

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137553.6 - phase: 0
(456 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
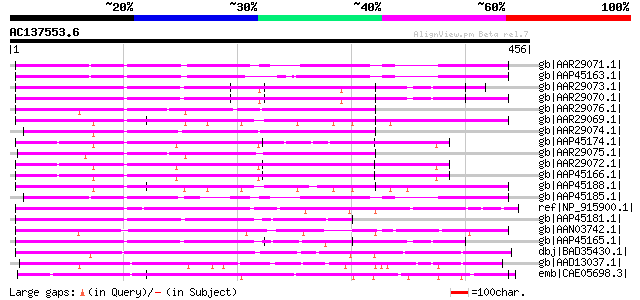
Score E
Sequences producing significant alignments: (bits) Value
gb|AAR29071.1| blight resistance protein RGA3 [Solanum bulbocast... 172 2e-41
gb|AAP45163.1| putative disease resistant protein RGA1 [Solanum ... 172 3e-41
gb|AAR29073.1| blight resistance protein B149 [Solanum bulbocast... 160 8e-38
gb|AAR29070.1| blight resistance protein RGA1 [Solanum bulbocast... 160 8e-38
gb|AAR29076.1| blight resistance protein T118 [Solanum tarijense] 159 2e-37
gb|AAR29069.1| blight resistance protein RPI [Solanum bulbocasta... 157 9e-37
gb|AAR29074.1| blight resistance protein SH10 [Solanum tuberosum] 156 1e-36
gb|AAP45174.1| putative disease resistant protein rga4 [Solanum ... 153 1e-35
gb|AAR29075.1| blight resistance protein SH20 [Solanum tuberosum] 152 2e-35
gb|AAR29072.1| blight resistance protein RGA4 [Solanum bulbocast... 151 4e-35
gb|AAP45166.1| putative disease resistant protein RGA4 [Solanum ... 151 4e-35
gb|AAP45188.1| putative disease resistant protein rga2 [Solanum ... 150 1e-34
gb|AAP45185.1| putative disease resistant protein rga1 [Solanum ... 142 2e-32
ref|NP_915900.1| putative NBS-LRR type resistance protein [Oryza... 141 4e-32
gb|AAP45181.1| putative disease resistant protein rga3 [Solanum ... 138 3e-31
gb|AAN03742.1| NBS-LRR-like protein [Oryza sativa (japonica cult... 138 3e-31
gb|AAP45165.1| putative disease resistant protein RGA3 [Solanum ... 136 1e-30
dbj|BAD35430.1| putative disease resistance protein [Oryza sativ... 130 1e-28
gb|AAD13037.1| NBS-LRR-like protein cD8 [Phaseolus vulgaris] 127 9e-28
emb|CAE05698.3| OSJNBa0083D01.14 [Oryza sativa (japonica cultiva... 124 6e-27
>gb|AAR29071.1| blight resistance protein RGA3 [Solanum bulbocastanum]
Length = 979
Score = 172 bits (436), Expect = 2e-41
Identities = 127/435 (29%), Positives = 201/435 (46%), Gaps = 86/435 (19%)
Query: 6 FTVRPNAGFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSM-RLNRLHLAWDRNTN 64
F + G+ L +L NL ++ I L ++ D +E NLS+ L+ L L+WD +
Sbjct: 627 FVIGKRKGYQLGELKNLNLYGSISITKLDRVKKDSDAKEANLSAKANLHSLCLSWDLDGK 686
Query: 65 STNSAEEVLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQLPP 124
+E VL AL+PH +L ++G+ G+ +P+WM S+L +V +++ C NCS LPP
Sbjct: 687 HRYDSE-VLEALKPHSNLKYLEINGFGGIRLPDWMNQ-SVLKNVVSIRIRGCENCSCLPP 744
Query: 125 LGKLPFLNTLYL-SQMTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNLERVLRIEGVEM 183
G+LP L +L L + +V+Y++D+ + FPSL ++ ++D NL+ +L+ EG +
Sbjct: 745 FGELPCLESLELHTGSADVEYVEDNVHP----GRFPSLRKLVIWDFSNLKGLLKKEGEKQ 800
Query: 184 LSQLSKLSIQSIPQFELPSLPSVKEVYVGGETEEDIDHEASFLRDIAGKMPNLKELMIDA 243
L +++ P F +P+L SVK + V I +A+ LR I
Sbjct: 801 FPVLEEMTFYWCPMFVIPTLSSVKTLKV-------IATDATVLRSI-------------- 839
Query: 244 FHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLNSLPQSV 303
S+LR+L L I + + S+P +F L +L+ L+ +L LP S+
Sbjct: 840 -----------SNLRALTSLDISNNVEATSLPEEMFKSLANLKYLNISFFRNLKELPTSL 888
Query: 304 TTLTSLQRLIIHYCPELILPANMNMLNSLREVSIMGGDRRRGIYNGLEDIPLLQNLSLRD 363
+L +L+ L +C N L SL E + G
Sbjct: 889 ASLNALKSLKFEFC---------NALESLPEEGVKG------------------------ 915
Query: 364 FPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCIDRCPRLVNRLARRT 423
SL EL +S L LP+ L L L I +CP + R R
Sbjct: 916 -------------LTSLTELSVSNCMMLKCLPEGLQHLTALTTLTITQCPIVFKRCERGI 962
Query: 424 GEDWYKIAHVPILSL 438
GEDW+KIAH+P L+L
Sbjct: 963 GEDWHKIAHIPYLTL 977
Score = 41.2 bits (95), Expect = 0.068
Identities = 30/116 (25%), Positives = 58/116 (49%), Gaps = 2/116 (1%)
Query: 247 LTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLNSLPQSVTTL 306
L LP+ + L L L + ++ ++P + L +L+ L C SL+ LP+ + L
Sbjct: 537 LNQLPSSIGDLVHLRYLDLSGNFRIRNLPKRLCR-LQNLQTLDLHYCDSLSCLPKQTSKL 595
Query: 307 TSLQRLIIHYCPELILPANMNMLNSLREVSIMGGDRRRGIYNG-LEDIPLLQNLSL 361
SL+ L++ C P + +L L+ +S +R+G G L+++ L ++S+
Sbjct: 596 GSLRNLLLDGCSLTSTPPRIGLLTCLKSLSCFVIGKRKGYQLGELKNLNLYGSISI 651
Score = 35.4 bits (80), Expect = 3.7
Identities = 22/66 (33%), Positives = 38/66 (57%), Gaps = 1/66 (1%)
Query: 350 LEDIPLLQNLSLRDFPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCI 409
L+ L+ L+LR+ +L LP +GD + L+ L++S ++ +LP +L+NLQ L +
Sbjct: 521 LQKFVSLRVLNLRN-SNLNQLPSSIGDLVHLRYLDLSGNFRIRNLPKRLCRLQNLQTLDL 579
Query: 410 DRCPRL 415
C L
Sbjct: 580 HYCDSL 585
>gb|AAP45163.1| putative disease resistant protein RGA1 [Solanum bulbocastanum]
gi|46576967|sp|Q7XA42|RGA1_SOLBU Putative disease
resistance protein RGA1 (RGA3-blb)
Length = 1025
Score = 172 bits (435), Expect = 3e-41
Identities = 128/435 (29%), Positives = 201/435 (45%), Gaps = 86/435 (19%)
Query: 6 FTVRPNAGFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSM-RLNRLHLAWDRNTN 64
F + G L +L NL ++ I L ++ D +E NLS+ L+ L L+WD +
Sbjct: 673 FVIGKRKGHQLGELKNLNLYGSISITKLDRVKKDTDAKEANLSAKANLHSLCLSWDLDGK 732
Query: 65 STNSAEEVLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQLPP 124
+E VL AL+PH +L ++G+ G+ +P+WM S+L +V +++ C NCS LPP
Sbjct: 733 HRYDSE-VLEALKPHSNLKYLEINGFGGIRLPDWMNQ-SVLKNVVSIRIRGCENCSCLPP 790
Query: 125 LGKLPFLNTLYL-SQMTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNLERVLRIEGVEM 183
G+LP L +L L + +V+Y++D+ + FPSL ++ ++D NL+ +L++EG +
Sbjct: 791 FGELPCLESLELHTGSADVEYVEDNVHP----GRFPSLRKLVIWDFSNLKGLLKMEGEKQ 846
Query: 184 LSQLSKLSIQSIPQFELPSLPSVKEVYVGGETEEDIDHEASFLRDIAGKMPNLKELMIDA 243
L +++ P F +P+L SVK LK ++ DA
Sbjct: 847 FPVLEEMTFYWCPMFVIPTLSSVK---------------------------TLKVIVTDA 879
Query: 244 FHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLNSLPQSV 303
TVL +S+LR+L L I D + S+P +F L +L+ L +L LP S+
Sbjct: 880 ----TVL-RSISNLRALTSLDISDNVEATSLPEEMFKSLANLKYLKISFFRNLKELPTSL 934
Query: 304 TTLTSLQRLIIHYCPELILPANMNMLNSLREVSIMGGDRRRGIYNGLEDIPLLQNLSLRD 363
+L +L+ L +C + L SL E + G
Sbjct: 935 ASLNALKSLKFEFC---------DALESLPEEGVKG------------------------ 961
Query: 364 FPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCIDRCPRLVNRLARRT 423
SL EL +S L LP+ L L L I +CP + R R
Sbjct: 962 -------------LTSLTELSVSNCMMLKCLPEGLQHLTALTTLTITQCPIVFKRCERGI 1008
Query: 424 GEDWYKIAHVPILSL 438
GEDW+KIAH+P L+L
Sbjct: 1009 GEDWHKIAHIPYLTL 1023
Score = 42.0 bits (97), Expect = 0.040
Identities = 40/153 (26%), Positives = 70/153 (45%), Gaps = 7/153 (4%)
Query: 247 LTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLNSLPQSVTTL 306
L LP+ + L L L + ++ ++P + L +L+ L C SL+ LP+ + L
Sbjct: 583 LNQLPSSIGDLVHLRYLDLSGNFRIRNLPKRLCK-LQNLQTLDLHYCDSLSCLPKQTSKL 641
Query: 307 TSLQRLIIHYCPELILPANMNMLNSLREVSIMGGDRRRGIYNGLEDIPLLQNLSLRDFPS 366
SL+ L++ C P + +L L+ +S +R+G + L+NL+L S
