

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135799.3 + phase: 0
(242 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
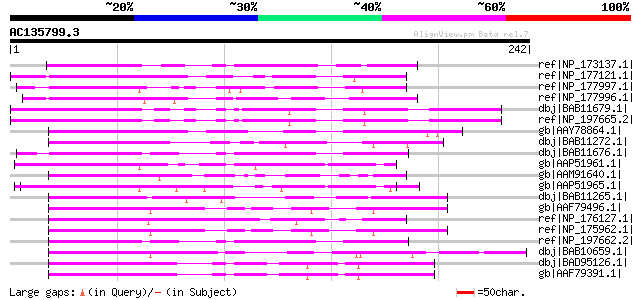
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_173137.1| F-box family protein [Arabidopsis thaliana] gi|... 86 1e-15
ref|NP_177121.1| F-box family protein [Arabidopsis thaliana] gi|... 78 2e-13
ref|NP_177997.1| F-box family protein [Arabidopsis thaliana] gi|... 75 2e-12
ref|NP_177996.1| F-box family protein [Arabidopsis thaliana] gi|... 74 4e-12
dbj|BAB11679.1| unnamed protein product [Arabidopsis thaliana] 73 6e-12
ref|NP_197665.2| F-box family protein [Arabidopsis thaliana] 73 6e-12
gb|AAY78864.1| F-box family protein [Arabidopsis thaliana] gi|18... 71 2e-11
dbj|BAB11272.1| unnamed protein product [Arabidopsis thaliana] g... 70 5e-11
dbj|BAB11676.1| unnamed protein product [Arabidopsis thaliana] 70 5e-11
gb|AAP51961.1| unknown protein [Oryza sativa (japonica cultivar-... 70 7e-11
gb|AAM91640.1| unknown protein [Arabidopsis thaliana] gi|1841414... 69 9e-11
gb|AAP51965.1| unknown protein [Oryza sativa (japonica cultivar-... 69 1e-10
dbj|BAB11265.1| unnamed protein product [Arabidopsis thaliana] g... 68 2e-10
gb|AAF79496.1| F20N2.9 [Arabidopsis thaliana] 68 3e-10
ref|NP_176127.1| F-box family protein [Arabidopsis thaliana] 68 3e-10
ref|NP_175962.1| F-box family protein [Arabidopsis thaliana] 68 3e-10
ref|NP_197662.2| F-box family protein [Arabidopsis thaliana] 68 3e-10
dbj|BAB10659.1| unnamed protein product [Arabidopsis thaliana] g... 67 4e-10
dbj|BAD95126.1| hypothetical protein [Arabidopsis thaliana] 67 6e-10
gb|AAF79391.1| F16A14.1 [Arabidopsis thaliana] gi|15222948|ref|N... 67 6e-10
>ref|NP_173137.1| F-box family protein [Arabidopsis thaliana] gi|25518753|pir||H86304
hypothetical protein F6I1.6 - Arabidopsis thaliana
gi|9802769|gb|AAF99838.1| Hypothetical protein
[Arabidopsis thaliana]
Length = 449
Score = 85.5 bits (210), Expect = 1e-15
Identities = 56/173 (32%), Positives = 93/173 (53%), Gaps = 33/173 (19%)
Query: 18 ADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKF 77
+D+IS LPD +L ILS +STKE+V TS+LSKRW NLWL +P +D ++SN+
Sbjct: 14 SDRISNLPDSLLCQILSDLSTKESVCTSVLSKRWRNLWLHVPVLD--------LDSNNFP 65
Query: 78 IDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWVNLV 137
D V+ F+NRF + N L + + + + W+N V
Sbjct: 66 DDDVF------------VSFVNRF---LGSENEQHLERFKLIYEVNEHDASRFKSWINAV 110
Query: 138 VQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRTLVSLDLHHFSV 190
++RR+ + +++H +VDD D L K+P+++++C LV+L L+ ++
Sbjct: 111 IKRRVCH--------FNVHNEVDDDDE--LVKMPLSLYSCERLVNLQLYRVAL 153
>ref|NP_177121.1| F-box family protein [Arabidopsis thaliana]
gi|12325183|gb|AAG52534.1| hypothetical protein;
77728-79262 [Arabidopsis thaliana]
