

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137553.4 - phase: 0 /pseudo
(921 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
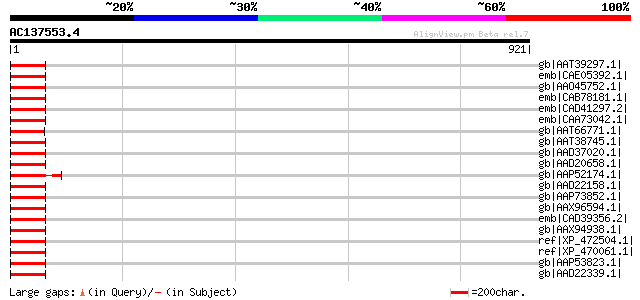
Score E
Sequences producing significant alignments: (bits) Value
gb|AAT39297.1| putative gag-pol protein [Solanum demissum] 96 4e-18
emb|CAE05392.1| OSJNBa0022F16.16 [Oryza sativa (japonica cultiva... 96 5e-18
gb|AAO45752.1| pol protein [Cucumis melo] 96 7e-18
emb|CAB78181.1| putative reverse-transcriptase-like protein [Ara... 94 2e-17
emb|CAD41297.2| OSJNBa0020J04.2 [Oryza sativa (japonica cultivar... 94 2e-17
emb|CAA73042.1| polyprotein [Ananas comosus] gi|7489851|pir||T07... 94 2e-17
gb|AAT66771.1| putative polyprotein [Solanum demissum] 94 2e-17
gb|AAT38745.1| putative polyprotein, 3'-partial [Solanum demissum] 93 5e-17
gb|AAD37020.1| putative retroelement pol polyprotein [Arabidopsi... 92 6e-17
gb|AAD20658.1| putative retroelement pol polyprotein [Arabidopsi... 92 8e-17
gb|AAP52174.1| putative polyprotein [Oryza sativa (japonica cult... 92 1e-16
gb|AAD22158.1| polyprotein [Sorghum bicolor] 92 1e-16
gb|AAP73852.1| putative gag-pol polyprotein [Oryza sativa (japon... 91 1e-16
gb|AAX96594.1| retrotransposon protein, putative, Ty3-gypsy sub-... 91 1e-16
emb|CAD39356.2| OSJNBa0059H15.7 [Oryza sativa (japonica cultivar... 91 1e-16
gb|AAX94938.1| retrotransposon protein, putative, Ty3-gypsy sub-... 91 1e-16
ref|XP_472504.1| OSJNBb0049I21.5 [Oryza sativa (japonica cultiva... 91 1e-16
ref|XP_470061.1| putative polyprotein [Oryza sativa (japonica cu... 91 1e-16
gb|AAP53823.1| putative polyprotein [Oryza sativa (japonica cult... 91 2e-16
gb|AAD22339.1| putative retroelement pol polyprotein [Arabidopsi... 90 4e-16
>gb|AAT39297.1| putative gag-pol protein [Solanum demissum]
Length = 1438
Score = 96.3 bits (238), Expect = 4e-18
Identities = 46/63 (73%), Positives = 55/63 (87%)
Query: 1 MSVSELAELKKQLEDLVEMKFFRPSVSPWGAPVLLVKKKDGSMRLRIDYRQLNKVTIKNR 60
M+ SEL ELK+QL+DL++ F RPSVSPWGAPVL V+KKDGS+R+ IDYRQLNKVTIKN+
Sbjct: 615 MAPSELKELKEQLKDLLDKGFIRPSVSPWGAPVLFVRKKDGSLRMCIDYRQLNKVTIKNK 674
Query: 61 YLL 63
Y L
Sbjct: 675 YPL 677
>emb|CAE05392.1| OSJNBa0022F16.16 [Oryza sativa (japonica cultivar-group)]
gi|50930029|ref|XP_474542.1| OSJNBa0022F16.16 [Oryza
sativa (japonica cultivar-group)]
Length = 1402
Score = 95.9 bits (237), Expect = 5e-18
Identities = 46/63 (73%), Positives = 53/63 (84%)
Query: 1 MSVSELAELKKQLEDLVEMKFFRPSVSPWGAPVLLVKKKDGSMRLRIDYRQLNKVTIKNR 60
MSV+EL ELKKQ+ +L E F RPS SPWGAPVL VKKKDGSMR+ +DYR LN+VTIKN+
Sbjct: 485 MSVNELEELKKQIRELQEKGFVRPSSSPWGAPVLFVKKKDGSMRMCVDYRSLNEVTIKNK 544
Query: 61 YLL 63
YLL
Sbjct: 545 YLL 547
