

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135311.8 + phase: 0
(189 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
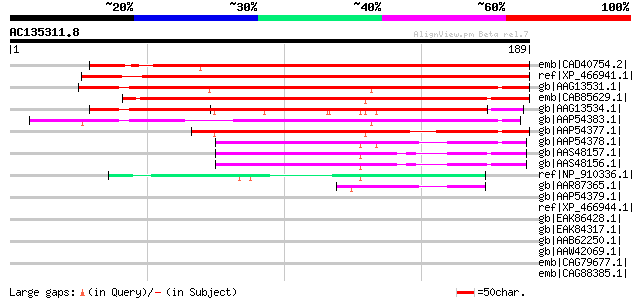
Score E
Sequences producing significant alignments: (bits) Value
emb|CAD40754.2| OSJNBa0081G05.7 [Oryza sativa (japonica cultivar... 189 3e-47
ref|XP_466941.1| ripening-related protein-like [Oryza sativa (ja... 186 2e-46
gb|AAG13531.1| hypothetical protein [Oryza sativa (japonica cult... 181 7e-45
emb|CAB85629.1| putative ripening-related protein [Vitis vinifera] 166 3e-40
gb|AAG13534.1| hypothetical protein [Oryza sativa (japonica cult... 164 2e-39
gb|AAP54383.1| hypothetical protein [Oryza sativa (japonica cult... 158 6e-38
gb|AAP54377.1| hypothetical protein [Oryza sativa (japonica cult... 115 8e-25
gb|AAP54378.1| hypothetical protein [Oryza sativa (japonica cult... 100 2e-20
gb|AAS48157.1| putative ripening related protein [Aegilops tausc... 80 4e-14
gb|AAS48156.1| putative ripening related protein [Aegilops tausc... 74 2e-12
ref|NP_910336.1| hypothetical protein~predicted by FGENESH etc. ... 63 5e-09
gb|AAR87365.1| hypothetical protein [Oryza sativa (japonica cult... 47 4e-04
gb|AAP54379.1| hypothetical protein [Oryza sativa (japonica cult... 45 0.001
ref|XP_466944.1| hypothetical protein [Oryza sativa (japonica cu... 42 0.009
gb|EAK86428.1| hypothetical protein UM05495.1 [Ustilago maydis 5... 42 0.011
gb|EAK84317.1| hypothetical protein UM03212.1 [Ustilago maydis 5... 39 0.074
gb|AAB62250.1| B2-aldehyde-forming enzyme [Schizophyllum commune] 38 0.17
gb|AAW42069.1| conserved hypothetical protein [Cryptococcus neof... 36 0.63
emb|CAG79677.1| unnamed protein product [Yarrowia lipolytica CLI... 35 0.82
emb|CAG88385.1| unnamed protein product [Debaryomyces hansenii C... 35 1.1
>emb|CAD40754.2| OSJNBa0081G05.7 [Oryza sativa (japonica cultivar-group)]
gi|50923499|ref|XP_472110.1| OSJNBa0081G05.7 [Oryza
sativa (japonica cultivar-group)]
Length = 183
Score = 189 bits (481), Expect = 3e-47
Identities = 92/161 (57%), Positives = 111/161 (68%), Gaps = 8/161 (4%)
Query: 30 CRPSGRIRGKKAPPGKCNQENDSDCCVQGKMYTTYVCSP-SVSTHTKAYLTLNSFQKGGD 88
C PSG IR +C DCC G+ Y TY CSP + + T A +TLN F GGD
Sbjct: 30 CNPSGAIRSSTTH--RCQ-----DCCKAGQSYPTYTCSPPTTGSSTDAVMTLNDFDAGGD 82
Query: 89 GGGPSECDKQYHSDDTPVVALSTGWFNHKSRCLNNITISANGRNVVAMVVDECDSRKGCD 148
GGGPSECD+ YHS+ VVALSTGW+ SRC ++ I+ANGR+V+A VVDECDS++GCD
Sbjct: 83 GGGPSECDEMYHSNTELVVALSTGWYAGGSRCGKSVRINANGRSVLAKVVDECDSQRGCD 142
