHHFIRDYVQKGILD 1557
+LWM L+DY ++ + +P+ CDNT+AI ++KNP+ HSR KHIEI++HF+RD V+KG +
Sbjct: 1701 VLWMISTLKDYGLSFSGVPLLCDNTSAINIAKNPVQHSRTKHIEIRYHFLRDNVEKGTIV 1760
Query: 1558 IQFIDIEHQWADIFTKPLSVERFDFIKKNLNM 1589
++F++ E Q ADIFTKPL RF+F++ L +
Sbjct: 1761 LEFVESEKQLADIFTKPLDRSRFEFLRSELGV 1792
>ref|XP_468929.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|28209450|gb|AAO37468.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1128
Score = 848 bits (2191), Expect = 0.0
Identities = 451/944 (47%), Positives = 600/944 (62%), Gaps = 13/944 (1%)
Query: 652 KGKRVENVYKINFSDLADQKVVCLLSMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPN 711
+GK E K+ F D + + V+ L++ W+WH+RL H +SK+SK LV GL +
Sbjct: 185 EGKEQE---KVTFGDNSKRNVIGLVAKTSFGWLWHRRLAHVGMNQLSKLSKRDLVVGLKD 241
Query: 712 IDYHSDALCGACQKGKIVKSSFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIV 771
+ + D LC ACQ K V S +K I+STSRPLELLH+DLFGP S+ G+ + LVIV
Sbjct: 242 VKFEKDKLCSACQASKQVACSHPTKSIMSTSRPLELLHMDLFGPTTYKSIGGNSHCLVIV 301
Query: 772 DDYSRWTWVKFIKSKDYACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKH 831
DDYS +TWV F+ K E+F F + Q+E ++K+RSD G +F+N E +C+
Sbjct: 302 DDYSCYTWVFFLHDKCIVAELFKKFAKRAQNEFSCTLVKIRSDNGSKFKNTNIEDYCDDL 361
Query: 832 GILHEFSSPRTPQQNGVVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIR 891
I HE S+ +PQQNGVVERKNRTL EMARTM+ E ++ FWAEA+NT+C+ NR+Y+
Sbjct: 362 SIKHELSATYSPQQNGVVERKNRTLIEMARTMLDEYGVSDSFWAEAINTACHATNRLYLH 421
Query: 892 PMLEKTTYELFKGRRPNISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYR 951
+L+KT+YEL GR+PN++YF FGC CYI L KF+++ G LGY+ SKAYR
Sbjct: 422 RLLKKTSYELIVGRKPNVAYFRVFGCKCYIYRKGVRLTKFESRCDEGFLLGYASNSKAYR 481
Query: 952 VYNSETQCVEESMHVKFDDREPGSKTSEQSESNAGTTDSEDASESDQPSDSEKYTEVESS 1011
VYN VEE+ V+FD+ GS+ ++ + G A ++ D K EVE
Sbjct: 482 VYNKNKGIVEETADVQFDETN-GSQEGHENLDDVGDEGLMRAMKNMSIGDV-KPIEVEDK 539
Query: 1012 PEAEYNSEAEPSPKVQNESASEDFQDNTQQ--VIQPKFKHKSS--HPEELIIGSKDSPRR 1067
P +++ EPS A + + Q + P+ S HP + ++G +
Sbjct: 540 PST--STQDEPSTSASPSQAQVEVEKEKAQDPPMPPRIYTALSKDHPIDQVLGDISKGVQ 597
Query: 1068 TRSHFRQEESLIGLLSIIEPKTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTL 1127
TRS +S +EPK V+EAL D W+ A+ EEL NF P +
Sbjct: 598 TRSPVASICEHYSFVSCLEPKHVDEALYDPDWMNAIHEELNNFARNKVWTLVERPRDHNV 657
Query: 1128 LEQNGYSKTS*MNKEK*PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINH 1187
+ + K VAQG++Q E +D+ ETF PVARLEAIR+LL++A
Sbjct: 658 IGTKWVFRNKQDENRLVVRNKARLVAQGFTQVEDLDFGETFGPVARLEAIRILLAFASCF 717
Query: 1188 GIILYQMDVKSAFLNGVIEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRL 1247
I L+QMDVKSAFLNG I E V+V+QPPGF+D K+P+HVYKL K+LYGLKQAPRAWY+RL
Sbjct: 718 DIKLFQMDVKSAFLNGEIAELVFVEQPPGFDDPKYPNHVYKLSKALYGLKQAPRAWYERL 777
Query: 1248 SNFLIKNDFERGQVDTTLFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEM 1307
+FL+ DF+ G+VDTTLF + + D + QIYVDDIIFGSTN CK F +M EFEM