>gb|AAO45752.1| pol protein [Cucumis melo]
Length = 923
Score = 95.5 bits (236), Expect = 7e-18
Identities = 46/63 (73%), Positives = 54/63 (85%)
Query: 1 MSVSELAELKKQLEDLVEMKFFRPSVSPWGAPVLLVKKKDGSMRLRIDYRQLNKVTIKNR 60
M+ +EL ELK QL++L++ F RPSVSPWGAPVL VKKKDGSMRL IDYR+LNKVT+KNR
Sbjct: 1 MAPAELKELKVQLQELLDKGFIRPSVSPWGAPVLFVKKKDGSMRLCIDYRELNKVTVKNR 60
Query: 61 YLL 63
Y L
Sbjct: 61 YPL 63
Score = 37.7 bits (86), Expect = 1.8
Identities = 14/23 (60%), Positives = 20/23 (86%)
Query: 350 RFEVFSDHKTLKYLFDQKELNVR 372
+ ++F+DHK+LKY F QKELN+R
Sbjct: 350 KIQIFTDHKSLKYFFTQKELNMR 372
>emb|CAB78181.1| putative reverse-transcriptase-like protein [Arabidopsis thaliana]
gi|4539436|emb|CAB40024.1| putative
reverse-transcriptase-like protein [Arabidopsis
thaliana] gi|7487640|pir||T04193 hypothetical protein
T4F9.40 - Arabidopsis thaliana
Length = 1240
Score = 94.4 bits (233), Expect = 2e-17
Identities = 45/63 (71%), Positives = 53/63 (83%)
Query: 1 MSVSELAELKKQLEDLVEMKFFRPSVSPWGAPVLLVKKKDGSMRLRIDYRQLNKVTIKNR 60
M+ +E+AELKKQL+DL+ F RPS SPWGAPVL VKKKDGS RL IDYR+LN+VT+KNR
Sbjct: 499 MAPAEMAELKKQLKDLLGKGFIRPSTSPWGAPVLFVKKKDGSFRLCIDYRELNRVTVKNR 558
Query: 61 YLL 63
Y L
Sbjct: 559 YPL 561
Score = 36.2 bits (82), Expect = 5.1
Identities = 14/23 (60%), Positives = 20/23 (86%)
Query: 350 RFEVFSDHKTLKYLFDQKELNVR 372
+ +VF+DHK+LKY+F Q ELN+R
Sbjct: 848 KVQVFTDHKSLKYIFTQPELNLR 870
>emb|CAD41297.2| OSJNBa0020J04.2 [Oryza sativa (japonica cultivar-group)]
gi|50928135|ref|XP_473595.1| OSJNBa0020J04.2 [Oryza
sativa (japonica cultivar-group)]
Length = 1537
Score = 94.0 bits (232), Expect = 2e-17
Identities = 45/63 (71%), Positives = 52/63 (82%)
Query: 1 MSVSELAELKKQLEDLVEMKFFRPSVSPWGAPVLLVKKKDGSMRLRIDYRQLNKVTIKNR 60
M V+EL ELKKQ+ +L E F RPS SPWGAPVL VKKKDGSMR+ +DYR LN+VTIKN+
Sbjct: 485 MPVNELEELKKQIRELQEKGFVRPSSSPWGAPVLFVKKKDGSMRMCVDYRSLNEVTIKNK 544
Query: 61 YLL 63
YLL
Sbjct: 545 YLL 547
>emb|CAA73042.1| polyprotein [Ananas comosus] gi|7489851|pir||T07863 probable
polyprotein - pineapple retrotransposon dea1
(fragment)
Length = 871
Score = 94.0 bits (232), Expect = 2e-17
Identities = 44/63 (69%), Positives = 54/63 (84%)
Query: 1 MSVSELAELKKQLEDLVEMKFFRPSVSPWGAPVLLVKKKDGSMRLRIDYRQLNKVTIKNR 60
M+ +EL EL+ QL+DL++ F RPSVSPWGAPVL VKKKDGS+RL +DYR+LNKVTIKN+
Sbjct: 25 MAPAELRELRAQLQDLLDKGFIRPSVSPWGAPVLFVKKKDGSLRLCVDYRELNKVTIKNK 84
Query: 61 YLL 63
Y L
Sbjct: 85 YPL 87
Score = 40.0 bits (92), Expect = 0.36
Identities = 17/23 (73%), Positives = 21/23 (90%)
Query: 350 RFEVFSDHKTLKYLFDQKELNVR 372
R EV++DHK+LKYLF QKELN+R
Sbjct: 374 RCEVYTDHKSLKYLFTQKELNLR 396
>gb|AAT66771.1| putative polyprotein [Solanum demissum]
Length = 1769
