

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135317.6 + phase: 0
(432 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
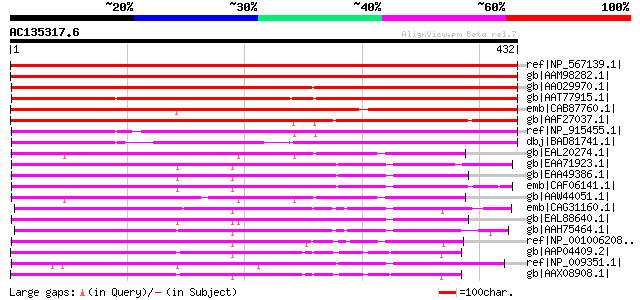
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_567139.1| GTP-binding protein-related [Arabidopsis thaliana] 457 e-127
gb|AAM98282.1| At3g63150/T20O10_250 [Arabidopsis thaliana] gi|16... 454 e-126
gb|AAO29970.1| unknown protein [Arabidopsis thaliana] gi|1969886... 441 e-122
gb|AAT77915.1| expressed protein [Oryza sativa (japonica cultiva... 439 e-122
emb|CAB87760.1| rac-GTP binding protein-like [Arabidopsis thalia... 418 e-115
gb|AAF27037.1| unknown protein [Arabidopsis thaliana] gi|1522996... 363 7e-99
ref|NP_915455.1| rac-GTP binding protein -like [Oryza sativa (ja... 340 7e-92
dbj|BAD81741.1| putative mitochondrial Rho 1 [Oryza sativa (japo... 323 5e-87
gb|EAL20274.1| hypothetical protein CNBF0860 [Cryptococcus neofo... 221 3e-56
gb|EAA71923.1| conserved hypothetical protein [Gibberella zeae P... 219 2e-55
gb|EAA49386.1| hypothetical protein MG01044.4 [Magnaporthe grise... 218 2e-55
emb|CAF06141.1| conserved hypothetical protein [Neurospora crass... 216 1e-54
gb|AAW44051.1| vesicle-mediated transport-related protein, putat... 207 4e-52
emb|CAG31160.1| hypothetical protein [Gallus gallus] 202 2e-50
gb|EAL88640.1| mitochondrial GTPase (Miro-2), putative [Aspergil... 198 2e-49
gb|AAH75464.1| Ras homolog gene family, member T1 [Xenopus tropi... 195 3e-48
ref|NP_001006208.1| similar to hypothetical protein [Gallus gall... 194 6e-48
gb|AAP04409.2| miro protein [Bos taurus] gi|31342304|ref|NP_8478... 194 6e-48
ref|NP_009351.1| Evolutionarily-conserved tail-anchored outer mi... 194 6e-48
gb|AAX08908.1| ras homolog gene family, member T2 [Bos taurus] 194 6e-48
>ref|NP_567139.1| GTP-binding protein-related [Arabidopsis thaliana]
Length = 643
Score = 457 bits (1176), Expect = e-127
Identities = 224/433 (51%), Positives = 311/433 (71%), Gaps = 1/433 (0%)
Query: 1 MDGTLTDHEVNEFQIRSFGASLQQPDITQLKTMVLRNVPEGVNSLGLTFPGFIEIHNMFL 60
+DG L D E+N+FQ+ FGA L ++ +K +V P+GV LGLT PGF+ + ++F+
Sbjct: 211 LDGALNDAELNDFQVNCFGAPLDPVELMGVKKVVQERQPDGVTDLGLTLPGFLFLFSLFI 270
Query: 61 KKGRTETFWAVLRKFGYGNDLKLRDDFLPVPSKKASDQSVELSGAAIEFLKGVFRLVDTD 120
++GR ET WA+LRK GY + L+L + LPVP+K++ DQS+EL+ A++FL G+F+L D D
Sbjct: 271 ERGRPETAWAILRKCGYNDSLELHAELLPVPAKQSPDQSIELTNEAMDFLSGIFQLYDLD 330
Query: 121 KDQLLRPAEVDKLFDAAPESPWKDAPYMDAAETTDMGYISLKGFLSRWALMTLLDPRYSL 180
D L+PAE+D LF AP+SPW + PY +AAE T G +++ GFLS WALMTLLDPR SL
Sbjct: 331 NDGALQPAELDDLFQTAPDSPWLEDPYKEAAEKTPGGSLTINGFLSEWALMTLLDPRKSL 390
Query: 181 ANLICIGYRDNPSAALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRALLGRPF 240
ANL IGY +P++ +T +RS DRK Q+TERNVFQC+VFG K +GKSALL + LGR F
Sbjct: 391 ANLTYIGYGHDPASTFSVTRKRSVDRKKQRTERNVFQCFVFGPKKSGKSALLDSFLGRKF 450
Query: 241 SNDYTPTTVERYAANIIELIAGTKKTLILREIPEDEVSNFLSNKDCLAACDVAAFVHDSS 300
SN Y T ERYAAN+I+ G+KKTLILREIPED V FL+NK+ LAACDVA V+DSS
Sbjct: 451 SNSYKATMGERYAANVIDQPGGSKKTLILREIPEDRVKKFLTNKESLAACDVAVVVYDSS 510
Query: 301 DGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDLTPFPRAVLDSVKVAQELKIDAPI 360
D YSW+++ ++L +V +GE G+ P LL+AAKDDL P+P +V +S +V EL ID P+
Sbjct: 511 DVYSWRKAREILMEVARRGEERGYGTPCLLVAAKDDLDPYPMSVQESDRVCMELGIDIPV 570
Query: 361 RLSIKSDDSNNVYSKIINAAEHPHLSIPETEFVRKRKQHQQLLH-TFIFALAGAAMALAG 419
LS+K + N+++S+I++ AE+PH+SIPETE R+ + +QL++ + +F G A+ AG
Sbjct: 571 SLSMKLGEPNSLFSRIVSTAENPHMSIPETESGRRSRNIRQLVNSSLLFVSVGTAVGFAG 630
Query: 420 FTARRARANRNSS 432
A RA + R ++
Sbjct: 631 LAAYRAYSARKNA 643
>gb|AAM98282.1| At3g63150/T20O10_250 [Arabidopsis thaliana]
gi|16648803|gb|AAL25592.1| AT3g63150/T20O10_250
[Arabidopsis thaliana]
Length = 643
Score = 454 bits (1168), Expect = e-126
Identities = 223/433 (51%), Positives = 310/433 (71%), Gaps = 1/433 (0%)
Query: 1 MDGTLTDHEVNEFQIRSFGASLQQPDITQLKTMVLRNVPEGVNSLGLTFPGFIEIHNMFL 60
+DG L D E+N+FQ+ FGA L ++ +K +V P+GV LGLT PGF+ + ++F+
Sbjct: 211 LDGALNDAELNDFQVNCFGAPLDPVELMGVKKVVQERQPDGVTDLGLTLPGFLFLFSLFI 270
Query: 61 KKGRTETFWAVLRKFGYGNDLKLRDDFLPVPSKKASDQSVELSGAAIEFLKGVFRLVDTD 120
++GR ET WA+LRK GY + L+L + LPVP+K++ DQS+EL+ A++FL G+F+L D D
Sbjct: 271 ERGRPETAWAILRKCGYNDSLELHAELLPVPAKQSPDQSIELTNEAMDFLSGIFQLYDLD 330
Query: 121 KDQLLRPAEVDKLFDAAPESPWKDAPYMDAAETTDMGYISLKGFLSRWALMTLLDPRYSL 180
D L+PAE+D LF AP+SPW + PY +AAE T G +++ GFLS WALMTLLDPR SL
Sbjct: 331 NDGALQPAELDDLFQTAPDSPWLEDPYKEAAEKTPGGSLTINGFLSEWALMTLLDPRKSL 390
Query: 181 ANLICIGYRDNPSAALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRALLGRPF 240
ANL IGY +P++ +T +RS DRK Q+TERNVFQC+VFG K + KSALL + LGR F
Sbjct: 391 ANLTYIGYGHDPASTFSVTRKRSVDRKKQRTERNVFQCFVFGPKKSRKSALLDSFLGRKF 450
Query: 241 SNDYTPTTVERYAANIIELIAGTKKTLILREIPEDEVSNFLSNKDCLAACDVAAFVHDSS 300
