

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137077.10 + phase: 0
(247 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
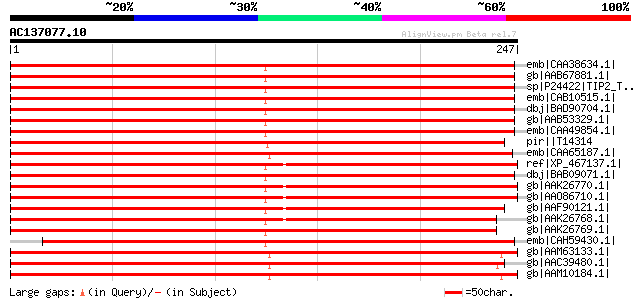
Score E
Sequences producing significant alignments: (bits) Value
emb|CAA38634.1| possible membrane channel protein [Nicotiana tab... 410 e-113
gb|AAB67881.1| membrane channel protein [Solanum tuberosum] gi|1... 409 e-113
sp|P24422|TIP2_TOBAC Probable aquaporin TIP-type RB7-18C (Tonopl... 408 e-113
emb|CAB10515.1| membrane channel like protein [Arabidopsis thali... 405 e-112
dbj|BAD90704.1| tonoplast intrinsic protein 2;1 [Mimosa pudica] 403 e-111
gb|AAB53329.1| Rb7 [Lycopersicon esculentum] 403 e-111
emb|CAA49854.1| integral membrane protein [Antirrhinum majus] gi... 401 e-110
pir||T14314 probable membrane protein - carrot gi|1794147|dbj|BA... 395 e-109
emb|CAA65187.1| aquaporin [Helianthus annuus] gi|7441272|pir||T1... 395 e-109
ref|XP_467137.1| tonoplast intrinsic protein [Oryza sativa (japo... 394 e-108
dbj|BAB09071.1| membrane channel protein-like; aquaporin (tonopl... 390 e-107
gb|AAK26770.1| tonoplast membrane integral protein ZmTIP2-3 [Zea... 390 e-107
gb|AAO86710.1| tonoplast water channel [Zea mays] 387 e-106
gb|AAF90121.1| tonoplast intrinsic protein 1 [Hordeum vulgare] 384 e-105
gb|AAK26768.1| tonoplast membrane integral protein ZmTIP2-1 [Zea... 383 e-105
gb|AAK26769.1| tonoplast membrane integral protein ZmTIP2-2 [Zea... 381 e-105
emb|CAH59430.1| aquaporin 1 [Plantago major] 378 e-104
gb|AAM63133.1| delta tonoplast integral protein delta-TIP [Arabi... 375 e-103
gb|AAC39480.1| aquaporin [Vernicia fordii] gi|11276920|pir||T488... 374 e-102
gb|AAM10184.1| delta tonoplast intrinsic protein [Arabidopsis th... 372 e-102
>emb|CAA38634.1| possible membrane channel protein [Nicotiana tabacum]
gi|7580481|gb|AAB23597.2| root-specific gene regulator
[Nicotiana tabacum] gi|126959|sp|P21653|TIP1_TOBAC
Probable aquaporin TIP-type RB7-5A (Tonoplast intrinsic
protein, root-specific RB7-5A) (TobRB7) (RT-TIP)
Length = 250
Score = 410 bits (1055), Expect = e-113
Identities = 206/247 (83%), Positives = 223/247 (89%), Gaps = 1/247 (0%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
MV+I FG+ DSFS SLKAY++EFIATL+FVFAGVGSAIAYN LT+DAALDPAGLVAVA
Sbjct: 1 MVRIAFGSIGDSFSVGSLKAYVAEFIATLLFVFAGVGSAIAYNKLTADAALDPAGLVAVA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
VAHAFALFVGV+IAANISGGHLNPAVT GLA+GGNITILTG FYWIAQLLGS VA LLL
Sbjct: 61 VAHAFALFVGVSIAANISGGHLNPAVTLGLAVGGNITILTGFFYWIAQLLGSTVACLLLK 120
Query: 121 YVT-AKSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
YVT +VPTHGVAAGLN + G+V EIIITF LVYTVYATAADPKKGS+GTIAPIAIGF+
Sbjct: 121 YVTNGLAVPTHGVAAGLNGLQGVVMEIIITFALVYTVYATAADPKKGSLGTIAPIAIGFI 180
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGSY 239
VGANILAAGPFSGGSMNPARSFGPAVV+G+F+ NWIYW GPLIGGGLAG IYGDVFIG +
Sbjct: 181 VGANILAAGPFSGGSMNPARSFGPAVVAGDFSQNWIYWAGPLIGGGLAGFIYGDVFIGCH 240
Query: 240 TPAPASE 246
TP P SE
Sbjct: 241 TPLPTSE 247
>gb|AAB67881.1| membrane channel protein [Solanum tuberosum]
gi|11276918|pir||T48884 membrane channel protein
[imported] - potato (fragment)
Length = 250
Score = 409 bits (1052), Expect = e-113
Identities = 204/247 (82%), Positives = 223/247 (89%), Gaps = 1/247 (0%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