gi|25404821|pir||A96718 hypothetical protein T6C23.17
[imported] - Arabidopsis thaliana
gi|10092292|gb|AAG12704.1| hypothetical protein;
7662-9196 [Arabidopsis thaliana]
Length = 451
Score = 77.8 bits (190), Expect = 2e-13
Identities = 56/186 (30%), Positives = 87/186 (46%), Gaps = 34/186 (18%)
Query: 1 MQRQKRRRRSSKASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPN 60
M + + +K S V D IS+LPD +L +L + TK+ V TS+LS+RW NLW +P
Sbjct: 1 MDEDREKHVRAKGSDEV-DWISKLPDCLLCEVLLNLPTKDVVKTSVLSRRWRNLWKHVPG 59
Query: 61 IDFNNIKIDSIESNSKFIDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPH 120
+D +N + F+DS F+ +F L Y D+
Sbjct: 60 LDLDNTDFQEFNTFLSFVDSFLD--------FNSESFLQKFILK---------YDCDDE- 101
Query: 121 LLYKRSCHNVVKWVNLVVQRRLKYLRLNLRLGYDLHLDV-DDYDNSYLPKLPITIFTCRT 179
Y + +W+N +V R+++ H+DV DD S+ +LP +I+TC +
Sbjct: 102 --YDPDIFLIGRWINTIVTRKVQ------------HIDVLDDSYGSWEVQLPSSIYTCES 147
Query: 180 LVSLDL 185
LVSL L
Sbjct: 148 LVSLKL 153
>ref|NP_177997.1| F-box family protein [Arabidopsis thaliana]
gi|3834318|gb|AAC83034.1| Similar to gi|2244754 heat
shock transcription factor HSF30 homolog from
Arabidopsis thaliana chromosome 4 contig gb|Z97335
gi|25406567|pir||F96816 hypothetical protein F9K20.20
[imported] - Arabidopsis thaliana
Length = 452
Score = 74.7 bits (182), Expect = 2e-12
Identities = 60/188 (31%), Positives = 98/188 (51%), Gaps = 50/188 (26%)
Query: 4 QKRRRRSSKASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLP--NI 61
+KR RRS + D++S LP+ ++ I+ + TK+ V +S+LSKRW NLW +P N+
Sbjct: 7 EKRARRSEE------DRLSNLPESLICQIMLNIPTKDVVKSSVLSKRWRNLWRYVPGLNV 60
Query: 62 DFNNIKIDSIESNSKFIDSVYSVLVSPDTAIGGSHFINRF-CLDVQ--FRNGNPLYGYDN 118
++N +F+D Y+ VS F++RF LD + F++ Y D
Sbjct: 61 EYN-----------QFLD--YNAFVS---------FVDRFLALDRESCFQSFRLRYDCDE 98
Query: 119 PHLLYKRSCHNVVKWVNLVVQRRLKYLRLNLRLGYDLHLDVDDYD-NSYLPKLPITIFTC 177
+R+ NV +W+N+VV ++LK LDV DY + ++P +++TC
Sbjct: 99 E----ERTISNVKRWINIVVDQKLKV------------LDVLDYTWGNEEVQIPPSVYTC 142
Query: 178 RTLVSLDL 185
+LVSL L
Sbjct: 143 ESLVSLKL 150
>ref|NP_177996.1| F-box family protein [Arabidopsis thaliana]
gi|3834319|gb|AAC83035.1| Similar to gi|2244754 heat
shock transcription factor HSF30 homolog from
Arabidopsis thaliana chromosome 4 contig gb|Z97335
gi|25406566|pir||E96816 hypothetical protein F9K20.21
[imported] - Arabidopsis thaliana
Length = 458
Score = 73.9 bits (180), Expect = 4e-12
Identities = 61/191 (31%), Positives = 91/191 (46%), Gaps = 39/191 (20%)
Query: 7 RRRSSKASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNI----- 61
+R +K S V D IS LP+ +L +L + TK+ V +S+LS RW NLW +P
Sbjct: 7 KRVRAKGSREV-DWISNLPETLLCQVLFYLPTKDVVKSSVLSSRWRNLWKYVPGFNLSYC 65
Query: 62 DFNNIKIDSIESNS--KFIDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNP 119
DF+ S + N+ +F+DS F ++ CL FR GY P
