

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136472.16 - phase: 0
(212 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
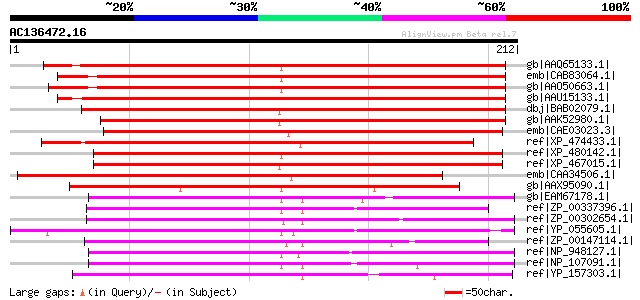
Score E
Sequences producing significant alignments: (bits) Value
gb|AAQ65133.1| At3g43660 [Arabidopsis thaliana] gi|7362791|emb|C... 258 1e-67
emb|CAB83064.1| nodulin-like protein [Arabidopsis thaliana] gi|1... 257 2e-67
gb|AAO50663.1| putative tonoplast intrinsic protein, alpha (alph... 253 2e-66
gb|AAU15133.1| At1g76800 [Arabidopsis thaliana] gi|15223732|ref|... 244 9e-64
dbj|BAB02079.1| nodulin-lile protein [Arabidopsis thaliana] gi|1... 223 4e-57
gb|AAK52980.1| AT3g25190/MJL12_14 [Arabidopsis thaliana] gi|1797... 214 1e-54
emb|CAE03023.3| OSJNBa0091D06.17 [Oryza sativa (japonica cultiva... 198 1e-49
ref|XP_474433.1| OSJNBa0070M12.11 [Oryza sativa (japonica cultiv... 194 1e-48
ref|XP_480142.1| putative nodulin-21 [Oryza sativa (japonica cul... 188 8e-47
ref|XP_467015.1| nodulin-21-like [Oryza sativa (japonica cultiva... 185 6e-46
emb|CAA34506.1| unnamed protein product [Glycine max] gi|99942|p... 183 3e-45
gb|AAX95090.1| Integral membrane protein, putative [Oryza sativa... 142 5e-33
gb|EAM67178.1| Protein of unknown function DUF125 [Jannaschia sp... 120 3e-26
ref|ZP_00337396.1| COG1814: Uncharacterized membrane protein [Si... 110 3e-23
ref|ZP_00302654.1| COG1814: Uncharacterized membrane protein [No... 109 5e-23
ref|YP_055605.1| conserved mebrane associated protein, DUF125 [P... 108 1e-22
ref|ZP_00147114.1| COG1814: Uncharacterized membrane protein [Ps... 108 1e-22
ref|NP_948127.1| possible nodulin-related protein [Rhodopseudomo... 107 3e-22
ref|NP_107091.1| similar to nodulin 21 [Mesorhizobium loti MAFF3... 103 3e-21
ref|YP_157303.1| hypothetical protein ebA544 [Azoarcus sp. EbN1]... 98 1e-19
>gb|AAQ65133.1| At3g43660 [Arabidopsis thaliana] gi|7362791|emb|CAB83067.1|
nodulin-like protein [Arabidopsis thaliana]
gi|15229736|ref|NP_189952.1| nodulin, putative
[Arabidopsis thaliana] gi|51970300|dbj|BAD43842.1|
nodulin - like protein [Arabidopsis thaliana]
gi|11282380|pir||T47402 nodulin-like protein -
Arabidopsis thaliana
Length = 198
Score = 258 bits (658), Expect = 1e-67
Identities = 134/194 (69%), Positives = 162/194 (83%), Gaps = 4/194 (2%)
Query: 15 SESKVYNMDVEKKGEEENDEKDYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVKT 74
S + N+D+EK ++ DY++RAQWLRAAVLGANDGL+STASLMMG+GAV +DV+
Sbjct: 3 SNNNNLNLDMEK---DQETTFDYSKRAQWLRAAVLGANDGLVSTASLMMGIGAVKQDVRI 59
Query: 75 MILTGIAGLVAGACSMAIGEFVSVYSQYDIEFAQMKRQ-GNISQKDKLPNPYYAAFASAI 133
M+LTG AGLVAGACSMAIGEF+SVYSQYDIE AQMKR+ G ++K+KLP+P AA ASA+
Sbjct: 60 MLLTGFAGLVAGACSMAIGEFISVYSQYDIEVAQMKRESGGETKKEKLPSPTQAAIASAL 119
Query: 134 AFAVGAFVPLLGAAFVKDYKVRLGVVVGVVSLALFGFGLLSAVLGKAPLVKSSLRVLIGG 193
AF +GA VPLL AAFVK+YKVR+GV+V V+LAL FG L AVLGKAP+VKS +RVLIGG
