

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137670.5 + phase: 0
(409 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
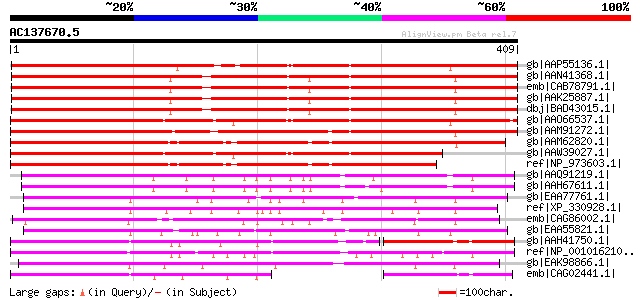
Score E
Sequences producing significant alignments: (bits) Value
gb|AAP55136.1| unknown protein [Oryza sativa (japonica cultivar-... 503 e-141
gb|AAN41368.1| unknown protein [Arabidopsis thaliana] gi|6232009... 500 e-140
emb|CAB78791.1| putative protein [Arabidopsis thaliana] gi|28945... 500 e-140
gb|AAK25887.1| unknown protein [Arabidopsis thaliana] 498 e-139
dbj|BAD43015.1| unknown protein [Arabidopsis thaliana] 498 e-139
gb|AAO66537.1| putative zinc finger protein [Oryza sativa (japon... 497 e-139
gb|AAM91272.1| zinc finger protein Glo3-like [Arabidopsis thalia... 474 e-132
gb|AAM62820.1| zinc finger protein Glo3-like [Arabidopsis thalia... 448 e-124
gb|AAW39027.1| putative zinc finger protein [Oryza sativa (japon... 410 e-113
ref|NP_973603.1| human Rev interacting-like family protein / hRI... 376 e-103
gb|AAQ91219.1| ADP-ribosylation factor GTPase activating protein... 196 1e-48
gb|AAH67611.1| Zgc:85765 protein [Danio rerio] 195 2e-48
gb|EAA77761.1| hypothetical protein FG09712.1 [Gibberella zeae P... 178 2e-43
ref|XP_330928.1| hypothetical protein [Neurospora crassa] gi|289... 176 2e-42
emb|CAG86002.1| unnamed protein product [Debaryomyces hansenii C... 173 8e-42
gb|EAA55821.1| hypothetical protein MG01472.4 [Magnaporthe grise... 167 4e-40
gb|AAH41750.1| Arfgap3-prov protein [Xenopus laevis] 158 3e-37
ref|NP_001016210.1| hypothetical protein LOC548964 [Xenopus trop... 157 4e-37
gb|EAK98866.1| potential ARF GAP [Candida albicans SC5314] gi|46... 157 7e-37
emb|CAG02441.1| unnamed protein product [Tetraodon nigroviridis] 155 2e-36
>gb|AAP55136.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|37537094|ref|NP_922849.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
gi|12643061|gb|AAK00450.1| unknown protein [Oryza
sativa]
Length = 407
Score = 503 bits (1294), Expect = e-141
Identities = 271/413 (65%), Positives = 321/413 (77%), Gaps = 16/413 (3%)
Query: 2 ASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISF 61
+S + DKN VFRKL+ KS+NK CFDCNAKNPTWASVTYG+FLCIDCSAVHRSLGVH+SF
Sbjct: 6 SSSAAADKNTVFRKLRAKSDNKMCFDCNAKNPTWASVTYGVFLCIDCSAVHRSLGVHVSF 65
Query: 62 VRSTNLDSWTPEQLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQLLSKEV 121
VRSTNLDSWTPEQLKMM +GGN+RAQ FF+QHGW GK+EAKYTSRAA+LY+QLL+K+V
Sbjct: 66 VRSTNLDSWTPEQLKMMVYGGNNRAQAFFKQHGWTDGGKIEAKYTSRAADLYRQLLAKDV 125
Query: 122 AKSMSEEAALSAP--PAASSQSAQGTNGLPDVKTNEVPIEKTVEKTVEKPEKTESSSSPR 179
AK+ +E+ S P P A+SQ + +PD+K E E EKT + E SPR
Sbjct: 126 AKNSTEDGNNSWPSSPVAASQPTNQADAIPDLKLAEASKEVANEKT-----EPEVIRSPR 180
Query: 180 AYTAVSNNLKKPIGAKKTG-KSGGLGARKLTRKPSESLYEQKPEELPAPVSSSTITKNNL 238
A T ++ KKPI AKK G K+GGLGARKLT KP+ESLYEQKPEEL AP + +T+N+
Sbjct: 181 APT---HSFKKPIVAKKPGNKTGGLGARKLTSKPNESLYEQKPEEL-AP-ALPPVTENST 235
Query: 239 PSGPPLTSRFEYTEDVQSSELNSGGSNVTGHVSVPKSSSSFFSDFGMDSGFQKKSGPSSS 298
TSRFEY E+ S+ NS + V GHV+ PK SS+FF +FGMDSG+ KKS P S
Sbjct: 236 AKSKSHTSRFEYVENTPSAGSNSEENQVIGHVAPPK-SSNFFGEFGMDSGYHKKSAPGPS 294
Query: 299 KVQIQESDEARKKFSNAKSISSSQFFGDQNKANADAQATLSKFSGSSAISSADLFG--DS 356
KVQI+ES EAR+KFSNAKSISSSQFFGDQ +AQ +L KFSGSSAISSADLFG +
Sbjct: 295 KVQIEESSEARQKFSNAKSISSSQFFGDQASFEKEAQVSLQKFSGSSAISSADLFGHPTN 354
Query: 357 SDNVDLAASDLINRISFQAQQDISSLKNIAGETGKKLTSLASSLMTDLQDRIL 409
S NVDL+ASDLINR+SFQA QD+SS+KN+AGETGKKLTSLAS++M+DLQDRIL
Sbjct: 355 SSNVDLSASDLINRLSFQASQDLSSIKNMAGETGKKLTSLASNIMSDLQDRIL 407
>gb|AAN41368.1| unknown protein [Arabidopsis thaliana] gi|62320091|dbj|BAD94263.1|
hypothetical protein [Arabidopsis thaliana]
gi|62319827|dbj|BAD93852.1| hypothetical protein
[Arabidopsis thaliana] gi|18414983|ref|NP_567543.1|
human Rev interacting-like family protein / hRIP family
protein [Arabidopsis thaliana]
gi|51971433|dbj|BAD44381.1| unknown protein [Arabidopsis
thaliana] gi|51970716|dbj|BAD44050.1| unknown protein
[Arabidopsis thaliana]
Length = 413
Score = 500 bits (1287), Expect = e-140
Identities = 263/418 (62%), Positives = 327/418 (77%), Gaps = 19/418 (4%)
Query: 2 ASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISF 61
++D+ TDKN VFRKLK+KSENK CFDC+AKNPTWASVTYGIFLCIDCSA HR+LGVHISF
Sbjct: 5 SADNLTDKNIVFRKLKSKSENKVCFDCSAKNPTWASVTYGIFLCIDCSATHRNLGVHISF 64
Query: 62 VRSTNLDSWTPEQLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQLLSKEV 121
VRSTNLDSW+PEQL+ M FGGN+RAQVFF+QHGW GK+EAKYTSRAA+LY+Q+L+KEV
Sbjct: 65 VRSTNLDSWSPEQLRTMMFGGNNRAQVFFKQHGWTDGGKIEAKYTSRAADLYRQILAKEV 124
Query: 122 AKSMSEE--AALSAPPAASSQSAQGTNGLPDVKTNEVPIEKTVEKTVEKPEKTESSSSPR 179
AK+++EE + L + P A+SQ + +NG+ E E + K E T ++SSP+
Sbjct: 125 AKAIAEETNSGLLSSPVATSQLPEVSNGVSSYSVKE-------ELPLSKHEATSATSSPK 177
