

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135796.7 - phase: 0
(277 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
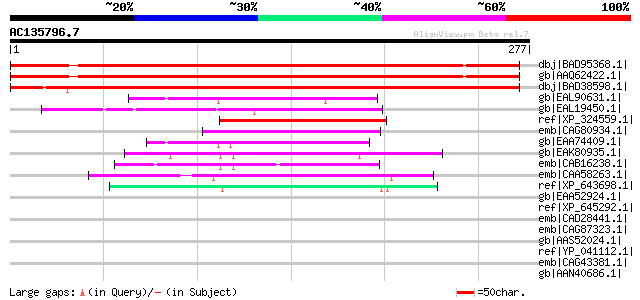
Sequences producing significant alignments: (bits) Value
dbj|BAD95368.1| hypothetical protein [Arabidopsis thaliana] gi|1... 305 9e-82
gb|AAQ62422.1| At1g53760 [Arabidopsis thaliana] 303 4e-81
dbj|BAD38598.1| unknown protein [Oryza sativa (japonica cultivar... 269 7e-71
gb|EAL90631.1| membrane protein, putative [Aspergillus fumigatus... 74 5e-12
gb|EAL19450.1| hypothetical protein CNBG3970 [Cryptococcus neofo... 62 1e-08
ref|XP_324559.1| hypothetical protein [Neurospora crassa] gi|289... 62 2e-08
emb|CAG80934.1| unnamed protein product [Yarrowia lipolytica CLI... 57 6e-07
gb|EAA74409.1| hypothetical protein FG05070.1 [Gibberella zeae P... 57 7e-07
gb|EAK80935.1| hypothetical protein UM00391.1 [Ustilago maydis 5... 56 1e-06
emb|CAB16238.1| SPAC23H3.12c [Schizosaccharomyces pombe] gi|1911... 54 6e-06
emb|CAA58263.1| ORF D1293 [Saccharomyces cerevisiae] gi|51012609... 49 1e-04
ref|XP_643698.1| hypothetical protein DDB0202593 [Dictyostelium ... 49 1e-04
gb|EAA52924.1| hypothetical protein MG06052.4 [Magnaporthe grise... 44 0.004
ref|XP_645292.1| hypothetical protein DDB0168800 [Dictyostelium ... 43 0.011
emb|CAD28441.1| hypothetical protein [Aspergillus fumigatus] 42 0.019
emb|CAG87323.1| unnamed protein product [Debaryomyces hansenii C... 41 0.032
gb|AAS52024.1| ADR104Cp [Ashbya gossypii ATCC 10895] gi|45187977... 40 0.072
ref|YP_041112.1| Spo0B-associated GTP-binding protein [Staphyloc... 35 1.8
emb|CAG43381.1| Spo0B-associated GTP-binding protein [Staphyloco... 35 1.8
gb|AAN40686.1| translationally controlled tumor protein-like pro... 35 2.3
>dbj|BAD95368.1| hypothetical protein [Arabidopsis thaliana]
gi|15220931|ref|NP_175779.1| expressed protein
[Arabidopsis thaliana] gi|12324017|gb|AAG51966.1|
hypothetical protein; 1267-3026 [Arabidopsis thaliana]
Length = 272
Score = 305 bits (781), Expect = 9e-82
Identities = 151/273 (55%), Positives = 197/273 (71%), Gaps = 6/273 (2%)
Query: 1 MKAKLIVFPIRGRNWCFTRSIDHTLPASSSTADFSQSPSTLKQLWKSINTGDKPFNAKAE 60
M+A+L+VFPI+G+ WCF+RS+D S S +P+T++ LWK I++ KP NA AE
Sbjct: 1 MRARLVVFPIKGKKWCFSRSVDPFAAQSPSGV----TPTTVRGLWKKISSESKPINANAE 56
Query: 61 LFTDYVANKMNNGWVTLENAPDGSFKKKIHGLGLWLLSRVKPSEIFLKSISKDVTGVEVV 120