Sbjct: 642 GSLRNLLLDGCSLTSTPPRIGLLTCLKSLSCFVIGKRKG-----HQLGELKNLNLYGSIS 696
Query: 367 LRSLPDWLGDTLSLQELEISKFPKLTSLPDNFD 399
+ L DT +E +S L SL ++D
Sbjct: 697 ITKLDRVKKDT-DAKEANLSAKANLHSLCLSWD 728
Score = 35.4 bits (80), Expect = 3.7
Identities = 22/66 (33%), Positives = 38/66 (57%), Gaps = 1/66 (1%)
Query: 350 LEDIPLLQNLSLRDFPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCI 409
L+ L+ L+LR+ +L LP +GD + L+ L++S ++ +LP +L+NLQ L +
Sbjct: 567 LQKFVSLRVLNLRN-SNLNQLPSSIGDLVHLRYLDLSGNFRIRNLPKRLCKLQNLQTLDL 625
Query: 410 DRCPRL 415
C L
Sbjct: 626 HYCDSL 631
>gb|AAR29073.1| blight resistance protein B149 [Solanum bulbocastanum]
Length = 971
Score = 160 bits (405), Expect = 8e-38
Identities = 109/336 (32%), Positives = 167/336 (49%), Gaps = 23/336 (6%)
Query: 6 FTVRPNAGFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSM-RLNRLHLAWDRNTN 64
F V G+ L +L NL + I L+ ++N + +E NLS+ L+ L ++WDR
Sbjct: 636 FVVGERKGYQLGELRNLNLRGAISITHLERVKNDMEAKEANLSAKANLHSLSMSWDRPNR 695
Query: 65 STNSAEEVLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQLPP 124
+ +VL AL+PH +L + + G +P+WM S+L +V + + C NCS LPP
Sbjct: 696 YESEEVKVLEALKPHPNLKYLEIIDFCGFCLPDWMNH-SVLKNVVSILISGCENCSCLPP 754
Query: 125 LGKLPFLNTLYLSQ-MTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNLERVLRIEGVEM 183
G+LP L +L L V+Y++DS + T FPSL ++ + NL+ + R++G E
Sbjct: 755 FGELPCLESLELQDGSVEVEYVEDSGF--LTRRRFPSLRKLHIGGFCNLKGLQRMKGAEQ 812
Query: 184 LSQLSKLSIQSIPQFELPSLPSVKEVYVGGETEED-----------------IDHEA-SF 225
L ++ I P F P+L SVK++ + GE + +H S
Sbjct: 813 FPVLEEMKISDCPMFVFPTLSSVKKLEIWGEADAGGLSSISNLSTLTSLKIFSNHTVTSL 872
Query: 226 LRDIAGKMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISL 285
L ++ + NL L + L LP L+SL +L+ L I C LES+P GL SL
Sbjct: 873 LEEMFKNLENLIYLSVSFLENLKELPTSLASLNNLKCLDIRYCYALESLPEEGLEGLSSL 932
Query: 286 RILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPELI 321
L C+ L LP+ + LT+L L I CP+LI
Sbjct: 933 TELFVEHCNMLKCLPEGLQHLTTLTSLKIRGCPQLI 968
Score = 53.1 bits (126), Expect = 2e-05
Identities = 63/229 (27%), Positives = 104/229 (44%), Gaps = 15/229 (6%)
Query: 195 IPQF-ELPSLPSVKEVYVGGETEEDIDHEASFLRDIAGKMPNLKELMIDAFHQLTVLPNE 253
+P F ELP L S++ G E + ++ FL + P+L++L I F L
Sbjct: 752 LPPFGELPCLESLE--LQDGSVEVEYVEDSGFLT--RRRFPSLRKLHIGGFCNL----KG 803
Query: 254 LSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSF---VICHSLNSLPQSVTTLTSLQ 310
L ++ E+ +++ K+ P VF L S++ L L+S+ +++TLTSL+
Sbjct: 804 LQRMKGAEQFPVLEEMKISDCPMFVFPTLSSVKKLEIWGEADAGGLSSI-SNLSTLTSLK 862
Query: 311 RLIIHYCPELILPANMNMLNSLREVSIMGGDRRRGIYNGLEDIPLLQNLSLRDFPSLRSL 370
H L+ N+ N L +S+ + + + L + L+ L +R +L SL
Sbjct: 863 IFSNHTVTSLLEEMFKNLEN-LIYLSVSFLENLKELPTSLASLNNLKCLDIRYCYALESL 921
Query: 371 PD-WLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCIDRCPRLVNR 418
P+ L SL EL + L LP+ L L L I CP+L+ R
Sbjct: 922 PEEGLEGLSSLTELFVEHCNMLKCLPEGLQHLTTLTSLKIRGCPQLIKR 970
Score = 52.8 bits (125), Expect = 2e-05
Identities = 48/176 (27%), Positives = 85/176 (48%), Gaps = 14/176 (7%)
Query: 225 FLRDIAGKMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLIS 284
F R ++ ++ NL F QL +L LR L+ + NK+ S+P + L +
Sbjct: 531 FKRFVSLRVLNLSN---SEFEQLPSSVGDLVHLRYLD----LSGNKICSLPKRLCK-LRN 582
Query: 285 LRILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPELILPANMNMLNSLREVSIMGGDRRR 344
L+ L C SL+ LP+ + L SL+ L++ +CP +P + +L L+ + R+
Sbjct: 583 LQTLDLYNCQSLSCLPKQTSKLCSLRNLVLDHCPLTSMPPRIGLLTCLKTLGYFVVGERK 642
Query: 345 GIYNGLEDIPLLQNLSLRDFPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQ 400
G G L+NL+LR S+ L + + + + +E +S L SL ++D+
Sbjct: 643 GYQLG-----ELRNLNLRGAISITHL-ERVKNDMEAKEANLSAKANLHSLSMSWDR 692
>gb|AAR29070.1| blight resistance protein RGA1 [Solanum bulbocastanum]
Length = 992
Score = 160 bits (405), Expect = 8e-38
Identities = 109/336 (32%), Positives = 167/336 (49%), Gaps = 23/336 (6%)
Query: 6 FTVRPNAGFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSM-RLNRLHLAWDRNTN 64
F V G+ L +L NL + I L+ ++N + +E NLS+ L+ L ++WDR
Sbjct: 636 FVVGERKGYQLGELRNLNLRGAISITHLERVKNDMEAKEANLSAKANLHSLSMSWDRPNR 695
Query: 65 STNSAEEVLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQLPP 124
+ +VL AL+PH +L + + G +P+WM S+L +V + + C NCS LPP
Sbjct: 696 YESEEVKVLEALKPHPNLKYLEIIDFCGFCLPDWMNH-SVLKNVVSILISGCENCSCLPP 754
Query: 125 LGKLPFLNTLYLSQ-MTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNLERVLRIEGVEM 183
G+LP L +L L V+Y++DS + T FPSL ++ + NL+ + R++G E
Sbjct: 755 FGELPCLESLELQDGSVEVEYVEDSGF--LTRRRFPSLRKLHIGGFCNLKGLQRMKGAEQ 812
Query: 184 LSQLSKLSIQSIPQFELPSLPSVKEVYVGGETEED-----------------IDHEA-SF 225
L ++ I P F P+L SVK++ + GE + +H S
Sbjct: 813 FPVLEEMKISDCPMFVFPTLSSVKKLEIWGEADAGGLSSISNLSTLTSLKIFSNHTVTSL 872
Query: 226 LRDIAGKMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISL 285
L ++ + NL L + L LP L+SL +L+ L I C LES+P GL SL
Sbjct: 873 LEEMFKNLENLIYLSVSFLENLKELPTSLASLNNLKCLDIRYCYALESLPEEGLEGLSSL 932
Query: 286 RILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPELI 321
L C+ L LP+ + LT+L L I CP+LI
Sbjct: 933 TELFVEHCNMLKCLPEGLQHLTTLTSLKIRGCPQLI 968
Score = 74.7 bits (182), Expect = 6e-12
Identities = 71/249 (28%), Positives = 119/249 (47%), Gaps = 15/249 (6%)
Query: 195 IPQF-ELPSLPSVKEVYVGGETEEDIDHEASFLRDIAGKMPNLKELMIDAFHQLTVLPNE 253
+P F ELP L S++ G E + ++ FL + P+L++L I F L
Sbjct: 752 LPPFGELPCLESLE--LQDGSVEVEYVEDSGFLT--RRRFPSLRKLHIGGFCNL----KG 803
Query: 254 LSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSF---VICHSLNSLPQSVTTLTSLQ 310
L ++ E+ +++ K+ P VF L S++ L L+S+ +++TLTSL+
Sbjct: 804 LQRMKGAEQFPVLEEMKISDCPMFVFPTLSSVKKLEIWGEADAGGLSSI-SNLSTLTSLK 862
Query: 311 RLIIHYCPELILPANMNMLNSLREVSIMGGDRRRGIYNGLEDIPLLQNLSLRDFPSLRSL 370
H L+ N+ N L +S+ + + + L + L+ L +R +L SL
Sbjct: 863 IFSNHTVTSLLEEMFKNLEN-LIYLSVSFLENLKELPTSLASLNNLKCLDIRYCYALESL 921
Query: 371 PD-WLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCIDRCPRLVNRLARRTGEDWYK 429
P+ L SL EL + L LP+ L L L I CP+L+ R + GEDW+K
Sbjct: 922 PEEGLEGLSSLTELFVEHCNMLKCLPEGLQHLTTLTSLKIRGCPQLIKRCEKGIGEDWHK 981
Query: 430 IAHVPILSL 438
I+H+P +++
Sbjct: 982 ISHIPNVNI 990
Score = 52.8 bits (125), Expect = 2e-05
Identities = 48/176 (27%), Positives = 85/176 (48%), Gaps = 14/176 (7%)
Query: 225 FLRDIAGKMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLIS 284
F R ++ ++ NL F QL +L LR L+ + NK+ S+P + L +
Sbjct: 531 FKRFVSLRVLNLSN---SEFEQLPSSVGDLVHLRYLD----LSGNKICSLPKRLCK-LQN 582
Query: 285 LRILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPELILPANMNMLNSLREVSIMGGDRRR 344
L+ L C SL+ LP+ + L SL+ L++ +CP +P + +L L+ + R+
Sbjct: 583 LQTLDLYNCQSLSCLPKQTSKLCSLRNLVLDHCPLTSMPPRIGLLTCLKTLGYFVVGERK 642
