

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136507.18 - phase: 1 /pseudo
(305 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
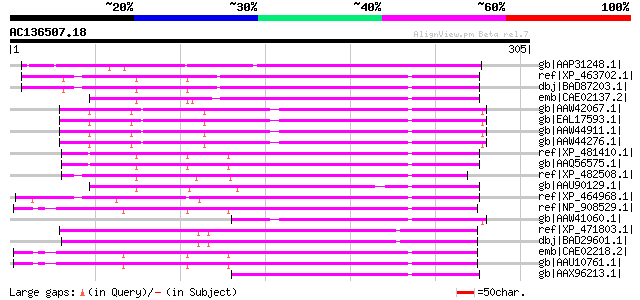
Score E
Sequences producing significant alignments: (bits) Value
gb|AAP31248.1| transposase [Fusarium oxysporum f. sp. melonis] 122 1e-26
ref|XP_463702.1| putative far-red impaired response protein [Ory... 110 7e-23
dbj|BAD87203.1| far-red impaired response-like [Oryza sativa (ja... 110 7e-23
emb|CAE02137.2| OSJNBa0074L08.5 [Oryza sativa (japonica cultivar... 109 1e-22
gb|AAW42067.1| conserved hypothetical protein [Cryptococcus neof... 103 5e-21
gb|EAL17593.1| hypothetical protein CNBM0450 [Cryptococcus neofo... 103 5e-21
gb|AAW44911.1| conserved hypothetical protein [Cryptococcus neof... 103 5e-21
gb|AAW44276.1| conserved hypothetical protein [Cryptococcus neof... 103 5e-21
ref|XP_481410.1| far-red impaired response protein -like [Oryza ... 103 8e-21
gb|AAQ56575.1| putative transposase [Oryza sativa (japonica cult... 103 8e-21
ref|XP_482508.1| putative far-red impaired response protein [Ory... 102 2e-20
gb|AAU90129.1| unknown protein [Oryza sativa (japonica cultivar-... 96 1e-18
ref|XP_464968.1| putative far-red impaired response protein [Ory... 96 2e-18
ref|NP_908529.1| P0702D12.15 [Oryza sativa (japonica cultivar-gr... 95 3e-18
gb|AAW41060.1| conserved hypothetical protein [Cryptococcus neof... 94 5e-18
ref|XP_471803.1| OSJNBa0050F15.2 [Oryza sativa (japonica cultiva... 94 6e-18
dbj|BAD29601.1| putative far-red impaired response protein [Oryz... 94 6e-18
emb|CAE02218.2| OSJNBb0002N06.8 [Oryza sativa (japonica cultivar... 93 8e-18
gb|AAU10761.1| unknown protein [Oryza sativa (japonica cultivar-... 92 1e-17
gb|AAX96213.1| transposon protein, putative, unclassified [Oryza... 92 2e-17
>gb|AAP31248.1| transposase [Fusarium oxysporum f. sp. melonis]
Length = 836
Score = 122 bits (306), Expect = 1e-26
Identities = 80/276 (28%), Positives = 139/276 (49%), Gaps = 11/276 (3%)
Query: 8 CERSGSHIVPKKKPKHAGTGSRKCGCLFMISGYQSKQTKEWRLNILNGVH--NHPMEPAL 65
C+R G+ +P + T +R+ GCLF + +S W L G H H EP+
Sbjct: 61 CDR-GAGRIPSLSDRRQTT-TRRTGCLFSVLAKESLCKTIWSLRHRPGPHFSQHNHEPSF 118
Query: 66 E--GHILAGRLKEDDKKIVRDLTKNKMLPRNILIHLKNQRPHCMTNVKQVYNERQQIWKA 123
H +L ++ V LT + P+ I L+ + + + +YN + +
Sbjct: 119 SEVAHPTLRQLSRQEEITVNQLTNAGIAPKEIGSFLRITS-NTLATQQDIYNCIAKGRRD 177
Query: 124 NRGDKKPLQYLISKLEEHKYTYYSRTQL-ESNTIEDIFWAHPTSIKLFNNFPTVLVMDST 182
+ + L +L E + ++R L ES+ + + +AHP S++ +P VL++DST
Sbjct: 178 LSKGQSNIHALADQLNEEGF--WNRICLDESSRVTAVLFAHPKSLEYLKTYPEVLILDST 235
Query: 183 YKTNIYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSS-KMNMPKVI 241
YKTN ++MP+ ++VGV + T+ + F F++ E+E +F W L+ LR + + +P VI
Sbjct: 236 YKTNRFKMPLLDIVGVDACQRTFCIAFAFLSGEEEGDFTWALQALRSVYEDHNIGLPSVI 295
Query: 242 VTDRDMSLMKAVAHVFPESYAMNCYFHVQANVKQRC 277
+TDR ++ M AV+ FP S C +H+ V+ C
Sbjct: 296 LTDRCLACMNAVSSCFPGSALFLCLWHINKAVQSYC 331
>ref|XP_463702.1| putative far-red impaired response protein [Oryza sativa (japonica
cultivar-group)]
Length = 1114
Score = 110 bits (274), Expect = 7e-23
Identities = 77/285 (27%), Positives = 125/285 (43%), Gaps = 23/285 (8%)
Query: 8 CERSGSHIVPKKKPKHAGTGSRK--------CGCLFMISGYQSKQTKEWRLNILNGVHNH 59