Query: 180 A-YTAVSNNLKKPIGAKKTGKSGGLGARKLTRKPSESLYEQKPEELPAPVSSSTITKNNL 238
A T V + KKPIGAK+TGK+GGLGARKLT KP ++LYEQKPEE+ APV + + NN
Sbjct: 178 ASNTVVPSTFKKPIGAKRTGKTGGLGARKLTTKPKDNLYEQKPEEV-APVIPAVSSTNNG 236
Query: 239 PS----GPPLTSRFEYTEDVQSSELNSGGSNVTGHVSVPKSSSSFFSDFGMDSGFQKKSG 294
S G SRFEY +D+QS + GG+ V HV+ PK SSSFFSDFGMDS F KKS
Sbjct: 237 ESKSSAGSSFASRFEYNDDLQSGGQSVGGTQVLNHVAPPK-SSSFFSDFGMDSSFPKKSS 295
Query: 295 PSSSKVQIQESDEARKKFSNAKSISSSQFFGDQNK-ANADAQATLSKFSGSSAISSADLF 353
+SSK Q++ESDEARKKF+NAKSISS+Q+FGDQNK A+ +++ATL KF+GS++ISSAD +
Sbjct: 296 SNSSKSQVEESDEARKKFTNAKSISSAQYFGDQNKNADLESKATLQKFAGSASISSADFY 355
Query: 354 GDSSD--NVDLAASDLINRISFQAQQDISSLKNIAGETGKKLTSLASSLMTDLQDRIL 409
G D N+D+ ASDLINR+SFQAQQD+SSL NIAGET KKL +LAS + +D+QDR+L
Sbjct: 356 GHDQDDSNIDITASDLINRLSFQAQQDLSSLVNIAGETKKKLGTLASGIFSDIQDRML 413
>emb|CAB78791.1| putative protein [Arabidopsis thaliana] gi|2894598|emb|CAA17132.1|
putative protein [Arabidopsis thaliana]
gi|7487780|pir||T05075 hypothetical protein T6K21.70 -
Arabidopsis thaliana
Length = 1082
Score = 500 bits (1287), Expect = e-140
Identities = 263/418 (62%), Positives = 327/418 (77%), Gaps = 19/418 (4%)
Query: 2 ASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISF 61
++D+ TDKN VFRKLK+KSENK CFDC+AKNPTWASVTYGIFLCIDCSA HR+LGVHISF
Sbjct: 5 SADNLTDKNIVFRKLKSKSENKVCFDCSAKNPTWASVTYGIFLCIDCSATHRNLGVHISF 64
Query: 62 VRSTNLDSWTPEQLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQLLSKEV 121
VRSTNLDSW+PEQL+ M FGGN+RAQVFF+QHGW GK+EAKYTSRAA+LY+Q+L+KEV
Sbjct: 65 VRSTNLDSWSPEQLRTMMFGGNNRAQVFFKQHGWTDGGKIEAKYTSRAADLYRQILAKEV 124
Query: 122 AKSMSEE--AALSAPPAASSQSAQGTNGLPDVKTNEVPIEKTVEKTVEKPEKTESSSSPR 179
AK+++EE + L + P A+SQ + +NG+ E E + K E T ++SSP+
Sbjct: 125 AKAIAEETNSGLLSSPVATSQLPEVSNGVSSYSVKE-------ELPLSKHEATSATSSPK 177
Query: 180 A-YTAVSNNLKKPIGAKKTGKSGGLGARKLTRKPSESLYEQKPEELPAPVSSSTITKNNL 238
A T V + KKPIGAK+TGK+GGLGARKLT KP ++LYEQKPEE+ APV + + NN
Sbjct: 178 ASNTVVPSTFKKPIGAKRTGKTGGLGARKLTTKPKDNLYEQKPEEV-APVIPAVSSTNNG 236
Query: 239 PS----GPPLTSRFEYTEDVQSSELNSGGSNVTGHVSVPKSSSSFFSDFGMDSGFQKKSG 294
S G SRFEY +D+QS + GG+ V HV+ PK SSSFFSDFGMDS F KKS
Sbjct: 237 ESKSSAGSSFASRFEYNDDLQSGGQSVGGTQVLNHVAPPK-SSSFFSDFGMDSSFPKKSS 295
Query: 295 PSSSKVQIQESDEARKKFSNAKSISSSQFFGDQNK-ANADAQATLSKFSGSSAISSADLF 353
+SSK Q++ESDEARKKF+NAKSISS+Q+FGDQNK A+ +++ATL KF+GS++ISSAD +
Sbjct: 296 SNSSKSQVEESDEARKKFTNAKSISSAQYFGDQNKNADLESKATLQKFAGSASISSADFY 355
Query: 354 GDSSD--NVDLAASDLINRISFQAQQDISSLKNIAGETGKKLTSLASSLMTDLQDRIL 409
G D N+D+ ASDLINR+SFQAQQD+SSL NIAGET KKL +LAS + +D+QDR+L
Sbjct: 356 GHDQDDSNIDITASDLINRLSFQAQQDLSSLVNIAGETKKKLGTLASGIFSDIQDRML 413
>gb|AAK25887.1| unknown protein [Arabidopsis thaliana]
Length = 413
Score = 498 bits (1283), Expect = e-139
Identities = 262/418 (62%), Positives = 327/418 (77%), Gaps = 19/418 (4%)
Query: 2 ASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISF 61
++D+ TDKN VFRKLK+KSENK CFDC+AKNPTWASVTYGIFLCIDCSA HR+LGVHISF
Sbjct: 5 SADNLTDKNIVFRKLKSKSENKVCFDCSAKNPTWASVTYGIFLCIDCSATHRNLGVHISF 64
Query: 62 VRSTNLDSWTPEQLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQLLSKEV 121
VRSTNLDSW+PEQL+ M FGGN+RAQVFF+QHGW GK+EAKYTSRAA+LY+Q+L+KEV
Sbjct: 65 VRSTNLDSWSPEQLRTMMFGGNNRAQVFFKQHGWTDGGKIEAKYTSRAADLYRQILAKEV 124
Query: 122 AKSMSEE--AALSAPPAASSQSAQGTNGLPDVKTNEVPIEKTVEKTVEKPEKTESSSSPR 179
AK+++EE + L + P A+SQ + +NG+ E E + K E T ++SSP+
Sbjct: 125 AKAIAEETNSGLLSSPVATSQLPEVSNGVSSYSVKE-------ELPLSKHEATSATSSPK 177
Query: 180 A-YTAVSNNLKKPIGAKKTGKSGGLGARKLTRKPSESLYEQKPEELPAPVSSSTITKNNL 238
A T V + KKPIGAK+TGK+GGLGARKLT KP ++LYEQKPEE+ APV + + NN
Sbjct: 178 ASNTVVPSTFKKPIGAKRTGKTGGLGARKLTTKPKDNLYEQKPEEV-APVIPAVSSTNNG 236
Query: 239 PS----GPPLTSRFEYTEDVQSSELNSGGSNVTGHVSVPKSSSSFFSDFGMDSGFQKKSG 294
S G SRFEY +D+QS + GG+ V HV+ PK SSSFFSDFGMDS F KKS
Sbjct: 237 ESKSSAGSSFASRFEYNDDLQSGGQSVGGTQVLNHVAPPK-SSSFFSDFGMDSSFPKKSS 295
Query: 295 PSSSKVQIQESDEARKKFSNAKSISSSQFFGDQNK-ANADAQATLSKFSGSSAISSADLF 353
+SSK Q++ESDEAR+KF+NAKSISS+Q+FGDQNK A+ +++ATL KF+GS++ISSAD +
Sbjct: 296 SNSSKSQVEESDEAREKFTNAKSISSAQYFGDQNKNADLESKATLQKFAGSASISSADFY 355
Query: 354 GDSSD--NVDLAASDLINRISFQAQQDISSLKNIAGETGKKLTSLASSLMTDLQDRIL 409
G D N+D+ ASDLINR+SFQAQQD+SSL NIAGET KKL +LAS + +D+QDR+L
Sbjct: 356 GHDQDDSNIDITASDLINRLSFQAQQDLSSLVNIAGETKKKLGTLASGIFSDIQDRML 413
>dbj|BAD43015.1| unknown protein [Arabidopsis thaliana]
Length = 413
Score = 498 bits (1283), Expect = e-139
Identities = 263/418 (62%), Positives = 326/418 (77%), Gaps = 19/418 (4%)
Query: 2 ASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISF 61
++D+ TDKN VFRKLK+KSENK CFDC+AKNPTWASVTYGIFLCIDCSA HR+LGVHISF
Sbjct: 5 SADNLTDKNIVFRKLKSKSENKVCFDCSAKNPTWASVTYGIFLCIDCSATHRNLGVHISF 64
Query: 62 VRSTNLDSWTPEQLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQLLSKEV 121
