

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134521.1 - phase: 0 /pseudo
(324 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
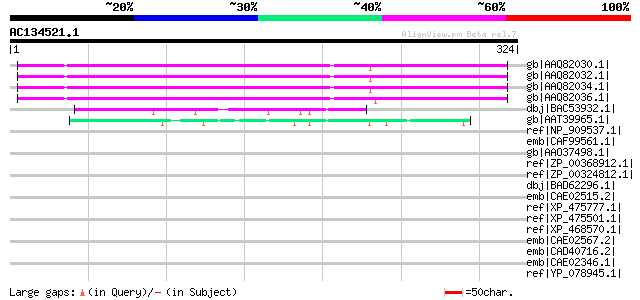
Score E
Sequences producing significant alignments: (bits) Value
gb|AAQ82030.1| unknown [Pisum sativum] 224 3e-57
gb|AAQ82032.1| unknown [Pisum sativum] 223 8e-57
gb|AAQ82034.1| unknown [Pisum sativum] 221 3e-56
gb|AAQ82036.1| unknown [Pisum sativum] 214 2e-54
dbj|BAC53932.1| hypothetical protein [Nicotiana tabacum] 54 6e-06
gb|AAT39965.1| hypothetical protein [Solanum demissum] 50 9e-05
ref|NP_909537.1| Putative mutator-like transposase [Oryza sativa... 36 1.3
emb|CAF99561.1| unnamed protein product [Tetraodon nigroviridis] 36 1.8
gb|AAO37498.1| retrotransposon protein, putative, Ty3-gypsy sub-... 35 2.3
ref|ZP_00368912.1| biotin sulfoxide reductase (bisC) [Campylobac... 35 3.0
ref|ZP_00324812.1| COG0384: Predicted epimerase, PhzC/PhzF homol... 35 3.9
dbj|BAD62296.1| Epstein-Barr virus EBNA-1-like protein [Oryza sa... 35 3.9
emb|CAE02515.2| OSJNBb0003A12.2 [Oryza sativa (japonica cultivar... 34 5.1
ref|XP_475777.1| unknown protein [Oryza sativa (japonica cultiva... 34 5.1
ref|XP_475501.1| putative mutator-like transposase [Oryza sativa... 34 5.1
ref|XP_468570.1| Putative mutator-like transposase [Oryza sativa... 34 5.1
emb|CAE02567.2| OSJNBa0006M15.10 [Oryza sativa (japonica cultiva... 34 5.1
emb|CAD40716.2| OSJNBb0042I07.13 [Oryza sativa (japonica cultiva... 34 5.1
emb|CAE02346.1| OSJNBb0072M01.7 [Oryza sativa (japonica cultivar... 34 5.1
ref|YP_078945.1| carbamoyl-phosphate synthetase (catalytic subun... 34 6.7
>gb|AAQ82030.1| unknown [Pisum sativum]
Length = 562
Score = 224 bits (571), Expect = 3e-57
Identities = 133/317 (41%), Positives = 182/317 (56%), Gaps = 7/317 (2%)
Query: 6 RRTHGYKFKAPDTESLEGLRGLTKMLKNPGHFQERYGNLLDILKTKVDVMLLNTLVQFYD 65
R+T Y F SL L L G F ++YG++L +LK VD + L TL+QFYD
Sbjct: 25 RKTCSYSFHREPLTSLIELSNLMTSGNQKG-FVDQYGDILTLLKMVVDPVPLQTLLQFYD 83
Query: 66 PIYHGFTFPDFQLFPTLEEYSHWVGLPVLDEVPFHAFGPIPKIPTIAKALHLEVADIKDK 125
P H FTF D+QL PTLEEYS + +PV +VPF +A+ALHL + ++ D
Sbjct: 84 PELHCFTFQDYQLAPTLEEYSILLSVPVRYQVPFLDVPKEVDFRVVARALHLGIKEVSDN 143
