

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135234.5 - phase: 0 /pseudo
(256 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
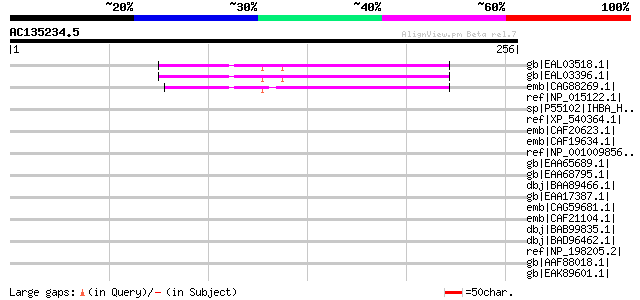
Score E
Sequences producing significant alignments: (bits) Value
gb|EAL03518.1| hypothetical protein CaO19.12424 [Candida albican... 60 8e-08
gb|EAL03396.1| hypothetical protein CaO19.4959 [Candida albicans... 60 8e-08
emb|CAG88269.1| unnamed protein product [Debaryomyces hansenii C... 57 7e-07
ref|NP_015122.1| Iron-regulated transcriptional activator, requi... 41 0.037
sp|P55102|IHBA_HORSE Inhibin beta A chain precursor (Activin bet... 39 0.18
ref|XP_540364.1| PREDICTED: similar to activin beta A subunit pr... 37 0.41
emb|CAF20623.1| conserved hypothetical protein [Corynebacterium ... 37 0.41
emb|CAF19634.1| conserved hypothetical protein [Corynebacterium ... 37 0.41
ref|NP_001009856.1| inhibin beta A [Felis catus] gi|32455208|gb|... 37 0.53
gb|EAA65689.1| hypothetical protein AN0859.2 [Aspergillus nidula... 37 0.53
gb|EAA68795.1| predicted protein [Gibberella zeae PH-1] gi|46111... 36 1.2
dbj|BAA89466.1| gag-pol polyprotein [Oryza sativa (indica cultiv... 36 1.2
gb|EAA17387.1| hypothetical protein [Plasmodium yoelii yoelii] 36 1.2
emb|CAG59681.1| unnamed protein product [Candida glabrata CBS138... 35 1.6
emb|CAF21104.1| conserved hypothetical protein [Corynebacterium ... 35 1.6
dbj|BAB99835.1| Hypothetical protein [Corynebacterium glutamicum... 35 1.6
dbj|BAD96462.1| inhibin beta A subunit precursor variant [Homo s... 35 2.0
ref|NP_198205.2| far-red impaired responsive protein, putative [... 35 2.0
gb|AAF88018.1| contains simlarity to Arabidopsis thaliana far-re... 35 2.0
gb|EAK89601.1| hypothetical protein, signal peptide [Cryptospori... 35 2.6
>gb|EAL03518.1| hypothetical protein CaO19.12424 [Candida albicans SC5314]
Length = 573
Score = 59.7 bits (143), Expect = 8e-08
Identities = 46/159 (28%), Positives = 67/159 (41%), Gaps = 14/159 (8%)
Query: 76 WREREELLTWVRRQGVKAGFGVCIDKSVLKRPYLTTQSERSGIYKPPKTRK----KPNLE 131
+ R+EL ++ GFGV I S K Y T E G Y+ K +K K
Sbjct: 211 FNSRDELNEFIAEFARDNGFGVVIAHSNKKAIYYTC--ELGGRYRHKKNKKIDVTKQIDV 268
Query: 132 GIGSM--------KCNCPFRLKGFFDKDTNDWWLAILCGMHNHDLEEKLQGHLIAGRLSA 183
G G M K CPF + + K N W L C HNH + L H + + S