L D++++KMN WV LE APDGS K KIHG GL LL+RVKPSEIFLKSISK+VT V+V
Sbjct: 57 LLVDFISDKMNKAWVGLEKAPDGSIKNKIHGFGLKLLARVKPSEIFLKSISKEVTSVQVT 116
Query: 121 YPSSMNARLVRRRLRHIAMRGTIIHRKFFYGSVSLIPLSSAFSILPLPNVPFFWILFRSY 180
YP S++ RLVRRRLRHIAM GTI+H+K+ GSV+L+PL+SAF +LPLPN+PFFW+LFR+Y
Sbjct: 117 YPPSLDPRLVRRRLRHIAMSGTILHKKYLVGSVTLLPLTSAFMVLPLPNIPFFWVLFRTY 176
Query: 181 SHWRALQGSEKLFQLVSDGSQSSNTYSGKKETEHEDSENE-SLGLDEPHWVLTPSKELEN 239
SHWRALQGSEKL +L+S+ + S E ++S + P +L PS+EL
Sbjct: 177 SHWRALQGSEKLLKLISNEANPDKPDSTDDADESKNSNTKPEQKSQSPTCILLPSEELYQ 236
Query: 240 IVRQEDGNDGLSRGTIEEICKIYDLNTQDVVKY 272
++R E +GL TI EICK +DLN DV+KY
Sbjct: 237 LIR-EASEEGLDEATIIEICKSFDLNKNDVLKY 268
>gb|AAQ62422.1| At1g53760 [Arabidopsis thaliana]
Length = 272
Score = 303 bits (776), Expect = 4e-81
Identities = 150/273 (54%), Positives = 196/273 (70%), Gaps = 6/273 (2%)
Query: 1 MKAKLIVFPIRGRNWCFTRSIDHTLPASSSTADFSQSPSTLKQLWKSINTGDKPFNAKAE 60
M+A+L+VFPI+G+ WCF+RS+D S S +P+T++ LWK I++ KP NA AE
Sbjct: 1 MRARLVVFPIKGKKWCFSRSVDPFAAQSPSGV----TPTTVRGLWKKISSESKPINANAE 56
Query: 61 LFTDYVANKMNNGWVTLENAPDGSFKKKIHGLGLWLLSRVKPSEIFLKSISKDVTGVEVV 120
L D++++KMN WV LE APDGS K KIHG GL LL+RVKPSEIFLKSISK+VT V+V
Sbjct: 57 LLVDFISDKMNKAWVGLEKAPDGSIKNKIHGFGLKLLARVKPSEIFLKSISKEVTSVQVT 116
Query: 121 YPSSMNARLVRRRLRHIAMRGTIIHRKFFYGSVSLIPLSSAFSILPLPNVPFFWILFRSY 180
YP S++ RLVRRRLRHIAM GTI+H+K+ GSV+L+PL+SAF +LPLPN+PFFW+LFR+Y
Sbjct: 117 YPPSLDPRLVRRRLRHIAMSGTILHKKYLVGSVTLLPLTSAFMVLPLPNIPFFWVLFRTY 176
Query: 181 SHWRALQGSEKLFQLVSDGSQSSNTYSGKKETEHEDSENE-SLGLDEPHWVLTPSKELEN 239
HWRALQGSEKL +L+S+ + S E ++S + P +L PS+EL
Sbjct: 177 PHWRALQGSEKLLKLISNEANPDKPDSTDDADESKNSNTKPEQKSQSPTCILLPSEELYQ 236
Query: 240 IVRQEDGNDGLSRGTIEEICKIYDLNTQDVVKY 272
++R E +GL TI EICK +DLN DV+KY
Sbjct: 237 LIR-EASEEGLDEATIIEICKSFDLNKNDVLKY 268
>dbj|BAD38598.1| unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 281
Score = 269 bits (687), Expect = 7e-71
Identities = 137/278 (49%), Positives = 183/278 (65%), Gaps = 7/278 (2%)
Query: 1 MKAKLIVFPIRGRNWCFTRSIDHTLPASS-----STADFSQSPSTLKQLWKSINTGDKPF 55
M+A+++VFP++GR WCF S T PA++ A P T+K LW+ I G +
Sbjct: 1 MQARVVVFPVKGRAWCFA-SPRATAPAAACGGGDGGALLPPPPPTVKDLWRGIAGGGRTA 59
Query: 56 NAKAELFTDYVANKMNNGWVTLENAPDGSFKKKIHGLGLWLLSRVKPSEIFLKSISKDVT 115
+ AE D+VA+KMN W+ +AP+GS K +IH GL LLSRV+PSE+ LKS++KDV+
Sbjct: 60 SENAEAVVDFVADKMNRAWIGFGSAPEGSMKNRIHSFGLKLLSRVRPSEVLLKSVTKDVS 119
Query: 116 GVEVVYPSSMNARLVRRRLRHIAMRGTIIHRKFFYGSVSLIPLSSAFSILPLPNVPFFWI 175