Query: 345 GIYNGLEDIPLLQNLSLRDFPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQ 400
G G L+NL+LR S+ L + + + + +E +S L SL ++D+
Sbjct: 643 GYQLG-----ELRNLNLRGAISITHL-ERVKNDMEAKEANLSAKANLHSLSMSWDR 692
>gb|AAR29076.1| blight resistance protein T118 [Solanum tarijense]
Length = 948
Score = 159 bits (401), Expect = 2e-37
Identities = 108/325 (33%), Positives = 171/325 (52%), Gaps = 15/325 (4%)
Query: 6 FTVRPNAGFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSMR-LNRLHLAWD---R 61
F V+ G+ L +L +L ++ I L+ ++N + +E NLS+ L+ L + WD R
Sbjct: 627 FVVKRKKGYQLGELGSLNLYGSIKISHLERVKNDKEAKEANLSAKENLHSLSMKWDDDER 686
Query: 62 NTNSTNSAEEVLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQ 121
+ EVL AL+PH +LT +SG+RG+ +P+WM S+L +V +++ C NCS
Sbjct: 687 PHRYESEEVEVLEALKPHSNLTCLTISGFRGIRLPDWMNH-SVLKNIVLIEISGCKNCSC 745
Query: 122 LPPLGKLPFLNTLYLSQMTNVKYIDDSPYEIS-----TENAFPSLTEMTLFDLPNLERVL 176
LPP G LP L +L L + + +Y+++ ++ T FPSL ++ + NL+ ++
Sbjct: 746 LPPFGDLPCLESLQLYR-GSAEYVEEVDIDVEDSGFPTRIRFPSLRKLCICKFDNLKGLV 804
Query: 177 RIEGVEMLSQLSKLSIQSIPQFELPSLPSVKEVYVGGETEEDIDHEASFLRDIAGKMPNL 236
+ EG E L ++ I+ P +P+L S + ++ + SF ++ + NL
Sbjct: 805 KKEGGEQFPVLEEMEIRYCP---IPTLSSNLKALTSLNISDNKE-ATSFPEEMFKSLANL 860
Query: 237 KELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSL 296
K L I F L LP L+SL +L+ L I C LESIP GL SL L C L
Sbjct: 861 KYLNISHFKNLKELPTSLASLNALKSLKIQWCCALESIPEEGVKGLTSLTELIVKFCKML 920
Query: 297 NSLPQSVTTLTSLQRLIIHYCPELI 321
LP+ + LT+L R+ I CP+LI
Sbjct: 921 KCLPEGLQHLTALTRVKIWGCPQLI 945
Score = 37.7 bits (86), Expect = 0.75
Identities = 29/88 (32%), Positives = 44/88 (49%), Gaps = 6/88 (6%)
Query: 256 SLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLNSLPQSVTTLTSLQRLIIH 315
SLR L Y +K E +P+++ L+ LR + + SLP+ + L +LQ L +
Sbjct: 525 SLRVLNLSY----SKFEELPSSIG-DLVHLRYMDLSNNIEIRSLPKQLCKLQNLQTLDLQ 579
Query: 316 YCPEL-ILPANMNMLNSLREVSIMGGDR 342
YC L LP + L SLR + + G R
Sbjct: 580 YCTRLCCLPKQTSKLGSLRNLLLHGCHR 607
Score = 36.6 bits (83), Expect = 1.7
Identities = 21/58 (36%), Positives = 32/58 (54%), Gaps = 3/58 (5%)
Query: 358 NLSLRDFPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCIDRCPRL 415
NLS F LP +GD + L+ +++S ++ SLP +L+NLQ L + C RL
Sbjct: 530 NLSYSKF---EELPSSIGDLVHLRYMDLSNNIEIRSLPKQLCKLQNLQTLDLQYCTRL 584
Score = 34.7 bits (78), Expect = 6.4
Identities = 47/193 (24%), Positives = 83/193 (42%), Gaps = 21/193 (10%)
Query: 232 KMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRIL-SF 290
K+ NL+ L + +L LP + S L SL L + C++L P + L L+ L F
Sbjct: 569 KLQNLQTLDLQYCTRLCCLPKQTSKLGSLRNLLLHGCHRLTRTPPRI-GSLTCLKTLGQF 627
Query: 291 VICHSLNSLPQSVTTLTSLQRLIIHYCPEL-----ILPANMNMLNSLREVSIMGGDRRRG 345
V+ + +L + I + + AN++ +L +S+ D R
Sbjct: 628 VVKRKKGYQLGELGSLNLYGSIKISHLERVKNDKEAKEANLSAKENLHSLSMKWDDDERP 687
Query: 346 IYNGLEDIPLLQN---------LSLRDFPSLRSLPDWLGDTL--SLQELEISKFPKLTSL 394
E++ +L+ L++ F +R LPDW+ ++ ++ +EIS + L
Sbjct: 688 HRYESEEVEVLEALKPHSNLTCLTISGFRGIR-LPDWMNHSVLKNIVLIEISGCKNCSCL 746
Query: 395 PDNFDQ--LENLQ 405
P D LE+LQ
Sbjct: 747 PPFGDLPCLESLQ 759
Score = 34.3 bits (77), Expect = 8.3
Identities = 24/82 (29%), Positives = 41/82 (49%), Gaps = 1/82 (1%)
Query: 231 GKMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSF 290
G + +L+ + + ++ LP +L L++L+ L + C +L +P L SLR L
Sbjct: 544 GDLVHLRYMDLSNNIEIRSLPKQLCKLQNLQTLDLQYCTRLCCLPKQT-SKLGSLRNLLL 602
Query: 291 VICHSLNSLPQSVTTLTSLQRL 312
CH L P + +LT L+ L
Sbjct: 603 HGCHRLTRTPPRIGSLTCLKTL 624
>gb|AAR29069.1| blight resistance protein RPI [Solanum bulbocastanum]
gi|32693281|gb|AAP86601.1| putative disease resistant
protein RGA2 [Solanum bulbocastanum]
gi|46576968|sp|Q7XBQ9|RGA2_SOLBU Disease resistance
protein RGA2 (RGA2-blb) (Blight resistance protein RPI)
Length = 970
Score = 157 bits (396), Expect = 9e-37
Identities = 111/328 (33%), Positives = 170/328 (50%), Gaps = 21/328 (6%)
Query: 6 FTVRPNAGFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSM-RLNRLHLAWDRNTN 64
F V G+ L +L NL ++ I L+ ++N D +E NLS+ L+ L ++W+
Sbjct: 628 FVVGRKKGYQLGELGNLNLYGSIKISHLERVKNDKDAKEANLSAKGNLHSLSMSWNNFGP 687
Query: 65 STNSAEEV--LGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQL 122
+EEV L AL+PH +LT ++ G+RG+++P WM S+L +V + + N NCS L
Sbjct: 688 HIYESEEVKVLEALKPHSNLTSLKIYGFRGIHLPEWMNH-SVLKNIVSILISNFRNCSCL 746
Query: 123 PPLGKLPFLNTLYLSQ-MTNVKYIDDSPYEIS----TENAFPSLTEMTLFDLPNLERVLR 177
PP G LP L +L L +V+Y+++ ++ T FPSL ++ ++D +L+ +L+
Sbjct: 747 PPFGDLPCLESLELHWGSADVEYVEEVDIDVHSGFPTRIRFPSLRKLDIWDFGSLKGLLK 806
Query: 178 IEGVEMLSQLSKLSIQSIPQFELPS----LPSVKEVYVGGETEEDIDHEASFLRDIAGKM 233
EG E L ++ I P L S L S++ Y T SF ++ +
Sbjct: 807 KEGEEQFPVLEEMIIHECPFLTLSSNLRALTSLRICYNKVAT--------SFPEEMFKNL 858
Query: 234 PNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVIC 293
NLK L I + L LP L+SL +L+ L I C LES+P GL SL L C
Sbjct: 859 ANLKYLTISRCNNLKELPTSLASLNALKSLKIQLCCALESLPEEGLEGLSSLTELFVEHC 918
Query: 294 HSLNSLPQSVTTLTSLQRLIIHYCPELI 321
+ L LP+ + LT+L L I CP+LI
Sbjct: 919 NMLKCLPEGLQHLTTLTSLKIRGCPQLI 946
Score = 72.0 bits (175), Expect = 4e-11
Identities = 90/356 (25%), Positives = 144/356 (40%), Gaps = 62/356 (17%)
Query: 121 QLPPLGKLPFLNTLYLSQMTNVKYIDDSPY-EISTENAFPSLTEMTLFDLPNL---ERVL 176
QL LG L ++ +S + VK D+ +S + SL+ P++ E V
Sbjct: 637 QLGELGNLNLYGSIKISHLERVKNDKDAKEANLSAKGNLHSLSMSWNNFGPHIYESEEVK 696
Query: 177 RIEGVEMLSQLSKLSIQSIPQFELPSLPSVKEVYVGGETEEDIDHEASFLRDIAGKMPNL 236
+E ++ S L+ L I LP E ++H + N+
Sbjct: 697 VLEALKPHSNLTSLKIYGFRGIHLP---------------EWMNHSV---------LKNI 732
Query: 237 KELMIDAFHQLTVLP--NELSSLRSLEELY-IIDCNKLESIPNNVFYGLI------SLRI 287
++I F + LP +L L SLE + D +E + +V G SLR
Sbjct: 733 VSILISNFRNCSCLPPFGDLPCLESLELHWGSADVEYVEEVDIDVHSGFPTRIRFPSLRK 792
Query: 288 LSFVICHSLNSL--PQSVTTLTSLQRLIIHYCPELILPANMNMLNSLR------------ 333
L SL L + L+ +IIH CP L L +N+ L SLR
Sbjct: 793 LDIWDFGSLKGLLKKEGEEQFPVLEEMIIHECPFLTLSSNLRALTSLRICYNKVATSFPE 852
Query: 334 ----------EVSIMGGDRRRGIYNGLEDIPLLQNLSLRDFPSLRSLPD-WLGDTLSLQE 382
++I + + + L + L++L ++ +L SLP+ L SL E
Sbjct: 853 EMFKNLANLKYLTISRCNNLKELPTSLASLNALKSLKIQLCCALESLPEEGLEGLSSLTE 912
Query: 383 LEISKFPKLTSLPDNFDQLENLQKLCIDRCPRLVNRLARRTGEDWYKIAHVPILSL 438
L + L LP+ L L L I CP+L+ R + GEDW+KI+H+P +++
Sbjct: 913 LFVEHCNMLKCLPEGLQHLTTLTSLKIRGCPQLIKRCEKGIGEDWHKISHIPNVNI 968
>gb|AAR29074.1| blight resistance protein SH10 [Solanum tuberosum]
Length = 948
Score = 156 bits (395), Expect = 1e-36
Identities = 105/313 (33%), Positives = 158/313 (49%), Gaps = 9/313 (2%)
Query: 13 GFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSM-RLNRLHLAWDRNTNSTNSAEE 71
G+ L L ++ ++ I L+ ++N D +E NLS+ L+ L + W R +EE
Sbjct: 638 GYQLGKLRDVNLYGSIEITHLERVKNVMDAKEANLSAKGNLHSLIMNWSRKGPHIYESEE 697