Query: 149 EQHDYQPPCPNNIVDASKAVWKALGVPKEQWGGLDITWSDA 189
E+H YQPPC N+VDAS+AVW ALG+ E G DITWSDA
Sbjct: 143 EEHAYQPPCRPNVVDASQAVWDALGITGEDVGEYDITWSDA 183
>ref|XP_466941.1| ripening-related protein-like [Oryza sativa (japonica
cultivar-group)] gi|49388698|dbj|BAD25879.1|
ripening-related protein-like [Oryza sativa (japonica
cultivar-group)] gi|49387973|dbj|BAD25081.1|
ripening-related protein-like [Oryza sativa (japonica
cultivar-group)]
Length = 192
Score = 186 bits (473), Expect = 2e-46
Identities = 88/163 (53%), Positives = 110/163 (66%), Gaps = 7/163 (4%)
Query: 27 AQNCRPSGRIRGKKAPPGKCNQENDSDCCVQGKMYTTYVCSPSVSTHTKAYLTLNSFQKG 86
A C PSG +R ++ + DCC G+ Y TY CSP+ + TKA +TLN F+ G
Sbjct: 37 AGGCNPSGTLRPSRS-------HSCQDCCKAGRSYPTYACSPATTGSTKAVMTLNDFEAG 89
Query: 87 GDGGGPSECDKQYHSDDTPVVALSTGWFNHKSRCLNNITISANGRNVVAMVVDECDSRKG 146
GDGG PSECD ++H + VVALSTGW+ + RC NI I+ANGR+V+A VVDECDS G
Sbjct: 90 GDGGDPSECDGKFHKNTERVVALSTGWYANGRRCNKNIRINANGRSVLAKVVDECDSLHG 149
Query: 147 CDEQHDYQPPCPNNIVDASKAVWKALGVPKEQWGGLDITWSDA 189
CD++H YQPPC N+VDAS+AVW AL + E G DITWSDA
Sbjct: 150 CDKEHAYQPPCRPNVVDASQAVWDALRITGEDVGEYDITWSDA 192
>gb|AAG13531.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|31432798|gb|AAP54385.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
gi|37535592|ref|NP_922098.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
gi|40539115|gb|AAR87371.1| expressed protein [Oryza
sativa (japonica cultivar-group)]
Length = 213
Score = 181 bits (460), Expect = 7e-45
Identities = 90/168 (53%), Positives = 113/168 (66%), Gaps = 8/168 (4%)
Query: 26 EAQNCRPSGRIRGKKAPPGKCNQENDSDCCVQGKMYTTYVCSPSVS---THTKAYLTLNS 82
++ +C+PSG I G G CN +N S+CC G Y TY CSP V+ T T A LTLNS
Sbjct: 50 QSSSCQPSGAITGTS---GDCNADNGSECCQDGVQYMTYACSPPVAAGGTGTAALLTLNS 106
Query: 83 FQKGGDGGGPSECDKQYHSDDTPVVALSTGWFNHKSRCLNNITISANG-RNVVAMVVDEC 141
F GGDGGG C +++ D VVALSTGWF+ +SRC ++ I A+G +V AMVVDEC
Sbjct: 107 FADGGDGGGAPSCTGRFYDDGQLVVALSTGWFDGRSRCEKDVVIRASGGASVTAMVVDEC 166
Query: 142 DSRKGCDEQHDYQPPCPNNIVDASKAVWKALGVPKEQWGGLDITWSDA 189
DS++GCD H+++PPC NNIVD S AVW ALG+ K+ G ITWSDA
Sbjct: 167 DSQRGCDSDHNFEPPCRNNIVDGSPAVWDALGLNKDD-GEAQITWSDA 213
>emb|CAB85629.1| putative ripening-related protein [Vitis vinifera]
Length = 220
Score = 166 bits (420), Expect = 3e-40
Identities = 83/149 (55%), Positives = 102/149 (67%), Gaps = 3/149 (2%)
Query: 42 PPGKCNQENDSDCCVQGKMYTTYVCSPSVSTHTKAYLTLNSFQKGGDGGGPSECDKQYHS 101
PP C Q + + C GK TY CSP +++ T A LT N+F+KGGDGGGPS CD +YH