+E+V+P+S+N RLVRRRLRHIA+RG +HRKF YGSV L+P++S F +LPLPN+PFFW+
Sbjct: 120 LLEIVHPASINPRLVRRRLRHIAVRGASVHRKFLYGSVCLLPVTSVFMVLPLPNIPFFWV 179
Query: 176 LFRSYSHWRALQGSEKLFQLVSDGS-QSSNTYSGKKETEHEDSENESLGLDEPHWVLTPS 234
LFR+YSHWRALQGSE+L LVSD S Q +KE N W L PS
Sbjct: 180 LFRAYSHWRALQGSERLQLLVSDSSDQWKILLEKQKEMSSRKDGNPCENTQYAPWNLQPS 239
Query: 235 KELENIVRQEDGNDGLSRGTIEEICKIYDLNTQDVVKY 272
K+L+ + N+GL TI IC+ YDL+ DV+KY
Sbjct: 240 KKLDGFLESRKLNEGLDCDTISRICQAYDLDKIDVLKY 277
>gb|EAL90631.1| membrane protein, putative [Aspergillus fumigatus Af293]
Length = 284
Score = 73.9 bits (180), Expect = 5e-12
Identities = 51/153 (33%), Positives = 78/153 (50%), Gaps = 21/153 (13%)
Query: 64 DYVANKMNNGWVTLENAPDGSFKKKIHGLGLWLLSRVKPSEIFLKSI-----------SK 112
D V NK W E A G +KK + G + R+ E LKSI S
Sbjct: 32 DRVTNKAAETWAKWEEADKG-WKKHLVTWGNKVQQRIPFEEWGLKSIPSLKAQRRLDKSS 90
Query: 113 DVTGVEVVYP-SSMNARLVRRRLRHIAMRGTIIHRKFFYGSVSLIPLSSAFSILPL---- 167
+ V+V++P +++ A +R LR IA +HRK + S+ PL++ +++P+
Sbjct: 91 ETKKVDVLFPGNAIKAEKIRSILRKIATERQDLHRKKMWWSLVAAPLTAPIALIPVYSLC 150
Query: 168 ----PNVPFFWILFRSYSHWRALQGSEKLFQLV 196
PN+PFF++ +R +SHWRAL GS+ L LV
Sbjct: 151 LTDVPNIPFFYLAYRGWSHWRALNGSKHLEYLV 183
>gb|EAL19450.1| hypothetical protein CNBG3970 [Cryptococcus neoformans var.
neoformans B-3501A] gi|57228046|gb|AAW44503.1|
hypothetical protein CNG00810 [Cryptococcus neoformans
var. neoformans JEC21] gi|58269308|ref|XP_571810.1|
hypothetical protein CNG00810 [Cryptococcus neoformans
var. neoformans JEC21]
Length = 354
Score = 62.4 bits (150), Expect = 1e-08
Identities = 51/200 (25%), Positives = 98/200 (48%), Gaps = 21/200 (10%)
Query: 18 TRSIDHTLPASSSTADFSQS-PSTLKQLWKSINTGDKPFNAKAELFTDYVANKMNNGWVT 76
+R++ T P S +A S + P T L+ ++ + ++ L T + +K + W+
Sbjct: 32 SRTLPTTPPPSLESASSSHTTPPTPLILFHALQP-EPSADSPPSLITKAL-DKSSETWLK 89
Query: 77 LENAPDGSFKKKIHGLGLWLLSRVKPSEIFLKSISKDVTGVEVVYPSSMNARL------- 129
L P GS+ + G L+ R++ E LK++ ++ GV+V + R+
Sbjct: 90 LGEKPKGSWMYWFYEKGEKLMDRIEYEEWALKAV-REGEGVKVAKDGQVLERIEIPLVLP 148
Query: 130 ---------VRRRLRHIAMRGTIIHRKFFYGSVSLIPLSSAFSILPL-PNVPFFWILFRS 179
+ +L ++ HRK Y S+ P++ F+I+P+ PN P F++L+R+
Sbjct: 149 RLPNQEMPSLLPKLHRFLLKRIPHHRKMMYRSILFSPVTWPFAIIPIIPNFPLFYVLWRA 208
Query: 180 YSHWRALQGSEKLFQLVSDG 199
+SH++A +G+ L QL+ G
Sbjct: 209 WSHYKAWRGATYLEQLIKAG 228
>ref|XP_324559.1| hypothetical protein [Neurospora crassa] gi|28923800|gb|EAA32965.1|
