

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136449.11 - phase: 0 /pseudo
(982 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
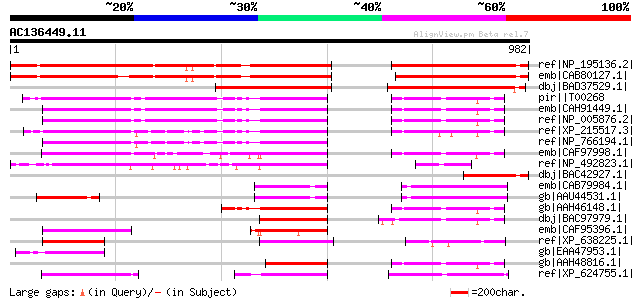
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_195136.2| zinc finger (C3HC4-type RING finger) family pro... 835 0.0
emb|CAB80127.1| putative protein [Arabidopsis thaliana] gi|29110... 795 0.0
dbj|BAD37529.1| zinc finger (C3HC4-type RING finger)protein-like... 402 e-110
pir||T00268 hypothetical protein KIAA0597 - human (fragment) gi|... 271 8e-71
emb|CAH91449.1| hypothetical protein [Pongo pygmaeus] 266 3e-69
ref|NP_005876.2| membrane-associated ring finger (C3HC4) 6 [Homo... 266 3e-69
ref|XP_215517.3| PREDICTED: similar to KIAA0597 protein [Rattus ... 260 1e-67
ref|NP_766194.1| membrane-associated ring finger (C3HC4) 6 [Mus ... 259 2e-67
emb|CAF97998.1| unnamed protein product [Tetraodon nigroviridis] 251 8e-65
ref|NP_492823.1| RING zinc finger containing protein (1L381) [Ca... 224 1e-56
dbj|BAC42927.1| unknown protein [Arabidopsis thaliana] 170 2e-40
emb|CAB79984.1| putative protein [Arabidopsis thaliana] gi|30637... 151 9e-35
gb|AAU44531.1| hypothetical protein AT4G32670 [Arabidopsis thali... 151 9e-35
gb|AAH46148.1| MARCH6 protein [Homo sapiens] 147 2e-33
dbj|BAC97979.1| mKIAA0597 protein [Mus musculus] 141 9e-32
emb|CAF95396.1| unnamed protein product [Tetraodon nigroviridis] 139 5e-31
ref|XP_638225.1| hypothetical protein DDB0186668 [Dictyostelium ... 132 4e-29
gb|EAA47953.1| hypothetical protein MG09083.4 [Magnaporthe grise... 131 1e-28
gb|AAH48816.1| March6 protein [Mus musculus] 129 5e-28
ref|XP_624755.1| PREDICTED: similar to Hypothetical protein CBG1... 129 5e-28
>ref|NP_195136.2| zinc finger (C3HC4-type RING finger) family protein [Arabidopsis
thaliana]
Length = 1092
Score = 835 bits (2156), Expect = 0.0
Identities = 444/665 (66%), Positives = 497/665 (73%), Gaps = 80/665 (12%)
Query: 1 MEIANEPPPSLDGTPIA--ATTPSSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSAKY 58
MEI+ S+ G + + PS SS SSSSS + S + + + +T ++Y
Sbjct: 1 MEISPADSLSISGAAASEVVSEPSVSSSSSSSSPNQASPNPFSNMDPAVSTATG---SRY 57
Query: 59 DDDDEDEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHP 118
DDDEDEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHP
Sbjct: 58 VDDDEDEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHP 117
Query: 119 FSFSPVYAENAPARLPFQEFVVGMAMKACHVLQFFVRLSFVLSVWLLIIPFITFWIWRLA 178
FSFSPVYA+NAP+RLPFQEFVVG+AMKACHVLQFF+RLSFVLSVWLL IPFITFWIWRLA
Sbjct: 118 FSFSPVYADNAPSRLPFQEFVVGIAMKACHVLQFFLRLSFVLSVWLLTIPFITFWIWRLA 177
Query: 179 FVRSFGEAQRLFFYHLFTAVILTDCLHGFLLSASIVFIFFGGTSLRDYFRHLREIGGQDA 238
FVR+FGEAQRLF H+ T VILTDCLHGFLLSASIVFIF G TSLRDYFRHLRE+GGQ+
Sbjct: 178 FVRTFGEAQRLFLSHISTTVILTDCLHGFLLSASIVFIFLGATSLRDYFRHLRELGGQE- 236
Query: 239 EREDEVDRNGARVARRPAGQANRNVNGDANGEDAVAAQGVAGAGQVIRRNAENVAARWEM 298
ER+D+VDRNGAR ARRPAGQANRN+ G+ NGEDA QG A GQ+ RRN ENV AR ++
Sbjct: 237 ERDDDVDRNGARAARRPAGQANRNLAGEGNGEDA-GDQG-AAVGQIARRNPENVLARLDI 294
Query: 299 QAARLEAHVEQMFDGLDDADGAEDVPFDELVGMQ-------------------------- 332
QAARLEA VEQMFDGLDDADGAEDVPFDELVGMQ
Sbjct: 295 QAARLEAQVEQMFDGLDDADGAEDVPFDELVGMQGPVFHLVENAFTVLASNMIFLGVVIF 354
Query: 333 ---------------------VPPTDASLSLA-------NITLKNALTAVQNLSTATQES 364
P ASL L NITLK+ALTAV NL++ Q +
Sbjct: 355 VPFTLGRIILYHVSWLFAAARGPAVAASLHLTDTGLSLENITLKSALTAVSNLTSEGQGN 414
Query: 365 GSIGQIAEMLKVNASELSEMSNNITASVSDDLLKGGSIGTSRISDVTTLAXGYIFLSTLI 424
G +GQ+ EM+KVN SEL+ +N T SV+ DLLKG ++G S++SD+TTLA GY+F+ L+
Sbjct: 415 GLLGQLTEMMKVNGSELNGANN--TLSVATDLLKGSTVGASKLSDITTLAVGYMFIVFLV 472
Query: 425 FCYFGVVALIRYTKGEPLTAGRFYGIASIAETIPSLFRQFLAAMRHLMTMVKVAFLLVIE 484
F Y G++ALIRY K E +PSL RQFLAAMRHLMTM+KVAFLLVIE
Sbjct: 473 FLYLGIIALIRYAK----------------EAVPSLLRQFLAAMRHLMTMIKVAFLLVIE 516
Query: 485 LGVFPLMCGWWLDVCTIQMFGKTMVHRAQFFSASPLASSLAHWVVGIVYMLQISIFVSLL 544
LGVFPLMCGWWLDVCT++MFGKTM HR QF S SPLASSL HWVVGI+YMLQISIFVSLL
Sbjct: 517 LGVFPLMCGWWLDVCTVRMFGKTMSHRVQFLSISPLASSLVHWVVGIMYMLQISIFVSLL 576
Query: 545 RGVLRNGVLYFLRDPADPNYNPSRDLIDNPVHKHARRVLLSVAVYGSLMVMLVILPR*LG 604
RGVLR GVLYFLRDPADPNYNP RDLID+PVHKHARRVLLSVAVYGSL+VMLV LP L
Sbjct: 577 RGVLRPGVLYFLRDPADPNYNPFRDLIDDPVHKHARRVLLSVAVYGSLIVMLVFLPVKLA 636
Query: 605 IADGP 609
I P
Sbjct: 637 IRMAP 641
Score = 413 bits (1062), Expect = e-113
Identities = 198/261 (75%), Positives = 221/261 (83%), Gaps = 6/261 (2%)
Query: 722 YVIWTAVAGVRYSIEQIRKRRTSVLLNQIWKWCSIVVKSSALLSIWIFVIPVLIGLLFEL 781
Y WT ++G RY+IE ++ +RTSVLLNQIWKWC IV KSS LL+IW+F+IPVLIGLLFEL
Sbjct: 835 YAFWTTISGARYAIEHVKSKRTSVLLNQIWKWCGIVFKSSVLLAIWVFIIPVLIGLLFEL 894
Query: 782 LVIVPMRVPVDESPVFLLYQDWALGLIFLKIWTRLVMLDHMMPLMDESWRVKFERVRDDG 841
LVIVPMRVPVDESPVFLLYQDWALGLIFLKIWTRLVMLDHM+P++D+SWR KFERVR+DG
Sbjct: 895 LVIVPMRVPVDESPVFLLYQDWALGLIFLKIWTRLVMLDHMLPIVDDSWRAKFERVREDG 954
Query: 842 FSRLQGLWVLREIVLPIIMKLLTALCVPYVLARGVFPALGYPLVVNSAVYRFAWLGCLSF 901
FSRLQGLWVLREIV PI+MKLLTALCVPYVLARGVFP LGYPLVVNSAVYRFAW+GCLS
Sbjct: 955 FSRLQGLWVLREIVFPIVMKLLTALCVPYVLARGVFPMLGYPLVVNSAVYRFAWIGCLSV 1014
Query: 902 SFVCFCAKRFHVWFTNLHNSIRDDRYLIGRRLHNFGE-HVEKANEAATSTGVQDAILLGP 960
S CFCAKR HVWF NLHNSIRDDRYLIGRRLHNFGE + N+ +S D +L+G
Sbjct: 1015 SLFCFCAKRCHVWFRNLHNSIRDDRYLIGRRLHNFGEAALANQNQNQSSEDAGDGVLIG- 1073
Query: 961 NINQQDRDADVGLRLRHINQQ 981
++ D D GLRLR QQ
Sbjct: 1074 ----REGDVDNGLRLRRAIQQ 1090
>emb|CAB80127.1| putative protein [Arabidopsis thaliana] gi|2911052|emb|CAA17562.1|
putative protein [Arabidopsis thaliana]
gi|7486281|pir||T05426 hypothetical protein F28A23.140 -
Arabidopsis thaliana
Length = 1051
Score = 795 bits (2053), Expect = 0.0
Identities = 428/665 (64%), Positives = 481/665 (71%), Gaps = 98/665 (14%)
Query: 1 MEIANEPPPSLDGTPIA--ATTPSSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSAKY 58
MEI+ S+ G + + PS SS SSSSS + S + + + +T ++Y
Sbjct: 1 MEISPADSLSISGAAASEVVSEPSVSSSSSSSSPNQASPNPFSNMDPAVSTATG---SRY 57
Query: 59 DDDDEDEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHP 118
DDDEDEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHP
Sbjct: 58 VDDDEDEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHP 117
Query: 119 FSFSPVYAENAPARLPFQEFVVGMAMKACHVLQFFVRLSFVLSVWLLIIPFITFWIWRLA 178
FSFSPVYA+NAP+RLPFQEFVVG+AMKACHVLQFF+RLSFVLSVWLL IPFITFWIWRLA
Sbjct: 118 FSFSPVYADNAPSRLPFQEFVVGIAMKACHVLQFFLRLSFVLSVWLLTIPFITFWIWRLA 177
Query: 179 FVRSFGEAQRLFFYHLFTAVILTDCLHGFLLSASIVFIFFGGTSLRDYFRHLREIGGQDA 238
FVR+FGEAQRLF H+ T VILTDCLHG DYFRHLRE+GGQ+
Sbjct: 178 FVRTFGEAQRLFLSHISTTVILTDCLHG------------------DYFRHLRELGGQE- 218
Query: 239 EREDEVDRNGARVARRPAGQANRNVNGDANGEDAVAAQGVAGAGQVIRRNAENVAARWEM 298
ER+D+VDRNGAR ARRPAGQANRN+ G+ NGEDA QG A GQ+ RRN ENV AR ++
Sbjct: 219 ERDDDVDRNGARAARRPAGQANRNLAGEGNGEDA-GDQG-AAVGQIARRNPENVLARLDI 276
Query: 299 QAARLEAHVEQMFDGLDDADGAEDVPFDELVGMQ-------------------------- 332
QAARLEA VEQMFDGLDDADGAEDVPFDELVGMQ
Sbjct: 277 QAARLEAQVEQMFDGLDDADGAEDVPFDELVGMQGPVFHLVENAFTVLASNMIFLGVVIF 336
Query: 333 ---------------------VPPTDASLSLA-------NITLKNALTAVQNLSTATQES 364
P ASL L NITLK+ALTAV NL++ Q +
Sbjct: 337 VPFTLGRIILYHVSWLFAAARGPAVAASLHLTDTGLSLENITLKSALTAVSNLTSEGQGN 396
Query: 365 GSIGQIAEMLKVNASELSEMSNNITASVSDDLLKGGSIGTSRISDVTTLAXGYIFLSTLI 424
G +GQ+ EM+KVN SEL+ +N T SV+ DLLKG ++G S++SD+TTLA GY+F+ L+
Sbjct: 397 GLLGQLTEMMKVNGSELNGANN--TLSVATDLLKGSTVGASKLSDITTLAVGYMFIVFLV 454
Query: 425 FCYFGVVALIRYTKGEPLTAGRFYGIASIAETIPSLFRQFLAAMRHLMTMVKVAFLLVIE 484
F Y G++ALIRY K E +PSL RQFLAAMRHLMTM+KVAFLLVIE
Sbjct: 455 FLYLGIIALIRYAK----------------EAVPSLLRQFLAAMRHLMTMIKVAFLLVIE 498
Query: 485 LGVFPLMCGWWLDVCTIQMFGKTMVHRAQFFSASPLASSLAHWVVGIVYMLQISIFVSLL 544
LGVFPLMCGWWLDVCT++MFGKTM HR QF S SPLASSL HWVVGI+YMLQISIFVSLL
Sbjct: 499 LGVFPLMCGWWLDVCTVRMFGKTMSHRVQFLSISPLASSLVHWVVGIMYMLQISIFVSLL 558
Query: 545 RGVLRNGVLYFLRDPADPNYNPSRDLIDNPVHKHARRVLLSVAVYGSLMVMLVILPR*LG 604
RGVLR GVLYFLRDPADPNYNP RDLID+PVHKHARRVLLSVAVYGSL+VMLV LP L
Sbjct: 559 RGVLRPGVLYFLRDPADPNYNPFRDLIDDPVHKHARRVLLSVAVYGSLIVMLVFLPVKLA 618
Query: 605 IADGP 609
I P
Sbjct: 619 IRMAP 623
Score = 403 bits (1035), Expect = e-110
Identities = 194/253 (76%), Positives = 217/253 (85%), Gaps = 6/253 (2%)
Query: 730 GVRYSIEQIRKRRTSVLLNQIWKWCSIVVKSSALLSIWIFVIPVLIGLLFELLVIVPMRV 789
G++Y+IE ++ +RTSVLLNQIWKWC IV KSS LL+IW+F+IPVLIGLLFELLVIVPMRV
Sbjct: 802 GIKYAIEHVKSKRTSVLLNQIWKWCGIVFKSSVLLAIWVFIIPVLIGLLFELLVIVPMRV 861
Query: 790 PVDESPVFLLYQDWALGLIFLKIWTRLVMLDHMMPLMDESWRVKFERVRDDGFSRLQGLW 849
PVDESPVFLLYQDWALGLIFLKIWTRLVMLDHM+P++D+SWR KFERVR+DGFSRLQGLW
Sbjct: 862 PVDESPVFLLYQDWALGLIFLKIWTRLVMLDHMLPIVDDSWRAKFERVREDGFSRLQGLW 921
Query: 850 VLREIVLPIIMKLLTALCVPYVLARGVFPALGYPLVVNSAVYRFAWLGCLSFSFVCFCAK 909
VLREIV PI+MKLLTALCVPYVLARGVFP LGYPLVVNSAVYRFAW+GCLS S CFCAK
Sbjct: 922 VLREIVFPIVMKLLTALCVPYVLARGVFPMLGYPLVVNSAVYRFAWIGCLSVSLFCFCAK 981
Query: 910 RFHVWFTNLHNSIRDDRYLIGRRLHNFGE-HVEKANEAATSTGVQDAILLGPNINQQDRD 968
R HVWF NLHNSIRDDRYLIGRRLHNFGE + N+ +S D +L+G ++ D
Sbjct: 982 RCHVWFRNLHNSIRDDRYLIGRRLHNFGEAALANQNQNQSSEDAGDGVLIG-----REGD 1036
Query: 969 ADVGLRLRHINQQ 981
D GLRLR QQ
Sbjct: 1037 VDNGLRLRRAIQQ 1049
>dbj|BAD37529.1| zinc finger (C3HC4-type RING finger)protein-like [Oryza sativa
(japonica cultivar-group)] gi|51536350|dbj|BAD37481.1|
zinc finger (C3HC4-type RING finger)protein-like [Oryza
sativa (japonica cultivar-group)]
Length = 681
Score = 402 bits (1032), Expect = e-110
Identities = 196/266 (73%), Positives = 223/266 (83%), Gaps = 11/266 (4%)
Query: 716 SSTLPCYVIWTAVAGVRYSIEQIRKRRTSVLLNQIWKWCSIVVKSSALLSIWIFVIPVLI 775
S ++ CY+IW+A AG RY+I+ IR RR + L+ QI KWCSIVVKSSALLSIWIFVIPVLI
Sbjct: 409 SFSIGCYIIWSAAAGTRYAIDYIRSRRLAFLVQQICKWCSIVVKSSALLSIWIFVIPVLI 468
Query: 776 GLLFELLVIVPMRVPVDESPVFLLYQDWALGLIFLKIWTRLVMLDHMMPLMDESWRVKFE 835
GLLFELLVIVPMRVP+DESPVFLLYQDWALGLIFLKIWTRLVMLD M PL+DESWR KFE
Sbjct: 469 GLLFELLVIVPMRVPIDESPVFLLYQDWALGLIFLKIWTRLVMLDQMAPLVDESWRTKFE 528
Query: 836 RVRDDGFSRLQGLWVLREIVLPIIMKLLTALCVPYVLARGVFPALGYPLVVNSAVYRFAW 895
RVR+DGFSRL+GLWVL EI++PI+ KLLTALCVPYVLARGVFP LGYPL+VNSAVYRFAW
Sbjct: 529 RVREDGFSRLRGLWVLHEIIMPIVTKLLTALCVPYVLARGVFPVLGYPLIVNSAVYRFAW 588
Query: 896 LGCLSFSFVCFCAKRFHVWFTNLHNSIRDDRYLIGRRLHNFGEHVEKANEAATSTGVQD- 954
LGCL FS + FC KRFHVWFTNLHNSIRDDRYLIGRRLHNFGE ++E T+T D
Sbjct: 589 LGCLIFSALFFCGKRFHVWFTNLHNSIRDDRYLIGRRLHNFGEDSLHSSEPGTTTASDDD 648
Query: 955 ----AILLGPNINQQDRDADVGLRLR 976
A++ +D++ ++GLR R
Sbjct: 649 EHEQALI------PRDQEGELGLRFR 668
Score = 321 bits (822), Expect = 9e-86
Identities = 158/220 (71%), Positives = 182/220 (81%)
Query: 390 ASVSDDLLKGGSIGTSRISDVTTLAXGYIFLSTLIFCYFGVVALIRYTKGEPLTAGRFYG 449
AS L+KG +IG+S +SD+TTLA GY+F+ L+F Y G +AL+RY +GE T GR YG
Sbjct: 3 ASGKSSLIKGTAIGSSYLSDLTTLAVGYMFIFCLVFLYIGSLALLRYARGERFTIGRLYG 62
Query: 450 IASIAETIPSLFRQFLAAMRHLMTMVKVAFLLVIELGVFPLMCGWWLDVCTIQMFGKTMV 509
IA+I E IPSL RQF A M+HLMTMVKVAFLLVIELGVFPLMCGWWLDVCT++M G T+
Sbjct: 63 IATILEAIPSLCRQFFAGMKHLMTMVKVAFLLVIELGVFPLMCGWWLDVCTLKMLGATIA 122
Query: 510 HRAQFFSASPLASSLAHWVVGIVYMLQISIFVSLLRGVLRNGVLYFLRDPADPNYNPSRD 569
R +FF+ SPLASS HW+VGI+YMLQISIFVSLLRGVLRNGVLYFLRDPADPNYNP RD
Sbjct: 123 QRVEFFTMSPLASSSIHWLVGIIYMLQISIFVSLLRGVLRNGVLYFLRDPADPNYNPFRD 182
Query: 570 LIDNPVHKHARRVLLSVAVYGSLMVMLVILPR*LGIADGP 609
LID+PVHKHARRVLLSVAVYGSL+V+LV LP L + P
Sbjct: 183 LIDDPVHKHARRVLLSVAVYGSLIVILVFLPVKLAMRVAP 222
>pir||T00268 hypothetical protein KIAA0597 - human (fragment)
gi|3043718|dbj|BAA25523.1| KIAA0597 protein [Homo
sapiens]
Length = 971
Score = 271 bits (693), Expect = 8e-71
Identities = 175/580 (30%), Positives = 277/580 (47%), Gaps = 63/580 (10%)
Query: 25 SPSSSSSSPRGSKGKEIDAEAVATASTAPPSAKYDDDDEDEEDVCRICRNPGDADNPLRY 84
SP S + PR +G + APP D D EED+CR+CR+ G + PL +
Sbjct: 39 SPPSFPARPREPRG---------CVTAAPP----DKMDTAEEDICRVCRSEGTPEKPLYH 85
Query: 85 PCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHPFSFSPVYAENAPARLPFQEFVVGMAM 144
PC C+GSIKF+HQ+CL+QWL HS CE+CKH F+F+P+Y+ + P+RLP Q+ G+
Sbjct: 86 PCVCTGSIKFIHQECLVQWLKHSRKEYCELCKHRFAFTPIYSPDMPSRLPIQDIFAGLVT 145
Query: 145 KACHVLQFFVRLSFVLSVWLLIIPFITFWIWRLAFVRSFGEAQRLFFYHLFTAVILTDCL 204
++++ + V WL ++P I++ F S L L T +L DCL
Sbjct: 146 SIGTAIRYWFHYTLVAFAWLGVVPLTACRIYKCLFTGSVSSLLTLPLDMLSTENLLADCL 205