SN Y T ERYAAN+I+ G+KKTLILREIPED V FL+NK+ LAACDVA V+DSS
Sbjct: 451 SNSYKATIGERYAANVIDQPGGSKKTLILREIPEDRVKKFLTNKESLAACDVAVVVYDSS 510
Query: 301 DGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDLTPFPRAVLDSVKVAQELKIDAPI 360
D YSW+++ ++L +V +GE G+ P LL+AAKDDL P+P +V +S +V EL ID P+
Sbjct: 511 DVYSWRKAREILMEVARRGEERGYGTPCLLVAAKDDLDPYPMSVQESDRVCMELGIDIPV 570
Query: 361 RLSIKSDDSNNVYSKIINAAEHPHLSIPETEFVRKRKQHQQLLH-TFIFALAGAAMALAG 419
LS+K + N+++S+I++ AE+PH+SIPETE R+ + +QL++ + +F G A+ AG
Sbjct: 571 SLSMKLGEPNSLFSRIVSTAENPHMSIPETESGRRSRNIRQLVNSSLLFVSVGTAVGFAG 630
Query: 420 FTARRARANRNSS 432
A RA + R ++
Sbjct: 631 LAAYRAYSARKNA 643
>gb|AAO29970.1| unknown protein [Arabidopsis thaliana] gi|19698867|gb|AAL91169.1|
unknown protein [Arabidopsis thaliana]
gi|15240981|ref|NP_198106.1| GTP-binding protein-related
[Arabidopsis thaliana]
Length = 648
Score = 441 bits (1134), Expect = e-122
Identities = 222/433 (51%), Positives = 303/433 (69%), Gaps = 3/433 (0%)
Query: 2 DGTLTDHEVNEFQIRSFGASLQQPDITQLKTMVLRNVPEGVNSLGLTFPGFIEIHNMFLK 61
DG L++ E+N+FQ++ F A LQ +I +K +V +PEGVN GLT GF+ +H +F++
Sbjct: 215 DGALSEAELNDFQVKCFHAPLQPSEIEGVKRVVQEKLPEGVNERGLTVTGFLFLHALFIE 274
Query: 62 KGRTETFWAVLRKFGYGNDLKLRDDFLPVPS-KKASDQSVELSGAAIEFLKGVFRLVDTD 120
KGR ET W VLRKFGY ND++L ++ LP K+A DQS EL+ AAI+FLKG++ L D D
Sbjct: 275 KGRLETTWTVLRKFGYNNDIRLAEELLPSAIFKRAPDQSFELTNAAIDFLKGMYMLFDDD 334
Query: 121 KDQLLRPAEVDKLFDAAPESPWKDAPYMDAAETTDMGYISLKGFLSRWALMTLLDPRYSL 180
+D LRP E++ LF APESPWK+APY DAAE T +G +S FLS W+LMTLL+P S+
Sbjct: 335 QDNNLRPQEIEDLFSTAPESPWKEAPYEDAAEKTALGGLSFDAFLSMWSLMTLLEPARSV 394
Query: 181 ANLICIGYRDNPSAALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRALLGRPF 240
NLI IG+ +PS A+R+T RR DRK Q+ ER VFQC+VFG AGKSALL LGR +
Sbjct: 395 ENLIYIGFPGDPSTAIRVTRRRRLDRKKQQCERKVFQCFVFGPNNAGKSALLNCFLGRSY 454
Query: 241 SNDYTPTTVERYAANIIELIAGTKKTLILREIPEDEVSNFLSNKDCLAACDVAAFVHDSS 300
+++ TT ERYA N+++ +G KKTLI+REIPED V S+K+ LAACD+A FV+DSS
Sbjct: 455 TDNQESTTDERYAVNMVD-ESGAKKTLIMREIPEDGVQGLFSSKESLAACDIAVFVYDSS 513
Query: 301 DGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDLTPFPRAVLDSVKVAQELKIDAPI 360
D SWKR+ LL +V N GE TG+ P L+++AKDDL P ++ +S ++ Q++ I+ P+
Sbjct: 514 DESSWKRATQLLVEVANYGEATGYEVPCLMVSAKDDLDSSPISIQESTRMTQDMGIEPPV 573
Query: 361 RLSIKSDDSNNVYSKIINAAEHPHLSIPETEFVRKRKQHQQLLHTFIFALA-GAAMALAG 419
+S K D NN++ KI+ AA+HPHLSIPETE + RK + +L++ + A++ GAA + G
Sbjct: 574 SISSKLGDFNNLFRKILTAAQHPHLSIPETEAGKSRKHYNRLINRSLMAVSIGAAAVVVG 633
Query: 420 FTARRARANRNSS 432
A R A R SS
Sbjct: 634 LAAYRVYATRKSS 646
>gb|AAT77915.1| expressed protein [Oryza sativa (japonica cultivar-group)]
Length = 642
Score = 439 bits (1129), Expect = e-122
Identities = 219/432 (50%), Positives = 310/432 (71%), Gaps = 4/432 (0%)
Query: 2 DGTLTDHEVNEFQIRSFGASLQQPDITQLKTMVLRNVPEGVNSLGLTFPGFIEIHNMFLK 61
DG L+D E+N+FQ++ F A LQ +I +K +V +PEGVN GLT GF+ +H +F++
Sbjct: 212 DGALSDVELNDFQVKCFNAPLQPTEIAGVKRVVQEKMPEGVNDNGLTLTGFLFLHALFIE 271
Query: 62 KGRTETFWAVLRKFGYGNDLKLRDDFLPVPSKKASDQSVELSGAAIEFLKGVFRLVDTDK 121
KGR ET W VLRKFGY N++KLRDD +P K+A DQ++EL+G AI+FL+G+F + DTD
Sbjct: 272 KGRLETTWTVLRKFGYDNEIKLRDDLIPT-IKRAPDQTLELTGQAIDFLRGIFNMFDTDN 330
Query: 122 DQLLRPAEVDKLFDAAPESPWKDAPYMDAAETTDMGYISLKGFLSRWALMTLLDPRYSLA 181
D L PAE+D LF APE+PW + PY+D AE +G +SL+GFLS+WALMTLLDP S A
Sbjct: 331 DDALLPAELDDLFSTAPENPWSNNPYVDCAERNVLGGLSLEGFLSKWALMTLLDPANSFA 390
Query: 182 NLICIGYRDNPSAALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRALLGRPFS 241
NLI +GY + +A +R DRK Q+T+RNVFQCYVFG + AGK+ALL++ LGR
Sbjct: 391 NLIYVGYSGDFGSAFTTMRKRRVDRKKQQTQRNVFQCYVFGPRGAGKTALLQSFLGRQ-P 449
Query: 242 NDYTPTTVERYAANIIELIAGTKKTLILREIPEDEVSNFLSNKDCLAACDVAAFVHDSSD 301
+D P ER+AAN +EL +G++KTL+ REIPED+V L++++ LA CDVA FV+DS D
Sbjct: 450 SDALPMNGERFAANTVEL-SGSRKTLVFREIPEDDVRPLLADRESLAPCDVAVFVYDSCD 508
Query: 302 GYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDLTPFPRAVLDSVKVAQELKIDAPIR 361
+SW+R+ DLL +V GE TG+ P L++AAKDDL P A+ +S +V+Q++ I+ PI
Sbjct: 509 EFSWQRTRDLLVEVATHGENTGYEVPCLIVAAKDDLDQSPLALQESTRVSQDMGIEMPIP 568
Query: 362 LSIKSDDSNNVYSKIINAAEHPHLSIPETEFVRKRKQHQQLLHTFIFALA-GAAMALAGF 420
+S++ D NN++ +I++AA+ PHLSIPETE + R+Q++QLL+ + ++ GAA+A+ G
Sbjct: 569 ISVRLRDLNNIFCRIVHAAQQPHLSIPETEAGKTRRQYRQLLNRSLMVVSVGAAVAVVGI 628
Query: 421 TARRARANRNSS 432
A R A R ++
Sbjct: 629 AAYRVYAARKNT 640
>emb|CAB87760.1| rac-GTP binding protein-like [Arabidopsis thaliana]
gi|11358744|pir||T48104 rac-GTP binding protein-like -
Arabidopsis thaliana
Length = 676
Score = 418 bits (1074), Expect = e-115
Identities = 218/473 (46%), Positives = 305/473 (64%), Gaps = 48/473 (10%)
Query: 1 MDGTLTDHEVNEFQIRSFGASLQQPDITQLKTMVLRNVPEGVNSLGLTFPGFIEIHNMFL 60
+DG L D E+N+FQ+ FGA L ++ +K +V P+GV LGLT PGF+ + ++F+