hypothetical protein [Neurospora crassa]
Length = 360
Score = 62.0 bits (149), Expect = 2e-08
Identities = 35/91 (38%), Positives = 56/91 (61%), Gaps = 2/91 (2%)
Query: 113 DVTGVEVVYPSSM-NARLVRRRLRHIAMRGTIIHRKFFYGSVSLIPLSSAFSILPL-PNV 170
DV VEVVYP + A V R L +A H+K+F+ ++ +P++ ILPL PN+
Sbjct: 125 DVDKVEVVYPGGLIKAEQVPRILHKLATEREGHHKKWFWWCLAGMPVTIPIGILPLVPNL 184
Query: 171 PFFWILFRSYSHWRALQGSEKLFQLVSDGSQ 201
PFF++++R++SH RAL G + + L+ S+
Sbjct: 185 PFFYLVYRAWSHHRALSGGKHISFLLQHASR 215
>emb|CAG80934.1| unnamed protein product [Yarrowia lipolytica CLIB99]
gi|50550547|ref|XP_502746.1| hypothetical protein
[Yarrowia lipolytica]
Length = 310
Score = 57.0 bits (136), Expect = 6e-07
Identities = 31/97 (31%), Positives = 56/97 (56%), Gaps = 2/97 (2%)
Query: 104 EIFLKSISKDVTGVEVVYPSSMNAR-LVRRRLRHIAMRGTIIHRKFFYGSVSLIPLSSAF 162
E K +++ V +V+P+ ++ + +L +A+ G HRK+ PL+
Sbjct: 119 EDIAKKHHEELLKVPLVFPNHLSTPDQAKAQLLKLAIDGAKHHRKYMIWCAIGAPLTLPV 178
Query: 163 SILP-LPNVPFFWILFRSYSHWRALQGSEKLFQLVSD 198
++LP LPN+P F+++FR +SHW+AL+G+ L L+ D
Sbjct: 179 ALLPVLPNLPGFYLVFRCWSHWKALEGANHLKHLIED 215
>gb|EAA74409.1| hypothetical protein FG05070.1 [Gibberella zeae PH-1]
gi|46121383|ref|XP_385246.1| hypothetical protein
FG05070.1 [Gibberella zeae PH-1]
Length = 281
Score = 56.6 bits (135), Expect = 7e-07
Identities = 36/132 (27%), Positives = 70/132 (52%), Gaps = 14/132 (10%)
Query: 74 WVTLENAPDGSFKKKIHGLGLWLLSRVKPSEIFLKSI--------SKDVTG---VEVVYP 122
W E G +K+K+ G + R+ E LKS+ +D+ G VE+ +P
Sbjct: 43 WAQWEKMESG-WKRKVVDYGNYAFRRIPYEEWGLKSVPPLSSRRRGEDIQGRGKVELCFP 101
Query: 123 SSM-NARLVRRRLRHIAMRGTIIHRKFFYGSVSLIPLSSAFSILPL-PNVPFFWILFRSY 180
SS+ L+ +A +H+K + +P++ F+++P+ PN+PFF++++R++
Sbjct: 102 SSVIPTNKAEGILQTLATERQALHKKRLVWCIVGMPITIPFALIPIIPNLPFFYLVYRAW 161
Query: 181 SHWRALQGSEKL 192
SH+RA+ G + +
Sbjct: 162 SHYRAIAGGKHI 173
>gb|EAK80935.1| hypothetical protein UM00391.1 [Ustilago maydis 521]
gi|49067432|ref|XP_398006.1| hypothetical protein
UM00391.1 [Ustilago maydis 521]
Length = 313
Score = 55.8 bits (133), Expect = 1e-06
Identities = 49/208 (23%), Positives = 93/208 (44%), Gaps = 38/208 (18%)
Query: 62 FTDYVANKMNNGWVTLENAPDGS---FKKKIHGLGLWLLSRVKPSEIFLKSIS------- 111
FT + NK + W+ L S +K++ + LG L+ R++ E LK I
Sbjct: 52 FTTRMLNKASAFWIDLGRTDQTSTFDWKRRTYVLGERLMDRIEYQEWALKGIDPAMGPHL 111
Query: 112 -----KDVTGVE-------------VVYPSSM-NARLVRRRLRHIAMRGTIIHRKFFYGS 152
