

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135311.2 + phase: 0
(164 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
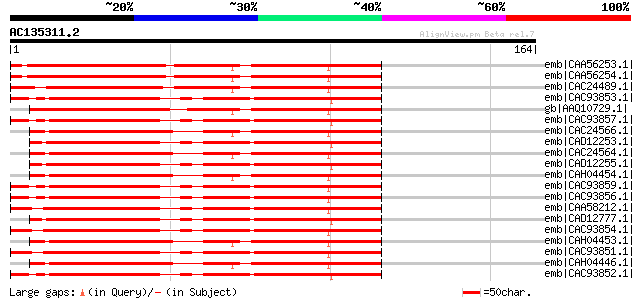
Score E
Sequences producing significant alignments: (bits) Value
emb|CAA56253.1| serine proteinase inhibitor [Medicago sativa] gi... 153 2e-36
emb|CAA56254.1| serine proteinase inhibitor [Medicago sativa] gi... 152 4e-36
emb|CAC24489.1| trypsin inhibitor [Pisum sativum] 141 6e-33
emb|CAC93853.1| trypsin/chymotrypsin inhibitor [Pisum sativum] 137 1e-31
gb|AAQ10729.1| trypsin inhibitor [Medicago scutellata] 137 1e-31
emb|CAC93857.1| trypsin/chymotrypsin inhibitor [Pisum sativum] 135 4e-31
emb|CAC24566.1| trypsin inhibitor [Pisum sativum] 135 4e-31
emb|CAD12253.1| Trypsin/chymotrypsin inhibitor [Lens culinaris] 132 4e-30
emb|CAC24564.1| trypsin inhibitor [Pisum sativum] 132 4e-30
emb|CAD12255.1| trypsin/chymotrypsin inhibitor [Lens ervoides] 131 8e-30
emb|CAH04454.1| putative trypsin inhibitor [Lens nigricans] 131 8e-30
emb|CAC93859.1| trypsin/chymotrypsin inhibitor [Pisum sativum] 130 1e-29
emb|CAC93856.1| trypsin/chymotrypsin inhibitor [Pisum sativum] 130 1e-29
emb|CAA58212.1| trypsin/chymotrypsin inhibitor [Pisum sativum] g... 130 2e-29
emb|CAD12777.1| trypsin/chymotrypsin inhibitor [Lens lamottei] g... 130 2e-29
emb|CAC93854.1| trypsin/chymotrypsin inhibitor [Pisum sativum] 130 2e-29
emb|CAH04453.1| putative trypsin inhibitor [Lens ervoides] 129 2e-29
emb|CAC93851.1| trypsin/chymotrypsin inhibitor [Pisum sativum] 129 2e-29
emb|CAH04446.1| putative trypsin inhibitor [Lens culinaris subsp... 129 3e-29
emb|CAC93852.1| trypsin/chymotrypsin inhibitor [Pisum sativum] 129 3e-29
>emb|CAA56253.1| serine proteinase inhibitor [Medicago sativa]
gi|2129858|pir||S56647 trypsin inhibitor precursor
(clone ATI18) - alfalfa
Length = 113
Score = 153 bits (386), Expect = 2e-36
Identities = 79/119 (66%), Positives = 90/119 (75%), Gaps = 9/119 (7%)
Query: 1 MELMNKNKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTAT 60
MELM NKKAM+MKLALLVFLLGFTSTVVDARFD SFI+Q++ NG++ NYD KST T
Sbjct: 1 MELM-MNKKAMMMKLALLVFLLGFTSTVVDARFDQASFISQLLFNGEAA--NYDVKSTTT 57
Query: 61 ACCNTCLC--SVTPELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCYKPCN 116
ACCN C C S+ P QCRC D+G+TCHSACK+C C S P C C DIT+FCY CN
Sbjct: 58 ACCNFCPCTRSIPP---QCRCTDIGETCHSACKSCLCTRSIPPQCRCTDITNFCYPKCN 113
>emb|CAA56254.1| serine proteinase inhibitor [Medicago sativa]
gi|1362056|pir||S56648 trypsin inhibitor precursor
(clone ATI21) - alfalfa
Length = 113
Score = 152 bits (383), Expect = 4e-36
Identities = 79/119 (66%), Positives = 90/119 (75%), Gaps = 9/119 (7%)
Query: 1 MELMNKNKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTAT 60
MELM NKKAM+MKLALLVFLLGFTST VDARFD SFITQ++ NG++ NYD KST T
Sbjct: 1 MELM-MNKKAMMMKLALLVFLLGFTSTGVDARFDRASFITQLLFNGEAA--NYDVKSTTT 57
Query: 61 ACCNTCLC--SVTPELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCYKPCN 116
