

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135848.1 + phase: 0
(64 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
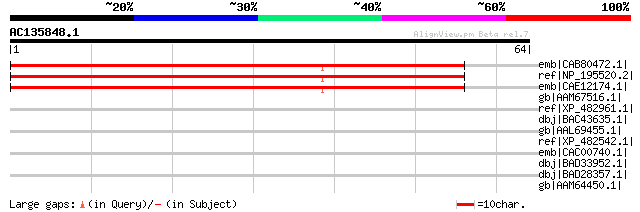
Score E
Sequences producing significant alignments: (bits) Value
emb|CAB80472.1| hypothetical protein [Arabidopsis thaliana] gi|4... 53 2e-06
ref|NP_195520.2| bHLH family protein [Arabidopsis thaliana] 53 2e-06
emb|CAE12174.1| putative bHLH131 transcription factor [Arabidops... 53 2e-06
gb|AAM67516.1| putative DNA-binding protein [Arabidopsis thalian... 40 0.012
ref|XP_482961.1| bHLH protein family-like [Oryza sativa (japonic... 39 0.036
dbj|BAC43635.1| putative bHLH transcription factor bHLH106 [Arab... 39 0.036
gb|AAL69455.1| At2g41130/T3K9.10 [Arabidopsis thaliana] 39 0.036
ref|XP_482542.1| DNA-binding protein-like [Oryza sativa (japonic... 36 0.23
emb|CAC00740.1| putative HLH DNA binding protein [Arabidopsis th... 33 2.6
dbj|BAD33952.1| bHLH-like protein [Oryza sativa (japonica cultiv... 32 3.4
dbj|BAD28357.1| DNA binding protein-like [Oryza sativa (japonica... 32 3.4
gb|AAM64450.1| putative HLH DNA-binding protein [Arabidopsis tha... 32 5.7
>emb|CAB80472.1| hypothetical protein [Arabidopsis thaliana]
gi|4467113|emb|CAB37547.1| hypothetical protein
[Arabidopsis thaliana] gi|7485848|pir||T05634
hypothetical protein F20D10.190 - Arabidopsis thaliana
Length = 1496
Score = 52.8 bits (125), Expect = 2e-06
Identities = 27/57 (47%), Positives = 42/57 (73%), Gaps = 1/57 (1%)
Query: 1 MSSISRALLSMKAKVVKVEMVTVGGRTRSVLWVQGVE-NESLGMLKSTLKVVMHKPS 56
MS ++ ++ ++KAK V+ E++TVGGRT+ L+VQGV NE L LK +LK+V++ S
Sbjct: 1422 MSEVAESMKAVKAKAVRAEIMTVGGRTKCALFVQGVNGNEGLVKLKKSLKLVVNGKS 1478
>ref|NP_195520.2| bHLH family protein [Arabidopsis thaliana]
Length = 1513
Score = 52.8 bits (125), Expect = 2e-06
Identities = 27/57 (47%), Positives = 42/57 (73%), Gaps = 1/57 (1%)
Query: 1 MSSISRALLSMKAKVVKVEMVTVGGRTRSVLWVQGVE-NESLGMLKSTLKVVMHKPS 56
MS ++ ++ ++KAK V+ E++TVGGRT+ L+VQGV NE L LK +LK+V++ S
Sbjct: 1439 MSEVAESMKAVKAKAVRAEIMTVGGRTKCALFVQGVNGNEGLVKLKKSLKLVVNGKS 1495
>emb|CAE12174.1| putative bHLH131 transcription factor [Arabidopsis thaliana]
Length = 256
Score = 52.8 bits (125), Expect = 2e-06
Identities = 27/57 (47%), Positives = 42/57 (73%), Gaps = 1/57 (1%)
Query: 1 MSSISRALLSMKAKVVKVEMVTVGGRTRSVLWVQGVE-NESLGMLKSTLKVVMHKPS 56
MS ++ ++ ++KAK V+ E++TVGGRT+ L+VQGV NE L LK +LK+V++ S
Sbjct: 182 MSEVAESMKAVKAKAVRAEIMTVGGRTKCALFVQGVNGNEGLVKLKKSLKLVVNGKS 238
>gb|AAM67516.1| putative DNA-binding protein [Arabidopsis thaliana]
gi|18176098|gb|AAL59983.1| putative DNA-binding protein