VRSTNLDSW+PEQL+ M FGGN+RAQVFF+QHGW GK+EAKYTSRAA+LY+Q+L+KEV
Sbjct: 65 VRSTNLDSWSPEQLRTMMFGGNNRAQVFFKQHGWTDGGKIEAKYTSRAADLYRQILAKEV 124
Query: 122 AKSMSEE--AALSAPPAASSQSAQGTNGLPDVKTNEVPIEKTVEKTVEKPEKTESSSSPR 179
AK+++EE + L + P A+SQ + +NG+ E E + K E T ++SSP+
Sbjct: 125 AKAIAEETNSGLLSSPVATSQLPEVSNGVSSYSVKE-------ELPLSKHEATSATSSPK 177
Query: 180 A-YTAVSNNLKKPIGAKKTGKSGGLGARKLTRKPSESLYEQKPEELPAPVSSSTITKNNL 238
A T V + KKPIGAK+TGK+GGLGARKLT KP ++LYEQKPEE+ APV + + NN
Sbjct: 178 ASNTVVPSTFKKPIGAKRTGKTGGLGARKLTTKPKDNLYEQKPEEV-APVIPAVSSTNNG 236
Query: 239 PS----GPPLTSRFEYTEDVQSSELNSGGSNVTGHVSVPKSSSSFFSDFGMDSGFQKKSG 294
S G SRFEY +D+QS + GG+ V HV+ PK SSSFFSDFGMDS F KKS
Sbjct: 237 ESKSSAGSSFASRFEYNDDLQSGGQSVGGTQVLNHVAPPK-SSSFFSDFGMDSSFPKKSS 295
Query: 295 PSSSKVQIQESDEARKKFSNAKSISSSQFFGDQNK-ANADAQATLSKFSGSSAISSADLF 353
+SSK Q++ESDEARKKF+NAKSISS+Q+FGDQNK A+ +++ATL KF+GS++ISSAD
Sbjct: 296 SNSSKSQVEESDEARKKFTNAKSISSAQYFGDQNKNADLESKATLQKFAGSASISSADFH 355
Query: 354 GDSSD--NVDLAASDLINRISFQAQQDISSLKNIAGETGKKLTSLASSLMTDLQDRIL 409
G D N+D+ ASDLINR+SFQAQQD+SSL NIAGET KKL +LAS + +D+QDR+L
Sbjct: 356 GHDQDDSNIDITASDLINRLSFQAQQDLSSLVNIAGETKKKLGTLASGIFSDIQDRML 413
>gb|AAO66537.1| putative zinc finger protein [Oryza sativa (japonica
cultivar-group)] gi|50920175|ref|XP_470448.1| putative
zinc finger protein [Oryza sativa (japonica
cultivar-group)]
Length = 412
Score = 497 bits (1279), Expect = e-139
Identities = 263/419 (62%), Positives = 328/419 (77%), Gaps = 17/419 (4%)
Query: 1 MASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 60
MA D+FTDKNAVFR+LK K ENK CFDC+AKNPTWASVTYGIFLC+DCSAVHRSLGVHI+
Sbjct: 1 MAFDAFTDKNAVFRRLKAKPENKMCFDCSAKNPTWASVTYGIFLCLDCSAVHRSLGVHIT 60
Query: 61 FVRSTNLDSWTPEQLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQLLSKE 120
FVRSTNLDSWTP+QLKMM+FGGN+RA FF+QHGW GKV+AKYTSRAAELY+Q+L KE
Sbjct: 61 FVRSTNLDSWTPDQLKMMAFGGNNRAHAFFKQHGWTDGGKVDAKYTSRAAELYRQILQKE 120
Query: 121 VAKSMSEEAALSAPPAASSQSAQGTNGLPDVKTNEVPIEKTVEKTVEKPEKTESSSSPR- 179
VAKS S + L + P A+SQ ++ P+ K E P E T K ++P+ T S +P
Sbjct: 121 VAKS-SADNVLPSSPVAASQPQNPSDDFPEFKLPEAPAENTNGK--QEPDVTNSQKAPTQ 177
Query: 180 -----AYTAVSNNLKKPIGAKKT-GKSGGLGARKLTRKPSESLYEQKPEELPAPVSSSTI 233
+ + ++KK IGAKK GK+GGLG +KLT KPSESLY+QKPEE P P ++ +
Sbjct: 178 TPKAPTHPTFATSVKKSIGAKKIGGKTGGLGVKKLTTKPSESLYDQKPEE-PKP-AAPVM 235
Query: 234 TKNNLPSGPPLTSRFEYTEDVQSSELNSGGSNVTGHVSVPKSSSSFFSDFGMDSGFQKKS 293
T + SGP L SRFEY E+ + + +GG+ +TGHV+ PK SS+FF ++GMD+GFQKK+
Sbjct: 236 TTSTTKSGPSLHSRFEYVENEPAVDSRNGGTQMTGHVAPPK-SSNFFQEYGMDNGFQKKT 294
Query: 294 GPSSSKVQIQESDEARKKFSNAKSISSSQFFGDQNKANADAQATLSKFSGSSAISSADLF 353
+++K QIQE+DEARKKFSNAK+ISSSQFFG+Q++ DAQ +L KF+GSS+ISSADLF
Sbjct: 295 STAATKTQIQETDEARKKFSNAKAISSSQFFGNQSREEKDAQMSLQKFAGSSSISSADLF 354
Query: 354 G--DSSD-NVDLAASDLINRISFQAQQDISSLKNIAGETGKKLTSLASSLMTDLQDRIL 409
G D D N+DL+A+DLINRISFQA QD+SSLKN+AGETGKKLTS+AS+ ++DL DRIL
Sbjct: 355 GRRDMDDSNLDLSAADLINRISFQASQDLSSLKNMAGETGKKLTSIASNFISDL-DRIL 412
>gb|AAM91272.1| zinc finger protein Glo3-like [Arabidopsis thaliana]
gi|9758511|dbj|BAB08919.1| zinc finger protein Glo3-like
[Arabidopsis thaliana] gi|20466454|gb|AAM20544.1| zinc
finger protein Glo3-like [Arabidopsis thaliana]
gi|15237500|ref|NP_199487.1| human Rev interacting-like
family protein / hRIP family protein [Arabidopsis
thaliana]
Length = 402
Score = 474 bits (1219), Expect = e-132
Identities = 249/412 (60%), Positives = 314/412 (75%), Gaps = 13/412 (3%)
Query: 1 MASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 60
MA+++ TDKN VFRKLK+KSENK CFDC+AKNPTWASV YGIFLCIDCSAVHRSLGVHIS
Sbjct: 1 MATENLTDKNVVFRKLKSKSENKVCFDCSAKNPTWASVPYGIFLCIDCSAVHRSLGVHIS 60
Query: 61 FVRSTNLDSWTPEQLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQLLSKE 120
FVRSTNLDSW+PEQL+ M FGGN+RAQVFF+QHGWN GK+EAKYTSRAA++Y+Q L+KE
Sbjct: 61 FVRSTNLDSWSPEQLRTMMFGGNNRAQVFFKQHGWNDGGKIEAKYTSRAADMYRQTLAKE 120
Query: 121 VAKSMSEEAALSAPPAASSQSAQGTNGLPDVKTNEVPIEKTVEKTVEKPEKTESSSSPRA 180
VAK+M+EE L P +S ++Q + T+E P E ++ K E SS +
Sbjct: 121 VAKAMAEETVL--PSLSSVATSQPVESSENGFTSESPKESSL-----KQEAAVVSSPKAS 173
Query: 181 YTAVSNNLKKPIGAKKTGKSGGLGARKLTRKPSESLYEQKPEELPAPVSSSTITKNNLPS 240
V++ KKP+ ++K+GK+GGLGARKLT K ++LYEQKPEE + +++ T + +
Sbjct: 174 QKVVASTFKKPLVSRKSGKTGGLGARKLTTKSKDNLYEQKPEEPVPVIPAASPTNDTSAA 233
Query: 241 GPPLTSRFEYTEDVQSSELNSGGSNVTGHVSVPKSSSSFFSDFGMDSGFQKKSGPSSSKV 300
G SRFEY +D QS G+ V HV+ PK SS+FF++FGMDS F KKS SSSK
Sbjct: 234 GSSFASRFEYFDDEQSG--GQSGTRVLSHVAPPK-SSNFFNEFGMDSAFPKKSSSSSSKA 290
Query: 301 QIQESDEARKKFSNAKSISSSQFFGDQNK-ANADAQATLSKFSGSSAISSADLFGDSSD- 358
Q++E+DEARKKFSNAKSISS+QFFG+QN+ A+ D++ATL KFSGS+AISS+DLFG D
Sbjct: 291 QVEETDEARKKFSNAKSISSAQFFGNQNRDADLDSKATLQKFSGSAAISSSDLFGHGPDD 350