Sbjct: 778 RDFLLSKDFKIGKVDTTLFTKIIGDDFFVCQIYVDDIIFGSTNEVFCKEFGDMMSREFEM 837
Query: 1308 SMMGELKFFLGIQINQSKEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTG 1367
SM+ EL FFLG+QI Q K+G +V QTKY K+LLK+F LED K + TPM L ++ G
Sbjct: 838 SMIEELSFFLGLQIKQLKDGTFVSQTKYIKDLLKRFGLEDAKPIKTPMATNWHLDLDEGG 897
Query: 1368 TVVDQKLYRGMIGSLLYLTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNL 1427
VD KLYR MIGSLLYLTASRP I+FSVC+ ARFQ+ P+E HL AVKRI RYLK ++ +
Sbjct: 898 KPVDLKLYRSMIGSLLYLTASRPDIMFSVCMYARFQAAPKECHLVAVKRILRYLKHSSTI 957
Query: 1428 GLLYRKSLDYKLIGFCDADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAE 1487
L Y K +KL+G+ D+DYAG +++RKSTSG+CQ LG +L+SW+SK+Q ++A+STAEAE
Sbjct: 958 SLWYPKGAKFKLVGYSDSDYAGYKVDRKSTSGSCQMLGRSLVSWSSKKQNSVALSTAEAE 1017
Query: 1488 YISAASCCTQLLWMKHQLEDYQIN--ANSIPIYCDNTAAICLSKNPILHSRAKHIEIKHH 1545
YISA SCC QLLWMK L DY I+ P+ C+N + I ++ NP+ H R KHI+I+HH
Sbjct: 1018 YISAGSCCAQLLWMKQILLDYGISFTETQTPLLCNNDSTIKIANNPVQHFRTKHIDIRHH 1077
Query: 1546 FIRDYVQKGILDIQFIDIEHQWADIFTKPLSVERFDFIKKNLNM 1589
F+ D+V K + I I E Q ADIFTKPL RF ++ LN+
Sbjct: 1078 FLTDHVAKCDIVISHIRTEDQLADIFTKPLDETRFCKLRNELNV 1121
Score = 38.9 bits (89), Expect = 1.4
Identities = 18/43 (41%), Positives = 27/43 (61%), Gaps = 2/43 (4%)
Query: 555 WYLDSGCSRHMTGEKALFLTLTM--KDGGEVKFGGNQTGKIIG 595
W LDS C++ MTG++A+F T + K+ +V FG N +IG
Sbjct: 162 WVLDSVCTQRMTGDRAMFTTFEVEGKEQEKVTFGDNSKRNVIG 204
>ref|NP_912905.1| unnamed protein product [Oryza sativa (japonica cultivar-group)]
Length = 940
Score = 846 bits (2186), Expect = 0.0
Identities = 453/919 (49%), Positives = 593/919 (64%), Gaps = 46/919 (5%)
Query: 698 SKISKLQLVKGLPNIDYHSDALCGACQKGKIVKSSFKSKDIVSTSRPLELLHIDLFGPVN 757
S+ SK + + GL NI + D +C ACQ GK + + K++++T+RPLELLH+DLFGP+
Sbjct: 34 SQTSKARHILGLTNIQFEKDRVCSACQAGKQIGAHHPVKNVMTTTRPLELLHMDLFGPIA 93
Query: 758 TASLYGSKYGLVIVDDYSRWTWVKFIKSKDYACEVFSSFCTQIQSEKELKILKVRSDYGG 817
S+ G+KYGLVIVDD+S +TWV F+ K +F F + Q+E +LKI +RSD
Sbjct: 94 YLSIGGNKYGLVIVDDFSCFTWVFFLHDKSETQAIFKKFARRAQNEFDLKIKNIRSDNVK 153
Query: 818 EFENEPFELFCEKHGILHEFSSPRTPQQNGVVERKNRTLQEMARTMIHENNLAKHFWAEA 877
EF+N E F ++ GI HEFS+P +PQQNGV ERKNRTL E+ARTM+ E + FWAEA
Sbjct: 154 EFKNTCIESFLDEEGIKHEFSAPYSPQQNGVAERKNRTLIEIARTMLDEYKTSDRFWAEA 213
Query: 878 VNTSCYIQNRIYIRPMLEKTTYELFKGRRPNISYFHQFGCTCYILNTKDYLKKFDAKAQR 937
VNT C+ NR+Y+ +L+KT YEL G +PN+SYF FG CYILN K KF K
Sbjct: 214 VNTVCHDINRLYLHRLLKKTPYELLTGNKPNVSYFRVFGSKCYILNKKARSSKFAPKVDG 273
Query: 938 GIFLGYSERSKAYRVYNSETQCVEESMHVKFDDREPGSKTSEQSESNAGTTDSEDASESD 997
G LGY AYRV+N + VE + V FD E + S DS E +
Sbjct: 274 GFLLGYGSNECAYRVFNKTSGIVEIAPDVTFD---------ETNGSQVEQVDSHVLGEEE 324
Query: 998 QPSDSEKYTEV------ESSPEAEYNSEAEPSPKVQNESASEDFQDNTQQ---------- 1041
P ++ K + E A +++ EP Q S D ++
Sbjct: 325 DPREAIKRLALGDVRPREPQQGASSSTQVEPPTSTQANDPSTSSLDQGEEGEQVPPSSIN 384
Query: 1042 VIQPKFKHKS---SHPEELIIGSKDSPRRTRSHFRQEESLIGLLSIIEPKTVEEALSDDG 1098