Sbjct: 66 DFHVRNTFSYDHNTFLRFVDSFMG-------------FNSQSCLQ-SFRLEYDSSGYGEP 111
Query: 120 HLLYKRSCHNVVKWVNLVVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRT 179
L R +W+N VV R++KYL + +DD ++Y ++P T++TC T
Sbjct: 112 KLALIR------RWINSVVSRKVKYLGV-----------LDDSCDNYEFEMPPTLYTCET 154
Query: 180 LVSLDLHHFSV 190
LV L L S+
Sbjct: 155 LVYLTLDGLSL 165
>dbj|BAB11679.1| unnamed protein product [Arabidopsis thaliana]
Length = 457
Score = 73.2 bits (178), Expect = 6e-12
Identities = 64/233 (27%), Positives = 108/233 (45%), Gaps = 49/233 (21%)
Query: 1 MQRQKRRRRSSKASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPN 60
M+ + R +A D IS+LPD ++ IL + K+ V TS LS RW +LWL +P
Sbjct: 1 MKAKYGERSQGTYFMAGEDLISKLPDSLITQILLYLPIKDIVRTSSLSSRWKSLWLLIPR 60
Query: 61 IDFNNIKIDSIESNSKFIDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPH 120
+D +DS E Y+ V F+N+F + F +G D
Sbjct: 61 LD-----LDSEEFQD------YNAFVG---------FMNKF---IDF-SGEEKICLDKLK 96
Query: 121 LLYKRSCHN---VVKWVNLVVQRRLKYLRLNLRLGYDLHLDVDDYDN-SYLPKLPITIFT 176
L +++ ++ V +W++ VV+R+LK HLDV+ N +L ++P++++
Sbjct: 97 LSSRKTVNDLPCVTRWIDFVVRRKLK------------HLDVECLVNRKFLEEMPLSLYV 144
Query: 177 CRTLVSLDLHHFSVKGVCYRFLLKALSTSNSLRLDTFKLYRSIYQVRFIMNTM 229
C TLV+L LH R LL + L T +L ++Y ++ ++
Sbjct: 145 CDTLVNLRLH---------RVLLGKFEAVSLPCLKTMRLEENVYANDVVLESL 188
>ref|NP_197665.2| F-box family protein [Arabidopsis thaliana]
Length = 466
Score = 73.2 bits (178), Expect = 6e-12
Identities = 64/233 (27%), Positives = 108/233 (45%), Gaps = 49/233 (21%)
Query: 1 MQRQKRRRRSSKASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPN 60
M+ + R +A D IS+LPD ++ IL + K+ V TS LS RW +LWL +P
Sbjct: 10 MKAKYGERSQGTYFMAGEDLISKLPDSLITQILLYLPIKDIVRTSSLSSRWKSLWLLIPR 69
Query: 61 IDFNNIKIDSIESNSKFIDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPH 120
+D +DS E Y+ V F+N+F + F +G D
Sbjct: 70 LD-----LDSEEFQD------YNAFVG---------FMNKF---IDF-SGEEKICLDKLK 105
Query: 121 LLYKRSCHN---VVKWVNLVVQRRLKYLRLNLRLGYDLHLDVDDYDN-SYLPKLPITIFT 176
L +++ ++ V +W++ VV+R+LK HLDV+ N +L ++P++++
Sbjct: 106 LSSRKTVNDLPCVTRWIDFVVRRKLK------------HLDVECLVNRKFLEEMPLSLYV 153
Query: 177 CRTLVSLDLHHFSVKGVCYRFLLKALSTSNSLRLDTFKLYRSIYQVRFIMNTM 229
C TLV+L LH R LL + L T +L ++Y ++ ++
Sbjct: 154 CDTLVNLRLH---------RVLLGKFEAVSLPCLKTMRLEENVYANDVVLESL 197
>gb|AAY78864.1| F-box family protein [Arabidopsis thaliana]
gi|18423560|ref|NP_568800.1| F-box family protein
(FBL13) [Arabidopsis thaliana]
Length = 444
Score = 71.2 bits (173), Expect = 2e-11
Identities = 53/200 (26%), Positives = 94/200 (46%), Gaps = 43/200 (21%)
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKFI 78
+++S+LPD ++ ILS +STK+AV TSILS RW NLW +P +DF++ ++ S F
Sbjct: 18 ERLSQLPDHLICVILSHLSTKDAVRTSILSTRWRNLWQLVPVLDFDSRELRSFSEFVSFA 77