Score = 94.0 bits (232), Expect = 2e-17
Identities = 45/61 (73%), Positives = 51/61 (82%)
Query: 1 MSVSELAELKKQLEDLVEMKFFRPSVSPWGAPVLLVKKKDGSMRLRIDYRQLNKVTIKNR 60
M+ +EL EL QLEDL+ F RPSVSPWGAPVL VKKKDG+MR+ IDYRQLNKVT+KNR
Sbjct: 856 MAPAELRELSAQLEDLLGKGFIRPSVSPWGAPVLFVKKKDGTMRMCIDYRQLNKVTVKNR 915
Query: 61 Y 61
Y
Sbjct: 916 Y 916
>gb|AAT38745.1| putative polyprotein, 3'-partial [Solanum demissum]
Length = 1622
Score = 92.8 bits (229), Expect = 5e-17
Identities = 45/63 (71%), Positives = 52/63 (82%)
Query: 1 MSVSELAELKKQLEDLVEMKFFRPSVSPWGAPVLLVKKKDGSMRLRIDYRQLNKVTIKNR 60
M+ +EL EL QL+DL+ F RPSVSPWGAPVL VKKKDG+MR+ IDYRQLNKVT+KNR
Sbjct: 819 MAPAELKELSVQLQDLLGKGFIRPSVSPWGAPVLFVKKKDGTMRMCIDYRQLNKVTVKNR 878
Query: 61 YLL 63
Y L
Sbjct: 879 YPL 881
>gb|AAD37020.1| putative retroelement pol polyprotein [Arabidopsis thaliana]
gi|25411315|pir||D84487 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana
Length = 949
Score = 92.4 bits (228), Expect = 6e-17
Identities = 45/63 (71%), Positives = 52/63 (82%)
Query: 1 MSVSELAELKKQLEDLVEMKFFRPSVSPWGAPVLLVKKKDGSMRLRIDYRQLNKVTIKNR 60
M+ +E+AELKKQLE+L+ F RPS SPWGAPVL VKKKDGS RL IDYR LNKVT+KN+
Sbjct: 162 MAPAEMAELKKQLEELLAKGFIRPSSSPWGAPVLFVKKKDGSFRLCIDYRGLNKVTVKNK 221
Query: 61 YLL 63
Y L
Sbjct: 222 YPL 224
>gb|AAD20658.1| putative retroelement pol polyprotein [Arabidopsis thaliana]
gi|25411368|pir||G84493 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana
Length = 1611
Score = 92.0 bits (227), Expect = 8e-17
Identities = 44/63 (69%), Positives = 53/63 (83%)
Query: 1 MSVSELAELKKQLEDLVEMKFFRPSVSPWGAPVLLVKKKDGSMRLRIDYRQLNKVTIKNR 60
M+ +E+A+LKKQLE+L++ F RPS SPWGAPVL VKKKDGS RL IDYR LNKVT+KN+
Sbjct: 667 MAPAEMAKLKKQLEELLDKGFIRPSSSPWGAPVLFVKKKDGSFRLCIDYRGLNKVTVKNK 726
Query: 61 YLL 63
Y L
Sbjct: 727 YPL 729
>gb|AAP52174.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|37531170|ref|NP_919887.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
gi|22725924|gb|AAN04934.1| Putative polyprotein [Oryza
sativa] gi|20177629|gb|AAM14684.1| Putative polyprotein
[Oryza sativa (japonica cultivar-group)]
Length = 1569
Score = 91.7 bits (226), Expect = 1e-16
Identities = 48/91 (52%), Positives = 60/91 (65%), Gaps = 9/91 (9%)
Query: 1 MSVSELAELKKQLEDLVEMKFFRPSVSPWGAPVLLVKKKDGSMRLRIDYRQLNKVTIKNR 60
M V+EL ELKKQ+ +L E F RPS SPWGAPVL VKKKDGSMR+ +DYR LN+VTIKN+
Sbjct: 649 MPVNELEELKKQIRELQEKGFVRPSSSPWGAPVLFVKKKDGSMRMCVDYRSLNEVTIKNK 708
Query: 61 YLLRELMI*WIS*WVQKFLARLI*GKVITRL 91
Y L W+ +L KV +++
Sbjct: 709 YPLP---------WIDDLFDQLKGAKVFSKI 730
>gb|AAD22158.1| polyprotein [Sorghum bicolor]
Length = 924
Score = 91.7 bits (226), Expect = 1e-16
Identities = 43/63 (68%), Positives = 51/63 (80%)