MV+I FG+ DS S SLKAYL+EFIATL+FVFAGVGSAIAYN LTSDAALDPAGLVA+A
Sbjct: 1 MVRIAFGSIGDSLSVGSLKAYLAEFIATLLFVFAGVGSAIAYNKLTSDAALDPAGLVAIA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
VAHAFALFVGV++AANISGGHLNPAVT GLA+GGNIT LTGLFYW+AQLLGS VA LLL
Sbjct: 61 VAHAFALFVGVSMAANISGGHLNPAVTLGLAVGGNITTLTGLFYWVAQLLGSTVACLLLK 120
Query: 121 YVT-AKSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
YVT +VPTHGVAAG+N G+V EI+ITF LVYTVYATAADPKKGS+GTIAPIAIGF+
Sbjct: 121 YVTNGLAVPTHGVAAGMNGAEGVVMEIVITFALVYTVYATAADPKKGSLGTIAPIAIGFI 180
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGSY 239
VGANILAAGPFSGGSMNPARSFGPAVV+G+F+ NWIYWVGPLIGGGLAG IYGDVFIGS+
Sbjct: 181 VGANILAAGPFSGGSMNPARSFGPAVVAGDFSQNWIYWVGPLIGGGLAGFIYGDVFIGSH 240
Query: 240 TPAPASE 246
TP P SE
Sbjct: 241 TPLPTSE 247
>sp|P24422|TIP2_TOBAC Probable aquaporin TIP-type RB7-18C (Tonoplast intrinsic protein,
root-specific RB7-18C) (TobRB7) (RT-TIP)
Length = 250
Score = 408 bits (1049), Expect = e-113
Identities = 206/247 (83%), Positives = 221/247 (89%), Gaps = 1/247 (0%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
MV I FG+ DSFS SLKAY++EFIATL+FVFAGVGSAIAYN LT+DAALDPAGLVAVA
Sbjct: 1 MVLIAFGSIGDSFSVGSLKAYVAEFIATLLFVFAGVGSAIAYNKLTADAALDPAGLVAVA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
VAHAFALFVGV+IAANISGGHLNPAVT GLA+GGNITILTG FYWIAQLLGS VA LLL
Sbjct: 61 VAHAFALFVGVSIAANISGGHLNPAVTLGLAVGGNITILTGFFYWIAQLLGSTVACLLLK 120
Query: 121 YVT-AKSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
YVT +VPTHGVAAGLN G+V EIIITF LVYTVYATAADPKKGS+GTIAPIAIGF+
Sbjct: 121 YVTNGLAVPTHGVAAGLNGFQGVVMEIIITFALVYTVYATAADPKKGSLGTIAPIAIGFI 180
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGSY 239
VGANILAAGPFSGGSMNPARSFGPAVV+G+F+ NWIYW GPLIGGGLAG IYGDVFIG +
Sbjct: 181 VGANILAAGPFSGGSMNPARSFGPAVVAGDFSQNWIYWAGPLIGGGLAGFIYGDVFIGCH 240
Query: 240 TPAPASE 246
TP P SE
Sbjct: 241 TPLPTSE 247
>emb|CAB10515.1| membrane channel like protein [Arabidopsis thaliana]
gi|7268486|emb|CAB78737.1| membrane channel like protein
[Arabidopsis thaliana] gi|21618185|gb|AAM67235.1|
membrane channel like protein [Arabidopsis thaliana]
gi|15810075|gb|AAL06963.1| AT4g17340/dl4705w
[Arabidopsis thaliana] gi|14194155|gb|AAK56272.1|
AT4g17340/dl4705w [Arabidopsis thaliana]
gi|32363277|sp|Q41975|TIP22_ARATH Probable aquaporin
TIP2.2 (Tonoplast intrinsic protein 2.2)
gi|15236043|ref|NP_193465.1| major intrinsic family
protein / MIP family protein [Arabidopsis thaliana]
gi|7441256|pir||F71442 probable membrane channel protein
- Arabidopsis thaliana
Length = 250
Score = 405 bits (1041), Expect = e-112
Identities = 204/247 (82%), Positives = 223/247 (89%), Gaps = 1/247 (0%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
MVKI G+ DSFS ASLKAYLSEFIATL+FVFAGVGSA+A+ LTSDAALDPAGLVAVA
Sbjct: 1 MVKIEIGSVGDSFSVASLKAYLSEFIATLLFVFAGVGSALAFAKLTSDAALDPAGLVAVA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
VAHAFALFVGV+IAANISGGHLNPAVT GLA+GGNIT++TG FYWIAQ LGSIVA LLL
Sbjct: 61 VAHAFALFVGVSIAANISGGHLNPAVTLGLAVGGNITVITGFFYWIAQCLGSIVACLLLV 120
Query: 121 YVT-AKSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
+VT +SVPTHGVAAGL I G+V EI++TF LVYTVYATAADPKKGS+GTIAPIAIGF+
Sbjct: 121 FVTNGESVPTHGVAAGLGAIEGVVMEIVVTFALVYTVYATAADPKKGSLGTIAPIAIGFI 180
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGSY 239