Sbjct: 74 PPSTC-QPSGTLACKGGKPKNTYTCSPPITSSTPAVLTNNNFEKGGDGGGPSACDNKYHD 132
Query: 102 DDTPVVALSTGWFNHKSRCLNNITISA-NGRNVVAMVVDECDSRKGCDEQHDYQPPCPNN 160
+ +VALSTGW+N SRC I I+A NGR+V+A VVDECDS GCD++H QPPC NN
Sbjct: 133 NSERIVALSTGWYNGGSRCGKMIRITAQNGRSVLAKVVDECDSMHGCDKEHAGQPPCDNN 192
Query: 161 IVDASKAVWKALGVPKEQWGGLDITWSDA 189
IVD S AVW ALG+ G +D+TWS A
Sbjct: 193 IVDGSNAVWNALGL-DINIGEVDVTWSMA 220
>gb|AAG13534.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|31432804|gb|AAP54391.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
gi|37535604|ref|NP_922104.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 377
Score = 164 bits (414), Expect = 2e-39
Identities = 82/150 (54%), Positives = 104/150 (68%), Gaps = 8/150 (5%)
Query: 30 CRPSGRIRGKKAPPGKCNQENDSDCCVQGKMYTTYVCSPSVSTH--TKAYLTLNSFQKGG 87
C SG + GK GKCN+ + SDCCV G+ Y + CSP VS+ T A LTLNSF +GG
Sbjct: 56 CHISGFLHGKA---GKCNRAHGSDCCVTGRRYPQFRCSPPVSSARPTPATLTLNSFARGG 112
Query: 88 DGGGPSECDKQYHSDDTPVVALSTGWF--NHKSRCLNNITISA-NGRNVVAMVVDECDSR 144
DGGG S CD ++H D VVALS+GW + SRC I ++A NGR+ +A VVDECDS
Sbjct: 113 DGGGRSSCDGRFHPDTAMVVALSSGWLRLDGASRCNRMIRVAAGNGRSALARVVDECDSV 172
Query: 145 KGCDEQHDYQPPCPNNIVDASKAVWKALGV 174
GCD +H+++PPCPN++VD S AVWKALG+
Sbjct: 173 NGCDAEHNFEPPCPNDVVDGSPAVWKALGL 202
Score = 91.7 bits (226), Expect = 1e-17
Identities = 53/124 (42%), Positives = 70/124 (55%), Gaps = 11/124 (8%)
Query: 74 TKAYLTLNSFQKGGDGGG-PSECDKQYHSDDTPVVALSTGWFN---HKSRCLNNITI--- 126
T A LTL F G D GG P+ CD ++H + VVALS+GW + RC I +
Sbjct: 247 TPAILTLKVFDHGEDDGGVPTSCDMRFHRNTELVVALSSGWLRLGGGRRRCHRRIRVFAV 306
Query: 127 --SANGRN-VVAMVVDECDSRKGCDEQHDYQPPCPNNIVDASKAVWKALGVPKEQWGGLD 183
+A+GR+ VVA VVD+CDS GC E+ + PPC NN V S VW+ LG+ G +
Sbjct: 307 AGAASGRSSVVARVVDDCDSVNGCREEDGFAPPCRNNAVGGSPVVWEKLGL-NASVGEFE 365
Query: 184 ITWS 187
+ WS
Sbjct: 366 VVWS 369
>gb|AAP54383.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|37535588|ref|NP_922096.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
gi|22726004|gb|AAN05004.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 276
Score = 158 bits (400), Expect = 6e-38
Identities = 91/186 (48%), Positives = 106/186 (56%), Gaps = 28/186 (15%)
Query: 8 VSFIFLAITLIFTSFVYS----EAQNCRPSGRIRGKKAPPGKCNQENDSDCCVQGKMYTT 63
++ IFL L T S A CRPSG + GK G C + ND DCC GK Y
Sbjct: 1 MAMIFLLAALSTTHLASSLRPVAAGACRPSGYLPGKS---GNCEKSNDPDCCEDGKAYPQ 57
Query: 64 YVCSPSVSTHTKAYLTLNSFQKGGDGGGPSECDKQYHSDDTPVVALSTGWFNHKSRCLNN 123
Y F+KG DGGGPSECD YHSD VVALSTGWF +RC +
