

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135230.4 - phase: 0
(360 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
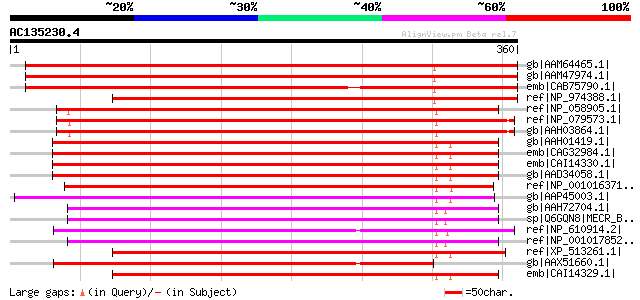
Score E
Sequences producing significant alignments: (bits) Value
gb|AAM64465.1| nuclear receptor binding factor-like protein [Ara... 513 e-144
gb|AAM47974.1| oxidoreductase of zinc-binding dehydrogenase fami... 511 e-143
emb|CAB75790.1| nuclear receptor binding factor-like protein [Ar... 492 e-138
ref|NP_974388.1| oxidoreductase, zinc-binding dehydrogenase fami... 446 e-124
ref|NP_058905.1| nuclear receptor-binding factor 1 [Rattus norve... 275 1e-72
ref|NP_079573.1| nuclear receptor-binding factor 1 [Mus musculus... 274 3e-72
gb|AAH03864.1| Nuclear receptor-binding factor 1 [Mus musculus] ... 272 1e-71
gb|AAH01419.1| Nuclear receptor-binding factor 1, isoform a [Hom... 267 3e-70
emb|CAG32984.1| CGI-63 [Homo sapiens] 267 3e-70
emb|CAI14330.1| OTTHUMP00000065098 [Homo sapiens] gi|67078404|re... 266 5e-70
gb|AAD34058.1| CGI-63 protein [Homo sapiens] 266 7e-70
ref|NP_001016371.1| hypothetical protein LOC549125 [Xenopus trop... 265 1e-69
gb|AAP45003.1| 2-enoyl thioester reductase [Bos taurus] gi|62900... 259 1e-67
gb|AAH72704.1| Unknown (protein for IMAGE:7036761) [Danio rerio] 252 1e-65
sp|Q6GQN8|MECR_BRARE Trans-2-enoyl-CoA reductase, mitochondrial ... 252 1e-65
ref|NP_610914.2| CG16935-PA [Drosophila melanogaster] gi|4544555... 251 3e-65
ref|NP_001017852.1| hypothetical protein LOC550550 [Danio rerio]... 249 7e-65
ref|XP_513261.1| PREDICTED: hypothetical protein XP_513261 [Pan ... 244 2e-63
gb|AAX51660.1| AT25977p [Drosophila melanogaster] 244 4e-63
emb|CAI14329.1| OTTHUMP00000065099 [Homo sapiens] gi|67078406|re... 243 6e-63
>gb|AAM64465.1| nuclear receptor binding factor-like protein [Arabidopsis thaliana]
gi|62900587|sp|Q8LCU7|MECR_ARATH Probable
trans-2-enoyl-CoA reductase, mitochondrial precursor
gi|18408069|ref|NP_566881.1| oxidoreductase,
zinc-binding dehydrogenase family protein [Arabidopsis
thaliana]
Length = 375
Score = 513 bits (1321), Expect = e-144
Identities = 258/360 (71%), Positives = 297/360 (81%), Gaps = 11/360 (3%)
Query: 12 SSLFNFTSLRSRILNTHT-HAFSSAVSPPSKAIIYESHGQPDAVTKLVDIPATEVKENDL 70
SS NF S+R T +FS+ +SPPSKAI+YE HG PD+VT+LV++P EVKEND+
Sbjct: 16 SSTANFRSIRRGETPTLCIKSFSTIMSPPSKAIVYEEHGSPDSVTRLVNLPPVEVKENDV 75
Query: 71 CVKMLAAPINPSDINRIQGVYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWVIPSPP 130
CVKM+AAPINPSDINRI+GVYPVRP PAVGGYEGVGEV++VGS V FSPGDWVIPSPP
Sbjct: 76 CVKMIAAPINPSDINRIEGVYPVRPPVPAVGGYEGVGEVYAVGSNVNGFSPGDWVIPSPP 135
Query: 131 SFGTWQTYIVKDQNVWHKVNKGVPMEYAATITVNPLTALLMLEDCVTLNSGDAIVQNGAT 190
S GTWQTY+VK+++VWHK++K PMEYAATITVNPLTAL MLED V LNSGD++VQNGAT
Sbjct: 136 SSGTWQTYVVKEESVWHKIDKECPMEYAATITVNPLTALRMLEDFVNLNSGDSVVQNGAT 195
Query: 191 SMVGQCVIQLAKSRGIHNINIIRDRPGVGEVKERLKDLGADEVFTESELEVKNVKSLLGG 250
S+VGQCVIQLA+ RGI IN+IRDR G E +E+LK LGADEVF+ES+L VKNVKSLLG
Sbjct: 196 SIVGQCVIQLARLRGISTINLIRDRAGSDEAREQLKALGADEVFSESQLNVKNVKSLLGN 255
Query: 251 IPEPALGFNCVGGNSASLVLKFLRRGGTMVTYGGMSKKPVTVSTSSFIFK---------- 300
+PEPALGFNCVGGN+ASLVLK+LR GGTMVTYGGMSKKP+TVST+SFIFK
Sbjct: 256 LPEPALGFNCVGGNAASLVLKYLREGGTMVTYGGMSKKPITVSTTSFIFKDLALRGFWLQ 315
Query: 301 NWLSTDKAEEGRRMIDRLLGLVQDGKLKYKMELTPFNDFNTALDKALGKLGSQPKQVIKF 360
+WLS K +E R MID LLGL +DGKLKY+ EL PF +F ALDKALGKLG QPKQVI F
Sbjct: 316 SWLSMGKVKECREMIDYLLGLARDGKLKYETELVPFEEFPVALDKALGKLGRQPKQVITF 375
>gb|AAM47974.1| oxidoreductase of zinc-binding dehydrogenase family [Arabidopsis
thaliana] gi|17064956|gb|AAL32632.1| oxidoreductase of
zinc-binding dehydrogenase family [Arabidopsis thaliana]
Length = 375
Score = 511 bits (1317), Expect = e-143
Identities = 257/360 (71%), Positives = 297/360 (82%), Gaps = 11/360 (3%)
Query: 12 SSLFNFTSLRSRILNTHT-HAFSSAVSPPSKAIIYESHGQPDAVTKLVDIPATEVKENDL 70