Query: 359 -NVDLAASDLINRISFQAQQDISSLKNIAGETGKKLTSLASSLMTDLQDRIL 409
N+D+ ASDLINRISFQAQQD+SS+ N+A ET KL + ASS+ +DLQDR+L
Sbjct: 351 SNIDITASDLINRISFQAQQDMSSIANLAEETKNKLGTFASSIFSDLQDRML 402
>gb|AAM62820.1| zinc finger protein Glo3-like [Arabidopsis thaliana]
gi|3668084|gb|AAC61816.1| expressed protein [Arabidopsis
thaliana] gi|25408361|pir||H84765 hypothetical protein
At2g35210 [imported] - Arabidopsis thaliana
gi|18403775|ref|NP_565801.1| human Rev interacting-like
family protein / hRIP family protein [Arabidopsis
thaliana]
Length = 395
Score = 448 bits (1153), Expect = e-124
Identities = 249/404 (61%), Positives = 305/404 (74%), Gaps = 15/404 (3%)
Query: 1 MASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 60
MAS++ DK +VF+KLK KS+NK CFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS
Sbjct: 1 MASENLNDKISVFKKLKAKSDNKICFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 60
Query: 61 FVRSTNLDSWTPEQLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQLLSKE 120
FVRSTNLDSW+ EQLKMM +GGN+RAQVFF+Q+GW+ GK EAKYTSRAA+LYKQ+L+KE
Sbjct: 61 FVRSTNLDSWSSEQLKMMIYGGNNRAQVFFKQYGWSDGGKTEAKYTSRAADLYKQILAKE 120
Query: 121 VAKSMSEEAALSAPPAASSQSAQGTNGLPDVKTNEVPIEKTVEKTVEKPEKTESSSSPRA 180
VAKS +EE L PP + S Q NGL +KT+E E K EKP+ SPR
Sbjct: 121 VAKSKAEE-ELDLPP-SPPDSTQVPNGLSSIKTSEALKESNTLKQQEKPDVV--PVSPR- 175
Query: 181 YTAVSNNLKKPIGAKKTGKSGGLGARKLTRKPSESLYEQKPEELPAPVSSSTITKNNLPS 240
+S ++KKP+GAKKTGK+GGLGARKLT K S +LY+QKPEE ++S ++ + S
Sbjct: 176 ---ISRSVKKPLGAKKTGKTGGLGARKLTTKSSGTLYDQKPEESVIIQATSPVSAKSARS 232
Query: 241 GPPLTSRFEYTEDVQSSELNSGGSNVTGHVSVPKSSSSFFSDFGMDSG-FQKKSGPSSSK 299
+SRF+Y ++VQ+ E + V HV+ PKSS F + M+ G FQKK SSSK
Sbjct: 233 S--FSSRFDYADNVQNRE-DYMSPQVVSHVAPPKSSGFFEEELEMNGGRFQKKPITSSSK 289
Query: 300 VQIQESDEARKKFSNAKSISSSQFFG-DQNKANADAQATLSKFSGSSAISSADLFGDSSD 358
+QIQE+DEARKKF+NAKSISS+Q+FG D N A+ +A+++L KFSGSSAISSADLFGD
Sbjct: 290 LQIQETDEARKKFTNAKSISSAQYFGNDNNSADLEAKSSLKKFSGSSAISSADLFGDGDG 349
Query: 359 N--VDLAASDLINRISFQAQQDISSLKNIAGETGKKLTSLASSL 400
+ +DL A DL+NR+S QAQQDISSLKN+A ET KKL S+ASSL
Sbjct: 350 DFPLDLTAGDLLNRLSLQAQQDISSLKNMAEETKKKLGSVASSL 393
>gb|AAW39027.1| putative zinc finger protein [Oryza sativa (japonica
cultivar-group)]
Length = 384
Score = 410 bits (1055), Expect = e-113
Identities = 212/356 (59%), Positives = 270/356 (75%), Gaps = 13/356 (3%)
Query: 1 MASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 60
MA D+FTDKNAVFR+LK K ENK CFDC+AKNPTWASVTYGIFLC+DCSAVHRSLGVHI+
Sbjct: 1 MAFDAFTDKNAVFRRLKAKPENKMCFDCSAKNPTWASVTYGIFLCLDCSAVHRSLGVHIT 60
Query: 61 FVRSTNLDSWTPEQLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQLLSKE 120
FVRSTNLDSWTP+QLKMM+FGGN+RA FF+QHGW GKV+AKYTSRAAELY+Q+L KE
Sbjct: 61 FVRSTNLDSWTPDQLKMMAFGGNNRAHAFFKQHGWTDGGKVDAKYTSRAAELYRQILQKE 120
Query: 121 VAKSMSEEAALSAPPAASSQSAQGTNGLPDVKTNEVPIEKTVEKTVEKPEKTESSSSPR- 179
VAKS S + L + P A+SQ ++ P+ K E P E T K ++P+ T S +P
Sbjct: 121 VAKS-SADNVLPSSPVAASQPQNPSDDFPEFKLPEAPAENTNGK--QEPDVTNSQKAPTQ 177
Query: 180 -----AYTAVSNNLKKPIGAKKT-GKSGGLGARKLTRKPSESLYEQKPEELPAPVSSSTI 233
+ + ++KK IGAKK GK+GGLG +KLT KPSESLY+QKPEE P P ++ +
Sbjct: 178 TPKAPTHPTFATSVKKSIGAKKIGGKTGGLGVKKLTTKPSESLYDQKPEE-PKP-AAPVM 235
Query: 234 TKNNLPSGPPLTSRFEYTEDVQSSELNSGGSNVTGHVSVPKSSSSFFSDFGMDSGFQKKS 293
T + SGP L SRFEY E+ + + +GG+ +TGHV+ PK SS+FF ++GMD+GFQKK+
Sbjct: 236 TTSTTKSGPSLHSRFEYVENEPAVDSRNGGTQMTGHVAPPK-SSNFFQEYGMDNGFQKKT 294
Query: 294 GPSSSKVQIQESDEARKKFSNAKSISSSQFFGDQNKANADAQATLSKFSGSSAISS 349
+++K QIQE+DEARKKFSNAK+ISSSQFFG+Q++ DAQ +L KF+ +++S
Sbjct: 295 STAATKTQIQETDEARKKFSNAKAISSSQFFGNQSREEKDAQMSLQKFAVRISLAS 350
>ref|NP_973603.1| human Rev interacting-like family protein / hRIP family protein
[Arabidopsis thaliana]
Length = 371
Score = 376 bits (966), Expect = e-103
Identities = 207/346 (59%), Positives = 257/346 (73%), Gaps = 13/346 (3%)
Query: 1 MASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 60
MAS++ DK +VF+KLK KS+NK CFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS
Sbjct: 1 MASENLNDKISVFKKLKAKSDNKICFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 60
Query: 61 FVRSTNLDSWTPEQLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQLLSKE 120
FVRSTNLDSW+ EQLKMM +GGN+RAQVFF+Q+GW+ GK EAKYTSRAA+LYKQ+L+KE
Sbjct: 61 FVRSTNLDSWSSEQLKMMIYGGNNRAQVFFKQYGWSDGGKTEAKYTSRAADLYKQILAKE 120
Query: 121 VAKSMSEEAALSAPPAASSQSAQGTNGLPDVKTNEVPIEKTVEKTVEKPEKTESSSSPRA 180
VAKS +EE L PP + S Q NGL +KT+E E K EKP+ SPR
Sbjct: 121 VAKSKAEE-ELDLPP-SPPDSTQVPNGLSSIKTSEALKESNTLKQQEKPDVV--PVSPR- 175
Query: 181 YTAVSNNLKKPIGAKKTGKSGGLGARKLTRKPSESLYEQKPEELPAPVSSSTITKNNLPS 240
+S ++KKP+GAKKTGK+GGLGARKLT K S +LY+QKPEE ++S ++ + S
Sbjct: 176 ---ISRSVKKPLGAKKTGKTGGLGARKLTTKSSGTLYDQKPEESVIIQATSPVSAKSARS 232
Query: 241 GPPLTSRFEYTEDVQSSELNSGGSNVTGHVSVPKSSSSFFSDFGMDSG-FQKKSGPSSSK 299