Sbjct: 120 AFTLGAIVPLLAAAFVKEYKVRIGVIVAAVTLALVMFGWLGAVLGKAPVVKSLVRVLIGG 179
Query: 194 WLAMSLTFGLTKLV 207
WLAM++TFG TKLV
Sbjct: 180 WLAMAITFGFTKLV 193
>emb|CAB83064.1| nodulin-like protein [Arabidopsis thaliana]
gi|15229728|ref|NP_189949.1| nodulin, putative
[Arabidopsis thaliana] gi|11282379|pir||T47399
nodulin-like protein - Arabidopsis thaliana
Length = 200
Score = 257 bits (656), Expect = 2e-67
Identities = 135/188 (71%), Positives = 160/188 (84%), Gaps = 4/188 (2%)
Query: 21 NMDVEKKGEEENDEKDYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVKTMILTGI 80
N+D+EK E+ D Y++RAQWLRAAVLGANDGL+STASLMMGVGAV ++VK MILTG
Sbjct: 8 NLDMEKDQEKAFD---YSKRAQWLRAAVLGANDGLVSTASLMMGVGAVKQNVKIMILTGF 64
Query: 81 AGLVAGACSMAIGEFVSVYSQYDIEFAQMKRQ-GNISQKDKLPNPYYAAFASAIAFAVGA 139
AGLVAGACSMAIGEFVSVYSQYDIE AQMKR+ G +K+KLP+P AA ASA+AF++GA
Sbjct: 65 AGLVAGACSMAIGEFVSVYSQYDIEVAQMKRETGGEIEKEKLPSPTQAAAASALAFSLGA 124
Query: 140 FVPLLGAAFVKDYKVRLGVVVGVVSLALFGFGLLSAVLGKAPLVKSSLRVLIGGWLAMSL 199
VPLL AAFVK+YKVR+G +V V+LAL FG L AVLGKAP+VKSSLRVL+GGWLAM++
Sbjct: 125 MVPLLAAAFVKEYKVRIGAIVAAVTLALVMFGWLGAVLGKAPVVKSSLRVLVGGWLAMAI 184
Query: 200 TFGLTKLV 207
T+G TKL+
Sbjct: 185 TYGFTKLI 192
>gb|AAO50663.1| putative tonoplast intrinsic protein, alpha (alpha-TIP)
[Arabidopsis thaliana] gi|28392889|gb|AAO41881.1|
putative tonoplast intrinsic protein, alpha (alpha-TIP)
[Arabidopsis thaliana] gi|30687198|ref|NP_173538.2|
nodulin, putative [Arabidopsis thaliana]
gi|25352447|pir||F86344 T22I11.3 protein - Arabidopsis
thaliana gi|8886987|gb|AAF80647.1| Contains similarity
to Nodulin 21 from Soybean gb|X16488. [Arabidopsis
thaliana]
Length = 200
Score = 253 bits (647), Expect = 2e-66
Identities = 135/192 (70%), Positives = 158/192 (81%), Gaps = 4/192 (2%)
Query: 17 SKVYNMDVEKKGEEENDEKDYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVKTMI 76
S N+D+E E+ D Y++RAQWLRAAVLGANDGL+STASLMMGVGAV +DVK MI
Sbjct: 7 SNSLNLDMEMDQEKAFD---YSKRAQWLRAAVLGANDGLVSTASLMMGVGAVKQDVKVMI 63
Query: 77 LTGIAGLVAGACSMAIGEFVSVYSQYDIEFAQMKRQ-GNISQKDKLPNPYYAAFASAIAF 135
L+G AGLVAGACSMAIGEFVSVYSQYDIE AQMKR+ G +K+KLP+P AA ASA+AF
Sbjct: 64 LSGFAGLVAGACSMAIGEFVSVYSQYDIEVAQMKRENGGQVEKEKLPSPMQAAAASALAF 123
Query: 136 AVGAFVPLLGAAFVKDYKVRLGVVVGVVSLALFGFGLLSAVLGKAPLVKSSLRVLIGGWL 195
++GA VPL+ AAFVKDY VR+G +V V+LAL FG L AVLGKAP+ KSS RVLIGGWL
Sbjct: 124 SLGAIVPLMAAAFVKDYHVRIGAIVAAVTLALVMFGWLGAVLGKAPVFKSSARVLIGGWL 183
Query: 196 AMSLTFGLTKLV 207
AM++TFGLTKL+
Sbjct: 184 AMAVTFGLTKLI 195
>gb|AAU15133.1| At1g76800 [Arabidopsis thaliana] gi|15223732|ref|NP_177806.1|
nodulin, putative [Arabidopsis thaliana]
gi|48958481|gb|AAT47793.1| At1g76800 [Arabidopsis
thaliana] gi|6143895|gb|AAF04441.1| nodulin-like
protein; 66117-66707 [Arabidopsis thaliana]
gi|25352449|pir||F96796 nodulin-like protein,
66117-66707 [imported] - Arabidopsis thaliana
Length = 196
Score = 244 bits (624), Expect = 9e-64