C R G + +KK K R C C+ + + WR+ +N HNH
Sbjct: 401 CYRYGKNDDGQKKKKKKTQQQRNTQVIDRTDCKCMLTVR----LEGDVWRILSMNLNHNH 456
Query: 60 PMEPALEGHILAGR--LKEDDKKIVRDLTKNKMLPRNILIHLKNQR------PHCMTNVK 111
+ P E L + +++KK++R L + + RN++ L R P+ +V
Sbjct: 457 TLTPNREAKFLRSHKHMTDEEKKLIRTLKQCNIPTRNMISILSFLRGGIAALPYTRKDVS 516
Query: 112 QVYNERQQIWKANRGDKKPLQYLISKLEEHKYTYYSRTQLESNTIEDIFWAHPTSIKLFN 171
V + N + ++YL K + YY T E+N ++ +FW S +L+
Sbjct: 517 NVGTAINSETR-NTDMNQVMEYLRQKEAKDPGYYYKLTLDENNKVKSMFWTDGRSTQLYE 575
Query: 172 NFPTVLVMDSTYKTNIYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLL 231
+ + D+TYKTN Y MP +VGVT F+ E E F WV + +
Sbjct: 576 QYGEFVSFDTTYKTNRYNMPFAPIVGVTGHGNICIFACAFLGDETTETFKWVFETFLTAM 635
Query: 232 SSKMNMPKVIVTDRDMSLMKAVAHVFPESYAMNCYFHVQANVKQR 276
K P+ I+TD+D+++ A+ VFP S NC FH+ ++R
Sbjct: 636 GGK--HPETIITDQDLAMRAAIRQVFPNSKHRNCLFHILKKCRER 678
>dbj|BAD87203.1| far-red impaired response-like [Oryza sativa (japonica
cultivar-group)]
Length = 1130
Score = 110 bits (274), Expect = 7e-23
Identities = 77/285 (27%), Positives = 125/285 (43%), Gaps = 23/285 (8%)
Query: 8 CERSGSHIVPKKKPKHAGTGSRK--------CGCLFMISGYQSKQTKEWRLNILNGVHNH 59
C R G + +KK K R C C+ + + WR+ +N HNH
Sbjct: 417 CYRYGKNDDGQKKKKKKTQQQRNTQVIDRTDCKCMLTVR----LEGDVWRILSMNLNHNH 472
Query: 60 PMEPALEGHILAGR--LKEDDKKIVRDLTKNKMLPRNILIHLKNQR------PHCMTNVK 111
+ P E L + +++KK++R L + + RN++ L R P+ +V
Sbjct: 473 TLTPNREAKFLRSHKHMTDEEKKLIRTLKQCNIPTRNMISILSFLRGGIAALPYTRKDVS 532
Query: 112 QVYNERQQIWKANRGDKKPLQYLISKLEEHKYTYYSRTQLESNTIEDIFWAHPTSIKLFN 171
V + N + ++YL K + YY T E+N ++ +FW S +L+
Sbjct: 533 NVGTAINSETR-NTDMNQVMEYLRQKEAKDPGYYYKLTLDENNKVKSMFWTDGRSTQLYE 591
Query: 172 NFPTVLVMDSTYKTNIYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLL 231
+ + D+TYKTN Y MP +VGVT F+ E E F WV + +
Sbjct: 592 QYGEFVSFDTTYKTNRYNMPFAPIVGVTGHGNICIFACAFLGDETTETFKWVFETFLTAM 651
Query: 232 SSKMNMPKVIVTDRDMSLMKAVAHVFPESYAMNCYFHVQANVKQR 276
K P+ I+TD+D+++ A+ VFP S NC FH+ ++R
Sbjct: 652 GGK--HPETIITDQDLAMRAAIRQVFPNSKHRNCLFHILKKCRER 694
>emb|CAE02137.2| OSJNBa0074L08.5 [Oryza sativa (japonica cultivar-group)]
gi|50926620|ref|XP_473257.1| OSJNBa0074L08.5 [Oryza
sativa (japonica cultivar-group)]
Length = 1061
Score = 109 bits (272), Expect = 1e-22
Identities = 69/240 (28%), Positives = 113/240 (46%), Gaps = 17/240 (7%)
Query: 48 WRLNILNGVHNHPMEPALEGHILAGR--LKEDDKKIVRDLTKNKMLPRNILIHLKNQR-- 103
WR+ +N HNH + P E L + +++KK++R L + + RN++ L R
Sbjct: 392 WRILSMNLNHNHTLTPNREAKFLRSHKHMTDEEKKLIRTLKQCNIPTRNMISILSFLRGG 451
Query: 104 ----PHC---MTNVKQVYNERQQIWKANRGDKKPLQYLISKLEEHKYTYYSRTQLESNTI 156
P+ ++NV N + N + ++YL K + YY T E+N +
Sbjct: 452 IAALPYTGKDVSNVGTAINSETR----NTDMNQVMEYLRQKEAKDPGYYYKLTLDENNKV 507
Query: 157 EDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMPMFEVVGVTSTDLTYSVGFGFMTHEK 216
+ +FW S +L+ + + D+TYKTN Y MP +VGVT F+ E
Sbjct: 508 KSMFWTDGRSTQLYEQYGEFVSFDTTYKTNRYNMPFAPIVGVTGHGNICIFACAFLGDET 567
Query: 217 EENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYAMNCYFHVQANVKQR 276
E F WV + + K P+ I+TD+D+++ A+ VFP S NC FH+ ++R
Sbjct: 568 TETFKWVFETFLTAMGGK--HPETIITDQDLAMRAAIRQVFPNSKHRNCLFHILKKCRER 625
>gb|AAW42067.1| conserved hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21] gi|50260537|gb|EAL23192.1|
hypothetical protein CNBA5360 [Cryptococcus neoformans
