

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136507.1 - phase: 0
(236 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
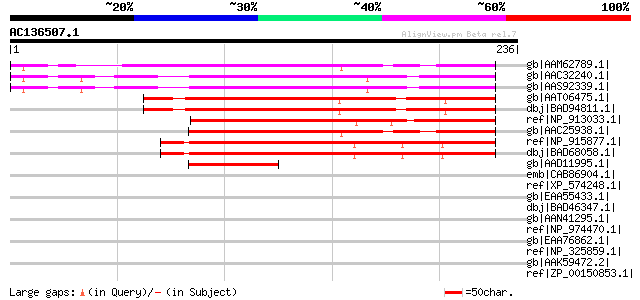
Score E
Sequences producing significant alignments: (bits) Value
gb|AAM62789.1| unknown [Arabidopsis thaliana] gi|20465913|gb|AAM... 158 1e-37
gb|AAC32240.1| unknown protein [Arabidopsis thaliana] gi|7459608... 152 7e-36
gb|AAS92339.1| At2g26850 [Arabidopsis thaliana] gi|45773808|gb|A... 152 7e-36
gb|AAT06475.1| At2g41170 [Arabidopsis thaliana] gi|30688559|ref|... 150 4e-35
dbj|BAD94811.1| hypothetical protein [Arabidopsis thaliana] 148 1e-34
ref|NP_913033.1| unnamed protein product [Oryza sativa (japonica... 143 5e-33
gb|AAC25938.1| unknown protein [Arabidopsis thaliana] gi|7487431... 141 1e-32
ref|NP_915877.1| P0702B09.33 [Oryza sativa (japonica cultivar-gr... 136 4e-31
dbj|BAD68058.1| F-box family protein-like [Oryza sativa (japonic... 136 4e-31
gb|AAD11995.1| unknown protein [Arabidopsis thaliana] gi|7459607... 54 4e-06
emb|CAB86904.1| putative protein [Arabidopsis thaliana] gi|15231... 45 0.001
ref|XP_574248.1| PREDICTED: similar to F-box only protein 31 [Ra... 39 0.16
gb|EAA55433.1| hypothetical protein MG09240.4 [Magnaporthe grise... 38 0.21
dbj|BAD46347.1| unknown protein [Oryza sativa (japonica cultivar... 38 0.27
gb|AAN41295.1| unknown protein [Arabidopsis thaliana] gi|8388622... 37 0.35
ref|NP_974470.1| F-box family protein / lectin-related [Arabidop... 37 0.35
gb|EAA76862.1| hypothetical protein FG07514.1 [Gibberella zeae P... 37 0.60
ref|NP_325859.1| ABC TRANSPORTER PERMEASE PROTEIN [Mycoplasma pu... 36 0.78
gb|AAK59472.2| unknown protein [Arabidopsis thaliana] 36 1.0
ref|ZP_00150853.1| COG0525: Valyl-tRNA synthetase [Dechloromonas... 36 1.0
>gb|AAM62789.1| unknown [Arabidopsis thaliana] gi|20465913|gb|AAM20109.1| unknown
protein [Arabidopsis thaliana]
gi|18491203|gb|AAL69504.1| unknown protein [Arabidopsis
thaliana] gi|30685347|ref|NP_850191.1| F-box family
protein [Arabidopsis thaliana]
Length = 371
Score = 158 bits (400), Expect = 1e-37
Identities = 98/235 (41%), Positives = 129/235 (54%), Gaps = 45/235 (19%)
Query: 1 MLLYF-ITCVSFFLFLKPLLLLKPLPSWTNEVRLISLLFHKDFCLKSFCKLFRKTLVSRM 59
MLLYF ITC+SFF F K L LP W +E KTL+S
Sbjct: 1 MLLYFLITCLSFFFFTKSL----SLPPWASET---------------------KTLLSFY 35
Query: 60 SLVKKTLVSKVENVEEEVDSLQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRER 119