Identities = 129/187 (68%), Positives = 152/187 (80%), Gaps = 4/187 (2%)
Query: 21 NMDVEKKGEEENDEKDYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVKTMILTGI 80
NMD+EK E+ DY++R+QWLRAAVLGANDGL+STASLMMGVGAV DVK MIL+G
Sbjct: 9 NMDIEK----ESTTFDYSKRSQWLRAAVLGANDGLVSTASLMMGVGAVKHDVKAMILSGF 64
Query: 81 AGLVAGACSMAIGEFVSVYSQYDIEFAQMKRQGNISQKDKLPNPYYAAFASAIAFAVGAF 140
AG+VAGACSMAIGEFVSVYSQYDIE AQM+R +K+KLP+P AA ASA+AF+ GA
Sbjct: 65 AGMVAGACSMAIGEFVSVYSQYDIEVAQMERDSVEIEKEKLPSPMQAAAASALAFSAGAI 124
Query: 141 VPLLGAAFVKDYKVRLGVVVGVVSLALFGFGLLSAVLGKAPLVKSSLRVLIGGWLAMSLT 200
VPLL AAFVK+YK+R+ VV V++AL FG L A LGKAP V+SS RVL GGWLAM++T
Sbjct: 125 VPLLAAAFVKEYKMRIISVVVAVTVALMVFGWLGAALGKAPAVRSSARVLFGGWLAMAVT 184
Query: 201 FGLTKLV 207
FGLTKL+
Sbjct: 185 FGLTKLI 191
>dbj|BAB02079.1| nodulin-lile protein [Arabidopsis thaliana]
gi|15230815|ref|NP_189156.1| nodulin, putative
[Arabidopsis thaliana]
Length = 219
Score = 223 bits (567), Expect = 4e-57
Identities = 117/190 (61%), Positives = 146/190 (76%), Gaps = 13/190 (6%)
Query: 31 ENDEKDYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVKTMILTGIAGLVAGACSM 90
E +E DY QRAQWLRAA+LGANDGL++ ASLMMGVG++ +DVK M+L G AGLVAGACSM
Sbjct: 23 EKEEVDYMQRAQWLRAALLGANDGLVTVASLMMGVGSIKEDVKAMLLVGFAGLVAGACSM 82
Query: 91 AIGEFVSVYSQYDIEFAQMKR-------------QGNISQKDKLPNPYYAAFASAIAFAV 137
AIGEFVSV +Q DIE AQMKR Q +K++LPNP AA ASA+AF+V
Sbjct: 83 AIGEFVSVCTQRDIETAQMKRAIEHKTSLSAIDEQEEEEKKERLPNPGQAAIASALAFSV 142
Query: 138 GAFVPLLGAAFVKDYKVRLGVVVGVVSLALFGFGLLSAVLGKAPLVKSSLRVLIGGWLAM 197
GA +PLLGA F++++KVR+ VV V ++AL FG+ AVLGK +VKSS+RV+IGGW+AM
Sbjct: 143 GAAMPLLGAVFIENHKVRMVVVAVVATIALVVFGVTGAVLGKTSVVKSSVRVVIGGWMAM 202
Query: 198 SLTFGLTKLV 207
+LTFGLTK +
Sbjct: 203 ALTFGLTKFI 212
>gb|AAK52980.1| AT3g25190/MJL12_14 [Arabidopsis thaliana]
gi|17978889|gb|AAL47414.1| AT3g25190/MJL12_14
[Arabidopsis thaliana]
Length = 190
Score = 214 bits (545), Expect = 1e-54
Identities = 113/182 (62%), Positives = 141/182 (77%), Gaps = 13/182 (7%)
Query: 39 QRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVKTMILTGIAGLVAGACSMAIGEFVSV 98
QRAQWLRAA+LGANDGL++ ASLMMGVG++ +DVK M+L G AGLVAGACSMAIGEFVSV
Sbjct: 2 QRAQWLRAALLGANDGLVTVASLMMGVGSIKEDVKAMLLVGFAGLVAGACSMAIGEFVSV 61
Query: 99 YSQYDIEFAQMKR-------------QGNISQKDKLPNPYYAAFASAIAFAVGAFVPLLG 145
+Q DIE AQMKR Q +K++LPNP AA ASA+AF+VGA +PLLG
Sbjct: 62 CTQRDIETAQMKRAIEHKTSLSAIDEQEEEEKKERLPNPGQAAIASALAFSVGAAMPLLG 121
Query: 146 AAFVKDYKVRLGVVVGVVSLALFGFGLLSAVLGKAPLVKSSLRVLIGGWLAMSLTFGLTK 205
A F++++KVR+ VV V ++AL FG+ AVLGK +VKSS+RV+IGGW+AM+LTFGLTK
Sbjct: 122 AVFIENHKVRMVVVAVVATIALVVFGVTGAVLGKTSVVKSSVRVVIGGWMAMALTFGLTK 181
Query: 206 LV 207
+
Sbjct: 182 FI 183
>emb|CAE03023.3| OSJNBa0091D06.17 [Oryza sativa (japonica cultivar-group)]
gi|50926906|ref|XP_473339.1| OSJNBa0091D06.17 [Oryza
sativa (japonica cultivar-group)]
Length = 189
Score = 198 bits (503), Expect = 1e-49