var. neoformans B-3501A] gi|50259285|gb|EAL21958.1|
hypothetical protein CNBC0980 [Cryptococcus neoformans
var. neoformans B-3501A] gi|50256296|gb|EAL19021.1|
hypothetical protein CNBH1230 [Cryptococcus neoformans
var. neoformans B-3501A] gi|57229113|gb|AAW45547.1|
conserved hypothetical protein [Cryptococcus neoformans
var. neoformans JEC21] gi|58271396|ref|XP_572854.1|
conserved hypothetical protein [Cryptococcus neoformans
var. neoformans JEC21] gi|58264436|ref|XP_569374.1|
conserved hypothetical protein [Cryptococcus neoformans
var. neoformans JEC21]
Length = 932
Score = 103 bits (258), Expect = 5e-21
Identities = 75/268 (27%), Positives = 125/268 (45%), Gaps = 25/268 (9%)
Query: 30 KCGCLFMISGYQSKQT--------KEWRLNILNGVHNHPMEPALEGHIL--AGRLKEDDK 79
K GC F ++ + K W ++N HNHP + +L A R E K
Sbjct: 169 KNGCPFAAVAWEMAEAEVKEHGGDKCWTWKVVNASHNHPPVANIGIALLRRASRTPEA-K 227
Query: 80 KIVRDLTKNKMLPRNILIHLKNQRPHCMTNVKQV----YNERQQIWKANRGDKKPLQYLI 135
+++ L ++ +I+ + + P M + + + R + + ++ L+
Sbjct: 228 ELINTLIDSRTPFPSIISTMSQRFPTVMLTSEDLHRLAFRRRADLRAGSSASAACIRQLV 287
Query: 136 SKLEEHKYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMPMFEV 195
S E H+ +R +L + + T+ L + FP VL D TYKTN+YRMPM +
Sbjct: 288 SNDELHRPFVNTRDELIG-----LVYTKRTARALIHRFPEVLFFDCTYKTNLYRMPMLHI 342
Query: 196 VGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLMKAVAH 255
VG TST +TY+ G M E + L +L+ KV++TDRD +L+ A+
Sbjct: 343 VGSTSTGMTYTAGVILMLRETTNWYTQALNSFFELVGKP--DVKVVITDRDPALINALMS 400
Query: 256 VFPESYAMNCYFHVQANVKQR---CVLD 280
V P++Y +C++H+Q NVK C +D
Sbjct: 401 VLPKAYRFSCFWHLQENVKSNIRPCFVD 428
>gb|EAL17593.1| hypothetical protein CNBM0450 [Cryptococcus neoformans var.
neoformans B-3501A] gi|57226908|gb|AAW43368.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21] gi|58267038|ref|XP_570675.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21]
Length = 580
Score = 103 bits (258), Expect = 5e-21
Identities = 75/268 (27%), Positives = 125/268 (45%), Gaps = 25/268 (9%)
Query: 30 KCGCLFMISGYQSKQT--------KEWRLNILNGVHNHPMEPALEGHIL--AGRLKEDDK 79
K GC F ++ + K W ++N HNHP + +L A R E K
Sbjct: 169 KNGCPFAAVAWEMAEAEVKEHGGDKCWTWKVVNASHNHPPVANIGIALLRRASRTPEA-K 227
Query: 80 KIVRDLTKNKMLPRNILIHLKNQRPHCMTNVKQV----YNERQQIWKANRGDKKPLQYLI 135
+++ L ++ +I+ + + P M + + + R + + ++ L+
Sbjct: 228 ELINTLIDSRTPFPSIISTMSQRFPTVMLTSEDLHRLAFRRRADLRAGSSASAACIRQLV 287
Query: 136 SKLEEHKYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMPMFEV 195
S E H+ +R +L + + T+ L + FP VL D TYKTN+YRMPM +
Sbjct: 288 SNDELHRPFVNTRDELIG-----LVYTKRTARALIHRFPEVLFFDCTYKTNLYRMPMLHI 342
Query: 196 VGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLMKAVAH 255
VG TST +TY+ G M E + L +L+ KV++TDRD +L+ A+
Sbjct: 343 VGSTSTGMTYTAGVILMLRETTNWYTQALNSFFELVGKP--DVKVVITDRDPALINALMS 400
Query: 256 VFPESYAMNCYFHVQANVKQR---CVLD 280
V P++Y +C++H+Q NVK C +D
Sbjct: 401 VLPKAYRFSCFWHLQENVKSNIRPCFVD 428
>gb|AAW44911.1| conserved hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21] gi|58270124|ref|XP_572218.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21]
Length = 920
Score = 103 bits (258), Expect = 5e-21
Identities = 75/268 (27%), Positives = 125/268 (45%), Gaps = 25/268 (9%)
Query: 30 KCGCLFMISGYQSKQT--------KEWRLNILNGVHNHPMEPALEGHIL--AGRLKEDDK 79
