

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137825.4 - phase: 0
(161 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
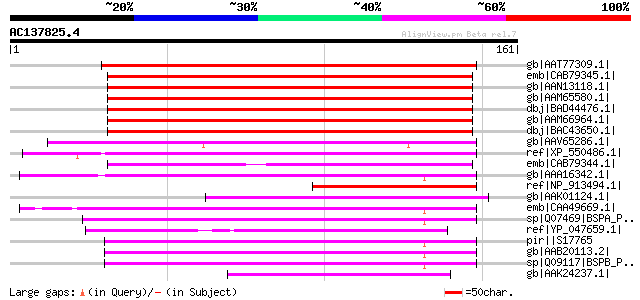
Score E
Sequences producing significant alignments: (bits) Value
gb|AAT77309.1| unknown protein [Oryza sativa (japonica cultivar-... 164 8e-40
emb|CAB79345.1| putative protein [Arabidopsis thaliana] gi|50517... 117 8e-26
gb|AAN13118.1| unknown protein [Arabidopsis thaliana] gi|1387803... 117 8e-26
gb|AAM65580.1| unknown [Arabidopsis thaliana] 117 8e-26
dbj|BAD44476.1| putative protein [Arabidopsis thaliana] 117 8e-26
gb|AAM66964.1| unknown [Arabidopsis thaliana] 109 3e-23
dbj|BAC43650.1| unknown protein [Arabidopsis thaliana] gi|289509... 109 3e-23
gb|AAV65286.1| bark protein-like protein [Thuja occidentalis] 102 5e-21
ref|XP_550486.1| putative vegetative storage protein win4.5 [Ory... 102 5e-21
emb|CAB79344.1| putative protein [Arabidopsis thaliana] gi|50517... 93 2e-18
gb|AAA16342.1| vegetative storage protein [Populus balsamifera s... 87 2e-16
ref|NP_913494.1| unnamed protein product [Oryza sativa (japonica... 83 3e-15
gb|AAK01124.1| vegetative storage protein PNI288 [Populus balsam... 79 6e-14
emb|CAA49669.1| bark storage protein [Populus deltoides] gi|4218... 75 8e-13
sp|Q07469|BSPA_POPDE BARK STORAGE PROTEIN A PRECURSOR 73 2e-12
ref|YP_047659.1| putative phosphorylase family protein [Acinetob... 71 9e-12
pir||S17765 major storage protein - Carolina poplar 71 1e-11
gb|AAB20113.2| major storage protein [Populus x canadensis] 71 1e-11
sp|Q09117|BSPB_POPDE Bark storage protein B precursor gi|169445|... 66 4e-10
gb|AAK24237.1| phosphorylase family protein [Caulobacter crescen... 61 9e-09
>gb|AAT77309.1| unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 360
Score = 164 bits (415), Expect = 8e-40
Identities = 70/119 (58%), Positives = 92/119 (76%)
Query: 30 REISNINNEGPSIGIVVPNAYELNPLLHSSSFVPHNKFPYFDFAGRHFRIGELEKKKVIV 89
R + +N GP +G+VVPN +E+ PLL S +F P K PY D AGR FR G + +KKVI+
Sbjct: 42 RGVRQVNRRGPYLGVVVPNGFEMEPLLRSPAFSPAKKLPYLDVAGRRFRFGSIGEKKVII 101
Query: 90 VMSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQIGDVTIPKYWAHTGLWHWQ 148
VM+G GMLN+G+ TQLLLTLF++EG++H+GIAGN + IGDVT+P+YWAHTGLW+WQ
Sbjct: 102 VMTGLGMLNSGVTTQLLLTLFDVEGIVHFGIAGNADPDLHIGDVTVPRYWAHTGLWNWQ 160
>emb|CAB79345.1| putative protein [Arabidopsis thaliana] gi|5051777|emb|CAB45070.1|
putative protein [Arabidopsis thaliana]
gi|7487318|pir||T09898 hypothetical protein T22A6.180 -