Query: 126 FTTKDGLPCLPYNFLHQKATTCFEKSKTDDFESILALLIYGIVLFPKVDKFVDMNAIRIF 185
+ + + LP FL + A EK + F + LA++IYGIVLFP + FVD A+ IF
Sbjct: 144 WKSSGDVVGLPLKFLLRVAREEAEKGSWEAFHAQLAVMIYGIVLFPSMPNFVDYAAVSIF 203
Query: 186 LTQNPVPVLLADTYVAIHDRTSHGK-GTIVCCTPLLHRWITSHLP--RPSIRPEPI-PWS 241
+ NPVP LLADTY AIH R HGK G I CC PLL RW S LP P + + W+
Sbjct: 204 IGGNPVPTLLADTYYAIHSR--HGKGGAIRCCLPLLLRWFLSLLPVSGPFVDAQSTHKWT 261
Query: 242 QKLMTWTPKDIIWFNPSCDPEFIIDSCGDFNNVPLLGTQGGISYSPILARRQFEYYTEMK 301
Q++M+ T DI W + D +I SCG F NVPL+GT+G I+Y+P+L+ RQ + + +
Sbjct: 262 QRVMSLTSYDIRWQSYRMDVRNVIMSCGGFRNVPLVGTRGCINYNPVLSLRQLGFVMKGR 321
Query: 302 PVYLILDRDFFLYKKDD 318
P+ + + K+ D
Sbjct: 322 PLEAEIAESVYFEKQSD 338
>gb|AAQ82032.1| unknown [Pisum sativum]
Length = 562
Score = 223 bits (567), Expect = 8e-57
Identities = 133/317 (41%), Positives = 181/317 (56%), Gaps = 7/317 (2%)
Query: 6 RRTHGYKFKAPDTESLEGLRGLTKMLKNPGHFQERYGNLLDILKTKVDVMLLNTLVQFYD 65
R+T Y F SL L L G F ++YG++L +LK VD + L TL+QFYD
Sbjct: 25 RKTCSYSFHREPLTSLIELSNLMTSGNQKG-FVDQYGDILTLLKMVVDPVPLQTLLQFYD 83
Query: 66 PIYHGFTFPDFQLFPTLEEYSHWVGLPVLDEVPFHAFGPIPKIPTIAKALHLEVADIKDK 125
P H FTF D+QL PTLEEYS + +PV +VPF +A+ALHL + ++ D
Sbjct: 84 PELHCFTFQDYQLAPTLEEYSILLSVPVRYQVPFLDVPKEVDFRVVARALHLGIKEVSDS 143
Query: 126 FTTKDGLPCLPYNFLHQKATTCFEKSKTDDFESILALLIYGIVLFPKVDKFVDMNAIRIF 185
+ + + LP FL + A EK + F + LA++IYGIVLFP + FVD A+ IF
Sbjct: 144 WKSSGDVVGLPLKFLLRVAREEAEKGSWEAFHAQLAVMIYGIVLFPSMPNFVDYAAVSIF 203
Query: 186 LTQNPVPVLLADTYVAIHDRTSHGK-GTIVCCTPLLHRWITSHLP--RPSIRPEPI-PWS 241
+ NPVP LLADTY AIH R HGK G I CC PLL RW S LP P + + W+
Sbjct: 204 IGGNPVPTLLADTYYAIHSR--HGKGGAIRCCLPLLLRWFLSLLPVSGPFVDAQSTHKWT 261
Query: 242 QKLMTWTPKDIIWFNPSCDPEFIIDSCGDFNNVPLLGTQGGISYSPILARRQFEYYTEMK 301
Q++M+ T DI W + D +I SCG F NVPL+GT+G I+Y+P+L+ RQ + + +
Sbjct: 262 QRVMSLTSYDIRWQSYRMDVRNVIMSCGGFRNVPLIGTRGCINYNPVLSLRQLGFVMKGR 321
Query: 302 PVYLILDRDFFLYKKDD 318
P+ + K+ D
Sbjct: 322 PLEAEVAESVCFEKRSD 338
>gb|AAQ82034.1| unknown [Pisum sativum]
Length = 562
Score = 221 bits (562), Expect = 3e-56
Identities = 132/317 (41%), Positives = 181/317 (56%), Gaps = 7/317 (2%)
Query: 6 RRTHGYKFKAPDTESLEGLRGLTKMLKNPGHFQERYGNLLDILKTKVDVMLLNTLVQFYD 65
R+T Y F SL L L G F +++G++L +LK VD + L TL+QFYD