K GC F ++ + K W ++N HNHP + +L A R E K
Sbjct: 169 KNGCPFAAVAWEMAEAEVKEHGGDKCWTWKVVNASHNHPPVANIGIALLRRASRTPEA-K 227
Query: 80 KIVRDLTKNKMLPRNILIHLKNQRPHCMTNVKQV----YNERQQIWKANRGDKKPLQYLI 135
+++ L ++ +I+ + + P M + + + R + + ++ L+
Sbjct: 228 ELINTLIDSRTPFPSIISTMSQRFPTVMLTSEDLHRLAFRRRADLRAGSSASAACIRQLV 287
Query: 136 SKLEEHKYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMPMFEV 195
S E H+ +R +L + + T+ L + FP VL D TYKTN+YRMPM +
Sbjct: 288 SNDELHRPFVNTRDELIG-----LVYTKRTARALIHRFPEVLFFDCTYKTNLYRMPMLHI 342
Query: 196 VGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLMKAVAH 255
VG TST +TY+ G M E + L +L+ KV++TDRD +L+ A+
Sbjct: 343 VGSTSTGMTYTAGVILMLRETTNWYTQALNSFFELVGKP--DVKVVITDRDPALINALMS 400
Query: 256 VFPESYAMNCYFHVQANVKQR---CVLD 280
V P++Y +C++H+Q NVK C +D
Sbjct: 401 VLPKAYRFSCFWHLQENVKSNIRPCFVD 428
>gb|AAW44276.1| conserved hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21] gi|58268854|ref|XP_571583.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21]
Length = 895
Score = 103 bits (258), Expect = 5e-21
Identities = 75/268 (27%), Positives = 125/268 (45%), Gaps = 25/268 (9%)
Query: 30 KCGCLFMISGYQSKQT--------KEWRLNILNGVHNHPMEPALEGHIL--AGRLKEDDK 79
K GC F ++ + K W ++N HNHP + +L A R E K
Sbjct: 169 KNGCPFAAVAWEMAEAEVKEHGGDKCWTWKVVNASHNHPPVANIGIALLRRASRTPEA-K 227
Query: 80 KIVRDLTKNKMLPRNILIHLKNQRPHCMTNVKQV----YNERQQIWKANRGDKKPLQYLI 135
+++ L ++ +I+ + + P M + + + R + + ++ L+
Sbjct: 228 ELINTLIDSRTPFPSIISTMSQRFPTVMLTSEDLHRLAFRRRADLRAGSSASAACIRQLV 287
Query: 136 SKLEEHKYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMPMFEV 195
S E H+ +R +L + + T+ L + FP VL D TYKTN+YRMPM +
Sbjct: 288 SNDELHRPFVNTRDELIG-----LVYTKRTARALIHRFPEVLFFDCTYKTNLYRMPMLHI 342
Query: 196 VGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLMKAVAH 255
VG TST +TY+ G M E + L +L+ KV++TDRD +L+ A+
Sbjct: 343 VGSTSTGMTYTAGVILMLRETTNWYTQALNSFFELVGKP--DVKVVITDRDPALINALMS 400
Query: 256 VFPESYAMNCYFHVQANVKQR---CVLD 280
V P++Y +C++H+Q NVK C +D
Sbjct: 401 VLPKAYRFSCFWHLQENVKSNIRPCFVD 428
>ref|XP_481410.1| far-red impaired response protein -like [Oryza sativa (japonica
cultivar-group)] gi|35215248|dbj|BAC92598.1| putative
far-red impaired response protein [Oryza sativa
(japonica cultivar-group)] gi|35215057|dbj|BAC92415.1|
putative far-red impaired response protein [Oryza sativa
(japonica cultivar-group)]
Length = 1132
Score = 103 bits (256), Expect = 8e-21
Identities = 71/253 (28%), Positives = 114/253 (44%), Gaps = 10/253 (3%)
Query: 31 CGCLFMISGYQSKQTKEWRLNILNGVHNHPMEPALEGHILAGR--LKEDDKKIVRDLTKN 88
C C+ M+ + W++ L+ HNH + P E L + +K+++R L +
Sbjct: 479 CKCMMMVEKMFALGDV-WQIATLDLKHNHALCPRREAKFLKSHKNMTIKEKRLIRTLKEC 537
Query: 89 KMLPRNILIHLKNQR---PHCMTNVKQVYNERQQIWKANRGD--KKPLQYLISKLEEHKY 143
+ R++++ L R P K V N I R + K+ L YL K E
Sbjct: 538 NIPTRSMIVILSFLRGGLPALPYTKKDVSNVGTAINSETRNNDMKQVLAYLRKKEIEDPG 597
Query: 144 TYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMPMFEVVGVTSTDL 203
Y E+N + +FW S +L+ + + D+TY+TN Y MP VGV
Sbjct: 598 ISYKFKLDENNKVTSMFWTDGRSTQLYEEYGDCISFDTTYRTNRYNMPFAPFVGVIGHGT 657
Query: 204 TYSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYAM 263
T G F+ E E F WV + + K PK I+TD+D ++ A+A VF +
Sbjct: 658 TCLFGCAFLGDETAETFKWVFETFATAMGGK--HPKTIITDQDNAMRSAIAQVFQNTKHR 715