VGANILAAGPFSGGSMNPARSFGPAVVSG+F+ WIYWVGPL+GG LAGLIYGDVFIGSY
Sbjct: 181 VGANILAAGPFSGGSMNPARSFGPAVVSGDFSQIWIYWVGPLVGGALAGLIYGDVFIGSY 240
Query: 240 TPAPASE 246
PAP +E
Sbjct: 241 APAPTTE 247
>dbj|BAD90704.1| tonoplast intrinsic protein 2;1 [Mimosa pudica]
Length = 250
Score = 403 bits (1036), Expect = e-111
Identities = 200/247 (80%), Positives = 222/247 (88%), Gaps = 1/247 (0%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
MVKI FG+ +DSFSAAS+KAY++EFIATL+FVFAGVGSAIAY LT DAALDPAGLVAVA
Sbjct: 1 MVKIAFGSLNDSFSAASIKAYVAEFIATLLFVFAGVGSAIAYGALTKDAALDPAGLVAVA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
VAHAFALFVGV++AANISGGHLNPAVTFGL IGGNIT+LT +FYWI QLLGS+VA LLL
Sbjct: 61 VAHAFALFVGVSVAANISGGHLNPAVTFGLVIGGNITVLTAIFYWIFQLLGSLVACLLLK 120
Query: 121 YVT-AKSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
+VT A +VP HG+A+G+ G+VFEI+ TFGLVYTVYATAADPKKGS+G IAPIAIGF+
Sbjct: 121 FVTSAPTVPVHGLASGVGIFEGIVFEIVATFGLVYTVYATAADPKKGSLGIIAPIAIGFI 180
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGSY 239
VGANILAAGPFSGGSMNPARSFGPA VSG+F + WIYWVGPLIGGGLAGLIYGDVFIGSY
Sbjct: 181 VGANILAAGPFSGGSMNPARSFGPAAVSGDFTNYWIYWVGPLIGGGLAGLIYGDVFIGSY 240
Query: 240 TPAPASE 246
P P SE
Sbjct: 241 APVPTSE 247
>gb|AAB53329.1| Rb7 [Lycopersicon esculentum]
Length = 250
Score = 403 bits (1036), Expect = e-111
Identities = 201/247 (81%), Positives = 222/247 (89%), Gaps = 1/247 (0%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
MV+I FG+ DS S SLKAYL+EFIATL+FVFAGVGSAIA+N LTS AALDPAGLVA+A
Sbjct: 1 MVRIAFGSIGDSLSVGSLKAYLAEFIATLLFVFAGVGSAIAFNKLTSGAALDPAGLVAIA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
VAHAFALFVGV++AANISGGHLNPAVT GLA+GGNITILTGLFYW+AQLLGS VA LLL
Sbjct: 61 VAHAFALFVGVSMAANISGGHLNPAVTLGLAVGGNITILTGLFYWVAQLLGSTVACLLLK 120
Query: 121 YVT-AKSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
YVT +VPTHGVAAG++ G+V EI+ITF LVYTVYATAADPKKGS+GTIAPIAIGF+
Sbjct: 121 YVTNGLAVPTHGVAAGMSGAEGVVMEIVITFALVYTVYATAADPKKGSLGTIAPIAIGFI 180
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGSY 239
VGANILAAGPFSGGSMNPARSFGPAVV+G+F+ NWIYWVGPLIGGGLAG IYGDVFIG +
Sbjct: 181 VGANILAAGPFSGGSMNPARSFGPAVVAGDFSQNWIYWVGPLIGGGLAGFIYGDVFIGCH 240
Query: 240 TPAPASE 246
TP P SE
Sbjct: 241 TPLPTSE 247
>emb|CAA49854.1| integral membrane protein [Antirrhinum majus]
gi|461929|sp|P33560|TIP_ANTMA Probable aquaporin
TIP-type (Tonoplast intrinsic protein DiP) (Dark
intrinsic protein)
Length = 250
Score = 401 bits (1030), Expect = e-110
Identities = 201/247 (81%), Positives = 220/247 (88%), Gaps = 1/247 (0%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
MVKI FG+ DSFS AS+KAY++EFIATL+FVFAGVGSAIAYN LTSDAALDPAGLVAVA
Sbjct: 1 MVKIAFGSIGDSFSVASIKAYVAEFIATLLFVFAGVGSAIAYNKLTSDAALDPAGLVAVA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
VAHAFALFVGV++AAN+SGGHLNPAVT GLA+GGNITILTGLFYWIAQ LGS VA LLL
Sbjct: 61 VAHAFALFVGVSMAANVSGGHLNPAVTLGLAVGGNITILTGLFYWIAQCLGSTVACLLLK 120
Query: 121 YVT-AKSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
+VT SVPTHGVAAG++ I G+V EIIITF LVYTVYATAADPKKGS+G IAPIAIGF+
Sbjct: 121 FVTNGLSVPTHGVAAGMDAIQGVVMEIIITFALVYTVYATAADPKKGSLGVIAPIAIGFI 180
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGSY 239
VGANILAAGPFSGGSMNPARSFGPAV SG+F+ NWIYW GPLIGG LAG IYGDVFI ++