ACCN C C S+ P QCRC+D+G+TCHSACK+C C S P C C DIT+FCY CN
Sbjct: 58 ACCNFCPCTKSIPP---QCRCSDIGETCHSACKSCICTRSYPPQCRCTDITNFCYPKCN 113
>emb|CAC24489.1| trypsin inhibitor [Pisum sativum]
Length = 110
Score = 141 bits (356), Expect = 6e-33
Identities = 73/119 (61%), Positives = 85/119 (71%), Gaps = 12/119 (10%)
Query: 1 MELMNKNKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTAT 60
MEL+N K +MKLAL+VFLLGFT+TVVDARFDSTSFITQ++SNGD+ Y KST T
Sbjct: 1 MELINTKK---MMKLALMVFLLGFTATVVDARFDSTSFITQLLSNGDA---GYSIKSTTT 54
Query: 61 ACCNTCLC--SVTPELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCYKPCN 116
ACC++C+C S+ P QC C DVGKTCHS C C C S P C C D DFCY+ CN
Sbjct: 55 ACCDSCICTKSIPP---QCHCTDVGKTCHSGCNLCLCTRSFPPQCHCTDTNDFCYQKCN 110
>emb|CAC93853.1| trypsin/chymotrypsin inhibitor [Pisum sativum]
Length = 104
Score = 137 bits (344), Expect = 1e-31
Identities = 74/117 (63%), Positives = 87/117 (74%), Gaps = 14/117 (11%)
Query: 1 MELMNKNKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTAT 60
MELMNK KAM MKLAL+VFLLGFT+ VV+AR DSTSFITQV+SNGD D KS
Sbjct: 1 MELMNK--KAM-MKLALMVFLLGFTANVVNARLDSTSFITQVLSNGD------DVKS--- 48
Query: 61 ACCNTCLCSVTPELTQCRCADVGKTCHSACKNCECGWSS-PLCTCYDITDFCYKPCN 116
ACC+TCLC+ + T CRCADVG+TCHSAC +C C +S+ P C C+D FCYK C+
Sbjct: 49 ACCDTCLCTASNPPT-CRCADVGETCHSACNSCICAYSNPPKCQCFDTHKFCYKACH 104
>gb|AAQ10729.1| trypsin inhibitor [Medicago scutellata]
Length = 110
Score = 137 bits (344), Expect = 1e-31
Identities = 70/113 (61%), Positives = 81/113 (70%), Gaps = 11/113 (9%)
Query: 7 NKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNTC 66
NKKAM+MKLALLVFLL F ST VDARFD SFITQ++SNG++ T KST TACC+ C
Sbjct: 6 NKKAMMMKLALLVFLLSFASTTVDARFDRASFITQLLSNGEANT-----KSTTTACCDFC 60
Query: 67 LC--SVTPELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCYKPCN 116
C S+ P QC+C DV + CHSACK+C C S P C CYDITDFCY C+
Sbjct: 61 PCTRSIPP---QCQCTDVREKCHSACKSCLCTRSFPPQCRCYDITDFCYPSCS 110
>emb|CAC93857.1| trypsin/chymotrypsin inhibitor [Pisum sativum]
Length = 104
Score = 135 bits (340), Expect = 4e-31
Identities = 73/117 (62%), Positives = 86/117 (73%), Gaps = 14/117 (11%)
Query: 1 MELMNKNKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTAT 60
MELMNK KAM MKLAL+VFLLGFT+ VV+AR DSTSFITQV+SNGD D KS
Sbjct: 1 MELMNK--KAM-MKLALMVFLLGFTANVVNARLDSTSFITQVLSNGD------DVKS--- 48
Query: 61 ACCNTCLCSVTPELTQCRCADVGKTCHSACKNCECGWSS-PLCTCYDITDFCYKPCN 116
ACC+TCLC+ + T CRC DVG+TCHSAC +C C +S+ P C C+D FCYK C+
Sbjct: 49 ACCDTCLCTASNPPT-CRCVDVGETCHSACNSCICAYSNPPKCQCFDTHKFCYKACH 104
>emb|CAC24566.1| trypsin inhibitor [Pisum sativum]
Length = 104
Score = 135 bits (340), Expect = 4e-31
Identities = 70/113 (61%), Positives = 84/113 (73%), Gaps = 16/113 (14%)
Query: 7 NKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNTC 66
NKKAM MKLAL++FLLGFT+TVVDARF STSFITQ++SNGD++ ACC++C
Sbjct: 5 NKKAM-MKLALMLFLLGFTATVVDARFGSTSFITQLLSNGDASNK---------ACCDSC 54
Query: 67 LC--SVTPELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCYKPCN 116