Query: 79 DSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWVNLVV 138
S + + +I + + + GN + + W++LV
Sbjct: 78 GSFFY--------LHKDSYIQKLRVCIYDLAGN----------------YYLTSWIDLVT 113
Query: 139 QRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRTLVSLDLHHFSVKGV----- 193
+ R++ H+D+ + S +P++++TC TLV L L ++ V
Sbjct: 114 RHRIQ------------HIDISVFTCSGFGVIPLSLYTCDTLVHLKLSRVTMVNVEFVSL 161
Query: 194 -CYRFL-LKALSTSNSLRLD 211
C + L L ++ +N LD
Sbjct: 162 PCLKILDLDFVNFTNETTLD 181
>dbj|BAB11272.1| unnamed protein product [Arabidopsis thaliana]
gi|15241216|ref|NP_200455.1| F-box family protein
[Arabidopsis thaliana]
Length = 430
Score = 70.1 bits (170), Expect = 5e-11
Identities = 55/192 (28%), Positives = 90/192 (46%), Gaps = 48/192 (25%)
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKFI 78
D+IS LPDDV+ ILS V TK V+T++LSKRW LW +P +D+ + + E
Sbjct: 2 DRISLLPDDVVFKILSFVPTKVVVSTNLLSKRWRYLWKHVPKLDYRDPSLVDTEHWR--- 58
Query: 79 DSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSC--HNVVKWVNL 136
S F+++F L + P+ + HL R+C ++ W+++
Sbjct: 59 ---------------ASRFVDKFLL----LHEAPVL--ETLHLSLSRNCPPSDIETWISV 97
Query: 137 VVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLP-----KLPITIFTCRTLVSLD-LHHFSV 190
+ RR++ L + Y+P +LP +++TC TLVSL L F+V
Sbjct: 98 AISRRVRNLHIY----------------RYIPSTGPIRLPRSLYTCETLVSLYLLLDFTV 141
Query: 191 KGVCYRFLLKAL 202
+ F ++L
Sbjct: 142 DDAPFMFCFRSL 153
>dbj|BAB11676.1| unnamed protein product [Arabidopsis thaliana]
Length = 452
Score = 70.1 bits (170), Expect = 5e-11
Identities = 55/183 (30%), Positives = 87/183 (47%), Gaps = 40/183 (21%)
Query: 4 QKRRRRSSKASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDF 63
QKR + S++ D+IS LPD++L ILS + TK AV TSILS RW ++WLS P +D
Sbjct: 11 QKRNQMSNRG-----DRISSLPDELLCQILSNLPTKNAVTTSILSTRWRSIWLSTPVLD- 64
Query: 64 NNIKIDSIESNSKFIDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLY 123
I ID+ + + F+ F +RF ++F + L+ +
Sbjct: 65 --IDIDAFDDATTFVS-----------------FASRF---LEFSKDSCLHKFKLSVERD 102
Query: 124 KRSCHNVVKWVNLVVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRTLVSL 183
++ W+ V RR++ HL+VD + + + +T++ TLVSL
Sbjct: 103 DVDMCTIMPWIQDAVNRRIQ------------HLEVDCRFSFHFEAVYLTLYLSETLVSL 150
Query: 184 DLH 186
LH
Sbjct: 151 RLH 153
>gb|AAP51961.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|37530744|ref|NP_919674.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
gi|20042913|gb|AAM08741.1| Unknown protein [Oryza sativa
(japonica cultivar-group)]
Length = 373
Score = 69.7 bits (169), Expect = 7e-11
Identities = 50/187 (26%), Positives = 95/187 (50%), Gaps = 26/187 (13%)
Query: 3 RQKRRRRSSKASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLP--N 60
R++RRR + A D IS LPD++LH+I+S ++ ++AV T +LS+RW++LW ++P N
Sbjct: 7 RRRRRRVKAPTRAAGMDWISGLPDEILHHIMSFLNARQAVQTCVLSRRWSDLWRTVPCIN 66