Sbjct: 25 RKTCSYSFHREPLTSLIELSNLMTSGNQKG-FVDQHGDILTLLKMVVDPVPLQTLLQFYD 83
Query: 66 PIYHGFTFPDFQLFPTLEEYSHWVGLPVLDEVPFHAFGPIPKIPTIAKALHLEVADIKDK 125
P H FTF D+QL PTLEEYS + +PV +VPF +A+ALHL + ++ D
Sbjct: 84 PELHCFTFQDYQLAPTLEEYSILLSVPVRYQVPFLDVPKEVDFGVVARALHLGIKEVSDS 143
Query: 126 FTTKDGLPCLPYNFLHQKATTCFEKSKTDDFESILALLIYGIVLFPKVDKFVDMNAIRIF 185
+ + LP FL + A EK + F + LA++IYGIVLFP + FVD A+ IF
Sbjct: 144 WKYSGDVVGLPLKFLLRVAREEAEKGSWEAFRAQLAVMIYGIVLFPSMPNFVDYAAVSIF 203
Query: 186 LTQNPVPVLLADTYVAIHDRTSHGK-GTIVCCTPLLHRWITSHLP--RPSIRPEPI-PWS 241
+ NPVP LLADTY AIH R HGK G I CC PLL RW S LP P + + W+
Sbjct: 204 IGGNPVPTLLADTYYAIHSR--HGKGGAIRCCLPLLLRWFLSLLPVSGPFVDAQSTHKWT 261
Query: 242 QKLMTWTPKDIIWFNPSCDPEFIIDSCGDFNNVPLLGTQGGISYSPILARRQFEYYTEMK 301
Q++M+ T DI W + D +I SCG F NVPL+GT+G I+Y+P+L+ RQ + + +
Sbjct: 262 QRVMSLTSYDIRWQSYRMDVRNVIMSCGGFRNVPLVGTRGCINYNPVLSLRQLGFVMKGR 321
Query: 302 PVYLILDRDFFLYKKDD 318
P+ + + K+ D
Sbjct: 322 PLEAEVAESVYFEKQSD 338
>gb|AAQ82036.1| unknown [Pisum sativum]
Length = 546
Score = 214 bits (546), Expect = 2e-54
Identities = 128/317 (40%), Positives = 178/317 (55%), Gaps = 7/317 (2%)
Query: 6 RRTHGYKFKAPDTESLEGLRGLTKMLKNPGHFQERYGNLLDILKTKVDVMLLNTLVQFYD 65
R+T Y F SL L L + G F ++G++L +LKT VD + L TL+QFYD
Sbjct: 9 RKTCSYSFYREPLTSLIELGSLMPSDQLKG-FVGQFGDILTLLKTVVDPVPLQTLLQFYD 67
Query: 66 PIYHGFTFPDFQLFPTLEEYSHWVGLPVLDEVPFHAFGPIPKIPTIAKALHLEVADIKDK 125
P FTF D+QL PTLEEYS + +P+ +VPF IA+AL + V ++ D
Sbjct: 68 PELRCFTFQDYQLAPTLEEYSILMSIPIQHQVPFLDVPKEVDFRVIARALRMSVKEVCDN 127
Query: 126 FTTKDGLPCLPYNFLHQKATTCFEKSKTDDFESILALLIYGIVLFPKVDKFVDMNAIRIF 185
+ + +P FL + A EK + F + LA +IYGIVLFP + F+D AI IF
Sbjct: 128 WKPSGEVVGMPLKFLLRVAREEAEKGNWEVFHAQLAAMIYGIVLFPSMPNFIDHAAISIF 187
Query: 186 LTQNPVPVLLADTYVAIHDRTSHGK-GTIVCCTPLLHRWITSHLPRPS---IRPEPIPWS 241
+ NPVP LLADTY AIH R HGK G I CC PLL RW S LP + W+
Sbjct: 188 IGGNPVPTLLADTYYAIHSR--HGKGGAIRCCLPLLLRWFMSLLPASGPFMDTQSTLKWT 245
Query: 242 QKLMTWTPKDIIWFNPSCDPEFIIDSCGDFNNVPLLGTQGGISYSPILARRQFEYYTEMK 301
Q++M+ T DI W + D + ++ SCG+F NVPL+GT+G I+Y+P+L+ RQ + +
Sbjct: 246 QRVMSLTSYDIRWQSYRMDVKDVVMSCGEFRNVPLVGTKGCINYNPVLSLRQLGFIMSRR 305
Query: 302 PVYLILDRDFFLYKKDD 318
P+ + + K+ D
Sbjct: 306 PLEAEIVESVYFEKRTD 322