Arabidopsis thaliana
Length = 232
Score = 117 bits (294), Expect = 8e-26
Identities = 54/116 (46%), Positives = 74/116 (63%)
Query: 32 ISNINNEGPSIGIVVPNAYELNPLLHSSSFVPHNKFPYFDFAGRHFRIGELEKKKVIVVM 91
I +N GP IG+V A E N L S F P P+ D +GR FRIG++ KKV+ V
Sbjct: 38 IRKLNRGGPYIGLVTVIATEENAFLRSVDFRPDPTHPFLDLSGRRFRIGKIHGKKVVYVR 97
Query: 92 SGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQIGDVTIPKYWAHTGLWHW 147
G GM+N ATQ ++ +FN++G++H+GIAGN+N+ IGDV+IPK + GLW W
Sbjct: 98 CGRGMVNGAAATQQMIDVFNVKGIVHFGIAGNMNNSMSIGDVSIPKQITNAGLWDW 153
>gb|AAN13118.1| unknown protein [Arabidopsis thaliana] gi|13878031|gb|AAK44093.1|
unknown protein [Arabidopsis thaliana]
gi|30686500|ref|NP_194166.2| phosphorylase family
protein [Arabidopsis thaliana]
Length = 336
Score = 117 bits (294), Expect = 8e-26
Identities = 54/116 (46%), Positives = 74/116 (63%)
Query: 32 ISNINNEGPSIGIVVPNAYELNPLLHSSSFVPHNKFPYFDFAGRHFRIGELEKKKVIVVM 91
I +N GP IG+V A E N L S F P P+ D +GR FRIG++ KKV+ V
Sbjct: 38 IRKLNRGGPYIGLVTVIATEENAFLRSVDFRPDPTHPFLDLSGRRFRIGKIHGKKVVYVR 97
Query: 92 SGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQIGDVTIPKYWAHTGLWHW 147
G GM+N ATQ ++ +FN++G++H+GIAGN+N+ IGDV+IPK + GLW W
Sbjct: 98 CGRGMVNGAAATQQMIDVFNVKGIVHFGIAGNMNNSMSIGDVSIPKQITNAGLWDW 153
>gb|AAM65580.1| unknown [Arabidopsis thaliana]
Length = 336
Score = 117 bits (294), Expect = 8e-26
Identities = 54/116 (46%), Positives = 74/116 (63%)
Query: 32 ISNINNEGPSIGIVVPNAYELNPLLHSSSFVPHNKFPYFDFAGRHFRIGELEKKKVIVVM 91
I +N GP IG+V A E N L S F P P+ D +GR FRIG++ KKV+ V
Sbjct: 38 IRKLNRGGPYIGLVTVIATEENAFLRSVDFRPDPTHPFLDLSGRRFRIGKIHGKKVVYVR 97
Query: 92 SGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQIGDVTIPKYWAHTGLWHW 147
G GM+N ATQ ++ +FN++G++H+GIAGN+N+ IGDV+IPK + GLW W
Sbjct: 98 CGRGMVNGAAATQQMIDVFNVKGIVHFGIAGNMNNSMSIGDVSIPKQITNAGLWDW 153
>dbj|BAD44476.1| putative protein [Arabidopsis thaliana]
Length = 313
Score = 117 bits (294), Expect = 8e-26
Identities = 54/116 (46%), Positives = 74/116 (63%)
Query: 32 ISNINNEGPSIGIVVPNAYELNPLLHSSSFVPHNKFPYFDFAGRHFRIGELEKKKVIVVM 91
I +N GP IG+V A E N L S F P P+ D +GR FRIG++ KKV+ V
Sbjct: 15 IRKLNRGGPYIGLVTVIATEENAFLRSVDFRPDPTHPFLDLSGRRFRIGKIHGKKVVYVR 74
Query: 92 SGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQIGDVTIPKYWAHTGLWHW 147
G GM+N ATQ ++ +FN++G++H+GIAGN+N+ IGDV+IPK + GLW W
Sbjct: 75 CGRGMVNGAAATQQMIDVFNVKGIVHFGIAGNMNNSMSIGDVSIPKQITNAGLWDW 130
>gb|AAM66964.1| unknown [Arabidopsis thaliana]