SS NF S+R T +FS+ +SPPSKAI+YE HG PD+VT+LV++P EVKEND+
Sbjct: 16 SSTANFRSIRRGETPTLCIKSFSTIMSPPSKAIVYEEHGSPDSVTRLVNLPPVEVKENDV 75
Query: 71 CVKMLAAPINPSDINRIQGVYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWVIPSPP 130
CVKM+AAPINPSDINRI+GVYPVRP PAVGGYEGVGEV++VGS V FSPGDWVIPSPP
Sbjct: 76 CVKMIAAPINPSDINRIEGVYPVRPPVPAVGGYEGVGEVYAVGSNVNGFSPGDWVIPSPP 135
Query: 131 SFGTWQTYIVKDQNVWHKVNKGVPMEYAATITVNPLTALLMLEDCVTLNSGDAIVQNGAT 190
S GTWQTY+VK+++VWHK++K PMEYAATITVNPLTAL MLED V LNSGD++VQNGAT
Sbjct: 136 SSGTWQTYVVKEESVWHKIDKECPMEYAATITVNPLTALRMLEDFVNLNSGDSVVQNGAT 195
Query: 191 SMVGQCVIQLAKSRGIHNINIIRDRPGVGEVKERLKDLGADEVFTESELEVKNVKSLLGG 250
S+VGQCVIQLA+ RGI IN+IRDR G E +E+LK LGADEVF+ES+L VKNVKSLLG
Sbjct: 196 SIVGQCVIQLARLRGISTINLIRDRAGSDEAREQLKALGADEVFSESQLNVKNVKSLLGN 255
Query: 251 IPEPALGFNCVGGNSASLVLKFLRRGGTMVTYGGMSKKPVTVSTSSFIFK---------- 300
+PEPALGFNCVGGN+ASLVLK+LR GGTMVTYGGMSK+P+TVST+SFIFK
Sbjct: 256 LPEPALGFNCVGGNAASLVLKYLREGGTMVTYGGMSKEPITVSTTSFIFKDLALRGFWLQ 315
Query: 301 NWLSTDKAEEGRRMIDRLLGLVQDGKLKYKMELTPFNDFNTALDKALGKLGSQPKQVIKF 360
+WLS K +E R MID LLGL +DGKLKY+ EL PF +F ALDKALGKLG QPKQVI F
Sbjct: 316 SWLSMGKVKECREMIDYLLGLARDGKLKYETELVPFEEFPVALDKALGKLGRQPKQVITF 375
>emb|CAB75790.1| nuclear receptor binding factor-like protein [Arabidopsis thaliana]
Length = 367
Score = 492 bits (1266), Expect = e-138
Identities = 250/360 (69%), Positives = 289/360 (79%), Gaps = 19/360 (5%)
Query: 12 SSLFNFTSLRSRILNTHT-HAFSSAVSPPSKAIIYESHGQPDAVTKLVDIPATEVKENDL 70
SS NF S+R T +FS+ +SPPSKAI+YE HG PD+VT+LV++P EVKEND+
Sbjct: 16 SSTANFRSIRRGETPTLCIKSFSTIMSPPSKAIVYEEHGSPDSVTRLVNLPPVEVKENDV 75
Query: 71 CVKMLAAPINPSDINRIQGVYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWVIPSPP 130
CVKM+AAPINPSDINRI+GVYPVRP PAVGGYEGVGEV++VGS V FSPGDWVIPSPP
Sbjct: 76 CVKMIAAPINPSDINRIEGVYPVRPPVPAVGGYEGVGEVYAVGSNVNGFSPGDWVIPSPP 135
Query: 131 SFGTWQTYIVKDQNVWHKVNKGVPMEYAATITVNPLTALLMLEDCVTLNSGDAIVQNGAT 190
S GTWQTY+VK+++VWHK++K PMEYAATITVNPLTAL MLED V LNSGD++VQNGAT
Sbjct: 136 SSGTWQTYVVKEESVWHKIDKECPMEYAATITVNPLTALRMLEDFVNLNSGDSVVQNGAT 195
Query: 191 SMVGQCVIQLAKSRGIHNINIIRDRPGVGEVKERLKDLGADEVFTESELEVKNVKSLLGG 250
S+VGQCVIQLA+ RGI IN+IRDR G E +E+LK LGADEVF+ES+L G
Sbjct: 196 SIVGQCVIQLARLRGISTINLIRDRAGSDEAREQLKALGADEVFSESQLN--------GN 247
Query: 251 IPEPALGFNCVGGNSASLVLKFLRRGGTMVTYGGMSKKPVTVSTSSFIFK---------- 300
+PEPALGFNCVGGN+ASLVLK+LR GGTMVTYGGMSKKP+TVST+SFIFK
Sbjct: 248 LPEPALGFNCVGGNAASLVLKYLREGGTMVTYGGMSKKPITVSTTSFIFKDLALRGFWLQ 307
Query: 301 NWLSTDKAEEGRRMIDRLLGLVQDGKLKYKMELTPFNDFNTALDKALGKLGSQPKQVIKF 360
+WLS K +E R MID LLGL +DGKLKY+ EL PF +F ALDKALGKLG QPKQVI F
Sbjct: 308 SWLSMGKVKECREMIDYLLGLARDGKLKYETELVPFEEFPVALDKALGKLGRQPKQVITF 367
>ref|NP_974388.1| oxidoreductase, zinc-binding dehydrogenase family protein
[Arabidopsis thaliana]
Length = 297
Score = 446 bits (1147), Expect = e-124
Identities = 222/297 (74%), Positives = 251/297 (83%), Gaps = 10/297 (3%)
Query: 74 MLAAPINPSDINRIQGVYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFG 133
M+AAPINPSDINRI+GVYPVRP PAVGGYEGVGEV++VGS V FSPGDWVIPSPPS G
Sbjct: 1 MIAAPINPSDINRIEGVYPVRPPVPAVGGYEGVGEVYAVGSNVNGFSPGDWVIPSPPSSG 60
Query: 134 TWQTYIVKDQNVWHKVNKGVPMEYAATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMV 193
TWQTY+VK+++VWHK++K PMEYAATITVNPLTAL MLED V LNSGD++VQNGATS+V
Sbjct: 61 TWQTYVVKEESVWHKIDKECPMEYAATITVNPLTALRMLEDFVNLNSGDSVVQNGATSIV 120
Query: 194 GQCVIQLAKSRGIHNINIIRDRPGVGEVKERLKDLGADEVFTESELEVKNVKSLLGGIPE 253
GQCVIQLA+ RGI IN+IRDR G E +E+LK LGADEVF+ES+L VKNVKSLLG +PE
Sbjct: 121 GQCVIQLARLRGISTINLIRDRAGSDEAREQLKALGADEVFSESQLNVKNVKSLLGNLPE 180
Query: 254 PALGFNCVGGNSASLVLKFLRRGGTMVTYGGMSKKPVTVSTSSFIFK----------NWL 303
PALGFNCVGGN+ASLVLK+LR GGTMVTYGGMSKKP+TVST+SFIFK +WL
Sbjct: 181 PALGFNCVGGNAASLVLKYLREGGTMVTYGGMSKKPITVSTTSFIFKDLALRGFWLQSWL 240