Query: 264 NCYFHVQANVKQR 276
NC FH++ N +++
Sbjct: 716 NCLFHIKKNCREK 728
>gb|AAQ56575.1| putative transposase [Oryza sativa (japonica cultivar-group)]
Length = 1037
Score = 103 bits (256), Expect = 8e-21
Identities = 71/253 (28%), Positives = 114/253 (44%), Gaps = 10/253 (3%)
Query: 31 CGCLFMISGYQSKQTKEWRLNILNGVHNHPMEPALEGHILAGR--LKEDDKKIVRDLTKN 88
C C+ M+ + W++ L+ HNH + P E L + +K+++R L +
Sbjct: 384 CKCMMMVEKMFALGDV-WQIATLDLKHNHALCPRREAKFLKSHKNMTIKEKRLIRTLKEC 442
Query: 89 KMLPRNILIHLKNQR---PHCMTNVKQVYNERQQIWKANRGD--KKPLQYLISKLEEHKY 143
+ R++++ L R P K V N I R + K+ L YL K E
Sbjct: 443 NIPTRSMIVILSFLRGGLPALPYTKKDVSNVGTAINSETRNNDMKQVLAYLRKKEIEDPG 502
Query: 144 TYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMPMFEVVGVTSTDL 203
Y E+N + +FW S +L+ + + D+TY+TN Y MP VGV
Sbjct: 503 ISYKFKLDENNKVTSMFWTDGRSTQLYEEYGDCISFDTTYRTNRYNMPFAPFVGVIGHGT 562
Query: 204 TYSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYAM 263
T G F+ E E F WV + + K PK I+TD+D ++ A+A VF +
Sbjct: 563 TCLFGCAFLGDETAETFKWVFETFATAMGGK--HPKTIITDQDNAMRSAIAQVFQNTKHR 620
Query: 264 NCYFHVQANVKQR 276
NC FH++ N +++
Sbjct: 621 NCLFHIKKNCREK 633
>ref|XP_482508.1| putative far-red impaired response protein [Oryza sativa (japonica
cultivar-group)] gi|25553698|dbj|BAC24942.1| putative
far-red impaired response protein [Oryza sativa
(japonica cultivar-group)]
Length = 1148
Score = 102 bits (253), Expect = 2e-20
Identities = 66/246 (26%), Positives = 116/246 (46%), Gaps = 13/246 (5%)
Query: 31 CGCLFMISGYQSKQTKEWRLNILNGVHNHPMEPALEGHILAGR--LKEDDKKIVRDLTKN 88
C C +IS ++ + W ++ + HNH + P E L + ++K ++R L +
Sbjct: 446 CKCALVIS----ERNEIWNISRVQLDHNHQLSPRDEVRFLKSHKHMTTEEKMLIRTLKEC 501
Query: 89 KMLPRNILIHLKNQRPHCMT---NVKQVYNERQQIWKANRGDK--KPLQYLISKLEEHKY 143
+ R++++ L R + K + N R I K + K L++ K E+ +
Sbjct: 502 NIPTRHMIVILSVLRGGLTSLPYTKKDISNVRTTINKETSSNDIMKTLEFFRRKKEKDPH 561
Query: 144 TYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMPMFEVVGVTSTDL 203
+Y ES ++++FW S + + + V+ D+TY TN Y +P VG+
Sbjct: 562 FFYEFDLDESKKVKNLFWTDGRSREWYEKYGDVVSFDTTYFTNKYNLPFAPFVGIYGHGN 621
Query: 204 TYSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYAM 263
T G F+ E E F W+ + K +S K PK I+TD+D ++ A+A VF +
Sbjct: 622 TIVFGCAFLHDETSETFKWLFRTFLKAMSQK--EPKTIITDQDGAMRSAIAQVFQNAKHR 679
Query: 264 NCYFHV 269
NC+FH+
Sbjct: 680 NCFFHI 685
>gb|AAU90129.1| unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 904
Score = 96.3 bits (238), Expect = 1e-18
Identities = 62/236 (26%), Positives = 110/236 (46%), Gaps = 14/236 (5%)
Query: 48 WRLNILNGVHNHPMEPALEGHILAGR--LKEDDKKIVRDLTKNKMLPRNILIHLKNQRP- 104
W + LN HNHP+ P E L + + +K ++R L K + R I+ L R
Sbjct: 431 WYITRLNLEHNHPLSPPEERKFLWSHKHMIDQEKLLIRTLNKINVPTRMIMSVLSYVRGG 490
Query: 105 -HCMTNVKQVYNERQQIWKANRGDKKPLQ---YLISKLEEHKYTYYSRTQLESNTIEDIF 160
+ K+ + + + G +Q + K+ E Y+ E+N ++ +F
Sbjct: 491 LFAVPYTKKAMSNYRDFVRRESGKNDMMQCLDFFEKKISEDPLFYFRFRTDENNVVKSLF 550
Query: 161 WAHPTSIKLFNNFPTVLVMDSTYKTNIYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENF 220
W+ S K + F ++ D+TYKTN Y +P VG+TS G+ F+ E
Sbjct: 551 WSDRNSRKFYEMFGDIVSFDTTYKTNRYDLPFAPFVGITSHGDNCLFGYAFLQDE----- 605
Query: 221 VWVLKMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYAMNCYFHVQANVKQR 276
V++ + + K +P I+TD+D+++ A+ VFP++ NC FH+ +N + +
Sbjct: 606 TMVVQYVLDCMGGK--VPATIITDQDLAMKAAIVIVFPDTVHRNCMFHMLSNARDK 659
>ref|XP_464968.1| putative far-red impaired response protein [Oryza sativa (japonica
cultivar-group)] gi|47848338|dbj|BAD22200.1| putative
far-red impaired response protein [Oryza sativa
(japonica cultivar-group)] gi|47847707|dbj|BAD21486.1|
putative far-red impaired response protein [Oryza sativa
(japonica cultivar-group)]
Length = 934
Score = 95.5 bits (236), Expect = 2e-18
Identities = 70/287 (24%), Positives = 129/287 (44%), Gaps = 21/287 (7%)
Query: 4 FRLVCERSG------SHIVPKKKPKHAGTGSRKCGCLFMISGYQSKQTKEWRLNILNGVH 57
F L C +SG S P KK + +C ++ EW + H
Sbjct: 314 FHLECNKSGKMKLTKSSENPMKKRRSNLVEKTQCKARVIVK----LDKGEWEFTAVRHEH 369
Query: 58 NHPM--EPALEGHILAGR-LKEDDKKIVRDLTKNKMLPRNILIHLKNQRPHCMTNV---- 110
NHP+ P L I+ + + +K +R L +N++ P+ I+ + R C ++
Sbjct: 370 NHPLCPSPLLARFIVDHKQMSTGEKSFLRVLQQNRVPPKKIMKIFRKLRV-CFGDIPFEN 428
Query: 111 KQVYNERQ-QIWKANRGDKKPLQYLISKLEEHKYTYYSRTQLESNTIEDIFWAHPTSIKL 169
K +N Q + KAN + L++ ++ Y + E NT+ IFW
Sbjct: 429 KDEHNIAQTEHRKANSDVESALKHFTELQIQNPEFLYVMQKDEDNTVTSIFWTDARLRIE 488
Query: 170 FNNFPTVLVMDSTYKTNIYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRK 229
++ F +++ D+ Y T++Y MP ++G+ S + +G + EK E F W+L+ +
Sbjct: 489 YDIFGDLIMFDAAYSTDMYNMPFVPIIGINSHATPFLLGCALLKDEKVETFEWMLRTFLQ 548
Query: 230 LLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYAMNCYFHVQANVKQR 276
++ K MP+ ++T++D S+ KA A + P C HV + +++
Sbjct: 549 VMGGK--MPRAVITNQDTSMEKAFAELMPHVRLRFCKRHVMSKAQEK 593
>ref|NP_908529.1| P0702D12.15 [Oryza sativa (japonica cultivar-group)]
Length = 832
Score = 94.7 bits (234), Expect = 3e-18
Identities = 72/282 (25%), Positives = 127/282 (44%), Gaps = 19/282 (6%)
Query: 3 MFRLVCERSGSHIVPK-KKPKHAGTGSRKCGCLFMISGYQSKQTKEWRLNILNGVHNHPM 61
M ++ C G + PK KKP S + GC M+ +S+ W + + HNHPM
Sbjct: 113 MRQICCSHQGRN--PKTKKP------SVRIGCPAMMKINRSRAGSGWSVTKVVSAHNHPM 164
Query: 62 EPAL---EGHILAGRLKEDDKKIVRDLTKNKMLPRNILIHLKNQR--PHCMTNVKQVYNE 116
+ ++ + + ++ E + I+ ++ + M N+ L P + ++ +
Sbjct: 165 KKSVGVTKNYQSHNQIDEGTRGIIEEMVDSSMSLTNMYGMLSGMHGGPSMVPFTRKAMDR 224
Query: 117 RQQIWKANRGD---KKPLQYLISKLEEHKYTYYSRTQLESNTIEDIFWAHPTSIKLFNNF 173
+ + +K L L + K +YS E+ +++IFW+H S F +F
Sbjct: 225 LAYAIRRDESSDDMQKTLDVLKDLQKRSKNFFYSIQVDEACRVKNIFWSHAVSRLNFEHF 284
Query: 174 PTVLVMDSTYKTNIYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSS 233
V+ D+TYKTN Y MP VGV + + G + E EE+F W+ ++ ++
Sbjct: 285 GDVITFDTTYKTNKYNMPFAPFVGVNNHFQSTFFGCALLREETEESFTWLFNTFKECMNG 344
Query: 234 KMNMPKVIVTDRDMSLMKAVAHVFPESYAMNCYFHVQANVKQ 275
K +P I+TD S+ A+ VFP + C +HV K+
Sbjct: 345 K--VPIGILTDNCPSMAAAIRTVFPNTIHRVCKWHVLKKAKE 384
>gb|AAW41060.1| conserved hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21] gi|58258933|ref|XP_566879.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21]
Length = 678
Score = 94.0 bits (232), Expect = 5e-18
Identities = 54/153 (35%), Positives = 83/153 (53%), Gaps = 10/153 (6%)
Query: 131 LQYLISKLEEHKYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRM 190
++ L+S E H+ +R +L + + T+ L + FP VL D TYKTN+YRM
Sbjct: 29 IRQLVSNDELHRPFVNTRDELIG-----LVYTKRTARALIHRFPEVLFFDCTYKTNLYRM 83
Query: 191 PMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLM 250