Identities = 100/173 (57%), Positives = 132/173 (75%), Gaps = 6/173 (3%)
Query: 40 RAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVKTMILTGIAGLVAGACSMAIGEFVSVY 99
RAQWLRAAVLGANDGL+S ASLM+G+GAV ++ K M+++G+AGLVAGACSMAIGEFVSVY
Sbjct: 3 RAQWLRAAVLGANDGLVSVASLMIGIGAVNENNKAMLVSGLAGLVAGACSMAIGEFVSVY 62
Query: 100 SQYDIEFAQMKRQGNI------SQKDKLPNPYYAAFASAIAFAVGAFVPLLGAAFVKDYK 153
+QYDIE Q++R G+I + ++KLP+P AAFASA+AFA+G +PLL + F+K +
Sbjct: 63 AQYDIEVTQIERDGDIDGADAAAAREKLPSPTQAAFASALAFAIGGLLPLLTSGFIKPWG 122
Query: 154 VRLGVVVGVVSLALFGFGLLSAVLGKAPLVKSSLRVLIGGWLAMSLTFGLTKL 206
R+GVV S+ L GFG LG A +V+S RVL+GGWLAM +T+ + +L
Sbjct: 123 PRVGVVCAASSVGLAGFGAAGGYLGGANMVRSGTRVLLGGWLAMLITYAVLRL 175
>ref|XP_474433.1| OSJNBa0070M12.11 [Oryza sativa (japonica cultivar-group)]
gi|38345828|emb|CAD41933.2| OSJNBa0070M12.11 [Oryza
sativa (japonica cultivar-group)]
Length = 199
Score = 194 bits (494), Expect = 1e-48
Identities = 103/183 (56%), Positives = 133/183 (72%), Gaps = 3/183 (1%)
Query: 14 RSESKVYNMDVEKKGEEENDEKDYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVK 73
RS S +D E DE D RA WLRAAVLGANDGL+STASLM+GVGAV + +
Sbjct: 9 RSSSNKLQVDAENPAAV-GDELDLAARANWLRAAVLGANDGLVSTASLMLGVGAVKAEAR 67
Query: 74 TMILTGIAGLVAGACSMAIGEFVSVYSQYDIEFAQMKRQGNISQKDK--LPNPYYAAFAS 131
M+++G AGL+AGACSMAIGEFVSV SQ D+E AQ++R G +++ LP+P AA AS
Sbjct: 68 AMVISGFAGLLAGACSMAIGEFVSVCSQRDVELAQLERDGKRGGEEEKALPSPAQAAAAS 127
Query: 132 AIAFAVGAFVPLLGAAFVKDYKVRLGVVVGVVSLALFGFGLLSAVLGKAPLVKSSLRVLI 191
A+AF+VGA VPLL A F+ +Y++R+ VVV V S+AL FG + AVLG+A + +SS RV++
Sbjct: 128 AMAFSVGAVVPLLAAGFIVNYRLRIAVVVAVASVALAAFGCVGAVLGRAAVARSSARVVL 187
Query: 192 GGW 194
GGW
Sbjct: 188 GGW 190
>ref|XP_480142.1| putative nodulin-21 [Oryza sativa (japonica cultivar-group)]
gi|37806250|dbj|BAC99767.1| putative nodulin-21 [Oryza
sativa (japonica cultivar-group)]
gi|27817911|dbj|BAC55676.1| putative nodulin-21 [Oryza
sativa (japonica cultivar-group)]
Length = 249
Score = 188 bits (478), Expect = 8e-47
Identities = 99/182 (54%), Positives = 127/182 (69%), Gaps = 11/182 (6%)
Query: 36 DYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVKTMILTGIAGLVAGACSMAIGEF 95
DY+ RAQWLRAAVLGANDGL+S ASLM+GVGAV++ + M+++G+AGLVAGACSMAIGEF
Sbjct: 63 DYSGRAQWLRAAVLGANDGLVSVASLMIGVGAVSESGRAMLVSGVAGLVAGACSMAIGEF 122
Query: 96 VSVYSQYDIEFAQMKRQ-----------GNISQKDKLPNPYYAAFASAIAFAVGAFVPLL 144
VSVY+QYDIE A +R+ G +LP+P+ AA ASA+AF VGA +PLL
Sbjct: 123 VSVYAQYDIEVAAARRRRRQRRRRCDGDGEEEGSGRLPSPFKAAAASALAFTVGALLPLL 182
Query: 145 GAAFVKDYKVRLGVVVGVVSLALFGFGLLSAVLGKAPLVKSSLRVLIGGWLAMSLTFGLT 204
FV+ + R+ V S AL GFG L A LG A +S+ RVL+GGW AM+ +G+
Sbjct: 183 AGGFVRPWAPRVAAVCAATSAALAGFGALGAALGGASPARSAARVLLGGWAAMAACYGVL 242
Query: 205 KL 206
+L
Sbjct: 243 RL 244
>ref|XP_467015.1| nodulin-21-like [Oryza sativa (japonica cultivar-group)]
gi|49388656|dbj|BAD25791.1| nodulin-21-like [Oryza
sativa (japonica cultivar-group)]
Length = 232