Query: 304 STDKAEEGRRMIDRLLGLVQDGKLKYKMELTPFNDFNTALDKALGKLGSQPKQVIKF 360
S K +E R MID LLGL +DGKLKY+ EL PF +F ALDKALGKLG QPKQVI F
Sbjct: 241 SMGKVKECREMIDYLLGLARDGKLKYETELVPFEEFPVALDKALGKLGRQPKQVITF 297
>ref|NP_058905.1| nuclear receptor-binding factor 1 [Rattus norvegicus]
gi|62900383|sp|Q9Z311|MECR_RAT Trans-2-enoyl-CoA
reductase, mitochondrial precursor (Nuclear receptor
binding factor 1) (NRBF-1) gi|3970880|dbj|BAA34804.1|
nuclear receptor binding factor-1 [Rattus norvegicus]
Length = 373
Score = 275 bits (704), Expect = 1e-72
Identities = 150/327 (45%), Positives = 206/327 (62%), Gaps = 13/327 (3%)
Query: 34 SAVSPPS--KAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVY 91
SA S PS +A++Y +HG P V +L ++ T V+ +D+ VKMLAAPINPSDIN IQG Y
Sbjct: 35 SAFSEPSHVRALVYGNHGDPAKVIQLKNLELTAVEGSDVHVKMLAAPINPSDINMIQGNY 94
Query: 92 PVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKVNK 151
+ P+ PAVGG EGVG+V +VGS+V+ PGDWVIP+ GTW+T V + V K
Sbjct: 95 GLLPKLPAVGGNEGVGQVIAVGSSVSGLKPGDWVIPANAGLGTWRTEAVFSEEALIGVPK 154
Query: 152 GVPMEYAATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINI 211
+P++ AAT+ VNP TA ML D L GD+++QN + S VGQ VIQ+A + G+ IN+
Sbjct: 155 DIPLQSAATLGVNPCTAYRMLVDFEQLQPGDSVIQNASNSGVGQAVIQIASALGLKTINV 214
Query: 212 IRDRPGVGEVKERLKDLGADEVFTESELEVKNVKSLLGGIPEPALGFNCVGGNSASLVLK 271
IRDRP + ++ +RLKDLGAD V TE EL + K++ +P P L NCVGG S++ +L+
Sbjct: 215 IRDRPDIKKLTDRLKDLGADYVLTEEELRMPETKNIFKDLPLPRLALNCVGGKSSTELLR 274
Query: 272 FLRRGGTMVTYGGMSKKPVTVSTSSFIFKN----------WLSTDKAEEGRRMIDRLLGL 321
L GGTMVTYGGM+K+PVT S S IFK+ W +E + +I L L
Sbjct: 275 HLAPGGTMVTYGGMAKQPVTASVSMLIFKDLKLRGFWLSQWKKNHSPDEFKELILILCNL 334
Query: 322 VQDGKLKY-KMELTPFNDFNTALDKAL 347
++ G+L P D+ AL+ ++
Sbjct: 335 IRQGQLTAPAWSGIPLQDYQQALEASM 361
>ref|NP_079573.1| nuclear receptor-binding factor 1 [Mus musculus]
gi|12832585|dbj|BAB22169.1| unnamed protein product [Mus
musculus]
Length = 373
Score = 274 bits (700), Expect = 3e-72
Identities = 149/338 (44%), Positives = 210/338 (62%), Gaps = 14/338 (4%)
Query: 34 SAVSPPSK--AIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVY 91
SA+S PS+ A++Y +HG P V +L ++ T VK +D+ V+MLAAPINPSDIN IQG Y
Sbjct: 35 SALSEPSRVRALVYGNHGDPAKVVQLKNLELTAVKGSDVHVRMLAAPINPSDINMIQGNY 94
Query: 92 PVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKVNK 151
+ P+ PAVGG EGVG+V +VGS+V+ PGDWVIP+ GTW+T V + + K
Sbjct: 95 GLLPKLPAVGGNEGVGQVIAVGSSVSALKPGDWVIPANAGLGTWRTEAVFSEEALIGIPK 154
Query: 152 GVPMEYAATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINI 211
+P++ AAT+ VNP TA ML D L GD+++QN + S VGQ VIQ+A + + IN+
Sbjct: 155 DIPLQSAATLGVNPCTAYRMLVDFEQLQPGDSVIQNASNSGVGQAVIQIASALRLKTINV 214
Query: 212 IRDRPGVGEVKERLKDLGADEVFTESELEVKNVKSLLGGIPEPALGFNCVGGNSASLVLK 271
+RDRP + ++ +RLKDLGAD V TE EL + K++ +P P L NCVGG S++ +L+
Sbjct: 215 VRDRPDIKKLTDRLKDLGADYVLTEEELRMPETKTIFKDLPLPRLALNCVGGKSSTELLR 274
Query: 272 FLRRGGTMVTYGGMSKKPVTVSTSSFIFKN----------WLSTDKAEEGRRMIDRLLGL 321
L GGTMVTYGGM+K+PVT S S IFK+ W +E + +I L L
Sbjct: 275 HLAPGGTMVTYGGMAKQPVTASVSLLIFKDLKLRGFWLSQWKKNHSPDEFKELILTLCNL 334
Query: 322 VQDGKLKY-KMELTPFNDFNTALDKALGKLGSQPKQVI 358
++ G+L P + AL+ ++ S KQ++
Sbjct: 335 IRQGRLTAPSCSEVPLQGYQQALEASMKPFVSS-KQIL 371
>gb|AAH03864.1| Nuclear receptor-binding factor 1 [Mus musculus]
gi|62900598|sp|Q9DCS3|MECR_MOUSE Trans-2-enoyl-CoA
reductase, mitochondrial precursor
Length = 373
Score = 272 bits (696), Expect = 1e-71
Identities = 148/338 (43%), Positives = 210/338 (61%), Gaps = 14/338 (4%)
Query: 34 SAVSPPSK--AIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVY 91
SA+S PS+ A++Y +HG P V +L ++ T V+ +D+ V+MLAAPINPSDIN IQG Y
Sbjct: 35 SALSEPSRVRALVYGNHGDPAKVVQLKNLELTAVEGSDVHVRMLAAPINPSDINMIQGNY 94
Query: 92 PVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKVNK 151
+ P+ PAVGG EGVG+V +VGS+V+ PGDWVIP+ GTW+T V + + K
Sbjct: 95 GLLPKLPAVGGNEGVGQVIAVGSSVSALKPGDWVIPANAGLGTWRTEAVFSEEALIGIPK 154