Sbjct: 269 GDGYMLDPDTKTKKLKCPFAMTASYKKSANAWTLRTTCNEHNHPQLDPLSNHPMLRKRSD 328
Query: 184 EEKKKVIDMTKSLTVPRNILTNLKENNKESVTTIKQVYN 222
E +++M K T P +I + +KE + + + +YN
Sbjct: 329 ELNVMILEMYKLGTKPSHIESKIKEEYPDVLIKREDIYN 367
>gb|EAL03396.1| hypothetical protein CaO19.4959 [Candida albicans SC5314]
Length = 573
Score = 59.7 bits (143), Expect = 8e-08
Identities = 46/159 (28%), Positives = 67/159 (41%), Gaps = 14/159 (8%)
Query: 76 WREREELLTWVRRQGVKAGFGVCIDKSVLKRPYLTTQSERSGIYKPPKTRK----KPNLE 131
+ R+EL ++ GFGV I S K Y T E G Y+ K +K K
Sbjct: 211 FNSRDELNEFIAEFARDNGFGVVIAHSNKKAIYYTC--ELGGRYRHKKNKKIDVTKQIDV 268
Query: 132 GIGSM--------KCNCPFRLKGFFDKDTNDWWLAILCGMHNHDLEEKLQGHLIAGRLSA 183
G G M K CPF + + K N W L C HNH + L H + + S
Sbjct: 269 GDGYMLDPDTKTKKLKCPFAMTASYKKSANAWTLRTTCNEHNHPQLDPLSNHPMLRKRSD 328
Query: 184 EEKKKVIDMTKSLTVPRNILTNLKENNKESVTTIKQVYN 222
E +++M K T P +I + +KE + + + +YN
Sbjct: 329 ELNVMILEMYKLGTKPSHIESKIKEEYPDVLIKREDIYN 367
>emb|CAG88269.1| unnamed protein product [Debaryomyces hansenii CBS767]
gi|50422877|ref|XP_460016.1| unnamed protein product
[Debaryomyces hansenii]
Length = 481
Score = 56.6 bits (135), Expect = 7e-07
Identities = 42/160 (26%), Positives = 67/160 (41%), Gaps = 21/160 (13%)
Query: 79 REELLTWVRRQGVKAGFGVCIDKSVLKRPYLTTQSERSGIYKPPKTRK------------ 126
R++L +++ GFGV I S K Y T E G Y+ K++K
Sbjct: 165 RDDLNEFIQEFARDNGFGVVIAHSNKKAIYYTC--ELGGRYRQKKSKKGMEDARHLEVDN 222
Query: 127 ----KPNLEGIGSMKCNCPFRLKGFFDKDTNDWWLAILCGMHNHDLEEKLQGHLIAGRLS 182
PN + + K CPF + + K T W L C HNH + L H + + S
Sbjct: 223 GYILDPNTK---TKKLRCPFSMTATYKKSTGMWTLKTTCNEHNHPQLDPLSNHPMLRKRS 279
Query: 183 AEEKKKVIDMTKSLTVPRNILTNLKENNKESVTTIKQVYN 222
E ++D+ K T P +I +K+ + + + +YN
Sbjct: 280 DELNLLILDLYKVGTKPSHIEAKIKDQYPDVLIKREDIYN 319
>ref|NP_015122.1| Iron-regulated transcriptional activator, required for iron
homeostasis and resistance to oxidative stress; similar
to Aft1p; Aft2p [Saccharomyces cerevisiae]
gi|1370420|emb|CAA97916.1| unnamed protein product
[Saccharomyces cerevisiae] gi|2132231|pir||S65221
hypothetical protein YPL202c - yeast (Saccharomyces
cerevisiae)
Length = 416
Score = 40.8 bits (94), Expect = 0.037
Identities = 21/93 (22%), Positives = 44/93 (46%), Gaps = 6/93 (6%)
Query: 76 WREREELLTWVRRQGVKAGFGVCIDKSVLKRPYLTTQSERSGIYKPPKTRKKPNLEGIGS 135