Sbjct: 181 VGANILAAGPFSGGSMNPARSFGPAVASGDFSQNWIYWAGPLIGGALAGFIYGDVFITAH 240
Query: 240 TPAPASE 246
P P SE
Sbjct: 241 APLPTSE 247
>pir||T14314 probable membrane protein - carrot gi|1794147|dbj|BAA19129.1|
similar to EMBL Accession Number : X54855 [Daucus
carota]
Length = 248
Score = 395 bits (1016), Expect = e-109
Identities = 196/242 (80%), Positives = 218/242 (89%), Gaps = 1/242 (0%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
MVK+ G+ DSFSA SLK+YL+EFIATL+FVFAGVGSAIAY LT+DAALDPAGLVA+A
Sbjct: 1 MVKLAIGSVGDSFSAVSLKSYLAEFIATLLFVFAGVGSAIAYGHLTADAALDPAGLVAIA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
+AHAFALFVGV++AANISGGHLNPAVT GLAIGGNITI+TGLFYWIAQ LGS VA LL
Sbjct: 61 IAHAFALFVGVSMAANISGGHLNPAVTLGLAIGGNITIITGLFYWIAQCLGSTVACFLLK 120
Query: 121 YVTA-KSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
+VTA K++PTHGV AGL G+VFEI+ITF LVYTVYATAADPKKGS+GTIAPIAIGF+
Sbjct: 121 FVTAGKAIPTHGVGAGLGAAEGVVFEIVITFALVYTVYATAADPKKGSLGTIAPIAIGFI 180
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGSY 239
VGANILAAGPFSGGSMNPARSFGPAV S +F+ +WIYWVGPLIGGGLAGL+YGDVFIGSY
Sbjct: 181 VGANILAAGPFSGGSMNPARSFGPAVASFDFSGHWIYWVGPLIGGGLAGLVYGDVFIGSY 240
Query: 240 TP 241
P
Sbjct: 241 AP 242
>emb|CAA65187.1| aquaporin [Helianthus annuus] gi|7441272|pir||T14000 aquaporin TIP7
- common sunflower
Length = 250
Score = 395 bits (1014), Expect = e-109
Identities = 196/246 (79%), Positives = 219/246 (88%), Gaps = 1/246 (0%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
MVK+ G+ DS SA S+K+YL+EFIATL+FVFAGVGSAIAY LT+DAALDPAGLVA+A
Sbjct: 1 MVKLAIGSIGDSLSAGSIKSYLAEFIATLLFVFAGVGSAIAYGKLTTDAALDPAGLVAIA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
+AHAFALFVGV++AANISGGHLNPAVTFGLAIGGNITI+TGLFYWIAQ LGSIVA LL
Sbjct: 61 IAHAFALFVGVSMAANISGGHLNPAVTFGLAIGGNITIITGLFYWIAQCLGSIVACFLLQ 120
Query: 121 YVTAK-SVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
+VT +VPTHGVA G+N + G+V EIIITF LVYTVYATA DPKKGS+GTIAP+AIGF+
Sbjct: 121 FVTGGLAVPTHGVADGMNGVQGVVMEIIITFALVYTVYATAVDPKKGSLGTIAPMAIGFI 180
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGSY 239
VGANILAAGPFSGGSMNPARSFGPAVVSG+F+ NWIYWVGPLIGGGLAG I+GDVFIGSY
Sbjct: 181 VGANILAAGPFSGGSMNPARSFGPAVVSGDFSQNWIYWVGPLIGGGLAGFIFGDVFIGSY 240
Query: 240 TPAPAS 245
P S
Sbjct: 241 ETLPDS 246
>ref|XP_467137.1| tonoplast intrinsic protein [Oryza sativa (japonica
cultivar-group)] gi|14140156|emb|CAC39073.1| putative
aquaporin [Oryza sativa] gi|32879774|dbj|BAC79359.1|
tonoplast intrinsic protein [Oryza sativa (japonica
cultivar-group)] gi|49388575|dbj|BAD25694.1| tonoplast
intrinsic protein [Oryza sativa (japonica
cultivar-group)] gi|49387590|dbj|BAD25765.1| tonoplast
intrinsic protein [Oryza sativa (japonica
cultivar-group)]
Length = 248
Score = 394 bits (1011), Expect = e-108
Identities = 192/248 (77%), Positives = 222/248 (89%), Gaps = 2/248 (0%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
MVK+ FG+ DSFSA S+KAY++EFIATL+FVFAGVGSAIAY LT+ ALDPAGLVA+A
Sbjct: 1 MVKLAFGSLGDSFSATSVKAYVAEFIATLLFVFAGVGSAIAYGQLTNGGALDPAGLVAIA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
+AHA ALFVGV++AANISGGHLNPAVTFGLA+GG+ITILTGLFYWIAQLLG+ +A LLL
Sbjct: 61 IAHALALFVGVSVAANISGGHLNPAVTFGLAVGGHITILTGLFYWIAQLLGASIACLLLK 120
Query: 121 YVT-AKSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
+VT K++PTHGVA G++ + G+V EI+ITF LVYTVYATAADPKKGS+GTIAPIAIGF+