KD E ++YP S+ + + L+++ T H + F+
Sbjct: 112 IRRERKDADSPEATNGGIPKLDHVPLLYPPSLLEPERLLKSLKNLTDHRTPHHYRRFWYC 171
Query: 153 VSLIPLSSAFSILPL-PNVPFFWILFRSYSHWRA--------LQGSEKLFQLVSDGSQSS 203
V +P++ F+++P+ PN+PFF++++R++SHW+ + + +F S +S
Sbjct: 172 VVGMPITIPFALVPVVPNLPFFYLVYRAWSHWKGQDRVRPTESKDLDAIFAQASPPVDAS 231
Query: 204 NTYSGKKETEHEDSENESLGLDEPHWVL 231
+ + KETE + + S H +L
Sbjct: 232 SPEASNKETETTATTSTSNPAQNEHMLL 259
>emb|CAB16238.1| SPAC23H3.12c [Schizosaccharomyces pombe]
gi|19114714|ref|NP_593802.1| hypothetical protein
SPAC23H3.12c [Schizosaccharomyces pombe 972h-]
gi|7490731|pir||T38305 hypothetical protein - fission
yeast (Schizosaccharomyces pombe)
gi|50401298|sp|O13942|YEPC_SCHPO Hypothetical protein
C23H3.12c in chromosome I
Length = 226
Score = 53.5 bits (127), Expect = 6e-06
Identities = 41/151 (27%), Positives = 80/151 (52%), Gaps = 13/151 (8%)
Query: 57 AKAELFTDYVANKMNNGWVTLENAPDGSFKKKIHGLGLWLLSRVKPSEIFLKSIS--KDV 114
AK D + N++ W + +A K+K+ LG +L E FL++I+ K +
Sbjct: 24 AKKVTIHDKIINRIYKYWDSW-SASKSYTKQKVVSLGNRILHATPYEENFLRAIAPVKKL 82
Query: 115 TGVE------VVYPSSMNARLVRRRL-RHIAMRGTIIHRKFFYGSVSLIPLSSAFSILPL 167
E + +P ++ + + L R ++ T H + G++ +PL+ F ++PL
Sbjct: 83 NDTELHQTLYIEHPPNLESSTILAELNRSKQLQKT--HTNYLIGNIIGLPLTIPFILIPL 140
Query: 168 -PNVPFFWILFRSYSHWRALQGSEKLFQLVS 197
PN+P F++ +R+Y ++RA+QGS +L +++S
Sbjct: 141 IPNIPGFYLCYRAYCNFRAIQGSIQLARVMS 171
>emb|CAA58263.1| ORF D1293 [Saccharomyces cerevisiae] gi|51012609|gb|AAT92598.1|
YDL183C [Saccharomyces cerevisiae]
gi|6320018|ref|NP_010098.1| Hypothetical ORF; Ydl183cp
[Saccharomyces cerevisiae] gi|1431297|emb|CAA98758.1|
unnamed protein product [Saccharomyces cerevisiae]
gi|1351811|sp|P48569|YDB3_YEAST Hypothetical 37.0 kDa
protein in RPL41A-INH1 intergenic region
Length = 320
Score = 49.3 bits (116), Expect = 1e-04
Identities = 47/203 (23%), Positives = 87/203 (42%), Gaps = 25/203 (12%)
Query: 43 QLWKSINTGDKPFNAKAELFTDYVANKMNNGWVTLENAPDGSF-KKKIHGLGLWLLSRVK 101
+LW+ + K +N K + N +L P S+ K+I G + K
Sbjct: 84 KLWRKLKKSPKSYNKKIVSMVQSLLNSTPWSENSLLTIPSESYILKRIKG------EKDK 137
Query: 102 PSEIFL-------KSISKDVTGVEVVYPSSMNAR-LVRRRLRHIAMRGTIIHRKFFYGSV 153
EI L K+ D + V YP +++ R+++ + G I H+K+ +
Sbjct: 138 TQEIRLTLKDYTVKAEQVDTQPLHVYYPPGISSPDECLRQMKKLYQEGLIYHKKWTLYCL 197
Query: 154 SLIPLSSAFSILPL-PNVPFFWILFRSYSHWRALQGSEKLFQLVSDGSQS---------S 203
+PL+ ++PL PNVP F++ +R+Y + +A G++ L L+ Q+ +