Sbjct: 211 LDGALNDAELNDFQVNCFGAPLDPVELMGVKKVVQERQPDGVTDLGLTLPGFLFLFSLFI 270
Query: 61 KKGRTETFWAVLRKFGYGNDLKLRDDFLPVPSKKASDQSVELSGAAIEFLKGVFRLVDTD 120
++GR ET WA+LRK GY + L+L + LPVP+K++ DQS+EL+ A++FL G+F+L D D
Sbjct: 271 ERGRPETAWAILRKCGYNDSLELHAELLPVPAKQSPDQSIELTNEAMDFLSGIFQLYDLD 330
Query: 121 KDQLLRPAEVDKLFDAAPES---------------------------------------- 140
D L+PAE+D LF AP+S
Sbjct: 331 NDGALQPAELDDLFQTAPDSELTFFLLNFLANFFNALVHEYVYYFRNMFLYTYNLLYDFS 390
Query: 141 PWKDAPYMDAAETTDMGYISLKGFLSRWALMTLLDPRYSLANLICIGYRDNPSAALRLTS 200
PW + PY +AAE T G +++ GFLS WALMTLLDPR SLANL IGY +P++ +T
Sbjct: 391 PWLEDPYKEAAEKTPGGSLTINGFLSEWALMTLLDPRKSLANLTYIGYGHDPASTFSVTR 450
Query: 201 RRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELI 260
+RS DRK Q+TERNVFQC+VFG K +GKSALL + LGR FSN Y T ERYAAN+I+
Sbjct: 451 KRSVDRKKQRTERNVFQCFVFGPKKSGKSALLDSFLGRKFSNSYKATMGERYAANVIDQP 510
Query: 261 AGTKKTLILREIPEDEVSNFLSNKDCLAACDVAAFVHDSSDGYSWKRSIDLLEKVVNQGE 320
G+KKTLILREIPED V FL+NK+ LAACDVA V+D +++ ++L +V +GE
Sbjct: 511 GGSKKTLILREIPEDRVKKFLTNKESLAACDVAVVVYD-------RKAREILMEVARRGE 563
Query: 321 LTGHRFPSLLIAAKDDLTPFPRAVLDSVKVAQELKIDAPIRLSIKSDDSNNVYSKIINAA 380
G+ P LL+AAKDDL P+P +V +S +V EL ID P+ LS+K + N+++S+I++ A
Sbjct: 564 ERGYGTPCLLVAAKDDLDPYPMSVQESDRVCMELGIDIPVSLSMKLGEPNSLFSRIVSTA 623
Query: 381 EHPHLSIPETEFVRKRKQHQQLLH-TFIFALAGAAMALAGFTARRARANRNSS 432
E+PH+SIPETE R+ + +QL++ + +F G A+ AG A RA + R ++
Sbjct: 624 ENPHMSIPETESGRRSRNIRQLVNSSLLFVSVGTAVGFAGLAAYRAYSARKNA 676
>gb|AAF27037.1| unknown protein [Arabidopsis thaliana] gi|15229963|ref|NP_187182.1|
GTP-binding protein-related [Arabidopsis thaliana]
Length = 648
Score = 363 bits (931), Expect = 7e-99
Identities = 198/441 (44%), Positives = 288/441 (64%), Gaps = 12/441 (2%)
Query: 1 MDGTLTDHEVNEFQIRSFGASLQQPDITQLKTMVLRNVPEGVNSLGLTFPGFIEIHNMFL 60
MDG L+D E+NE Q + F L +I Q+K ++ P+GVN GLT GF+ ++ +
Sbjct: 211 MDGILSDEELNELQKKCFDTPLVPCEIKQMKNVMQVTFPQGVNERGLTLDGFLFLNTRLI 270
Query: 61 KKGRTETFWAVLRKFGYGNDLKLRDDFLPVPS-KKASDQSVELSGAAIEFLKGVFRLVDT 119
++ R +T W +LRKFGY NDL+L DD +P S K+ +DQSVEL+ AIEFL+ V+ D+
Sbjct: 271 EEARIQTLWTMLRKFGYSNDLRLGDDLVPYSSFKRQADQSVELTNVAIEFLREVYEFFDS 330
Query: 120 DKDQLLRPAEVDKLFDAAPESPWKDAPYMDAAETTDMGYISLKGFLSRWALMTLLDPRYS 179
+ D L P E+ LF+ APESPW Y D E G +SL+ FLS W+LMTL+DP S
Sbjct: 331 NGDNNLEPHEMGYLFETAPESPWTKPLYKDVTEENMDGGLSLEAFLSLWSLMTLIDPPRS 390
Query: 180 LANLICIGY-RDNPSAALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRALLGR 238
L L+ I + D+PS+A+R+T +R DRK +K+ER V QC+VFG K AGKSALL +GR
Sbjct: 391 LEYLMYIRFPSDDPSSAVRVTRKRVLDRKEKKSERKVVQCFVFGPKNAGKSALLNQFIGR 450
Query: 239 PF---SNDYTPTTVERYAANIIE---LIAGTKKTLILREIPEDEVSNFLSNKDCLAACDV 292
+ SN+ +T E YA N+++ +I+ T KTL+L+E+ + F+ +K+ LAACDV
Sbjct: 451 SYDDDSNNNNGSTDEHYAVNMVKEPGVISDTDKTLVLKEVRIKD-DGFMLSKEALAACDV 509
Query: 293 AAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDLTPFPRAVLDSVKVAQ 352
A F++DSSD YSW R++D+L +V + +G+ FP L++AAK DL PFP A+ +S +V Q
Sbjct: 510 AIFIYDSSDEYSWNRAVDMLAEVATIAKDSGYVFPCLMVAAKTDLDPFPVAIQESTRVTQ 569
Query: 353 ELKIDAPIRLSIKSDDSNNVYSKIINAAEHPHLSIPETEFVRKRKQHQQLLHTFIFALA- 411
++ IDAPI +S K D +N++ KI+ AAE+PHL+IPE E K+K+ +L + + A++
Sbjct: 570 DIGIDAPIPISSKLGDVSNLFRKILTAAENPHLNIPEIE--SKKKRSCKLNNRSLMAVSI 627
Query: 412 GAAMALAGFTARRARANRNSS 432
G A+ +AG + R R S
Sbjct: 628 GTAVLIAGLASFRLYTARKQS 648
>ref|NP_915455.1| rac-GTP binding protein -like [Oryza sativa (japonica
cultivar-group)]
Length = 634
Score = 340 bits (871), Expect = 7e-92
Identities = 187/458 (40%), Positives = 276/458 (59%), Gaps = 35/458 (7%)
Query: 2 DGTLTDHEVNEFQIRSFGASLQQPDITQLKTMVLRNVPEGVNSLGLTFPGFIEIHNMFLK 61
DG +D E+N+FQ+ F A LQ +I +K + + EGVN GLT GF+ +H + +
Sbjct: 183 DGAFSDVELNDFQVICFNAPLQPNEIIGVKRTIQEKLTEGVNENGLTLTGFLFLHTLIIG 242
Query: 62 KGRTETFWAVLRKFGYGNDLKLRDDFLPVPSKKASDQSVELSGAAIEFLKGVFRLVDTDK 121
G+ ET W VLRKFGY N+LKLRDD +P K+A DQ + G+F T
Sbjct: 243 NGKLETTWTVLRKFGYDNELKLRDDLIPA-IKRAPDQFIYTPF-------GIFITNITTV 294
Query: 122 DQLLRPAEVDKLFDAAPESPWKDAPYMDAAETTDMGYISLKGFLSRWALMTLLDPRYSLA 181
D L+PAE++ LF APE+PW Y + AE +G +S +GF+S+W LMTL+ P S A
Sbjct: 295 DGALQPAEINDLFSTAPENPWSSHLYENCAENNVLGGLSFEGFISKWTLMTLIHPSNSFA 354
Query: 182 NLICIGYRDNPSAALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRALLGRPFS 241
NLI +GY + +A T +R DRK ++T+RNVFQCYVFG + AGK+ALL++ L R S
Sbjct: 355 NLIYVGYPGDFDSAFTTTRKRRVDRKKKQTQRNVFQCYVFGPRHAGKTALLQSFLKRYHS 414
Query: 242 ----------NDYTPTTVERYAANIIEL----------------IAGTKKTLILREIPED 275
+ P + N+ ++ + GT+KTL++REI E
Sbjct: 415 IVFYIMNFVCHGNLPMLLLSMVNNLQQILLNCLIFILNSATFTHVFGTRKTLVMREISEG 474
Query: 276 EVSNFLSNKDCLAACDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKD 335
+V LS+K+ LA CDVA V+DS D SW+R+ +LL +V +G+ TG+ P L++AAKD