Query: 1 MSVSELAELKKQLEDLVEMKFFRPSVSPWGAPVLLVKKKDGSMRLRIDYRQLNKVTIKNR 60
MSV EL ELKKQL++L + + RPS SPWG+PVL VKKKDGSMR+ IDYR LN VT+KN+
Sbjct: 1 MSVDELEELKKQLKELADKGYIRPSASPWGSPVLFVKKKDGSMRMCIDYRNLNAVTVKNK 60
Query: 61 YLL 63
Y L
Sbjct: 61 YPL 63
>gb|AAP73852.1| putative gag-pol polyprotein [Oryza sativa (japonica
cultivar-group)]
Length = 823
Score = 91.3 bits (225), Expect = 1e-16
Identities = 43/63 (68%), Positives = 52/63 (82%)
Query: 1 MSVSELAELKKQLEDLVEMKFFRPSVSPWGAPVLLVKKKDGSMRLRIDYRQLNKVTIKNR 60
M V++L ELKKQ+ +L E +F RPS SPWGAPVL VKKKDGSMR+ +DYR LN+VTIKN+
Sbjct: 1 MPVNDLEELKKQIRELQEKEFVRPSSSPWGAPVLFVKKKDGSMRMCVDYRSLNEVTIKNK 60
Query: 61 YLL 63
Y L
Sbjct: 61 YPL 63
>gb|AAX96594.1| retrotransposon protein, putative, Ty3-gypsy sub-class [Oryza
sativa (japonica cultivar-group)]
gi|62734535|gb|AAX96644.1| retrotransposon protein,
putative, Ty3-gypsy sub-class [Oryza sativa (japonica
cultivar-group)]
Length = 1158
Score = 91.3 bits (225), Expect = 1e-16
Identities = 44/63 (69%), Positives = 51/63 (80%)
Query: 1 MSVSELAELKKQLEDLVEMKFFRPSVSPWGAPVLLVKKKDGSMRLRIDYRQLNKVTIKNR 60
M V+EL ELKKQ+ +L E F RPS SPWGAPVL VKKKDGSMR+ +DYR LN+VTIKN+
Sbjct: 430 MPVNELEELKKQIRELQEKGFVRPSSSPWGAPVLFVKKKDGSMRMCVDYRSLNEVTIKNK 489
Query: 61 YLL 63
Y L
Sbjct: 490 YPL 492
>emb|CAD39356.2| OSJNBa0059H15.7 [Oryza sativa (japonica cultivar-group)]
gi|50921661|ref|XP_471191.1| OSJNBa0059H15.7 [Oryza
sativa (japonica cultivar-group)]
Length = 920
Score = 91.3 bits (225), Expect = 1e-16
Identities = 44/63 (69%), Positives = 51/63 (80%)
Query: 1 MSVSELAELKKQLEDLVEMKFFRPSVSPWGAPVLLVKKKDGSMRLRIDYRQLNKVTIKNR 60
M V+EL ELKKQ+ +L E F RPS SPWGAPVL VKKKDGSMR+ +DYR LN+VTIKN+
Sbjct: 131 MPVNELEELKKQIRELQEKGFVRPSSSPWGAPVLFVKKKDGSMRMCVDYRSLNEVTIKNK 190
Query: 61 YLL 63
Y L
Sbjct: 191 YPL 193
>gb|AAX94938.1| retrotransposon protein, putative, Ty3-gypsy sub-class [Oryza
sativa (japonica cultivar-group)]
Length = 1436
Score = 91.3 bits (225), Expect = 1e-16
Identities = 44/63 (69%), Positives = 51/63 (80%)
Query: 1 MSVSELAELKKQLEDLVEMKFFRPSVSPWGAPVLLVKKKDGSMRLRIDYRQLNKVTIKNR 60
M V+EL ELKKQ+ +L E F RPS SPWGAPVL VKKKDGSMR+ +DYR LN+VTIKN+
Sbjct: 550 MPVNELEELKKQIRELQEKGFVRPSSSPWGAPVLFVKKKDGSMRMCVDYRSLNEVTIKNK 609
Query: 61 YLL 63
Y L
Sbjct: 610 YPL 612
>ref|XP_472504.1| OSJNBb0049I21.5 [Oryza sativa (japonica cultivar-group)]
gi|38346969|emb|CAE02265.2| OSJNBb0049I21.5 [Oryza
sativa (japonica cultivar-group)]
Length = 1644
Score = 91.3 bits (225), Expect = 1e-16
Identities = 44/63 (69%), Positives = 51/63 (80%)
Query: 1 MSVSELAELKKQLEDLVEMKFFRPSVSPWGAPVLLVKKKDGSMRLRIDYRQLNKVTIKNR 60
M V+EL ELKKQ+ +L E F RPS SPWGAPVL VKKKDGSMR+ +DYR LN+VTIKN+