+ P+ H+S HP + I+G + TRSH +S +EP VEEAL+D
Sbjct: 385 LAHPRI-HQSIQRDHPTDNILGDINKGVSTRSHIANFCEHYSFVSSLEPLRVEEALNDPD 443
Query: 1099 WILAMQEELINFKGMMCGIWYPYPFRRTLLEQNGYSK--TS*MNKEK*PET------KPD 1150
W++AMQEEL NF +W TL+E++ + T + + K E K
Sbjct: 444 WVMAMQEELNNFTRNE--VW-------TLVERSRQNVIGTKWIFRNKQDEAGVVIRNKAR 494
Query: 1151 FVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQMDVKSAFLNGVIEEEVY 1210
VAQG++Q EG+D+ ETFAPVARLE+IR+LL++A N LYQMDVKSAFLNG+I E VY
Sbjct: 495 LVAQGFTQIEGLDFGETFAPVARLESIRILLTFATNLNFKLYQMDVKSAFLNGLINELVY 554
Query: 1211 VKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDFERGQVDTTLFRRTL 1270
V+QPPGF+D K+P+HVYKL K+LY LKQAPRAWY+ L NFL+KN FE G+ D+TLF +
Sbjct: 555 VEQPPGFKDPKYPNHVYKLHKALYELKQAPRAWYECLRNFLVKNGFEIGKADSTLFTKRH 614
Query: 1271 KKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELKFFLGIQINQSKEGVYV 1330
DI + QIYVDDIIFGSTN S + FS++M FEMSMMGELKFFLG+QI Q KEG ++
Sbjct: 615 DNDIFVCQIYVDDIIFGSTNKSFSEEFSRMMTKRFEMSMMGELKFFLGLQIKQLKEGTFI 674
Query: 1331 HQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGTVVDQKLYRGMIGSLLYLTASRP 1390
QTKY K++LKKF +E+ K ++TPM L + G VDQK+YR +IGSLLYL ASRP
Sbjct: 675 CQTKYLKDMLKKFGMENAKPIHTPMPSNGHLDLNEQGKDVDQKVYRSIIGSLLYLCASRP 734
Query: 1391 AILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGLLYRKSLDYKLIGFCDADYAGD 1450
I+ SVC+CARFQ+ P+E HL AVKRI RYL T NLGL Y K + LIG+ DADYAG
Sbjct: 735 DIMLSVCMCARFQAAPKECHLVAVKRILRYLVHTPNLGLWYPKGARFDLIGYADADYAGC 794
Query: 1451 RIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEYISAASCCTQLLWMKHQLEDYQI 1510
+++RKSTSG CQFLG +L+SW+SK+Q ++A+STAEAEYIS SCC QL+WMK L DY +
Sbjct: 795 KVDRKSTSGTCQFLGRSLVSWSSKKQNSVALSTAEAEYISTGSCCAQLIWMKQTLRDYGL 854
Query: 1511 NANSIPIYCDNTAAICLSKNPILHSRAKHIEIKHHFIRDYVQKGILDIQFIDIEHQWADI 1570
N + IP+ CDN +AI ++ NP+ HSR KHI+I+HHF+RD+ +G +DIQ + I+ Q ADI
Sbjct: 855 NVSKIPLLCDNESAIKIANNPVQHSRTKHIDIRHHFLRDHSTRGDIDIQHVRIDKQLADI 914
Query: 1571 FTKPLSVERFDFIKKNLNM 1589
FTKPL RF ++ LN+
Sbjct: 915 FTKPLDEARFCELRSELNI 933
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.332 0.143 0.445
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,483,860,782
Number of Sequences: 2540612
Number of extensions: 101492621
Number of successful extensions: 397116
Number of sequences better than 10.0: 4360
Number of HSP's better than 10.0 without gapping: 2699
Number of HSP's successfully gapped in prelim test: 1703
Number of HSP's that attempted gapping in prelim test: 373307
Number of HSP's gapped (non-prelim): 14581
length of query: 1594
length of database: 863,360,394
effective HSP length: 142
effective length of query: 1452
effective length of database: 502,593,490
effective search space: 729765747480
effective search space used: 729765747480
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 82 (36.2 bits)
Medicago: description of AC137996.10