LC S+ P QC+C D+G+TCHSACK C C S P C+C DITDFCYK CN
Sbjct: 55 LCTRSIPP---QCQCNDIGETCHSACKACLCTRSLPPKCSCTDITDFCYKKCN 104
>emb|CAD12253.1| Trypsin/chymotrypsin inhibitor [Lens culinaris]
Length = 104
Score = 132 bits (331), Expect = 4e-30
Identities = 68/111 (61%), Positives = 82/111 (73%), Gaps = 12/111 (10%)
Query: 7 NKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNTC 66
NKKA +MKLAL++FLLGFT+ VVDARFDSTSFITQV+SNGD D KS ACC+TC
Sbjct: 5 NKKA-IMKLALMLFLLGFTANVVDARFDSTSFITQVLSNGD------DVKS---ACCDTC 54
Query: 67 LCSVTPELTQCRCADVGKTCHSACKNCECGWSS-PLCTCYDITDFCYKPCN 116
LC+ + T CRC DV ++CHSAC C C +S+ P C CYD +FCYK C+
Sbjct: 55 LCTRSQPPT-CRCVDVRESCHSACDKCVCAYSNPPQCQCYDTHNFCYKTCH 104
>emb|CAC24564.1| trypsin inhibitor [Pisum sativum]
Length = 104
Score = 132 bits (331), Expect = 4e-30
Identities = 68/113 (60%), Positives = 82/113 (72%), Gaps = 16/113 (14%)
Query: 7 NKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNTC 66
NKKAM MKLAL++FLLGFT+TVVDARFDS SFITQ++SNG ++ ACC++C
Sbjct: 5 NKKAM-MKLALVLFLLGFTATVVDARFDSNSFITQLLSNGGASNK---------ACCDSC 54
Query: 67 LC--SVTPELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCYKPCN 116
LC S+ P +CRC D+G+TCHSACK C C S P C C DITDFCY+ CN
Sbjct: 55 LCTRSIPP---RCRCNDIGETCHSACKTCICTRSLPPQCRCIDITDFCYEKCN 104
>emb|CAD12255.1| trypsin/chymotrypsin inhibitor [Lens ervoides]
Length = 104
Score = 131 bits (329), Expect = 8e-30
Identities = 68/111 (61%), Positives = 82/111 (73%), Gaps = 12/111 (10%)
Query: 7 NKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNTC 66
NKKA +MKLAL+VFLLGFT+ VV+ARFDSTSFITQV+SNGD D KS ACC+TC
Sbjct: 5 NKKA-IMKLALMVFLLGFTANVVNARFDSTSFITQVLSNGD------DVKS---ACCDTC 54
Query: 67 LCSVTPELTQCRCADVGKTCHSACKNCECGWSS-PLCTCYDITDFCYKPCN 116
LC+ + T CRC DV ++CHSAC C C +S+ P C CYD +FCYK C+
Sbjct: 55 LCTRSQPPT-CRCVDVRESCHSACDKCVCAYSNPPQCQCYDTHNFCYKTCH 104
>emb|CAH04454.1| putative trypsin inhibitor [Lens nigricans]
Length = 104
Score = 131 bits (329), Expect = 8e-30
Identities = 67/113 (59%), Positives = 81/113 (71%), Gaps = 16/113 (14%)
Query: 7 NKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNTC 66
NKK M MKLAL++FLLGFT+TVVDARFDST FITQ+ SNGD++ ACCN+C
Sbjct: 5 NKKTM-MKLALMLFLLGFTATVVDARFDSTFFITQLFSNGDASNK---------ACCNSC 54
Query: 67 LC--SVTPELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCYKPCN 116
C S+ P +CRC D+G+TCHSACK+C C S P C C D+T+FCYK CN
Sbjct: 55 PCTRSIPP---KCRCTDIGETCHSACKSCLCTRSIPPQCRCTDVTNFCYKNCN 104
>emb|CAC93859.1| trypsin/chymotrypsin inhibitor [Pisum sativum]
Length = 114
Score = 130 bits (328), Expect = 1e-29
Identities = 72/117 (61%), Positives = 83/117 (70%), Gaps = 14/117 (11%)
Query: 1 MELMNKNKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTAT 60
MELMNK KAM MKLAL+VFLL F + VV+ARFDSTSFITQV+SNGD D KS
Sbjct: 1 MELMNK--KAM-MKLALMVFLLSFAANVVNARFDSTSFITQVLSNGD------DVKS--- 48
Query: 61 ACCNTCLCSVTPELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCYKPCN 116
ACC+TCLC+ + T CRC DVG+TCHSAC +C C S P C C+D FCYK C+