Sbjct: 198 LGLPLTIPLILIPLIPNVPGFYLSYRAYVNIKAYLGAKHLKSLLESSKQNLEFRELLGYT 257
Query: 204 NTYSGKKETEHEDSENESLGLDE 226
Y + + ++ ES G E
Sbjct: 258 EVYKRGTSSRTQGNQKESKGAPE 280
>ref|XP_643698.1| hypothetical protein DDB0202593 [Dictyostelium discoideum]
gi|60471799|gb|EAL69754.1| hypothetical protein
DDB0202593 [Dictyostelium discoideum]
Length = 312
Score = 49.3 bits (116), Expect = 1e-04
Identities = 50/239 (20%), Positives = 87/239 (35%), Gaps = 64/239 (26%)
Query: 54 PFNAKAELFTDYVANKMNNGWVTLENAPDG-SFKKKIHGLGLWLLSRVKPSEIFLKSISK 112
P +K L + + N ++NA DG K K+ L +L S++ P E L+ SK
Sbjct: 47 PTQSKGVLILNKLKNIAIESKNEIDNAQDGVGLKGKLKSLLSYLESKIHPEEAVLQQFSK 106
Query: 113 --------------------------------------------DVTGVEVVYPSSMNAR 128
D + YPS++ R
Sbjct: 107 INDNNLIKKHNHHNHHNHIHHNNNNNNNQNETVKHQFEHFNEQRDKASFTIYYPSNIGYR 166
Query: 129 LVRRRLRHIAMRGTIIHRKFFYGSVSLIPLSSAFSILPLPNVPFFWILFRSYSHWRALQG 188
R + + + H+K+ + LIP++ A +ILP PN+ + L+R Y H++AL+G
Sbjct: 167 RANRFSQVLMRQKIRFHQKWMIINSLLIPITFAATILPGPNIFLAYNLYRLYGHFQALKG 226
Query: 189 SEKLFQLVS-----------------DGS--QSSNTYSGKKETEHEDSENESLGLDEPH 228
L + DG Q + +++ + + E E +D P+
Sbjct: 227 CTNLSRYTKSHGRLLFKPSLEMFECLDGELLQQQQQFKQQQQQQQQQQEEEEQSIDNPN 285
>gb|EAA52924.1| hypothetical protein MG06052.4 [Magnaporthe grisea 70-15]
gi|39976043|ref|XP_369412.1| hypothetical protein
MG06052.4 [Magnaporthe grisea 70-15]
Length = 276
Score = 44.3 bits (103), Expect = 0.004
Identities = 41/151 (27%), Positives = 63/151 (41%), Gaps = 24/151 (15%)
Query: 54 PFNAKAELFTDYVANKMNNGWVTLENAPDGSFKKKIHGLGLWLLSRVKPSEIFLKSISKD 113
P K + ++ V +K W E P G ++K + G G L R+ E LKS+
Sbjct: 21 PAIPKKQTISERVQSKAARTWADWEKKP-GGWQKLVVGYGNQALRRIPYEEWGLKSVPPL 79
Query: 114 VTG-----------VEVVYPSSM-NARLVRRRLRHIAMRGTIIHRKFFYGSVSLIPLSSA 161
T VEVVYP S+ A V L ++ +H+K +PL+
Sbjct: 80 STARKEDEQAGKVKVEVVYPKSLVKAERVTSILHTLSTGRESLHKKRLVLCFLGMPLTIP 139
Query: 162 FSILPLPNVPFFWILFRSYSHWRALQGSEKL 192
+I+P+ ++SHWRAL G +
Sbjct: 140 VAIIPI-----------AWSHWRALAGGRHI 159
>ref|XP_645292.1| hypothetical protein DDB0168800 [Dictyostelium discoideum]
gi|42761552|gb|AAS45370.1| similar to Dictyostelium
discoideum (Slime mold). R1062 protein (Fragment)
gi|60473316|gb|EAL71262.1| hypothetical protein
DDB0168800 [Dictyostelium discoideum]
Length = 871
Score = 42.7 bits (99), Expect = 0.011
Identities = 37/130 (28%), Positives = 56/130 (42%), Gaps = 15/130 (11%)
Query: 118 EVVYPSSMNARLVRRRLRHIAMRGTIIHRKFFYGSVSLIPLSSAFSILPLPNVPFFWILF 177