+ + + + + + +SVLDLPEL L+CIL+ LPPS LC MA VC SLRER
Sbjct: 36 FIKNPFMNTLHQTKHDPASPVIDQMSVLDLPELALDCILDLLPPSGLCSMARVCSSLRER 95
Query: 120 CVSDHLWERHMKQKWGRVIGSVAYREWKWHVATK--------KNIGSLRHGKPRSLIKFM 171
CVSDHLWE+H+K KWG+++G A+REW+ ++++ G+L K SLI+ +
Sbjct: 96 CVSDHLWEKHLKTKWGKILGPAAHREWQCYISSSTYHLDSPHHQTGNLGFAKIISLIRSL 155
Query: 172 SLYWPFSWMRMKVDAAYSNSANKQNSYLPPDSFMTWYLALETGNFSFPAQVYNRE 226
S + K Y++S LP DS M+ YL+LETG F FPAQVYNRE
Sbjct: 156 SSV----FREDKQRRGYASS-------LPLDSSMSCYLSLETGRFWFPAQVYNRE 199
>gb|AAC32240.1| unknown protein [Arabidopsis thaliana] gi|7459608|pir||T02650
hypothetical protein At2g26850 [imported] - Arabidopsis
thaliana
Length = 346
Score = 152 bits (384), Expect = 7e-36
Identities = 95/229 (41%), Positives = 128/229 (55%), Gaps = 34/229 (14%)
Query: 1 MLLYF-ITCVSFFLFLKPLLLLKPLPSWTNEVR-LISLLFHKDFCLKSFCKLFRKTLVSR 58
MLLY ITC+SFF F K L P W ++ + L+S F K LF +L
Sbjct: 1 MLLYLLITCLSFFFFSKSL----SFPQWASKTKNLLSFYFSK--------YLFTNSLHQI 48
Query: 59 MSLVKKTLVSKVENVEEEVDSLQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRE 118
+ ++ K +S+LDLP+L L+CILE LPPS LC MA VC SLRE
Sbjct: 49 TPDLASPVLGK--------------MSILDLPDLPLDCILELLPPSELCTMARVCSSLRE 94
Query: 119 RCVSDHLWERHMKQKWGRVIGSVAYREWKWHVATKKNIGSLRHGKPR-SLIKFMSLYWPF 177
RCVSDHLWE+H+K KWG+++G A++EW+ ++++ ++ S H L K +SL
Sbjct: 95 RCVSDHLWEKHLKTKWGKILGPSAHKEWQCYLSSPYHLDSPHHKTSHLGLAKIISLMRSL 154
Query: 178 SWMRMKVDAAYSNSANKQNSYLPPDSFMTWYLALETGNFSFPAQVYNRE 226
S + D + S +P DS M +YL+LETG F FPAQVYNRE
Sbjct: 155 SSIFRDDD-----HRRRYPSSIPLDSTMNFYLSLETGRFWFPAQVYNRE 198
>gb|AAS92339.1| At2g26850 [Arabidopsis thaliana] gi|45773808|gb|AAS76708.1|
At2g26850 [Arabidopsis thaliana]
gi|42569359|ref|NP_180253.2| F-box family protein
[Arabidopsis thaliana]
Length = 371
Score = 152 bits (384), Expect = 7e-36
Identities = 95/229 (41%), Positives = 128/229 (55%), Gaps = 34/229 (14%)
Query: 1 MLLYF-ITCVSFFLFLKPLLLLKPLPSWTNEVR-LISLLFHKDFCLKSFCKLFRKTLVSR 58
MLLY ITC+SFF F K L P W ++ + L+S F K LF +L
Sbjct: 1 MLLYLLITCLSFFFFSKSL----SFPQWASKTKNLLSFYFSK--------YLFTNSLHQI 48
Query: 59 MSLVKKTLVSKVENVEEEVDSLQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRE 118
+ ++ K +S+LDLP+L L+CILE LPPS LC MA VC SLRE
Sbjct: 49 TPDLASPVLGK--------------MSILDLPDLPLDCILELLPPSELCTMARVCSSLRE 94
Query: 119 RCVSDHLWERHMKQKWGRVIGSVAYREWKWHVATKKNIGSLRHGKPR-SLIKFMSLYWPF 177
RCVSDHLWE+H+K KWG+++G A++EW+ ++++ ++ S H L K +SL