Score = 185 bits (470), Expect = 6e-46
Identities = 96/177 (54%), Positives = 129/177 (72%), Gaps = 6/177 (3%)
Query: 36 DYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVKTMILTGIAGLVAGACSMAIGEF 95
+Y RAQWLRAAVLGANDGL+S ASLM+GVGA + M+++G+AGLVAGACSMAIGEF
Sbjct: 50 NYVARAQWLRAAVLGANDGLVSVASLMVGVGAANGTRRAMLVSGLAGLVAGACSMAIGEF 109
Query: 96 VSVYSQYDIEFAQMKR------QGNISQKDKLPNPYYAAFASAIAFAVGAFVPLLGAAFV 149
VSVY+Q DI+ AQ++R ++++LP+P AA ASA++FA GA +PLL FV
Sbjct: 110 VSVYAQCDIQAAQIERARGGKDADGGEEEEELPSPTMAAVASALSFAAGAALPLLAGGFV 169
Query: 150 KDYKVRLGVVVGVVSLALFGFGLLSAVLGKAPLVKSSLRVLIGGWLAMSLTFGLTKL 206
+ + R+ V SL L GFG+ SA LG A + +S +R+L+GGWLAM++T+G+ KL
Sbjct: 170 RPWAARVAAVCAASSLGLAGFGVASAYLGGAGVARSGVRMLVGGWLAMAVTYGVLKL 226
>emb|CAA34506.1| unnamed protein product [Glycine max] gi|99942|pir||S08632
nodulin-21 - soybean gi|128405|sp|P16313|NO21_SOYBN
NODULIN 21 (N-21)
Length = 206
Score = 183 bits (464), Expect = 3e-45
Identities = 96/189 (50%), Positives = 125/189 (65%), Gaps = 11/189 (5%)
Query: 4 IQHHHAATLARSESKVYNMDVEKKGEEENDEKDYTQRAQWLRAAVLGANDGLLSTASLMM 63
+ H+H + + + + E++ DY QRAQWLRAA+LGANDGL+S ASLMM
Sbjct: 17 VPHNHVGAVLLTIPTIKIDGKQTLATEDHTSIDYLQRAQWLRAAILGANDGLVSVASLMM 76
Query: 64 GVGAVTKDVKTMILTGIAGLVAGACSMAIGEFVSVYSQYDIEFAQMKRQGNIS------- 116
GVGAV +D K M+L G AGLVAGAC MAIGEFV+VY+QY++E QMKR N+S
Sbjct: 77 GVGAVKRDAKAMLLAGFAGLVAGACGMAIGEFVAVYTQYEVEVGQMKRDMNMSVGGERDL 136
Query: 117 ----QKDKLPNPYYAAFASAIAFAVGAFVPLLGAAFVKDYKVRLGVVVGVVSLALFGFGL 172
++ LPNP A ASA+ F++GA VPLL AAF+++Y+ R+ VVV + LAL FG
Sbjct: 137 EMEMERRTLPNPLQATLASALCFSIGALVPLLSAAFIENYRTRIIVVVAMSCLALVVFGW 196
Query: 173 LSAVLGKAP 181
+ A LGK P
Sbjct: 197 VGAKLGKTP 205
>gb|AAX95090.1| Integral membrane protein, putative [Oryza sativa (japonica
cultivar-group)]
Length = 207
Score = 142 bits (359), Expect = 5e-33
Identities = 86/176 (48%), Positives = 115/176 (64%), Gaps = 13/176 (7%)
Query: 26 KKGEEENDEKDYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTK-DVKTMILTGIAGLV 84
K+ EE D+ +QWLRAAVLGA+DGL+STA+LM+G+GA D + ++L+G+AGLV
Sbjct: 17 KEEEEATDDGGGGVSSQWLRAAVLGASDGLVSTAALMLGIGAARPADARAVLLSGLAGLV 76
Query: 85 AGACSMAIGEFVSVYSQYDIEFAQMKRQ-----------GNISQKDKLPNPYYAAFASAI 133
AGACSMAIGE+VSV+ Q D+E A ++R+ G + + P AA ASA+
Sbjct: 77 AGACSMAIGEYVSVHVQLDVELADLERRRRRGGPAPAGLGLHAAAAAVSRPGQAAAASAL 136
Query: 134 AFAVGAFVPLLGAAFVKD-YKVRLGVVVGVVSLALFGFGLLSAVLGKAPLVKSSLR 188
+FA GA +PLL A FV Y+VR+ VVV SLAL FG A LG+AP ++ LR
Sbjct: 137 SFAAGAALPLLAAWFVAGAYRVRVVVVVATASLALAAFGAAGARLGRAPGGRAGLR 192
>gb|EAM67178.1| Protein of unknown function DUF125 [Jannaschia sp. CCS1]
Length = 239
Score = 120 bits (300), Expect = 3e-26
Identities = 87/228 (38%), Positives = 116/228 (50%), Gaps = 53/228 (23%)
Query: 34 EKDYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVKTMILTGIAGLVAGACSMAIG 93
E + RA WLRAAV+GANDG+LSTASL+ GV A + D T++L G+AGLVAGA SMA G