Query: 205 HGFLLSASIVFIFFGGTSLRDYFRHLREIGGQDAEREDEVDRNGARVARRPAGQANRNVN 264
G + + F LR+ H GG E A AG
Sbjct: 206 QGCFVVTCTLCAFISLVWLREQIVH----GGAPIWLEHAAPPFNA------AGHHQNEAP 255
Query: 265 GDANGEDAVAAQGVAGAGQVIRRNAENVAARWEMQAARLEAHVEQMFDGLDDADGAEDVP 324
NG + VAA A EN A+ + E + E+ G++DA A +
Sbjct: 256 AGGNGAENVAADQPANPPAENAVVGENPDAQDDQAEEEEEDNEEEDDAGVEDAADANNGA 315
Query: 325 FDELVGMQVPPTDASLSLA---NITLKNALTAVQNLSTATQESGSIGQIAEMLKVNASEL 381
D++ + A+ L + L +L ++++ + + +
Sbjct: 316 QDDMNWNALEWDRAAEELTWERMLGLDGSLVFLEHVFWVVSLNTLFILVFAFCPYHIGHF 375
Query: 382 SEMSNNITASVSDDLLKGGSIGTSRISDVTTLAXGYIFLS-TLIFCYFGVVALIRYTKGE 440
S + V S + T GYI L+ TLI C+ G+ L+++ +
Sbjct: 376 SLVGLGFEEHVQ----------ASHFEGLITTIVGYILLAITLIICH-GLATLVKFHRSR 424
Query: 441 PLTAGRFYGIASIAETIPSLFRQFLAAMRHLMTMVKVAFLLVIELGVFPLMCGWWLDVCT 500
R G+ I +VKV+ L+V+E+GVFPL+CGWWLD+C+
Sbjct: 425 -----RLLGVCYI--------------------VVKVSLLVVVEIGVFPLICGWWLDICS 459
Query: 501 IQMFGKTMVHRAQFFSASPLASSLAHWVVGIVYMLQISIFVSLLRGVLRNGVLYFLRDPA 560
++MF T+ R F ++P + HW+VG+VY+ + F+ LLR VLR GVL+FLR+
Sbjct: 460 LEMFDATLKDRELSFQSAPGTTMFLHWLVGMVYVFYFASFILLLREVLRPGVLWFLRNLN 519
Query: 561 DPNYNPSRDLIDNPVHKHARRVLLSVAVYGSLMVMLVILP 600
DP++NP +++I P+++H RR +LSV V+GS++++++ LP
Sbjct: 520 DPDFNPVQEMIHLPIYRHLRRFILSVIVFGSIVLLMLWLP 559
Score = 127 bits (320), Expect = 1e-27
Identities = 70/221 (31%), Positives = 127/221 (56%), Gaps = 18/221 (8%)
Query: 722 YVIWTAVAGVRYSIEQIRKRRTSVLLNQIWKWCSIVVKSSALLSIWIFVIPVLIGLLFEL 781
YV W + V + + + R V+ ++ +W +++K+ + + V+P+L+GLLFEL
Sbjct: 744 YVCWLTIRAVTVMVAWMPQGRR-VIFQKVKEWSLMIMKTLIVAVLLAGVVPLLLGLLFEL 802
Query: 782 LVIVPMRVPVDESPVFLLYQDWALGLIFLKIWTRLVMLDHMMPLMDESWRVK--FERVRD 839
+++ P+RVP+D++P+F +QDWALG++ KI + LM W +K E+V
Sbjct: 803 VIVAPLRVPLDQTPLFYPWQDWALGVLHAKIIAAIT-------LMGPQWWLKTVIEQVYA 855
Query: 840 DGFSRLQGLWVLREIVLPIIMKLLTALCVPYVLARGVFPALGYPL----VVNSAVYRFAW 895
+G + +++R++ P+I LL +LCVPYV+A GV P LG +V+ +Y F
Sbjct: 856 NGIRNIDLHYIVRKLAAPVISVLLLSLCVPYVIASGVVPLLGVTAEMQNLVHRRIYPFLL 915
Query: 896 LGCLSFSFVCFCAKRFHVWFTNLHNSIRDDRYLIGRRLHNF 936
+ + + + F ++ F L+ I++D+YL+G+RL N+
Sbjct: 916 MVVVLMAILSFQVRQ----FKRLYEHIKNDKYLVGQRLVNY 952
>emb|CAH91449.1| hypothetical protein [Pongo pygmaeus]
Length = 910
Score = 266 bits (680), Expect = 3e-69
Identities = 165/543 (30%), Positives = 264/543 (48%), Gaps = 50/543 (9%)
Query: 62 DEDEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHPFSF 121
D EED+CR+CR+ G + PL +PC C+GSIKF+HQ+CL+QWL HS CE+CKH F+F
Sbjct: 2 DTAEEDICRVCRSEGTPEKPLYHPCVCTGSIKFIHQECLVQWLKHSRKEYCELCKHRFAF 61
Query: 122 SPVYAENAPARLPFQEFVVGMAMKACHVLQFFVRLSFVLSVWLLIIPFITFWIWRLAFVR 181
+P+Y+ + P+RLP Q+ G+ ++++ + V WL ++P I++ F
Sbjct: 62 TPIYSPDMPSRLPIQDIFAGLVTSIGTAIRYWFHYTLVAFAWLGVVPLTACRIYKCLFTG 121
Query: 182 SFGEAQRLFFYHLFTAVILTDCLHGFLLSASIVFIFFGGTSLRDYFRHLREIGGQDAERE 241
S L L T +L DCL G + + F LR+ H GG E
Sbjct: 122 SVSSLLTLPLDMLSTENLLADCLQGCFVVTCTLCAFISLVWLREQIVH----GGAPIWLE 177
Query: 242 DEVDRNGARVARRPAGQANRNVNGDANGEDAVAAQGVAGAGQVIRRNAENVAARWEMQAA 301
A AG NG + VAA A EN A+ +
Sbjct: 178 HAAPPFNA------AGHHQNEAPAGGNGAENVAADQPANPPAENAVVGENPDAQDDQAEE 231
Query: 302 RLEAHVEQMFDGLDDADGAEDVPFDELVGMQVPPTDASLSLA---NITLKNALTAVQNLS 358
E + E+ G++DA A + D++ + A+ L + L +L ++++
Sbjct: 232 EEEDNEEEDDAGVEDAADANNGAQDDMNWNALEWDRAAEELTWERMLGLDGSLVFLEHVF 291
Query: 359 TATQESGSIGQIAEMLKVNASELSEMSNNITASVSDDLLKGGSIGTSRISDVTTLAXGYI 418
+ + + S + V S + T GYI
Sbjct: 292 WVVSLNTLFILVFAFCPYHIGHFSLVGLGFEEHVQ----------ASHFEGLITTIVGYI 341
Query: 419 FLS-TLIFCYFGVVALIRYTKGEPLTAGRFYGIASIAETIPSLFRQFLAAMRHLMTMVKV 477
L+ TLI C+ G+ L+++ + R G+ I +VKV
Sbjct: 342 LLAITLIICH-GLATLVKFHRSR-----RLLGVCYI--------------------VVKV 375
Query: 478 AFLLVIELGVFPLMCGWWLDVCTIQMFGKTMVHRAQFFSASPLASSLAHWVVGIVYMLQI 537
+ L+V+E+GVFPL+CGWWLD+C+++MF T+ R F ++P + HW+VG+VY+
Sbjct: 376 SLLVVVEIGVFPLICGWWLDICSLEMFDATLKDRELSFQSAPGTTMFLHWLVGMVYVFYF 435
Query: 538 SIFVSLLRGVLRNGVLYFLRDPADPNYNPSRDLIDNPVHKHARRVLLSVAVYGSLMVMLV 597
+ F+ LLR VLR GVL+FLR+ DP++NP +++I P+++H RR +LSV V+GS++++++
Sbjct: 436 ASFILLLREVLRPGVLWFLRNLNDPDFNPVQEMIHLPIYRHLRRFILSVIVFGSIVLLML 495
Query: 598 ILP 600
LP
Sbjct: 496 WLP 498
Score = 127 bits (319), Expect = 2e-27
Identities = 70/221 (31%), Positives = 127/221 (56%), Gaps = 18/221 (8%)
Query: 722 YVIWTAVAGVRYSIEQIRKRRTSVLLNQIWKWCSIVVKSSALLSIWIFVIPVLIGLLFEL 781
YV W + V + + + R V+ ++ +W +++K+ + + V+P+L+GLLFEL
Sbjct: 683 YVCWLTIRAVTVMVAWMPQGRR-VVFQKVKEWSLMIMKTLIVAVLLAGVVPLLLGLLFEL 741
Query: 782 LVIVPMRVPVDESPVFLLYQDWALGLIFLKIWTRLVMLDHMMPLMDESWRVK--FERVRD 839
+++ P+RVP+D++P+F +QDWALG++ KI + LM W +K E+V
Sbjct: 742 VIVAPLRVPLDQTPLFYPWQDWALGVLHAKIIAAIT-------LMGPQWWLKTVIEQVYA 794
Query: 840 DGFSRLQGLWVLREIVLPIIMKLLTALCVPYVLARGVFPALGYPL----VVNSAVYRFAW 895
+G + +++R++ P+I LL +LCVPYV+A GV P LG +V+ +Y F
Sbjct: 795 NGIRNIDLHYIVRKLAAPVISVLLLSLCVPYVIASGVVPLLGVTAEMQNLVHRRIYPFLL 854
Query: 896 LGCLSFSFVCFCAKRFHVWFTNLHNSIRDDRYLIGRRLHNF 936
+ + + + F ++ F L+ I++D+YL+G+RL N+
Sbjct: 855 MVVVLMAILSFQVRQ----FKRLYEHIKNDKYLVGQRLVNY 891
>ref|NP_005876.2| membrane-associated ring finger (C3HC4) 6 [Homo sapiens]
Length = 910
Score = 266 bits (680), Expect = 3e-69
Identities = 165/543 (30%), Positives = 264/543 (48%), Gaps = 50/543 (9%)
Query: 62 DEDEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHPFSF 121
D EED+CR+CR+ G + PL +PC C+GSIKF+HQ+CL+QWL HS CE+CKH F+F
Sbjct: 2 DTAEEDICRVCRSEGTPEKPLYHPCVCTGSIKFIHQECLVQWLKHSRKEYCELCKHRFAF 61
Query: 122 SPVYAENAPARLPFQEFVVGMAMKACHVLQFFVRLSFVLSVWLLIIPFITFWIWRLAFVR 181
+P+Y+ + P+RLP Q+ G+ ++++ + V WL ++P I++ F
Sbjct: 62 TPIYSPDMPSRLPIQDIFAGLVTSIGTAIRYWFHYTLVAFAWLGVVPLTACRIYKCLFTG 121
Query: 182 SFGEAQRLFFYHLFTAVILTDCLHGFLLSASIVFIFFGGTSLRDYFRHLREIGGQDAERE 241
S L L T +L DCL G + + F LR+ H GG E
Sbjct: 122 SVSSLLTLPLDMLSTENLLADCLQGCFVVTCTLCAFISLVWLREQIVH----GGAPIWLE 177
Query: 242 DEVDRNGARVARRPAGQANRNVNGDANGEDAVAAQGVAGAGQVIRRNAENVAARWEMQAA 301
A AG NG + VAA A EN A+ +
Sbjct: 178 HAAPPFNA------AGHHQNEAPAGGNGAENVAADQPANPPAENAVVGENPDAQDDQAEE 231
Query: 302 RLEAHVEQMFDGLDDADGAEDVPFDELVGMQVPPTDASLSLA---NITLKNALTAVQNLS 358
E + E+ G++DA A + D++ + A+ L + L +L ++++
