

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136451.2 - phase: 0 /pseudo
(355 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
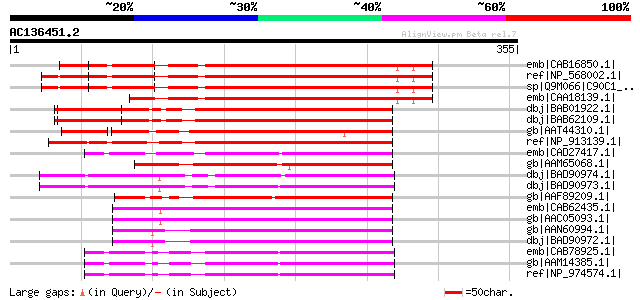
Score E
Sequences producing significant alignments: (bits) Value
emb|CAB16850.1| cytochrome P450 like protein [Arabidopsis thalia... 248 2e-64
ref|NP_568002.1| cytochrome P450 90C1 (CYP90C1) / rotundifolia3 ... 248 2e-64
sp|Q9M066|C90C1_ARATH Cytochrome P450 90C1 (ROTUNDIFOLIA3) gi|41... 248 2e-64
emb|CAA18139.1| cytochrome P450 like protein (fragment) [Arabido... 226 8e-58
dbj|BAB01922.1| cytochrome P450-like protein [Arabidopsis thaliana] 201 2e-50
dbj|BAB62109.1| CYP90D [Arabidopsis thaliana] gi|28827564|gb|AAO... 201 2e-50
gb|AAT44310.1| putative cytochrome P450 [Oryza sativa (japonica ... 191 2e-47
ref|NP_913139.1| cytochrome P450-like protein [Oryza sativa (jap... 187 5e-46
emb|CAD27417.1| cytochrome P450 [Nicotiana tabacum] 144 5e-33
gb|AAM65068.1| cytochrome P450 90A1 [Arabidopsis thaliana] gi|97... 141 3e-32
dbj|BAD90974.1| cytochrome P450 [Oryza sativa (japonica cultivar... 137 6e-31
dbj|BAD90973.1| cytochrome P450 [Oryza sativa (japonica cultivar... 137 6e-31
gb|AAF89209.1| cytochrome P450 [Vigna radiata] 134 5e-30
emb|CAB62435.1| steroid 22-alpha-hydroxylase (DWF4) [Arabidopsis... 130 6e-29
gb|AAC05093.1| steroid 22-alpha-hydroxylase; DWF4; CYP90B1 [Arab... 130 6e-29
gb|AAN60994.1| Putative steroid 22-alpha-hydroxylase [Oryza sati... 124 3e-27
dbj|BAD90972.1| cytochrome P450 [Oryza sativa (japonica cultivar... 124 3e-27
emb|CAB78925.1| cytochrome P450 [Arabidopsis thaliana] gi|304681... 124 4e-27
gb|AAM14385.1| putative cytochrome P450 protein [Arabidopsis tha... 124 4e-27
ref|NP_974574.1| cytochrome P450 family protein [Arabidopsis tha... 124 4e-27
>emb|CAB16850.1| cytochrome P450 like protein [Arabidopsis thaliana]
gi|7270586|emb|CAB80304.1| cytochrome P450 like protein
[Arabidopsis thaliana] gi|25282635|pir||D85429
cytochrome P450 like protein [imported] - Arabidopsis
thaliana
Length = 457
Score = 248 bits (634), Expect = 2e-64
Identities = 134/247 (54%), Positives = 170/247 (68%), Gaps = 21/247 (8%)
Query: 56 GYSSCPVTFMEKRKSILYKSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVNDVV 115
G P+ F R LYKSLKAKER++KMV ++VEER + + NDVV
Sbjct: 177 GLICIPIKFPGTR---LYKSLKAKERLIKMVKKVVEERQVAMTTTSP--------ANDVV 225
Query: 116 DALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKL 175
D LLRD G+ + S S + +S I+E MIPGEET+PTAMTLA+KFL+D P+AL+KL
Sbjct: 226 DVLLRDGGDSEKQSQPS----DFVSGKIVEMMIPGEETMPTAMTLAVKFLSDNPVALAKL 281
Query: 176 MEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYL 235
+EENME+K++K + Y WTDYMSL FTQNVI+ETLRMANI+NG+WRKA+KDVEIKGYL
Sbjct: 282 VEENMEMKRRKLELGEEYKWTDYMSLSFTQNVINETLRMANIINGVWRKALKDVEIKGYL 341
Query: 236 IPKDWGVMASLTSVHMDSKNYENPYKFDPWRWEII--YLHEAIIYTMF----YACRKLEL 289
IPK W V+AS SVHMD Y+NPY+FDPWRW+ I + +I +T F C LEL
Sbjct: 342 IPKGWCVLASFISVHMDEDIYDNPYQFDPWRWDRINGSANSSICFTPFGGGQRLCPGLEL 401
Query: 290 CLVTIAL 296
+ I++
Sbjct: 402 SKLEISI 408
Score = 62.4 bits (150), Expect = 2e-08
Identities = 31/66 (46%), Positives = 43/66 (64%)
Query: 36 IPKGNSGWPLLGETLDFIASGYSSCPVTFMEKRKSILYKSLKAKERMMKMVGRIVEERMK 95
IP G+ GWP++GETL+FIA GYSS PVTFM+KRKS+ K K ++ E K
Sbjct: 2 IPNGSLGWPVIGETLNFIACGYSSRPVTFMDKRKSLYGKVFKTNIIGTPIIISTDAEVNK 61
Query: 96 LIMDNN 101
+++ N+
Sbjct: 62 VVLQNH 67
>ref|NP_568002.1| cytochrome P450 90C1 (CYP90C1) / rotundifolia3 (ROT3) [Arabidopsis
thaliana]
Length = 524
Score = 248 bits (634), Expect = 2e-64