+++YP S ++ + H+ M GT F+YGS+ LIP F ILF
Sbjct: 707 QLLYPMSFRRTMIALLIYHLLMIGTFNVYSFYYGSLILIPF-------------FLTILF 753
Query: 178 RSYSHWRALQGSEKLFQLVSDGSQSSNTYSGKKETEHEDSENESLGLDEPHWVLTPSKEL 237
Y + S+ S+SSN S + ++ N+S L+E +L SKE
Sbjct: 754 WVYCEYTFYSVSKNGLLDHYQSSKSSNQKSQLNSINNNNNNNKS-NLNEKSSLL-QSKER 811
Query: 238 ENIVRQEDGN 247
N + ED N
Sbjct: 812 NNYIDIEDSN 821
>emb|CAD28441.1| hypothetical protein [Aspergillus fumigatus]
Length = 154
Score = 42.0 bits (97), Expect = 0.019
Identities = 34/116 (29%), Positives = 55/116 (47%), Gaps = 13/116 (11%)
Query: 64 DYVANKMNNGWVTLENAPDGSFKKKIHGLGLWLLSRVKPSEIFLKSI-----------SK 112
D V NK W E A G +KK + G + R+ E LKSI S
Sbjct: 32 DRVTNKAAETWAKWEEADKG-WKKHLVTWGNKVQQRIPFEEWGLKSIPSLKAQRRLDKSS 90
Query: 113 DVTGVEVVYP-SSMNARLVRRRLRHIAMRGTIIHRKFFYGSVSLIPLSSAFSILPL 167
+ V+V++P +++ A +R LR IA +HRK + S+ PL++ +++P+
Sbjct: 91 ETKKVDVLFPGNAIKAEKIRSILRKIATERQDLHRKKMWWSLVAAPLTAPIALIPV 146
>emb|CAG87323.1| unnamed protein product [Debaryomyces hansenii CBS767]
gi|50421213|ref|XP_459152.1| unnamed protein product
[Debaryomyces hansenii]
Length = 364
Score = 41.2 bits (95), Expect = 0.032
Identities = 38/166 (22%), Positives = 73/166 (43%), Gaps = 8/166 (4%)
Query: 107 LKSISKDVTGVEVVYPSSMNARLVRRRLRHIAMRGTIIHRKFFYGSVSLIPLSSAFSILP 166
LK S + + + +P + + +L H K+ IP+S F+++P
Sbjct: 172 LKIPSSQIKPIPLYHPKFQSPTTILNQLYLFRDDAYKTHLKYSILCAIGIPISLPFALVP 231
Query: 167 L-PNVPFFWILFRSYSHWRALQGSEKLFQLVSD----GSQSSNTYSGKKETEHEDSENES 221
+ PNVP F+I +R Y + +AL G + L L+ D S +N S KK + ++ +
Sbjct: 232 IVPNVPGFYIAYRLYCNVKALLGVKHLDYLLEDCTHKPSAPANASSEKKAIQSTVNDTKH 291
Query: 222 LG---LDEPHWVLTPSKELENIVRQEDGNDGLSRGTIEEICKIYDL 264
+ +D + + L + D ++ I+ +C+ +DL
Sbjct: 292 IAFEHVDHLDAIYLSNTSLSDHNLLSDEKIIITPDIIDGLCQNFDL 337
>gb|AAS52024.1| ADR104Cp [Ashbya gossypii ATCC 10895] gi|45187977|ref|NP_984200.1|
ADR104Cp [Eremothecium gossypii]
Length = 306
Score = 40.0 bits (92), Expect = 0.072
Identities = 22/84 (26%), Positives = 44/84 (52%), Gaps = 6/84 (7%)
Query: 119 VVYPSSMNARLVRRRLRHIAMRGTIIHRKFFYGSVSLIPLSSAFSILPL-PNVPFFWILF 177
V+ S +N LVR + G H ++ + +PL+ +++PL PN+P F++ +
Sbjct: 156 VISESVLNDELVR-----LLREGKQTHMRYLIYCLLALPLTIPIALVPLVPNIPGFYLAY 210
Query: 178 RSYSHWRALQGSEKLFQLVSDGSQ 201
R+Y +++A G+ L ++ Q
Sbjct: 211 RAYCNFKAYSGARHLESMMQGDMQ 234
>ref|YP_041112.1| Spo0B-associated GTP-binding protein [Staphylococcus aureus subsp.