Query: 72 V--LGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQLPPLGKLP 129
V + AL+PH +LT +SG+RG P WM S+L +V +++ C NCS LPP G+LP
Sbjct: 698 VRVIEALKPHPNLTCLTISGFRGFRFPEWMNH-SVLKNVVSIEISGCKNCSCLPPFGELP 756
Query: 130 FLNTLYLSQ-MTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNLERVLRIEGVEMLSQLS 188
L L L + V+Y+D T FPSL ++ + + PNL+ +L+ EG E L
Sbjct: 757 CLKRLELQKGSAEVEYVDSG---FPTRRRFPSLRKLFIGEFPNLKGLLKKEGEEKFPVLE 813
Query: 189 KLSIQSIPQFELPSLPSVKEVYVGGETEEDIDHEASFLRDIAGKMPNLKELMIDAFHQLT 248
+++I F +L S + + S +I NLK L I F+ L
Sbjct: 814 RMTIFYCHMFVYTTLSSNFRALTSLHISHN-NEATSLPEEIFKSFANLKYLKISLFYNLK 872
Query: 249 VLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLNSLPQSVTTLTS 308
LP+ L+ L +L+ L I C+ LES+P GL SL L C L LP+ + LT+
Sbjct: 873 ELPSSLACLNALKTLEIHSCSALESLPEEGVKGLTSLTELFVYDCEMLKFLPEGLQHLTA 932
Query: 309 LQRLIIHYCPELI 321
L L + CP+LI
Sbjct: 933 LTSLKLRRCPQLI 945
Score = 40.0 bits (92), Expect = 0.15
Identities = 46/193 (23%), Positives = 86/193 (43%), Gaps = 20/193 (10%)
Query: 232 KMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFV 291
K+ NL+ L + + L+ LP E S L SL L+ C++L S+P + L L+ L ++
Sbjct: 572 KLQNLQTLDLHNCYSLSCLPKEPSKLGSLRNLFFHGCDELNSMPPRI-GSLTFLKTLKWI 630
Query: 292 IC------HSLNSLPQ-------SVTTLTSLQRLIIHYCPELILPANMN--MLNSLREVS 336
C + L L +T L ++ ++ L N++ ++N R+
Sbjct: 631 CCGIQKKGYQLGKLRDVNLYGSIEITHLERVKNVMDAKEANLSAKGNLHSLIMNWSRKGP 690
Query: 337 IMGGDRRRGIYNGLEDIPLLQNLSLRDFPSLRSLPDWLGDTL--SLQELEISKFPKLTSL 394
+ + L+ P L L++ F R P+W+ ++ ++ +EIS + L
Sbjct: 691 HIYESEEVRVIEALKPHPNLTCLTISGFRGFR-FPEWMNHSVLKNVVSIEISGCKNCSCL 749
Query: 395 PDNFDQLENLQKL 407
P F +L L++L
Sbjct: 750 PP-FGELPCLKRL 761
>gb|AAP45174.1| putative disease resistant protein rga4 [Solanum bulbocastanum]
Length = 988
Score = 153 bits (386), Expect = 1e-35
Identities = 108/339 (31%), Positives = 169/339 (48%), Gaps = 26/339 (7%)
Query: 6 FTVRPNAGFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSM-RLNRLHLAWDRNTN 64
F V G+ L +L NL ++ I L+ ++N D E NLS+ L L ++WD +
Sbjct: 628 FIVGSKKGYQLGELKNLNLCGSISITHLERVKNDTDA-EANLSAKANLQSLSMSWDNDGP 686
Query: 65 STNSAEEV--LGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQL 122
+ +EEV L AL+PH +L + + G P+W+ S+L +++ V++ +C NC L
Sbjct: 687 NRYESEEVKVLEALKPHPNLKYLEIIAFGGFRFPSWINH-SVLEKVISVRIKSCKNCLCL 745
Query: 123 PPLGKLPFLNTLYLSQ-MTNVKYI--DDSPYEISTENAFPSLTEMTLFDLPNLERVLRIE 179
PP G+LP L L L V+Y+ DD ST +FPSL ++ ++ +L+ +++ E
Sbjct: 746 PPFGELPCLENLELQNGSAEVEYVEEDDVHSRFSTRRSFPSLKKLRIWFFRSLKGLMKEE 805
Query: 180 GVEMLSQLSKLSIQSIPQFELPSLPSVKEVYVGGETEE-----------------DIDHE 222
G E L +++I P F P+L SVK++ V G T ++
Sbjct: 806 GEEKFPMLEEMAILYCPLFVFPTLSSVKKLEVHGNTNTRGLSSISNLSTLTSLRIGANYR 865
Query: 223 ASFL-RDIAGKMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYG 281
A+ L ++ + NL+ L F L LP L+SL +L+ L I C+ LES P G
Sbjct: 866 ATSLPEEMFTSLTNLEFLSFFDFKNLKDLPTSLTSLNALKRLQIESCDSLESFPEQGLEG 925
Query: 282 LISLRILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPEL 320
L SL L C L LP+ + LT+L L + CPE+
Sbjct: 926 LTSLTQLFVKYCKMLKCLPEGLQHLTALTNLGVSGCPEV 964
Score = 52.4 bits (124), Expect = 3e-05
Identities = 43/168 (25%), Positives = 83/168 (48%), Gaps = 7/168 (4%)
Query: 223 ASFLRDIAGKMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGL 282
+S+ + K +L+ L + ++ +L LP+ + L L L + CN S+P + L
Sbjct: 516 SSYSPSLLKKFVSLRVLNL-SYSKLEQLPSSIGDLLHLRYLDL-SCNNFRSLPERLCK-L 572
Query: 283 ISLRILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPELILPANMNMLNSLREVSIMGGDR 342
+L+ L C+SLN LP+ + L+SL+ L++ CP P + +L L+ +
Sbjct: 573 QNLQTLDVHNCYSLNCLPKQTSKLSSLRHLVVDGCPLTSTPPRIGLLTCLKTLGFFIVGS 632
Query: 343 RRGIYNG-LEDIPLLQNLSLRDFPSLRSLPDW---LGDTLSLQELEIS 386
++G G L+++ L ++S+ +++ D L +LQ L +S
Sbjct: 633 KKGYQLGELKNLNLCGSISITHLERVKNDTDAEANLSAKANLQSLSMS 680
Score = 34.7 bits (78), Expect = 6.4
Identities = 22/62 (35%), Positives = 34/62 (54%), Gaps = 2/62 (3%)
Query: 364 FPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCIDRCPRLVNRLARRT 423
+ L LP +GD L L+ L++S SLP+ +L+NLQ L + C L N L ++T
Sbjct: 536 YSKLEQLPSSIGDLLHLRYLDLS-CNNFRSLPERLCKLQNLQTLDVHNCYSL-NCLPKQT 593
Query: 424 GE 425
+
Sbjct: 594 SK 595
>gb|AAR29075.1| blight resistance protein SH20 [Solanum tuberosum]
Length = 947
Score = 152 bits (384), Expect = 2e-35
Identities = 106/323 (32%), Positives = 170/323 (51%), Gaps = 16/323 (4%)
Query: 8 VRPNAGFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSMR-LNRLHLAWDRNTNS- 65
V+ G+ L +L +L ++ I L+ ++N + +E NLS+ L+ L + WD + +
Sbjct: 629 VKRKKGYQLGELGSLNLYGSIKISHLERVKNDKEAKEANLSAKENLHSLSMKWDDDEHPH 688
Query: 66 --TNSAEEVLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQLP 123
+ EVL AL+PH +LT ++SG+RG+ +P+WM S+L +V +++ C NCS LP
Sbjct: 689 RYESEEVEVLEALKPHSNLTCLKISGFRGIRLPDWMNH-SVLKNIVLIEISGCKNCSCLP 747
Query: 124 PLGKLPFLNTLYLSQMTNVKYIDDSPYEIS----TENAFPSLTEMTLFDLPNLERVLRIE 179
P G LP L +L L + + +Y+++ ++ T PSL ++ + NL+ +L+ E
Sbjct: 748 PFGDLPCLESLELYR-GSAEYVEEVDIDVDSGFPTRIRLPSLRKLCICKFDNLKGLLKKE 806
Query: 180 GVEMLSQLSKLSIQSIPQFEL-PSLPSVKEVYVGGETEEDIDHEASFLRDIAGKMPNLKE 238
G E L ++ I+ P L P+L ++ + + E SF ++ + NLK
Sbjct: 807 GGEQFPVLEEMEIRYCPIPTLSPNLKALTSLNISDNKEA-----TSFPEEMFKSLANLKY 861
Query: 239 LMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLNS 298
L I F L LP L+SL +L+ L I C LE+IP GL SL L L
Sbjct: 862 LNISHFKNLKELPTSLASLNALKSLKIQWCCALENIPKEGVKGLTSLTELIVKFSKVLKC 921
Query: 299 LPQSVTTLTSLQRLIIHYCPELI 321
LP+ + LT+L RL I CP+LI
Sbjct: 922 LPEGLHHLTALTRLKIWGCPQLI 944
Score = 38.1 bits (87), Expect = 0.58
Identities = 31/113 (27%), Positives = 51/113 (44%), Gaps = 11/113 (9%)
Query: 313 IIHYCPELILPANMNMLNSLREVSIMGGDRRRGIYNGLEDIPLLQNLSLRDFPSLR---- 368
+IH +L AN + N +RE+++ E + L+ F SLR
Sbjct: 473 LIHDLAXSLLSANTSSSN-IREINVESYTHMMMSIGFSEVVSSYSPSLLQKFVSLRVLNL 531
Query: 369 ------SLPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCIDRCPRL 415
LP +GD + L+ +++S ++ SLP +L+NLQ L + C RL
Sbjct: 532 SYSKFEELPSSIGDLVHLRYMDLSNNIEIRSLPKQLCKLQNLQTLDLQYCTRL 584
Score = 35.8 bits (81), Expect = 2.9
Identities = 30/128 (23%), Positives = 60/128 (46%), Gaps = 3/128 (2%)
Query: 223 ASFLRDIAGKMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGL 282
+S+ + K +L+ L + ++ + LP+ + L L + + + ++ S+P + L
Sbjct: 513 SSYSPSLLQKFVSLRVLNL-SYSKFEELPSSIGDLVHLRYMDLSNNIEIRSLPKQLCK-L 570
Query: 283 ISLRILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPELI-LPANMNMLNSLREVSIMGGD 341
+L+ L C L LP+ + L SL+ L++H C L P + L L+ +
Sbjct: 571 QNLQTLDLQYCTRLCCLPKQTSKLGSLRNLLLHGCHRLTRTPPRIGSLTCLKTLGQSVVK 630
Query: 342 RRRGIYNG 349
R++G G
Sbjct: 631 RKKGYQLG 638