Sbjct: 49 ACCDTCLCTKSNPPT-CRCVDVGETCHSACNSCICALSYPPKCQCFDTQKFCYKACH 104
>emb|CAC93856.1| trypsin/chymotrypsin inhibitor [Pisum sativum]
Length = 104
Score = 130 bits (328), Expect = 1e-29
Identities = 72/117 (61%), Positives = 83/117 (70%), Gaps = 14/117 (11%)
Query: 1 MELMNKNKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTAT 60
MELMNK KAM MKLAL+VFLL F + VV+ARFDSTSFITQV+SNGD D KS
Sbjct: 1 MELMNK--KAM-MKLALMVFLLSFAANVVNARFDSTSFITQVLSNGD------DVKS--- 48
Query: 61 ACCNTCLCSVTPELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCYKPCN 116
ACC+TCLC+ + T CRC DVG+TCHSAC +C C S P C C+D FCYK C+
Sbjct: 49 ACCDTCLCTKSNPPT-CRCVDVGETCHSACNSCICALSYPPKCQCFDTQKFCYKACH 104
>emb|CAA58212.1| trypsin/chymotrypsin inhibitor [Pisum sativum]
gi|6016305|sp|Q41066|IBB2_PEA Seed trypsin/chymotrypsin
inhibitor TI5-72 precursor
Length = 114
Score = 130 bits (326), Expect = 2e-29
Identities = 69/117 (58%), Positives = 82/117 (69%), Gaps = 14/117 (11%)
Query: 1 MELMNKNKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTAT 60
MELMNK ++MKLAL+VFLL F + VV+ARFDSTSFITQV+SNGD D KS
Sbjct: 1 MELMNKK---VMMKLALMVFLLSFAANVVNARFDSTSFITQVLSNGD------DVKS--- 48
Query: 61 ACCNTCLCSVTPELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCYKPCN 116
ACC+TCLC+ + T CRC DVG+TCHSAC +C C S P C C+D FCYK C+
Sbjct: 49 ACCDTCLCTKSDPPT-CRCVDVGETCHSACDSCICALSYPPQCQCFDTHKFCYKACH 104
>emb|CAD12777.1| trypsin/chymotrypsin inhibitor [Lens lamottei]
gi|17148535|emb|CAD12776.1| trypsin/chymotrypsin
inhibitor [Lens odemensis] gi|17059628|emb|CAD12252.1|
trypsin/chymotrypsin inhibitor [Lens culinaris]
Length = 104
Score = 130 bits (326), Expect = 2e-29
Identities = 67/111 (60%), Positives = 82/111 (73%), Gaps = 12/111 (10%)
Query: 7 NKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNTC 66
NKKA +MKLAL++FLLGFT+ VV+ARFDSTSFITQV+SNGD D KS ACC+TC
Sbjct: 5 NKKA-IMKLALMLFLLGFTANVVNARFDSTSFITQVLSNGD------DVKS---ACCDTC 54
Query: 67 LCSVTPELTQCRCADVGKTCHSACKNCECGWSS-PLCTCYDITDFCYKPCN 116
LC+ + T CRC DV ++CHSAC C C +S+ P C CYD +FCYK C+
Sbjct: 55 LCTRSQPPT-CRCVDVRESCHSACDKCVCAYSNPPQCQCYDTHNFCYKTCH 104
>emb|CAC93854.1| trypsin/chymotrypsin inhibitor [Pisum sativum]
Length = 104
Score = 130 bits (326), Expect = 2e-29
Identities = 69/117 (58%), Positives = 82/117 (69%), Gaps = 14/117 (11%)
Query: 1 MELMNKNKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTAT 60
MELMNK ++MKLAL+VFLL F + VV+ARFDSTSFITQV+SNGD D KS
Sbjct: 1 MELMNKK---VMMKLALMVFLLSFAANVVNARFDSTSFITQVLSNGD------DVKS--- 48
Query: 61 ACCNTCLCSVTPELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCYKPCN 116
ACC+TCLC+ + T CRC DVG+TCHSAC +C C S P C C+D FCYK C+
Sbjct: 49 ACCDTCLCTKSDPPT-CRCVDVGETCHSACDSCICALSYPPQCQCFDTHKFCYKACH 104
>emb|CAH04453.1| putative trypsin inhibitor [Lens ervoides]
Length = 104
Score = 129 bits (325), Expect = 2e-29
Identities = 66/113 (58%), Positives = 81/113 (71%), Gaps = 16/113 (14%)
Query: 7 NKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNTC 66