Sbjct: 58 Y-----------------RFEKGKDGGGPSECDNAYHSDGELVVALSTGWFAGTARCGHR 100
Query: 124 ITISANG---RNVVAMVVDECDSRKGCDEQHDYQPPCPNNIVDASKAVWKALGVPKEQWG 180
+ I+A+G R+VVA VVDECDS GCD +H+Y+ PC NNIVDAS AVW ALG+ K G
Sbjct: 101 VRITASGGGGRSVVAKVVDECDSVHGCDGEHNYEAPCGNNIVDASPAVWDALGLDKNV-G 159
Query: 181 GLDITW 186
ITW
Sbjct: 160 MEHITW 165
>gb|AAP54377.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|37535576|ref|NP_922090.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
gi|16905188|gb|AAL31058.1| hypothetical protein [Oryza
sativa] gi|40539110|gb|AAR87366.1| hypothetical protein
[Oryza sativa (japonica cultivar-group)]
Length = 167
Score = 115 bits (287), Expect = 8e-25
Identities = 65/126 (51%), Positives = 81/126 (63%), Gaps = 13/126 (10%)
Query: 67 SPSVSTH-TKAYLTLNSFQKGGDGGGPSECDKQYHSDDTPVVALSTGWFNHKSRCLNNIT 125
SP+VS T A +T+N F++G DGGGP+ CD +YHSD + V ALSTGWF RC I
Sbjct: 52 SPAVSEDGTPAVMTVNGFEEGEDGGGPAACDGRYHSDRSLVAALSTGWFAGGRRCHRGIR 111
Query: 126 ISA--NGRNVVAMVVDECDSRKGCDEQHDYQPPCPNNIVDASKAVWKALGVPKEQWGGLD 183
I++ NGR+VVA VVDECDSR G C ++IVD S AVW ALG+ G +
Sbjct: 112 ITSRQNGRSVVATVVDECDSRHG---------GCKDDIVDTSAAVWSALGLDTNV-GEVP 161
Query: 184 ITWSDA 189
+TWSDA
Sbjct: 162 VTWSDA 167
>gb|AAP54378.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|37535578|ref|NP_922091.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
gi|16905186|gb|AAL31056.1| hypothetical protein [Oryza
sativa] gi|40539111|gb|AAR87367.1| hypothetical protein
[Oryza sativa (japonica cultivar-group)]
Length = 162
Score = 100 bits (250), Expect = 2e-20
Identities = 57/123 (46%), Positives = 73/123 (59%), Gaps = 21/123 (17%)
Query: 76 AYLTLNSFQKGGDGGGPSECDKQYHSDDTPVVALSTGWFNHKSRCLNNITI--SANGRN- 132
A +T+N F+KG DGGGP+ CD YHSD +VALST WF RC I I S +GR
Sbjct: 50 AVMTVNGFEKGEDGGGPAACDGHYHSDGELIVALSTEWFAGGRRCHRRIRITPSEHGRRG 109
Query: 133 -------VVAMVVDECDSRKGCDEQHDYQPPCPNNIVDASKAVWKALGVPKEQWGGLDIT 185
V A VVDECDSR+GC + ++VD+S AVW+ALG+ + G + +T
Sbjct: 110 GGGGRRAVEATVVDECDSRRGCKD----------DVVDSSPAVWRALGLDTDS-GEVRVT 158
Query: 186 WSD 188
WSD
Sbjct: 159 WSD 161
>gb|AAS48157.1| putative ripening related protein [Aegilops tauschii]
Length = 145
Score = 79.7 bits (195), Expect = 4e-14
Identities = 50/116 (43%), Positives = 62/116 (53%), Gaps = 18/116 (15%)
Query: 76 AYLTLNSFQKGGDGGGPSECDKQYHSDDTPVVALSTGWFNHKSRCLNNITI---SANGRN 132
A +T+N FQ+G GGGP+ CD QYHSDD VV+LS+ W+ +RC I I S +
Sbjct: 42 AAMTVNGFQQGEGGGGPAACDGQYHSDDDLVVSLSSEWYAGGARCGRIIRIADPSNSNFG 101