Query: 61 IDFNNIKIDSIESNSKFIDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPL------- 113
DFN + + D Y + + F+NR ++ R+ +
Sbjct: 67 ADFNEFDFIDYQGD----DEDY------NDEVAFKRFVNRM---LELRDPATMMDKFWLK 113
Query: 114 YGYDNPHLLYKRSCHNVVKWVNLVVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPIT 173
Y + + YK S + +W++ +Q++ + + + + + L LD + + YL K+
Sbjct: 114 YKISDGYNEYKDSNVDANRWISHALQKQARVMEV-VVFSFPLELDHSVFTSCYLRKIG-- 170
Query: 174 IFTCRTL 180
F+C +L
Sbjct: 171 -FSCVSL 176
>gb|AAM91640.1| unknown protein [Arabidopsis thaliana] gi|18414142|ref|NP_567422.1|
F-box family protein [Arabidopsis thaliana]
Length = 381
Score = 69.3 bits (168), Expect = 9e-11
Identities = 55/171 (32%), Positives = 91/171 (53%), Gaps = 38/171 (22%)
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKI----DSIESN 74
D IS LPDD+ +ILS + TKEA +TS+LSK+W L+ +PN+D ++ + E +
Sbjct: 8 DVISSLPDDISSHILSFLPTKEAASTSVLSKKWRYLFAFVPNLDLDDSVYLNPENETEIS 67
Query: 75 SKFIDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWV 134
+ F+D V VL A+ G+ +++F L + G+ G D ++ W+
Sbjct: 68 TSFMDFVDRVL-----ALQGNSPLHKFSLKI----GD---GIDPV---------RIIPWI 106
Query: 135 NLVVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRTLVSLDL 185
N V++R + L DLHL++ ++ +L LP ++ C+TLV L L
Sbjct: 107 NNVLERGVSDL--------DLHLNL---ESEFL--LPSQVYLCKTLVWLKL 144
>gb|AAP51965.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|37530752|ref|NP_919678.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
gi|20042917|gb|AAM08745.1| Unknown protein [Oryza sativa
(japonica cultivar-group)]
Length = 949
Score = 68.9 bits (167), Expect = 1e-10
Identities = 53/188 (28%), Positives = 95/188 (50%), Gaps = 28/188 (14%)
Query: 3 RQKRRRRSSKASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLP--N 60
R K RR + A D+IS LPD +LH+I+S ++T++AV T +LS+RW NLW ++P N
Sbjct: 74 RHKLRRVKAHVRAARMDRISGLPDKILHHIMSFLNTRQAVQTCVLSRRWRNLWRTMPCIN 133
Query: 61 IDFNNIKIDSIESNSK-------FIDSVYSVLVSPD-TAIGGSHFINRFCLDVQFRNGNP 112
DF+ + + + + F V +L D TA+ + +++ D
Sbjct: 134 ADFDEFDFIAYQGDDEDYNDEVAFKRFVNQMLELRDPTAMIDTFWLSYIIWD-------- 185
Query: 113 LYGYDNPHLLYKRSCHNVVKWVNLVVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPI 172
GY+ YK S + +W++ +Q++ + + + + + L LD + + YL K+
Sbjct: 186 --GYNE----YKDSNVDANRWISHALQKQARVIEV-VVFAFPLELDHSVFTSCYLRKIG- 237
Query: 173 TIFTCRTL 180
F+C +L
Sbjct: 238 --FSCVSL 243
Score = 58.5 bits (140), Expect = 2e-07
Identities = 49/204 (24%), Positives = 92/204 (45%), Gaps = 37/204 (18%)
Query: 6 RRRRSSKASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNN 65
RR+ + A A D IS LPD++LH I+S ++ ++AV T +LS+RW NLW ++P I+ +
Sbjct: 536 RRKVKTPAHAAGMDWISDLPDEILHRIMSFLNARQAVQTCVLSRRWRNLWRTVPCINADC 595
Query: 66 IKIDSIESNSKFIDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKR 125