Sbjct: 640 MPVNELEELKKQIRELQEKGFVRPSSSPWGAPVLFVKKKDGSMRMCVDYRSLNEVTIKNK 699
Query: 61 YLL 63
Y L
Sbjct: 700 YPL 702
>ref|XP_470061.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|40786577|gb|AAR89852.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1312
Score = 91.3 bits (225), Expect = 1e-16
Identities = 43/63 (68%), Positives = 52/63 (82%)
Query: 1 MSVSELAELKKQLEDLVEMKFFRPSVSPWGAPVLLVKKKDGSMRLRIDYRQLNKVTIKNR 60
M V++L ELKKQ+ +L E +F RPS SPWGAPVL VKKKDGSMR+ +DYR LN+VTIKN+
Sbjct: 465 MPVNDLEELKKQIRELQEKEFVRPSSSPWGAPVLFVKKKDGSMRMCVDYRSLNEVTIKNK 524
Query: 61 YLL 63
Y L
Sbjct: 525 YPL 527
>gb|AAP53823.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|37534468|ref|NP_921536.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1093
Score = 90.5 bits (223), Expect = 2e-16
Identities = 43/63 (68%), Positives = 53/63 (83%)
Query: 1 MSVSELAELKKQLEDLVEMKFFRPSVSPWGAPVLLVKKKDGSMRLRIDYRQLNKVTIKNR 60
M+ +ELAE+K+Q+E+L + RPS SPWGAPVLLVKKKDGS R+ IDYR LN+VTIKN+
Sbjct: 255 MAANELAEVKRQIEELESKGYVRPSSSPWGAPVLLVKKKDGSERMVIDYRALNEVTIKNK 314
Query: 61 YLL 63
YLL
Sbjct: 315 YLL 317
>gb|AAD22339.1| putative retroelement pol polyprotein [Arabidopsis thaliana]
gi|25411103|pir||A84460 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana
Length = 1411
Score = 89.7 bits (221), Expect = 4e-16
Identities = 44/63 (69%), Positives = 50/63 (78%)
Query: 1 MSVSELAELKKQLEDLVEMKFFRPSVSPWGAPVLLVKKKDGSMRLRIDYRQLNKVTIKNR 60
M+ +E+ ELKKQLEDL+ F RPS SPWGAPVL VKKKDGS RL IDYR LN VT+KN+
Sbjct: 525 MAPAEMTELKKQLEDLLGKGFIRPSTSPWGAPVLFVKKKDGSFRLCIDYRGLNWVTVKNK 584
Query: 61 YLL 63
Y L
Sbjct: 585 YPL 587
Score = 37.0 bits (84), Expect = 3.0
Identities = 14/23 (60%), Positives = 21/23 (90%)
Query: 350 RFEVFSDHKTLKYLFDQKELNVR 372
+ +VF+DHK+LKY+F+Q ELN+R
Sbjct: 874 KVQVFTDHKSLKYIFNQPELNLR 896
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.366 0.163 0.618
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,204,760,221
Number of Sequences: 2540612
Number of extensions: 40623132
Number of successful extensions: 255389
Number of sequences better than 10.0: 2126
Number of HSP's better than 10.0 without gapping: 967
Number of HSP's successfully gapped in prelim test: 1159
Number of HSP's that attempted gapping in prelim test: 252839
Number of HSP's gapped (non-prelim): 2941
length of query: 921
length of database: 863,360,394
effective HSP length: 137
effective length of query: 784
effective length of database: 515,296,550
effective search space: 403992495200
effective search space used: 403992495200
T: 11
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.6 bits)
S2: 80 (35.4 bits)
Medicago: description of AC137553.4