Sbjct: 232 EEEDNEEEDDAGVEDAADANNGAQDDMNWNALEWDRAAEELTWERMLGLDGSLVFLEHVF 291
Query: 359 TATQESGSIGQIAEMLKVNASELSEMSNNITASVSDDLLKGGSIGTSRISDVTTLAXGYI 418
+ + + S + V S + T GYI
Sbjct: 292 WVVSLNTLFILVFAFCPYHIGHFSLVGLGFEEHVQ----------ASHFEGLITTIVGYI 341
Query: 419 FLS-TLIFCYFGVVALIRYTKGEPLTAGRFYGIASIAETIPSLFRQFLAAMRHLMTMVKV 477
L+ TLI C+ G+ L+++ + R G+ I +VKV
Sbjct: 342 LLAITLIICH-GLATLVKFHRSR-----RLLGVCYI--------------------VVKV 375
Query: 478 AFLLVIELGVFPLMCGWWLDVCTIQMFGKTMVHRAQFFSASPLASSLAHWVVGIVYMLQI 537
+ L+V+E+GVFPL+CGWWLD+C+++MF T+ R F ++P + HW+VG+VY+
Sbjct: 376 SLLVVVEIGVFPLICGWWLDICSLEMFDATLKDRELSFQSAPGTTMFLHWLVGMVYVFYF 435
Query: 538 SIFVSLLRGVLRNGVLYFLRDPADPNYNPSRDLIDNPVHKHARRVLLSVAVYGSLMVMLV 597
+ F+ LLR VLR GVL+FLR+ DP++NP +++I P+++H RR +LSV V+GS++++++
Sbjct: 436 ASFILLLREVLRPGVLWFLRNLNDPDFNPVQEMIHLPIYRHLRRFILSVIVFGSIVLLML 495
Query: 598 ILP 600
LP
Sbjct: 496 WLP 498
Score = 127 bits (320), Expect = 1e-27
Identities = 70/221 (31%), Positives = 127/221 (56%), Gaps = 18/221 (8%)
Query: 722 YVIWTAVAGVRYSIEQIRKRRTSVLLNQIWKWCSIVVKSSALLSIWIFVIPVLIGLLFEL 781
YV W + V + + + R V+ ++ +W +++K+ + + V+P+L+GLLFEL
Sbjct: 683 YVCWLTIRAVTVMVAWMPQGRR-VIFQKVKEWSLMIMKTLIVAVLLAGVVPLLLGLLFEL 741
Query: 782 LVIVPMRVPVDESPVFLLYQDWALGLIFLKIWTRLVMLDHMMPLMDESWRVK--FERVRD 839
+++ P+RVP+D++P+F +QDWALG++ KI + LM W +K E+V
Sbjct: 742 VIVAPLRVPLDQTPLFYPWQDWALGVLHAKIIAAIT-------LMGPQWWLKTVIEQVYA 794
Query: 840 DGFSRLQGLWVLREIVLPIIMKLLTALCVPYVLARGVFPALGYPL----VVNSAVYRFAW 895
+G + +++R++ P+I LL +LCVPYV+A GV P LG +V+ +Y F
Sbjct: 795 NGIRNIDLHYIVRKLAAPVISVLLLSLCVPYVIASGVVPLLGVTAEMQNLVHRRIYPFLL 854
Query: 896 LGCLSFSFVCFCAKRFHVWFTNLHNSIRDDRYLIGRRLHNF 936
+ + + + F ++ F L+ I++D+YL+G+RL N+
Sbjct: 855 MVVVLMAILSFQVRQ----FKRLYEHIKNDKYLVGQRLVNY 891
>ref|XP_215517.3| PREDICTED: similar to KIAA0597 protein [Rattus norvegicus]
Length = 1583
Score = 260 bits (665), Expect = 1e-67
Identities = 177/593 (29%), Positives = 283/593 (46%), Gaps = 83/593 (13%)
Query: 26 PSSSSSSPRGSKGKEIDAEAVATASTAPPSAKYDDDDEDEEDVCRICRNPGDADNPLRYP 85
PSS P G + + A A PP+ D EED+CR+CR+ G + PL +P
Sbjct: 116 PSSLPPFPSGPRLCRV-AGASRLRHRRPPNKM----DTAEEDICRVCRSEGTPEKPLYHP 170
Query: 86 CACSGSIKFVHQDCLLQWLNHSNARQCEVCKHPFSFSPVYAENAPARLPFQEFVVGMAMK 145
C C+GSIKF+HQ+CL+QWL HS CE+CKH F+F+P+Y+ + P+RLP Q+ G+
Sbjct: 171 CVCTGSIKFIHQECLVQWLKHSRKEYCELCKHRFAFTPIYSPDMPSRLPIQDIFAGLVTS 230
Query: 146 ACHVLQFFVRLSFVLSVWLLIIPFITFWIWRLAFVRSFGEAQRLFFYHLFTAVILTDCLH 205
++++ + V WL ++P I++ F S L L T +L DCL
Sbjct: 231 IGTAIRYWFHYTLVAFAWLGVVPLTACRIYKCLFTGSVSSLLTLPLDMLSTENLLADCLQ 290
Query: 206 GFLLSASIVFIFFGGTSLRDYFRHLREIGGQDA---------------EREDEVDRNGAR 250
G + + F LR+ H GG + E V NGA
Sbjct: 291 GCFVVTCTLCAFISLVWLREQIVH----GGAPIWLEHAAPPFNAAGHHQNEAPVGGNGAE 346
Query: 251 --VARRPAGQANRNVNGDANGEDAVAAQGVAGAGQVIRRNAENVAARWEMQAARLEAHVE 308
A +PA A N GE+ A G A + ++ + A A +
Sbjct: 347 NPAADQPANPAGENA---VLGENPDAQDGQAEEEEEDNEEEDDAGVE-DAADANNGAQDD 402
Query: 309 QMFDGLDDADGAEDVPFDELVGMQVPPTDASLSLANITLKNALTAVQNLSTATQESGSIG 368
++ L+ AE++ ++ ++G+ D SL + L++ V +L+T
Sbjct: 403 MNWNALEWDRAAEELTWERMLGL-----DGSL----VFLEHVFWVV-SLNTL------FI 446
Query: 369 QIAEMLKVNASELSEMSNNITASVSDDLLKGGSIGTSRISDVTTLAXGYIFLS-TLIFCY 427
+ + S + V S + T GYI L+ TLI C+
Sbjct: 447 LVFAFCPYHIGHFSLVGLGFEEHVQ----------ASHFEGLITTIVGYILLAITLIICH 496
Query: 428 FGVVALIRYTKGEPLTAGRFYGIASIAETIPSLFRQFLAAMRHLMTMVKVAFLLVIELGV 487
+ L+++ + R G+ I +VKV+ L+V+E+GV
Sbjct: 497 -ALATLVKFHRSR-----RLLGVCYI--------------------VVKVSLLVVVEIGV 530
Query: 488 FPLMCGWWLDVCTIQMFGKTMVHRAQFFSASPLASSLAHWVVGIVYMLQISIFVSLLRGV 547
FPL+CGWWLD+C+++MF T+ R F ++P + HW+VG+VY+ + F+ LLR V
Sbjct: 531 FPLICGWWLDICSLEMFDATLKDRELSFQSAPGTTMFLHWLVGMVYVFYFASFILLLREV 590
Query: 548 LRNGVLYFLRDPADPNYNPSRDLIDNPVHKHARRVLLSVAVYGSLMVMLVILP 600
LR GVL+FLR+ DP++NP +++I P+++H RR +LSV V+GS++++++ LP
Sbjct: 591 LRPGVLWFLRNLNDPDFNPVQEMIHLPIYRHLRRFILSVIVFGSIVLLMLWLP 643
Score = 109 bits (273), Expect = 4e-22
Identities = 68/258 (26%), Positives = 125/258 (48%), Gaps = 48/258 (18%)
Query: 722 YVIWTAVAGVRYSIEQIRKRRTSVLLNQIWKWCSIVVKSSALLSIWIFVIPVLIGLLFEL 781
YV W + V + + + R V+ ++ +W +++K+ + + V+P+L+GLLFEL
Sbjct: 827 YVCWLTIRAVTVLVAWMPQGRR-VIFQKVKEWSLMIMKTLIVAVLLAGVVPLLLGLLFEL 885
Query: 782 LVIVPMRVPVDESPVFLLYQDWALGLIFLKI----------WTRLVMLDHMMPLMDESWR 831
+++ P+RVP+D++P+F +QDWALG++ KI W +++ R
Sbjct: 886 VIVAPLRVPLDQTPLFYPWQDWALGVLHAKIIAAITLMGPQWWLKTVIEQGQEGSSAFLR 945
Query: 832 VKF-----------------------------ERVRDDGFSRLQGLWVLREIVLPIIMKL 862
V F V +G + +++R++ P+I L
Sbjct: 946 VAFSPPKTSNARGVKCSVFVLVSRFCQSLLTDSEVYANGIRNIDLHYIIRKLAAPVISVL 1005
Query: 863 LTALCVPYVLARGVFPALGYPL----VVNSAVYRFAWLGCLSFSFVCFCAKRFHVWFTNL 918
L +LC+PYV+A GV P LG +V+ +Y F + + + F ++ F L
Sbjct: 1006 LLSLCIPYVIASGVVPLLGVTAEMQNLVHRRIYPFLLMVVVLMGILSFQVRQ----FKRL 1061
Query: 919 HNSIRDDRYLIGRRLHNF 936
+ I++D+YL+G+RL N+
Sbjct: 1062 YEHIKNDKYLVGQRLVNY 1079
>ref|NP_766194.1| membrane-associated ring finger (C3HC4) 6 [Mus musculus]
gi|26354689|dbj|BAC40971.1| unnamed protein product [Mus
musculus]
Length = 661
Score = 259 bits (663), Expect = 2e-67
Identities = 168/557 (30%), Positives = 271/557 (48%), Gaps = 78/557 (14%)
Query: 62 DEDEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHPFSF 121
D EED+CR+CR+ G + PL +PC C+GSIKF+HQ+CL+QWL HS CE+CKH F+F
Sbjct: 2 DTAEEDICRVCRSEGTPEKPLYHPCVCTGSIKFIHQECLVQWLKHSRKEYCELCKHRFAF 61
Query: 122 SPVYAENAPARLPFQEFVVGMAMKACHVLQFFVRLSFVLSVWLLIIPFITFWIWRLAFVR 181
+P+Y+ + P+RLP Q+ G+ ++++ + V WL ++P I++ F
Sbjct: 62 TPIYSPDMPSRLPIQDIFAGLVTSIGTAIRYWFHYTLVAFAWLGVVPLTACRIYKCLFTG 121
Query: 182 SFGEAQRLFFYHLFTAVILTDCLHGFLLSASIVFIFFGGTSLRDYFRHLREIGGQDA--- 238
S L L T +L DCL G + + F LR+ H GG
Sbjct: 122 SVSSLLTLPLDMLSTENLLADCLQGCFVVTCTLCAFISLVWLREQIVH----GGAPIWLE 177
Query: 239 ------------EREDEVDRNGAR--VARRPAGQANRNVNGDANGEDAVAAQGVAGAGQV 284
+ E V NGA A +PA A N GE+ A G A +
Sbjct: 178 HAAPPFNAAGHHQNEAPVGGNGAENPAADQPANPAGENA---VLGENPDAQDGQAEEEEE 234
Query: 285 IRRNAENVAARWEMQAARLEAHVEQMFDGLDDADGAEDVPFDELVGMQVPPTDASLSLAN 344
++ + A A + ++ L+ AE++ ++ ++G+ D SL