+ D ++ F+NR L+++ +P+ D Y +
Sbjct: 596 KEFDFFGFRRSEVEF--------------KRFVNRL-LELR----DPIAMMDAFWFRYHK 636
Query: 126 ------SCHNVVKWVNLVVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFT--- 176
S + +W++ +Q++ + L + + L LD + + YL ++ +
Sbjct: 637 LDTDTTSSADTNRWISHALQKQARVLEAVMYPCHLLDLDHSSFTSRYLRRIGFSGVRLDQ 696
Query: 177 ---------CRTLVSLDLHHFSVK 191
C L L LHH +++
Sbjct: 697 GFFKQLEAGCPALEDLFLHHCTIE 720
>dbj|BAB11265.1| unnamed protein product [Arabidopsis thaliana]
gi|15241200|ref|NP_200448.1| F-box family protein
[Arabidopsis thaliana] gi|42573692|ref|NP_974942.1|
F-box family protein [Arabidopsis thaliana]
gi|46518439|gb|AAS99701.1| At5g56370 [Arabidopsis
thaliana] gi|51970526|dbj|BAD43955.1| unknown protein
[Arabidopsis thaliana]
Length = 421
Score = 68.2 bits (165), Expect = 2e-10
Identities = 57/188 (30%), Positives = 85/188 (44%), Gaps = 38/188 (20%)
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKFI 78
D IS LPDD L ILSL+ TK+ + TS+LSKRW LW +P + ++ ID + F+
Sbjct: 2 DSISLLPDDFLLRILSLLPTKDVLNTSVLSKRWRYLWKLVPKLQYS--LIDKNADHGTFV 59
Query: 79 DSV-YSVLVSPDTAIGGSHF-INRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWVNL 136
V S+L+S + H + R C +V ++ WV +
Sbjct: 60 RFVDRSLLLSMAPVLESLHLKLGRQCSEV-----------------------DIGFWVRI 96
Query: 137 VVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRTLVSLDLHHFSVKGVCYR 196
V++ L +L D + Y +LP ++FTC TL L L + S+K V +
Sbjct: 97 AVEKGL----------CELDFDYEHYKTEPC-RLPQSLFTCGTLTVLKLKNVSLKDVQFP 145
Query: 197 FLLKALST 204
K L T
Sbjct: 146 VCFKLLKT 153
>gb|AAF79496.1| F20N2.9 [Arabidopsis thaliana]
Length = 526
Score = 67.8 bits (164), Expect = 3e-10
Identities = 59/190 (31%), Positives = 94/190 (49%), Gaps = 36/190 (18%)
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFN--NIKIDSIESNSK 76
DKIS+LPD++L +LS +STK+AV+TSILS RW +LW+ LP +++N + + + ++
Sbjct: 87 DKISQLPDELLVKVLSFLSTKDAVSTSILSMRWKSLWMWLPKLEYNFRHYSVSEGQGLAR 146
Query: 77 FIDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWVNL 136
FI S V +P L ++FR G G P +Y WV+L
Sbjct: 147 FITSSLRVHKAPAIE----------SLSLKFRYG--AIGSIKPKDIY--------LWVSL 186
Query: 137 VVQ-RRLKYLRLNLRLGYDLHLDVDDYDNSYLP-KLPITIFTCRTLVSLDLHHFSVKGVC 194
V ++ L L L ++ + LP KLP ++ C+++V L L + V
Sbjct: 187 AVHVSNVRELSLKL------------FNFAELPTKLPKSLCKCKSIVILKLKDEILVDVP 234
Query: 195 YRFLLKALST 204
+ L +L T
Sbjct: 235 RKVCLPSLKT 244
>ref|NP_176127.1| F-box family protein [Arabidopsis thaliana]
Length = 505
Score = 67.8 bits (164), Expect = 3e-10
Identities = 51/171 (29%), Positives = 90/171 (51%), Gaps = 21/171 (12%)
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDF-NNIKIDSIESNSKF 77
D IS LPD +L +ILS ++TKEA +TS+L+K+W L+ S+PN+DF +++ + + N
Sbjct: 8 DIISGLPDSLLCHILSFLNTKEAASTSVLAKKWRYLFASVPNLDFDDSVHLRLGKRNPAV 67