>dbj|BAC53932.1| hypothetical protein [Nicotiana tabacum]
Length = 464
Score = 53.9 bits (128), Expect = 6e-06
Identities = 53/209 (25%), Positives = 85/209 (40%), Gaps = 29/209 (13%)
Query: 42 GNLLDILKTKVDVMLLNTLVQFYDPIYHGFTFPDFQLFPTLEEYSHWVG---------LP 92
G L DI+K K L+ LV F+DP+++ F F DF+L PTLEE + G L
Sbjct: 39 GALTDIMKIKPRDDLIEALVTFWDPVHNVFCFSDFELTPTLEEIDGYSGFGRDLRNQELI 98
Query: 93 VLDEVPFHAFGPIPKIPTIAKALHL-----EVADIKDKFTTKDGLPCLPYNFLHQKATTC 147
+ H F + I + ++ + +F +G +H+K
Sbjct: 99 FPRALSVHRFFDLLNISKQIRKTNVVKGCCSFYFLYSRFGQPNGFE------MHEKGLNN 152
Query: 148 FEKSKTDDFESILALLI--YGIVLFPKVDKFVDMNAIRI--FLTQNP----VPVLLADTY 199
+ T A ++ GI++FP ++ +D R+ LT P++L+D Y
Sbjct: 153 KQNKDTWHIHRCFAFIMAFLGIMVFPNRERTIDTRIARVVQVLTTKEHHTLAPIILSDIY 212
Query: 200 VAIHDRTSHGKGTIVCCTPLLHRWITSHL 228
A+ G C LL W+ HL
Sbjct: 213 RAL-TLCKSGAKFFEGCNILLQMWLIEHL 240
>gb|AAT39965.1| hypothetical protein [Solanum demissum]
Length = 607
Score = 50.1 bits (118), Expect = 9e-05
Identities = 79/311 (25%), Positives = 121/311 (38%), Gaps = 66/311 (21%)
Query: 39 ERYGNLLDILKTKVDVMLLNTLVQFYDPIYHGFTFPDFQLFPTLEEYSHWVGL-PVLDE- 96
+R G L +L + L+ LV F+DPI + F F DF+L PTLEE ++G L E
Sbjct: 46 KRLGFLRSLLYVEPRRDLIEALVHFWDPIKNVFHFSDFELTPTLEEIGAFIGRGKNLHEG 105
Query: 97 ---VPFHAFGPIPKIPTIAKALHLEVADI--------------KDKFTTKDGLPCLPYNF 139
+P H G + LH+ +I ++ KDG L
Sbjct: 106 EPMIPKHING-----RRFLELLHINENEIGGCLDNGWVPLEFLYKRYGRKDGFE-LYGKK 159
Query: 140 LHQKATTCFEKSKTDDFESILALLIYGIVLFPKVDKFVDMN--AIRIFLTQNP----VPV 193
LH C +T +++I + GI++F K +++N A+ + P VP+
Sbjct: 160 LHNNG--CRLTWETHSYDAITVAFL-GIMVFLKRGGKININLAAVITAFEKKPNITLVPM 216
Query: 194 LLADTYVAIHDRTSHGKGTIVCCTPLLHRWITSHL-------------------PRPSIR 234
+LAD Y A+ +G C LL W+ H+ I+
Sbjct: 217 ILADIYRAL-TICKNGGDYFEGCNMLLQLWMIEHIRHHPYVVDFKVECNDYIGGHEERIK 275
Query: 235 PEPIP-----WSQKLMTWTPKDIIWFNPSCDPEFIIDSCGDFNN-VPLLGTQGGISYSPI 288
P W + L T I+W N P + F + + L+G +G Y P+
Sbjct: 276 DHSFPKGIEAWKKYLNNLTADKIVW-NYHWFPSAEVIYMSTFRSFIVLMGLRGVQPYMPL 334
Query: 289 -----LARRQF 294
L RRQF
Sbjct: 335 RVMRQLGRRQF 345
>ref|NP_909537.1| Putative mutator-like transposase [Oryza sativa (japonica
cultivar-group)] gi|19697447|gb|AAL93082.1| Putative
mutator-like transposase [Oryza sativa (japonica
cultivar-group)]