+SRF+Y ++VQ+ E + V HV+ PKSS F + M+ G FQKK SSSK
Sbjct: 233 S--FSSRFDYADNVQNRE-DYMSPQVVSHVAPPKSSGFFEEELEMNGGRFQKKPITSSSK 289
Query: 300 VQIQESDEARKKFSNAKSISSSQFFG-DQNKANADAQATLSKFSGS 344
+QIQE+DEARKKF+NAKSISS+Q+FG D N A+ +A+++L KFS S
Sbjct: 290 LQIQETDEARKKFTNAKSISSAQYFGNDNNSADLEAKSSLKKFSQS 335
>gb|AAQ91219.1| ADP-ribosylation factor GTPase activating protein 3 [Danio rerio]
gi|45387621|ref|NP_991160.1| ADP-ribosylation factor
GTPase activating protein 3 [Danio rerio]
Length = 498
Score = 196 bits (497), Expect = 1e-48
Identities = 163/493 (33%), Positives = 231/493 (46%), Gaps = 102/493 (20%)
Query: 10 NAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDS 69
+AV ++L+ + NK CFDC+AKNP+WAS+TYG+FLCIDCS HRSLGVH+SF+RST LDS
Sbjct: 10 SAVLKRLRAAAANKVCFDCSAKNPSWASITYGVFLCIDCSGTHRSLGVHLSFIRSTELDS 69
Query: 70 -WTPEQLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYK---QLLSKEVAKSM 125
W+ QL+ M GGN+ A FF QHG + AKY+SRAA LY+ + L+ + +
Sbjct: 70 NWSWFQLRCMQVGGNASANAFFSQHGCSSSSAANAKYSSRAAALYRDKIRALANQATRQH 129
Query: 126 SEEAALSAPPAASSQS---------AQGT-NGLPD-VKTNEVPIEKTVE------KTVEK 168
E L A S S Q T + LPD V+ N +K V+ K+
Sbjct: 130 GTELWLDAQAPLSPSSPLDKQEDFFTQHTQSALPDTVQLNISQSQKRVQDNNNNAKSEAG 189
Query: 169 PEKTESSSSPRAYTAVSNNL--KKPIGAKKT---GKSGGLGARKLT----------RKPS 213
P + S SP A +L KKP +K+ G+ GLGA+K++ +
Sbjct: 190 PSVEQLSVSPVQSAAEPLSLLKKKPAAVRKSAGGGRRAGLGAQKVSSQSFTLQQRRAQAE 249
Query: 214 ESLYEQKPEELP-------------------APVSSSTITK----NNLPSG--------- 241
+ L EQ+ P A SSS+ K L G
Sbjct: 250 DRLTEQRTSTAPDHSVWSVSRVQQAVDVQRSAQDSSSSRMKQQQEERLGMGLSGRSGLSH 309
Query: 242 --------------PPLTSRFEYTEDVQSSELNSGGSNVTGHVSVPKSSSSFFSDFGMDS 287
P L+SR Y ED Q + + + SS FFS + +
Sbjct: 310 SVTEDMQTIIQQDCPSLSSRRVYEEDEQQEDTHFCSRALD---EGRGSSDLFFSQWSSEK 366
Query: 288 GFQKK-----------SGPSSSKVQIQESDEARKKFSNAKSISSSQFFGDQNKANADAQA 336
+ K +S K D+A++KF +AK+ISS FFG Q+++ + +A
Sbjct: 367 QSRMKPESDYCVEDEHRASASRKEPQSVCDDAQRKFGDAKAISSDMFFGTQDRSEYEVRA 426
Query: 337 TLSKFSGSSAISSADLFGDSSDNVDLAASDLINRIS--FQAQQDISSLKNIAGETGKKLT 394
L FS SSAISSADLF D AA R+S + D++ L++ KL+
Sbjct: 427 RLENFSSSSAISSADLF----DEQKKAAGSSSYRLSSVLSSVPDMTQLRSGVRSVAGKLS 482
Query: 395 SLASSLMTDLQDR 407
+AS +++ +QDR
Sbjct: 483 GMASGVVSTIQDR 495
>gb|AAH67611.1| Zgc:85765 protein [Danio rerio]
Length = 498
Score = 195 bits (495), Expect = 2e-48
Identities = 165/495 (33%), Positives = 231/495 (46%), Gaps = 106/495 (21%)
Query: 10 NAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDS 69
+AV ++L+ + NK CFDC+AKNP+WAS+TYG+FLCIDCS HRSLGVH+SF+RST LDS
Sbjct: 10 SAVLKRLRAAAANKVCFDCSAKNPSWASITYGVFLCIDCSGTHRSLGVHLSFIRSTELDS 69
Query: 70 -WTPEQLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYK---QLLSKEVAKSM 125
W+ QL+ M GGN+ A FF QHG + AKY+SRAA LY+ + L+ + +
Sbjct: 70 NWSWFQLRCMQVGGNASANAFFSQHGCSSSSAANAKYSSRAAALYRDKIRALANQATRQH 129
Query: 126 SEEAALSAPPAASSQS---------AQGT-NGLPD-VKTNEVPIEKTVE------KTVEK 168
E L A S S Q T + LPD V+ N +K V+ K+
Sbjct: 130 GTELWLDAQAPLSPSSPLDKQEDFFTQHTQSALPDTVQLNISQSQKRVQDNNNNAKSEAG 189
Query: 169 PEKTESSSSPRAYTAVSNNL--KKPIGAKKT---GKSGGLGARKLT----------RKPS 213
P + S SP A +L KKP +K+ G+ GLGA+K++ +
Sbjct: 190 PSVEQLSVSPVQSAAEPLSLLKKKPAAVRKSAGGGRRAGLGAQKVSSQSFTLQQRRAQAE 249
Query: 214 ESLYEQKPEELP-------------------APVSSSTITK----NNLPSG--------- 241
+ L EQ+ P A SSS+ K L G
Sbjct: 250 DRLTEQRTSTAPDHSVWSVSRVQQAVDVQRSAQDSSSSRMKQQQEERLGMGLSGRSGLSH 309
Query: 242 --------------PPLTSRFEYTEDVQSSELNSGGSNVTGHVSVPKSSSSFFSDFGMDS 287
P L+SR Y ED Q + + V SS FFS + +
Sbjct: 310 SVTEDMQTIIQQDCPSLSSRRVYEEDEQQEDTHFCSRAVD---EGRGSSDLFFSQWSSEK 366
Query: 288 GFQKKSGPSSS-------------KVQIQESDEARKKFSNAKSISSSQFFGDQNKANADA 334
Q + P S K D+A++KF +AK+ISS FFG Q+++ +
Sbjct: 367 --QGRMKPESDYCVEDEHRASARRKEPQSVCDDAQRKFGDAKAISSDMFFGTQDRSEYEV 424
Query: 335 QATLSKFSGSSAISSADLFGDSSDNVDLAASDLINRIS--FQAQQDISSLKNIAGETGKK 392
+A L FS SSAISSADLF D AA R+S + D++ L++ K
Sbjct: 425 RARLENFSSSSAISSADLF----DEQKKAAGSSSYRLSSVLSSVPDMTQLRSGVRSVAGK 480
Query: 393 LTSLASSLMTDLQDR 407
L+ +AS +++ +QDR
Sbjct: 481 LSGMASGVVSTIQDR 495
>gb|EAA77761.1| hypothetical protein FG09712.1 [Gibberella zeae PH-1]
gi|46136393|ref|XP_389888.1| hypothetical protein
FG09712.1 [Gibberella zeae PH-1]
Length = 479
Score = 178 bits (452), Expect = 2e-43
Identities = 144/470 (30%), Positives = 205/470 (42%), Gaps = 85/470 (18%)
Query: 12 VFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWT 71
+F KLKTK NK CFDC KNPTW SV +GI+LC+DCS+ HR+LGVHISFVRSTNLD W
Sbjct: 13 IFEKLKTKPANKICFDCGQKNPTWTSVPFGIYLCLDCSSNHRNLGVHISFVRSTNLDQWQ 72
Query: 72 PEQLKMMSFGGNSRAQVFFRQHGWN---GDGKVEAKYTSRAAELYKQLLSKEVAKSMSEE 128
+QL++M GGN A FF+Q+G + KY S AA YK L + A+ +
Sbjct: 73 WDQLRVMKVGGNESATKFFQQNGGTAALNSKDPKTKYQSNAATKYKDELKRRAARDAQDY 132