Sbjct: 95 RCVSDHLWEKHLKTKWGKILGPSAHKEWQCYLSSPYHLDSPHHKTSHLGLAKIISLMRSL 154
Query: 178 SWMRMKVDAAYSNSANKQNSYLPPDSFMTWYLALETGNFSFPAQVYNRE 226
S + D + S +P DS M +YL+LETG F FPAQVYNRE
Sbjct: 155 SSIFRDDD-----HRRRYPSSIPLDSTMNFYLSLETGRFWFPAQVYNRE 198
>gb|AAT06475.1| At2g41170 [Arabidopsis thaliana] gi|30688559|ref|NP_850347.1| F-box
family protein [Arabidopsis thaliana]
Length = 371
Score = 150 bits (378), Expect = 4e-35
Identities = 80/170 (47%), Positives = 103/170 (60%), Gaps = 15/170 (8%)
Query: 63 KKTLVSKVENVEEEVDSLQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVS 122
+K K +N E+E +S+LDLP+L L+CILEKL PS LC M VC LR++CVS
Sbjct: 41 RKETKRKKKNQEDE-----NKMSLLDLPDLTLDCILEKLSPSELCAMTSVCSELRDKCVS 95
Query: 123 DHLWERHMKQKWGRVIGSVAYREWKWHVAT-----KKNIGSLRHGKPRSLIKFMSLYWPF 177
DHLWE+HM+ KWGR++G A +EWK HVAT + S R KP +F++ PF
Sbjct: 96 DHLWEKHMETKWGRLMGDAAIQEWKSHVATIMRCLTSSSSSSRKSKPNWSSRFVANLKPF 155
Query: 178 SWMRMKVDAAYSNSANKQNSYLPP-DSFMTWYLALETGNFSFPAQVYNRE 226
+W+ + + +SYL P DS M WY LE G F FPAQVYNRE
Sbjct: 156 AWL----SSNHGCENRGSSSYLAPIDSVMYWYSNLENGKFWFPAQVYNRE 201
>dbj|BAD94811.1| hypothetical protein [Arabidopsis thaliana]
Length = 371
Score = 148 bits (374), Expect = 1e-34
Identities = 79/170 (46%), Positives = 103/170 (60%), Gaps = 15/170 (8%)
Query: 63 KKTLVSKVENVEEEVDSLQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVS 122
+K K +N E+E +S+LDLP+L L+CILEKL PS LC M VC LR++CVS
Sbjct: 41 RKETKRKKKNQEDE-----NKMSLLDLPDLTLDCILEKLSPSELCAMTSVCSELRDKCVS 95
Query: 123 DHLWERHMKQKWGRVIGSVAYREWKWHVAT-----KKNIGSLRHGKPRSLIKFMSLYWPF 177
DHLWE+HM+ KWGR++G A ++WK HVAT + S R KP +F++ PF
Sbjct: 96 DHLWEKHMETKWGRLMGDAAIQKWKSHVATIMRCLTSSSSSSRKSKPNWSSRFVANLKPF 155
Query: 178 SWMRMKVDAAYSNSANKQNSYLPP-DSFMTWYLALETGNFSFPAQVYNRE 226
+W+ + + +SYL P DS M WY LE G F FPAQVYNRE
Sbjct: 156 AWL----SSNHGCENRGSSSYLAPIDSVMYWYSNLENGKFWFPAQVYNRE 201
>ref|NP_913033.1| unnamed protein product [Oryza sativa (japonica cultivar-group)]
gi|11041564|dbj|BAB00648.2| unnamed protein product
[Oryza sativa (japonica cultivar-group)]
gi|13873014|dbj|BAB44118.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
gi|6498442|dbj|BAA87845.1| unknown protein [Oryza sativa
(japonica cultivar-group)] gi|11138071|dbj|BAB17744.1|
contains ESTs AU081301(E20138),C99280(E10593)~similar to
Arabidopsis thaliana chromosome 2, F12C20.11~unknown
protein [Oryza sativa (japonica cultivar-group)]
Length = 415
Score = 143 bits (360), Expect = 5e-33