>gb|AAR29072.1| blight resistance protein RGA4 [Solanum bulbocastanum]
Length = 1040
Score = 151 bits (382), Expect = 4e-35
Identities = 107/339 (31%), Positives = 169/339 (49%), Gaps = 26/339 (7%)
Query: 6 FTVRPNAGFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSM-RLNRLHLAWDRNTN 64
F V G+ L +L NL ++ I L+ ++N D E NLS+ L L ++WD +
Sbjct: 680 FIVGSKKGYQLGELKNLNLCGSISITHLERVKNDTDA-EANLSAKANLQSLSMSWDNDGP 738
Query: 65 STNSAEEV--LGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQL 122
+ ++EV L AL+PH +L + + G P+W+ S+L +++ V++ +C NC L
Sbjct: 739 NRYESKEVKVLEALKPHPNLKYLEIIAFGGFRFPSWINH-SVLEKVISVRIKSCKNCLCL 797
Query: 123 PPLGKLPFLNTLYLSQ-MTNVKYI--DDSPYEISTENAFPSLTEMTLFDLPNLERVLRIE 179
PP G+LP L L L V+Y+ DD ST +FPSL ++ ++ +L+ +++ E
Sbjct: 798 PPFGELPCLENLELQNGSAEVEYVEEDDVHSRFSTRRSFPSLKKLRIWFFRSLKGLMKEE 857
Query: 180 GVEMLSQLSKLSIQSIPQFELPSLPSVKEVYVGGETEE-----------------DIDHE 222
G E L +++I P F P+L SVK++ V G T ++
Sbjct: 858 GEEKFPMLEEMAILYCPLFVFPTLSSVKKLEVHGNTNTRGLSSISNLSTLTSLRIGANYR 917
Query: 223 ASFL-RDIAGKMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYG 281
A+ L ++ + NL+ L F L LP L+SL +L+ L I C+ LES P G
Sbjct: 918 ATSLPEEMFTSLTNLEFLSFFDFKNLKDLPTSLTSLNALKRLQIESCDSLESFPEQGLEG 977
Query: 282 LISLRILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPEL 320
L SL L C L LP+ + LT+L L + CPE+
Sbjct: 978 LTSLTQLFVKYCKMLKCLPEGLQHLTALTNLGVSGCPEV 1016
Score = 52.4 bits (124), Expect = 3e-05
Identities = 43/168 (25%), Positives = 83/168 (48%), Gaps = 7/168 (4%)
Query: 223 ASFLRDIAGKMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGL 282
+S+ + K +L+ L + ++ +L LP+ + L L L + CN S+P + L
Sbjct: 568 SSYSPSLLKKFVSLRVLNL-SYSKLEQLPSSIGDLLHLRYLDL-SCNNFRSLPERLCK-L 624
Query: 283 ISLRILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPELILPANMNMLNSLREVSIMGGDR 342
+L+ L C+SLN LP+ + L+SL+ L++ CP P + +L L+ +
Sbjct: 625 QNLQTLDVHNCYSLNCLPKQTSKLSSLRHLVVDGCPLTSTPPRIGLLTCLKTLGFFIVGS 684
Query: 343 RRGIYNG-LEDIPLLQNLSLRDFPSLRSLPDW---LGDTLSLQELEIS 386
++G G L+++ L ++S+ +++ D L +LQ L +S
Sbjct: 685 KKGYQLGELKNLNLCGSISITHLERVKNDTDAEANLSAKANLQSLSMS 732
Score = 34.7 bits (78), Expect = 6.4
Identities = 22/62 (35%), Positives = 34/62 (54%), Gaps = 2/62 (3%)
Query: 364 FPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCIDRCPRLVNRLARRT 423
+ L LP +GD L L+ L++S SLP+ +L+NLQ L + C L N L ++T
Sbjct: 588 YSKLEQLPSSIGDLLHLRYLDLS-CNNFRSLPERLCKLQNLQTLDVHNCYSL-NCLPKQT 645
Query: 424 GE 425
+
Sbjct: 646 SK 647
>gb|AAP45166.1| putative disease resistant protein RGA4 [Solanum bulbocastanum]
gi|46576965|sp|Q7XA39|RGA4_SOLBU Putative disease
resistance protein RGA4 (RGA4-blb)
Length = 988
Score = 151 bits (382), Expect = 4e-35
Identities = 107/339 (31%), Positives = 169/339 (49%), Gaps = 26/339 (7%)
Query: 6 FTVRPNAGFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSM-RLNRLHLAWDRNTN 64
F V G+ L +L NL ++ I L+ ++N D E NLS+ L L ++WD +
Sbjct: 628 FIVGSKKGYQLGELKNLNLCGSISITHLERVKNDTDA-EANLSAKANLQSLSMSWDNDGP 686
Query: 65 STNSAEEV--LGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQL 122
+ ++EV L AL+PH +L + + G P+W+ S+L +++ V++ +C NC L
Sbjct: 687 NRYESKEVKVLEALKPHPNLKYLEIIAFGGFRFPSWINH-SVLEKVISVRIKSCKNCLCL 745
Query: 123 PPLGKLPFLNTLYLSQ-MTNVKYI--DDSPYEISTENAFPSLTEMTLFDLPNLERVLRIE 179
PP G+LP L L L V+Y+ DD ST +FPSL ++ ++ +L+ +++ E
Sbjct: 746 PPFGELPCLENLELQNGSAEVEYVEEDDVHSRFSTRRSFPSLKKLRIWFFRSLKGLMKEE 805
Query: 180 GVEMLSQLSKLSIQSIPQFELPSLPSVKEVYVGGETEE-----------------DIDHE 222
G E L +++I P F P+L SVK++ V G T ++
Sbjct: 806 GEEKFPMLEEMAILYCPLFVFPTLSSVKKLEVHGNTNTRGLSSISNLSTLTSLRIGANYR 865
Query: 223 ASFL-RDIAGKMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYG 281
A+ L ++ + NL+ L F L LP L+SL +L+ L I C+ LES P G
Sbjct: 866 ATSLPEEMFTSLTNLEFLSFFDFKNLKDLPTSLTSLNALKRLQIESCDSLESFPEQGLEG 925
Query: 282 LISLRILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPEL 320
L SL L C L LP+ + LT+L L + CPE+
Sbjct: 926 LTSLTQLFVKYCKMLKCLPEGLQHLTALTNLGVSGCPEV 964
Score = 52.4 bits (124), Expect = 3e-05
Identities = 43/168 (25%), Positives = 83/168 (48%), Gaps = 7/168 (4%)
Query: 223 ASFLRDIAGKMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGL 282
+S+ + K +L+ L + ++ +L LP+ + L L L + CN S+P + L
Sbjct: 516 SSYSPSLLKKFVSLRVLNL-SYSKLEQLPSSIGDLLHLRYLDL-SCNNFRSLPERLCK-L 572
Query: 283 ISLRILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPELILPANMNMLNSLREVSIMGGDR 342
+L+ L C+SLN LP+ + L+SL+ L++ CP P + +L L+ +
Sbjct: 573 QNLQTLDVHNCYSLNCLPKQTSKLSSLRHLVVDGCPLTSTPPRIGLLTCLKTLGFFIVGS 632
Query: 343 RRGIYNG-LEDIPLLQNLSLRDFPSLRSLPDW---LGDTLSLQELEIS 386
++G G L+++ L ++S+ +++ D L +LQ L +S
Sbjct: 633 KKGYQLGELKNLNLCGSISITHLERVKNDTDAEANLSAKANLQSLSMS 680
Score = 34.7 bits (78), Expect = 6.4
Identities = 22/62 (35%), Positives = 34/62 (54%), Gaps = 2/62 (3%)
Query: 364 FPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCIDRCPRLVNRLARRT 423
+ L LP +GD L L+ L++S SLP+ +L+NLQ L + C L N L ++T
Sbjct: 536 YSKLEQLPSSIGDLLHLRYLDLS-CNNFRSLPERLCKLQNLQTLDVHNCYSL-NCLPKQT 593
Query: 424 GE 425
+
Sbjct: 594 SK 595
>gb|AAP45188.1| putative disease resistant protein rga2 [Solanum bulbocastanum]
Length = 872
Score = 150 bits (378), Expect = 1e-34
Identities = 109/328 (33%), Positives = 168/328 (50%), Gaps = 27/328 (8%)
Query: 6 FTVRPNAGFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSM-RLNRLHLAWDRNTN 64
F V G+ L +L NL ++ I L+ ++N D +E NLS+ L+ L ++W+
Sbjct: 536 FVVGRKKGYQLGELGNLNLYGSIKISHLERVKNDKDAKEANLSAKGNLHSLSMSWNNFGP 595
Query: 65 STNSAEEV--LGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQL 122
+EEV L AL+PH +LT ++ G+RG+++P WM S+L +V + + N NCS L
Sbjct: 596 HIYESEEVKVLEALKPHSNLTSLKIYGFRGIHLPEWMNH-SVLKNIVSILISNFRNCSCL 654
Query: 123 PPLGKLPFLNTLYLS-QMTNVKYIDDSPYEI----STENAFPSLTEMTLFDLPNLERVLR 177
PP G LP L +L L +V+Y+++ ++ T FPSL ++ ++D +L+ +L+
Sbjct: 655 PPFGDLPCLESLELHWGSADVEYVEEVDIDVHSGFPTRIRFPSLRKLDIWDFGSLKGLLK 714
Query: 178 IEGVEMLSQLSKLSIQSIPQFELPS----LPSVKEVYVGGETEEDIDHEASFLRDIAGKM 233
EG E L ++ I P L S L S++ Y T SF ++ +
Sbjct: 715 KEGEEQFPVLEEMIIHECPFLTLSSNLRALTSLRICYNKVAT--------SFPEEMFKNL 766
Query: 234 PNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVIC 293
NLK L I + L LP L+SL +L+ L LES+P GL SL L C
Sbjct: 767 ANLKYLTISRCNNLKELPTSLASLNALKSL------ALESLPEEGLEGLSSLTELFVEHC 820
Query: 294 HSLNSLPQSVTTLTSLQRLIIHYCPELI 321
+ L LP+ + LT+L L I CP+LI
Sbjct: 821 NMLKCLPEGLQHLTTLTSLKIRGCPQLI 848
Score = 70.5 bits (171), Expect = 1e-10
Identities = 91/350 (26%), Positives = 144/350 (41%), Gaps = 56/350 (16%)
Query: 121 QLPPLGKLPFLNTLYLSQMTNVKYIDDSPY-EISTENAFPSLTEMTLFDLPNL---ERVL 176
QL LG L ++ +S + VK D+ +S + SL+ P++ E V
Sbjct: 545 QLGELGNLNLYGSIKISHLERVKNDKDAKEANLSAKGNLHSLSMSWNNFGPHIYESEEVK 604