+ +R E+ W+++ G + I++S + +S RS + PK++ G S
Sbjct: 51 FEDRHEIKPWLQKIFYPQGIDIVIERSDSSKVTFKCRSVRSKVGLNPKSK------GSSS 104
Query: 136 MKCNCPFRLKGFFDKDTNDWWLAILCGMHNHDL 168
CPFR++ + W + ++ +H+H+L
Sbjct: 105 RSHACPFRIRAAYSVRLQKWNVVVMNNIHSHEL 137
>sp|P55102|IHBA_HORSE Inhibin beta A chain precursor (Activin beta-A chain)
gi|786171|dbj|BAA08862.1| inhibin beta A subunit [Equus
caballus]
Length = 426
Score = 38.5 bits (88), Expect = 0.18
Identities = 34/140 (24%), Positives = 66/140 (46%), Gaps = 7/140 (5%)
Query: 41 PMQARIGKIVEPKR--VPIGVVCE-ARDTASFFTTNARWREREELLTWVRRQGVKAGFGV 97
P+ + I ++++ + + I + C+ +T + + +++EE ++ G +AG GV
Sbjct: 222 PVSSSIQRLLDQGKSSLDIRIACDQCHETGASLVLLGKKKKKEEEGEGKKKDGGEAGAGV 281
Query: 98 CIDKSVLKRPYLTTQSERSGIYKPPKTRKKPNLEGIGSMKCNCPFR-LKGFFDKDTNDWW 156
+K RP+L Q+ +S + P R++ LE G + C + F D NDW
Sbjct: 282 DEEKEQSHRPFLMLQARQSEDH--PHRRRRRGLECDGKVNICCKKQFFVSFKDIGWNDWI 339
Query: 157 LAILCGMHNHDLEEKLQGHL 176
+A G H + E + H+
Sbjct: 340 IA-PSGYHANYCEGECPSHI 358
>ref|XP_540364.1| PREDICTED: similar to activin beta A subunit precursor [Canis
familiaris]
Length = 547
Score = 37.4 bits (85), Expect = 0.41
Identities = 35/140 (25%), Positives = 63/140 (45%), Gaps = 7/140 (5%)
Query: 41 PMQARIGKIVEPKR--VPIGVVCE-ARDTASFFTTNARWREREELLTWVRRQGVKAGFGV 97
P+ + I ++++ R + + + CE +T + + +++EE ++ G AG G
Sbjct: 343 PVSSSIQRLLDQGRSSLDVRIACEQCHETGASLVLLGKKKKKEEEGEGKKKDGGDAGAGG 402
Query: 98 CIDKSVLKRPYLTTQSERSGIYKPPKTRKKPNLEGIGSMKCNCPFR-LKGFFDKDTNDWW 156
DK RP+L Q+ +S + P R++ LE G + C + F D NDW
Sbjct: 403 DEDKEQSHRPFLMLQARQSEDH--PHRRRRRGLECDGKVNICCKKQFFVSFKDIGWNDWI 460
Query: 157 LAILCGMHNHDLEEKLQGHL 176
+A G H + E H+
Sbjct: 461 IA-PSGYHANYCEGGCPSHI 479
>emb|CAF20623.1| conserved hypothetical protein [Corynebacterium glutamicum ATCC
13032] gi|21325052|dbj|BAB99674.1| Hypothetical protein
[Corynebacterium glutamicum ATCC 13032]
gi|62391122|ref|YP_226524.1| hypothetical protein cg2504
[Corynebacterium glutamicum ATCC 13032]
gi|19553479|ref|NP_601481.1| hypothetical protein
NCgl2201 [Corynebacterium glutamicum ATCC 13032]
Length = 517
Score = 37.4 bits (85), Expect = 0.41
Identities = 31/108 (28%), Positives = 47/108 (42%), Gaps = 8/108 (7%)
Query: 3 DNVADDDVEPKPAKTEASGEGSGDVKPTINLKPFKVQIPMQARIGKIVEPKRVPIGVVCE 62
D+ DDD +P+P K E D KP + KP + +I + E R+