[Arabidopsis thaliana] gi|18409132|ref|NP_564944.1|
basic helix-loop-helix (bHLH) family protein
[Arabidopsis thaliana] gi|12324140|gb|AAG52041.1|
putative DNA-binding protein; 36199-34606 [Arabidopsis
thaliana] gi|12323209|gb|AAG51581.1| putative
DNA-binding protein [Arabidopsis thaliana]
gi|25404781|pir||H96712 probable DNA-binding protein
T6L1.1 [imported] - Arabidopsis thaliana
Length = 368
Score = 40.4 bits (93), Expect = 0.012
Identities = 21/57 (36%), Positives = 33/57 (57%), Gaps = 8/57 (14%)
Query: 6 RALLSMKAKVVKVEMVTVGGRTRSVLWVQGVENES--------LGMLKSTLKVVMHK 54
+ L +M+ K +K E+ TVGGR ++VL+V G E+ +G ++ LK VM K
Sbjct: 277 KTLKAMRLKTLKAEITTVGGRVKNVLFVTGEESSGEEVEEEYCIGTIEEALKAVMEK 333
>ref|XP_482961.1| bHLH protein family-like [Oryza sativa (japonica cultivar-group)]
gi|42407861|dbj|BAD09003.1| bHLH protein family-like
[Oryza sativa (japonica cultivar-group)]
Length = 223
Score = 38.9 bits (89), Expect = 0.036
Identities = 24/61 (39%), Positives = 38/61 (61%), Gaps = 9/61 (14%)
Query: 1 MSSISRALLSMKAKVVKVEMVTVGGRTRSVLWVQGV------ENE---SLGMLKSTLKVV 51
MS + RA+ S+ A+ V+ E+ TVGGRTRSVL + V +N+ +L L++ L+ V
Sbjct: 132 MSDLGRAVRSVSARPVRAEVATVGGRTRSVLELDVVVASDAADNDRAVALSALRAALRTV 191
Query: 52 M 52
+
Sbjct: 192 L 192
>dbj|BAC43635.1| putative bHLH transcription factor bHLH106 [Arabidopsis thaliana]
gi|3402704|gb|AAD11998.1| unknown protein [Arabidopsis
thaliana] gi|15226839|ref|NP_181646.1| basic
helix-loop-helix (bHLH) family protein [Arabidopsis
thaliana] gi|7487617|pir||T02106 hypothetical protein
At2g41130 [imported] - Arabidopsis thaliana
Length = 253
Score = 38.9 bits (89), Expect = 0.036
Identities = 22/60 (36%), Positives = 35/60 (57%), Gaps = 4/60 (6%)
Query: 1 MSSISRALLSMKAKVVKVEMVTVGGRTRSVLWVQGVEN----ESLGMLKSTLKVVMHKPS 56
+ + L S+ K ++ EMVT+GGRTRSVL V + ES+ L++ LK ++ + S
Sbjct: 166 LPDLMEILKSLNMKTLRAEMVTIGGRTRSVLVVAADKEMHGVESVHFLQNALKSLLERSS 225
>gb|AAL69455.1| At2g41130/T3K9.10 [Arabidopsis thaliana]
Length = 245
Score = 38.9 bits (89), Expect = 0.036
Identities = 22/60 (36%), Positives = 35/60 (57%), Gaps = 4/60 (6%)
Query: 1 MSSISRALLSMKAKVVKVEMVTVGGRTRSVLWVQGVEN----ESLGMLKSTLKVVMHKPS 56
+ + L S+ K ++ EMVT+GGRTRSVL V + ES+ L++ LK ++ + S
Sbjct: 158 LPDLMEILKSLNMKTLRAEMVTIGGRTRSVLVVAADKEMHGVESVHFLQNALKSLLERSS 217
>ref|XP_482542.1| DNA-binding protein-like [Oryza sativa (japonica cultivar-group)]
gi|42409474|dbj|BAD09830.1| DNA-binding protein-like
[Oryza sativa (japonica cultivar-group)]
Length = 345
Score = 36.2 bits (82), Expect = 0.23
Identities = 16/36 (44%), Positives = 25/36 (69%)