PM +VG TST +TY+ G M E + L +L+ KV++TDRD +L+
Sbjct: 84 PMLHIVGSTSTGMTYTAGVILMLRETTNWYTQALNSFFELVGKP--DVKVVITDRDPALI 141
Query: 251 KAVAHVFPESYAMNCYFHVQANVKQR---CVLD 280
A+ V P++Y +C++H+Q NVK C +D
Sbjct: 142 NALMSVLPKAYRFSCFWHLQENVKSNIRPCFVD 174
>ref|XP_471803.1| OSJNBa0050F15.2 [Oryza sativa (japonica cultivar-group)]
gi|38345091|emb|CAD40514.2| OSJNBa0050F15.2 [Oryza
sativa (japonica cultivar-group)]
Length = 688
Score = 93.6 bits (231), Expect = 6e-18
Identities = 63/254 (24%), Positives = 119/254 (46%), Gaps = 10/254 (3%)
Query: 30 KCGCLFMISGYQSKQTKEWRLNILNGVHNHPM-EPALEGHILAGRLKEDDKKIVR-DLTK 87
+CGCL + ++++ W + + H H + P + R+ D KK +L
Sbjct: 87 RCGCLAKLEIERNEEKGVWFVKEFDNQHTHELANPDHVAFLGVHRVMSDSKKAQAVELRM 146
Query: 88 NKMLPRNILIHLKNQRPHCMTN---VKQVYN--ERQQIWKANRGDKKP-LQYLISKLEEH 141
+ + P ++ ++N +K +YN R + D + L+YL K EE
Sbjct: 147 SGLRPFQVMEVMENNHDELDEVGFVMKDLYNFFTRYDMKNIKGHDAEDVLKYLTRKQEED 206
Query: 142 KYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMPMFEVVGVTST 201
++ T E + ++FWA S + F V++ DSTY+ N Y +P +GV
Sbjct: 207 AEFFFKYTTDEEGRLRNVFWADAESRLDYAAFGGVVIFDSTYRVNKYNLPFIPFIGVNHH 266
Query: 202 DLTYSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESY 261
T G G +++E ++ W+L+ L + + PK ++TD D+++ KA++ V P +Y
Sbjct: 267 RSTTIFGCGILSNESVNSYCWLLETF--LEAMRQVHPKSLITDGDLAMAKAISKVMPGAY 324
Query: 262 AMNCYFHVQANVKQ 275
C +H++ N+ +
Sbjct: 325 HRLCTWHIEENMSR 338
>dbj|BAD29601.1| putative far-red impaired response protein [Oryza sativa (japonica
cultivar-group)] gi|50251625|dbj|BAD29488.1| putative
far-red impaired response protein [Oryza sativa
(japonica cultivar-group)]
Length = 688
Score = 93.6 bits (231), Expect = 6e-18
Identities = 63/253 (24%), Positives = 119/253 (46%), Gaps = 10/253 (3%)
Query: 31 CGCLFMISGYQSKQTKEWRLNILNGVHNHPM-EPALEGHILAGRLKEDDKKIVR-DLTKN 88
CGCL + ++++ W + + H H + P + R+ D KK +L +
Sbjct: 88 CGCLAKLEIERNEEKGVWFVKEFDNQHTHELANPDHVAFLGVHRVMSDSKKAQAVELRMS 147
Query: 89 KMLPRNILIHLKNQRPHCMTN---VKQVYN--ERQQIWKANRGDKKP-LQYLISKLEEHK 142
+ P ++ ++N +K +YN R ++ D + L+YL K EE
Sbjct: 148 GLRPFQVMEVMENNHDELDEVGFVMKDLYNFFTRYEMKNIKGHDAEDVLKYLTRKQEEDA 207
Query: 143 YTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMPMFEVVGVTSTD 202
++ T E + ++FWA S + F V++ DSTY+ N Y +P +GV
Sbjct: 208 EFFFKYTTDEEGRLRNVFWADAESRLDYAAFGGVVIFDSTYRVNKYNLPFIPFIGVNHHR 267
Query: 203 LTYSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYA 262
T G G +++E ++ W+L+ L + + PK ++TD D+++ KA++ V P +Y
Sbjct: 268 STTIFGCGILSNESVNSYCWLLETF--LEAMRQVHPKSLITDGDLAMAKAISKVMPGAYH 325
Query: 263 MNCYFHVQANVKQ 275
C +H++ N+ +
Sbjct: 326 RLCTWHIEENMSR 338
>emb|CAE02218.2| OSJNBb0002N06.8 [Oryza sativa (japonica cultivar-group)]
gi|50923349|ref|XP_472035.1| OSJNBb0002N06.8 [Oryza
sativa (japonica cultivar-group)]
Length = 885
Score = 93.2 bits (230), Expect = 8e-18
Identities = 72/282 (25%), Positives = 126/282 (44%), Gaps = 19/282 (6%)
Query: 3 MFRLVCERSGSHIVPK-KKPKHAGTGSRKCGCLFMISGYQSKQTKEWRLNILNGVHNHPM 61
M ++ C G + PK KKP S + GC M+ +S W + + HNHPM
Sbjct: 166 MRQICCSHQGRN--PKTKKP------SVRIGCPAMMKINRSGAGSGWSVTKVVSTHNHPM 217
Query: 62 EPAL---EGHILAGRLKEDDKKIVRDLTKNKMLPRNILIHLKNQR--PHCMTNVKQVYNE 116
+ ++ + + ++ E + I+ ++ + M N+ L P + ++ +
Sbjct: 218 KKSVGVTKNYQSHNQIDEGTRGIIEEMVDSSMSLTNMYGMLSGMHGGPSMVPFTRKAMDR 277