Sbjct: 235 DNEEEDDAGVE-DAADANNGAQDDMNWNALEWDRAAEELTWERMLGL-----DGSL---- 284
Query: 345 ITLKNALTAVQNLSTATQESGSIGQIAEMLKVNASELSEMSNNITASVSDDLLKGGSIGT 404
+ L++ V +L+T + + S + V
Sbjct: 285 VFLEHVFWVV-SLNTL------FILVFAFCPYHIGHFSLVGLGFEEHVQ----------A 327
Query: 405 SRISDVTTLAXGYIFLS-TLIFCYFGVVALIRYTKGEPLTAGRFYGIASIAETIPSLFRQ 463
S + T GYI L+ TLI C+ + L+++ + R G+ I
Sbjct: 328 SHFEGLITTIVGYILLAITLIICH-ALATLVKFHRSR-----RLLGVCYI---------- 371
Query: 464 FLAAMRHLMTMVKVAFLLVIELGVFPLMCGWWLDVCTIQMFGKTMVHRAQFFSASPLASS 523
+VKV+ L+V+E+GVFPL+CGWWLD+C+++MF T+ R F ++P +
Sbjct: 372 ----------VVKVSLLVVVEIGVFPLICGWWLDICSLEMFDATLKDRELSFQSAPGTTM 421
Query: 524 LAHWVVGIVYMLQISIFVSLLRGVLRNGVLYFLRDPADPNYNPSRDLIDNPVHKHARRVL 583
HW+VG+VY+ + F+ LLR VLR GVL+FLR+ DP++NP +++I P+++H RR +
Sbjct: 422 FLHWLVGMVYVFYFASFILLLREVLRPGVLWFLRNLNDPDFNPVQEMIHLPIYRHLRRFI 481
Query: 584 LSVAVYGSLMVMLVILP 600
LSV V+GS++++++ LP
Sbjct: 482 LSVIVFGSIVLLMLWLP 498
>emb|CAF97998.1| unnamed protein product [Tetraodon nigroviridis]
Length = 972
Score = 251 bits (641), Expect = 8e-65
Identities = 177/578 (30%), Positives = 280/578 (47%), Gaps = 60/578 (10%)
Query: 61 DDEDEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHPFS 120
D +E D+CR+CR+ G D PL +PC C+GSIKF+HQ+CL+QWL HS CE+CKH F+
Sbjct: 4 DTAEEADICRVCRSEGTPDKPLYHPCVCTGSIKFIHQECLVQWLKHSRKEYCELCKHRFA 63
Query: 121 FSPVYAENAPARLPFQEFVVGMAMKACHVLQFFVRLSFVLSVWLLIIPFITFWIWRLAFV 180
F+P+Y+ + P+RLP Q+ G+ ++++ + V WL ++P I++ F
Sbjct: 64 FTPIYSPDMPSRLPIQDICAGLLTSVGTAIRYWFHYTLVAFAWLGVVPLTACRIYKCLFT 123
Query: 181 RSFGEAQRLFFYHLFTAVILTDCLHGFLLSASIVFIFFGGTSLRDYFRHLREIGG--QDA 238
S L L T +L DCL G + + F LR+ H GG Q
Sbjct: 124 GSVSSLLTLPLDMLSTENLLADCLQGCFVVTCTLCAFISLVWLREQIVH----GGAPQWL 179
Query: 239 EREDEVDRNGARVARRPAGQANRNV-NGDANGEDA-----------VAAQGVAGAGQVIR 286
E+ +N A A + R + NG + + V AQ AG G
Sbjct: 180 EQHQPPPQNAAGQANEVSSLVQRWLGNGFVRLQSSPVLSFFFVFPCVQAQ-AAGQGVAQE 238
Query: 287 RNAENVAARWEMQAARLEAHVEQMFDGLDDADGAEDVPFDELVGMQVPPTDASLSLANIT 346
R A A A+ EA E DG + A+D +E DA+ +
Sbjct: 239 RPAAQPAP--PDPPAQNEAEPEPP-DG--PPEQADDPDLEEEEEAAAEDADANNGAQDDM 293
Query: 347 LKNALTAVQNLSTATQESGSIGQIAEMLKVNASELSEMSNNITASVSDDL-------LKG 399
NAL + T E ++ ++ +SN L L G
Sbjct: 294 NWNALEWDRAAEELTWERVTL----------SARRCSISNTPNVGALKGLPSFQMLGLDG 343
Query: 400 GSIGTSRISDVTTLAXGYIFLSTLIFC--YFGVVALIRYTKGEPLTAGRFYGIAS----- 452
+ + V +L +F+ FC + G +++ E + A F G+ +
Sbjct: 344 SLVFLEHVFWVVSL--NTLFILVFAFCPYHIGHFSVVGLGFEEYVQASHFEGLITTIVGY 401
Query: 453 IAETIPSLFRQFLAAM------RHLM----TMVKVAFLLVIELGVFPLMCGWWLDVCTIQ 502
I + + LAA+ R L+ +VKV+ L+V+E+GVFPL+CGWWLD+C+++
Sbjct: 402 ILLAMTLILCHGLAALVRFQRSRRLLGVCYIVVKVSLLVVVEIGVFPLICGWWLDICSLE 461
Query: 503 MFGKTMVHRAQFFSASPLASSLAHWVVGIVYMLQISIFVSLLRGVLRNGVLYFLRDPADP 562
MF ++ R F ++P + HW+VG+VY+ + F+ LLR VLR GVL+FLR+ DP
Sbjct: 462 MFDASLKDRELSFKSAPGTTMFLHWLVGMVYVFYFASFILLLREVLRPGVLWFLRNLNDP 521
Query: 563 NYNPSRDLIDNPVHKHARRVLLSVAVYGSLMVMLVILP 600
++NP +++I P+++H RR +LSV V+GS++++++ LP
Sbjct: 522 DFNPVQEMIHLPIYRHLRRFILSVVVFGSIVLLMLWLP 559
Score = 135 bits (339), Expect = 9e-30
Identities = 74/221 (33%), Positives = 131/221 (58%), Gaps = 18/221 (8%)
Query: 722 YVIWTAVAGVRYSIEQIRKRRTSVLLNQIWKWCSIVVKSSALLSIWIFVIPVLIGLLFEL 781
YV W ++ GV + + + RT V+++++ +W +++K+ + + VIP+L+GLLFEL
Sbjct: 752 YVCWLSIRGVTVLLAWMPQGRT-VIVHKVQEWTLMILKTLVVALLVAGVIPLLLGLLFEL 810
Query: 782 LVIVPMRVPVDESPVFLLYQDWALGLIFLKIWTRLVMLDHMMPLMDESWRVK--FERVRD 839
+++ P+RVP+D++P+F +QDWALG++ KI + LM W +K E+V
Sbjct: 811 VIVAPLRVPLDQTPLFYPWQDWALGVLHAKIIAAIT-------LMGPQWWLKTVIEQVYA 863
Query: 840 DGFSRLQGLWVLREIVLPIIMKLLTALCVPYVLARGVFPALG----YPLVVNSAVYRFAW 895
+G + +++R++ P+I LL ALCVPYV+A GV PA G +++ +Y F
Sbjct: 864 NGIRNIDLQFIIRKLAAPVISVLLLALCVPYVIAAGVVPAAGVTPEMEILMQRRIYPFLL 923
Query: 896 LGCLSFSFVCFCAKRFHVWFTNLHNSIRDDRYLIGRRLHNF 936
+ L + F ++ F L+ I++D+YL+G+RL N+
Sbjct: 924 MVVLLIGILSFQIRQ----FKRLYEHIKNDKYLVGQRLVNY 960
>ref|NP_492823.1| RING zinc finger containing protein (1L381) [Caenorhabditis
elegans]
Length = 1017
Score = 224 bits (571), Expect = 1e-56
Identities = 177/653 (27%), Positives = 286/653 (43%), Gaps = 81/653 (12%)
Query: 1 MEIANEPPPSLDGTPIAATTPSSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSAKYDD 60
ME+ +E PP + A + +S+ PS+SS +P + PPS D
Sbjct: 1 MEVEHEQPPEHE----AGSVDASNRPSTSSENPNPEE-----------PCPQPPSDPIID 45
Query: 61 DDEDEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHPFS 120
D++D +CR+CR + L YPC C+GSIK+VHQ+CL++WL +S CE+C H +S
Sbjct: 46 DNDDHL-MCRVCRGN---EGSLYYPCLCTGSIKYVHQECLVEWLKYSKKEVCELCNHKYS 101
Query: 121 FSPVYAENAPARLPFQEFVVGMAMKACHVLQFFVRLSFVLSVWLLIIPFITFWIWRLAFV 180
F P+Y ++ P LP E + G+ + +++ ++ + V++ WL I+P I+ F
Sbjct: 102 FQPIYRQDMPKALPILEILRGVLISGAIMVRTWIIYTIVMTTWLGIVPLTAARIYNCIFY 161
Query: 181 RSFGEAQRLFFYHLFTAVILTDCLHGFLLSASIVFIFFGGTSLRDYF-----RHLREIGG 235
SF + F D L G LL V F LR+ +H I
Sbjct: 162 LSFHDIVNAPFQLFKAEHFFNDILKGSLLLLVFVCTFISLVWLREQIIIGGPQHFLNIEA 221
Query: 236 QDAEREDE-VDRNGARVARRPAGQANRNVNGDAN--------------GEDAVAAQGVAG 280
++ E ED+ V+ N + + + D + ED V G
Sbjct: 222 EENEGEDDVVNENNESSDESEDDEEDEDDEEDDDEDDEEEEEDSESGYEEDPVPEFREEG 281
Query: 281 AGQVIRRNAENVAARWEMQAARLEAHVEQ----MFDGLDDADG--------AEDVPFDEL 328
V RR R ++ AR + ++D + D + ++V D +
Sbjct: 282 EAYVARRRVPFEERRRDILEARHGFRPQMPNLGIYDDMRDDENEVLENDHVLQNVQEDRI 341
Query: 329 VGMQVP----PTDASLSLANITLKNALTAVQNLSTATQESGSIGQIAEMLKVNASELSEM 384
P P A+ + I + + A QE G G + N E
Sbjct: 342 PPPMEPEPIVPRAAARANRRIRRQRENEEAAAAAAAAQEQGGAGDQED----NWREWDRF 397
Query: 385 SNNITASVSDDLLKGGSIGTSRISDVTTLAXGYIFLSTLIFCYF------------GVVA 432
+ +T L G I + V +L L T F YF G+
Sbjct: 398 GDELTWQRLLG-LDGSLIFLEHVFWVISLNT----LFTATFAYFPFRIGNWFLSVIGLHG 452
Query: 433 LIRYTKGEPLTAGRFYGIASIAETIPSLFRQFLAAMRH-----LMTMVKVAFLLVIELGV 487
I Y + IA I I + QF M + + ++KV L+ +E+G
Sbjct: 453 KISYFPSIVAMLLGYIQIAIITYVIHQIMLQFKMKMMYRFLGIMFLIIKVFLLVFLEIGF 512
Query: 488 FPLMCGWWLDVCTIQMFGKTMVHRAQFFSASPLASSLAHWVVGIVYMLQISIFVSLLRGV 547
FP+MCG W+DVCT+ +F T+ R F+ +P S HW+VG+VY+ + FV LLR +
Sbjct: 513 FPVMCGCWMDVCTLPLFNITLSQRIATFATAPFMSIFLHWMVGMVYVFYSASFVILLREI 572