Query: 152 GVPMEYAATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINI 211
+P++ AAT+ VNP TA ML D L GD+++QN + S VGQ VIQ+A + + IN+
Sbjct: 155 DIPLQSAATLGVNPCTAYRMLVDFEQLQPGDSVIQNASNSGVGQAVIQIASALRLKTINV 214
Query: 212 IRDRPGVGEVKERLKDLGADEVFTESELEVKNVKSLLGGIPEPALGFNCVGGNSASLVLK 271
+RDRP + ++ +RLKDLGAD V TE EL + K++ +P P L NCVGG S++ +L+
Sbjct: 215 VRDRPDIKKLTDRLKDLGADYVLTEEELRMPETKTIFKDLPLPRLALNCVGGKSSTELLR 274
Query: 272 FLRRGGTMVTYGGMSKKPVTVSTSSFIFKN----------WLSTDKAEEGRRMIDRLLGL 321
L GGTMVTYGGM+K+PVT S S IFK+ W +E + +I L L
Sbjct: 275 HLAPGGTMVTYGGMAKQPVTASVSLLIFKDLKLRGFWLSQWKKNHSPDEFKELILTLCNL 334
Query: 322 VQDGKLKY-KMELTPFNDFNTALDKALGKLGSQPKQVI 358
++ G+L P + AL+ ++ S KQ++
Sbjct: 335 IRQGRLTAPSCSEVPLQGYQQALEASMKPFVSS-KQIL 371
>gb|AAH01419.1| Nuclear receptor-binding factor 1, isoform a [Homo sapiens]
gi|62900596|sp|Q9BV79|MECR_HUMAN Trans-2-enoyl-CoA
reductase, mitochondrial precursor (HsNrbf-1) (NRBF-1)
Length = 373
Score = 267 bits (683), Expect = 3e-70
Identities = 145/329 (44%), Positives = 205/329 (62%), Gaps = 12/329 (3%)
Query: 31 AFSSAVSPPS-KAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQG 89
++S++ P +A++Y HG P V +L ++ V+ +D+ VKMLAAPINPSDIN IQG
Sbjct: 33 SYSASAEPARVRALVYGHHGDPAKVVELKNLELAAVRGSDVRVKMLAAPINPSDINMIQG 92
Query: 90 VYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKV 149
Y + PE PAVGG EGV +V +VGS VT PGDWVIP+ GTW+T V + +V
Sbjct: 93 NYGLLPELPAVGGNEGVAQVVAVGSNVTGLKPGDWVIPANAGLGTWRTEAVFSEEALIQV 152
Query: 150 NKGVPMEYAATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNI 209
+P++ AAT+ VNP TA ML D L GD+++QN + S VGQ VIQ+A + G+ I
Sbjct: 153 PSDIPLQSAATLGVNPCTAYRMLMDFEQLQPGDSVIQNASNSGVGQAVIQIAAALGLRTI 212
Query: 210 NIIRDRPGVGEVKERLKDLGADEVFTESELEVKNVKSLLGGIPEPALGFNCVGGNSASLV 269
N++RDRP + ++ +RLK LGA+ V TE EL +K+ +P+P L NCVGG S++ +
Sbjct: 213 NVVRDRPDIQKLSDRLKSLGAEHVITEEELRRPEMKNFFKDMPQPRLALNCVGGKSSTEL 272
Query: 270 LKFLRRGGTMVTYGGMSKKPVTVSTSSFIFKN------WLSTDKAEEG----RRMIDRLL 319
L+ L RGGTMVTYGGM+K+PV S S IFK+ WLS K + + +I L
Sbjct: 273 LRQLARGGTMVTYGGMAKQPVVASVSLLIFKDLKLRGFWLSQWKKDHSPDQFKELILTLC 332
Query: 320 GLVQDGKLKY-KMELTPFNDFNTALDKAL 347
L++ G+L P D+ +AL+ ++
Sbjct: 333 DLIRRGQLTAPACSQVPLQDYQSALEASM 361
>emb|CAG32984.1| CGI-63 [Homo sapiens]
Length = 373
Score = 267 bits (683), Expect = 3e-70
Identities = 145/329 (44%), Positives = 205/329 (62%), Gaps = 12/329 (3%)
Query: 31 AFSSAVSPPS-KAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQG 89
++S++ P +A++Y HG P V +L ++ V+ +D+ VKMLAAPINPSDIN IQG
Sbjct: 33 SYSASAEPARVRALVYGHHGDPAKVVELKNLELAAVRGSDVRVKMLAAPINPSDINMIQG 92
Query: 90 VYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKV 149
Y + PE PAVGG EGV +V +VGS VT PGDWVIP+ GTW+T V + +V
Sbjct: 93 NYGLLPELPAVGGNEGVAQVVAVGSNVTGLKPGDWVIPANAGLGTWRTEAVFSEEALIQV 152
Query: 150 NKGVPMEYAATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNI 209
+P++ AAT+ VNP TA ML D L GD+++QN + S VGQ VIQ+A + G+ I
Sbjct: 153 PSDIPLQSAATLGVNPCTAYRMLMDFEQLQPGDSVIQNASNSGVGQAVIQIAAALGLRTI 212
Query: 210 NIIRDRPGVGEVKERLKDLGADEVFTESELEVKNVKSLLGGIPEPALGFNCVGGNSASLV 269
N++RDRP + ++ +RLK LGA+ V TE EL +K+ +P+P L NCVGG S++ +
Sbjct: 213 NVVRDRPDIQKLSDRLKSLGAEHVITEEELRRPEMKNFFKDMPQPRLALNCVGGKSSTEL 272
Query: 270 LKFLRRGGTMVTYGGMSKKPVTVSTSSFIFKN------WLSTDKAEEG----RRMIDRLL 319
L+ L RGGTMVTYGGM+K+PV S S IFK+ WLS K + + +I L
Sbjct: 273 LRQLARGGTMVTYGGMAKQPVVASVSLLIFKDLKLRGFWLSQWKKDHSPDQFKELILTLC 332
Query: 320 GLVQDGKLKY-KMELTPFNDFNTALDKAL 347
L++ G+L P D+ +AL+ ++
Sbjct: 333 DLIRRGQLTAPACSQVPLQDYQSALEASM 361
>emb|CAI14330.1| OTTHUMP00000065098 [Homo sapiens] gi|67078404|ref|NP_057095.2|
nuclear receptor-binding factor 1 isoform a [Homo
sapiens]
Length = 373
Score = 266 bits (681), Expect = 5e-70
Identities = 145/329 (44%), Positives = 204/329 (61%), Gaps = 12/329 (3%)