aureus MRSA252] gi|49242017|emb|CAG40715.1|
Spo0B-associated GTP-binding protein [Staphylococcus
aureus subsp. aureus MRSA252]
Length = 430
Score = 35.4 bits (80), Expect = 1.8
Identities = 19/62 (30%), Positives = 33/62 (52%)
Query: 210 KETEHEDSENESLGLDEPHWVLTPSKELENIVRQEDGNDGLSRGTIEEICKIYDLNTQDV 269
K+ + E ES+G++ + TPS++ I R +DG +S IE + K+ D N+
Sbjct: 332 KDVDFTVEEEESVGINRVLYKHTPSQDKFTISRDDDGAYVVSGNAIERMFKMTDFNSDPA 391
Query: 270 VK 271
V+
Sbjct: 392 VR 393
>emb|CAG43381.1| Spo0B-associated GTP-binding protein [Staphylococcus aureus subsp.
aureus MSSA476] gi|14247416|dbj|BAB57806.1|
Spo0B-associated GTP-binding protein [Staphylococcus
aureus subsp. aureus Mu50] gi|21204763|dbj|BAB95459.1|
Spo0B-associated GTP-binding protein [Staphylococcus
aureus subsp. aureus MW2] gi|15927224|ref|NP_374757.1|
Spo0B-associated GTP-binding protein [Staphylococcus
aureus subsp. aureus N315] gi|13701442|dbj|BAB42736.1|
Spo0B-associated GTP-binding protein [Staphylococcus
aureus subsp. aureus N315] gi|49486477|ref|YP_043698.1|
Spo0B-associated GTP-binding protein [Staphylococcus
aureus subsp. aureus MSSA476]
gi|21283323|ref|NP_646411.1| Spo0B-associated
GTP-binding protein [Staphylococcus aureus subsp. aureus
MW2] gi|25295786|pir||C89947 Spo0B-associated
GTP-binding protein [imported] - Staphylococcus aureus
(strain N315) gi|15924634|ref|NP_372168.1|
Spo0B-associated GTP-binding protein [Staphylococcus
aureus subsp. aureus Mu50]
Length = 430
Score = 35.4 bits (80), Expect = 1.8
Identities = 19/62 (30%), Positives = 33/62 (52%)
Query: 210 KETEHEDSENESLGLDEPHWVLTPSKELENIVRQEDGNDGLSRGTIEEICKIYDLNTQDV 269
K+ + E ES+G++ + TPS++ I R +DG +S IE + K+ D N+
Sbjct: 332 KDVDFTVEEEESVGINRVLYKHTPSQDKFTISRDDDGAYVVSGNAIERMFKMTDFNSDPA 391
Query: 270 VK 271
V+
Sbjct: 392 VR 393
>gb|AAN40686.1| translationally controlled tumor protein-like protein [Zea mays]
Length = 167
Score = 35.0 bits (79), Expect = 2.3
Identities = 21/68 (30%), Positives = 31/68 (44%), Gaps = 2/68 (2%)
Query: 54 PFNAKAELFTDYVANKMNNGWVTLENAPDGSFKKKIHGLGLWLLSRVKPSEIFLKSISKD 113
PF+ K+ F Y+ + N LE FKK + G +LLS++K + F+ KD
Sbjct: 80 PFDKKS--FVSYIKKYIKNLTAVLEPEKADEFKKGVEGATKFLLSKLKDLQFFVGESMKD 137
Query: 114 VTGVEVVY 121
V Y
Sbjct: 138 DASVAFAY 145
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.317 0.134 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 478,581,355
Number of Sequences: 2540612
Number of extensions: 20178931
Number of successful extensions: 53843
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 16
Number of HSP's successfully gapped in prelim test: 20
Number of HSP's that attempted gapping in prelim test: 53810
Number of HSP's gapped (non-prelim): 40
length of query: 277
length of database: 863,360,394
effective HSP length: 126
effective length of query: 151
effective length of database: 543,243,282
effective search space: 82029735582
effective search space used: 82029735582
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 74 (33.1 bits)
Medicago: description of AC135796.7