Query: 117 RQQIWKANRGD---KKPLQYLISKLEEHKYTYYSRTQLESNTIEDIFWAHPTSIKLFNNF 173
+ + +K L L + K +YS E+ +++IFW+H S F +F
Sbjct: 278 LAYAIRRDESSDDMQKTLDVLKDLQKRSKNFFYSIQVDEACRVKNIFWSHAVSRLNFEHF 337
Query: 174 PTVLVMDSTYKTNIYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSS 233
V+ D+TYKTN Y MP VGV + + G + E EE+F W+ ++ ++
Sbjct: 338 GDVITFDTTYKTNKYNMPFAPFVGVNNHFQSTFFGCALLREETEESFTWLFNTFKECMNG 397
Query: 234 KMNMPKVIVTDRDMSLMKAVAHVFPESYAMNCYFHVQANVKQ 275
K +P I+TD S+ A+ VFP + C +HV K+
Sbjct: 398 K--VPIGILTDNCPSMAAAIRTVFPNTIHRVCKWHVLKKAKE 437
>gb|AAU10761.1| unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 822
Score = 92.4 bits (228), Expect = 1e-17
Identities = 72/282 (25%), Positives = 125/282 (43%), Gaps = 19/282 (6%)
Query: 3 MFRLVCERSGSHIVPK-KKPKHAGTGSRKCGCLFMISGYQSKQTKEWRLNILNGVHNHPM 61
M ++ C G + PK KKP S + GC M+ S W + + HNHPM
Sbjct: 166 MRQICCSHQGRN--PKTKKP------SVRIGCPAMMKINMSGAGSGWSVTKVVSAHNHPM 217
Query: 62 EPAL---EGHILAGRLKEDDKKIVRDLTKNKMLPRNILIHLKNQR--PHCMTNVKQVYNE 116
+ ++ + + ++ E + I+ ++ + M N+ L P + ++ +
Sbjct: 218 KKSVGVTKNYQSHNQIDEGTRGIIEEMVDSSMSLTNMYGMLSGMHGGPSMVPFTRKAMDR 277
Query: 117 RQQIWKANRGD---KKPLQYLISKLEEHKYTYYSRTQLESNTIEDIFWAHPTSIKLFNNF 173
+ + +K L L + K +YS E+ +++IFW+H S F +F
Sbjct: 278 VAYAIRRDESSDDMQKTLDVLKDLQKRSKNFFYSIQVDEACRVKNIFWSHAVSRLNFEHF 337
Query: 174 PTVLVMDSTYKTNIYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSS 233
V+ D+TYKTN Y MP VGV + + G + E EE+F W+ ++ ++
Sbjct: 338 GDVITFDTTYKTNKYNMPFAPFVGVNNHFQSTFFGCALLREETEESFTWLFNTFKECMNG 397
Query: 234 KMNMPKVIVTDRDMSLMKAVAHVFPESYAMNCYFHVQANVKQ 275
K +P I+TD S+ A+ VFP + C +HV K+
Sbjct: 398 K--VPIGILTDNCPSMAAAIRTVFPNTIHRVCKWHVLKKAKE 437
>gb|AAX96213.1| transposon protein, putative, unclassified [Oryza sativa (japonica
cultivar-group)]
Length = 1318
Score = 92.0 bits (227), Expect = 2e-17
Identities = 44/146 (30%), Positives = 75/146 (51%), Gaps = 2/146 (1%)
Query: 131 LQYLISKLEEHKYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRM 190
L + K E Y+ E+N ++ +FW+ + K + F ++ D+ YKTN Y +
Sbjct: 739 LDFFEKKRSEDPLFYFRFRTDENNVVKSLFWSDGNNRKFYEMFDDIVSFDTMYKTNRYDL 798
Query: 191 PMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLM 250
P VG+TS G+ F+ E E F W+ K + K +P I+TD+D+++
Sbjct: 799 PFAPFVGITSHGDNCLFGYAFLQDETSETFQWLFKTFLDCMGGK--VPATIITDQDLAMK 856
Query: 251 KAVAHVFPESYAMNCYFHVQANVKQR 276
A+A VFP++ NC FH+ +N + +
Sbjct: 857 AAIAIVFPDTVHRNCLFHMLSNARDK 882
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.321 0.136 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 513,503,343
Number of Sequences: 2540612
Number of extensions: 20313063
Number of successful extensions: 46068
Number of sequences better than 10.0: 429
Number of HSP's better than 10.0 without gapping: 297
Number of HSP's successfully gapped in prelim test: 132
Number of HSP's that attempted gapping in prelim test: 45537
Number of HSP's gapped (non-prelim): 472
length of query: 305
length of database: 863,360,394
effective HSP length: 127
effective length of query: 178
effective length of database: 540,702,670
effective search space: 96245075260
effective search space used: 96245075260
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 75 (33.5 bits)
Medicago: description of AC136507.18