Query: 31 AFSSAVSPPS-KAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQG 89
++S++ P +A++Y HG P V +L ++ V+ +D+ VKMLAAPINPSDIN IQG
Sbjct: 33 SYSASAEPARVRALVYGHHGDPAKVVELKNLELAAVRGSDVRVKMLAAPINPSDINMIQG 92
Query: 90 VYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKV 149
Y PE PAVGG EGV +V +VGS VT PGDWVIP+ GTW+T V + +V
Sbjct: 93 NYGFLPELPAVGGNEGVAQVVAVGSNVTGLKPGDWVIPANAGLGTWRTEAVFSEEALIQV 152
Query: 150 NKGVPMEYAATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNI 209
+P++ AAT+ VNP TA ML D L GD+++QN + S VGQ VIQ+A + G+ I
Sbjct: 153 PSDIPLQSAATLGVNPCTAYRMLMDFEQLQPGDSVIQNASNSGVGQAVIQIAAALGLRTI 212
Query: 210 NIIRDRPGVGEVKERLKDLGADEVFTESELEVKNVKSLLGGIPEPALGFNCVGGNSASLV 269
N++RDRP + ++ +RLK LGA+ V TE EL +K+ +P+P L NCVGG S++ +
Sbjct: 213 NVVRDRPDIQKLSDRLKSLGAEHVITEEELRRPEMKNFFKDMPQPRLALNCVGGKSSTEL 272
Query: 270 LKFLRRGGTMVTYGGMSKKPVTVSTSSFIFKN------WLSTDKAEEG----RRMIDRLL 319
L+ L RGGTMVTYGGM+K+PV S S IFK+ WLS K + + +I L
Sbjct: 273 LRQLARGGTMVTYGGMAKQPVVASVSLLIFKDLKLRGFWLSQWKKDHSPDQFKELILTLC 332
Query: 320 GLVQDGKLKY-KMELTPFNDFNTALDKAL 347
L++ G+L P D+ +AL+ ++
Sbjct: 333 DLIRRGQLTAPACSQVPLQDYQSALEASM 361
>gb|AAD34058.1| CGI-63 protein [Homo sapiens]
Length = 373
Score = 266 bits (680), Expect = 7e-70
Identities = 144/329 (43%), Positives = 204/329 (61%), Gaps = 12/329 (3%)
Query: 31 AFSSAVSPPS-KAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQG 89
++S++ P + ++Y HG P V +L ++ V+ +D+ VKMLAAPINPSDIN IQG
Sbjct: 33 SYSASAEPARVRGVVYGHHGDPAKVVELKNLELAAVRGSDVRVKMLAAPINPSDINMIQG 92
Query: 90 VYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKV 149
Y + PE PAVGG EGV +V +VGS VT PGDWVIP+ GTW+T V + +V
Sbjct: 93 NYGLLPELPAVGGNEGVAQVVAVGSNVTGLKPGDWVIPANAGLGTWRTEAVFSEEALIQV 152
Query: 150 NKGVPMEYAATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNI 209
+P++ AAT+ VNP TA ML D L GD+++QN + S VGQ VIQ+A + G+ I
Sbjct: 153 PSDIPLQSAATLGVNPCTAYRMLMDFEQLQPGDSVIQNASNSGVGQAVIQIAAALGLRTI 212
Query: 210 NIIRDRPGVGEVKERLKDLGADEVFTESELEVKNVKSLLGGIPEPALGFNCVGGNSASLV 269
N++RDRP + ++ +RLK LGA+ V TE EL +K+ +P+P L NCVGG S++ +
Sbjct: 213 NVVRDRPDIQKLSDRLKSLGAEHVITEEELRRPEMKNFFKDMPQPRLALNCVGGKSSTEL 272
Query: 270 LKFLRRGGTMVTYGGMSKKPVTVSTSSFIFKN------WLSTDKAEEG----RRMIDRLL 319
L+ L RGGTMVTYGGM+K+PV S S IFK+ WLS K + + +I L
Sbjct: 273 LRQLARGGTMVTYGGMAKQPVVASVSLLIFKDLKLRGFWLSQWKKDHSPDQFKELILTLC 332
Query: 320 GLVQDGKLKY-KMELTPFNDFNTALDKAL 347
L++ G+L P D+ +AL+ ++
Sbjct: 333 DLIRRGQLTAPACSQVPLQDYQSALEASM 361
>ref|NP_001016371.1| hypothetical protein LOC549125 [Xenopus tropicalis]
Length = 350
Score = 265 bits (678), Expect = 1e-69
Identities = 145/318 (45%), Positives = 193/318 (60%), Gaps = 14/318 (4%)
Query: 40 SKAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPA 99
++ ++YE HG+P V +L ++ T +N++ VKMLAAPINPSDIN +QG Y + P+ PA
Sbjct: 17 ARGLVYEKHGEPLQVLRLKNVNITHPADNEVRVKMLAAPINPSDINMVQGTYALLPQLPA 76
Query: 100 VGGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKVNKGVPMEYAA 159
VGG EGVG V +G V+ PGDWV+P GTW T V ++ +V +P+ AA
Sbjct: 77 VGGNEGVGVVVEIGRHVSSMRPGDWVVPVDAGLGTWCTEAVFSEDSLVRVPSDIPVAGAA 136
Query: 160 TITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRDRPGVG 219
T++VNP TA +L D TL GD I+QN + S VGQ VIQ+A S GI IN++RDR +
Sbjct: 137 TVSVNPCTAYRLLSDFETLRPGDTIIQNASNSGVGQAVIQIATSLGITTINVVRDREDLS 196
Query: 220 EVKERLKDLGADEVFTESELEVKNVKSLLGGIPEPALGFNCVGGNSASLVLKFLRRGGTM 279
+ +RL+DLGAD V TE +L +K L P P L NCVGG S + +L+ L GGTM
Sbjct: 197 SLIQRLRDLGADHVITEEQLRKPEMKDLFKNCPRPRLALNCVGGKSTTEMLRHLDYGGTM 256
Query: 280 VTYGGMSKKPVTVSTSSFIFKN------WLSTDKAEEGR-------RMIDRLLGLVQDGK 326
VTYGGMSK+PVTV S+ IFKN W++ K E + +MI L L++ GK
Sbjct: 257 VTYGGMSKQPVTVPVSALIFKNVKLCGFWVTQWKKERAQTDREEIVKMIRDLCDLIRRGK 316
Query: 327 L-KYKMELTPFNDFNTAL 343
L P DF+ AL
Sbjct: 317 LVPPPSTQRPLEDFSRAL 334
>gb|AAP45003.1| 2-enoyl thioester reductase [Bos taurus]