Query: 129 AALSAPPAASSQSAQGTNGLPD-----------VKTNEVPIEKT---------------- 161
A + G PD +K P+ +T
Sbjct: 133 PTEVIITDAIDDGSATPAGEPDDDFFSSWDKPAIKRPTPPVSRTGTPPVVGRTPSPFLNS 192
Query: 162 -----VEKTVEKPEKTES-SSSPRAYTAVSNNLKKP-----------IGAKKTGKSGGLG 204
+ +T +T + + P + S L+K +GAKKT K LG
Sbjct: 193 GNGKDIARTASPLSRTSTGENKPASRITTSAALRKTPASTGPRKANVLGAKKTTK---LG 249
Query: 205 ARKLT----------RKPSESL-------YEQKPEELPAPVSSSTITK--NNLPSGPPLT 245
A+K+T RK E Y+ EE PA +S + + P P
Sbjct: 250 AKKVTADIIDFDEAERKAKEEADRIAKLGYDPDAEEDPATKNSGSAAAIISPTPVAPSRG 309
Query: 246 SRFEYTEDVQSSELNSGGSNVT----GHVSVPKSSSSFFSDFGMDSGFQKKSGPSSSKVQ 301
S +T +E+ G + G V PK+++S S ++G GP ++
Sbjct: 310 SASSHTRQKSDAEVERLGMGMNRLGFGQVGGPKAAAS--SAPKRNAGGFGSVGPVRAQDV 367
Query: 302 IQESDEARKKFSNAKSISSSQFFGD---QNKANADAQATLSKFSGSSAISSADLFGDSSD 358
AR+KF K ISS +FFG ++A+ L F G++AISS FG D
Sbjct: 368 DDSERYAREKFGTQKGISSDEFFGKGAFDPSQQSEAKTRLQGFEGATAISSNAYFGRPED 427
Query: 359 -------NVDLAASDLINRISFQAQQDISSLKNIAGETGKKLTSLASSLM 401
+++ AA D + + A D+ +L +AGE +L S +
Sbjct: 428 EPEEEYGDLESAAKDFVRKFGITAGDDLENLTQMAGEVSTRLQGAIRSYL 477
>ref|XP_330928.1| hypothetical protein [Neurospora crassa] gi|28923588|gb|EAA32762.1|
hypothetical protein [Neurospora crassa]
Length = 496
Score = 176 bits (445), Expect = 2e-42
Identities = 150/474 (31%), Positives = 213/474 (44%), Gaps = 92/474 (19%)
Query: 12 VFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWT 71
+F KLK K NK CFDC KNPTW SV +GI+LC+DCSA HR+LGVHISFVRSTNLD W
Sbjct: 13 LFEKLKAKPANKICFDCGQKNPTWTSVPFGIYLCLDCSANHRNLGVHISFVRSTNLDQWQ 72
Query: 72 PEQLKMMSFGGNSRAQVFFRQHGWN---GDGKVEAKYTSRAAELYKQLLSKEVAKSM--- 125
+QL++M GGN A FF+Q+G + + KY S AA YK+ L K A+
Sbjct: 73 WDQLRIMKVGGNESATKFFQQNGGSAALNSKDPKTKYQSAAATKYKEELKKRAARDAREY 132
Query: 126 -SEEAALSAPPAASSQSAQG--------TNGLPDVKTNEVPIEKT--------------- 161
E A SS + G + P +K PI +T
Sbjct: 133 PEEVVITDGDDATSSNTPAGEPDDDFFSSWDKPAIKKPTPPISRTSTPPVIGRTASPLLG 192
Query: 162 ----VEKTVEKPEKTESSSSPRAYTA----VSNNLKKPI-GAKKTG---KSGG------- 202
+++T KT+S + A A S L+K G+ TG K GG
Sbjct: 193 NGKDIQRTSSPLSKTDSDAPTPAPAASRITTSAALRKTTPGSSTTGGPRKVGGGILGAKK 252
Query: 203 ----LGARKLT----------RKPSESL-------YEQKPEELPAPVSSSTITKNNL-PS 240
LG +K++ +K E Y+ + EE A + T T N + P+
Sbjct: 253 PAAKLGVKKISADLIDFDEAEKKAKEEAERIEKLGYDPEAEEEKAAKKAETKTNNIITPA 312
Query: 241 GPPLTSRFEYT--EDVQSSELNSGGSNVT----GHVSVPKSSSSFFSDFGMDSGFQKKSG 294
P++S + ++ ++E+ G V G V P + + G + + G
Sbjct: 313 AAPVSSSSSRSAAQEKSAAEVERLGMGVRKLGFGMVGKPGGAGAAAGAGGAAAAKKNAGG 372
Query: 295 -PSSSKVQIQESDE---ARKKFSNAKSISSSQFFGDQN---KANADAQATLSKFSGSSAI 347
S ++ E+DE AR KF+N K+ISS +FF N A+ +A L F G+ AI
Sbjct: 373 FGSVGPIKASEADEEQYARNKFANQKAISSDEFFNKGNYDPSVKAETKARLQGFEGAQAI 432
Query: 348 SSADLFGDSSD--------NVDLAASDLINRISFQAQQDISSLKNIAGETGKKL 393
SS FG D +++ AA D I + A D+ +L + GE +L
Sbjct: 433 SSNAYFGRPEDDAPAEDYGDLESAAKDFIRKFGITASDDLENLTQMVGEGAGRL 486
>emb|CAG86002.1| unnamed protein product [Debaryomyces hansenii CBS767]
gi|50418851|ref|XP_457946.1| unnamed protein product
[Debaryomyces hansenii]
Length = 461
Score = 173 bits (439), Expect = 8e-42
Identities = 145/467 (31%), Positives = 221/467 (47%), Gaps = 88/467 (18%)
Query: 3 SDSFTDKNA---VFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHI 59
SDS K VF KL+ + N+ CFDC+ KNPTW S+ +GI LC++CSA HR+LGVHI
Sbjct: 2 SDSMATKEEATKVFNKLRQQPANQVCFDCSNKNPTWTSIPFGILLCLECSAAHRNLGVHI 61
Query: 60 SFVRSTNLDSWTPEQLKMMSFGGNSRAQVFFRQHG-----WNGDG-KVEAKYTSRAAELY 113
SFV+S+NLDSW QL+ FGGN A+ F+ ++G N DG + AKYT+ A Y
Sbjct: 62 SFVKSSNLDSWQRIQLRHFKFGGNQVAKEFYTKNGGSKFLGNKDGIDINAKYTAPVALKY 121
Query: 114 KQLLSKEVAKSMSEEAALSAPPAASSQSAQGTNGLPDVKTNEVPIEKTVEKTVEKPEKTE 173
K+ L + K+ +EA P S + + GL +N V + + T
Sbjct: 122 KEKLKQ---KAQQDEA--KHPDEVSIDDLEESGGLLTDSSNNVSTDDFFSNWTKPINSTP 176
Query: 174 SSSSPRAYTAVSNN----LKKPIGAKKTG------------------------KSGGLGA 205
S S + T ++N +KKP+ + T +S L
Sbjct: 177 SPLSSKNITPNASNDDLSVKKPVTTRTTSRLVNNSSGANQPKGSILSSKGNGPRSSRLAN 236
Query: 206 RKLTRKPSESLYE-------QKPEELPA----PVSSSTI-TKNNLPS-------GPPLTS 246
+++ ++ E +E Q+ EE A P + ST T N P+ G PL S
Sbjct: 237 KRINKEAEEIDFEEIERKAKQEAEEAKALGYNPTTESTSDTSNAKPAKSEPKKPGLPLGS 296
Query: 247 RFEYTEDVQSSELNSGGSNVTGHVSVPKSSSSFFS-DFGMDSGFQKKSGPSSSKVQ-IQE 304
R + TE S +L++ +V + +++ F FGM +G + S + ++
Sbjct: 297 RKQDTEG--SEDLSA------KNVPIKEATQKFQKLGFGMTAGSNEVVNEKSKNYKPVKY 348
Query: 305 SDEARKKFSNAKSISSSQFFG-----DQNKANADAQATLSKFSGSSAISSADLFGDSSD- 358
+ E KF K ISS ++FG D+N ANA+A+ L F+G+ +ISS+ FG+
Sbjct: 349 TGEVSNKFGTQKGISSDEYFGRGPRFDEN-ANAEARDKLQSFNGAQSISSSSYFGEDESA 407
Query: 359 ----------NVDLAASDLINRISFQAQQDISSLKNIAGETGKKLTS 395
+++ +A + +R S A QD+ LK+ E KL S
Sbjct: 408 QNNTQGAGLTDIESSAREFASRFSGNANQDLDVLKDAFEEGANKLGS 454
>gb|EAA55821.1| hypothetical protein MG01472.4 [Magnaporthe grisea 70-15]
