

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
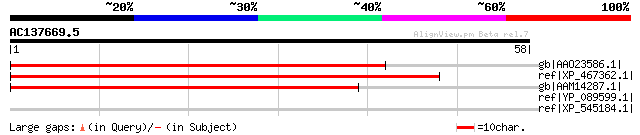
Query= AC137669.5 + phase: 0
(58 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
gb|AAO23586.1| At3g50910/F18B3_190 [Arabidopsis thaliana] gi|145... 62 3e-09
ref|XP_467362.1| unknown protein [Oryza sativa (japonica cultiva... 57 1e-07
gb|AAM14287.1| unknown protein [Arabidopsis thaliana] gi|1581031... 55 5e-07
ref|YP_089599.1| NADH dehydrogenase subunit 4 [Rhinatrema bivitt... 31 7.6
ref|XP_545184.1| PREDICTED: similar to a disintegrin-like and me... 31 10.0
>gb|AAO23586.1| At3g50910/F18B3_190 [Arabidopsis thaliana]
gi|14517540|gb|AAK62660.1| AT3g50910/F18B3_190
[Arabidopsis thaliana] gi|18409208|ref|NP_566941.1|
expressed protein [Arabidopsis thaliana]
Length = 447
Score = 62.4 bits (150), Expect = 3e-09
Identities = 25/42 (59%), Positives = 34/42 (80%)
Query: 1 MARRERQRKSKKQKWIWGSLTTVIALGTAAVAWSYLPAGGES 42
+ARRERQ++ K+Q+WIWGS+ I LG+AA+AWSY+PA S
Sbjct: 398 VARRERQKRKKRQRWIWGSIAATITLGSAALAWSYIPAAKPS 439
>ref|XP_467362.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|41053140|dbj|BAD08083.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 456
Score = 57.0 bits (136), Expect = 1e-07
Identities = 23/48 (47%), Positives = 33/48 (67%)
Query: 1 MARRERQRKSKKQKWIWGSLTTVIALGTAAVAWSYLPAGGESYSAEDH 48
M+R++R + K+QKW WGS+ + LGTAA+AWSYLPA S + +
Sbjct: 405 MSRQQRNIRKKRQKWFWGSVGLAVTLGTAAIAWSYLPAAQPQASQDSN 452
>gb|AAM14287.1| unknown protein [Arabidopsis thaliana] gi|15810313|gb|AAL07044.1|
unknown protein [Arabidopsis thaliana]
gi|18425078|ref|NP_569035.1| expressed protein
[Arabidopsis thaliana]
Length = 444
Score = 55.1 bits (131), Expect = 5e-07
Identities = 20/39 (51%), Positives = 30/39 (76%)
Query: 1 MARRERQRKSKKQKWIWGSLTTVIALGTAAVAWSYLPAG 39
+ARR+ Q++ ++Q+WIWGS+ I LG+ +AWSYLP G
Sbjct: 390 VARRDGQKRKRRQRWIWGSIAATITLGSGVLAWSYLPPG 428
>ref|YP_089599.1| NADH dehydrogenase subunit 4 [Rhinatrema bivittatum]
gi|42406130|gb|AAS13727.1| NADH dehydrogenase subunit 4
[Rhinatrema bivittatum]
Length = 458
Score = 31.2 bits (69), Expect = 7.6
Identities = 12/28 (42%), Positives = 19/28 (67%), Gaps = 1/28 (3%)
Query: 12 KQKWIWGSLTT-VIALGTAAVAWSYLPA 38
KQKW+W +LTT + + A++ W Y P+
Sbjct: 19 KQKWLWPALTTHSLIIAAASLVWFYSPS 46
>ref|XP_545184.1| PREDICTED: similar to a disintegrin-like and metalloprotease
(reprolysin type) with thrombospondin type 1 motif, 16
preproprotein [Canis familiaris]
Length = 2887
Score = 30.8 bits (68), Expect = 10.0
Identities = 14/35 (40%), Positives = 19/35 (54%)
Query: 5 ERQRKSKKQKWIWGSLTTVIALGTAAVAWSYLPAG 39
ER K KK +W+ + + V A G A AW +P G
Sbjct: 2364 ERCHKHKKLQWLLSAWSQVCAPGPAGTAWPAVPRG 2398
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.313 0.126 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 109,827,663
Number of Sequences: 2540612
Number of extensions: 3281655
Number of successful extensions: 7265
Number of sequences better than 10.0: 5
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 7261
Number of HSP's gapped (non-prelim): 5
length of query: 58
length of database: 863,360,394
effective HSP length: 34
effective length of query: 24
effective length of database: 776,979,586
effective search space: 18647510064
effective search space used: 18647510064
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 68 (30.8 bits)
Medicago: description of AC137669.5