Identities = 74/150 (49%), Positives = 93/150 (61%), Gaps = 11/150 (7%)
Query: 85 SVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWGRVIGSVAYR 144
SVLDLPEL +ECIL +LPPS L MAGVC S+RERC DHLWERHM +KWG V+G A
Sbjct: 96 SVLDLPELAIECILARLPPSELRNMAGVCRSMRERCRGDHLWERHMSEKWGGVLGHAARE 155
Query: 145 EWKWHVATKKNIGSLR-------HGKPRSLIKFMSLYWP-FSWMRMKVDAAYSNSANKQN 196
EW+ ++A+ + G G+ R + +S P SWMR + D S K
Sbjct: 156 EWRTYLASAADTGGAAASCSLAGGGRHRRWLAALSCVCPVVSWMRPRAD---GGSGGKSA 212
Query: 197 SYLPPDSFMTWYLALETGNFSFPAQVYNRE 226
+ DS M+WYL++E+G F FPAQVYNRE
Sbjct: 213 GPVLDDSIMSWYLSMESGKFWFPAQVYNRE 242
>gb|AAC25938.1| unknown protein [Arabidopsis thaliana] gi|7487431|pir||T02555
hypothetical protein At2g32560 [imported] - Arabidopsis
thaliana
Length = 228
Score = 141 bits (356), Expect = 1e-32
Identities = 76/151 (50%), Positives = 98/151 (64%), Gaps = 19/151 (12%)
Query: 84 LSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWGRVIGSVAY 143
+SVLDLPEL L+CIL+ LPPS LC MA VC SLRERCVSDHLWE+H+K KWG+++G A+
Sbjct: 1 MSVLDLPELALDCILDLLPPSGLCSMARVCSSLRERCVSDHLWEKHLKTKWGKILGPAAH 60
Query: 144 REWKWHVATK--------KNIGSLRHGKPRSLIKFMSLYWPFSWMRMKVDAAYSNSANKQ 195
REW+ ++++ G+L K SLI+ +S + K Y++S
Sbjct: 61 REWQCYISSSTYHLDSPHHQTGNLGFAKIISLIRSLSSV----FREDKQRRGYASS---- 112
Query: 196 NSYLPPDSFMTWYLALETGNFSFPAQVYNRE 226
LP DS M+ YL+LETG F FPAQVYNRE
Sbjct: 113 ---LPLDSSMSCYLSLETGRFWFPAQVYNRE 140
>ref|NP_915877.1| P0702B09.33 [Oryza sativa (japonica cultivar-group)]
Length = 397
Score = 136 bits (343), Expect = 4e-31
Identities = 75/163 (46%), Positives = 104/163 (63%), Gaps = 9/163 (5%)
Query: 71 ENVEEEVDSLQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHM 130
E E E+++ L +LDLPEL LE +LE+L P SL MA VC +LR+RC +D LW RH+
Sbjct: 94 EGTEAELNAAA--LPLLDLPELALERVLEELEPPSLAAMACVCVALRDRCSADTLWGRHV 151
Query: 131 KQKWGRVIGSVAYREWKWHVATKKNIGSL-RHGKPRSLIKFMSLYWPFSWMR---MKVDA 186
+KWGRV+G+ A +EW+ +A +++ G+L R + RSL ++ WPFSW+ +K +A
Sbjct: 152 NRKWGRVLGAAARKEWEAELAARRSSGALPRPARRRSLADSLACAWPFSWITCRWLKGNA 211
Query: 187 AYSNSANKQNSYLP---PDSFMTWYLALETGNFSFPAQVYNRE 226
+ A S LP D+ WY A+E G FSFPAQVYNRE
Sbjct: 212 VAAEPAAATPSPLPSPATDTVAAWYRAVECGEFSFPAQVYNRE 254
>dbj|BAD68058.1| F-box family protein-like [Oryza sativa (japonica cultivar-group)]
Length = 427
Score = 136 bits (343), Expect = 4e-31
Identities = 75/163 (46%), Positives = 104/163 (63%), Gaps = 9/163 (5%)
Query: 71 ENVEEEVDSLQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHM 130