gi|62900582|sp|Q7YS70|MECR_BOVIN Trans-2-enoyl-CoA
reductase, mitochondrial precursor (BtNrbf-1) (NRBF-1)
gi|31982403|ref|NP_858055.1| nuclear receptor binding
factor 1 [Bos taurus]
Length = 373
Score = 259 bits (661), Expect = 1e-67
Identities = 150/353 (42%), Positives = 208/353 (58%), Gaps = 12/353 (3%)
Query: 4 SLSHKPSPSSLFNFTSLRSRILNTHTHAFSSAVSPPS-KAIIYESHGQPDAVTKLVDIPA 62
+L +P+ L SR + +FS++ P +A++Y HG P V +L ++
Sbjct: 6 ALCRTRAPAQLGQRLLPESRRRRPASASFSASAEPSRVRALVYGHHGDPAKVVELKNLEL 65
Query: 63 TEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPG 122
V + + VKMLAAPINPSDIN IQG Y + P+ PAVGG EGVG+V +VGS VT PG
Sbjct: 66 AAVGGSHVHVKMLAAPINPSDINMIQGNYGLLPQLPAVGGNEGVGQVVAVGSGVTGVKPG 125
Query: 123 DWVIPSPPSFGTWQTYIVKDQNVWHKVNKGVPMEYAATITVNPLTALLMLEDCVTLNSGD 182
DWVIP+ P GTW+T V + V +P++ AAT+ VNP TA ML D L D
Sbjct: 126 DWVIPANPGLGTWRTEAVFGEEELITVPSDIPLQSAATLGVNPCTAYRMLVDFERLRPRD 185
Query: 183 AIVQNGATSMVGQCVIQLAKSRGIHNINIIRDRPGVGEVKERLKDLGADEVFTESELEVK 242
+I+QN + S VGQ VIQ+A +RG+ IN++RD P + ++ + LK+LGA+ V TE EL
Sbjct: 186 SIIQNASNSGVGQAVIQIAAARGLRTINVLRDTPDLQKLTDTLKNLGANHVVTEEELRKP 245
Query: 243 NVKSLLGGIPEPALGFNCVGGNSASLVLKFLRRGGTMVTYGGMSKKPVTVSTSSFIFKN- 301
+KS +P+P L NCVGG S++ +L+ L GGTMVTYGGM+K+PV S S IFK+
Sbjct: 246 EMKSFFKDVPQPRLALNCVGGKSSTELLRHLAPGGTMVTYGGMAKQPVIASVSQLIFKDL 305
Query: 302 -----WLSTDKAEEG----RRMIDRLLGLVQDGKLKY-KMELTPFNDFNTALD 344
WLS K + + +I L L++ G+L P D+ AL+
Sbjct: 306 KLRGFWLSQWKKDHSPDQFKELILTLCDLIRRGQLTAPACSEVPLQDYLCALE 358
>gb|AAH72704.1| Unknown (protein for IMAGE:7036761) [Danio rerio]
Length = 413
Score = 252 bits (644), Expect = 1e-65
Identities = 140/319 (43%), Positives = 189/319 (58%), Gaps = 13/319 (4%)
Query: 42 AIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAVG 101
A++Y +HG+P V +L + +V + VKMLAAPINPSD+N +QG Y + PE PAVG
Sbjct: 83 ALLYRNHGEPSQVVQLESLDLPQVGAECVLVKMLAAPINPSDLNMLQGTYAILPELPAVG 142
Query: 102 GYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKVNKGVPMEYAATI 161
G EGV +V VG V GDWVIP GTW+T V + + K +P+ AAT+
Sbjct: 143 GNEGVAQVMEVGDKVKTLKVGDWVIPKDAGIGTWRTAAVLKADDLVTLPKDIPVLSAATL 202
Query: 162 TVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRDRPGVGEV 221
VNP TA ML D L +GD ++QN A S VGQ VIQ+A ++GIH IN+IRDRP + ++
Sbjct: 203 GVNPCTAYRMLTDFEELKAGDTVIQNAANSGVGQAVIQIAAAKGIHTINVIRDRPDLRQL 262
Query: 222 KERLKDLGADEVFTESELEVKNVKSLLGGIPEPALGFNCVGGNSASLVLKFLRRGGTMVT 281
+RL +GA V TE L +K L P P L N VGG SA+ +L+ L+ GG++VT
Sbjct: 263 SDRLTAMGATHVITEETLRRPEMKELFKSCPRPKLALNGVGGKSATELLRHLQSGGSLVT 322
Query: 282 YGGMSKKPVTVSTSSFIFKN------WLSTDKA------EEGRRMIDRLLGLVQDGKLKY 329
YGGM+K+PVTV S+ IFK+ W++ K E R M+D L L++ GKL
Sbjct: 323 YGGMAKQPVTVPVSALIFKDVRVRGFWVTQWKRDNRHDDEALRHMLDELCILIRAGKLSA 382
Query: 330 KM-ELTPFNDFNTALDKAL 347
+ DF AL+ A+
Sbjct: 383 PICTQVQLQDFRKALENAM 401
>sp|Q6GQN8|MECR_BRARE Trans-2-enoyl-CoA reductase, mitochondrial precursor
Length = 377
Score = 252 bits (644), Expect = 1e-65
Identities = 140/319 (43%), Positives = 189/319 (58%), Gaps = 13/319 (4%)
Query: 42 AIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAVG 101
A++Y +HG+P V +L + +V + VKMLAAPINPSD+N +QG Y + PE PAVG
Sbjct: 47 ALLYRNHGEPSQVVQLESLDLPQVGAECVLVKMLAAPINPSDLNMLQGTYAILPELPAVG 106
Query: 102 GYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKVNKGVPMEYAATI 161
G EGV +V VG V GDWVIP GTW+T V + + K +P+ AAT+
Sbjct: 107 GNEGVAQVMEVGDKVKTLKVGDWVIPKDAGIGTWRTAAVLKADDLVTLPKDIPVLSAATL 166
Query: 162 TVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRDRPGVGEV 221
VNP TA ML D L +GD ++QN A S VGQ VIQ+A ++GIH IN+IRDRP + ++
Sbjct: 167 GVNPCTAYRMLTDFEELKAGDTVIQNAANSGVGQAVIQIAAAKGIHTINVIRDRPDLRQL 226
Query: 222 KERLKDLGADEVFTESELEVKNVKSLLGGIPEPALGFNCVGGNSASLVLKFLRRGGTMVT 281
+RL +GA V TE L +K L P P L N VGG SA+ +L+ L+ GG++VT
Sbjct: 227 SDRLTAMGATHVITEETLRRPEMKELFKSCPRPKLALNGVGGKSATELLRHLQSGGSLVT 286