Identities = 134/247 (54%), Positives = 170/247 (68%), Gaps = 21/247 (8%)
Query: 56 GYSSCPVTFMEKRKSILYKSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVNDVV 115
G P+ F R LYKSLKAKER++KMV ++VEER + + NDVV
Sbjct: 244 GLICIPIKFPGTR---LYKSLKAKERLIKMVKKVVEERQVAMTTTSP--------ANDVV 292
Query: 116 DALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKL 175
D LLRD G+ + S S + +S I+E MIPGEET+PTAMTLA+KFL+D P+AL+KL
Sbjct: 293 DVLLRDGGDSEKQSQPS----DFVSGKIVEMMIPGEETMPTAMTLAVKFLSDNPVALAKL 348
Query: 176 MEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYL 235
+EENME+K++K + Y WTDYMSL FTQNVI+ETLRMANI+NG+WRKA+KDVEIKGYL
Sbjct: 349 VEENMEMKRRKLELGEEYKWTDYMSLSFTQNVINETLRMANIINGVWRKALKDVEIKGYL 408
Query: 236 IPKDWGVMASLTSVHMDSKNYENPYKFDPWRWEII--YLHEAIIYTMF----YACRKLEL 289
IPK W V+AS SVHMD Y+NPY+FDPWRW+ I + +I +T F C LEL
Sbjct: 409 IPKGWCVLASFISVHMDEDIYDNPYQFDPWRWDRINGSANSSICFTPFGGGQRLCPGLEL 468
Query: 290 CLVTIAL 296
+ I++
Sbjct: 469 SKLEISI 475
Score = 65.1 bits (157), Expect = 3e-09
Identities = 34/79 (43%), Positives = 52/79 (65%), Gaps = 1/79 (1%)
Query: 23 KEKNNIKLRNGIKIPKGNSGWPLLGETLDFIASGYSSCPVTFMEKRKSILYKSLKAKERM 82
+E++N + + G+ IP G+ GWP++GETL+FIA GYSS PVTFM+KRKS+ K K
Sbjct: 57 EEEDNEEKKKGM-IPNGSLGWPVIGETLNFIACGYSSRPVTFMDKRKSLYGKVFKTNIIG 115
Query: 83 MKMVGRIVEERMKLIMDNN 101
++ E K+++ N+
Sbjct: 116 TPIIISTDAEVNKVVLQNH 134
>sp|Q9M066|C90C1_ARATH Cytochrome P450 90C1 (ROTUNDIFOLIA3) gi|4176420|dbj|BAA37167.1|
cytochrome P450 [Arabidopsis thaliana]
Length = 524
Score = 248 bits (634), Expect = 2e-64
Identities = 134/247 (54%), Positives = 170/247 (68%), Gaps = 21/247 (8%)
Query: 56 GYSSCPVTFMEKRKSILYKSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVNDVV 115
G P+ F R LYKSLKAKER++KMV ++VEER + + NDVV
Sbjct: 244 GLICIPIKFPGTR---LYKSLKAKERLIKMVKKVVEERQVAMTTTSP--------ANDVV 292
Query: 116 DALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKL 175
D LLRD G+ + S S + +S I+E MIPGEET+PTAMTLA+KFL+D P+AL+KL
Sbjct: 293 DVLLRDGGDSEKQSQPS----DFVSGKIVEMMIPGEETMPTAMTLAVKFLSDNPVALAKL 348
Query: 176 MEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYL 235
+EENME+K++K + Y WTDYMSL FTQNVI+ETLRMANI+NG+WRKA+KDVEIKGYL
Sbjct: 349 VEENMEMKRRKLELGEEYKWTDYMSLSFTQNVINETLRMANIINGVWRKALKDVEIKGYL 408
Query: 236 IPKDWGVMASLTSVHMDSKNYENPYKFDPWRWEII--YLHEAIIYTMF----YACRKLEL 289
IPK W V+AS SVHMD Y+NPY+FDPWRW+ I + +I +T F C LEL
Sbjct: 409 IPKGWCVLASFISVHMDEDIYDNPYQFDPWRWDRINGSANSSICFTPFGGGQRLCPGLEL 468
Query: 290 CLVTIAL 296
+ I++
Sbjct: 469 SKLEISI 475
Score = 65.1 bits (157), Expect = 3e-09
Identities = 34/79 (43%), Positives = 52/79 (65%), Gaps = 1/79 (1%)
Query: 23 KEKNNIKLRNGIKIPKGNSGWPLLGETLDFIASGYSSCPVTFMEKRKSILYKSLKAKERM 82
+E++N + + G+ IP G+ GWP++GETL+FIA GYSS PVTFM+KRKS+ K K
Sbjct: 57 EEEDNEEKKKGM-IPNGSLGWPVIGETLNFIACGYSSRPVTFMDKRKSLYGKVFKTNIIG 115
Query: 83 MKMVGRIVEERMKLIMDNN 101
++ E K+++ N+
Sbjct: 116 TPIIISTDAEVNKVVLQNH 134
>emb|CAA18139.1| cytochrome P450 like protein (fragment) [Arabidopsis thaliana]
gi|7484897|pir||T04602 cytochrome P450 homolog
F23E13.220 - Arabidopsis thaliana
Length = 255
Score = 226 bits (576), Expect = 8e-58
Identities = 119/218 (54%), Positives = 152/218 (69%), Gaps = 18/218 (8%)
Query: 85 MVGRIVEERMKLIMDNNAEDDEKVGVVNDVVDALLRDKGELSQSSSSSNLMVEMISQNII 144
MV ++VEER + + NDVVD LLRD G+ + S S + +S I+
Sbjct: 1 MVKKVVEERQVAMTTTSP--------ANDVVDVLLRDGGDSEKQSQPS----DFVSGKIV 48
Query: 145 EFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGYTWTDYMSLPFT 204
E MIPGEET+PTAMTLA+KFL+D P+AL+KL+EENME+K++K + Y WTDYMSL FT