Query: 133 VVAMVVDECDSRKGCDEQHDYQPPCPNNIVDASKAVWKALGVPKEQWGGLDITWSD 188
AMVVDEC GCD N V AS +WK + G +D TWSD
Sbjct: 102 TNAMVVDEC---AGCD-----------NEVGASSGIWKNFDL-DTSLGQVDTTWSD 142
>gb|AAS48156.1| putative ripening related protein [Aegilops tauschii]
Length = 141
Score = 74.3 bits (181), Expect = 2e-12
Identities = 46/114 (40%), Positives = 59/114 (51%), Gaps = 16/114 (14%)
Query: 76 AYLTLNSFQKGGDGGGPSECDKQYHSDDTPVVALSTGWFNHKSRCLNNITI-SANGRNVV 134
A +T+N FQ+G GGGP+ CD QYH+DD VV+LS+ W+ +RC I I +
Sbjct: 40 AVMTVNGFQEGEGGGGPASCDGQYHNDDEFVVSLSSEWYAGGARCGRTIRIVDTFNIGIN 99
Query: 135 AMVVDECDSRKGCDEQHDYQPPCPNNIVDASKAVWKALGVPKEQWGGLDITWSD 188
A VVDEC GCD N V AS +W + G +DI WSD
Sbjct: 100 AQVVDEC---AGCD-----------NEVGASSHIWNNFHL-DTSLGQVDIRWSD 138
>ref|NP_910336.1| hypothetical protein~predicted by FGENESH etc. [Oryza sativa
(japonica cultivar-group)] gi|24413966|dbj|BAC22218.1|
hypothetical protein [Oryza sativa (japonica
cultivar-group)]
Length = 253
Score = 62.8 bits (151), Expect = 5e-09
Identities = 47/159 (29%), Positives = 62/159 (38%), Gaps = 50/159 (31%)
Query: 37 RGKKAPPGKCNQENDSDCCVQGKMYTTYVCSPSVSTHTKAYLTLNS-FQKG--------- 86
RG + P G DCCV GK Y ++ + H A TL+ F++
Sbjct: 57 RGARCPKGS------PDCCVVGKRYPRFMWLTAGIGHAPAIPTLHMCFRRNMELVVELYS 110
Query: 87 -------GDGGGPSECDKQYHSDDTPVVALSTGWFNHKSRCLNNITI-----SANGRNVV 134
DGGG + C RC I + + NGR+VV
Sbjct: 111 GSGWLCLDDGGGGARC----------------------RRCNGRIRVVVTAAAVNGRSVV 148
Query: 135 AMVVDECDSRKGCDEQHDYQPPCPNNIVDASKAVWKALG 173
VVDECD GC + + PPC +N+ S VWK LG
Sbjct: 149 VRVVDECDYVNGCCNEDGFAPPCQDNVGGESLTVWKKLG 187
>gb|AAR87365.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
Length = 116
Score = 46.6 bits (109), Expect = 4e-04
Identities = 25/58 (43%), Positives = 34/58 (58%), Gaps = 14/58 (24%)
Query: 120 CLNN----ITISANGRNVVAMVVDECDSRKGCDEQHDYQPPCPNNIVDASKAVWKALG 173
CLN + I+ G V A VVDECDSR+GC + ++VD+S AVW+A+G
Sbjct: 62 CLNESHRKVRITGGGGAVEATVVDECDSRRGCKD----------DVVDSSPAVWRAMG 109
>gb|AAP54379.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|37535580|ref|NP_922092.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
gi|16905195|gb|AAL31065.1| hypothetical protein [Oryza
sativa]
Length = 61
Score = 45.1 bits (105), Expect = 0.001
Identities = 22/50 (44%), Positives = 31/50 (62%), Gaps = 10/50 (20%)
Query: 124 ITISANGRNVVAMVVDECDSRKGCDEQHDYQPPCPNNIVDASKAVWKALG 173
+ I+ G V A VVDECDSR+GC + ++VD+S AVW+A+G
Sbjct: 15 VRITGGGGAVEATVVDECDSRRGCKD----------DVVDSSPAVWRAMG 54