E E E+++ L +LDLPEL LE +LE+L P SL MA VC +LR+RC +D LW RH+
Sbjct: 94 EGTEAELNAAA--LPLLDLPELALERVLEELEPPSLAAMACVCVALRDRCSADTLWGRHV 151
Query: 131 KQKWGRVIGSVAYREWKWHVATKKNIGSL-RHGKPRSLIKFMSLYWPFSWMR---MKVDA 186
+KWGRV+G+ A +EW+ +A +++ G+L R + RSL ++ WPFSW+ +K +A
Sbjct: 152 NRKWGRVLGAAARKEWEAELAARRSSGALPRPARRRSLADSLACAWPFSWITCRWLKGNA 211
Query: 187 AYSNSANKQNSYLP---PDSFMTWYLALETGNFSFPAQVYNRE 226
+ A S LP D+ WY A+E G FSFPAQVYNRE
Sbjct: 212 VAAEPAAATPSPLPSPATDTVAAWYRAVECGEFSFPAQVYNRE 254
>gb|AAD11995.1| unknown protein [Arabidopsis thaliana] gi|7459607|pir||T02102
hypothetical protein At2g41170 [imported] - Arabidopsis
thaliana
Length = 371
Score = 53.9 bits (128), Expect = 4e-06
Identities = 24/42 (57%), Positives = 31/42 (73%)
Query: 84 LSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHL 125
+S+LDLP+L L+CILEKL PS LC M VC LR++ S +L
Sbjct: 1 MSLLDLPDLTLDCILEKLSPSELCAMTSVCSELRDKGSSSYL 42
Score = 44.3 bits (103), Expect = 0.003
Identities = 22/32 (68%), Positives = 23/32 (71%), Gaps = 1/32 (3%)
Query: 196 NSYLPP-DSFMTWYLALETGNFSFPAQVYNRE 226
+SYL P DS M WY LE G F FPAQVYNRE
Sbjct: 39 SSYLAPIDSVMYWYSNLENGKFWFPAQVYNRE 70
>emb|CAB86904.1| putative protein [Arabidopsis thaliana]
gi|15231726|ref|NP_190868.1| F-box family protein / SKP1
interacting partner 3-related [Arabidopsis thaliana]
gi|11291045|pir||T47557 hypothetical protein F8J2.170 -
Arabidopsis thaliana
Length = 300
Score = 45.4 bits (106), Expect = 0.001
Identities = 22/89 (24%), Positives = 45/89 (49%), Gaps = 2/89 (2%)
Query: 68 SKVENVEEEVDSLQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWE 127
S + N+ + + L + D+PE + C+ L P +C +AG+ S R SD +WE
Sbjct: 3 SSLSNLNDGTNGLAMGPGLGDIPESCVACVFMYLTPPEICNLAGLNRSFRGAASSDSVWE 62
Query: 128 RHMKQKWGRVIGSVAYREWKWHVATKKNI 156
+ + + + ++ + ++H +KK+I
Sbjct: 63 KKLPENYQDLLDLLPPE--RYHSLSKKDI 89
>ref|XP_574248.1| PREDICTED: similar to F-box only protein 31 [Rattus norvegicus]
Length = 238
Score = 38.5 bits (88), Expect = 0.16
Identities = 19/62 (30%), Positives = 33/62 (52%)
Query: 74 EEEVDSLQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQK 133
EE +++ S+L+LP +L I LP + L +A VC R +D +W R +++
Sbjct: 115 EERIEAGPARCSLLELPPELLVEIFASLPGTDLPSLAQVCSRFRRILHTDTIWRRRCREE 174
Query: 134 WG 135
+G
Sbjct: 175 YG 176
>gb|EAA55433.1| hypothetical protein MG09240.4 [Magnaporthe grisea 70-15]
gi|39958250|ref|XP_364395.1| hypothetical protein
MG09240.4 [Magnaporthe grisea 70-15]