Sbjct: 49 EMMIPGEETMPTAMTLAVKFLSDNPVALAKLVEENMEMKRRKLELGEEYKWTDYMSLSFT 108
Query: 205 QNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDP 264
QNVI+ETLRMANI+NG+WRKA+KDVEIKGYLIPK W V+AS SVHMD Y+NPY+FDP
Sbjct: 109 QNVINETLRMANIINGVWRKALKDVEIKGYLIPKGWCVLASFISVHMDEDIYDNPYQFDP 168
Query: 265 WRWEII--YLHEAIIYTMF----YACRKLELCLVTIAL 296
WRW+ I + +I +T F C LEL + I++
Sbjct: 169 WRWDRINGSANSSICFTPFGGGQRLCPGLELSKLEISI 206
>dbj|BAB01922.1| cytochrome P450-like protein [Arabidopsis thaliana]
Length = 464
Score = 201 bits (512), Expect = 2e-50
Identities = 112/235 (47%), Positives = 153/235 (64%), Gaps = 16/235 (6%)
Query: 34 IKIPKGNSGWPLLGETLDFIASGYSSCPVTFMEKRKSILYKSLKAKERMMKMVGRIVEER 93
I + KG L E +FI SG S P+ F + L++SL+AK+ M+K V RI+E +
Sbjct: 205 ISVEKGEDLEELKREFENFI-SGLMSLPINFPGTQ---LHRSLQAKKNMVKQVERIIEGK 260
Query: 94 MKLIMDNNAEDDEKVGVVNDVVDALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEET 153
++ N EDD + DVVD LL+D SS +L +I+ N+I+ MIPG ++
Sbjct: 261 IRKTK-NKEEDDV---IAKDVVDVLLKD--------SSEHLTHNLIANNMIDMMIPGHDS 308
Query: 154 LPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLR 213
+P +TLA+KFL+D P AL+ L EENM+LK K + W DY+SLPFTQ VI+ETLR
Sbjct: 309 VPVLITLAVKFLSDSPAALNLLTEENMKLKSLKELTGEPLYWNDYLSLPFTQKVITETLR 368
Query: 214 MANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRWE 268
M N++ G+ RKA+KDVEIKGY+IPK W +A L SVH+D YE+PYKF+PWRW+
Sbjct: 369 MGNVIIGVMRKAMKDVEIKGYVIPKGWCFLAYLRSVHLDKLYYESPYKFNPWRWQ 423
Score = 51.2 bits (121), Expect = 5e-05
Identities = 20/47 (42%), Positives = 34/47 (71%)
Query: 32 NGIKIPKGNSGWPLLGETLDFIASGYSSCPVTFMEKRKSILYKSLKA 78
+G K P G+ GWP++GET++F++S YS P +FM+KR+ + + K+
Sbjct: 47 HGPKFPHGSLGWPVIGETIEFVSSAYSDRPESFMDKRRLMYGRVFKS 93
>dbj|BAB62109.1| CYP90D [Arabidopsis thaliana] gi|28827564|gb|AAO50626.1| putative
cytochrome P450 [Arabidopsis thaliana]
gi|28393374|gb|AAO42111.1| putative cytochrome P450
[Arabidopsis thaliana] gi|18400142|ref|NP_566462.1|
cytochrome P450, putative [Arabidopsis thaliana]
Length = 491
Score = 201 bits (512), Expect = 2e-50
Identities = 112/235 (47%), Positives = 153/235 (64%), Gaps = 16/235 (6%)
Query: 34 IKIPKGNSGWPLLGETLDFIASGYSSCPVTFMEKRKSILYKSLKAKERMMKMVGRIVEER 93
I + KG L E +FI SG S P+ F + L++SL+AK+ M+K V RI+E +
Sbjct: 205 ISVEKGEDLEELKREFENFI-SGLMSLPINFPGTQ---LHRSLQAKKNMVKQVERIIEGK 260
Query: 94 MKLIMDNNAEDDEKVGVVNDVVDALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEET 153
++ N EDD + DVVD LL+D SS +L +I+ N+I+ MIPG ++
Sbjct: 261 IRKTK-NKEEDDV---IAKDVVDVLLKD--------SSEHLTHNLIANNMIDMMIPGHDS 308
Query: 154 LPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLR 213
+P +TLA+KFL+D P AL+ L EENM+LK K + W DY+SLPFTQ VI+ETLR
Sbjct: 309 VPVLITLAVKFLSDSPAALNLLTEENMKLKSLKELTGEPLYWNDYLSLPFTQKVITETLR 368
Query: 214 MANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRWE 268
M N++ G+ RKA+KDVEIKGY+IPK W +A L SVH+D YE+PYKF+PWRW+
Sbjct: 369 MGNVIIGVMRKAMKDVEIKGYVIPKGWCFLAYLRSVHLDKLYYESPYKFNPWRWQ 423
Score = 51.2 bits (121), Expect = 5e-05
Identities = 20/47 (42%), Positives = 34/47 (71%)
Query: 32 NGIKIPKGNSGWPLLGETLDFIASGYSSCPVTFMEKRKSILYKSLKA 78
+G K P G+ GWP++GET++F++S YS P +FM+KR+ + + K+
Sbjct: 47 HGPKFPHGSLGWPVIGETIEFVSSAYSDRPESFMDKRRLMYGRVFKS 93
>gb|AAT44310.1| putative cytochrome P450 [Oryza sativa (japonica cultivar-group)]
Length = 503
Score = 191 bits (486), Expect = 2e-47
Identities = 96/200 (48%), Positives = 140/200 (70%), Gaps = 13/200 (6%)
Query: 72 LYKSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVNDVVDALLRDKGELSQSSSS 131
LY+SLKAK+RM ++ I++E+ + I E + V D++D L+ S S