Query: 548 LRNGVLYFLRDPADPNYNPSRDLIDNPVHKHARRVLLSVAVYGSLMVMLVILP 600
LR GVL+F+R+ DP++NP +++ID P +H RR+++S ++ S ++++ +P
Sbjct: 573 LRPGVLWFMRNLNDPDFNPIQEMIDLPFTRHFRRLVVSTTLFFSTILLIFYIP 625
Score = 57.0 bits (136), Expect = 3e-06
Identities = 33/109 (30%), Positives = 57/109 (52%), Gaps = 9/109 (8%)
Query: 768 IFVIPVLIGLLFELLVIVPMRVPVDESPVFLLYQDWALGLIFLKIWTRLVMLDHMMPLMD 827
I V+P+L+G F+L+ PMR+ ++ + YQDWA+G++ +KI+ ++ +M
Sbjct: 851 IIVLPLLMGTYFQLVFFTPMRLGYQQTALMFPYQDWAMGVVQIKIF-------GVIAVMG 903
Query: 828 ESWRVKFE--RVRDDGFSRLQGLWVLREIVLPIIMKLLTALCVPYVLAR 874
W +K E + G L V IV+P I+ L T + P V+ +
Sbjct: 904 PDWWLKTELDLLVQRGIENFAALHVFVRIVVPCILYLSTFIAFPVVVIK 952
>dbj|BAC42927.1| unknown protein [Arabidopsis thaliana]
Length = 120
Score = 170 bits (431), Expect = 2e-40
Identities = 86/123 (69%), Positives = 94/123 (75%), Gaps = 6/123 (4%)
Query: 860 MKLLTALCVPYVLARGVFPALGYPLVVNSAVYRFAWLGCLSFSFVCFCAKRFHVWFTNLH 919
MKLLTALCVPYVLARGVFP LGYPLVVNSAVYRFAW+GCLS S CFCAKR HVWF NLH
Sbjct: 1 MKLLTALCVPYVLARGVFPMLGYPLVVNSAVYRFAWIGCLSVSLFCFCAKRCHVWFRNLH 60
Query: 920 NSIRDDRYLIGRRLHNFGE-HVEKANEAATSTGVQDAILLGPNINQQDRDADVGLRLRHI 978
NSIRDDRYLIGRRLHNFGE + N+ +S D +L+G ++ D D GLRLR
Sbjct: 61 NSIRDDRYLIGRRLHNFGEAALANQNQNQSSEDAGDGVLIG-----REGDVDNGLRLRRA 115
Query: 979 NQQ 981
QQ
Sbjct: 116 IQQ 118
>emb|CAB79984.1| putative protein [Arabidopsis thaliana] gi|3063703|emb|CAA18594.1|
putative protein [Arabidopsis thaliana]
gi|15233875|ref|NP_194993.1| expressed protein
[Arabidopsis thaliana] gi|7486406|pir||T04459
hypothetical protein F4D11.130 - Arabidopsis thaliana
Length = 808
Score = 151 bits (382), Expect = 9e-35
Identities = 78/200 (39%), Positives = 116/200 (58%), Gaps = 4/200 (2%)
Query: 742 RTSVLLNQIWKWCSIVVKSSALLSIWIFVIPVLIGLLFELLVIVPMRVPVDESPVFLLYQ 801
RT +LLN + + +++ L SIWI VIP ++GLL +L++I+P +VP+ ESPV+ L
Sbjct: 613 RTDLLLNHVLMF----IRNVLLFSIWISVIPGVLGLLIDLMIIIPSQVPLGESPVYNLLH 668
Query: 802 DWALGLIFLKIWTRLVMLDHMMPLMDESWRVKFERVRDDGFSRLQGLWVLREIVLPIIMK 861
DW +G++ L IW L ML + +WR K +R+R +RL W++R+++ II+
Sbjct: 669 DWLIGVVVLHIWIFLTMLTRINCFATVAWREKLQRIRSVTINRLPFTWLIRDVIGSIIVS 728
Query: 862 LLTALCVPYVLARGVFPALGYPLVVNSAVYRFAWLGCLSFSFVCFCAKRFHVWFTNLHNS 921
LL LCVPYV+ +FP LG+ VN V RF W L+ + F K LH
Sbjct: 729 LLFTLCVPYVVVNSLFPILGFSSAVNLTVQRFIWPAILALIPIWFSVKLIRDLILYLHKL 788
Query: 922 IRDDRYLIGRRLHNFGEHVE 941
D+RY +G RL +F E +E
Sbjct: 789 EFDNRYKVGERLVDFTEDLE 808
Score = 83.6 bits (205), Expect = 3e-14
Identities = 42/137 (30%), Positives = 78/137 (56%), Gaps = 9/137 (6%)
Query: 464 FLAAMRHLMTMVKVAFLLVIELGVFPLMCGWWLDVCTIQMFGKTMVHRAQFFSASPLASS 523
+L ++R + VK F+L +LGV P + G WL CT + GKT H + S PL +
Sbjct: 259 YLTSVRTFLPSVKDTFILSFKLGVLPWLLGCWLHFCTFPILGKTASHTVEVLSDYPLMAD 318
Query: 524 LAHWVVGIVYMLQISIFVSLLRGVLRNGVLYFLRDPADPNYNPSRDLIDNPVHKHARRVL 583
HW++G +Y++ + L++ +++ L++L D A+PNY ++ H +L
Sbjct: 319 -KHWLMGTLYLVSALSCMELIQKIVQKRALWYLLDVAEPNYKVTK--------LHLGPIL 369
Query: 584 LSVAVYGSLMVMLVILP 600
L+ A++G+++V+++ LP
Sbjct: 370 LAFALHGTMVVIVLHLP 386
Score = 46.2 bits (108), Expect = 0.005
Identities = 21/51 (41%), Positives = 34/51 (66%), Gaps = 7/51 (13%)
Query: 119 FSFSPVYAENAPARLPFQEFVVGMAMKACHVLQFFVRLSFVLSVWLLIIPF 169
+S PVY+ENAP RLP+ EF++G+ M+A +R ++ W+L++PF
Sbjct: 5 YSIVPVYSENAPERLPWHEFLMGLLMRA-------LRFMNLILPWILMMPF 48
>gb|AAU44531.1| hypothetical protein AT4G32670 [Arabidopsis thaliana]
Length = 860
Score = 151 bits (382), Expect = 9e-35
Identities = 78/200 (39%), Positives = 116/200 (58%), Gaps = 4/200 (2%)
Query: 742 RTSVLLNQIWKWCSIVVKSSALLSIWIFVIPVLIGLLFELLVIVPMRVPVDESPVFLLYQ 801
RT +LLN + + +++ L SIWI VIP ++GLL +L++I+P +VP+ ESPV+ L
Sbjct: 665 RTDLLLNHVLMF----IRNVLLFSIWISVIPGVLGLLIDLMIIIPSQVPLGESPVYNLLH 720
Query: 802 DWALGLIFLKIWTRLVMLDHMMPLMDESWRVKFERVRDDGFSRLQGLWVLREIVLPIIMK 861
DW +G++ L IW L ML + +WR K +R+R +RL W++R+++ II+
Sbjct: 721 DWLIGVVVLHIWIFLTMLTRINCFATVAWREKLQRIRSVTINRLPFTWLIRDVIGSIIVS 780
Query: 862 LLTALCVPYVLARGVFPALGYPLVVNSAVYRFAWLGCLSFSFVCFCAKRFHVWFTNLHNS 921
LL LCVPYV+ +FP LG+ VN V RF W L+ + F K LH
Sbjct: 781 LLFTLCVPYVVVNSLFPILGFSSAVNLTVQRFIWPAILALIPIWFSVKLIRDLILYLHKL 840
Query: 922 IRDDRYLIGRRLHNFGEHVE 941
D+RY +G RL +F E +E
Sbjct: 841 EFDNRYKVGERLVDFTEDLE 860
Score = 123 bits (308), Expect = 3e-26
Identities = 52/118 (44%), Positives = 77/118 (65%), Gaps = 7/118 (5%)
Query: 52 APPSAKYDDDDEDEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQ 111
A + + D + D+CRIC++P + DNPLR+PCAC GS+K++H DCL WLN
Sbjct: 16 AVTTEEVSDINNKAVDICRICQSPEEPDNPLRHPCACRGSLKYIHSDCLFLWLNRRKRNH 75
Query: 112 CEVCKHPFSFSPVYAENAPARLPFQEFVVGMAMKACHVLQFFVRLSFVLSVWLLIIPF 169
CE+CK +S PVY+ENAP RLP+ EF++G+ M+A +R ++ W+L++PF
Sbjct: 76 CEICKRSYSIVPVYSENAPERLPWHEFLMGLLMRA-------LRFMNLILPWILMMPF 126
Score = 83.6 bits (205), Expect = 3e-14
Identities = 42/137 (30%), Positives = 78/137 (56%), Gaps = 9/137 (6%)
Query: 464 FLAAMRHLMTMVKVAFLLVIELGVFPLMCGWWLDVCTIQMFGKTMVHRAQFFSASPLASS 523
+L ++R + VK F+L +LGV P + G WL CT + GKT H + S PL +
Sbjct: 337 YLTSVRTFLPSVKDTFILSFKLGVLPWLLGCWLHFCTFPILGKTASHTVEVLSDYPLMAD 396
Query: 524 LAHWVVGIVYMLQISIFVSLLRGVLRNGVLYFLRDPADPNYNPSRDLIDNPVHKHARRVL 583
HW++G +Y++ + L++ +++ L++L D A+PNY ++ H +L
Sbjct: 397 -KHWLMGTLYLVSALSCMELIQKIVQKRALWYLLDVAEPNYKVTK--------LHLGPIL 447
Query: 584 LSVAVYGSLMVMLVILP 600
L+ A++G+++V+++ LP
Sbjct: 448 LAFALHGTMVVIVLHLP 464
>gb|AAH46148.1| MARCH6 protein [Homo sapiens]
Length = 635
Score = 147 bits (371), Expect = 2e-33
Identities = 75/200 (37%), Positives = 122/200 (60%), Gaps = 27/200 (13%)
Query: 402 IGTSRISDVTTLAXGYIFLS-TLIFCYFGVVALIRYTKGEPLTAGRFYGIASIAETIPSL 460
+ S + T GYI L+ TLI C+ G+ L+++ + R G+ I
Sbjct: 50 VQASHFEGLITTIVGYILLAITLIICH-GLATLVKFHRSR-----RLLGVCYI------- 96
Query: 461 FRQFLAAMRHLMTMVKVAFLLVIELGVFPLMCGWWLDVCTIQMFGKTMVHRAQFFSASPL 520
+VKV+ L+V+E+GVFPL+CGWWLD+C+++MF T+ R F ++P
Sbjct: 97 -------------VVKVSLLVVVEIGVFPLICGWWLDICSLEMFDATLKDRELSFQSAPG 143
Query: 521 ASSLAHWVVGIVYMLQISIFVSLLRGVLRNGVLYFLRDPADPNYNPSRDLIDNPVHKHAR 580
+ HW+VG+VY+ + F+ LLR VLR GVL+FLR+ DP++NP +++I P+++H R
Sbjct: 144 TTMFLHWLVGMVYVFYFASFILLLREVLRPGVLWFLRNLNDPDFNPVQEMIHLPIYRHLR 203
Query: 581 RVLLSVAVYGSLMVMLVILP 600