Sbjct: 475 DVGPLLSDKESLAPCDVAVIVYDSGDEVSWQRARELLVQVATRGKNTGYEVPCLIVAAKD 534
Query: 336 DLTPFPRAVLDSVKVAQELKIDAPIRLSIKSDDSNNVYSKIINAAEHPHLSIPETEFVRK 395
DL P A+ DS +V+ ++ I+ PI +S++ D NN++ +I++AA+ PHLSIPETE +
Sbjct: 535 DLDQSPLALQDSTRVSHDMGIETPIPISVRLRDLNNIFCRIVHAAQQPHLSIPETEAGKT 594
Query: 396 RKQHQQLLHTFIFALA-GAAMALAGFTARRARANRNSS 432
+Q++QLL+ + ++ GAA+A+ G A R A R ++
Sbjct: 595 HRQYRQLLNRSLTVVSVGAAVAVVGVAAYRVYAARKNA 632
>dbj|BAD81741.1| putative mitochondrial Rho 1 [Oryza sativa (japonica
cultivar-group)]
Length = 594
Score = 323 bits (829), Expect = 5e-87
Identities = 177/432 (40%), Positives = 259/432 (58%), Gaps = 47/432 (10%)
Query: 2 DGTLTDHEVNEFQIRSFGASLQQPDITQLKTMVLRNVPEGVNSLGLTFPGFIEIHNMFLK 61
DG +D E+N+FQ+ F A LQ +I +K + + EGVN GLT GF+ +H + +
Sbjct: 207 DGAFSDVELNDFQVICFNAPLQPNEIIGVKRTIQEKLTEGVNENGLTLTGFLFLHTLIIG 266
Query: 62 KGRTETFWAVLRKFGYGNDLKLRDDFLPVPSKKASDQSVELSGAAIEFLKGVFRLVDTDK 121
G+ ET W VLRKFGY N+LKLRDD +P K+A DQ
Sbjct: 267 NGKLETTWTVLRKFGYDNELKLRDDLIPA-IKRAPDQ----------------------- 302
Query: 122 DQLLRPAEVDKLFDAAPESPWKDAPYMDAAETTDMGYISLKGFLSRWALMTLLDPRYSLA 181
D L+PAE++ LF APE+PW Y + AE +G +S +GF+S+W LMTL+ P S A
Sbjct: 303 DGALQPAEINDLFSTAPENPWSSHLYENCAENNVLGGLSFEGFISKWTLMTLIHPSNSFA 362
Query: 182 NLICIGYRDNPSAALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRALLGRPFS 241
NLI +GY + +A T +R DRK ++T+RNVFQ +P
Sbjct: 363 NLIYVGYPGDFDSAFTTTRKRRVDRKKKQTQRNVFQ--------------------QP-- 400
Query: 242 NDYTPTTVERYAANIIELIAGTKKTLILREIPEDEVSNFLSNKDCLAACDVAAFVHDSSD 301
+D P E++AAN +EL GT+KTL++REI E +V LS+K+ LA CDVA V+DS D
Sbjct: 401 SDAPPVNGEQFAANTVELPDGTRKTLVMREISEGDVGPLLSDKESLAPCDVAVIVYDSGD 460
Query: 302 GYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDLTPFPRAVLDSVKVAQELKIDAPIR 361
SW+R+ +LL +V +G+ TG+ P L++AAKDDL P A+ DS +V+ ++ I+ PI
Sbjct: 461 EVSWQRARELLVQVATRGKNTGYEVPCLIVAAKDDLDQSPLALQDSTRVSHDMGIETPIP 520
Query: 362 LSIKSDDSNNVYSKIINAAEHPHLSIPETEFVRKRKQHQQLLHTFIFALA-GAAMALAGF 420
+S++ D NN++ +I++AA+ PHLSIPETE + +Q++QLL+ + ++ GAA+A+ G
Sbjct: 521 ISVRLRDLNNIFCRIVHAAQQPHLSIPETEAGKTHRQYRQLLNRSLTVVSVGAAVAVVGV 580
Query: 421 TARRARANRNSS 432
A R A R ++
Sbjct: 581 AAYRVYAARKNA 592
>gb|EAL20274.1| hypothetical protein CNBF0860 [Cryptococcus neoformans var.
neoformans B-3501A]
Length = 686
Score = 221 bits (563), Expect = 3e-56
Identities = 144/433 (33%), Positives = 212/433 (48%), Gaps = 53/433 (12%)
Query: 2 DGTLTDHEVNEFQIRSFGASLQQPDITQLKTMVLRNVPEGVNSL---------------- 45
DG L HE+N+FQ + F LQ ++ + +V P V L
Sbjct: 205 DGLLNAHELNQFQQKCFSTPLQSQELDGILEIVRSYDPYAVQPLPSSSPNTPLSRDSSYG 264
Query: 46 -----------------GLTFPGFIEIHNMFLKKGRTETFWAVLRKFGYGNDLKLRDDFL 88
G+T GF+ +H MF+++GR ET W VLRKFGYG L LR+DFL
Sbjct: 265 QLHYFNNNVVPPSPPQEGITELGFLYLHTMFIQQGRMETTWTVLRKFGYGESLDLREDFL 324
Query: 89 PVPSKKASDQSVELSGAAIEFLKGVFRLVDTDKDQLLRPAEVDKLFDAAPESPWKDAPYM 148
SD SVELS +FL +F D D+D L E+D LF +P +PW +
Sbjct: 325 APKFDVPSDCSVELSPLGNQFLTDIFEAYDKDQDGALSQNELDDLFSTSPGNPWLSQGFP 384
Query: 149 DAAETTDMGYISLKGFLSRWALMTLLDPRYSLANLICIGYRDNPS------AALRLTSRR 202
D T DMG ++L+G+L++W++ TLL+ R +L L +GY +P+ AL +T R
Sbjct: 385 DTTITDDMGRVTLQGWLAQWSMTTLLNHRTTLNYLAYLGYSSSPATDLPTPTALHVTRPR 444
Query: 203 SEDRKNQKTERNVFQCYVFGSKTAGKSALLRALLGRPFS------NDYTPTTVERYAANI 256
+DR+ +K RNVF CYV G+ +GK++LLR+ + RPF Y PTT N
Sbjct: 445 KQDRRQRKVTRNVFLCYVLGATGSGKTSLLRSFVNRPFKGGEDGLGGYEPTTKVLSVVNS 504
Query: 257 IELIAGTKKTLILREIPEDEVSNFLSNKDCLAACDVAAFVHDSSDGYSWKRSIDLLEKVV 316
+E+ G +K L+L+E S L N L D+ +VHDSSD S+ +L +
Sbjct: 505 VEM-EGVEKYLVLQEFGSKYESEILRNSKRLDMADIIIYVHDSSDTNSFSYISNLRQ--- 560
Query: 317 NQGELTGHRFPSLLIAAKDDL-TPFPRAVLDSVKVAQELKIDAPIRLSIKSDDSNNVYSK 375
+ + PS+ +A K DL R + + L + AP+ +S + +N++
Sbjct: 561 ---QYSLDHIPSIFVATKSDLDLAQQRHEVQPDVYCRRLGLQAPMAVSSRLGPLHNLWVA 617
Query: 376 IINAAEHPHLSIP 388
I A P S+P
Sbjct: 618 ITRVALDPTSSLP 630
>gb|EAA71923.1| conserved hypothetical protein [Gibberella zeae PH-1]
gi|46128137|ref|XP_388622.1| conserved hypothetical
protein [Gibberella zeae PH-1]
Length = 627
Score = 219 bits (557), Expect = 2e-55
Identities = 138/435 (31%), Positives = 219/435 (49%), Gaps = 19/435 (4%)
Query: 2 DGTLTDHEVNEFQIRSFGASLQQPDITQLKTMVLRNVPEGVNSLGLTFPGFIEIHNMFLK 61
DG L + E+ +FQ R F L D+ +K + +++P G+ PGF++++ ++ +
Sbjct: 202 DGYLNEQEMRDFQARCFDKPLTTDDLDNIKLSIAKSLPASDLEKGIDLPGFLQLNKLYAE 261
Query: 62 KGRTETFWAVLRKFGYGNDLKLRDDFLPVPSKKASDQSVELSGAAIEFLKGVFRLVDTDK 121
KGR ET W +LRKF Y + L L D F+ + S ELS A F +F + D D
Sbjct: 262 KGRHETIWIILRKFHYTDSLSLEDKFIRPKFEVPEYSSAELSPAGYRFFVDLFLIFDKDN 321
Query: 122 DQLLRPAEVDKLFDAAPESP--WKDAPYMDAAETTDMGYISLKGFLSRWALMTLLDPRYS 179
D L E++ LF AP P W D+ + + + G+++L+G+L++W++ T ++P+ +
Sbjct: 322 DGGLNDEELEALFAPAPGLPSSWTDSSFPSSTVRNEAGHVTLQGWLAQWSMTTFIEPKTT 381