Length = 1530
Score = 36.2 bits (82), Expect = 1.3
Identities = 21/54 (38%), Positives = 31/54 (56%), Gaps = 4/54 (7%)
Query: 52 VDVMLLNTLVQFYDPIYHGFTFPDFQLFPTLEEYSHWVGLPVLDEVPFHAFGPI 105
VD LL TLV + P H F P ++ PTL++ S+ +GLP+ + A GP+
Sbjct: 965 VDRSLLATLVDRWRPETHTFHLPCGEVAPTLQDVSYLLGLPLAGD----AVGPV 1014
>emb|CAF99561.1| unnamed protein product [Tetraodon nigroviridis]
Length = 391
Score = 35.8 bits (81), Expect = 1.8
Identities = 33/113 (29%), Positives = 45/113 (39%), Gaps = 10/113 (8%)
Query: 2 EVVGRRTHGYKFKAPDTESLEGLRGLTKMLKNPGHFQERYGNLLDILKTKVDVMLLNTLV 61
E+ GRR + P S +G G + GHFQ + I+KT V LLNTLV
Sbjct: 18 ELEGRRVAFILYLVPPWRSSDG--GTLDLYSTDGHFQPQ-----SIVKTLVP--LLNTLV 68
Query: 62 QF-YDPIYHGFTFPDFQLFPTLEEYSHWVGLPVLDEVPFHAFGPIPKIPTIAK 113
F P+ S W P L+ P H P+P+ P + +
Sbjct: 69 LFEVSPVSFHQVAEVLTKDKCRLSLSGWFHGPCLERPPHHTEAPVPRSPHLPR 121
>gb|AAO37498.1| retrotransposon protein, putative, Ty3-gypsy sub-class [Oryza
sativa (japonica cultivar-group)]
gi|50916509|ref|XP_468651.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 966
Score = 35.4 bits (80), Expect = 2.3
Identities = 35/130 (26%), Positives = 56/130 (42%), Gaps = 16/130 (12%)
Query: 52 VDVMLLNTLVQFYDPIYHGFTFPDFQLFPTLEEYSHWVGLPVLDEVPFHAFGPI------ 105
VD LL LV + P H F P ++ PTL++ S+ +GLP+ + A GP+
Sbjct: 405 VDRSLLAALVDRWRPETHTFHLPCGEVAPTLQDVSYLLGLPLAGD----AVGPVTTGVDW 460
Query: 106 -----PKIPTIAKALHLEVADIKDKFTTKDGLPCLPYNFLHQKATTCFEKSKTDDFESIL 160
+ + +A HL + + T L + Q E S + E+ L
Sbjct: 461 KDDLTARFTPVHRAPHLPLQPLAHHRNTGPTKRWLLQFIVEQLQAEADEYSYSRCLEAYL 520
Query: 161 ALLIYGIVLF 170
LL++G V+F
Sbjct: 521 -LLLFGWVMF 529
>ref|ZP_00368912.1| biotin sulfoxide reductase (bisC) [Campylobacter lari RM2100]
gi|57018583|gb|EAL55357.1| biotin sulfoxide reductase
(bisC) [Campylobacter lari RM2100]
Length = 799
Score = 35.0 bits (79), Expect = 3.0
Identities = 15/48 (31%), Positives = 25/48 (51%), Gaps = 1/48 (2%)
Query: 61 VQFYDPIYHGFTFPDFQLFPTLEEYSHWVG-LPVLDEVPFHAFGPIPK 107
+Q Y P + + DF+ PT E + W+G ++ + PFH P P+
Sbjct: 615 IQIYSPKFEKYNLEDFKAHPTWFEPAEWLGNEKLVKKYPFHLLSPHPR 662
>ref|ZP_00324812.1| COG0384: Predicted epimerase, PhzC/PhzF homolog [Trichodesmium
erythraeum IMS101]
Length = 290
Score = 34.7 bits (78), Expect = 3.9