>ref|XP_466944.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|49388701|dbj|BAD25882.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
gi|49387976|dbj|BAD25084.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 328
Score = 42.0 bits (97), Expect = 0.009
Identities = 21/45 (46%), Positives = 25/45 (54%)
Query: 124 ITISANGRNVVAMVVDECDSRKGCDEQHDYQPPCPNNIVDASKAV 168
I I N V A+V+DECDSR GC+ Y PC N + AS V
Sbjct: 284 IQIYNNDNIVAALVIDECDSRNGCNLGTGYLLPCSPNTIAASPGV 328
>gb|EAK86428.1| hypothetical protein UM05495.1 [Ustilago maydis 521]
gi|49078784|ref|XP_403110.1| hypothetical protein
UM05495.1 [Ustilago maydis 521]
Length = 385
Score = 41.6 bits (96), Expect = 0.011
Identities = 31/98 (31%), Positives = 43/98 (43%), Gaps = 26/98 (26%)
Query: 100 HSDDTPVVALSTGWF----------NHKSRCLNNITISANGRNVVAMVVDECDSRKGCDE 149
+SDD P+VA+S F NH S C + IS G+ A DEC
Sbjct: 300 NSDDQPIVAISRDLFEKYNPSSGNPNHNSLCGKKVEISWKGKTAYAFATDEC-------- 351
Query: 150 QHDYQPPCPNNIVDASKAVWKALGVPKEQWGGLD-ITW 186
P C ++ +D S +V++ L + G LD ITW
Sbjct: 352 -----PTCTDHALDCSPSVFEQL--DNKDKGVLDGITW 382
>gb|EAK84317.1| hypothetical protein UM03212.1 [Ustilago maydis 521]
gi|49073190|ref|XP_400827.1| hypothetical protein
UM03212.1 [Ustilago maydis 521]
Length = 147
Score = 38.9 bits (89), Expect = 0.074
Identities = 30/113 (26%), Positives = 45/113 (39%), Gaps = 28/113 (24%)
Query: 71 STHTKAYLTLNSFQKGGDGGGPSECDKQYHSDDTPVVALSTGWFNHKSRCLNNITISANG 130
+ H + Y L G P C Q++SD P+VA+++ H C I I +NG
Sbjct: 53 ANHVRCYDVLEQ------NGNPGSCG-QWNSDSRPIVAVNSAQM-HDGLCGKPIWIQSNG 104
Query: 131 RNVVAMVVDECDSRKGCDEQHDYQPPCPNNIVDASKAVWKALGVPKEQWGGLD 183
+ + A + D C P C + +D S + EQ GLD
Sbjct: 105 KTIEATIADTC-------------PTCSSGSLDLSTGAF-------EQLSGLD 137
>gb|AAB62250.1| B2-aldehyde-forming enzyme [Schizophyllum commune]
Length = 200
Score = 37.7 bits (86), Expect = 0.17
Identities = 17/43 (39%), Positives = 27/43 (62%), Gaps = 3/43 (6%)
Query: 106 VVALSTGWFNHKSRCLNNITISANGRNVVAMVVDECDSRKGCD 148
+VAL+ G ++ + C ITI+ NG++ A +VDEC GC+
Sbjct: 14 IVALNQGMWDGGAHCFKMITITVNGKSTQAQIVDEC---PGCE 53
>gb|AAW42069.1| conserved hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21] gi|50258933|gb|EAL21614.1|
hypothetical protein CNBC6500 [Cryptococcus neoformans
var. neoformans B-3501A] gi|58264440|ref|XP_569376.1|
conserved hypothetical protein [Cryptococcus neoformans
var. neoformans JEC21]
Length = 317
Score = 35.8 bits (81), Expect = 0.63
Identities = 38/128 (29%), Positives = 54/128 (41%), Gaps = 31/128 (24%)