Sbjct: 132 DDSGDDDPDPEPDKPE-------DGKPDSD-KPRRPRISAEKHAIITDELARLNPNTTPS 183
Query: 63 ARDTASFFTTNARWREREELLTWVRRQGVKAGFGVCIDKSVLKRPYLT 110
A + + + A WR E+L TW+R Q A + +KR YL+
Sbjct: 184 AEELRTQALSQAIWRTPEDLRTWLRHQVTTANKNNPNPITAMKRRYLS 231
>emb|CAF19634.1| conserved hypothetical protein [Corynebacterium glutamicum ATCC
13032] gi|21323694|dbj|BAB98321.1| Hypothetical protein
[Corynebacterium glutamicum ATCC 13032]
gi|62389818|ref|YP_225220.1| hypothetical protein cg1059
[Corynebacterium glutamicum ATCC 13032]
gi|19552154|ref|NP_600156.1| hypothetical protein
NCgl0891 [Corynebacterium glutamicum ATCC 13032]
Length = 517
Score = 37.4 bits (85), Expect = 0.41
Identities = 31/108 (28%), Positives = 47/108 (42%), Gaps = 8/108 (7%)
Query: 3 DNVADDDVEPKPAKTEASGEGSGDVKPTINLKPFKVQIPMQARIGKIVEPKRVPIGVVCE 62
D+ DDD +P+P K E D KP + KP + +I + E R+
Sbjct: 132 DDSGDDDPDPEPDKPE-------DGKPDSD-KPRRPRISAEKHAIITDELARLNPNTTPS 183
Query: 63 ARDTASFFTTNARWREREELLTWVRRQGVKAGFGVCIDKSVLKRPYLT 110
A + + + A WR E+L TW+R Q A + +KR YL+
Sbjct: 184 AEELRNQALSQAIWRTPEDLRTWLRHQVTTANKNNPNPITAMKRRYLS 231
>ref|NP_001009856.1| inhibin beta A [Felis catus] gi|32455208|gb|AAP83318.1| activin
beta A subunit precursor [Felis catus]
Length = 424
Score = 37.0 bits (84), Expect = 0.53
Identities = 33/140 (23%), Positives = 63/140 (44%), Gaps = 7/140 (5%)
Query: 41 PMQARIGKIVEPKR--VPIGVVCE-ARDTASFFTTNARWREREELLTWVRRQGVKAGFGV 97
P+ + I ++++ + + + + CE +T + + +++EE ++ G G G
Sbjct: 220 PVSSSIQRLLDQGKSSLDVRIACEQCHETGASLVLLGKKKKKEEEGEGKKKDGGDGGAGA 279
Query: 98 CIDKSVLKRPYLTTQSERSGIYKPPKTRKKPNLEGIGSMKCNCPFR-LKGFFDKDTNDWW 156
DK RP+L Q+ +S + P R++ LE G + C + F D NDW
Sbjct: 280 DEDKEQSHRPFLMLQARQSEDH--PHRRRRRGLECDGKVNICCKKQFFVSFKDIGWNDWI 337
Query: 157 LAILCGMHNHDLEEKLQGHL 176
+A G H + E + H+
Sbjct: 338 IA-PSGYHANYCEGECPSHI 356
>gb|EAA65689.1| hypothetical protein AN0859.2 [Aspergillus nidulans FGSC A4]
gi|67517161|ref|XP_658463.1| hypothetical protein
AN0859_2 [Aspergillus nidulans FGSC A4]
gi|49085806|ref|XP_404996.1| hypothetical protein
AN0859.2 [Aspergillus nidulans FGSC A4]
Length = 404
Score = 37.0 bits (84), Expect = 0.53
Identities = 25/98 (25%), Positives = 43/98 (43%), Gaps = 2/98 (2%)
Query: 135 SMKCNCPFRLKGFFDKDTNDWWLAILCGMHNHDLEEKLQGHLIAGRLSAEEKKKVIDMTK 194