Query: 180 LANLICIGY-----RDNPSAALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRA 234
+ L +G+ +D+ +AAL++T R + + ERNV CYV G+ AGKSALL +
Sbjct: 382 IEYLAYLGFEPSNPKDSITAALKITKPRKRRSRLGRVERNVVLCYVLGASGAGKSALLDS 441
Query: 235 LLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDEVSNFLSNKDCLAACDVAA 294
L RPF Y PT R A N +EL G + LIL E+ E E + L N+ L ACD+
Sbjct: 442 FLNRPFYGLYHPTIKPRRAVNSVELPGGKQVYLILEELGELEPA-ILENRAKLDACDLIC 500
Query: 295 FVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDLTPF-PRAVLDSVKVAQE 353
+ +DSSD S+ +DL +K + EL PS+ A K D R L +
Sbjct: 501 YAYDSSDPDSFSHIVDLRKKYPHLDEL-----PSIYTALKADKDKTNQRCELQPDQYTSS 555
Query: 354 LKIDAPIRLSIKSDDSNNVYSKIINAAEHPHLSIPETEFVRKRKQHQQLLHTFIFALAGA 413
L + P+ +S+ + ++ +AA + P T F R ++ +I A A
Sbjct: 556 LSMSLPLHVSVTWGSISELFVAYADAA-----TTPSTAFPRSNEEGPDRTSLYIALGATA 610
Query: 414 AMALAGFTARRARAN 428
+A T R N
Sbjct: 611 CAGVAALTIWRRATN 625
>gb|EAA49386.1| hypothetical protein MG01044.4 [Magnaporthe grisea 70-15]
gi|39973619|ref|XP_368200.1| hypothetical protein
MG01044.4 [Magnaporthe grisea 70-15]
Length = 634
Score = 218 bits (556), Expect = 2e-55
Identities = 132/398 (33%), Positives = 209/398 (52%), Gaps = 14/398 (3%)
Query: 2 DGTLTDHEVNEFQIRSFGASLQQPDITQLKTMVLRNVPEGVNSLGLTFPGFIEIHNMFLK 61
DG L D E++EFQ RSF L+ ++ +KT + + +P LG+ PGF++++ ++ +
Sbjct: 209 DGYLNDREMHEFQARSFDKPLKPEELENIKTTIAKAIPGSRIDLGVDLPGFLQLNKLYAE 268
Query: 62 KGRTETFWAVLRKFGYGNDLKLRDDFLPVPSKKASDQSVELSGAAIEFLKGVFRLVDTDK 121
KGR ET W +LR++ Y + L L+D FL S ELS A F +F L D D
Sbjct: 269 KGRHETIWTILRQYHYTDSLSLQDSFLHPKFDVPEYASAELSPAGYRFFVDLFLLFDKDN 328
Query: 122 DQLLRPAEVDKLFDAAPESP--WKDAPYMDAAETTDMGYISLKGFLSRWALMTLLDPRYS 179
D L AE+ +F P P W + + + + G+I+L+G+L++W++ T ++P+ +
Sbjct: 329 DGGLNDAELAAMFAPTPGLPHSWSETSFPSSTVRNEAGHITLQGWLAQWSMTTFVEPKTT 388
Query: 180 LANLICIGY-----RDNPSAALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRA 234
L L +G+ RD +AAL++T R RK + ERNV CY+ G+ AGKS+LL A
Sbjct: 389 LEYLAYLGFEPPTPRDTITAALKITKPRKRRRKPGRVERNVVLCYIIGASGAGKSSLLDA 448
Query: 235 LLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDEVSNFLSNKDCLAACDVAA 294
L RPF Y PT R A N +EL G + LIL E+ E E + L N+ L ACD+
Sbjct: 449 FLNRPFEPLYHPTIKPRRAVNSVELQGGKQCYLILEELGELEPA-ILENQAKLDACDLIC 507
Query: 295 FVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDL-TPFPRAVLDSVKVAQE 353
+ +DSSD S+ +L + +L P++ A K D R+ L
Sbjct: 508 YAYDSSDPDSFSHIENLRRRHPQLDDL-----PAIYTALKADRDKTTQRSELQPDAYTSS 562
Query: 354 LKIDAPIRLSIKSDDSNNVYSKIINAAEHPHLSIPETE 391
L + P+ +S+ + ++ + AA +P + P++E
Sbjct: 563 LNMSTPLHVSVTWSSISELFVALAEAATNPSTAFPKSE 600
>emb|CAF06141.1| conserved hypothetical protein [Neurospora crassa]
gi|32405344|ref|XP_323285.1| hypothetical protein
[Neurospora crassa] gi|28918698|gb|EAA28369.1|
hypothetical protein [Neurospora crassa]
Length = 629
Score = 216 bits (549), Expect = 1e-54
Identities = 141/435 (32%), Positives = 221/435 (50%), Gaps = 18/435 (4%)
Query: 2 DGTLTDHEVNEFQIRSFGASLQQPDITQLKTMVLRNVPEGVNSLGLTFPGFIEIHNMFLK 61
DG L D E+ +FQ +SF L Q D+ +K V ++VP GL GF++++ ++ +
Sbjct: 203 DGYLNDQEMQDFQQKSFDKPLSQEDLDNIKLTVSKSVPSSSTDKGLDLRGFLQLNKLYAE 262
Query: 62 KGRTETFWAVLRKFGYGNDLKLRDDFLPVPSKKASDQSVELSGAAIEFLKGVFRLVDTDK 121
KGR ET W +LRK+ Y + L L D FL S ELS A F +F D D
Sbjct: 263 KGRHETIWIILRKYHYTDSLSLEDSFLHPRFDVPDYASAELSPAGYRFFMDLFLTFDKDN 322
Query: 122 DQLLRPAEVDKLFDAAPESP--WKDAPYMDAAETTDMGYISLKGFLSRWALMTLLDPRYS 179
D L E+ LF P P W + + + G+I+L+G+L++W++ T L+P+ +
Sbjct: 323 DGGLNDRELAALFAPTPGLPHSWAETSFPSTTVRNEAGHITLQGWLAQWSMTTFLEPKTT 382
Query: 180 LANLICIGY-----RDNPSAALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRA 234
L L +G+ R+ +AAL++T R R+ + +RNV CY+ GS AGKS+LL
Sbjct: 383 LEYLAYLGFETPNARETTTAALKITKPRKRRRRPGRVDRNVVLCYILGSSGAGKSSLLDV 442
Query: 235 LLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDEVSNFLSNKDCLAACDVAA 294
L RPF Y PT R A N +EL G + LIL E+ E E + L N+ L ACD+
Sbjct: 443 FLNRPFDTLYHPTIKPRQAVNSVELQGGKQCYLILEELGELEPA-ILENQAKLDACDLIC 501
Query: 295 FVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDL-TPFPRAVLDSVKVAQE 353
+ +DSS+ S+ ++L ++ EL P++ A K D R+ L
Sbjct: 502 YAYDSSEPDSFSHIVELRKRYPQLDEL-----PAVYTALKADRDKTTQRSELQPDAYTAA 556
Query: 354 LKIDAPIRLSIKSDDSNNVYSKIINAAEHPHLSIPETEFVRKRKQHQQLLHTFIFALAGA 413
L + AP+ +S+ + + ++ + AA +P + P +E + + L+ + A A A
Sbjct: 557 LNMSAPLHVSVTWNSISELFVALAEAATNPSTAFPRSE---EPPADRASLYMALGATACA 613
Query: 414 AMALAGFTARRARAN 428
A+A A RR+ +N
Sbjct: 614 ALA-AFMIWRRSTSN 627
>gb|AAW44051.1| vesicle-mediated transport-related protein, putative [Cryptococcus
neoformans var. neoformans JEC21]
gi|58268404|ref|XP_571358.1| vesicle-mediated
transport-related protein, putative [Cryptococcus
neoformans var. neoformans JEC21]
Length = 681
Score = 207 bits (528), Expect = 4e-52