Query: 4 ISRALLSMKAKVVKVEMVTVGGRTRSVLWVQGVENE 39
I+RAL +++ + + E+ T+GGR RSVL + VE E
Sbjct: 231 IARALAALRLRARRAEIATLGGRVRSVLLIAAVEEE 266
>emb|CAC00740.1| putative HLH DNA binding protein [Arabidopsis thaliana]
gi|15229004|ref|NP_191236.1| basic helix-loop-helix
(bHLH) family protein [Arabidopsis thaliana]
gi|11292134|pir||T51265 probable HLH DNA binding protein
- Arabidopsis thaliana
Length = 230
Score = 32.7 bits (73), Expect = 2.6
Identities = 18/60 (30%), Positives = 34/60 (56%), Gaps = 4/60 (6%)
Query: 1 MSSISRALLSMKAKVVKVEMVTVGGRTRSVLWVQGVEN----ESLGMLKSTLKVVMHKPS 56
+ + L S++ + + +M TVGGRTR+VL V + +S+ L++ LK ++ + S
Sbjct: 143 LKDLMETLKSLQMETLFADMTTVGGRTRNVLVVAADKEHHGVQSVNFLQNALKSLLERSS 202
>dbj|BAD33952.1| bHLH-like protein [Oryza sativa (japonica cultivar-group)]
Length = 215
Score = 32.3 bits (72), Expect = 3.4
Identities = 16/31 (51%), Positives = 21/31 (67%)
Query: 1 MSSISRALLSMKAKVVKVEMVTVGGRTRSVL 31
MS + A S+ A+ V+ E+ TVGGRTRS L
Sbjct: 132 MSELGDAERSVSARAVRAEIATVGGRTRSDL 162
>dbj|BAD28357.1| DNA binding protein-like [Oryza sativa (japonica cultivar-group)]
Length = 363
Score = 32.3 bits (72), Expect = 3.4
Identities = 14/36 (38%), Positives = 24/36 (65%)
Query: 4 ISRALLSMKAKVVKVEMVTVGGRTRSVLWVQGVENE 39
I+RAL +++ + + E+ T+GGR RSVL + E +
Sbjct: 230 IARALAALRLRARRAEITTLGGRVRSVLLITADEQQ 265
>gb|AAM64450.1| putative HLH DNA-binding protein [Arabidopsis thaliana]
gi|7939557|dbj|BAA95758.1| DNA-binding protein-like
[Arabidopsis thaliana] gi|18958020|gb|AAL79583.1|
AT3g25710/K13N2_1 [Arabidopsis thaliana]
gi|16604444|gb|AAL24228.1| AT3g25710/K13N2_1
[Arabidopsis thaliana] gi|15230869|ref|NP_189199.1|
basic helix-loop-helix (bHLH) family protein
[Arabidopsis thaliana]
Length = 344
Score = 31.6 bits (70), Expect = 5.7
Identities = 12/33 (36%), Positives = 23/33 (69%)
Query: 1 MSSISRALLSMKAKVVKVEMVTVGGRTRSVLWV 33
M + AL S++ + +K E+ TVGGR +++L++
Sbjct: 228 MHDVINALKSLRLRTLKAEIATVGGRVKNILFL 260
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.319 0.129 0.343
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 77,000,062
Number of Sequences: 2540612
Number of extensions: 1764623
Number of successful extensions: 3447
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 3436
Number of HSP's gapped (non-prelim): 12
length of query: 64
length of database: 863,360,394
effective HSP length: 40
effective length of query: 24
effective length of database: 761,735,914
effective search space: 18281661936
effective search space used: 18281661936
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 68 (30.8 bits)
Medicago: description of AC135848.1