Length = 642
Score = 38.1 bits (87), Expect = 0.21
Identities = 17/42 (40%), Positives = 26/42 (61%)
Query: 86 VLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWE 127
+L LP +L+ IL L P L ++G CH+L R S++LW+
Sbjct: 97 LLQLPAELLDNILTYLSPLDLAAVSGTCHALHTRANSNYLWQ 138
>dbj|BAD46347.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|52076038|dbj|BAD46491.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 287
Score = 37.7 bits (86), Expect = 0.27
Identities = 16/55 (29%), Positives = 29/55 (52%)
Query: 88 DLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWGRVIGSVA 142
DLPE + +L +L P +C+MA + + R D +WE + + + R++ A
Sbjct: 21 DLPESCVAEVLRRLDPPEICRMARLSRTFRGAASGDGVWEAKLPRNYARLLAVAA 75
>gb|AAN41295.1| unknown protein [Arabidopsis thaliana] gi|8388622|emb|CAB94142.1|
putative protein [Arabidopsis thaliana]
gi|30695337|ref|NP_567108.2| F-box family protein /
lectin-related [Arabidopsis thaliana]
gi|11291047|pir||T50527 hypothetical protein T27I15_150
- Arabidopsis thaliana
Length = 290
Score = 37.4 bits (85), Expect = 0.35
Identities = 21/79 (26%), Positives = 38/79 (47%), Gaps = 1/79 (1%)
Query: 78 DSLQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWGRV 137
D + L ++DLPE + I+ +L P +C++A + R +D +WE + + RV
Sbjct: 17 DVYSRKLRLVDLPENCVALIMTRLDPPEICRLARLNRMFRRASSADFIWESKLPANY-RV 75
Query: 138 IGSVAYREWKWHVATKKNI 156
I + E KK++
Sbjct: 76 IAHKVFDEITLTKLIKKDL 94
>ref|NP_974470.1| F-box family protein / lectin-related [Arabidopsis thaliana]
Length = 291
Score = 37.4 bits (85), Expect = 0.35
Identities = 21/79 (26%), Positives = 38/79 (47%), Gaps = 1/79 (1%)
Query: 78 DSLQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWGRV 137
D + L ++DLPE + I+ +L P +C++A + R +D +WE + + RV
Sbjct: 17 DVYSRKLRLVDLPENCVALIMTRLDPPEICRLARLNRMFRRASSADFIWESKLPANY-RV 75
Query: 138 IGSVAYREWKWHVATKKNI 156
I + E KK++
Sbjct: 76 IAHKVFDEITLTKLIKKDL 94
>gb|EAA76862.1| hypothetical protein FG07514.1 [Gibberella zeae PH-1]
gi|46126273|ref|XP_387690.1| hypothetical protein
FG07514.1 [Gibberella zeae PH-1]
Length = 501
Score = 36.6 bits (83), Expect = 0.60
Identities = 16/42 (38%), Positives = 25/42 (59%)
Query: 86 VLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWE 127
+LDLP +++ IL L P L ++ C LR+ +SD LW+
Sbjct: 28 LLDLPPELIDNILSHLSPYDLSAISATCRDLRKHALSDILWQ 69
>ref|NP_325859.1| ABC TRANSPORTER PERMEASE PROTEIN [Mycoplasma pulmonis UAB CTIP]
gi|14089441|emb|CAC13201.1| ABC TRANSPORTER PERMEASE