Sbjct: 239 LYRSLKAKKRMTSLIQNIIQEKRRRIF----EGKDLCAVSRDLIDVLM------SNGSDE 288
Query: 132 SNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPD 191
+L E+IS N+I+FMIP E+++P +TLA+K+L++ PLAL +L EENMELK+QK++ +
Sbjct: 289 LSLTDELISDNMIDFMIPAEDSVPVLITLAIKYLSECPLALQQLEEENMELKRQKSDVGE 348
Query: 192 GYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKG---YLIPKDWGVMASLTS 248
WTDYMSL FTQ+VI+ETLR+ NI++GI RKAV+DVE+KG +IPK W V+ S
Sbjct: 349 TLEWTDYMSLTFTQHVITETLRIGNIISGIMRKAVRDVEVKGQGDVVIPKGWCVLVYFRS 408
Query: 249 VHMDSKNYENPYKFDPWRWE 268
VH+D+ Y++PY F+PWRW+
Sbjct: 409 VHLDANIYDDPYAFNPWRWK 428
Score = 47.8 bits (112), Expect = 5e-04
Identities = 20/32 (62%), Positives = 26/32 (80%)
Query: 37 PKGNSGWPLLGETLDFIASGYSSCPVTFMEKR 68
P G+ GWPL+GETL FI++ YSS P +F+EKR
Sbjct: 46 PAGSLGWPLVGETLQFISAAYSSRPESFVEKR 77
>ref|NP_913139.1| cytochrome P450-like protein [Oryza sativa (japonica
cultivar-group)] gi|14209594|dbj|BAB56089.1| putative
cytochrome P450 90C1 [Oryza sativa (japonica
cultivar-group)]
Length = 490
Score = 187 bits (474), Expect = 5e-46
Identities = 103/241 (42%), Positives = 147/241 (60%), Gaps = 20/241 (8%)
Query: 28 IKLRNGIKIPKGNSGWPLLGETLDFIASGYSSCPVTFMEKRKSILYKSLKAKERMMKMVG 87
I +R I + G L + +FI G S P+ R LY+SL+AK++M +++
Sbjct: 198 ILVRGLIGLEAGEEMQQLKQQFQEFIV-GLMSLPIKLPGTR---LYRSLQAKKKMARLIQ 253
Query: 88 RIVEERMKLIMDNNAEDDEKVGVVNDVVDALLRDKGELSQSSSSSNLMVEMISQNIIEFM 147
RI+ E+ D +D L+ D S L E+IS N+I+ M
Sbjct: 254 RIIREKRAR--------RAAASPPRDAIDVLIGD--------GSDELTDELISDNMIDLM 297
Query: 148 IPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGYTWTDYMSLPFTQNV 207
IP E+++P +TLA+KFL++ PLAL +L EEN++LK++KT+ + WTDYMSL FTQ+V
Sbjct: 298 IPAEDSVPVLITLAVKFLSECPLALHQLEEENIQLKRRKTDMGETLQWTDYMSLSFTQHV 357
Query: 208 ISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRW 267
I+ETLR+ NI+ GI RKAV+DVE+KG+LIPK W V SVH+D Y+ PYKF+PWRW
Sbjct: 358 ITETLRLGNIIGGIMRKAVRDVEVKGHLIPKGWCVFVYFRSVHLDDTLYDEPYKFNPWRW 417
Query: 268 E 268
+
Sbjct: 418 K 418
Score = 45.8 bits (107), Expect = 0.002
Identities = 16/37 (43%), Positives = 28/37 (75%)
Query: 35 KIPKGNSGWPLLGETLDFIASGYSSCPVTFMEKRKSI 71
++P G+ GWP++GETL+F++ YS P F++KR+ +
Sbjct: 48 RLPPGSFGWPVVGETLEFVSCAYSPRPEAFVDKRRKL 84
>emb|CAD27417.1| cytochrome P450 [Nicotiana tabacum]
Length = 483
Score = 144 bits (362), Expect = 5e-33
Identities = 78/216 (36%), Positives = 128/216 (59%), Gaps = 18/216 (8%)
Query: 53 IASGYSSCPVTFMEKRKSILYKSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVN 112
+ G+ + P+ F S K+++A+ ++ + +G +V+ER K E + N
Sbjct: 206 VIEGFFTIPLPFFS---STYRKAIQARRKVAEALGLVVKERRK-------ERGGGERLKN 255
Query: 113 DVVDALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLAL 172
D+++AL G S E+I ++ ++ G ET T MTLA+KFLT+ P AL
Sbjct: 256 DMLEALFEGDGVEGFSD-------EVIVDFMLALLVAGYETTSTIMTLAVKFLTETPHAL 308
Query: 173 SKLMEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIK 232
S L EE+ E++ +K + + W DY S+PFTQ V++ETLR+ NI++G++R+ + D+ IK
Sbjct: 309 SLLKEEHEEIRLRKGDV-ESLLWEDYKSMPFTQCVVNETLRVGNIISGVFRRTMTDINIK 367
Query: 233 GYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRWE 268
GY IPK W V A +VH+D +++++ FDPWRW+
Sbjct: 368 GYTIPKGWKVFACFRAVHLDHEHFKDARTFDPWRWQ 403
Score = 33.9 bits (76), Expect = 7.6
Identities = 18/70 (25%), Positives = 36/70 (50%)
Query: 1 MEWFIGVYVSIIVLLMSLWFVKKEKNNIKLRNGIKIPKGNSGWPLLGETLDFIASGYSSC 60
M++ I +++S + L++ ++ R ++P G G P +GETL I++ +
Sbjct: 1 MDFIIYLFLSFSISLITFLLLRAAAAAHFRRRKTRLPPGTLGLPFIGETLQLISAYKTEN 60
Query: 61 PVTFMEKRKS 70
P F++ R S