S + CPF ++ K N W + + HNH + H R +K + I
Sbjct: 234 SRRTGCPFSIRISHHKTDNLWHVKVRDPSHNHG-PSRPGSHPSIRREEISQKTQYIKALL 292
Query: 195 SLTV-PRNILTNLKENNKESVTTIKQVYNVRTRWCKGE 231
+ + P IL L+ + +V I+ +YN+R R +G+
Sbjct: 293 DINMPPSQILKQLRHVDPTTVIQIRDIYNLRHRIYQGK 330
>gb|EAA68795.1| predicted protein [Gibberella zeae PH-1]
gi|46111341|ref|XP_382728.1| predicted protein
[Gibberella zeae PH-1]
Length = 392
Score = 35.8 bits (81), Expect = 1.2
Identities = 30/102 (29%), Positives = 42/102 (40%), Gaps = 8/102 (7%)
Query: 23 GSGDVKPTINLKPFKVQIPMQARIGKIVEPKRV--PIGVVCEARDTASFFTTNARWRERE 80
GS PT L P +A+ + P + +GV E RD + TT R RE
Sbjct: 21 GSNKRPPTTKLTALAKHRP-RAKFNLLDLPNELIYAVGVAAERRDARALSTTCRRLRENL 79
Query: 81 ELLTWVRRQGVKAGFGVCIDKSVLKR--PYLTTQSERSGIYK 120
+ W R VK G+C+ S + YL E+ G+ K
Sbjct: 80 ASIVWSR---VKLSIGLCVTSSDIDTFVAYLEEHCEKFGLIK 118
>dbj|BAA89466.1| gag-pol polyprotein [Oryza sativa (indica cultivar-group)]
Length = 1587
Score = 35.8 bits (81), Expect = 1.2
Identities = 23/72 (31%), Positives = 34/72 (46%), Gaps = 2/72 (2%)
Query: 100 DKSVLKRPYLTTQSERSGIYKPPKTRKKPNLEGIGSMKCNCPFRLKGFFDKDTN-DWWLA 158
D+ VL RP + +R G+ PP+ + N + +G +K P G +D D W LA
Sbjct: 119 DEEVLGRPQRQQRRQRHGMGAPPRREVRDNDDSLGKIKFTIPC-FDGKYDPDAYLTWELA 177
Query: 159 ILCGMHNHDLEE 170
I+ HD E
Sbjct: 178 IVQKFACHDFPE 189
>gb|EAA17387.1| hypothetical protein [Plasmodium yoelii yoelii]
Length = 1730
Score = 35.8 bits (81), Expect = 1.2
Identities = 16/46 (34%), Positives = 28/46 (60%)
Query: 185 EKKKVIDMTKSLTVPRNILTNLKENNKESVTTIKQVYNVRTRWCKG 230
+KKK+ID L + + NL +NNKE++ IK++ N+ ++ G
Sbjct: 968 DKKKLIDKKNELDYIKEMYLNLYKNNKENIKIIKKLKNIILKYTDG 1013
>emb|CAG59681.1| unnamed protein product [Candida glabrata CBS138]
gi|50288649|ref|XP_446754.1| unnamed protein product
[Candida glabrata]
Length = 437
Score = 35.4 bits (80), Expect = 1.6
Identities = 19/94 (20%), Positives = 42/94 (44%), Gaps = 3/94 (3%)
Query: 76 WREREELLTWVRRQGVKAGFGVCIDKSVLKRPYLTTQSERSGIYKPPKTRKKPNLEGIGS 135
++E+ ++ W++ G + I++S + + ++P R+K E +
Sbjct: 42 FQEKIDVKVWLQEMFFPMGIDIVIERSDKTKIIFKCKPSDYRSHEPKDLRQK---EPVDD 98
Query: 136 MKCNCPFRLKGFFDKDTNDWWLAILCGMHNHDLE 169
+ CPFR++ F W + I+ H+H L+
Sbjct: 99 KRHICPFRVRCTFSTQLKKWKIVIINNSHSHPLK 132
>emb|CAF21104.1| conserved hypothetical protein [Corynebacterium glutamicum ATCC