Sbjct: 121 FVTHGKAIPTHGVA-GISELEGVVMEIVITFALVYTVYATAADPKKGSLGTIAPIAIGFI 179
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGSY 239
VGANILAAGPFSGGSMNPARSFGPAV +GNFA NW+YWVGPLIGGGLAGL+YGDVFIGSY
Sbjct: 180 VGANILAAGPFSGGSMNPARSFGPAVAAGNFAGNWVYWVGPLIGGGLAGLVYGDVFIGSY 239
Query: 240 TPAPASEF 247
P ++
Sbjct: 240 QPVADQDY 247
>dbj|BAB09071.1| membrane channel protein-like; aquaporin (tonoplast intrinsic
protein)-like [Arabidopsis thaliana]
gi|15238100|ref|NP_199556.1| major intrinsic family
protein / MIP family protein [Arabidopsis thaliana]
gi|32363406|sp|Q9FGL2|TIP23_ARATH Probable aquaporin
TIP2.3 (Tonoplast intrinsic protein 2.3)
gi|44681458|gb|AAS47669.1| At5g47450 [Arabidopsis
thaliana] gi|40822901|gb|AAR92248.1| At5g47450
[Arabidopsis thaliana]
Length = 250
Score = 390 bits (1003), Expect = e-107
Identities = 196/247 (79%), Positives = 217/247 (87%), Gaps = 1/247 (0%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
MVKI G+ DSFS +SLKAYLSEFIATL+FVFAGVGSA+A+ LTSD ALDPAGLVA+A
Sbjct: 1 MVKIEVGSVGDSFSVSSLKAYLSEFIATLLFVFAGVGSAVAFAKLTSDGALDPAGLVAIA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
+AHAFALFVGV+IAANISGGHLNPAVT GLAIGGNIT++TG FYWIAQ LGSIVA LLL
Sbjct: 61 IAHAFALFVGVSIAANISGGHLNPAVTLGLAIGGNITLITGFFYWIAQCLGSIVACLLLV 120
Query: 121 YVT-AKSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
+VT KSVPTHGV+AGL + G+V EI++TF LVYTVYATAADPKKGS+GTIAPIAIGF+
Sbjct: 121 FVTNGKSVPTHGVSAGLGAVEGVVMEIVVTFALVYTVYATAADPKKGSLGTIAPIAIGFI 180
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGSY 239
VGANILAAGPFSGGSMNPARSFGPAVVSG+ + WIYWVGPL+GG LAGLIYGDVFIGSY
Sbjct: 181 VGANILAAGPFSGGSMNPARSFGPAVVSGDLSQIWIYWVGPLVGGALAGLIYGDVFIGSY 240
Query: 240 TPAPASE 246
E
Sbjct: 241 EAVETRE 247
>gb|AAK26770.1| tonoplast membrane integral protein ZmTIP2-3 [Zea mays]
gi|3264596|gb|AAC24569.1| putative tonoplast aquaporin
[Zea mays] gi|7441255|pir||T01648 probable tonoplast
aquaporin - maize
Length = 248
Score = 390 bits (1001), Expect = e-107
Identities = 190/248 (76%), Positives = 221/248 (88%), Gaps = 2/248 (0%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
MVK+ FG+F DS SAASLKAY++EFIATL+FVFAGVGSAIAY+ LT ALDPAGLVA+A
Sbjct: 1 MVKLAFGSFRDSLSAASLKAYVAEFIATLLFVFAGVGSAIAYSQLTKGGALDPAGLVAIA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
+AHAFALFVGV++AANISGGHLNPAVTFGLA+GG+ITILTG+ YW+AQLLG+ VA LL
Sbjct: 61 IAHAFALFVGVSMAANISGGHLNPAVTFGLAVGGHITILTGILYWVAQLLGASVACFLLQ 120
Query: 121 YVT-AKSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
YVT +++PTHGV+ G++ I G+V EI+ITF LVYTVYATAADPKKGS+GTIAP+AIGF+
Sbjct: 121 YVTHGQAIPTHGVS-GISEIEGVVMEIVITFALVYTVYATAADPKKGSLGTIAPMAIGFI 179
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGSY 239
VGANILAAGPFSGGSMNPARSFGPAV +GNFA NW+YWVGPL+GGGLAGL+YGDVFI SY
Sbjct: 180 VGANILAAGPFSGGSMNPARSFGPAVAAGNFAGNWVYWVGPLVGGGLAGLVYGDVFIASY 239
Query: 240 TPAPASEF 247
P E+
Sbjct: 240 QPVGQQEY 247
>gb|AAO86710.1| tonoplast water channel [Zea mays]
Length = 248
Score = 387 bits (994), Expect = e-106
Identities = 189/248 (76%), Positives = 220/248 (88%), Gaps = 2/248 (0%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
MVK+ FG+F DS SAASLKAY++EFIATL+FVFAGVGSAIAY+ LT ALDPAGLVA+A
Sbjct: 1 MVKLAFGSFRDSLSAASLKAYVAEFIATLLFVFAGVGSAIAYSQLTKGGALDPAGLVAIA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