Query: 78 IDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVK---WV 134
Y +++ + + F++ ++ ++ +PL+ + L R C ++V+ W+
Sbjct: 68 SGEDYLKMINERSDQLSTSFMDFVDQVLRLQDNSPLHKFS----LKIRDCVDIVRIICWI 123
Query: 135 NLVVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRTLVSLDL 185
V++R + L L D+HL + LP IF TLV L L
Sbjct: 124 LKVLERGVSDLEL------DMHL-------KWKSSLPSKIFLSETLVRLKL 161
>ref|NP_175962.1| F-box family protein [Arabidopsis thaliana]
Length = 492
Score = 67.8 bits (164), Expect = 3e-10
Identities = 59/190 (31%), Positives = 94/190 (49%), Gaps = 36/190 (18%)
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFN--NIKIDSIESNSK 76
DKIS+LPD++L +LS +STK+AV+TSILS RW +LW+ LP +++N + + + ++
Sbjct: 53 DKISQLPDELLVKVLSFLSTKDAVSTSILSMRWKSLWMWLPKLEYNFRHYSVSEGQGLAR 112
Query: 77 FIDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWVNL 136
FI S V +P L ++FR G G P +Y WV+L
Sbjct: 113 FITSSLRVHKAPAIE----------SLSLKFRYG--AIGSIKPKDIY--------LWVSL 152
Query: 137 VVQ-RRLKYLRLNLRLGYDLHLDVDDYDNSYLP-KLPITIFTCRTLVSLDLHHFSVKGVC 194
V ++ L L L ++ + LP KLP ++ C+++V L L + V
Sbjct: 153 AVHVSNVRELSLKL------------FNFAELPTKLPKSLCKCKSIVILKLKDEILVDVP 200
Query: 195 YRFLLKALST 204
+ L +L T
Sbjct: 201 RKVCLPSLKT 210
>ref|NP_197662.2| F-box family protein [Arabidopsis thaliana]
Length = 437
Score = 67.8 bits (164), Expect = 3e-10
Identities = 51/168 (30%), Positives = 80/168 (47%), Gaps = 35/168 (20%)
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKFI 78
D+IS LPD++L ILS + TK AV TSILS RW ++WLS P +D I ID+ + + F+
Sbjct: 6 DRISSLPDELLCQILSNLPTKNAVTTSILSTRWRSIWLSTPVLD---IDIDAFDDATTFV 62
Query: 79 DSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWVNLVV 138
F +RF ++F + L+ + ++ W+ V
Sbjct: 63 S-----------------FASRF---LEFSKDSCLHKFKLSVERDDVDMCTIMPWIQDAV 102
Query: 139 QRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRTLVSLDLH 186
RR++ HL+VD + + + +T++ TLVSL LH
Sbjct: 103 NRRIQ------------HLEVDCRFSFHFEAVYLTLYLSETLVSLRLH 138
>dbj|BAB10659.1| unnamed protein product [Arabidopsis thaliana]
gi|15238204|ref|NP_198999.1| F-box family protein
[Arabidopsis thaliana]
Length = 540
Score = 67.0 bits (162), Expect = 4e-10
Identities = 67/241 (27%), Positives = 110/241 (44%), Gaps = 56/241 (23%)
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFN-----------NIK 67
D+IS LPD ++ +ILS + TKEA +T++L+KRW L +PN++F+ N
Sbjct: 14 DRISGLPDALICHILSFLPTKEAASTTVLAKRWKPLLAFVPNLNFDDSIYFHPRARRNKY 73
Query: 68 IDSIESNSKFIDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSC 127
S ES F+DSV ++ T + H C DV ++
Sbjct: 74 SKSYESFMSFVDSVLALQAKTKTPLKRFHV---KCEDVVDQSW----------------- 113
Query: 128 HNVVKWVNLVVQRRLKYLRLNLRLGYDLHLDVD-DY-DNSYLPKLPITIFTCRTLVSLDL 185
V++W+ V++R + L DLH+ +Y +NS LP IF +TLV L +
Sbjct: 114 --VLEWIPKVLKRGV--------LDIDLHITSSRNYCENSSFYSLPSKIFVSKTLVRLKI 163