Query: 282 YGGMSKKPVTVSTSSFIFKN------WLSTDKA------EEGRRMIDRLLGLVQDGKLKY 329
YGGM+K+PVTV S+ IFK+ W++ K E R M+D L L++ GKL
Sbjct: 287 YGGMAKQPVTVPVSALIFKDVRVRGFWVTQWKRDNRHDDEALRHMLDELCILIRAGKLSA 346
Query: 330 KM-ELTPFNDFNTALDKAL 347
+ DF AL+ A+
Sbjct: 347 PICTQVQLQDFRKALENAM 365
>ref|NP_610914.2| CG16935-PA [Drosophila melanogaster] gi|45445556|gb|AAF58322.2|
CG16935-PA [Drosophila melanogaster]
gi|62900602|sp|Q9V6U9|MECR_DROME Probable
trans-2-enoyl-CoA reductase, mitochondrial precursor
Length = 357
Score = 251 bits (640), Expect = 3e-65
Identities = 149/340 (43%), Positives = 204/340 (59%), Gaps = 15/340 (4%)
Query: 32 FSSAVSPPSKAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVY 91
+S +S +K++ Y HG+P V +LV+ + K+N + VK+LAAPINP+DIN IQG Y
Sbjct: 15 WSRQMSVVAKSLKYTQHGEPQEVLQLVEDKLPDPKDNQVLVKILAAPINPADINTIQGKY 74
Query: 92 PVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKVNK 151
PV+P+ PAVGG E V EV VG V F G VIP GTW T+ V ++ V+K
Sbjct: 75 PVKPKFPAVGGNECVAEVICVGDKVKGFEAGQHVIPLASGLGTWTTHAVYKEDQLLIVSK 134
Query: 152 GVPMEYAATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINI 211
V + AAT TVNP TA ML+D V L GD ++QNGA S VGQ V QL ++ GI+++ I
Sbjct: 135 KVGLAEAATSTVNPTTAYRMLKDFVQLCPGDTVIQNGANSAVGQAVHQLCRAWGINSVGI 194
Query: 212 IRDRPGVGEVKERLKDLGADEVFTESELEVKNV-KSLLGGIPEPALGFNCVGGNSASLVL 270
+RDRP + E+K+ L+ LGA EV TE+E+ ++ KS G + +P L FNCVGG SA+ V
Sbjct: 195 VRDRPEIAELKQMLQCLGATEVLTEAEIRTSDIFKS--GKLKKPRLAFNCVGGKSATEVS 252
Query: 271 KFLRRGGTMVTYGGMSKKPVTVSTSSFIFKN------WLSTDKAE-----EGRRMIDRLL 319
+ L GG +VTYGGMS++PVTV+T IFK+ W++ E E +M +
Sbjct: 253 RHLDNGGVLVTYGGMSREPVTVATGPLIFKDIAFRGFWMTRWSKENYSSPERSKMFKEIF 312
Query: 320 GLVQDGK-LKYKMELTPFNDFNTALDKALGKLGSQPKQVI 358
L++ GK + E+ P F A AL G K+ I
Sbjct: 313 ELMEQGKFVAPNHEMVPLAKFKDAAAAALSFKGFTGKKYI 352
>ref|NP_001017852.1| hypothetical protein LOC550550 [Danio rerio]
gi|62203488|gb|AAH92759.1| Hypothetical protein
LOC550550 [Danio rerio]
Length = 377
Score = 249 bits (637), Expect = 7e-65
Identities = 139/319 (43%), Positives = 188/319 (58%), Gaps = 13/319 (4%)
Query: 42 AIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAVG 101
A++Y +HG+P V +L + +V + VKMLAAPINPSD+N +QG Y + PE PAVG
Sbjct: 47 ALLYRNHGEPSQVVQLESLDLPQVGAECVLVKMLAAPINPSDLNMLQGTYAILPELPAVG 106
Query: 102 GYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKVNKGVPMEYAATI 161
G EGV +V VG V GDWVIP GTW+T V + + K +P+ AAT+
Sbjct: 107 GNEGVAQVMEVGDKVKTLKVGDWVIPKDAGIGTWRTAAVLKADDLVTLPKDIPVLSAATL 166
Query: 162 TVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRDRPGVGEV 221
VNP TA ML D +GD ++QN A S VGQ VIQ+A ++GIH IN+IRDRP + ++
Sbjct: 167 GVNPCTAYRMLTDFEEPKAGDTVIQNAANSGVGQAVIQIAAAKGIHTINVIRDRPDLRQL 226
Query: 222 KERLKDLGADEVFTESELEVKNVKSLLGGIPEPALGFNCVGGNSASLVLKFLRRGGTMVT 281
+RL +GA V TE L +K L P P L N VGG SA+ +L+ L+ GG++VT
Sbjct: 227 SDRLTAMGATHVITEETLRRPEMKELFKSCPRPKLALNGVGGKSATELLRHLQSGGSLVT 286
Query: 282 YGGMSKKPVTVSTSSFIFKN------WLSTDKA------EEGRRMIDRLLGLVQDGKLKY 329
YGGM+K+PVTV S+ IFK+ W++ K E R M+D L L++ GKL
Sbjct: 287 YGGMAKQPVTVPVSALIFKDVRVRGFWVTQWKRDNRHDDEALRHMLDELCILIRAGKLSA 346
Query: 330 KM-ELTPFNDFNTALDKAL 347
+ DF AL+ A+
Sbjct: 347 PICTSVQLQDFRKALENAM 365
>ref|XP_513261.1| PREDICTED: hypothetical protein XP_513261 [Pan troglodytes]
Length = 584
Score = 244 bits (624), Expect = 2e-63
Identities = 134/290 (46%), Positives = 181/290 (62%), Gaps = 11/290 (3%)
Query: 74 MLAAPINPSDINRIQGVYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFG 133
MLAAPINPSDIN IQG Y + PE PAVGG EGV +V +VGS VT PGDWVIP+ G
Sbjct: 1 MLAAPINPSDINMIQGNYGLLPELPAVGGNEGVAQVVAVGSNVTGLKPGDWVIPANAGLG 60
Query: 134 TWQTYIVKDQNVWHKVNKGVPMEYAATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMV 193
TW+T V + +V +P++ AAT+ VNP TA ML D L GD+++QN + S V
Sbjct: 61 TWRTEAVFSEEALIQVPSDIPLQSAATLGVNPCTAYRMLMDFEQLQPGDSVIQNASNSGV 120