13032] gi|62391282|ref|YP_226684.1| hypothetical protein
cg2683 [Corynebacterium glutamicum ATCC 13032]
gi|19553638|ref|NP_601640.1| hypothetical protein
NCgl2356 [Corynebacterium glutamicum ATCC 13032]
Length = 517
Score = 35.4 bits (80), Expect = 1.6
Identities = 30/109 (27%), Positives = 45/109 (40%), Gaps = 12/109 (11%)
Query: 3 DNVADDDVEPKPAKTEASGEGSGDVK--PTINLKPFKVQIPMQARIGKIVEPKRVPIGVV 60
D+ DDD +P+P K E G+ GD P I+ + + AR+ P
Sbjct: 132 DDSGDDDPDPEPDKPE-DGKPDGDKPRGPRISAEKHAIITDELARLNPNTTPS------- 183
Query: 61 CEARDTASFFTTNARWREREELLTWVRRQGVKAGFGVCIDKSVLKRPYL 109
A + + + A WR E+L TW+R A + +KR YL
Sbjct: 184 --AEELRTQALSQAIWRTPEDLRTWLRHHVTTANKNNPNPITAMKRRYL 230
>dbj|BAB99835.1| Hypothetical protein [Corynebacterium glutamicum ATCC 13032]
Length = 528
Score = 35.4 bits (80), Expect = 1.6
Identities = 30/109 (27%), Positives = 45/109 (40%), Gaps = 12/109 (11%)
Query: 3 DNVADDDVEPKPAKTEASGEGSGDVK--PTINLKPFKVQIPMQARIGKIVEPKRVPIGVV 60
D+ DDD +P+P K E G+ GD P I+ + + AR+ P
Sbjct: 143 DDSGDDDPDPEPDKPE-DGKPDGDKPRGPRISAEKHAIITDELARLNPNTTPS------- 194
Query: 61 CEARDTASFFTTNARWREREELLTWVRRQGVKAGFGVCIDKSVLKRPYL 109
A + + + A WR E+L TW+R A + +KR YL
Sbjct: 195 --AEELRTQALSQAIWRTPEDLRTWLRHHVTTANKNNPNPITAMKRRYL 241
>dbj|BAD96462.1| inhibin beta A subunit precursor variant [Homo sapiens]
Length = 426
Score = 35.0 bits (79), Expect = 2.0
Identities = 31/140 (22%), Positives = 65/140 (46%), Gaps = 7/140 (5%)
Query: 41 PMQARIGKIVEPKR--VPIGVVCE-ARDTASFFTTNARWREREELLTWVRRQGVKAGFGV 97
P+ + I ++++ + + + + CE +++ + + +++EE ++ G + G G
Sbjct: 222 PVSSSIQRLLDQGKSSLDVRIACEQCQESGASLVLLGKKKKKEEEGEGKKKGGGEGGAGA 281
Query: 98 CIDKSVLKRPYLTTQSERSGIYKPPKTRKKPNLEGIGSMKCNCPFR-LKGFFDKDTNDWW 156
+K RP+L Q+ +S + P R++ LE G + C + F D NDW
Sbjct: 282 DEEKEQSHRPFLMLQARQSEDH--PHRRRRRGLECDGKVNICCKKQFFVSFKDIGWNDWI 339
Query: 157 LAILCGMHNHDLEEKLQGHL 176
+A G H + E + H+
Sbjct: 340 IA-PSGYHANYCESECPSHI 358
>ref|NP_198205.2| far-red impaired responsive protein, putative [Arabidopsis
thaliana]
Length = 685
Score = 35.0 bits (79), Expect = 2.0
Identities = 34/142 (23%), Positives = 67/142 (46%), Gaps = 29/142 (20%)
Query: 92 KAGFGVCIDKSVLKRPYLTTQSERSGIYKPP---------KTRKKPNLEGIG---SMKCN 139
K+GF + R +T+S+ G+Y+ + RKK N+E S++C
Sbjct: 77 KSGFSI--------RKARSTESQNLGVYRRDFVCYRSGFNQPRKKANVEHPRERKSVRCG 128