NKK M MKLAL++FLLGFT+TVVDARFDST FITQ+ SNGD++ ACCN+C
Sbjct: 5 NKKTM-MKLALMLFLLGFTATVVDARFDSTFFITQLFSNGDASNK---------ACCNSC 54
Query: 67 LC--SVTPELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCYKPCN 116
C S+ P +C C+D+G+TCHSACK+C C S P C C D+T+FCYK CN
Sbjct: 55 PCTRSIPP---KCSCSDIGETCHSACKSCLCTRSIPPQCRCTDVTNFCYKKCN 104
>emb|CAC93851.1| trypsin/chymotrypsin inhibitor [Pisum sativum]
Length = 104
Score = 129 bits (325), Expect = 2e-29
Identities = 69/117 (58%), Positives = 83/117 (69%), Gaps = 14/117 (11%)
Query: 1 MELMNKNKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTAT 60
MELMNK ++MKLAL+VFLLGFT+ VV+AR DSTSFITQV+SNG+ D KS
Sbjct: 1 MELMNKK---VMMKLALMVFLLGFTANVVNARLDSTSFITQVLSNGN------DVKS--- 48
Query: 61 ACCNTCLCSVTPELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCYKPCN 116
ACC+TCLC+ + T CRC DVG+TCHSAC +C C S P C C+D FCYK C+
Sbjct: 49 ACCDTCLCTKSNPPT-CRCVDVGETCHSACDSCICALSYPPKCQCFDTHKFCYKACH 104
>emb|CAH04446.1| putative trypsin inhibitor [Lens culinaris subsp. orientalis]
Length = 104
Score = 129 bits (324), Expect = 3e-29
Identities = 66/113 (58%), Positives = 81/113 (71%), Gaps = 16/113 (14%)
Query: 7 NKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNTC 66
NKK M MKLAL++FLLGFT+TVVDARFDST FITQ+ SNGD++ ACCN+C
Sbjct: 5 NKKTM-MKLALMLFLLGFTATVVDARFDSTFFITQLFSNGDASNK---------ACCNSC 54
Query: 67 LC--SVTPELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCYKPCN 116
C S+ P +C C+D+G+TCHSACK+C C S P C C D+T+FCYK CN
Sbjct: 55 PCTRSIPP---KCSCSDIGETCHSACKSCLCTRSIPPQCRCTDVTNFCYKNCN 104
>emb|CAC93852.1| trypsin/chymotrypsin inhibitor [Pisum sativum]
Length = 104
Score = 129 bits (324), Expect = 3e-29
Identities = 71/117 (60%), Positives = 84/117 (71%), Gaps = 14/117 (11%)
Query: 1 MELMNKNKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTAT 60
MELMNK KAM MKLAL+VFLL F + VV+ARFDSTSFITQV+SNGD D KS
Sbjct: 1 MELMNK--KAM-MKLALMVFLLSFAANVVNARFDSTSFITQVLSNGD------DVKS--- 48
Query: 61 ACCNTCLCSVTPELTQCRCADVGKTCHSACKNCECGWSS-PLCTCYDITDFCYKPCN 116
ACC+TCLC+ + T CRC DV +TCHSAC +C C +S+ P C C+D FCYK C+
Sbjct: 49 ACCDTCLCTKSNPPT-CRCVDVRETCHSACDSCICAYSNPPKCQCFDTHKFCYKACH 104
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.325 0.131 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 253,088,000
Number of Sequences: 2540612
Number of extensions: 9184155
Number of successful extensions: 35635
Number of sequences better than 10.0: 352
Number of HSP's better than 10.0 without gapping: 88
Number of HSP's successfully gapped in prelim test: 266
Number of HSP's that attempted gapping in prelim test: 34782
Number of HSP's gapped (non-prelim): 702
length of query: 164
length of database: 863,360,394
effective HSP length: 118
effective length of query: 46
effective length of database: 563,568,178
effective search space: 25924136188
effective search space used: 25924136188
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 70 (31.6 bits)
Medicago: description of AC135311.2