Sbjct: 61 PEPFIDDRVS 70
>gb|AAM65068.1| cytochrome P450 90A1 [Arabidopsis thaliana]
gi|9759094|dbj|BAB09663.1| cytochrome P450 90A1
[Arabidopsis thaliana] gi|871988|emb|CAA60794.1| CYP90
protein [Arabidopsis thaliana] gi|853719|emb|CAA60793.1|
CYP90 protein [Arabidopsis thaliana]
gi|20148303|gb|AAM10042.1| cytochrome P450 90A1
[Arabidopsis thaliana] gi|17380618|gb|AAL36072.1|
AT5g05690/MJJ3_9 [Arabidopsis thaliana]
gi|15239203|ref|NP_196188.1| cytochrome P450 90A1
(CYP90A1) (CYP90) (CPD) [Arabidopsis thaliana]
gi|15450717|gb|AAK96630.1| AT5g05690/MJJ3_9 [Arabidopsis
thaliana] gi|14596099|gb|AAK68777.1| cytochrome P450
90A1 [Arabidopsis thaliana]
gi|5915851|sp|Q42569|C90A1_ARATH Cytochrome P450 90A1
Length = 472
Score = 141 bits (355), Expect = 3e-32
Identities = 72/183 (39%), Positives = 115/183 (62%), Gaps = 14/183 (7%)
Query: 88 RIVEERMKLIMDNNAEDDEKVGVVNDVVDALLRDKGELSQSSSSSNLMVEMISQNIIEFM 147
++ E ++M E++E D++ ALL ++ E I ++ +
Sbjct: 226 KVAEALTVVVMKRREEEEEGAERKKDMLAALL---------AADDGFSDEEIVDFLVALL 276
Query: 148 IPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGYT--WTDYMSLPFTQ 205
+ G ET T MTLA+KFLT+ PLAL++L EE+ +++ K+ D Y+ W+DY S+PFTQ
Sbjct: 277 VAGYETTSTIMTLAVKFLTETPLALAQLKEEHEKIRAMKS---DSYSLEWSDYKSMPFTQ 333
Query: 206 NVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPW 265
V++ETLR+ANI+ G++R+A+ DVEIKGY IPK W V +S +VH+D ++++ F+PW
Sbjct: 334 CVVNETLRVANIIGGVFRRAMTDVEIKGYKIPKGWKVFSSFRAVHLDPNHFKDARTFNPW 393
Query: 266 RWE 268
RW+
Sbjct: 394 RWQ 396
>dbj|BAD90974.1| cytochrome P450 [Oryza sativa (japonica cultivar-group)]
Length = 501
Score = 137 bits (344), Expect = 6e-31
Identities = 88/252 (34%), Positives = 141/252 (55%), Gaps = 15/252 (5%)
Query: 22 KKEKNNIKLRNGIKIPKGNSGWPLLGETLDFIASGYSSCPVTFMEKRKSILY-KSLKAKE 80
KK N+ ++ + I G L E + I G+ S P Y ++LKA++
Sbjct: 183 KKITFNLTVKQLVSIEPGPWTESLRREYVKLI-DGFFSIPFPLAYFLPFTTYGQALKARK 241
Query: 81 RMMKMVGRIVEERMKLIMDNNAE--DDEKVGVVNDVVDALLRDKGELSQSSSSSNLMVEM 138
++ + ++++RM+ +N DDE D+V+ LL+ +G S S MV+
Sbjct: 242 KVAGALREVIKKRMEEKAENGGSIGDDEGKKEKKDMVEELLQAEG----GSFSEEEMVDF 297
Query: 139 ISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQK-TNCPDGYTWTD 197
+ ++ G ET MTLA+KFLT+ P AL++L EE+ ++ K N P W+D
Sbjct: 298 C----LSLLVAGYETTSVLMTLAVKFLTETPAALAELKEEHANIRDMKGKNQP--LEWSD 351
Query: 198 YMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYE 257
Y S+PFTQ VI+ETLR+ NI++G++R+A D+ K Y IPK + AS +VH+++++YE
Sbjct: 352 YKSMPFTQCVINETLRVGNIISGVFRRANTDIHYKDYTIPKGCKIFASFRAVHLNNEHYE 411
Query: 258 NPYKFDPWRWEI 269
N F+PWRW+I
Sbjct: 412 NARTFNPWRWQI 423
>dbj|BAD90973.1| cytochrome P450 [Oryza sativa (japonica cultivar-group)]
Length = 501
Score = 137 bits (344), Expect = 6e-31
Identities = 86/251 (34%), Positives = 138/251 (54%), Gaps = 13/251 (5%)
Query: 22 KKEKNNIKLRNGIKIPKGNSGWPLLGETLDFIASGYSSCPVTFMEKRKSILY-KSLKAKE 80
KK N+ ++ + I G L E + I G+ S P Y ++LKA++
Sbjct: 183 KKITFNLTVKQLVSIEPGPWTESLRREYVKLI-DGFFSIPFPLANLLPFTTYGQALKARK 241
Query: 81 RMMKMVGRIVEERMKLIMDNNAE--DDEKVGVVNDVVDALLRDKGELSQSSSSSNLMVEM 138
++ + ++++RM+ +N DDE D+V+ LL +G S S MV+
Sbjct: 242 KVAGALREVIKKRMEEKAENGGSIGDDEGKKEKKDMVEELLEAEG----GSFSEEEMVDF 297
Query: 139 ISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGYTWTDY 198
+ ++ G ET MTLA+KFLT+ P AL++L EE+ ++ K W+DY
Sbjct: 298 C----LSLLVAGYETTSMLMTLAVKFLTETPAALAELKEEHANIRDMKGK-KQPLEWSDY 352
Query: 199 MSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYEN 258
S+PFTQ VI+ETLR+ NI++G++R+A D+ K Y IPK + AS +VH+++++YEN
Sbjct: 353 KSMPFTQCVINETLRVGNIISGVFRRANTDIHYKDYTIPKGCKIFASFRAVHLNNEHYEN 412