Query: 177 RIEGVEMLSQLSKLSIQSIPQFELPSLPSVKEVYVGGETEEDIDHEASFLRDIAGKMPNL 236
+E ++ S L+ L I LP E ++H + N+
Sbjct: 605 VLEALKPHSNLTSLKIYGFRGIHLP---------------EWMNHSV---------LKNI 640
Query: 237 KELMIDAFHQLTVLP--NELSSLRSLEELY-IIDCNKLESIPNNVFYGLI------SLRI 287
++I F + LP +L L SLE + D +E + +V G SLR
Sbjct: 641 VSILISNFRNCSCLPPFGDLPCLESLELHWGSADVEYVEEVDIDVHSGFPTRIRFPSLRK 700
Query: 288 LSFVICHSLNSL--PQSVTTLTSLQRLIIHYCPELILPANMNMLNSLR-----EVSIMGG 340
L SL L + L+ +IIH CP L L +N+ L SLR +
Sbjct: 701 LDIWDFGSLKGLLKKEGEEQFPVLEEMIIHECPFLTLSSNLRALTSLRICYNKVATSFPE 760
Query: 341 DRRRGIYN----------GLEDIPL-LQNLSLRDFPSLRSLPD-WLGDTLSLQELEISKF 388
+ + + N L+++P L +L+ +L SLP+ L SL EL +
Sbjct: 761 EMFKNLANLKYLTISRCNNLKELPTSLASLNALKSLALESLPEEGLEGLSSLTELFVEHC 820
Query: 389 PKLTSLPDNFDQLENLQKLCIDRCPRLVNRLARRTGEDWYKIAHVPILSL 438
L LP+ L L L I CP+L+ R + GEDW+KI+H+P +++
Sbjct: 821 NMLKCLPEGLQHLTTLTSLKIRGCPQLIKRCEKGIGEDWHKISHIPNVNI 870
>gb|AAP45185.1| putative disease resistant protein rga1 [Solanum bulbocastanum]
Length = 923
Score = 142 bits (358), Expect = 2e-32
Identities = 120/428 (28%), Positives = 183/428 (42%), Gaps = 112/428 (26%)
Query: 13 GFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSM-RLNRLHLAWDRNTNSTNSAEE 71
G+ L +L NL ++ I L ++ D +E NLS+ L+ L L+WD + +E
Sbjct: 604 GYQLGELKNLNLYGSISITKLDRVKKDTDAKEANLSAKANLHSLCLSWDLDGKHRYDSE- 662
Query: 72 VLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQLPPLGKLPFL 131
VL AL+PH +L ++G+ G+ +P+WM S+L +V +++ C NCS LPP G+LP L
Sbjct: 663 VLEALKPHSNLKYLEINGFGGILLPDWMNQ-SVLKNVVSIRIRGCENCSCLPPFGELPCL 721
Query: 132 NTLYL-SQMTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNLERVLRIEGVEMLSQLSKL 190
+L L + V+Y++D+ E FP L EMT +
Sbjct: 722 ESLELHTGSAEVEYVEDN----EGEKQFPVLEEMTFY----------------------- 754
Query: 191 SIQSIPQFELPSLPSVKEVYVGGETEEDIDHEASFLRDIAGKMPNLKELMIDAFHQLTVL 250
P F +P+L SVK + V I +A+ LR I
Sbjct: 755 ---WCPMFVIPTLSSVKTLKV-------IATDATVLRSI--------------------- 783
Query: 251 PNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLNSLPQSVTTLTSLQ 310
S+LR+L L I + + S+P +F L +L+ L+ +L LP S+ +L +L+
Sbjct: 784 ----SNLRALTSLDISNNVEATSLPEEMFKSLANLKYLNISFFRNLKELPTSLASLNALK 839
Query: 311 RLIIHYCPELILPANMNMLNSLREVSIMGGDRRRGIYNGLEDIPLLQNLSLRDFPSLRSL 370
L +C + L SL E + G
Sbjct: 840 SLKFEFC---------DALESLPEEGVKG------------------------------- 859
Query: 371 PDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCIDRCPRLVNRLARRTGEDWYKI 430
SL EL +S L LP+ L L L I +CP + R R GEDW+KI
Sbjct: 860 ------LTSLTELSVSNCMMLKCLPEGLQHLTALTTLTITQCPIVFKRCERGIGEDWHKI 913
Query: 431 AHVPILSL 438
+H+P L+L
Sbjct: 914 SHIPYLTL 921
Score = 37.0 bits (84), Expect = 1.3
Identities = 23/66 (34%), Positives = 38/66 (56%), Gaps = 1/66 (1%)
Query: 350 LEDIPLLQNLSLRDFPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCI 409
L+ L+ L+LR+ +L LP +GD + L+ L++S ++ SLP +L+NLQ L +
Sbjct: 521 LQKFVSLRVLNLRN-SNLNQLPSSIGDLVHLRYLDLSGNVRIRSLPRRLCKLQNLQTLDL 579
Query: 410 DRCPRL 415
C L
Sbjct: 580 HYCDSL 585
>ref|NP_915900.1| putative NBS-LRR type resistance protein [Oryza sativa (japonica
cultivar-group)] gi|13872974|dbj|BAB44079.1| putative
NBS-LRR type resistance protein [Oryza sativa (japonica
cultivar-group)]
Length = 1110
Score = 141 bits (356), Expect = 4e-32
Identities = 135/454 (29%), Positives = 213/454 (46%), Gaps = 25/454 (5%)
Query: 6 FTVRPNAGFSLADLPNLQP-GDNLHIKGLQNLRNGGDVREPNLSSMR-LNRLHLAWDRNT 63
F V+ +G ++ +L N+ L I+GL N+ NG D L + L LHL WD +
Sbjct: 667 FVVQKRSGHNVTELNNMDELQGQLSIRGLNNVPNGQDAVCAKLRNKEHLRTLHLIWDEDC 726
Query: 64 NSTNSAE-EVLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQL 122
S S + EVL L+PH DL + G+ G+ P+W+ S L +L + + NC ++L
Sbjct: 727 ESNPSEQQEVLEGLQPHLDLKELVIKGFPGVRFPSWLAS-SFLPKLQTIHICNC-RSTRL 784
Query: 123 PPLGKLPFLNTLYLSQMTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNLERVLRIEGVE 182
P LG+LPFL L ++ +T V + FP+L ++ L D+PNL + +
Sbjct: 785 PALGQLPFLKYLVIAGVTEVTQLSSEFTGFGQPKGFPALEDLLLEDMPNLSEWIFDVADQ 844
Query: 183 MLSQLSKLSIQSIPQF-ELPSLPS-VKEVYVGGETEEDIDHEASFLRDIAGKMPNLKELM 240
+ QL++L + PQ +LP +PS ++ +++ +E ++ + P L
Sbjct: 845 LFPQLTELGLIKCPQLKKLPPIPSTLRTLWI---SESGLESLPELQNNSCPSSPT--SLY 899
Query: 241 IDAFHQLTVLPNELSSLR--SLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLN- 297
I+ LT L L + R +L+ L I C L S+P F LISLR L C L
Sbjct: 900 INDCPNLTSLRVGLLAYRPTALKSLTIAHCEGLVSLPEECFRPLISLRSLHIYECPCLVP 959
Query: 298 -SLPQSVTTLTSLQRLIIHYCPEL--ILPANMNMLNSLREVSIMG-GDRRRGIYNGLEDI 353
+ + TS++ + ++ C L +L ++ L LR I D GL
Sbjct: 960 WTALEGGLLPTSIEDIRLNSCTPLASVLLNGLSYLPHLRHFEIADCPDINNFPAEGLPH- 1018
Query: 354 PLLQNLSLRDFPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCIDRCP 413
LQ L + L+ LP L + SL+ L IS P + SLP + L +L I CP
Sbjct: 1019 -TLQFLEISCCDDLQCLPPGLHNISSLETLRISNCPGVESLPKEGLPM-GLNELYIKGCP 1076
Query: 414 RLVNRLARRTGEDWYKIAHVPILSLRLESDVVHP 447
+ + + + GE KIAH I + ++ DV+ P
Sbjct: 1077 Q-IKQQCQEGGEYHAKIAH--IRDIEIDGDVIVP 1107
Score = 37.4 bits (85), Expect = 0.98
Identities = 43/158 (27%), Positives = 71/158 (44%), Gaps = 7/158 (4%)
Query: 252 NELSSLRSLEELYIIDCNK--LESIPNNVFYGLISLRILSFVICHSLNSLPQSVTTLTSL 309
N L R L L II K + +P+ +F L LR+L + L LP+S+ L L
Sbjct: 534 NPLYGFRKLRTLTIIHGYKSRMSQLPHGLFMKLEYLRVLD-MHGQGLKELPESIGNLKQL 592
Query: 310 QRLIIHYCPELILPANMNMLNSLREVSIMGGDRRRGIYNGLEDIPLLQNLSLRDFPSLRS 369
+ L + LPA++ L +L+ + + + R + G+ + L++L L S
Sbjct: 593 RFLDLSSTEIETLPASLVKLYNLQILKLSDCNFLREVPQGITRLINLRHLEAS--TRLLS 650
Query: 370 LPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKL 407
+G + LQELE +F N +L N+ +L
Sbjct: 651 RIHGIGSLVCLQELE--EFVVQKRSGHNVTELNNMDEL 686
>gb|AAP45181.1| putative disease resistant protein rga3 [Solanum bulbocastanum]
Length = 908
Score = 138 bits (348), Expect = 3e-31
Identities = 95/299 (31%), Positives = 156/299 (51%), Gaps = 14/299 (4%)
Query: 6 FTVRPNAGFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSM-RLNRLHLAWDRNTN 64
F V G+ L +L NL + I L+ ++N + +E NLS+ L+ L ++WDR
Sbjct: 597 FVVGERKGYQLGELRNLNLRGAISITHLERVKNDMEAKEANLSAKANLHSLSMSWDRPNR 656
Query: 65 STNSAEEVLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQLPP 124
+ +VL AL+PH +L + + G +P+WM S+L +V + + C NCS LPP
Sbjct: 657 YESEEVKVLEALKPHPNLKYLEIIDFCGFCLPDWMNH-SVLKNVVSILISGCENCSCLPP 715
Query: 125 LGKLPFLNTLYLSQ-MTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNLERVLRIEGVEM 183
G+LP L +L L V++++DS + T FPSL ++ + NL+ + R+EG E
Sbjct: 716 FGELPCLESLELQDGSVEVEFVEDSGFP--TRRRFPSLRKLHIGGFCNLKGLQRMEGEEQ 773
Query: 184 LSQLSKLSIQSIPQFELPSLPSVKEVYVGGETEEDIDHEASFLRDIAGKMPNLKELMIDA 243
L ++ I P F P+L SVK++ + GE +A L I+ + L L I +