Length = 338
Score = 109 bits (272), Expect = 3e-23
Identities = 52/116 (44%), Positives = 73/116 (62%)
Query: 32 ISNINNEGPSIGIVVPNAYELNPLLHSSSFVPHNKFPYFDFAGRHFRIGELEKKKVIVVM 91
I IN GP IG+V E + L S F P+ D +GR FRIG++ KKV+ V
Sbjct: 43 IRKINRRGPYIGLVTVFETEEDAFLGSVDFRLDPTHPFLDLSGRRFRIGKIHGKKVVYVR 102
Query: 92 SGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQIGDVTIPKYWAHTGLWHW 147
G GM+NA ATQ ++ +F+++G++H+GIAGN+N+ IGDV+IPK + GLW W
Sbjct: 103 CGIGMVNAAAATQQMIDVFSVKGIVHFGIAGNINNSMSIGDVSIPKQITNAGLWDW 158
>dbj|BAC43650.1| unknown protein [Arabidopsis thaliana] gi|28950909|gb|AAO63378.1|
At4g24340 [Arabidopsis thaliana]
gi|18416327|ref|NP_567699.1| phosphorylase family
protein [Arabidopsis thaliana]
Length = 338
Score = 109 bits (272), Expect = 3e-23
Identities = 52/116 (44%), Positives = 73/116 (62%)
Query: 32 ISNINNEGPSIGIVVPNAYELNPLLHSSSFVPHNKFPYFDFAGRHFRIGELEKKKVIVVM 91
I IN GP IG+V E + L S F P+ D +GR FRIG++ KKV+ V
Sbjct: 43 IRKINRRGPYIGLVTVFETEEDAFLGSVDFRLDPTHPFLDLSGRRFRIGKIHGKKVVYVR 102
Query: 92 SGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQIGDVTIPKYWAHTGLWHW 147
G GM+NA ATQ ++ +F+++G++H+GIAGN+N+ IGDV+IPK + GLW W
Sbjct: 103 CGIGMVNAAAATQQMIDVFSVKGIVHFGIAGNINNSMSIGDVSIPKQITNAGLWDW 158
>gb|AAV65286.1| bark protein-like protein [Thuja occidentalis]
Length = 355
Score = 102 bits (253), Expect = 5e-21
Identities = 52/139 (37%), Positives = 78/139 (55%), Gaps = 3/139 (2%)
Query: 13 LLGNSIYAYGAINQLSWREISNINNEGPSIGIVVPNAYELNPLLHSSS--FVPHNKFPYF 70
L+ ++Y I Q I +N G G+VV EL +L SS F P + P
Sbjct: 17 LIALPFHSYAEIPQEIAERIERVNKRGEHYGLVVSTTAELEVVLEKSSGTFRPDKELPTI 76
Query: 71 DFAGRHFRIGELEKKKVIVVMSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLN-SRFQ 129
D +GR F +G++ ++ ++VM G GM+NA TQL++TLF + V+ YG + N +
Sbjct: 77 DISGRRFHVGDIGGRRTLLVMCGRGMVNAAQTTQLMVTLFRVRAVVQYGRGSSANPHKLN 136
Query: 130 IGDVTIPKYWAHTGLWHWQ 148
IGDV IP+ +AHTGL +W+
Sbjct: 137 IGDVAIPRQFAHTGLLYWE 155
>ref|XP_550486.1| putative vegetative storage protein win4.5 [Oryza sativa (japonica
cultivar-group)] gi|55295909|dbj|BAD67777.1| putative
vegetative storage protein win4.5 [Oryza sativa
(japonica cultivar-group)]
Length = 343
Score = 102 bits (253), Expect = 5e-21
Identities = 50/147 (34%), Positives = 84/147 (57%), Gaps = 4/147 (2%)
Query: 5 LGVLVMIVLLGNSIYA---YGAINQLSWREISNINNEGPSIGIVVPNAYELNPLLHSSSF 61
L +L++ +L G+S A + A++++ W ++ +N G SIG+V+ E L S F
Sbjct: 13 LPLLLLFLLAGSSSMALPVHPAMDRVRW-QVDKVNRRGHSIGLVMSYIDEATALESSGYF 71