Sbjct: 15 EPHLSGRAGWLRAAVMGANDGILSTASLIAGVAAGSGDKATILLAGLAGLVAGALSMAAG 74
Query: 94 EFVSVYSQYDIEFAQMKRQ---------------------------------GNISQKDK 120
E+VSV SQ D E A ++R+ ++++ D
Sbjct: 75 EYVSVSSQADAERADVERERSELARNPEAELAELTAIYVERGLTPDLADRVARDLTEVDA 134
Query: 121 L---------------PNPYYAAFASAIAFAVGAFVPLLGA--AFVKDYKVRLGVVVGVV 163
L P P AA SA+ FA GA VPL A A V D + +G G
Sbjct: 135 LTAHLRDEIGLTDLAPPRPVQAALVSALTFAAGASVPLAMAWLAPVDDILIWVG---GAT 191
Query: 164 SLALFGFGLLSAVLGKAPLVKSSLRVLIGGWLAMSLTFGLTKLVNHVV 211
AL G L A +G AP V+++ RV++ G LAM++T + LV +
Sbjct: 192 LAALGSLGALGATVGGAPRVRAAARVMVWGALAMAITTAIGALVGQAI 239
>ref|ZP_00337396.1| COG1814: Uncharacterized membrane protein [Silicibacter sp. TM1040]
Length = 234
Score = 110 bits (275), Expect = 3e-23
Identities = 77/216 (35%), Positives = 108/216 (49%), Gaps = 49/216 (22%)
Query: 33 DEKDYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVKTMILTGIAGLVAGACSMAI 92
D+ Y R+ WLRAAVLGANDG++S +SL++GV A + +++ GIAGLVAGA SMA
Sbjct: 9 DDPHYVNRSGWLRAAVLGANDGIVSVSSLVVGVAAADPGPRPVLIAGIAGLVAGAMSMAA 68
Query: 93 GEFVSVYSQYDIEFAQMKRQ---------------------------------GNISQKD 119
GE+VSV SQ D+E A ++R+ +++KD
Sbjct: 69 GEYVSVSSQSDVEQADIERERRALSETPEAELEELIEIYEGRGLSRPTAEAVAKELTEKD 128
Query: 120 KL---------------PNPYYAAFASAIAFAVGAFVPLLGAAFVKDYKVRLGVVVGVVS 164
L NP AA AS F+V A +P+L AA + + VV+ V
Sbjct: 129 ALGAHIRDELGLSEVHTANPLQAALASGATFSVAAAIPVL-AAILSPASAIIPVVLIVTI 187
Query: 165 LALFGFGLLSAVLGKAPLVKSSLRVLIGGWLAMSLT 200
AL G L A+ G AP+ + +RV+ G AM +T
Sbjct: 188 FALAILGALGAIAGAAPIAPAVIRVVGWGVFAMVVT 223
>ref|ZP_00302654.1| COG1814: Uncharacterized membrane protein [Novosphingobium
aromaticivorans DSM 12444]
Length = 230
Score = 109 bits (273), Expect = 5e-23
Identities = 84/227 (37%), Positives = 109/227 (48%), Gaps = 49/227 (21%)
Query: 33 DEKDYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVKTMILTGIAGLVAGACSMAI 92
+E +R WLRAAVLGANDG++STASL++GV A D + +I+ +AGLVAGA SMA
Sbjct: 5 EESHLVERIGWLRAAVLGANDGIVSTASLIIGVAASGADRQAIIVAAMAGLVAGAMSMAA 64
Query: 93 GEFVSVYSQYDIEFAQMKRQG-----------------------NISQKDKL-------- 121
GE+VSV SQ D E A + R+ + DK+
Sbjct: 65 GEYVSVSSQADTEKADLARETAELAADPEFEHRELAAIYVARGVDADTADKVAAQMMAHD 124
Query: 122 -----------------PNPYYAAFASAIAFAVGAFVPLLGAAFVKDYKVRLGVVVGVVS 164
P AA ASA F GA +P+L AA + + G VG +
Sbjct: 125 PLTTHARDELHITSTTAARPVVAALASAATFTAGALLPVLLAALLPMSLLAAGEAVGSI- 183
Query: 165 LALFGFGLLSAVLGKAPLVKSSLRVLIGGWLAMSLTFGLTKLVNHVV 211
L L G L A G AP + RV+ G LAM+LT G+ +LV VV
Sbjct: 184 LFLALLGTLGARAGGAPAGRPVARVVFWGALAMALTAGIGRLVGAVV 230
>ref|YP_055605.1| conserved mebrane associated protein, DUF125 [Propionibacterium
acnes KPA171202] gi|50839980|gb|AAT82647.1| conserved
mebrane associated protein, DUF125 [Propionibacterium
acnes KPA171202]
Length = 352
Score = 108 bits (270), Expect = 1e-22
Identities = 84/263 (31%), Positives = 119/263 (44%), Gaps = 57/263 (21%)