Identities = 142/433 (32%), Positives = 206/433 (46%), Gaps = 58/433 (13%)
Query: 2 DGTLTDHEVNEFQIRSFGASLQQPDITQLKTMVLRNVPEGVNSL---------------- 45
DG L HE+N+FQ + F LQ ++ + +V P V L
Sbjct: 205 DGLLNAHELNQFQQKCFSTPLQSQELDGILEIVRSYDPYAVQPLPSSSPNTPLSRDSSYG 264
Query: 46 -----------------GLTFPGFIEIHNMFLKKGRTETFWAVLRKFGYGNDLKLRDDFL 88
G+T GF+ +H MF+++GR ET W VLRKFGYG L LR+DFL
Sbjct: 265 QLHYFNNNVVPPSPPQEGITELGFLYLHTMFIQQGRMETTWTVLRKFGYGESLDLREDFL 324
Query: 89 PVPSKKASDQSVELSGAAIEFLKGVFRLVDTDKDQLLRPAEVDKLFDAAPESPWKDAPYM 148
SD SVELS +FL +F D D+D L E+D LF +P +PW +
Sbjct: 325 APKFDVPSDCSVELSPLGNQFLTDIFEAYDKDQDGALSQNELDDLFSTSPGNPWLSQGFP 384
Query: 149 DAAETTDMGYISLKGFLSRWALMTLLDPRYSLANLICIGYRDNPS------AALRLTSRR 202
D T DMG ++L+G + TLL+ R +L L +GY +P+ AL +T R
Sbjct: 385 DTTITDDMGRVTLQG-----CMTTLLNHRTTLNYLAYLGYSSSPATDLPTPTALHVTRPR 439
Query: 203 SEDRKNQKTERNVFQCYVFGSKTAGKSALLRALLGRPFS------NDYTPTTVERYAANI 256
+DR+ +K RNVF CYV G+ +GK++LLR+ + RPF Y PTT N
Sbjct: 440 KQDRRQRKVTRNVFLCYVLGATGSGKTSLLRSFVNRPFKGGEDGLGGYEPTTKVLSVVNS 499
Query: 257 IELIAGTKKTLILREIPEDEVSNFLSNKDCLAACDVAAFVHDSSDGYSWKRSIDLLEKVV 316
+E+ G +K L+L+E S L N L D+ +VHDSSD S+ +L +
Sbjct: 500 VEM-EGVEKYLVLQEFGSKYESEILRNSKRLDMADIIIYVHDSSDTNSFSYISNLRQ--- 555
Query: 317 NQGELTGHRFPSLLIAAKDDL-TPFPRAVLDSVKVAQELKIDAPIRLSIKSDDSNNVYSK 375
+ + PS+ +A K DL R + + L + AP+ +S + +N++
Sbjct: 556 ---QYSLDHIPSIFVATKSDLDLAQQRHEVQPDVYCRRLGLQAPMAVSSRLGPLHNLWVA 612
Query: 376 IINAAEHPHLSIP 388
I A P S+P
Sbjct: 613 ITRVALDPTSSLP 625
>emb|CAG31160.1| hypothetical protein [Gallus gallus]
Length = 618
Score = 202 bits (514), Expect = 2e-50
Identities = 142/432 (32%), Positives = 214/432 (48%), Gaps = 29/432 (6%)
Query: 5 LTDHEVNEFQIRSFGASLQQPDITQLKTMVLRNVPEGVNSLGLTFPGFIEIHNMFLKKGR 64
L+D E+N FQ FG L + +K +V +N +GV GLT GF+ ++ +F+++GR
Sbjct: 204 LSDDELNYFQKSCFGNPLAPQALEDVKMVVWKNTTDGVQDNGLTLNGFLFLNTLFIQRGR 263
Query: 65 TETFWAVLRKFGYGNDLKLRDDFLPVPSKKASDQSVELSGAAIEFLKGVFRLVDTDKDQL 124
ET W +LR+FGY ++L+L DD+L + S EL+ +FL+ +F D D+D
Sbjct: 264 HETTWTILRRFGYDDELELTDDYLYPQFRLPPGCSTELNHLGYQFLQRLFEKHDKDQDGA 323
Query: 125 LRPAEVDKLFDAAPESPWKDAPYMDAAETTDMGYISLKGFLSRWALMTLLDPRYSLANLI 184
L PAE+ F P PW Y + TTD G +SL GFL +W L+ LD R+ L L
Sbjct: 324 LSPAELQNFFSVFPCMPWGPELY-NTVCTTDKGLLSLHGFLCQWTLIAYLDVRHCLECLG 382
Query: 185 CIGY-----RDNPSAALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRALLGRP 239
+GY +D+ + AL +T + D + +T+RNVF C V G++ AGKSA L+A LGR
Sbjct: 383 YLGYPILSEQDSQTQALTVTREKRIDLEKGQTQRNVFLCKVLGARGAGKSAFLQAFLGRS 442
Query: 240 F-SNDYTPTTVERYAANIIELIAGTKKTLILREIPEDEVSNFLSNKDCLAACDVAAFVHD 298
+ +P Y N ++ + G +K LIL E+ + + F D AACDVA ++D
Sbjct: 443 LAAQRESPGEPSPYTINTVQ-VNGQEKYLILHEVSAE--TQFTKPSD--AACDVACLIYD 497
Query: 299 SSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDLTPFPRAV-LDSVKVAQELKID 357
SD S+ + ++ ++ P + +A+K DL + L + + +
Sbjct: 498 LSDPKSFSYCASIYKQHYMDSQI-----PCVFVASKTDLPEASQQPGLSPAEFCYKHCLP 552
Query: 358 APIRLSIKSD--DSNNVYSKIINAAEHPHLSIPETEFVRKRKQHQQLLHTFIFALAGAAM 415
P S S +Y+K+ AA PHL+ E AL A
Sbjct: 553 PPFLFSCHSQGPPGTAIYTKLATAATFPHLNAVELGAAS---------FWLRVALGAAVT 603
Query: 416 ALAGFTARRARA 427
AL GFT R A
Sbjct: 604 ALVGFTLYRVLA 615
>gb|EAL88640.1| mitochondrial GTPase (Miro-2), putative [Aspergillus fumigatus
Af293]
Length = 632
Score = 198 bits (504), Expect = 2e-49
Identities = 134/400 (33%), Positives = 201/400 (49%), Gaps = 15/400 (3%)
Query: 2 DGTLTDHEVNEFQIRSFGASLQQPDITQLKTMVLRNVPEGVNSLGLTFPGFIEIHNMFLK 61
DG L+D E+ +FQ+R F L + D+ +K + + P+ V G+ GFI ++ M+ +
Sbjct: 203 DGYLSDKEIEDFQMRCFDKPLSKEDLVHIKETIQKTHPDSVTPFGIDCRGFIHLNKMYAE 262
Query: 62 KGRTETFWAVLRKFGYGNDLKLRDDFLPVPSKKASDQSVELSGAAIEFLKGVFRLVDTDK 121
KGR ET W +LR F Y ++L L++ FL + S ELS F +F L D D
Sbjct: 263 KGRHETVWIILRAFQYTDNLSLQESFLHPRFEVPPFASAELSPEGYRFFVNLFLLSDKDN 322
Query: 122 DQLLRPAEVDKLFDAAPESP--WKDAPYMDAAETTDMGYISLKGFLSRWALMTLLDPRYS 179
D L AE+ LF P P W D + + + G+++L+G+L++W++ T P+ +
Sbjct: 323 DGGLNDAELASLFAPTPGLPASWADGSFPSSTVRNEAGHVTLQGWLAQWSMTTFQSPKTT 382
Query: 180 LANLICIGY----RDNPS--AALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLR 233
L L +G+ R NPS AAL++T R + ++ + RNV +V G+ +GKSALL
Sbjct: 383 LEYLAYLGFESSDRSNPSTTAALKVTRPRKKRKRPGRVGRNVVLGHVLGAPGSGKSALLD 442
Query: 234 ALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDEVSNFLSNKDCLAACDVA 293
A L R FS+ Y PT R A N +EL G + LIL E+ E E + + L CDV
Sbjct: 443 AFLARGFSHTYRPTIQPRTAVNTVELPGGKQCYLILDELGELEPAILENQAKLLDQCDVI 502
Query: 294 AFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDL-TPFPRAVLDSVKVAQ 352
+ +DSSD S+ L K + EL PS+ IA K DL RA +
Sbjct: 503 VYTYDSSDPDSFAYIPQLRSKYPHLEEL-----PSVFIALKADLDRTTQRAEHQPHEYTA 557
Query: 353 ELKIDA-PIRLSIKSDDSNNVYSKIINAAEHPHLSIPETE 391
L + + P+ +S V+ I AA P + P +E
Sbjct: 558 MLNMPSPPLHVSATWSSIQEVFVHIAEAAMEPSTAFPRSE 597