PROTEIN [Mycoplasma pulmonis] gi|25308843|pir||D90515
ABC transporter permease protein [imported] - Mycoplasma
pulmonis (strain UAB CTIP)
Length = 334
Score = 36.2 bits (82), Expect = 0.78
Identities = 33/110 (30%), Positives = 47/110 (42%), Gaps = 16/110 (14%)
Query: 1 MLLYFITCVSFFLFLKPLLLLKP-LPSWTNEVRLISLL--------FHKDFCLKSFCKLF 51
+LL+F T + F + ++ KP +P TNE+R ISLL F K F F +F
Sbjct: 53 VLLFFATIIVFPFYFLIVIAFKPDIPERTNEIRTISLLLDNLKWENFSKAFQAGYFKAVF 112
Query: 52 RKTLVSRMSLVKKTLVSKVENVEEEVDSLQQN-------LSVLDLPELVL 94
TL + S+ K S + LS+L LPE+ L
Sbjct: 113 LSTLTTVFSIFAKIFFSMTLGYAFSFHKWRGKKIVWAIFLSILILPEVAL 162
>gb|AAK59472.2| unknown protein [Arabidopsis thaliana]
Length = 269
Score = 35.8 bits (81), Expect = 1.0
Identities = 20/73 (27%), Positives = 36/73 (48%), Gaps = 1/73 (1%)
Query: 84 LSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWGRVIGSVAY 143
L ++DLPE + I+ +L P +C++A + R +D +WE + + RVI +
Sbjct: 2 LRLVDLPENCVALIMTRLDPPEICRLARLNRMFRRASSADFIWESKLPANY-RVIAHKVF 60
Query: 144 REWKWHVATKKNI 156
E KK++
Sbjct: 61 DEITLTKLIKKDL 73
>ref|ZP_00150853.1| COG0525: Valyl-tRNA synthetase [Dechloromonas aromatica RCB]
Length = 943
Score = 35.8 bits (81), Expect = 1.0
Identities = 24/99 (24%), Positives = 42/99 (42%), Gaps = 4/99 (4%)
Query: 114 HSLRERCVSDHLWERHMKQKW----GRVIGSVAYREWKWHVATKKNIGSLRHGKPRSLIK 169
+++++ C+S LW H W G++ + + E K A + G+L+ +
Sbjct: 404 NNIQDWCISRQLWWGHQIPAWYGDNGQIFVAHSEAEAKAEAAKQGYTGTLKRDEDVLDTW 463
Query: 170 FMSLYWPFSWMRMKVDAAYSNSANKQNSYLPPDSFMTWY 208
F S WPFS + D A + YLP +T +
Sbjct: 464 FSSALWPFSTLDWTGDEAIDAANPLLKQYLPSSVLVTGF 502
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.325 0.137 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 402,633,335
Number of Sequences: 2540612
Number of extensions: 15736663
Number of successful extensions: 53436
Number of sequences better than 10.0: 52
Number of HSP's better than 10.0 without gapping: 36
Number of HSP's successfully gapped in prelim test: 16
Number of HSP's that attempted gapping in prelim test: 53380
Number of HSP's gapped (non-prelim): 53
length of query: 236
length of database: 863,360,394
effective HSP length: 124
effective length of query: 112
effective length of database: 548,324,506
effective search space: 61412344672
effective search space used: 61412344672
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 73 (32.7 bits)
Medicago: description of AC136507.1