gi|39951659|ref|XP_363546.1| hypothetical protein
MG01472.4 [Magnaporthe grisea 70-15]
Length = 490
Score = 167 bits (424), Expect = 4e-40
Identities = 148/483 (30%), Positives = 207/483 (42%), Gaps = 100/483 (20%)
Query: 12 VFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWT 71
+F KLKTK NK CFDC KNPTW SV +GI+LC+DCS+ HR+LGVHISFVRSTNLD W
Sbjct: 13 IFEKLKTKQANKICFDCGQKNPTWTSVPFGIYLCLDCSSNHRNLGVHISFVRSTNLDQWQ 72
Query: 72 PEQLKMMSFGGNSRAQVFFRQHGWN---GDGKVEAKYTSRAAELYKQLLSKEV---AKSM 125
+QL++M GGN A FF+Q+G + + KY S A YK+ L K AK
Sbjct: 73 WDQLRVMKVGGNESATKFFQQNGGSAALNSKDPKTKYHSAVATKYKEELKKRAARDAKEY 132
Query: 126 SEEAALSAPPAASSQSAQGTN---GLPD-----------VKTNEVPIEKTVEKTV----- 166
EE ++ + + GTN G PD +K P+ +T V
Sbjct: 133 PEEVVIT----DGTDAGDGTNTPAGEPDDDFFSSWDKPAIKKPTPPVSRTATPPVVGRTP 188
Query: 167 -----------EKPEKTESSSSPRAYTAV---------SNNLKKPIGAKKTGKSGGLGAR 206
+ +T S + A +AV S+ LKK GA K+ LGA+
Sbjct: 189 SPFLTAGGANGKDIARTPSPLAKSADSAVKPAASRITHSSALKKTTGAGPK-KANVLGAK 247
Query: 207 KLT----RKPSESLYEQKPEELPAPVSSSTITKNNLPSGPPLTSRFEYTEDVQSSELNS- 261
K T +K + + + E A + I K T+ + V++ +
Sbjct: 248 KTTKLGVKKVNAEVIDFDEAEKKAKEEAERIEKLGYDPDDIDTTAAKKVGGVKAESSGAG 307
Query: 262 -----------GGSNVTGHVSVPKSSSSFFSDFGMDSG---------------FQKKS-- 293
GG +GH + S+S GM G Q K
Sbjct: 308 ILAPTPVSPARGGYGASGHSR--EKSASEMERLGMGMGRLGFGQVGGNKPAAAAQPKKGG 365
Query: 294 -----GPSSSKVQIQESDEARKKFSNAKSISSSQFFGDQN---KANADAQATLSKFSGSS 345
GP + + E AR KF K+ISS +FFG + A A+A+ L F G+S
Sbjct: 366 GFGSVGPIKATPEADEEKYARSKFGAQKAISSDEFFGKGSYDPNAQAEAKTRLQGFEGAS 425
Query: 346 AISSADLFG-------DSSDNVDLAASDLINRISFQAQQDISSLKNIAGETGKKLTSLAS 398
AISS FG + +++ AA D I + D+ +L + G+ KL
Sbjct: 426 AISSNAYFGRPEEEEVEEYGDLETAAKDFIRKFGLTRGDDLENLTAVLGDGATKLQGAIR 485
Query: 399 SLM 401
S +
Sbjct: 486 SYL 488
>gb|AAH41750.1| Arfgap3-prov protein [Xenopus laevis]
Length = 517
Score = 158 bits (400), Expect = 3e-37
Identities = 115/325 (35%), Positives = 163/325 (49%), Gaps = 37/325 (11%)
Query: 1 MASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 60
MA D ++FR+L++ NK CFDC AKNP+WAS+TYG+FLCIDCS HRSLGVH+S
Sbjct: 1 MAEPHKQDIASIFRRLRSVPTNKVCFDCGAKNPSWASITYGVFLCIDCSGSHRSLGVHLS 60
Query: 61 FVRSTNLDS-WTPEQLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQLLSK 119
F+RST LDS W+ QL+ M GGNS A +FFRQHG + AKY SRA++LY++ +
Sbjct: 61 FIRSTELDSNWSWFQLRCMQVGGNSNATIFFRQHGCS-TNDTNAKYNSRASQLYREKIKS 119
Query: 120 EVAKSMSEEA------ALSAPP------------AASSQSAQGTNGLPDVKTNEVPIEKT 161
++ + APP A+ S+ T + E P+ +
Sbjct: 120 LATQATRKHGTELWIDVYGAPPLSPQQQEEEDFFASHSEQVNNTAQGAAPQVEETPLTIS 179
Query: 162 VEKTVEK---PEKTESSSSPRAYTAVSNNLKKPIGAKKTG---KSGGLGARKLTRKPSES 215
++ E P S SP+A + LKK A K G K GGLGA+K++ K S S
Sbjct: 180 NKEAGEPDVGPSVDYLSMSPKASLESVSVLKKKSTAAKKGLGAKKGGLGAQKVSSK-SFS 238
Query: 216 LYEQKPEELPAPVSSSTITKNNLPSGPPLTS-RFEYTE-DVQSSELNSGGSNVTGHVSVP 273
E+ + + +T P +TS R Y + ++Q + + N +P
Sbjct: 239 EMEKHAQAVDKQNEQEAVTTTKKEEEPVVTSLRLAYKQLEIQKQKDDEKLKN------LP 292
Query: 274 KSSSSFFSDFGMDSGFQKKSGPSSS 298
++ GM GF +SG S S
Sbjct: 293 PKKAAEAERLGM--GFGTRSGISHS 315
Score = 71.2 bits (173), Expect = 5e-11
Identities = 40/106 (37%), Positives = 65/106 (60%), Gaps = 3/106 (2%)
Query: 302 IQESDEARKKFSNAKSISSSQFFGDQNKANADAQATLSKFSGSSAISSADLFGDSSDNVD 361
+ +D+A+KKF NAK+ISS FFG Q+ A+ + ++ L + SG+S+ISSADLF + D
Sbjct: 412 VPTTDDAQKKFGNAKAISSDMFFGKQDNADYETRSRLERLSGNSSISSADLFDEHKK--D 469
Query: 362 LAASDLINRISFQAQQDISSLKNIAGETGKKLTSLASSLMTDLQDR 407
A + + R+ + D+ + K +L+ LA+ +MT +QDR
Sbjct: 470 PAGNYNLTRV-LPSAPDMGNFKQGVKSVAGRLSVLANGVMTTIQDR 514
>ref|NP_001016210.1| hypothetical protein LOC548964 [Xenopus tropicalis]
Length = 535
Score = 157 bits (398), Expect = 4e-37
Identities = 155/541 (28%), Positives = 228/541 (41%), Gaps = 143/541 (26%)
Query: 1 MASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 60
MA D A+FR+L++ NK CFDC AKNP+WAS+TYG+FLCIDCS HRSLGVH+S
Sbjct: 1 MAEPHKQDIAAIFRRLRSIPSNKVCFDCGAKNPSWASITYGVFLCIDCSGTHRSLGVHLS 60
Query: 61 FVRSTNLDS-WTPEQLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQLLSK 119
F+RST LDS W+ QL+ M GGN+ A V+FRQHG AKY SRA++LY++ +
Sbjct: 61 FIRSTELDSNWSWFQLRCMQVGGNANATVYFRQHGC-ATNDTNAKYNSRASQLYRERVKS 119
Query: 120 EVAKSMSEEA------ALSAPP-----------AASSQSAQGTNGLPDVKTNEVPIEKTV 162
+ ++ A APP AA S+ T+ P + + PI +
Sbjct: 120 QATQATRRHGTDLWIDACGAPPLSPQHQEEDFFAAHSEQDNNTSQAPP-QAEDTPINVSS 178
Query: 163 E----KTVEKPEKTES----SSSPRAYTAVSNNL----------------KKPIGAKK-- 196
E +PE S S SP+A A + NL KKP AKK
Sbjct: 179 ESGPANNEGEPEVGPSVDYLSVSPKAALADAENLISAGNAPARESVSVLKKKPGAAKKGL 238
Query: 197 TGKSGGLGARKLTRKPSESLYEQKPEELPAPVSSSTITKNNLPSGPPLTS-RFEYTE-DV 254
K GGLGA+K++ K S S E++ + + T P + S R Y + ++
Sbjct: 239 GAKKGGLGAQKVSGK-SFSEMERQAQAVDKQKEQEAATVTKRAEEPVVASLRLAYKDLEI 297