+AHAFALFVGV++AANISGGHLNPAVTFGLA+GG+ITILTG+ YW+AQLLG+ VA LL
Sbjct: 61 IAHAFALFVGVSMAANISGGHLNPAVTFGLAVGGHITILTGILYWVAQLLGASVACFLLQ 120
Query: 121 YVT-AKSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
YVT +++PTHGV+ G++ I G+V EI+ITF LVYTVYATAADPKKGS+GTIAP+AIGF+
Sbjct: 121 YVTHGQAIPTHGVS-GISEIEGVVMEIVITFALVYTVYATAADPKKGSLGTIAPMAIGFI 179
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGSY 239
VGANILAAGPFSGGSMNPARSFGPAV +GNFA NW+YWVGPL+GGGLAG +YGDVFI SY
Sbjct: 180 VGANILAAGPFSGGSMNPARSFGPAVAAGNFAGNWVYWVGPLVGGGLAGPVYGDVFIASY 239
Query: 240 TPAPASEF 247
P E+
Sbjct: 240 QPVGQQEY 247
>gb|AAF90121.1| tonoplast intrinsic protein 1 [Hordeum vulgare]
Length = 249
Score = 384 bits (985), Expect = e-105
Identities = 185/242 (76%), Positives = 220/242 (90%), Gaps = 2/242 (0%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
MVK+ FG+F DSFSA S+++Y++EFIATL+FVFAGVGSAI+Y LT ALDPAGLVA+A
Sbjct: 1 MVKLAFGSFGDSFSATSIRSYVAEFIATLLFVFAGVGSAISYGQLTQGGALDPAGLVAIA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
+AHAFALFVGVA+AANISGGHLNPAVTFGLA+GG++TILTGLFYW+AQLLG+ VA LLL
Sbjct: 61 IAHAFALFVGVAMAANISGGHLNPAVTFGLAVGGHVTILTGLFYWVAQLLGASVACLLLQ 120
Query: 121 YVT-AKSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
+VT A+++PTH V+ G++ + G+V EI+ITF LVYTVYATAADPKKGS+GTIAP+AIGF+
Sbjct: 121 FVTHAQAMPTHAVS-GISEVEGVVMEIVITFALVYTVYATAADPKKGSLGTIAPMAIGFI 179
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGSY 239
VGANILAAGPFSGGSMNPARSFGPAV +GNF+ +W+YWVGPLIGGGLAGL+YGDVFI SY
Sbjct: 180 VGANILAAGPFSGGSMNPARSFGPAVAAGNFSGHWVYWVGPLIGGGLAGLVYGDVFIASY 239
Query: 240 TP 241
P
Sbjct: 240 QP 241
>gb|AAK26768.1| tonoplast membrane integral protein ZmTIP2-1 [Zea mays]
Length = 249
Score = 383 bits (983), Expect = e-105
Identities = 185/238 (77%), Positives = 217/238 (90%), Gaps = 2/238 (0%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
MVK+ FG+ DSFSA S+KAY++EFIATL+FVFAGVGSAIAY LT+ ALDPAGLVA+A
Sbjct: 1 MVKLAFGSVGDSFSATSIKAYVAEFIATLLFVFAGVGSAIAYGQLTNGGALDPAGLVAIA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
+AHA ALFVGV++AANISGGHLNPAVTFGLA+GG+ITILTG+FYW+AQLLG+ VA LLL
Sbjct: 61 IAHALALFVGVSVAANISGGHLNPAVTFGLAVGGHITILTGVFYWVAQLLGATVACLLLG 120
Query: 121 YVT-AKSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
+VT K++PTH VA G++ + G+VFE++ITF LVYTVYATAADPKKGS+GTIAPIAIGF+
Sbjct: 121 FVTHGKAIPTHAVA-GISELEGVVFEVVITFALVYTVYATAADPKKGSLGTIAPIAIGFI 179
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIG 237
VGANILAAGPFSGGSMNPARSFGPAV +G+FA NW+YWVGPL+GGGLAGL+YGDVFIG
Sbjct: 180 VGANILAAGPFSGGSMNPARSFGPAVAAGDFAGNWVYWVGPLVGGGLAGLVYGDVFIG 237
Score = 39.3 bits (90), Expect = 0.10
Identities = 30/86 (34%), Positives = 39/86 (44%), Gaps = 4/86 (4%)
Query: 160 AADPKKGSIGTIAPIAIGFVVGANILAAGPFSGGSMNPARSFGPAVVSG-NFADNWIYWV 218
A DP G + A+ VG ++ A SGG +NPA +FG AV YWV
Sbjct: 50 ALDPA-GLVAIAIAHALALFVGVSV--AANISGGHLNPAVTFGLAVGGHITILTGVFYWV 106
Query: 219 GPLIGGGLAGLIYGDVFIGSYTPAPA 244
L+G +A L+ G V G P A
Sbjct: 107 AQLLGATVACLLLGFVTHGKAIPTHA 132
>gb|AAK26769.1| tonoplast membrane integral protein ZmTIP2-2 [Zea mays]
Length = 250
Score = 381 bits (979), Expect = e-105
Identities = 184/238 (77%), Positives = 213/238 (89%), Gaps = 1/238 (0%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