Query: 140 CPFRLKGFFDKDTND----WWLAILCGMHNHDLEEKLQGHLIAG--RLSAEEKKKVIDMT 193
C +L + K+ D W+++ +HNH+L E Q L+ ++ ++++++ ++
Sbjct: 129 CDGKL--YLTKEVVDGVSHWYVSQFSNVHNHELLEDDQVRLLPAYRKIQQSDQERILLLS 186
Query: 194 KSLTVPRNILTNLKENNKESVT 215
K+ P N + L E K V+
Sbjct: 187 KA-GFPVNRIVKLLELEKGVVS 207
>gb|AAF88018.1| contains simlarity to Arabidopsis thaliana far-red impaired
response protein (GB:AAD51282.1)
Length = 694
Score = 35.0 bits (79), Expect = 2.0
Identities = 34/142 (23%), Positives = 67/142 (46%), Gaps = 29/142 (20%)
Query: 92 KAGFGVCIDKSVLKRPYLTTQSERSGIYKPP---------KTRKKPNLEGIG---SMKCN 139
K+GF + R +T+S+ G+Y+ + RKK N+E S++C
Sbjct: 77 KSGFSI--------RKARSTESQNLGVYRRDFVCYRSGFNQPRKKANVEHPRERKSVRCG 128
Query: 140 CPFRLKGFFDKDTND----WWLAILCGMHNHDLEEKLQGHLIAG--RLSAEEKKKVIDMT 193
C +L + K+ D W+++ +HNH+L E Q L+ ++ ++++++ ++
Sbjct: 129 CDGKL--YLTKEVVDGVSHWYVSQFSNVHNHELLEDDQVRLLPAYRKIQQSDQERILLLS 186
Query: 194 KSLTVPRNILTNLKENNKESVT 215
K+ P N + L E K V+
Sbjct: 187 KA-GFPVNRIVKLLELEKGVVS 207
>gb|EAK89601.1| hypothetical protein, signal peptide [Cryptosporidium parvum]
gi|66361656|ref|XP_627351.1| hypothetical protein
cgd8_4660 [Cryptosporidium parvum]
Length = 397
Score = 34.7 bits (78), Expect = 2.6
Identities = 23/67 (34%), Positives = 35/67 (51%), Gaps = 9/67 (13%)
Query: 186 KKKVIDMTKSLTVPRNILTNLKENNKE----SVTTIKQVY--NVRTRWCK---GERGDMT 236
+KK++D + T+PRN + +LKE N++ S T Y N +W K E G +T
Sbjct: 125 RKKIMDKLRQKTLPRNNIRSLKEENQDENNGSKTLPTSFYWPNETEKWAKISFLESGSIT 184
Query: 237 ELQFLIS 243
E F +S
Sbjct: 185 ETNFTMS 191
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.318 0.136 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 446,816,587
Number of Sequences: 2540612
Number of extensions: 18603507
Number of successful extensions: 44371
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 29
Number of HSP's that attempted gapping in prelim test: 44358
Number of HSP's gapped (non-prelim): 35
length of query: 256
length of database: 863,360,394
effective HSP length: 125
effective length of query: 131
effective length of database: 545,783,894
effective search space: 71497690114
effective search space used: 71497690114
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 74 (33.1 bits)
Medicago: description of AC135234.5