Query: 259 PYKFDPWRWEI 269
F+PWRW+I
Sbjct: 413 ARTFNPWRWQI 423
>gb|AAF89209.1| cytochrome P450 [Vigna radiata]
Length = 474
Score = 134 bits (336), Expect = 5e-30
Identities = 71/195 (36%), Positives = 124/195 (63%), Gaps = 17/195 (8%)
Query: 74 KSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVNDVVDALLRDKGELSQSSSSSN 133
+++KA+ ++ + + +V +R + + N ++K +D++ ALL +S +
Sbjct: 219 RAIKARTKVAEALTLVVRQRRE---EYNQGKEKK----SDMLGALL---------ASGDH 262
Query: 134 LMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGY 193
+ I ++ ++ G ET T MTLA+KFLT+ PLAL++L EE+ +++ +++
Sbjct: 263 FSDDQIVDFLLALLVAGYETTSTIMTLAVKFLTETPLALAQLKEEHDQIRA-RSDPGAPL 321
Query: 194 TWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDS 253
WTDY S+ FTQ+V++ETLR+ANI+ GI+R+A D++IKGY IPK W V AS +VH++
Sbjct: 322 EWTDYKSMVFTQHVVNETLRVANIIGGIFRRATTDIDIKGYTIPKGWKVFASFRAVHLNP 381
Query: 254 KNYENPYKFDPWRWE 268
+ Y++ F+PWRW+
Sbjct: 382 EYYKDARTFNPWRWQ 396
Score = 35.4 bits (80), Expect = 2.6
Identities = 16/38 (42%), Positives = 24/38 (63%)
Query: 31 RNGIKIPKGNSGWPLLGETLDFIASGYSSCPVTFMEKR 68
R ++P G+ G P +GETL I++ SS P FM++R
Sbjct: 26 RRKFRLPPGSYGLPFIGETLQLISAYKSSNPEPFMDER 63
>emb|CAB62435.1| steroid 22-alpha-hydroxylase (DWF4) [Arabidopsis thaliana]
gi|19699122|gb|AAL90927.1| AT3g50660/T3A5_40
[Arabidopsis thaliana] gi|15724348|gb|AAL06567.1|
AT3g50660/T3A5_40 [Arabidopsis thaliana]
gi|15229822|ref|NP_190635.1| steroid
22-alpha-hydroxylase (CYP90B1) (DWF4) [Arabidopsis
thaliana] gi|11249706|pir||T46143 steroid
22-alpha-hydroxylase (DWF4) - Arabidopsis thaliana
Length = 513
Score = 130 bits (327), Expect = 6e-29
Identities = 73/203 (35%), Positives = 112/203 (54%), Gaps = 7/203 (3%)
Query: 73 YKSLKAKERMMKMVGRIVEERMKLIMDNNAED------DEKVGVVNDVVDALLRDKGELS 126
+K+L+++ ++K + R +EER I + + E+ DE +D V D L
Sbjct: 229 HKALQSRATILKFIERKMEERKLDIKEEDQEEEEVKTEDEAEMSKSDHVRKQRTDDDLLG 288
Query: 127 QSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQK 186
SNL E I I+ + G ET A+ LA+ FL P A+ +L EE++E+ + K
Sbjct: 289 WVLKHSNLSTEQILDLILSLLFAGHETSSVAIALAIFFLQACPKAVEELREEHLEIARAK 348
Query: 187 TNCPDG-YTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMAS 245
+ W DY + FTQ VI+ETLR+ N+V + RKA+KDV KGY IP W V+
Sbjct: 349 KELGESELNWDDYKKMDFTQCVINETLRLGNVVRFLHRKALKDVRYKGYDIPSGWKVLPV 408
Query: 246 LTSVHMDSKNYENPYKFDPWRWE 268
+++VH+D+ Y+ P F+PWRW+
Sbjct: 409 ISAVHLDNSRYDQPNLFNPWRWQ 431
Score = 35.4 bits (80), Expect = 2.6
Identities = 18/58 (31%), Positives = 32/58 (55%), Gaps = 2/58 (3%)
Query: 13 VLLMSLWFVKKEKNNIKLRNGIKIPKGNSGWPLLGETLDFIASGYSSCPVTFMEKRKS 70
+L + L+ + ++ N K R +P G SGWP LGET+ ++ ++ FM++ S
Sbjct: 18 LLSLLLFLILLKRRNRKTR--FNLPPGKSGWPFLGETIGYLKPYTATTLGDFMQQHVS 73
>gb|AAC05093.1| steroid 22-alpha-hydroxylase; DWF4; CYP90B1 [Arabidopsis thaliana]
Length = 513
Score = 130 bits (327), Expect = 6e-29
Identities = 73/203 (35%), Positives = 112/203 (54%), Gaps = 7/203 (3%)
Query: 73 YKSLKAKERMMKMVGRIVEERMKLIMDNNAED------DEKVGVVNDVVDALLRDKGELS 126
+K+L+++ ++K + R +EER I + + E+ DE +D V D L
Sbjct: 229 HKALQSRATILKFIERKMEERKLDIKEEDQEEEEVKTEDEAEMSKSDHVRKQRTDDDLLG 288
Query: 127 QSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQK 186
SNL E I I+ + G ET A+ LA+ FL P A+ +L EE++E+ + K
Sbjct: 289 WVLKHSNLSTEQILDLILSLLFAGHETSSVAIALAIFFLQACPKAVEELREEHLEIARAK 348
Query: 187 TNCPDG-YTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMAS 245
+ W DY + FTQ VI+ETLR+ N+V + RKA+KDV KGY IP W V+
Sbjct: 349 KELGESELNWDDYKKMDFTQCVINETLRLGNVVRFLHRKALKDVRYKGYDIPSGWKVLPV 408