MVK+ FG+ DSFS S+KAY++EFIATL+FVFAGVGSAIA+ LT+ ALDPAGLVA+A
Sbjct: 1 MVKLAFGSVGDSFSVTSIKAYVAEFIATLLFVFAGVGSAIAFGQLTNGGALDPAGLVAIA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
VAHA ALFVGV++AAN SGGHLNPAVTFGLA+GG+IT+LTGLFYW+AQLLG+ VA LLL
Sbjct: 61 VAHALALFVGVSVAANTSGGHLNPAVTFGLAVGGHITVLTGLFYWVAQLLGASVACLLLR 120
Query: 121 YVT-AKSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
+VT K++PTHGV+ G + G+VFEI+ITF LVYTVYATAADPKKGS+GTIAPIAIGF+
Sbjct: 121 FVTHGKAIPTHGVSGGTTELEGVVFEIVITFALVYTVYATAADPKKGSLGTIAPIAIGFI 180
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIG 237
VGANILAAGPFSGGSMNPARSFGPAV + +FA NW+YWVGPLIGGGLAGL+YGDVFIG
Sbjct: 181 VGANILAAGPFSGGSMNPARSFGPAVAAADFAGNWVYWVGPLIGGGLAGLVYGDVFIG 238
Score = 36.2 bits (82), Expect = 0.85
Identities = 29/84 (34%), Positives = 39/84 (45%), Gaps = 6/84 (7%)
Query: 160 AADPKKGSIGTIAPIAIGFVVGANILAAGPFSGGSMNPARSFGPAVVSGNFA--DNWIYW 217
A DP G + A+ VG ++ A SGG +NPA +FG AV G+ YW
Sbjct: 50 ALDPA-GLVAIAVAHALALFVGVSV--AANTSGGHLNPAVTFGLAV-GGHITVLTGLFYW 105
Query: 218 VGPLIGGGLAGLIYGDVFIGSYTP 241
V L+G +A L+ V G P
Sbjct: 106 VAQLLGASVACLLLRFVTHGKAIP 129
>emb|CAH59430.1| aquaporin 1 [Plantago major]
Length = 234
Score = 378 bits (971), Expect = e-104
Identities = 185/231 (80%), Positives = 211/231 (91%), Gaps = 1/231 (0%)
Query: 17 SLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVAVAHAFALFVGVAIAAN 76
S+K+Y++EFIATL+FVFAGVGSAIAYN LTSDA+LDPAGLVA+A+AHAFALFVGV++AAN
Sbjct: 1 SIKSYVAEFIATLLFVFAGVGSAIAYNKLTSDASLDPAGLVAIAIAHAFALFVGVSMAAN 60
Query: 77 ISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLNYVT-AKSVPTHGVAAG 135
ISGGHLNPAVT GLA+GGNITI+TGLFYWIAQ LGSIVA LL++VT +VPTHGV+AG
Sbjct: 61 ISGGHLNPAVTLGLAVGGNITIITGLFYWIAQCLGSIVACFLLSFVTNGLAVPTHGVSAG 120
Query: 136 LNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFVVGANILAAGPFSGGSM 195
+ G+V EIIITF LVYTVYATAADPKKGS+GTIAP+AIGF+VGANILAAGPFSGGSM
Sbjct: 121 QTALQGVVMEIIITFALVYTVYATAADPKKGSLGTIAPMAIGFIVGANILAAGPFSGGSM 180
Query: 196 NPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGSYTPAPASE 246
NPARSFGPAV +G+F+ NWIYWVGPLIGGGLAGL+YGDV+I SY PASE
Sbjct: 181 NPARSFGPAVAAGDFSQNWIYWVGPLIGGGLAGLVYGDVYIASYAALPASE 231
>gb|AAM63133.1| delta tonoplast integral protein delta-TIP [Arabidopsis thaliana]
gi|9279707|dbj|BAB01264.1| delta tonoplast intrinsic
protein [Arabidopsis thaliana]
gi|32363275|sp|Q41951|TIP21_ARATH Aquaporin TIP2.1
(Tonoplast intrinsic protein 2.1) (Delta-tonoplast
intrinsic protein) (Delta-TIP) gi|1145697|gb|AAC49281.1|
delta tonoplast integral protein
gi|15233320|ref|NP_188245.1| delta tonoplast integral
protein (delta-TIP) [Arabidopsis thaliana]
Length = 250
Score = 375 bits (964), Expect = e-103
Identities = 187/250 (74%), Positives = 215/250 (85%), Gaps = 3/250 (1%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
M + FG+FDDSFS ASL+AYL+EFI+TL+FVFAGVGSAIAY LTSDAALD GLVA+A
Sbjct: 1 MAGVAFGSFDDSFSLASLRAYLAEFISTLLFVFAGVGSAIAYAKLTSDAALDTPGLVAIA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
V H FALFV VAI ANISGGH+NPAVTFGLA+GG IT++TG+FYWIAQLLGS A LL
Sbjct: 61 VCHGFALFVAVAIGANISGGHVNPAVTFGLAVGGQITVITGVFYWIAQLLGSTAACFLLK 120
Query: 121 YVTAK-SVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
YVT +VPTH VAAGL I G+V EIIITF LVYTVYATAADPKKGS+GTIAP+AIG +
Sbjct: 121 YVTGGLAVPTHSVAAGLGSIEGVVMEIIITFALVYTVYATAADPKKGSLGTIAPLAIGLI 180