Query: 69 SVSTHTKAYLTLNSFQKGGDGGGPSECDKQYHSDDTPVVAL-STGWF------NHKSRCL 121
S +T Y T S GG EC HSDD V+A+ S GW+ + C
Sbjct: 207 SSEVYTGGYATFYS-----QGGVAGECGTV-HSDDDYVIAIDSNGWWQDYESNDSSPYCG 260
Query: 122 NNITI--SANGRNVVAMVVDECDSRKGCDEQHDYQPPCPN-NIVDASKAVWKALGVPKEQ 178
+IT+ + NG++V A+V D C P C N +D S + + E+
Sbjct: 261 KSITLTNTNNGKSVTAVVADVC-------------PTCETANSLDLSIGAFNQIAT--EE 305
Query: 179 WGGLDITW 186
G + ITW
Sbjct: 306 DGMVPITW 313
>emb|CAG79677.1| unnamed protein product [Yarrowia lipolytica CLIB99]
gi|50553346|ref|XP_504084.1| hypothetical protein
[Yarrowia lipolytica]
Length = 290
Score = 35.4 bits (80), Expect = 0.82
Identities = 21/73 (28%), Positives = 31/73 (41%), Gaps = 15/73 (20%)
Query: 115 NHKSRCLNNITISANGRNVVAMVVDECDSRKGCDEQHDYQPPCPNNIVDASKAVWKALGV 174
NH + C +T G++V V D C P C + +D S A + AL
Sbjct: 231 NHNTLCGKKLTAHRGGKSVTVTVTDMC-------------PGCASGDLDLSPAAFNALAS 277
Query: 175 PKEQWGGLDITWS 187
P E G + ++WS
Sbjct: 278 PSE--GRVGVSWS 288
>emb|CAG88385.1| unnamed protein product [Debaryomyces hansenii CBS767]
gi|50423073|ref|XP_460115.1| unnamed protein product
[Debaryomyces hansenii]
Length = 232
Score = 35.0 bits (79), Expect = 1.1
Identities = 37/137 (27%), Positives = 49/137 (35%), Gaps = 33/137 (24%)
Query: 67 SPSVSTHTKAYLTLNSFQKGGDGGGPSECDK-------QYHSDDTPVVALS--------- 110
SP+ T + T S GD G + + D +VA+S
Sbjct: 108 SPATGASTSSTSTGTSLPSSGDHSGEGTFYSTGLGACGETNQDTDYIVAVSHILYENNQV 167
Query: 111 TGWFNHKSRCLNNITISANGRNVVAMVVDECDSRKGCDEQH-DYQPPCPNNIVDASKAVW 169
G N S C I S G++V VVD C+ GC E D+ P + I D S
Sbjct: 168 NGNSNDNSLCGKKIKASYEGKSVEVTVVDSCE---GCAENDLDFSPSAFSQIADQS---- 220
Query: 170 KALGVPKEQWGGLDITW 186
G +DITW
Sbjct: 221 ---------LGRIDITW 228
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.318 0.134 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 369,247,072
Number of Sequences: 2540612
Number of extensions: 15821107
Number of successful extensions: 27869
Number of sequences better than 10.0: 29
Number of HSP's better than 10.0 without gapping: 15
Number of HSP's successfully gapped in prelim test: 14
Number of HSP's that attempted gapping in prelim test: 27830
Number of HSP's gapped (non-prelim): 30
length of query: 189
length of database: 863,360,394
effective HSP length: 121
effective length of query: 68
effective length of database: 555,946,342
effective search space: 37804351256
effective search space used: 37804351256
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 71 (32.0 bits)
Medicago: description of AC135311.8