Query: 62 VPHNKFPYFDFAGRHFRIGELEKKKVIVVMSGEGMLNAGLATQLLLTLFNIEGVLHYGIA 121
P + P+ D GR + IG + VI ++G+ LNA + Q L+ +F + G++HYG A
Sbjct: 72 RPWHVLPFVDLYGRRYHIGSIRGVNVIYALTGQRRLNAAVTVQTLIDVFTVSGIVHYGTA 131
Query: 122 GNLNSRFQIGDVTIPKYWAHTGLWHWQ 148
G+ N GDV++PK+ A+T W W+
Sbjct: 132 GSSNDSMSFGDVSVPKFVAYTSAWTWK 158
>emb|CAB79344.1| putative protein [Arabidopsis thaliana] gi|5051776|emb|CAB45069.1|
putative protein [Arabidopsis thaliana]
gi|7487317|pir||T09897 hypothetical protein T22A6.170 -
Arabidopsis thaliana
Length = 329
Score = 93.2 bits (230), Expect = 2e-18
Identities = 47/116 (40%), Positives = 68/116 (58%), Gaps = 6/116 (5%)
Query: 32 ISNINNEGPSIGIVVPNAYELNPLLHSSSFVPHNKFPYFDFAGRHFRIGELEKKKVIVVM 91
I IN GP IG+V E + L S F P+ D +G+ + KKV+ V
Sbjct: 40 IRKINRRGPYIGLVTVFETEEDAFLGSVDFRLDPTHPFLDLSGK------IHGKKVVYVR 93
Query: 92 SGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQIGDVTIPKYWAHTGLWHW 147
G GM+NA ATQ ++ +F+++G++H+GIAGN+N+ IGDV+IPK + GLW W
Sbjct: 94 CGIGMVNAAAATQQMIDVFSVKGIVHFGIAGNINNSMSIGDVSIPKQITNAGLWDW 149
>gb|AAA16342.1| vegetative storage protein [Populus balsamifera subsp. trichocarpa
x Populus deltoides] gi|481815|pir||S39502 vegetative
storage protein win4.5 - western balsam poplar x
cottonwood (fragment)
Length = 324
Score = 86.7 bits (213), Expect = 2e-16
Identities = 47/146 (32%), Positives = 80/146 (54%), Gaps = 3/146 (2%)
Query: 4 LLGVLVMIVLLGNSIYAYGAINQLSWREISNINNEGPSIGIVVPNAYELNPLLHSSSFVP 63
L+ ++V +++L + N ++ E +N + +G+V+ + L S F P
Sbjct: 4 LVVMVVGLLVLAQQSFQMSLRNPVA--ETNNCKIDFTRLGLVLTSDTNEKALQDSGLFTP 61
Query: 64 HNKFPYFDFAGRHFRIGELEKKKVIVVMSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGN 123
+ PY D AGR F IG L + ++ V G +NA +A Q+LL F I+G++H+G AG+
Sbjct: 62 DAETPYVDIAGRRFHIGTLNARYIVYVKIGGNSVNAAIAVQILLNRFRIQGIIHFGSAGS 121
Query: 124 LNSRFQI-GDVTIPKYWAHTGLWHWQ 148
L+ + GDV++P A TG W+W+
Sbjct: 122 LDKESIVPGDVSVPLAVAFTGAWNWK 147
>ref|NP_913494.1| unnamed protein product [Oryza sativa (japonica cultivar-group)]
gi|7630247|dbj|BAA94780.1| putative bark storage protein
[Oryza sativa (japonica cultivar-group)]
Length = 331
Score = 82.8 bits (203), Expect = 3e-15
Identities = 34/52 (65%), Positives = 44/52 (84%)
Query: 97 LNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQIGDVTIPKYWAHTGLWHWQ 148
LNAGL TQLLL+LF ++G++H+GIAGN + QIGDVTIP++WAH LW+WQ
Sbjct: 75 LNAGLTTQLLLSLFRVKGIVHWGIAGNADEGLQIGDVTIPEHWAHLSLWNWQ 126
>gb|AAK01124.1| vegetative storage protein PNI288 [Populus balsamifera subsp.