Query: 1 MAEIQHHHAATLAR----SESKVYNMDVEKKGEEENDEKDYTQRAQWLRAAVLGANDGLL 56
M+ I + A LA +ES ++ + E + + WLRAAVLGANDG++
Sbjct: 91 MSSIPNTSALALAHDASSTESTRFSTSSRQPHEPDKGTGSLNSKLNWLRAAVLGANDGII 150
Query: 57 STASLMMGVGAVTKDVKTMILTGIAGLVAGACSMAIGEFVSVYSQYDIEFAQMKRQ---- 112
STA ++MGV T D ++ + G+AGL AGA SMA GE+VSV SQ DIE M ++
Sbjct: 151 STAGIVMGVAGATVDRSSLFIAGLAGLAAGALSMAGGEYVSVSSQRDIEKTVMAKEAAEL 210
Query: 113 ----------------------GNISQ----------------------KDKLPNPYYAA 128
G Q D+ NP+YAA
Sbjct: 211 RDFPDEELEELTEIYTEKGLSRGTARQVALELTAHDPLRAHAEAELGLDPDEYTNPWYAA 270
Query: 129 FASAIAFAVGAFVPLLGAAFVKDYKVRLGVVVGVVSLALFGFGLLSAVLGKAPLVKSSLR 188
FAS AF VGA VPLL R+ + + + LF GL SA+ + + R
Sbjct: 271 FASMAAFTVGALVPLL-VMVCSPTATRVYITIAATIVGLFLTGLGSALASGSGKTRPVAR 329
Query: 189 VLIGGWLAMSLTFGLTKLVNHVV 211
+I G +M++T+ L+ H+V
Sbjct: 330 NIIVGICSMAITY----LIGHLV 348
>ref|ZP_00147114.1| COG1814: Uncharacterized membrane protein [Psychrobacter sp. 273-4]
Length = 233
Score = 108 bits (270), Expect = 1e-22
Identities = 76/218 (34%), Positives = 113/218 (50%), Gaps = 51/218 (23%)
Query: 32 NDEKDYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVKTMILTGIAGLVAGACSMA 91
+DE + R WLRAAVLGANDGL+STASL++GV A + +T++LTG+A L AGA SMA
Sbjct: 6 HDEAHLSNRNHWLRAAVLGANDGLISTASLLVGVAAASISSQTLLLTGMAALTAGALSMA 65
Query: 92 IGEFVSVYSQYDIEFAQMKRQGN---------------------------------ISQK 118
GE++SV SQ D E A + ++ + ++Q
Sbjct: 66 AGEYISVSSQADTEKADLDKELHELTHNAEHELNELTKIYETRGLDHVLAHQVAVALTQH 125
Query: 119 DKL---------------PNPYYAAFASAIAFAVGAFVPLLGAAFVKDYKVRLGV-VVGV 162
D L P +A+ AS ++F GA +P++G + + + + +
Sbjct: 126 DALEAHARDEIGLTDLSQAKPIHASVASGLSFIAGAILPIIGILLLPVQSLVWSLSSLTI 185
Query: 163 VSLALFGFGLLSAVLGKAPLVKSSLRVLIGGWLAMSLT 200
V LAL G++SA LG AP++ ++ RV+I G LAM T
Sbjct: 186 VGLAL--LGIISARLGGAPVIPATARVVIWGVLAMVAT 221
>ref|NP_948127.1| possible nodulin-related protein [Rhodopseudomonas palustris
CGA009] gi|39649705|emb|CAE28226.1| possible
nodulin-related protein [Rhodopseudomonas palustris
CGA009]
Length = 231
Score = 107 bits (266), Expect = 3e-22
Identities = 80/226 (35%), Positives = 107/226 (46%), Gaps = 49/226 (21%)
Query: 34 EKDYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVKTMILTGIAGLVAGACSMAIG 93
E R WLRAAVLGANDG++STASL++GV A + ++L G+AGLVAGA SMA G
Sbjct: 7 ENHLINRIGWLRAAVLGANDGIISTASLVVGVAAAATSSEEVLLAGVAGLVAGAMSMAAG 66
Query: 94 EFVSVYSQYDIEFAQMKRQ---------------------------------GNISQKD- 119
E+VSV SQ D E A + R+ +S KD
Sbjct: 67 EYVSVSSQSDTEQADLARERKELADAPDSELDELTKIYVDRGLEPALARQVAEQLSAKDV 126
Query: 120 --------------KLPNPYYAAFASAIAFAVGAFVPLLGAAFVKDYKVRLGVVVGVVSL 165
+ P AA SA+ F+VGA +P +G + VV G +
Sbjct: 127 FAAHARDELGLSAHVVARPVQAALTSALTFSVGAALP-IGIVLLAPTGSTSMVVSGGSLI 185
Query: 166 ALFGFGLLSAVLGKAPLVKSSLRVLIGGWLAMSLTFGLTKLVNHVV 211
L G +SA +G A L+K +LRV G LAM+ + + LV H +
Sbjct: 186 CLAILGAVSAHIGGAGLLKPTLRVTFWGALAMAASAAIGALVGHTI 231