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGS- 238
VGANILAAGPFSGGSMNPARSFGPAV +G+F+ +W+YWVGPLIGGGLAGLIYG+VF+GS
Sbjct: 181 VGANILAAGPFSGGSMNPARSFGPAVAAGDFSGHWVYWVGPLIGGGLAGLIYGNVFMGSS 240
Query: 239 -YTPAPASEF 247
+ P +++F
Sbjct: 241 EHVPLASADF 250
>gb|AAC39480.1| aquaporin [Vernicia fordii] gi|11276920|pir||T48886 aquaporin
[imported] - Vernicia fordii
Length = 248
Score = 374 bits (961), Expect = e-102
Identities = 187/243 (76%), Positives = 211/243 (85%), Gaps = 2/243 (0%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
M +I FG FDDSFS S KAYL+EFI+TL++VFAGVGSAIAYN LT++AALDPAGLVA+A
Sbjct: 1 MARIAFGRFDDSFSLGSFKAYLAEFISTLLYVFAGVGSAIAYNKLTANAALDPAGLVAIA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
+ H FALFV VA+ ANISGGH+NPAV FGLA+GG ITILTG+FYWIAQLLGSIVA LL
Sbjct: 61 ICHGFALFVAVAVGANISGGHVNPAVAFGLALGGQITILTGIFYWIAQLLGSIVACFLLK 120
Query: 121 YVTAK-SVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
VT ++PTH VA + I G+V EIIITF LVYTVYATAADPKKGS+GTIAPIAIGF+
Sbjct: 121 LVTGGLAIPTHSVAGRVGAIEGVVMEIIITFALVYTVYATAADPKKGSLGTIAPIAIGFI 180
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFI-GS 238
VGANILAAGPFSGGSM PARSFGPAVVSG+F DNWIYWVGPLIGGGLAGLIYG+++I G
Sbjct: 181 VGANILAAGPFSGGSMYPARSFGPAVVSGDFHDNWIYWVGPLIGGGLAGLIYGNLYISGD 240
Query: 239 YTP 241
+TP
Sbjct: 241 HTP 243
>gb|AAM10184.1| delta tonoplast intrinsic protein [Arabidopsis thaliana]
gi|17473872|gb|AAL38357.1| delta tonoplast intrinsic
protein [Arabidopsis thaliana]
Length = 250
Score = 372 bits (956), Expect = e-102
Identities = 186/250 (74%), Positives = 214/250 (85%), Gaps = 3/250 (1%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
M + FG+FDDSFS ASL+AYL+EFI+TL+FVFAGVGSAIAY LTSDAALD GLVA+A
Sbjct: 1 MAGVAFGSFDDSFSLASLRAYLAEFISTLLFVFAGVGSAIAYAKLTSDAALDTPGLVAIA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
V H FALFV VAI ANISGGH+NPAVTFGLA+GG IT++TG+FYWIAQLLGS A LL
Sbjct: 61 VCHGFALFVAVAIGANISGGHVNPAVTFGLAVGGQITVITGVFYWIAQLLGSTAACFLLK 120
Query: 121 YVTAK-SVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
YVT +VPTH VAAGL I G+V EIIITF LVYTVYATAADPKKGS+GTIAP+AIG +
Sbjct: 121 YVTGGLAVPTHSVAAGLGSIEGVVMEIIITFALVYTVYATAADPKKGSLGTIAPLAIGLI 180
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGS- 238
VGANILAAGPFSGGSMNPARSFGPAV +G+F+ +W+YWVGPLIGG LAGLIYG+VF+GS
Sbjct: 181 VGANILAAGPFSGGSMNPARSFGPAVAAGDFSGHWVYWVGPLIGGELAGLIYGNVFMGSS 240
Query: 239 -YTPAPASEF 247
+ P +++F
Sbjct: 241 EHVPLASADF 250
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.323 0.141 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 430,740,214
Number of Sequences: 2540612
Number of extensions: 18730627
Number of successful extensions: 65401
Number of sequences better than 10.0: 1466
Number of HSP's better than 10.0 without gapping: 1140
Number of HSP's successfully gapped in prelim test: 328
Number of HSP's that attempted gapping in prelim test: 60355
Number of HSP's gapped (non-prelim): 1862
length of query: 247
length of database: 863,360,394
effective HSP length: 125
effective length of query: 122
effective length of database: 545,783,894
effective search space: 66585635068
effective search space used: 66585635068
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 73 (32.7 bits)
Medicago: description of AC137077.10