trichocarpa x Populus deltoides]
Length = 267
Score = 78.6 bits (192), Expect = 6e-14
Identities = 35/90 (38%), Positives = 54/90 (59%)
Query: 63 PHNKFPYFDFAGRHFRIGELEKKKVIVVMSGEGMLNAGLATQLLLTLFNIEGVLHYGIAG 122
P PY D GR F IG+++ V+VV G + N L TQ+L L +I G++H+G AG
Sbjct: 2 PSTHVPYIDLVGRRFNIGKIKDVHVVVVNVGGEIPNVVLGTQVLFDLLSIRGIIHFGSAG 61
Query: 123 NLNSRFQIGDVTIPKYWAHTGLWHWQVHTS 152
+++ ++GDV +P+ A TG W W+ + S
Sbjct: 62 SVSDSLRLGDVAVPESVAFTGNWEWKSNAS 91
>emb|CAA49669.1| bark storage protein [Populus deltoides] gi|421813|pir||S31580
storage protein, bark - cottonwood
Length = 329
Score = 74.7 bits (182), Expect = 8e-13
Identities = 47/146 (32%), Positives = 78/146 (53%), Gaps = 4/146 (2%)
Query: 4 LLGVLVMIVLLGNSIYAYGAINQLSWREISNINNEGPSIGIVVPNAYELNPLLHSSSFVP 63
LLG+L +V+ S+ A I+ ++ E SN +G+V + L S + P
Sbjct: 11 LLGLL--LVMPQQSMQA-SLIDPIAEIERSNCKIAHLRLGLVFTSDNNERALQDSGLYSP 67
Query: 64 HNKFPYFDFAGRHFRIGELEKKKVIVVMSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGN 123
++ D AGR F G L ++ V +G +N Q+LL FNI GV+++G AG+
Sbjct: 68 DSEDSSVDIAGRRFHSGTLNGSSIVYVKTGSPSVNMATTLQILLVRFNIHGVIYFGNAGS 127
Query: 124 LNSRFQI-GDVTIPKYWAHTGLWHWQ 148
L+ + + GDV++P+ A TG+W+W+
Sbjct: 128 LDKKTMVPGDVSVPQAVAFTGVWNWK 153
>sp|Q07469|BSPA_POPDE BARK STORAGE PROTEIN A PRECURSOR
Length = 312
Score = 73.2 bits (178), Expect = 2e-12
Identities = 40/126 (31%), Positives = 67/126 (52%), Gaps = 1/126 (0%)
Query: 24 INQLSWREISNINNEGPSIGIVVPNAYELNPLLHSSSFVPHNKFPYFDFAGRHFRIGELE 83
I+ ++ E SN +G+V + L S + P ++ D AGR F G L
Sbjct: 11 IDPIAEIERSNCKIAHLRLGLVFTSDNNERALQDSGLYSPDSEDSSVDIAGRRFHSGTLN 70
Query: 84 KKKVIVVMSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQI-GDVTIPKYWAHT 142
++ V +G +N Q+LL FNI GV+++G AG+L+ + + GDV++P+ A T
Sbjct: 71 GSSIVYVKTGSPSVNMATTLQILLVRFNIHGVIYFGNAGSLDKKTMVPGDVSVPQAVAFT 130
Query: 143 GLWHWQ 148
G+W+W+
Sbjct: 131 GVWNWK 136
>ref|YP_047659.1| putative phosphorylase family protein [Acinetobacter sp. ADP1]
gi|49532125|emb|CAG69837.1| putative phosphorylase
family protein [Acinetobacter sp. ADP1]
Length = 349
Score = 71.2 bits (173), Expect = 9e-12
Identities = 39/115 (33%), Positives = 63/115 (53%), Gaps = 5/115 (4%)
Query: 25 NQLSWREISNINNEGPSIGIVVPNAYELNPLLHSSSFVPHNKFPYFDFAGRHFRIGELEK 84
NQL ++++ + P + I+ E + L+ S+ +K Y GR F G LE
Sbjct: 32 NQLMTNSMNHLTDSTPRLAIIAAFGQEADLLIESTK----DKKEYI-INGRKFVTGTLEG 86
Query: 85 KKVIVVMSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQIGDVTIPKYW 139
+V++ +SG M NA + TQ++ FN++G++ GIAG+LN +GDV I K W
Sbjct: 87 AEVVITLSGISMTNAAMTTQMITDHFNLKGIVFSGIAGSLNPDLHVGDVVIAKSW 141
>pir||S17765 major storage protein - Carolina poplar
Length = 329
Score = 70.9 bits (172), Expect = 1e-11
Identities = 38/119 (31%), Positives = 63/119 (52%), Gaps = 1/119 (0%)
Query: 31 EISNINNEGPSIGIVVPNAYELNPLLHSSSFVPHNKFPYFDFAGRHFRIGELEKKKVIVV 90
E SN +G+V + L S + P ++ D AGR F G L ++ V
Sbjct: 35 ERSNCKIAHLRLGLVFTSDKNERALQDSGLYSPDSEDSSVDIAGRRFHSGTLNGSSIVYV 94