Query: 246 LTSVHMDSKNYENPYKFDPWRWE 268
+++VH+D+ Y+ P F+PWRW+
Sbjct: 409 ISAVHLDNSRYDQPNLFNPWRWQ 431
Score = 35.4 bits (80), Expect = 2.6
Identities = 18/58 (31%), Positives = 32/58 (55%), Gaps = 2/58 (3%)
Query: 13 VLLMSLWFVKKEKNNIKLRNGIKIPKGNSGWPLLGETLDFIASGYSSCPVTFMEKRKS 70
+L + L+ + ++ N K R +P G SGWP LGET+ ++ ++ FM++ S
Sbjct: 18 LLSLLLFLILLKRRNRKTR--FNLPPGKSGWPFLGETIGYLKPYTATTLGDFMQQHVS 73
>gb|AAN60994.1| Putative steroid 22-alpha-hydroxylase [Oryza sativa (japonica
cultivar-group)] gi|34902330|ref|NP_912511.1| Putative
steroid 22-alpha-hydroxylase [Oryza sativa (japonica
cultivar-group)]
Length = 502
Score = 124 bits (312), Expect = 3e-27
Identities = 68/199 (34%), Positives = 112/199 (56%), Gaps = 20/199 (10%)
Query: 73 YKSLKAKERMMKMVGRIVEERMKLIM--DNNAEDDEKVGVVNDVVDALLRDKGELSQSSS 130
+K+LK++ ++ ++ R +EER++ + D + E D+ +G +
Sbjct: 240 WKALKSRAAILGVIERKMEERVEKLSKEDASVEQDDLLG-----------------WALK 282
Query: 131 SSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMEL-KKQKTNC 189
SNL E I ++ + G ET A+ LA+ FL P A+ +L EE++ + ++Q+
Sbjct: 283 QSNLSKEQILDLLLSLLFAGHETSSMALALAIFFLEGCPKAVQELREEHLGIARRQRLRG 342
Query: 190 PDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSV 249
+W DY + FTQ VI+ETLR+ N+V + RK +KDV KGY IP W ++ L +V
Sbjct: 343 ECKLSWEDYKEMVFTQCVINETLRLGNVVRFLHRKVIKDVHYKGYDIPSGWKILPVLAAV 402
Query: 250 HMDSKNYENPYKFDPWRWE 268
H+DS YE+P +F+PWRW+
Sbjct: 403 HLDSSLYEDPQRFNPWRWK 421
Score = 33.9 bits (76), Expect = 7.6
Identities = 18/54 (33%), Positives = 30/54 (55%), Gaps = 3/54 (5%)
Query: 31 RNGIKIPKGNSGWPLLGETLDFIASGYSSCPVTFMEK---RKSILYKSLKAKER 81
R + +P G +GWPL+GET ++ + ++ FME+ R +Y+S ER
Sbjct: 45 RKRMNLPPGAAGWPLVGETFGYLRAHPATSVGRFMEQHIARYGKIYRSSLFGER 98
>dbj|BAD90972.1| cytochrome P450 [Oryza sativa (japonica cultivar-group)]
Length = 506
Score = 124 bits (312), Expect = 3e-27
Identities = 68/199 (34%), Positives = 112/199 (56%), Gaps = 20/199 (10%)
Query: 73 YKSLKAKERMMKMVGRIVEERMKLIM--DNNAEDDEKVGVVNDVVDALLRDKGELSQSSS 130
+K+LK++ ++ ++ R +EER++ + D + E D+ +G +
Sbjct: 244 WKALKSRAAILGVIERKMEERVEKLSKEDASVEQDDLLG-----------------WALK 286
Query: 131 SSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMEL-KKQKTNC 189
SNL E I ++ + G ET A+ LA+ FL P A+ +L EE++ + ++Q+
Sbjct: 287 QSNLSKEQILDLLLSLLFAGHETSSMALALAIFFLEGCPKAVQELREEHLGIARRQRLRG 346
Query: 190 PDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSV 249
+W DY + FTQ VI+ETLR+ N+V + RK +KDV KGY IP W ++ L +V
Sbjct: 347 ECKLSWEDYKEMVFTQCVINETLRLGNVVRFLHRKVIKDVHYKGYDIPSGWKILPVLAAV 406
Query: 250 HMDSKNYENPYKFDPWRWE 268
H+DS YE+P +F+PWRW+
Sbjct: 407 HLDSSLYEDPQRFNPWRWK 425
Score = 33.9 bits (76), Expect = 7.6
Identities = 18/54 (33%), Positives = 30/54 (55%), Gaps = 3/54 (5%)
Query: 31 RNGIKIPKGNSGWPLLGETLDFIASGYSSCPVTFMEK---RKSILYKSLKAKER 81
R + +P G +GWPL+GET ++ + ++ FME+ R +Y+S ER
Sbjct: 49 RKRMNLPPGAAGWPLVGETFGYLRAHPATSVGRFMEQHIARYGKIYRSSLFGER 102
>emb|CAB78925.1| cytochrome P450 [Arabidopsis thaliana] gi|3046815|emb|CAA16713.1|
cytochrome P450 [Arabidopsis thaliana]
gi|7430728|pir||T04444 cytochrome P450 - Arabidopsis
thaliana
Length = 457
Score = 124 bits (311), Expect = 4e-27
Identities = 73/217 (33%), Positives = 120/217 (54%), Gaps = 24/217 (11%)
Query: 53 IASGYSSCPVTFMEKRKSILYKSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVN 112
+ GY+S PV ++ +KS+KA++ + +++ RI+ ER + N + N
Sbjct: 202 LEKGYNSMPVNLPG---TLFHKSMKARKELSQILARILSERRQ----NGSSH-------N 247
Query: 113 DVVDALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLAL 172