Query: 186 H-----HFSVKGVCYRFLLKALSTSNSLRLDTFKLYRSIYQVRFIMNTMHVIYFLFLSEI 240
H V+G +L +L LD FK+ S+ + +++ H + L L+ +
Sbjct: 164 QFQDGVHIDVEGGV------SLPKLKTLHLDYFKIETSM--LNKLLSGCHALEELVLANL 215
Query: 241 L 241
+
Sbjct: 216 M 216
>dbj|BAD95126.1| hypothetical protein [Arabidopsis thaliana]
Length = 456
Score = 66.6 bits (161), Expect = 6e-10
Identities = 51/184 (27%), Positives = 89/184 (47%), Gaps = 31/184 (16%)
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKFI 78
D+IS LP+ ++ IL + TK +V TS+LS RW NLWL++P +D N +N K +
Sbjct: 10 DRISELPESLISQILLHLPTKASVKTSVLSTRWKNLWLNVPGLDLNCRDFPFQNNNEKLL 69
Query: 79 DSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWVNLV- 137
FI+RF +QF N + L + + S ++K+ + +
Sbjct: 70 ----------------IDFIDRF---LQFNNESRLQKFKVDY-----SRDKIIKFSDRIG 105
Query: 138 --VQRRLKYLRLNLRLGYDLHLDVDD-YDNSYLPKLPITIFTCRTLVSLDLHHFSVKGVC 194
+ R ++ L + Y D DD D + +P+ +++C+TLVSL L + ++
Sbjct: 106 DAISRGIRVLDVESNTYY---RDADDCIDYPCIEFMPLNLYSCKTLVSLKLSYSGLEDPG 162
Query: 195 YRFL 198
+ +L
Sbjct: 163 FVYL 166
>gb|AAF79391.1| F16A14.1 [Arabidopsis thaliana] gi|15222948|ref|NP_172833.1| F-box
family protein [Arabidopsis thaliana]
gi|46518457|gb|AAS99710.1| At1g13780 [Arabidopsis
thaliana]
Length = 451
Score = 66.6 bits (161), Expect = 6e-10
Identities = 51/184 (27%), Positives = 89/184 (47%), Gaps = 31/184 (16%)
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKFI 78
D+IS LP+ ++ IL + TK +V TS+LS RW NLWL++P +D N +N K +
Sbjct: 5 DRISELPESLISQILLHLPTKASVKTSVLSTRWKNLWLNVPGLDLNCRDFPFQNNNEKLL 64
Query: 79 DSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWVNLV- 137
FI+RF +QF N + L + + S ++K+ + +
Sbjct: 65 ----------------IDFIDRF---LQFNNESRLQKFKVDY-----SRDKIIKFSDRIG 100
Query: 138 --VQRRLKYLRLNLRLGYDLHLDVDD-YDNSYLPKLPITIFTCRTLVSLDLHHFSVKGVC 194
+ R ++ L + Y D DD D + +P+ +++C+TLVSL L + ++
Sbjct: 101 DAISRGIRVLDVESNTYY---RDADDCIDYPCIEFMPLNLYSCKTLVSLKLSYSGLEDPG 157
Query: 195 YRFL 198
+ +L
Sbjct: 158 FVYL 161
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.326 0.141 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 389,893,204
Number of Sequences: 2540612
Number of extensions: 15982942
Number of successful extensions: 47611
Number of sequences better than 10.0: 484
Number of HSP's better than 10.0 without gapping: 442
Number of HSP's successfully gapped in prelim test: 42
Number of HSP's that attempted gapping in prelim test: 47001
Number of HSP's gapped (non-prelim): 555
length of query: 242
length of database: 863,360,394
effective HSP length: 124
effective length of query: 118
effective length of database: 548,324,506
effective search space: 64702291708
effective search space used: 64702291708
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 73 (32.7 bits)
Medicago: description of AC135799.3