Query: 194 GQCVIQLAKSRGIHNINIIRDRPGVGEVKERLKDLGADEVFTESELEVKNVKSLLGGIPE 253
GQ VIQ+A + G+ IN++RDRP + ++ +RLK LGA+ V TE EL +K+ +P+
Sbjct: 121 GQAVIQIAAALGLRTINVVRDRPDIQKLSDRLKSLGAEHVITEEELRRPKMKNFFKDMPQ 180
Query: 254 PALGFNCVGGNSASLVLKFLRRGGTMVTYGGMSKKPVTVSTSSFIFKN------WLSTDK 307
P L NCVGG S++ +L+ L RGGTMVTYGGM+K+PV S S IFK+ WLS K
Sbjct: 181 PRLALNCVGGKSSTELLRQLARGGTMVTYGGMAKQPVVASVSLLIFKDLKLRGFWLSQWK 240
Query: 308 AEEG----RRMIDRLLGLVQDGKLKY-KMELTPFNDFNTALDKALGKLGS 352
+ + +I L L++ G+L P D+ +AL+ ++ L S
Sbjct: 241 KDHSPDQFKELILTLCDLIRRGQLTAPACSQVPLQDYQSALEASMKPLFS 290
>gb|AAX51660.1| AT25977p [Drosophila melanogaster]
Length = 325
Score = 244 bits (622), Expect = 4e-63
Identities = 133/271 (49%), Positives = 179/271 (65%), Gaps = 3/271 (1%)
Query: 32 FSSAVSPPSKAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVY 91
+S +S +K++ Y HG+P V +LV+ + K+N + VK+LAAPINP+DIN IQG Y
Sbjct: 15 WSRQMSVVAKSLKYTQHGEPQEVLQLVEDKLPDPKDNQVLVKILAAPINPADINTIQGKY 74
Query: 92 PVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKVNK 151
PV+P+ PAVGG E V EV VG V F G VIP GTW T+ V ++ V+K
Sbjct: 75 PVKPKFPAVGGNECVAEVICVGDKVKGFEAGQHVIPLASGLGTWTTHAVYKEDQLLIVSK 134
Query: 152 GVPMEYAATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINI 211
V + AAT TVNP TA ML+D V L GD ++QNGA S VGQ V QL ++ GI+++ I
Sbjct: 135 KVGLAEAATSTVNPTTAYRMLKDFVQLCPGDTVIQNGANSAVGQAVHQLCRAWGINSVGI 194
Query: 212 IRDRPGVGEVKERLKDLGADEVFTESELEVKNV-KSLLGGIPEPALGFNCVGGNSASLVL 270
+RDRP + E+K+ L+ LGA EV TE+E+ ++ KS G + +P L FNCVGG SA+ V
Sbjct: 195 VRDRPEIAELKQMLQCLGATEVLTEAEIRTSDIFKS--GKLKKPRLAFNCVGGKSATEVS 252
Query: 271 KFLRRGGTMVTYGGMSKKPVTVSTSSFIFKN 301
+ L GG +VTYGGMS++PVTV+T IFK+
Sbjct: 253 RHLDNGGVLVTYGGMSREPVTVATGPLIFKD 283
>emb|CAI14329.1| OTTHUMP00000065099 [Homo sapiens] gi|67078406|ref|NP_001019903.1|
nuclear receptor-binding factor 1 isoform b [Homo
sapiens]
Length = 297
Score = 243 bits (620), Expect = 6e-63
Identities = 132/285 (46%), Positives = 178/285 (62%), Gaps = 11/285 (3%)
Query: 74 MLAAPINPSDINRIQGVYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFG 133
MLAAPINPSDIN IQG Y PE PAVGG EGV +V +VGS VT PGDWVIP+ G
Sbjct: 1 MLAAPINPSDINMIQGNYGFLPELPAVGGNEGVAQVVAVGSNVTGLKPGDWVIPANAGLG 60
Query: 134 TWQTYIVKDQNVWHKVNKGVPMEYAATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMV 193
TW+T V + +V +P++ AAT+ VNP TA ML D L GD+++QN + S V
Sbjct: 61 TWRTEAVFSEEALIQVPSDIPLQSAATLGVNPCTAYRMLMDFEQLQPGDSVIQNASNSGV 120
Query: 194 GQCVIQLAKSRGIHNINIIRDRPGVGEVKERLKDLGADEVFTESELEVKNVKSLLGGIPE 253
GQ VIQ+A + G+ IN++RDRP + ++ +RLK LGA+ V TE EL +K+ +P+
Sbjct: 121 GQAVIQIAAALGLRTINVVRDRPDIQKLSDRLKSLGAEHVITEEELRRPEMKNFFKDMPQ 180
Query: 254 PALGFNCVGGNSASLVLKFLRRGGTMVTYGGMSKKPVTVSTSSFIFKN------WLSTDK 307
P L NCVGG S++ +L+ L RGGTMVTYGGM+K+PV S S IFK+ WLS K
Sbjct: 181 PRLALNCVGGKSSTELLRQLARGGTMVTYGGMAKQPVVASVSLLIFKDLKLRGFWLSQWK 240
Query: 308 AEEG----RRMIDRLLGLVQDGKLKY-KMELTPFNDFNTALDKAL 347
+ + +I L L++ G+L P D+ +AL+ ++
Sbjct: 241 KDHSPDQFKELILTLCDLIRRGQLTAPACSQVPLQDYQSALEASM 285
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.316 0.135 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 636,774,901
Number of Sequences: 2540612
Number of extensions: 27908107
Number of successful extensions: 92530
Number of sequences better than 10.0: 2436
Number of HSP's better than 10.0 without gapping: 895
Number of HSP's successfully gapped in prelim test: 1542
Number of HSP's that attempted gapping in prelim test: 90326
Number of HSP's gapped (non-prelim): 2722
length of query: 360
length of database: 863,360,394
effective HSP length: 129
effective length of query: 231
effective length of database: 535,621,446
effective search space: 123728554026
effective search space used: 123728554026
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 76 (33.9 bits)
Medicago: description of AC135230.4