Sbjct: 774 FPVLEEMKISDCPMFVFPTLSSVKKLEIWGEA------DARGLSSIS-NLSTLTSLKIFS 826
Query: 244 FHQLTVLPNEL-SSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLNSLPQ 301
H +T L E+ SL +L+ L + L+ +P ++ L +L+ L C++L SLP+
Sbjct: 827 NHTVTSLLEEMFKSLENLKYLSVSYLENLKELPTSL-ASLNNLKCLDIRYCYALESLPE 884
Score = 46.6 bits (109), Expect = 0.002
Identities = 33/119 (27%), Positives = 60/119 (49%), Gaps = 6/119 (5%)
Query: 282 LISLRILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPELILPANMNMLNSLREVSIMGGD 341
L +L+ L C SL+ LP+ + L SL+ L++ +CP +P + +L L+ +
Sbjct: 541 LQNLQTLDLYNCQSLSCLPKQTSKLCSLRNLVLDHCPLTSMPPRIGLLTCLKTLGYFVVG 600
Query: 342 RRRGIYNGLEDIPLLQNLSLRDFPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQ 400
R+G G L+NL+LR S+ L + + + + +E +S L SL ++D+
Sbjct: 601 ERKGYQLG-----ELRNLNLRGAISITHL-ERVKNDMEAKEANLSAKANLHSLSMSWDR 653
Score = 35.4 bits (80), Expect = 3.7
Identities = 45/189 (23%), Positives = 82/189 (42%), Gaps = 17/189 (8%)
Query: 233 MPNLKELMIDAFHQLTVLP--NELSSLRSLE------ELYIIDCNKLESIPNNVFYGLIS 284
+ N+ ++I + LP EL L SLE E+ ++ + + F L
Sbjct: 696 LKNVVSILISGCENCSCLPPFGELPCLESLELQDGSVEVEFVEDSGFPT--RRRFPSLRK 753
Query: 285 LRILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPELILPANMNMLNSLREVSIMGGDRRR 344
L I F L + + L+ + I CP + P L+S++++ I G R
Sbjct: 754 LHIGGFCNLKGLQRM-EGEEQFPVLEEMKISDCPMFVFPT----LSSVKKLEIWGEADAR 808
Query: 345 GIYNGLEDIPLLQNLSLRDFPSLRSLPDWLGDTL-SLQELEISKFPKLTSLPDNFDQLEN 403
G+ + + ++ L +L + ++ SL + + +L +L+ L +S L LP + L N
Sbjct: 809 GL-SSISNLSTLTSLKIFSNHTVTSLLEEMFKSLENLKYLSVSYLENLKELPTSLASLNN 867
Query: 404 LQKLCIDRC 412
L+ L I C
Sbjct: 868 LKCLDIRYC 876
>gb|AAN03742.1| NBS-LRR-like protein [Oryza sativa (japonica cultivar-group)]
Length = 1108
Score = 138 bits (348), Expect = 3e-31
Identities = 134/475 (28%), Positives = 201/475 (42%), Gaps = 85/475 (17%)
Query: 6 FTVRPNAGFSLADLPNLQP-GDNLHIKGLQNLRNGGDVREPNLSSM-RLNRLHLAW---- 59
F V + G+ +++L + G ++ IK L+++ + + E LS ++ L L W
Sbjct: 661 FVVHKDKGYKVSELKAMNKIGGHICIKNLESVSSAEEADEALLSEKAHISILDLIWSSSR 720
Query: 60 DRNTNSTNSAEEVLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINC 119
D + N E L +L PH +L + + G P+W IL L + L +C NC
Sbjct: 721 DFTSEEANQDIETLTSLEPHDELKELTVKAFAGFEFPHW-----ILSHLQTIHLSDCTNC 775
Query: 120 SQLPPLGKLPFLNTLYLSQMTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNLERVLRIE 179
S LP LG+LP L + + + I D S FPSL E+ D PNLER +
Sbjct: 776 SILPALGQLPLLKVIIIGGFPTIIKIGDEFSGSSEVKGFPSLKELVFEDTPNLERWTSTQ 835
Query: 180 GVEMLSQLSKLSIQSIPQF-ELPSLPSV---KEVYVGGETEEDIDHEASFLRDIAGKMPN 235
E L L +L + P+ ELP LPS ++ G + H FL P+
Sbjct: 836 DGEFLPFLRELQVLDCPKVTELPLLPSTLVELKISEAGFSVLPEVHAPRFL-------PS 888
Query: 236 LKELMIDAFHQLTVLPNELSS--LRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVIC 293
L L I LT L L S L +L++L I +C +L P
Sbjct: 889 LTRLQIHKCPNLTSLQQGLLSQQLSALQQLTITNCPELIHPPT----------------- 931
Query: 294 HSLNSLPQSVTTLTSLQRLIIHYCPEL-------ILPANMNMLNSLREVSIMGGDRRRGI 346
+ + TLT+LQ L I+ CP L +LP M+ LR S + +
Sbjct: 932 -------EGLRTLTALQSLHIYDCPRLATAEHRGLLP---RMIEDLRITSC--SNIINPL 979
Query: 347 YNGLEDIPLLQNLSLRDFPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQLE---- 402
+ L ++ L+NL + D SL + P+ L T L++LEI L SLP +
Sbjct: 980 LDELNELFALKNLVIADCVSLNTFPEKLPAT--LKKLEIFNCSNLASLPACLQEASCLKT 1037
Query: 403 -------------------NLQKLCIDRCPRLVNRLARRTGEDWYKIAHVPILSL 438
+L++L I CP L R +GEDW KI+H+ I+ +
Sbjct: 1038 MTILNCVSIKCLPAHGLPLSLEELYIKECPFLAERCQENSGEDWPKISHIAIIEI 1092
>gb|AAP45165.1| putative disease resistant protein RGA3 [Solanum bulbocastanum]
gi|46576966|sp|Q7XA40|RGA3_SOLBU Putative disease
resistance protein RGA3 (RGA1-blb) (Blight resistance
protein B149)
Length = 947
Score = 136 bits (343), Expect = 1e-30
Identities = 94/300 (31%), Positives = 154/300 (51%), Gaps = 16/300 (5%)
Query: 6 FTVRPNAGFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSM-RLNRLHLAWDRNTN 64
F V G+ L +L NL + I L+ ++N + +E NLS+ L+ L ++WDR
Sbjct: 636 FVVGERKGYQLGELRNLNLRGAISITHLERVKNDMEAKEANLSAKANLHSLSMSWDRPNR 695
Query: 65 STNSAEEVLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQLPP 124
+ +VL AL+PH +L + + G +P+WM S+L +V + + C NCS LPP
Sbjct: 696 YESEEVKVLEALKPHPNLKYLEIIDFCGFCLPDWMNH-SVLKNVVSILISGCENCSCLPP 754
Query: 125 LGKLPFLNTLYLSQ-MTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNLERVLRIEGVEM 183
G+LP L +L L V+Y++DS + T FPSL ++ + NL+ + R++G E
Sbjct: 755 FGELPCLESLELQDGSVEVEYVEDSGF--LTRRRFPSLRKLHIGGFCNLKGLQRMKGAEQ 812
Query: 184 LSQLSKLSIQSIPQFELPSLPSVKEVYVGGETEEDIDHEASFLRDIAGKMPNLKELMIDA 243
L ++ I P F P+L SVK++ + GE +A L I+ + L L I +
Sbjct: 813 FPVLEEMKISDCPMFVFPTLSSVKKLEIWGEA------DAGGLSSIS-NLSTLTSLKIFS 865
Query: 244 FHQLTVLPNELSSLRSLEELYIIDCNKLESIPN--NVFYGLISLRILSFVICHSLNSLPQ 301
H +T L E+ ++LE L + + LE++ L +L+ L C++L SLP+
Sbjct: 866 NHTVTSLLEEM--FKNLENLIYLSVSFLENLKELPTSLASLNNLKCLDIRYCYALESLPE 923
Score = 52.8 bits (125), Expect = 2e-05
Identities = 48/176 (27%), Positives = 85/176 (48%), Gaps = 14/176 (7%)
Query: 225 FLRDIAGKMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLIS 284
F R ++ ++ NL F QL +L LR L+ + NK+ S+P + L +
Sbjct: 531 FKRFVSLRVLNLSN---SEFEQLPSSVGDLVHLRYLD----LSGNKICSLPKRLCK-LQN 582
Query: 285 LRILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPELILPANMNMLNSLREVSIMGGDRRR 344
L+ L C SL+ LP+ + L SL+ L++ +CP +P + +L L+ + R+
Sbjct: 583 LQTLDLYNCQSLSCLPKQTSKLCSLRNLVLDHCPLTSMPPRIGLLTCLKTLGYFVVGERK 642
Query: 345 GIYNGLEDIPLLQNLSLRDFPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQ 400
G G L+NL+LR S+ L + + + + +E +S L SL ++D+
Sbjct: 643 GYQLG-----ELRNLNLRGAISITHL-ERVKNDMEAKEANLSAKANLHSLSMSWDR 692
>dbj|BAD35430.1| putative disease resistance protein [Oryza sativa (japonica
cultivar-group)]
Length = 1171
Score = 130 bits (326), Expect = 1e-28
Identities = 135/478 (28%), Positives = 206/478 (42%), Gaps = 52/478 (10%)
Query: 6 FTVRPNAGFSLADLPNLQP-GDNLHIKGLQNLRNGGDVREPNLSSMR-LNRLHLAWDRNT 63
F V+ +G L +L + L ++ L+N+ + + +L + L L W
Sbjct: 691 FHVKKESGHRLGELKEMNNIRGRLSVRFLENVEHQQQAVDAHLDCKEHVKHLQLEWSDLP 750
Query: 64 NSTNSA--EEVLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQ 121
S +VL ALRPH DL ++GY+G+ P W + + + L V L NC+ Q
Sbjct: 751 RPITSELDSDVLEALRPHPDLDRLNITGYKGLRSPTWF-ETNWMKALTSVILENCMGWVQ 809
Query: 122 LPPLGKLPFLNTLYLSQMTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNLERVLRIEGV 181
LPPLG+LP L L L M V I + Y FP L E+ +PN E+ IE
Sbjct: 810 LPPLGQLPLLEDLVLRNMHAVGQIGEEFYGNGEMKGFPKLEEIVFDGMPNWEKWSGIEDG 869
Query: 182 EMLSQLSKLSIQSIPQF-ELPSLPSVKEVYVGGETEEDIDHEASFLRDIAGKMPNLKELM 240
+L L++L I P+ E P L + +V V ++ +S L D + L+
Sbjct: 870 SLLPCLTRLYIAKCPKLQEAPPLNARPKVEVAITSD---SLPSSCLFDSLMASASYLILL 926