D++ + + DK EL+ E I+ NII + +T + M+ LK+L + P L
Sbjct: 248 DLLGSFMGDKEELTD---------EQIADNIIGVIFAARDTTASVMSWILKYLAENPNVL 298
Query: 173 SKLMEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIK 232
+ EE M ++K K + TW D +P T VI ETLR+A+I++ +R+AV+DVE +
Sbjct: 299 EAVTEEQMAIRKDKEE-GESLTWGDTKKMPLTSRVIQETLRVASILSFTFREAVEDVEYE 357
Query: 233 GYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRWEI 269
GYLIPK W V+ ++H + + NP KFDP R+E+
Sbjct: 358 GYLIPKGWKVLPLFRNIHHSADIFSNPGKFDPSRFEV 394
>gb|AAM14385.1| putative cytochrome P450 protein [Arabidopsis thaliana]
gi|15293093|gb|AAK93657.1| putative cytochrome P450
protein [Arabidopsis thaliana]
gi|18415271|ref|NP_567581.1| cytochrome P450 family
protein [Arabidopsis thaliana]
gi|46401564|dbj|BAD16629.1| cytochrome P450
monooxygenase [Arabidopsis thaliana]
Length = 467
Score = 124 bits (311), Expect = 4e-27
Identities = 73/217 (33%), Positives = 120/217 (54%), Gaps = 24/217 (11%)
Query: 53 IASGYSSCPVTFMEKRKSILYKSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVN 112
+ GY+S PV ++ +KS+KA++ + +++ RI+ ER + N + N
Sbjct: 202 LEKGYNSMPVNLPG---TLFHKSMKARKELSQILARILSERRQ----NGSSH-------N 247
Query: 113 DVVDALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLAL 172
D++ + + DK EL+ E I+ NII + +T + M+ LK+L + P L
Sbjct: 248 DLLGSFMGDKEELTD---------EQIADNIIGVIFAARDTTASVMSWILKYLAENPNVL 298
Query: 173 SKLMEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIK 232
+ EE M ++K K + TW D +P T VI ETLR+A+I++ +R+AV+DVE +
Sbjct: 299 EAVTEEQMAIRKDKEE-GESLTWGDTKKMPLTSRVIQETLRVASILSFTFREAVEDVEYE 357
Query: 233 GYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRWEI 269
GYLIPK W V+ ++H + + NP KFDP R+E+
Sbjct: 358 GYLIPKGWKVLPLFRNIHHSADIFSNPGKFDPSRFEV 394
>ref|NP_974574.1| cytochrome P450 family protein [Arabidopsis thaliana]
Length = 484
Score = 124 bits (311), Expect = 4e-27
Identities = 73/217 (33%), Positives = 120/217 (54%), Gaps = 24/217 (11%)
Query: 53 IASGYSSCPVTFMEKRKSILYKSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVN 112
+ GY+S PV ++ +KS+KA++ + +++ RI+ ER + N + N
Sbjct: 202 LEKGYNSMPVNLPG---TLFHKSMKARKELSQILARILSERRQ----NGSSH-------N 247
Query: 113 DVVDALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLAL 172
D++ + + DK EL+ E I+ NII + +T + M+ LK+L + P L
Sbjct: 248 DLLGSFMGDKEELTD---------EQIADNIIGVIFAARDTTASVMSWILKYLAENPNVL 298
Query: 173 SKLMEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIK 232
+ EE M ++K K + TW D +P T VI ETLR+A+I++ +R+AV+DVE +
Sbjct: 299 EAVTEEQMAIRKDKEE-GESLTWGDTKKMPLTSRVIQETLRVASILSFTFREAVEDVEYE 357
Query: 233 GYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRWEI 269
GYLIPK W V+ ++H + + NP KFDP R+E+
Sbjct: 358 GYLIPKGWKVLPLFRNIHHSADIFSNPGKFDPSRFEV 394
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.328 0.142 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 546,086,878
Number of Sequences: 2540612
Number of extensions: 21268013
Number of successful extensions: 74575
Number of sequences better than 10.0: 3898
Number of HSP's better than 10.0 without gapping: 1724
Number of HSP's successfully gapped in prelim test: 2174
Number of HSP's that attempted gapping in prelim test: 70820
Number of HSP's gapped (non-prelim): 4544
length of query: 355
length of database: 863,360,394
effective HSP length: 129
effective length of query: 226
effective length of database: 535,621,446
effective search space: 121050446796
effective search space used: 121050446796
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 75 (33.5 bits)
Medicago: description of AC136451.2