Query: 255 QSSELNSGGSNVTGHVSVPKSSSS---FFSDFGMDSGFQ----------KKSGPSSSKV- 300
Q + N+ PK ++ FG SG ++ P S K
Sbjct: 298 QKQKDEEKLKNLP-----PKKAAEAERLGMGFGARSGISHSVLSEMTTIEQEAPKSVKAK 352
Query: 301 QIQESDEARKKFSNA----------KSISSSQFFGDQ--------NKANADAQATLS--- 339
++ + +EA + +A + ++SS D+ + AD + L+
Sbjct: 353 KLYQEEEAEDFYFSAPRSRYREDPPEPVTSSFLSWDERSDDYWKMDSPGADTEVALTSKT 412
Query: 340 --------------------------KFSGSSAISSADLFGDSSDNVDLAASDLINRIS- 372
KF + AISS D+F D+ D A ++R+S
Sbjct: 413 TSYDDRPATRRKHDLEPAPSTDDAQKKFGNAKAISS-DMFFGKQDSADYEARSRLDRLSG 471
Query: 373 --------------------------FQAQQDISSLKNIAGETGKKLTSLASSLMTDLQD 406
A D+++ K +L+ LA+ +MT +QD
Sbjct: 472 NSSISSADLFDEQKKDPTGNYTLTRVLPAAPDMANFKQGVKSVAGRLSVLANGVMTTIQD 531
Query: 407 R 407
R
Sbjct: 532 R 532
>gb|EAK98866.1| potential ARF GAP [Candida albicans SC5314]
gi|46439448|gb|EAK98766.1| potential ARF GAP [Candida
albicans SC5314]
Length = 451
Score = 157 bits (396), Expect = 7e-37
Identities = 121/443 (27%), Positives = 201/443 (45%), Gaps = 63/443 (14%)
Query: 8 DKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNL 67
+ + +F +LK N+ CFDC+ KNPTW S+ +GIFLC+ CSAVHR+LGVHISFV+S+NL
Sbjct: 10 EASKIFDRLKKDPANQVCFDCSNKNPTWTSIPFGIFLCLQCSAVHRNLGVHISFVKSSNL 69
Query: 68 DSWTPEQLKMMSFGGNSRAQVFFRQHGW-----NGDG-KVEAKYTSRAAELYKQLLSKEV 121
DSW QL+ FGGN +A+ FF ++G N +G AKYTS A YK+ L ++
Sbjct: 70 DSWQRIQLRNFKFGGNQQAKDFFLKNGGSQFVNNKNGVDATAKYTSPCANKYKEKLKQKA 129
Query: 122 AK------------SMSEEAALSAPPAASSQSAQGTNGLPDVKTN---------EVPIEK 160
A+ +++ +LS P+ S+ P +N P
Sbjct: 130 AQDAAKHPDIVTLDDVTDVMSLSDSPSESTDDFFSNWNKPVNNSNTASPLSSRAATPNAS 189
Query: 161 TVEKTVEKPEKTESSSSPRAYTAVSNNLKKPIGAKKTG-KSGGLGARKLTRKPS------ 213
T + T +KP +++S R + + K + K G ++ L A+++ +
Sbjct: 190 TDDLTKKKPVVRTTTTSARLKSNNTAVKKSILSGKGNGPRTSRLAAKRINKTEDDIDFDE 249
Query: 214 -ESLYEQKPEELPAPVSSSTITKNNLPSGPPLTSRFEYTEDVQSSELNSGGSNVTGHVSV 272
E +Q+ EE T T P ++ ++ + +L +
Sbjct: 250 IEKKAKQEAEEAKKLGYKPTETVEPTVKKQPTSTSLSLKKENEEVKLTP--------API 301
Query: 273 PKSSSSFFS-DFGMDSGFQKKSGPSSSKVQIQESDEARKKFSNAKSISSSQFFGD----Q 327
+++ F FGM G + +++ + E K+ K ISS +FFG
Sbjct: 302 QETTQQFQKLGFGMTQGDNIVGSSTKKYKEVKYTGEVSNKYGTQKGISSDEFFGRGPRFD 361
Query: 328 NKANADAQATLSKFSGSSAISSADLFGDSS---------------DNVDLAASDLINRIS 372
+A +A+ L F+G+ +ISS+ FG+ +++ +A + ++ S
Sbjct: 362 EQAKTEARTKLQAFNGAQSISSSSYFGEEEGGARGGRSNSGAGGLGDLEASAREFASKFS 421
Query: 373 FQAQQDISSLKNIAGETGKKLTS 395
A QD+ LK+ + KL S
Sbjct: 422 GNANQDLEVLKDALEDGATKLGS 444
>emb|CAG02441.1| unnamed protein product [Tetraodon nigroviridis]
Length = 1054
Score = 155 bits (392), Expect = 2e-36
Identities = 94/237 (39%), Positives = 136/237 (56%), Gaps = 27/237 (11%)
Query: 1 MASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 60
M+ S D +A+F++L++ NK CFDC+ KNP+WAS+TYG+FLCIDCS +HRSLGVH+S
Sbjct: 1 MSEPSKQDISAIFKRLRSIPTNKICFDCSVKNPSWASITYGVFLCIDCSGIHRSLGVHLS 60
Query: 61 FVRSTNLD-SWTPEQLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQ---L 116
F+RST LD +W+ QL+ M GGN+ A FF QHG AKY SRAA+LY++
Sbjct: 61 FIRSTELDFNWSWFQLRCMQVGGNTNAIAFFNQHGCT-TSAANAKYNSRAAQLYREKMRT 119
Query: 117 LSKEVAKSMSEEAALSAPPAAS------------SQSAQGTNGLPDVKTNEVPIEKTVEK 164
L+ + + + L + S S ++ N D+ + + T EK
Sbjct: 120 LATQATRRHGTDLWLDSQGPLSPTTPEPKQVDFFSLHSEAENLNADMSLSIPQMAATAEK 179
Query: 165 TVEKPEKTES-------SSSPRAYTAVSNNL-KKPIGAKKT--GKSGGLGARKLTRK 211
+K KTE ++SP+A + + + KKP KKT K GGLGA+K++R+
Sbjct: 180 EDDKNGKTEEGPSVDLLATSPKANPELPSLIKKKPAATKKTLASKKGGLGAQKVSRE 236
Score = 58.9 bits (141), Expect = 3e-07
Identities = 35/104 (33%), Positives = 60/104 (57%), Gaps = 3/104 (2%)
Query: 302 IQESDEARKKFSNAKSISSSQFFGDQNKANADAQATLSKFSGSSAISSADLFGDSSDNVD 361
+ + EA+KKF + K+ISS +FG Q+K+ + ++ L + +GS++ISSADLF D
Sbjct: 412 VTNTGEAQKKFGDMKAISSDMYFGKQDKSEYETRSRLERLAGSASISSADLFEDPKKQTG 471
Query: 362 LAASDLINRISFQAQQDISSLKNIAGETGKKLTSLASSLMTDLQ 405
++ L N + + D+S LK KL+ +AS ++ +Q
Sbjct: 472 -SSYRLTNML--PSAPDMSQLKLGVRSVAGKLSVMASGVVNTIQ 512
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.307 0.123 0.332
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 634,999,299
Number of Sequences: 2540612
Number of extensions: 25734223
Number of successful extensions: 101690
Number of sequences better than 10.0: 1754
Number of HSP's better than 10.0 without gapping: 708
Number of HSP's successfully gapped in prelim test: 1090
Number of HSP's that attempted gapping in prelim test: 93369
Number of HSP's gapped (non-prelim): 4279
length of query: 409
length of database: 863,360,394
effective HSP length: 130
effective length of query: 279
effective length of database: 533,080,834
effective search space: 148729552686
effective search space used: 148729552686
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.6 bits)
S2: 76 (33.9 bits)
Medicago: description of AC137670.5