Identities = 23/93 (24%), Positives = 51/93 (54%), Gaps = 9/93 (9%)
Query: 107 KIPTIAKALHLEVADIKDKFTTKDGLPCLPYNFLHQKATTCFEKSKTD---DFESILALL 163
++ T+A+ L+LEV DI +F ++ +P+ + K+ +K+K + ++ I
Sbjct: 129 QVETVAEVLNLEVVDIDTRFPIQEVSTGIPFIIVPLKSLVALKKAKVNLDKYYQFIQTTK 188
Query: 164 IYGIVLF-PKVDKFVDMNAIRIF-----LTQNP 190
+ I++F P+ K +D ++R+F +T++P
Sbjct: 189 VTEILIFCPETYKDIDDLSVRMFAPYLGITEDP 221
>dbj|BAD62296.1| Epstein-Barr virus EBNA-1-like protein [Oryza sativa (japonica
cultivar-group)]
Length = 93
Score = 34.7 bits (78), Expect = 3.9
Identities = 24/68 (35%), Positives = 32/68 (46%), Gaps = 4/68 (5%)
Query: 41 YGNLLDILKTKVDVMLLNTLVQFYDPIYHGFTFPDFQLFPTLEEYSHWVGLPVLDEVPFH 100
+GNLL I K +D L N + + YD FT + TL++ +GLP E F
Sbjct: 22 FGNLLKINKIYIDRNLCNAITRSYDKEKKAFTINGTFVMMTLDDVDCLLGLPSKGEEIFE 81
Query: 101 AFGPIPKI 108
A PKI
Sbjct: 82 A----PKI 85
>emb|CAE02515.2| OSJNBb0003A12.2 [Oryza sativa (japonica cultivar-group)]
gi|38347084|emb|CAD39471.2| OSJNBa0001M07.4 [Oryza
sativa (japonica cultivar-group)]
gi|50930333|ref|XP_474694.1| OSJNBa0001M07.4 [Oryza
sativa (japonica cultivar-group)]
Length = 1031
Score = 34.3 bits (77), Expect = 5.1
Identities = 30/98 (30%), Positives = 42/98 (42%), Gaps = 19/98 (19%)
Query: 53 DVMLLNTLVQFYDPIYHGFTFPDFQLFPTLEEYSHWVGLPVLDEVPFHAFGPIPKIPTIA 112
D LL LV + P H F P ++ PTL++ S+ +GLP+ + A GP+
Sbjct: 426 DRSLLAALVDRWRPETHTFHLPCGEVAPTLQDVSYLLGLPLAGD----AVGPVDM----- 476
Query: 113 KALHLEVADIKDKFTTK----DGLPCLPYNFLHQKATT 146
D KD+ T + LP LP L Q T
Sbjct: 477 ------GVDWKDELTARFAPVQRLPGLPLEPLQQHRNT 508
>ref|XP_475777.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|47900426|gb|AAT39220.1| unknown protein [Oryza sativa
(japonica cultivar-group)]
Length = 896
Score = 34.3 bits (77), Expect = 5.1
Identities = 20/54 (37%), Positives = 30/54 (55%), Gaps = 4/54 (7%)
Query: 52 VDVMLLNTLVQFYDPIYHGFTFPDFQLFPTLEEYSHWVGLPVLDEVPFHAFGPI 105
VD LL LV + P H F P ++ PTL++ S+ +GLP+ + A GP+
Sbjct: 291 VDRSLLAALVDRWRPETHTFHLPCGEVAPTLQDVSYLLGLPLAGD----AVGPV 340
>ref|XP_475501.1| putative mutator-like transposase [Oryza sativa (japonica
cultivar-group)] gi|46981280|gb|AAT07598.1| putative
mutator-like transposase [Oryza sativa (japonica
cultivar-group)]
Length = 1725