R +LSV V+GS++++++ LP
Sbjct: 204 RFILSVIVFGSIVLLMLWLP 223
Score = 127 bits (320), Expect = 1e-27
Identities = 70/221 (31%), Positives = 127/221 (56%), Gaps = 18/221 (8%)
Query: 722 YVIWTAVAGVRYSIEQIRKRRTSVLLNQIWKWCSIVVKSSALLSIWIFVIPVLIGLLFEL 781
YV W + V + + + R V+ ++ +W +++K+ + + V+P+L+GLLFEL
Sbjct: 408 YVCWLTIRAVTVMVAWMPQGRR-VIFQKVKEWSLMIMKTLIVAVLLAGVVPLLLGLLFEL 466
Query: 782 LVIVPMRVPVDESPVFLLYQDWALGLIFLKIWTRLVMLDHMMPLMDESWRVK--FERVRD 839
+++ P+RVP+D++P+F +QDWALG++ KI + LM W +K E+V
Sbjct: 467 VIVAPLRVPLDQTPLFYPWQDWALGVLHAKIIAAIT-------LMGPQWWLKTVIEQVYA 519
Query: 840 DGFSRLQGLWVLREIVLPIIMKLLTALCVPYVLARGVFPALGYPL----VVNSAVYRFAW 895
+G + +++R++ P+I LL +LCVPYV+A GV P LG +V+ +Y F
Sbjct: 520 NGIRNIDLHYIVRKLAAPVISVLLLSLCVPYVIASGVVPLLGVTAEMQNLVHRRIYPFLL 579
Query: 896 LGCLSFSFVCFCAKRFHVWFTNLHNSIRDDRYLIGRRLHNF 936
+ + + + F ++ F L+ I++D+YL+G+RL N+
Sbjct: 580 MVVVLMAILSFQVRQ----FKRLYEHIKNDKYLVGQRLVNY 616
>dbj|BAC97979.1| mKIAA0597 protein [Mus musculus]
Length = 536
Score = 141 bits (356), Expect = 9e-32
Identities = 60/127 (47%), Positives = 97/127 (76%)
Query: 474 MVKVAFLLVIELGVFPLMCGWWLDVCTIQMFGKTMVHRAQFFSASPLASSLAHWVVGIVY 533
+VKV+ L+V+E+GVFPL+CGWWLD+C+++MF T+ R F ++P + HW+VG+VY
Sbjct: 25 VVKVSLLVVVEIGVFPLICGWWLDICSLEMFDATLKDRELSFQSAPGTTMFLHWLVGMVY 84
Query: 534 MLQISIFVSLLRGVLRNGVLYFLRDPADPNYNPSRDLIDNPVHKHARRVLLSVAVYGSLM 593
+ + F+ LLR VLR GVL+FLR+ DP++NP +++I P+++H RR +LSV V+GS++
Sbjct: 85 VFYFASFILLLREVLRPGVLWFLRNLNDPDFNPVQEMIHLPIYRHLRRFILSVIVFGSIV 144
Query: 594 VMLVILP 600
++++ LP
Sbjct: 145 LLMLWLP 151
Score = 123 bits (309), Expect = 3e-26
Identities = 79/258 (30%), Positives = 136/258 (52%), Gaps = 37/258 (14%)
Query: 698 PLHFAL-------FFCWSLLG*LSLSSTLPCYV------IWTAVAGVRYSIEQIRKRRTS 744
PL+F L F C +LL + TLP + WT A +I + T+
Sbjct: 278 PLNFPLRIFLLIVFMCITLLIASLICLTLPVFAGRWLMSFWTGTA-------KIHELYTA 330
Query: 745 VLLNQIWKWCSIVVKSSALLSIWIFVIPVLIGLLFELLVIVPMRVPVDESPVFLLYQDWA 804
+ +W +++K+ + + V+P+L+GLLFEL+++ P+RVP+D++P+F +QDWA
Sbjct: 331 ACGLYVKEWSLMIMKTLIVAVLLAGVVPLLLGLLFELVIVAPLRVPLDQTPLFYPWQDWA 390
Query: 805 LGLIFLKIWTRLVMLDHMMPLMDESWRVK--FERVRDDGFSRLQGLWVLREIVLPIIMKL 862
LG++ KI + LM W +K E+V +G + +++R++ P+I L
Sbjct: 391 LGVLHAKIIAAIT-------LMGPQWWLKTVIEQVYANGIRNIDLHYIIRKLAAPVISVL 443
Query: 863 LTALCVPYVLARGVFPALGYPL----VVNSAVYRFAWLGCLSFSFVCFCAKRFHVWFTNL 918
L +LCVPYV+A G P LG +V+ +Y F + + + F ++ F L
Sbjct: 444 LLSLCVPYVIASGAVPLLGVTAEMQNLVHRRIYPFLLMVVVLMGILSFQVRQ----FKRL 499
Query: 919 HNSIRDDRYLIGRRLHNF 936
+ I++D+YL+G+RL N+
Sbjct: 500 YEHIKNDKYLVGQRLVNY 517
>emb|CAF95396.1| unnamed protein product [Tetraodon nigroviridis]
Length = 1441
Score = 139 bits (350), Expect = 5e-31
Identities = 64/168 (38%), Positives = 94/168 (55%)
Query: 62 DEDEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHPFSF 121
D EED+CR+CR+ G D PL +PC C+GSIKF+HQ+CLLQWL HS CE+C+H F+F
Sbjct: 9 DTAEEDICRVCRSEGTQDRPLYHPCVCTGSIKFIHQECLLQWLKHSRKEYCELCQHRFAF 68
Query: 122 SPVYAENAPARLPFQEFVVGMAMKACHVLQFFVRLSFVLSVWLLIIPFITFWIWRLAFVR 181
+P+Y+ + P+RLP Q+ G+ ++++ + V WL ++P I++ F
Sbjct: 69 TPIYSPDMPSRLPVQDIFTGLLTSVGTAVRYWFHYTLVAFSWLGLVPLTACRIYKCLFAG 128
Query: 182 SFGEAQRLFFYHLFTAVILTDCLHGFLLSASIVFIFFGGTSLRDYFRH 229
S L L T +L DCL G + A + F LR+ H
Sbjct: 129 SVSSLLTLPLDLLSTQNLLADCLQGCFVVACTLGAFISLVWLREQIVH 176
Score = 127 bits (320), Expect = 1e-27
Identities = 69/181 (38%), Positives = 111/181 (61%), Gaps = 39/181 (21%)
Query: 456 TIPSLFRQFLAAM------RHLM----TMVKVAFLLVIELGVFPLMCGWWLDVCTIQMFG 505
T+PS Q LA++ RHL+ +VKV+ L+V+E+G+FPL+CGWWLD+C+++MF
Sbjct: 412 TVPS---QALASLVSFQRSRHLLGVCYIVVKVSLLVVMEIGLFPLICGWWLDICSLEMFD 468
Query: 506 KTMVHRAQFFSASPLASSLAHWVVGIVYMLQISIFVSLLR-------------------- 545
T+ R Q ++SP + HW+VG+VY+ ++ F+ LLR
Sbjct: 469 ATLKDRQQSLASSPGTTMFLHWLVGMVYVFYLASFILLLREVGQGLQPGWRLLGSTALQQ 528
Query: 546 ------GVLRNGVLYFLRDPADPNYNPSRDLIDNPVHKHARRVLLSVAVYGSLMVMLVIL 599
VLR GVL+FLR+ DP++NP +++I V++H RR +LSV V+GS++++++ L
Sbjct: 529 VLSLCSQVLRPGVLWFLRNLNDPDFNPVQEMIHLAVYRHLRRFILSVVVFGSIVLLMLWL 588
Query: 600 P 600
P
Sbjct: 589 P 589
Score = 37.4 bits (85), Expect = 2.5
Identities = 18/66 (27%), Positives = 38/66 (57%), Gaps = 3/66 (4%)
Query: 754 CSIVVKSSALLSIWIFVIPVLIGLLFELLVIVPMRVPVDESPVFLLYQDWALGLIFLKIW 813
C + ++++ ++ W+ P I L ++ VP+D++P+F +QDWALG++ KI
Sbjct: 794 CWLAIRAATVVLSWMPQGPSAIMLKVHEWMLT---VPLDQTPLFSPWQDWALGVLHAKII 850
Query: 814 TRLVML 819
+ ++
Sbjct: 851 AAITLM 856
>ref|XP_638225.1| hypothetical protein DDB0186668 [Dictyostelium discoideum]
gi|60466638|gb|EAL64690.1| hypothetical protein
DDB0186668 [Dictyostelium discoideum]
Length = 1088
Score = 132 bits (333), Expect = 4e-29
Identities = 52/118 (44%), Positives = 78/118 (66%)
Query: 62 DEDEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHPFSF 121
+++EED CR+CRN DNPL YPC CSGSIK++HQ+CLL+W+ HS + CE+C HPF F
Sbjct: 6 NQEEEDFCRVCRNGSTPDNPLSYPCKCSGSIKYIHQNCLLEWIQHSKSSSCELCGHPFRF 65
Query: 122 SPVYAENAPARLPFQEFVVGMAMKACHVLQFFVRLSFVLSVWLLIIPFITFWIWRLAF 179
+P+Y+ NAP +P E ++ ++ R+ +++ WL I+P +T WI+ F
Sbjct: 66 TPIYSPNAPEFIPSHELFYEALIRFKWYIKKISRILYIVFCWLFIVPTVTCWIFNFFF 123
Score = 104 bits (259), Expect = 2e-20
Identities = 62/208 (29%), Positives = 104/208 (49%), Gaps = 23/208 (11%)
Query: 750 IWKWCSIVVKSSALLSIWIFVIPVLIGLLFELLVIVPMRVPVDESPVFL----LYQDWAL 805
I +W SI +K + I + V+P+L G LF+ + +VP+ P DES F+ ++Q+W L
Sbjct: 867 IIQWVSIGLKVLLIGFIMVIVMPLLTGFLFDFIFMVPIMAPYDES-FFIHFGDIFQNWCL 925
Query: 806 GLIFLKIWTRLVMLDHMMPLMDES------------WRVKFERVRDDGFSRLQGLWVLRE 853
G + LK W R + + P + + W +F++ + +G S + W +
Sbjct: 926 GALLLKFWYRWINATNQNPDNNRNNVIEDLDQPRDRWIDRFKQFKRNGISNIDLKWTFSK 985
Query: 854 IVLPIIMKLLTALCVPYVLARGVFPALGYPLVVNSAVYRFAWLGCLSFSFVCFCAKRFH- 912
I+ PI L T L VP ++ + P G L++ S +R+ G + F+ K H
Sbjct: 986 IIFPICHYLFTLLTVPIFFSKFLVPLFGGSLILESISFRY---GFAVYCFILLFEKILHK 1042
Query: 913 --VWFTNLHNSIRDDRYLIGRRLHNFGE 938
W + N IRDD+YL+G++LHN +
Sbjct: 1043 IKQWSSRFPNMIRDDKYLVGKQLHNIDQ 1070
Score = 94.4 bits (233), Expect = 2e-17
Identities = 41/142 (28%), Positives = 83/142 (57%), Gaps = 1/142 (0%)