Query: 91 MSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQI-GDVTIPKYWAHTGLWHWQ 148
+G +N Q+LL F+I GV+++G AG+L+ + + GDV++P+ A TG+W+W+
Sbjct: 95 KTGSHSVNMATTLQILLARFSIHGVIYFGNAGSLDKKTMVPGDVSVPEAVAFTGVWNWK 153
>gb|AAB20113.2| major storage protein [Populus x canadensis]
Length = 329
Score = 70.9 bits (172), Expect = 1e-11
Identities = 38/119 (31%), Positives = 63/119 (52%), Gaps = 1/119 (0%)
Query: 31 EISNINNEGPSIGIVVPNAYELNPLLHSSSFVPHNKFPYFDFAGRHFRIGELEKKKVIVV 90
E SN +G+V + L S + P ++ D AGR F G L ++ V
Sbjct: 35 ERSNCKIAHLRLGLVFTSDKNERALQDSGLYSPDSEDSSVDIAGRRFHSGTLNGSSIVYV 94
Query: 91 MSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQI-GDVTIPKYWAHTGLWHWQ 148
+G +N Q+LL F+I GV+++G AG+L+ + + GDV++P+ A TG+W+W+
Sbjct: 95 KTGSHSVNMATTLQILLARFSIHGVIYFGNAGSLDKKTMVPGDVSVPEAVAFTGVWNWK 153
>sp|Q09117|BSPB_POPDE Bark storage protein B precursor gi|169445|gb|AAA50504.1| bark
storage protein gi|445117|prf||1908423A bark storage
protein
Length = 312
Score = 65.9 bits (159), Expect = 4e-10
Identities = 37/119 (31%), Positives = 62/119 (52%), Gaps = 1/119 (0%)
Query: 31 EISNINNEGPSIGIVVPNAYELNPLLHSSSFVPHNKFPYFDFAGRHFRIGELEKKKVIVV 90
E SN +G+V + L S + P ++ D AGR F G L ++ V
Sbjct: 18 ERSNCKIAHLRLGLVFTSDNNERALQDSGLYSPDSEDSSVDIAGRRFHSGTLNGSSIVYV 77
Query: 91 MSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQI-GDVTIPKYWAHTGLWHWQ 148
+G +N Q+LL F+I GV+++G AG+L+ + + GDV++P+ A TG+ +W+
Sbjct: 78 KTGSHSVNMATTLQILLARFSIHGVIYFGNAGSLDKKTMVPGDVSVPQAVAFTGVCNWK 136
>gb|AAK24237.1| phosphorylase family protein [Caulobacter crescentus CB15]
gi|16126505|ref|NP_421069.1| phosphorylase family
protein [Caulobacter crescentus CB15]
gi|25348455|pir||A87530 phosphorylase family protein
[imported] - Caulobacter crescentus
Length = 299
Score = 61.2 bits (147), Expect = 9e-09
Identities = 29/71 (40%), Positives = 42/71 (58%)
Query: 70 FDFAGRHFRIGELEKKKVIVVMSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQ 129
++ G F G+LE K V+V +SG M+NA + TQ+ L FNI ++ GIAG ++
Sbjct: 52 YEVNGVRFMTGKLEGKPVVVFLSGVSMVNAAMTTQMALERFNITRIVFSGIAGGVDESLD 111
Query: 130 IGDVTIPKYWA 140
IGDV + WA
Sbjct: 112 IGDVVVADQWA 122
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.324 0.142 0.448
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 293,775,449
Number of Sequences: 2540612
Number of extensions: 12332085
Number of successful extensions: 30040
Number of sequences better than 10.0: 170
Number of HSP's better than 10.0 without gapping: 148
Number of HSP's successfully gapped in prelim test: 22
Number of HSP's that attempted gapping in prelim test: 29875
Number of HSP's gapped (non-prelim): 177
length of query: 161
length of database: 863,360,394
effective HSP length: 117
effective length of query: 44
effective length of database: 566,108,790
effective search space: 24908786760
effective search space used: 24908786760
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 70 (31.6 bits)
Medicago: description of AC137825.4