>gb|AAH75464.1| Ras homolog gene family, member T1 [Xenopus tropicalis]
gi|55742344|ref|NP_001006725.1| ras homolog gene family,
member T1 [Xenopus tropicalis]
Length = 616
Score = 195 bits (495), Expect = 3e-48
Identities = 131/431 (30%), Positives = 204/431 (46%), Gaps = 33/431 (7%)
Query: 5 LTDHEVNEFQIRSFGASLQQPDITQLKTMVLRNVPEGVNSLGLTFPGFIEIHNMFLKKGR 64
L+D E+N FQ FG L + +K +V +N +GV GLT GF+ ++ +F+++GR
Sbjct: 204 LSDEELNFFQQSCFGNPLAPQALEDVKMVVKKNTADGVRDNGLTLNGFLFLNTLFIQRGR 263
Query: 65 TETFWAVLRKFGYGNDLKLRDDFLPVPSKKASDQSVELSGAAIEFLKGVFRLVDTDKDQL 124
ET W +LR+FGY + L+L DD+L P + + S EL+ +FL+ F D D+D
Sbjct: 264 HETTWTILRRFGYDDALELTDDYLYPPLRIPHESSTELNHFGYQFLQKAFEKHDLDEDGA 323
Query: 125 LRPAEVDKLFDAAPESPWKDAPYMDAAETTDMGYISLKGFLSRWALMTLLDPRYSLANLI 184
L P+E+ F P +PW T GY+ L G+L +W L+ LD L +L
Sbjct: 324 LSPSELQSFFSVFPYTPW-GPELASTVCTAQGGYLPLHGYLCQWTLVAYLDVHRCLEHLG 382
Query: 185 CIGY-----RDNPSAALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRALLGRP 239
+GY +++ + A+ +T +S D + +T+RNVF C V G + GKSA LRA LG+
Sbjct: 383 YLGYPILCEQESQTHAITVTREKSIDLEKGQTQRNVFLCRVIGPRGTGKSAFLRAFLGQS 442
Query: 240 FSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDEVSNFLSNKDCLAACDVAAFVHDS 299
+ + L+ G +K LIL E+ D + FL D A CDVA ++D
Sbjct: 443 LEEQQQSNKPPSFYSVNTVLVGGQEKYLILFEVDVD--TEFLKTSD--APCDVACLMYDV 498
Query: 300 SDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDLTPFPRA-VLDSVKVAQELKIDA 358
SD S+ + ++ + + P L + K D + + + + ++
Sbjct: 499 SDSKSFNYCASIYKQHYMESQT-----PCLFVGCKYDQGEVKQQHGISPAEFCHKHRLPP 553
Query: 359 PIRLSIKSDDSNNVYSKIINAAEHPHLSIPETEFVRKRKQHQQLLHTFIF----ALAGAA 414
P + + +YSK+ AA PHL H L T F AL
Sbjct: 554 PYHFTCQGTPDRTIYSKLATAAAFPHL-------------HDTELSTASFWLRVALGATV 600
Query: 415 MALAGFTARRA 425
A+ GFT +A
Sbjct: 601 AAVVGFTLYKA 611
>ref|NP_001006208.1| similar to hypothetical protein [Gallus gallus]
gi|53127682|emb|CAG31170.1| hypothetical protein [Gallus
gallus]
Length = 619
Score = 194 bits (492), Expect = 6e-48
Identities = 127/395 (32%), Positives = 208/395 (52%), Gaps = 21/395 (5%)
Query: 2 DGTLTDHEVNEFQIRSFGASLQQPDITQLKTMVLRNVPEGVNSLGLTFPGFIEIHNMFLK 61
DGTL D E+N FQ F L + +K +V +NV +GV GLT GF+ +H +F++
Sbjct: 201 DGTLNDAELNFFQRICFNTPLAPQALEDVKNVVRKNVSDGVADNGLTLKGFLFLHTLFIQ 260
Query: 62 KGRTETFWAVLRKFGYGNDLKLRDDFLPVPSKKASDQSVELSGAAIEFLKGVFRLVDTDK 121
+GR ET W VLR+FGY +DL+L ++L K D + EL+ A FL+ +F D D+
Sbjct: 261 RGRHETTWTVLRRFGYDDDLELTPEYLFPLLKIPPDCTTELNHHAYLFLQSIFDKHDLDR 320
Query: 122 DQLLRPAEVDKLFDAAPESPWKDAPYMDAAETTDMGYISLKGFLSRWALMTLLDPRYSLA 181
D L P E+ LF P PW G+I+ +GFLS+W L T LD + L
Sbjct: 321 DCALSPDELKDLFKVFPYMPWGPDVNNTVCTNGKGGWITYQGFLSQWTLTTYLDVQRCLE 380
Query: 182 NLICIGY-----RDNPSAALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRALL 236
L +GY +++ ++A+ +T + D + ++T+RNVF+C V G K GKS +L+ALL
Sbjct: 381 YLGYLGYSILAEQESQASAITVTRDKKIDLQKKQTQRNVFRCNVVGMKGCGKSGVLQALL 440
Query: 237 GRPFSNDYTPTTVER--YAANIIELIAGTKKTLILREIPEDEVSNFLSNKDCLAACDVAA 294
GR + YA N + + G +K L+L ++ + E FL++ + + CDV
Sbjct: 441 GRNLMRQRQIRAEHKSYYAINTV-YVYGQEKYLLLHDVSDSE---FLTDAETI--CDVVC 494
Query: 295 FVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDLTPFPRAV-LDSVKVAQE 353
V+D S+ S++ + + ++ R P L++AAK DL + + + ++
Sbjct: 495 LVYDVSNPKSFEYCVRIFKQ-----HFMDSRIPCLVVAAKSDLHEVRQEYSISPAEFCKK 549
Query: 354 LKIDAPIRLSIKSDD--SNNVYSKIINAAEHPHLS 386
K+ P + + D S +++ K+ A +PH++
Sbjct: 550 HKMPPPQAFTCNTVDMPSKDIFVKLTTMAMYPHVT 584
>gb|AAP04409.2| miro protein [Bos taurus] gi|31342304|ref|NP_847886.2|
mitochondrial Rho 2 [Bos taurus]
Length = 618
Score = 194 bits (492), Expect = 6e-48
Identities = 130/394 (32%), Positives = 203/394 (50%), Gaps = 23/394 (5%)
Query: 1 MDGTLTDHEVNEFQIRSFGASLQQPDITQLKTMVLRNVPEGVNSLGLTFPGFIEIHNMFL 60
MD L+D E+N FQ FG L + +K +V +NV GV LT GF+ ++ +F+
Sbjct: 200 MDQALSDQELNAFQTSCFGHPLAPQALEDVKMVVSKNVVGGVRDDPLTLEGFLFLNTLFI 259
Query: 61 KKGRTETFWAVLRKFGYGNDLKLRDDFLPVPSKKASDQSVELSGAAIEFLKGVFRLVDTD 120
++GR ET W +LR+FGYG+ L+L D+L P + S EL+ +F++ +F D D
Sbjct: 260 QRGRHETTWTILRRFGYGDSLELTADYLCPPLRVPPGCSAELNHRGYQFVQRMFEKHDQD 319
Query: 121 KDQLLRPAEVDKLFDAAPESPWKDAPYMDAAETTDMGYISLKGFLSRWALMTLLDPRYSL 180
+D L PAE+ LF P +PW P++ + T G + L G+L +W L+T LD R SL
Sbjct: 320 RDGALSPAELQSLFSVFPAAPW--GPHLPSTVRTKAGRLPLHGYLCQWTLVTYLDVRRSL 377
Query: 181 ANLICIGY-----RDNPSAALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRAL 235
+L +GY +D+ + A+ +T + D++ +T+RNV C V G++ GKS+ LRA
Sbjct: 378 EHLGYLGYPTLCEQDSQAHAITVTREKRLDQEKGQTQRNVLLCKVVGARGVGKSSFLRAF 437
Query: 236 LGRPFSN-DYTPTTVERYAANIIELIAGTKKTLILREIPEDEVSNFLSNKDCLAACDVAA 294
LG + D +V YA + ++ + G +K LIL E+ D L A+CDVA
Sbjct: 438 LGHSLGHQDAGEPSV--YAIDTVQ-VNGQEKYLILCEVAADS----LLTASADASCDVAC 490