>ref|NP_107091.1| similar to nodulin 21 [Mesorhizobium loti MAFF303099]
gi|14026279|dbj|BAB52877.1| mlr6622 [Mesorhizobium loti
MAFF303099]
Length = 231
Score = 103 bits (257), Expect = 3e-21
Identities = 80/227 (35%), Positives = 105/227 (46%), Gaps = 51/227 (22%)
Query: 34 EKDYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVKTMILTGIAGLVAGACSMAIG 93
E R WLRAAVLGANDG++STASL++GV A +++ GIAGLVAGA SMA G
Sbjct: 7 ENHLISRIGWLRAAVLGANDGIVSTASLIVGVAAANAAASNVLVAGIAGLVAGAMSMAAG 66
Query: 94 EFVSVYSQYDIEFAQMKRQ---------------------------------GNISQKDK 120
E+VSV SQ D E A + R+ + KD
Sbjct: 67 EYVSVSSQSDTERADLDRERRELATQPSFERQELADIYVKRGVEPKLALQVADQLMAKDA 126
Query: 121 L---------------PNPYYAAFASAIAFAVGAFVPLLGAAFVKDYKVRLGVVVGVVSL 165
L P AA ASA AF+VGA +PL A + L V V SL
Sbjct: 127 LGAHARDELGISEMTTARPIQAALASAAAFSVGAAMPL--AMVLVSPAAWLAATVSVASL 184
Query: 166 ALFG-FGLLSAVLGKAPLVKSSLRVLIGGWLAMSLTFGLTKLVNHVV 211
G + A G A ++++++RV G LAM+LT + +V +
Sbjct: 185 LFLAVLGAIGAKAGGANVLRATVRVTFWGALAMALTASIGAVVGTAI 231
>ref|YP_157303.1| hypothetical protein ebA544 [Azoarcus sp. EbN1]
gi|56311757|emb|CAI06402.1| conserved hypothetical
protein [Azoarcus sp. EbN1]
Length = 233
Score = 98.2 bits (243), Expect = 1e-19
Identities = 77/235 (32%), Positives = 106/235 (44%), Gaps = 55/235 (23%)
Query: 27 KGEEENDEKDYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVKTMILTGIAGLVAG 86
K + E+ T R W+RAAVLGANDG++STASL++GV A ++ G+AGLVAG
Sbjct: 2 KFARRHTERHRTDRLGWMRAAVLGANDGIVSTASLVVGVAAAGSGQGAALVAGVAGLVAG 61
Query: 87 ACSMAIGEFVSVYSQYDIEFAQMKRQGNISQKDKL------------------------- 121
A SMA GE+VSV+SQ D E A + R+ ++ +
Sbjct: 62 AMSMAAGEYVSVHSQADAENADLSRERTELERQPIAEHRELAAIYIARGLEPGLAHQVAD 121
Query: 122 -----------------------PNPYYAAFASAIAFAVGAFVPLLGAAFVKDYKVRLGV 158
P AA +SA +FAVGA +PL A G+
Sbjct: 122 QLMAHDALGAHARDELGISATLSARPVQAALSSAASFAVGAALPLAVTALAP----AQGL 177
Query: 159 VVGVVSLALFGFGLLSAV---LGKAPLVKSSLRVLIGGWLAMSLTFGLTKLVNHV 210
+ V +L LL AV +G A + + RV G LAM++T G+ L V
Sbjct: 178 IAWVSGTSLAFLALLGAVAARVGGASVPTGAWRVTFWGALAMAVTAGVGALFGAV 232
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.321 0.135 0.380
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 314,919,959
Number of Sequences: 2540612
Number of extensions: 11794014
Number of successful extensions: 48863
Number of sequences better than 10.0: 203
Number of HSP's better than 10.0 without gapping: 129
Number of HSP's successfully gapped in prelim test: 74
Number of HSP's that attempted gapping in prelim test: 48554
Number of HSP's gapped (non-prelim): 311
length of query: 212
length of database: 863,360,394
effective HSP length: 122
effective length of query: 90
effective length of database: 553,405,730
effective search space: 49806515700
effective search space used: 49806515700
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 72 (32.3 bits)
Medicago: description of AC136472.16