Query: 241 IDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLNS-- 298
++ L+ L + L +EEL + C + +P F GL SL++L C +L S
Sbjct: 927 VNCCSFLSSLNTD--QLSHVEELNVKSCT--DPMPACGFIGLSSLKVLRISNCSALLSSV 982
Query: 299 -------------------------------LPQSVTTLTSLQRLIIHYCPE---LILPA 324
LP+ + LT+L L+I+ C L L
Sbjct: 983 CVEAGEELDTCFFPQSLSELEIVDSNIQSSLLPRYLQGLTNLSVLVINSCDSMDLLSLAY 1042
Query: 325 NMNMLNSLREVSIMGGDRRRGIYNGLEDIPLLQNLSLRDFPSLRSLPDWLGDTLSLQELE 384
+ L SL + I + +G E++ L+ L + D + LP L +SL+ L
Sbjct: 1043 GTHHLTSLEAIIIKDCIFLSSL-DGFENLIALRKLVVADCKNFCFLPADLNALISLKTLA 1101
Query: 385 ISKFPKLTSLPDNFDQLENLQKLCIDRC-PRLVNRLARRTGEDWYKIAHVPILSLRLE 441
I PK+ LP N +LQ + + P L +L RR G +W KIAHVP L +E
Sbjct: 1102 IYGCPKMKFLPQN-GVPASLQLILLSLLHPELDRQLQRREGTEWDKIAHVPEKKLEVE 1158
>gb|AAD13037.1| NBS-LRR-like protein cD8 [Phaseolus vulgaris]
Length = 900
Score = 127 bits (318), Expect = 9e-28
Identities = 144/514 (28%), Positives = 211/514 (41%), Gaps = 101/514 (19%)
Query: 9 RPNAGFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSM-RLNRLHLAWD--RNTNS 65
+ ++ F++ L L L IK L+N+ N D +L + L L L W+ RN
Sbjct: 387 KSSSEFNIQQLGQLDLHGELSIKNLENIVNPCDALAADLKNKTHLVMLDLKWNLKRNNED 446
Query: 66 TNSAEEVLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQLPPL 125
EVL L+P + L ++GY G P W++D +L +V + C C LP L
Sbjct: 447 PIKEREVLENLQPSKHLEHLSINGYSGTQFPRWLSDTFVLN-VVSLSFYKCKYCQWLPSL 505
Query: 126 GKLPFLNTLYLSQMTNVKYIDDSPYEISTE-----------------------NAFPSLT 162
G L L L + + + ID Y S+ AFP L
Sbjct: 506 GLLTSLKHLKVRSLDEIVRIDADFYGNSSSAFASLETLIFYDMKEWEEWQCMTGAFPCLQ 565
Query: 163 EMTLFDLPNLERVL-------------------------RIEGVEMLSQ--------LSK 189
+++L D P L+ L IEGVEM + L
Sbjct: 566 DLSLHDCPKLKGHLPDLPHLKDRFITCCRQLVASTPSGVEIEGVEMETSSFDMIGHHLQS 625
Query: 190 LSIQSIPQFELPSLPSVKEVYVGGETEEDIDHEASFLRDIAGKMPNLKELMIDAFHQLTV 249
L I S P +P + V E + D +F D+ P L EL++ L +
Sbjct: 626 LRIISCPGMNIP-INYCYHFLVNLEISKCCDSLTNFPLDL---FPKLHELILSNCRNLQI 681
Query: 250 LPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVIC--HSLNSLPQSVTTLT 307
+ E L+ L I C++ ES PN GL++ +I IC L S+P+ ++ L
Sbjct: 682 ISQEHPH-HHLKSLSIYHCSEFESFPNE---GLLAPQIQEIYICAMEKLKSMPKRMSDLL 737
Query: 308 -SLQRLIIHYCPEL-----ILPAN---MNMLN-----------------SLREVSIMGGD 341
SL L I+ CPEL LP+N M +LN S++ +SI D
Sbjct: 738 PSLDYLFIYDCPELELSEGCLPSNIKEMCLLNCSKLVASLKKGGWGTNPSIQVLSINEVD 797
Query: 342 RRRGIYNGLEDIPLLQNLSLRDFPSLRSLPDWLG--DTLSLQELEISKFPKLTSLPDNFD 399
G + + Q L ++D P L+ L D+ G SLQ+L I P L LP+
Sbjct: 798 GECFPDEGFLPLSITQ-LEIKDCPKLKKL-DYRGLCHLSSLQKLGIENCPILQCLPEE-G 854
Query: 400 QLENLQKLCIDRCPRLVNRLARRTGEDWYKIAHV 433
E++ +L I+ CP L R + GEDW KIAH+
Sbjct: 855 LPESISELRIESCPLLNQRCKKEEGEDWKKIAHI 888
>emb|CAE05698.3| OSJNBa0083D01.14 [Oryza sativa (japonica cultivar-group)]
gi|50923705|ref|XP_472213.1| OSJNBa0083D01.14 [Oryza
sativa (japonica cultivar-group)]
Length = 1079
Score = 124 bits (311), Expect = 6e-27
Identities = 124/442 (28%), Positives = 194/442 (43%), Gaps = 18/442 (4%)
Query: 8 VRPNAGFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSMR-LNRLHLAWDRNTNST 66
V P GF ++L +L+ L I+ L+N+ N + L L L L W + +
Sbjct: 525 VPPKCGFIASELMDLKDLRYLCIRCLENV-NADEATLAKLGEKENLIMLSLTWKNSQQES 583
Query: 67 NSAEEVLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQLPPLG 126
++ E VL L+PH +LT ++ GY G P W+ + +I+ L + + NC LPPLG
Sbjct: 584 DTEERVLNNLQPHMNLTKLKIKGYNGSRSPCWLGNTTIIN-LTYLYISNCSYWQHLPPLG 642
Query: 127 KLPFLNTLYLSQMTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNLERVLRIEGVEMLSQ 186
+LP L LYL + +VK ID S Y FPSL + + LP LE + +EG + +
Sbjct: 643 ELPSLKYLYLICLNSVKRIDSSFYGCERPFGFPSLEYLFIEHLPALEEWVEMEGEHLFPR 702
Query: 187 LSKLSIQSIPQF-ELPSLPSVKEVYVGGETEEDIDHEASFLRDIA-GKMPNLKELMIDAF 244
L L ++ + +P+LPS HE + A + P+L L I
Sbjct: 703 LKALVVRHCKELRNVPTLPSTVNYLEMDSVGLTTLHEPYVPNENAEPQKPSLSRLKICHC 762
Query: 245 HQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLNSLPQSVT 304
L L +L+ SLEEL+I C L +P + L L+ ++ + C L P ++
Sbjct: 763 PYLETL-EQLNQFLSLEELHIEHCENLVQLPMDHLQMLSFLKHMTVLGCPKLMVPPATIR 821
Query: 305 TLTSLQRLIIHYCP--ELILPANMNMLNSLREVSIMGGDRRRGIYNGLEDIPLLQNLSLR 362
++L + C E L ++ L SL + + G D +E L LS
Sbjct: 822 LPLPTKKLHVGSCGTYETCLVNSLCGLTSLTTLMLYGCD--IAALPPVEVCKSLIALSCL 879
Query: 363 DFPSLRSLPDWLG--DTLSLQELEISKFPKLTSLP----DNFDQLENLQKLCIDRCPRLV 416
+ S L D G + SL EL++ KL LP F E+ Q + C +
Sbjct: 880 EIVSCHELADLNGMEELTSLTELKVIGCNKLEELPVVSSQRFQASEHNQ--VVTACTSYL 937
Query: 417 NRLARRTGEDWYKIAHVPILSL 438
+L R D + + P+ S+
Sbjct: 938 RKLKRLQISDPFVLQWAPLRSV 959
Score = 72.8 bits (177), Expect = 2e-11
Identities = 91/346 (26%), Positives = 149/346 (42%), Gaps = 48/346 (13%)
Query: 121 QLPPLGKLPFLNTLYLSQMTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNLERVLRIEG 180
Q P L +L + YL + + N F SL E+ + NL + L ++
Sbjct: 750 QKPSLSRLKICHCPYLETLEQL-------------NQFLSLEELHIEHCENLVQ-LPMDH 795
Query: 181 VEMLSQLSKLSIQSIPQFELPS------LPSVKEVYVGGETEEDIDHEASFLRDIAGKMP 234
++MLS L +++ P+ +P LP+ K+++VG +E + + G +
Sbjct: 796 LQMLSFLKHMTVLGCPKLMVPPATIRLPLPT-KKLHVGSCGT----YETCLVNSLCG-LT 849
Query: 235 NLKELMIDAFHQLTVLPNEL-SSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVIC 293
+L LM+ + P E+ SL +L L I+ C++L + N L SL L + C
Sbjct: 850 SLTTLMLYGCDIAALPPVEVCKSLIALSCLEIVSCHELADL--NGMEELTSLTELKVIGC 907
Query: 294 HSLNSLP-------------QSVTTLTSLQRLIIHYCPELILPANMNMLNSLREVSIMGG 340
+ L LP Q VT TS R + L S+ V+ M
Sbjct: 908 NKLEELPVVSSQRFQASEHNQVVTACTSYLRKLKRLQISDPFVLQWAPLRSVTSVTNMTI 967
Query: 341 DRRRGIYNG--LEDIPLLQNLSLRDFPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNF 398
+ R + +++ LQ + +RD L LP + SL+ LE ++ + SLP+
Sbjct: 968 NSCRCLPEEWLMQNCNNLQRIGVRDASHLEFLPSIMASLTSLESLEFTRVMLIQSLPELP 1027
Query: 399 DQLENLQKLCIDRCPRLVNRLARRTGEDWYKIAHVPILSLRLESDV 444
L LQ L + P L+ R + G DW+KIAH+P LR+ D+
Sbjct: 1028 SSLRRLQILGCN--PVLMRRCRKSRGRDWHKIAHIP--DLRIVEDI 1069
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.320 0.139 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 760,313,844
Number of Sequences: 2540612
Number of extensions: 31548843
Number of successful extensions: 89364
Number of sequences better than 10.0: 1913
Number of HSP's better than 10.0 without gapping: 347
Number of HSP's successfully gapped in prelim test: 1599
Number of HSP's that attempted gapping in prelim test: 79568
Number of HSP's gapped (non-prelim): 6190
length of query: 456
length of database: 863,360,394
effective HSP length: 131
effective length of query: 325
effective length of database: 530,540,222
effective search space: 172425572150
effective search space used: 172425572150
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 77 (34.3 bits)
Medicago: description of AC137553.6