Query: 295 FVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDLT-PFPRAVLDSVKVAQE 353
+ D SD RS L V Q + G + P L + +K DL P L + +
Sbjct: 491 LMFDGSD----LRSFALCASVYKQHYMDG-QTPCLFVCSKADLPGGVPLPGLSPAEFCRR 545
Query: 354 LKIDAPIRLSIKS--DDSNNVYSKIINAAEHPHL 385
++ P S + +++++ A PHL
Sbjct: 546 HRLPTPTLFSCAGPVEPCMGIFTRLATMATFPHL 579
>ref|NP_009351.1| Evolutionarily-conserved tail-anchored outer mitochondrial membrane
GTPase which regulates mitochondrial morphology; cells
lacking Gem1p contain collapsed, globular, or grape-like
mitochondria; not required for pheromone-induced cell
death; Gem1p [Saccharomyces cerevisiae]
gi|731284|sp|P39722|YAE8_YEAST Hypothetical 75.2 kDa
protein in ACS1-GCV3 intergenic region
gi|595536|gb|AAC04983.1| Yal048cp [Saccharomyces
cerevisiae]
Length = 662
Score = 194 bits (492), Expect = 6e-48
Identities = 138/439 (31%), Positives = 216/439 (48%), Gaps = 25/439 (5%)
Query: 2 DGTLTDHEVNEFQIRSFGASLQQPDITQLKTMVL---RNVPEGVN------SLGLTFPGF 52
D L D+E+ Q + F S+ ++ +K ++L ++ E +N G+T GF
Sbjct: 218 DSYLDDNEILGLQKKCFNKSIDVNELNFIKDLLLDISKHDQEYINRKLYVPGKGITKDGF 277
Query: 53 IEIHNMFLKKGRTETFWAVLRKFGYGNDLKLRDDFLPVPSKKASDQSVELSGAAIEFLKG 112
+ ++ ++ ++GR ET WA+LR F Y + L + D L SVELS FL
Sbjct: 278 LVLNKIYAERGRHETTWAILRTFHYTDSLCINDKILHPRLVVPDTSSVELSPKGYRFLVD 337
Query: 113 VFRLVDTDKDQLLRPAEVDKLFDAAPESP--WKDAPYMDAAETTDMGYISLKGFLSRWAL 170
+F D D D L E+ +LF P P W + + + G I+L+G+L++W++
Sbjct: 338 IFLKFDIDNDGGLNNQELHRLFKCTPGLPKLWTSTNFPFSTVVNNKGCITLQGWLAQWSM 397
Query: 171 MTLLDPRYSLANLICIGYRDNPSAALRLTSRRSEDRKNQK------TERNVFQCYVFGSK 224
T L+ + A L+ G++++ AL++T R R++ K +R VF C+V G
Sbjct: 398 TTFLNYSTTTAYLVYFGFQEDARLALQVTKPRKMRRRSGKLYRSNINDRKVFNCFVIGKP 457
Query: 225 TAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDEVSNFLSNK 284
GKS+LL A LGR FS +Y+PT R A N +EL G + LIL+E+ E E + L NK
Sbjct: 458 CCGKSSLLEAFLGRSFSEEYSPTIKPRIAVNSLELKGGKQYYLILQELGEQEYA-ILENK 516
Query: 285 DCLAACDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDL-TPFPRA 343
D L CDV +DSSD S+ + LL+K + +L P + +A+K DL R
Sbjct: 517 DKLKECDVICLTYDSSDPESFSYLVSLLDKFTHLQDL-----PLVFVASKADLDKQQQRC 571
Query: 344 VLDSVKVAQELKIDAPIRLSIKSDDS-NNVYSKIINAAEHPHLSIPETEFVRKRKQHQQL 402
+ ++A EL ++ P+ +S + S N ++ KI AA P + P K
Sbjct: 572 QIQPDELADELFVNHPLHISSRWLSSLNELFIKITEAALDPGKNTPGLPEETAAKDVDYR 631
Query: 403 LHTFIFALAGAAMALAGFT 421
IF +AL FT
Sbjct: 632 QTALIFGSTVGFVALCSFT 650
>gb|AAX08908.1| ras homolog gene family, member T2 [Bos taurus]
Length = 618
Score = 194 bits (492), Expect = 6e-48
Identities = 130/394 (32%), Positives = 203/394 (50%), Gaps = 23/394 (5%)
Query: 1 MDGTLTDHEVNEFQIRSFGASLQQPDITQLKTMVLRNVPEGVNSLGLTFPGFIEIHNMFL 60
MD L+D E+N FQ FG L + +K +V +NV GV LT GF+ ++ +F+
Sbjct: 200 MDQALSDQELNAFQTSCFGHPLAPQALEDVKMVVSKNVVGGVRDDQLTLDGFLFLNTLFI 259
Query: 61 KKGRTETFWAVLRKFGYGNDLKLRDDFLPVPSKKASDQSVELSGAAIEFLKGVFRLVDTD 120
++GR ET W +LR+FGYG+ L+L D+L P + S EL+ +F++ +F D D
Sbjct: 260 QRGRHETTWTILRRFGYGDSLELTADYLCPPLRVPPGCSAELNHRGYQFVQRMFEKHDQD 319
Query: 121 KDQLLRPAEVDKLFDAAPESPWKDAPYMDAAETTDMGYISLKGFLSRWALMTLLDPRYSL 180
+D L PAE+ LF P +PW P++ + T G + L G+L +W L+T LD R SL
Sbjct: 320 RDGALSPAELQSLFSVFPAAPW--GPHLPSTVRTKAGRLPLHGYLCQWTLVTYLDVRRSL 377
Query: 181 ANLICIGY-----RDNPSAALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRAL 235
+L +GY +D+ + A+ +T + D++ +T+RNV C V G++ GKS+ LRA
Sbjct: 378 EHLGYLGYPTLCEQDSQAHAITVTREKRLDQEKGQTQRNVLLCKVVGARGVGKSSFLRAF 437
Query: 236 LGRPFSN-DYTPTTVERYAANIIELIAGTKKTLILREIPEDEVSNFLSNKDCLAACDVAA 294
LG + D +V YA + ++ + G +K LIL E+ D L A+CDVA
Sbjct: 438 LGHSLGHQDAGEPSV--YAIDTVQ-VNGQEKYLILCEVAADS----LLTASADASCDVAC 490
Query: 295 FVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDLT-PFPRAVLDSVKVAQE 353
+ D SD RS L V Q + G + P L + +K DL P L + +
Sbjct: 491 LMFDGSD----LRSFALCASVYKQHYMDG-QTPCLFVCSKADLPGGVPLPGLSPAEFCRR 545
Query: 354 LKIDAPIRLSIKS--DDSNNVYSKIINAAEHPHL 385
++ P S + +++++ A PHL
Sbjct: 546 HRLPTPTLFSCAGPVEPCMGIFTRLATMATFPHL 579
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.319 0.135 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 702,374,067
Number of Sequences: 2540612
Number of extensions: 28890469
Number of successful extensions: 67533
Number of sequences better than 10.0: 283
Number of HSP's better than 10.0 without gapping: 205
Number of HSP's successfully gapped in prelim test: 78
Number of HSP's that attempted gapping in prelim test: 66979
Number of HSP's gapped (non-prelim): 382
length of query: 432
length of database: 863,360,394
effective HSP length: 131
effective length of query: 301
effective length of database: 530,540,222
effective search space: 159692606822
effective search space used: 159692606822
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 77 (34.3 bits)
Medicago: description of AC135317.6