Query: 473 TMVKVAFLLVIELGVFPLMCGWWLDVCTIQMFGKTMVHRAQFFSASPLASSLAHWVVGIV 532
+ +K+ + + ELG+ P++ G ++D C++++FG ++ R F + + + W+ GI
Sbjct: 529 SFIKIGVITIFELGILPIVVGCFIDFCSLRIFGGSIEARLSFALSQKMTFLFSRWIFGIF 588
Query: 533 YMLQISIFVSLLRGVLRNGVLYFLRDPADPNYNPSRDLIDNPVHKHARRVLLSVAVYGSL 592
+M+ + S+ + R GV++FL+DP+DP+++P +D+I +H +V +S+ Y +
Sbjct: 589 FMVNFTNLCSIFHQIFRKGVIWFLKDPSDPDFDPFKDMIKLSFKRHLFKVFVSLCAYAII 648
Query: 593 MVMLVILPR-*LGIADGPLPIS 613
++ V LP L G LPI+
Sbjct: 649 GLLFVFLPALFLSTIPGFLPIN 670
>gb|EAA47953.1| hypothetical protein MG09083.4 [Magnaporthe grisea 70-15]
gi|39956377|ref|XP_364238.1| hypothetical protein
MG09083.4 [Magnaporthe grisea 70-15]
Length = 437
Score = 131 bits (330), Expect = 1e-28
Identities = 66/168 (39%), Positives = 91/168 (53%), Gaps = 16/168 (9%)
Query: 12 DGTPIAATTPSSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSAKYDDDDEDEEDVCRI 71
D P+A S S + +P G+ G A A AST P D CRI
Sbjct: 13 DPEPVAHARRRFSGTSQHNKAPFGAAG----ANAADPASTLDP------------DTCRI 56
Query: 72 CRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHPFSFSPVYAENAPA 131
CR AD PL YPC CSGSIK+VHQDCL++WL+HS + CE+CK PF F+ +Y P
Sbjct: 57 CRGEATADEPLFYPCKCSGSIKYVHQDCLMEWLSHSQKKHCELCKTPFRFTKLYDRKMPK 116
Query: 132 RLPFQEFVVGMAMKACHVLQFFVRLSFVLSVWLLIIPFITFWIWRLAF 179
+LPF F+ + + + ++R V+S+WL+ +P++ IW L F
Sbjct: 117 KLPFVVFITHIVKYMVNNVLVWLRAGLVVSIWLIWLPYLMRSIWSLMF 164
>gb|AAH48816.1| March6 protein [Mus musculus]
Length = 527
Score = 129 bits (324), Expect = 5e-28
Identities = 54/116 (46%), Positives = 87/116 (74%)
Query: 485 LGVFPLMCGWWLDVCTIQMFGKTMVHRAQFFSASPLASSLAHWVVGIVYMLQISIFVSLL 544
+GVFPL+CGWWLD+C+++MF T+ R F ++P + HW+VG+VY+ + F+ LL
Sbjct: 1 IGVFPLICGWWLDICSLEMFDATLKDRELSFQSAPGTTMFLHWLVGMVYVFYFASFILLL 60
Query: 545 RGVLRNGVLYFLRDPADPNYNPSRDLIDNPVHKHARRVLLSVAVYGSLMVMLVILP 600
R VLR GVL+FLR+ DP++NP +++I P+++H RR +LSV V+GS++++++ LP
Sbjct: 61 REVLRPGVLWFLRNLNDPDFNPVQEMIHLPIYRHLRRFILSVIVFGSIVLLMLWLP 116
Score = 125 bits (315), Expect = 5e-27
Identities = 69/221 (31%), Positives = 125/221 (56%), Gaps = 18/221 (8%)
Query: 722 YVIWTAVAGVRYSIEQIRKRRTSVLLNQIWKWCSIVVKSSALLSIWIFVIPVLIGLLFEL 781
YV W + V + + + R V+ ++ +W +++K+ + + V+P+L+GLLFEL
Sbjct: 300 YVCWLTIRAVTVLVAWMPQGRR-VIFQKVKEWSLMIMKTLIVAVLLAGVVPLLLGLLFEL 358
Query: 782 LVIVPMRVPVDESPVFLLYQDWALGLIFLKIWTRLVMLDHMMPLMDESWRVK--FERVRD 839
+++ P+RVP+D++P+F +QDWALG++ KI + LM W +K E+V
Sbjct: 359 VIVAPLRVPLDQTPLFYPWQDWALGVLHAKIIAAIT-------LMGPQWWLKTVIEQVYA 411
Query: 840 DGFSRLQGLWVLREIVLPIIMKLLTALCVPYVLARGVFPALGYPL----VVNSAVYRFAW 895
+G + +++R++ P+I LL +LCVPYV+A G P LG +V+ +Y F
Sbjct: 412 NGIRNIDLHYIIRKLAAPVISVLLLSLCVPYVIASGAVPLLGVTAEMQNLVHRRIYPFLL 471
Query: 896 LGCLSFSFVCFCAKRFHVWFTNLHNSIRDDRYLIGRRLHNF 936
+ + + F ++ F L+ I++D+YL+G+RL N+
Sbjct: 472 MVVVLMGILSFQVRQ----FKRLYEHIKNDKYLVGQRLVNY 508
>ref|XP_624755.1| PREDICTED: similar to Hypothetical protein CBG12203 [Apis
mellifera]
Length = 956
Score = 129 bits (324), Expect = 5e-28
Identities = 77/230 (33%), Positives = 126/230 (54%), Gaps = 13/230 (5%)
Query: 718 TLPC--YVIWTAVAGVRYSIEQIRKRRTSVLLNQIWKWCSIVVKSSALLSIWIFVIPVLI 775
T+ C YV W A G+ + + + R + L+++I W I VK+S + I +IP+L
Sbjct: 734 TIACGMYVCWAAARGLALAFSWLPRGRRA-LIDRIKHWAIIGVKASVAFFLLIGLIPLLF 792
Query: 776 GLLFELLVIVPMRVPVDESPVFLLYQDWALGLIFLKIWTRLVMLDHMMPLMDESWRVK-- 833
GLL EL+V+VP+RVP++++ + ++QDWALG+++ KI T ++M M WR++
Sbjct: 793 GLLLELVVVVPLRVPLEQNAILFIWQDWALGVLYTKIATAIIM-------MGPDWRIRLA 845
Query: 834 FERVRDDGFSRLQGLWVLREIVLPIIMKLLTALCVPYVLARGVFPALGYPLVVNSAVYRF 893
ER +DG + +++ E+ P+I AL VPY +A G+ P L L + R
Sbjct: 846 IERAYNDGIREIDLKFIVTELAAPVICCFGLALAVPYAVAYGIVPLLITNLQTQILIARH 905
Query: 894 AWLGCLSFSFVCFCAKRFHVWFTNLHNSIRDDRYLIGRRLHNFGEHVEKA 943
+ L VC F L+ I++D+YL+G+RL N+ EH K+
Sbjct: 906 LYPFLLLVHVVCIVIYFQIRQFKKLYEHIKNDKYLVGQRLVNY-EHQNKS 954
Score = 129 bits (323), Expect = 6e-28
Identities = 62/176 (35%), Positives = 103/176 (58%), Gaps = 17/176 (9%)
Query: 425 FCYFGVVALIRYTKGEPLTAGRFYGIASIAETIPSLFRQFLAAMRHLMTMVKVAFLLVIE 484
+C GV ++ +T PL R + + I VKV+ L V+E
Sbjct: 430 YCVTGVGLVVLHTLAAPLGFQRSQRLLGLCYVI-----------------VKVSLLSVVE 472
Query: 485 LGVFPLMCGWWLDVCTIQMFGKTMVHRAQFFSASPLASSLAHWVVGIVYMLQISIFVSLL 544
+GV PL+CGWWLD+C++ MF T+ R F +P S HW+VG++Y+ + F+ LL
Sbjct: 473 IGVLPLVCGWWLDICSLAMFDATLRDRESSFKLAPGTSMFLHWLVGMIYIYYFASFILLL 532
Query: 545 RGVLRNGVLYFLRDPADPNYNPSRDLIDNPVHKHARRVLLSVAVYGSLMVMLVILP 600
R VLR GVL+FLR+ DP+++P +++I P+ +H RR++ S ++G+ +++++ LP
Sbjct: 533 REVLRPGVLWFLRNLNDPDFSPIQEMIHLPILRHVRRLVASAVIFGTAILLMLWLP 588
Score = 120 bits (301), Expect = 2e-25
Identities = 61/186 (32%), Positives = 96/186 (50%), Gaps = 5/186 (2%)
Query: 61 DDEDEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHPFS 120
+D D+CR+CR+ G AD PL +PC C+GSIK++HQ+CL+QW+ +S CE+C + FS
Sbjct: 3 EDMSGADICRVCRSEGLADRPLFHPCICTGSIKWIHQECLVQWMRYSRKEYCELCGYRFS 62
Query: 121 FSPVYAENAPARLPFQEFVVGMAMKACHVLQFFVRLSFVLSVWLLIIPFITFWIWRLAFV 180
F+P+Y+ + P RLP ++ + G+ +++++ + V WL ++P +R F
Sbjct: 63 FTPIYSPDMPRRLPLRDVIGGLFSNIVTAVKYWLHYTLVAIAWLGVVPLTACRTYRALFS 122
Query: 181 RSFG--EAQRLFFYHLFTAVILTDCLHGFLLSASIVFIFFGGTSLRDYFRHLREIGGQDA 238
L L I +D HG + +F F G LR+ H GG D
Sbjct: 123 GPLDLVRIMSLPMDMLSAENISSDVFHGCFVVTCTLFAFIGLVWLREQILH---AGGPDW 179
Query: 239 EREDEV 244
D V
Sbjct: 180 LDRDNV 185
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.328 0.141 0.441
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,583,846,863
Number of Sequences: 2540612
Number of extensions: 65992925
Number of successful extensions: 385042
Number of sequences better than 10.0: 524
Number of HSP's better than 10.0 without gapping: 290
Number of HSP's successfully gapped in prelim test: 234
Number of HSP's that attempted gapping in prelim test: 383664
Number of HSP's gapped (non-prelim): 1031
length of query: 982
length of database: 863,360,394
effective HSP length: 138
effective length of query: 844
effective length of database: 512,755,938
effective search space: 432766011672
effective search space used: 432766011672
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 80 (35.4 bits)
Medicago: description of AC136449.11