Score = 34.3 bits (77), Expect = 5.1
Identities = 30/98 (30%), Positives = 42/98 (42%), Gaps = 19/98 (19%)
Query: 53 DVMLLNTLVQFYDPIYHGFTFPDFQLFPTLEEYSHWVGLPVLDEVPFHAFGPIPKIPTIA 112
D LL LV + P H F P ++ PTL++ S+ +GLP+ + A GP+
Sbjct: 1122 DRSLLAALVDRWRPETHTFHLPCGEVAPTLQDVSYLLGLPLAGD----AVGPVDM----- 1172
Query: 113 KALHLEVADIKDKFTTK----DGLPCLPYNFLHQKATT 146
D KD+ T + LP LP L Q T
Sbjct: 1173 ------GVDWKDELTARFAPVQRLPGLPLEPLQQHRNT 1204
>ref|XP_468570.1| Putative mutator-like transposase [Oryza sativa (japonica
cultivar-group)] gi|24431599|gb|AAN61479.1| Putative
mutator-like transposase [Oryza sativa (japonica
cultivar-group)]
Length = 1596
Score = 34.3 bits (77), Expect = 5.1
Identities = 30/98 (30%), Positives = 42/98 (42%), Gaps = 19/98 (19%)
Query: 53 DVMLLNTLVQFYDPIYHGFTFPDFQLFPTLEEYSHWVGLPVLDEVPFHAFGPIPKIPTIA 112
D LL LV + P H F P ++ PTL++ S+ +GLP+ + A GP+
Sbjct: 956 DRSLLAALVDRWRPETHTFHLPCGEVAPTLQDVSYLLGLPLAGD----AVGPVDM----- 1006
Query: 113 KALHLEVADIKDKFTTK----DGLPCLPYNFLHQKATT 146
D KD+ T + LP LP L Q T
Sbjct: 1007 ------GVDWKDELTARFAPVQRLPGLPLEPLQQHRNT 1038
>emb|CAE02567.2| OSJNBa0006M15.10 [Oryza sativa (japonica cultivar-group)]
gi|32489202|emb|CAE04387.1| OSJNBa0027G07.29 [Oryza
sativa (japonica cultivar-group)]
gi|50924718|ref|XP_472713.1| OSJNBa0027G07.29 [Oryza
sativa (japonica cultivar-group)]
Length = 1620
Score = 34.3 bits (77), Expect = 5.1
Identities = 30/98 (30%), Positives = 42/98 (42%), Gaps = 19/98 (19%)
Query: 53 DVMLLNTLVQFYDPIYHGFTFPDFQLFPTLEEYSHWVGLPVLDEVPFHAFGPIPKIPTIA 112
D LL LV + P H F P ++ PTL++ S+ +GLP+ + A GP+
Sbjct: 956 DRSLLAALVDRWRPETHTFHLPCGEVAPTLQDVSYLLGLPLAGD----AVGPVDM----- 1006
Query: 113 KALHLEVADIKDKFTTK----DGLPCLPYNFLHQKATT 146
D KD+ T + LP LP L Q T
Sbjct: 1007 ------GVDWKDELTARFAPVQRLPGLPLEPLQQHRNT 1038
>emb|CAD40716.2| OSJNBb0042I07.13 [Oryza sativa (japonica cultivar-group)]
gi|50923575|ref|XP_472148.1| OSJNBb0042I07.13 [Oryza
sativa (japonica cultivar-group)]
Length = 1596
Score = 34.3 bits (77), Expect = 5.1
Identities = 30/98 (30%), Positives = 42/98 (42%), Gaps = 19/98 (19%)
Query: 53 DVMLLNTLVQFYDPIYHGFTFPDFQLFPTLEEYSHWVGLPVLDEVPFHAFGPIPKIPTIA 112
D LL LV + P H F P ++ PTL++ S+ +GLP+ + A GP+
Sbjct: 956 DRSLLAALVDRWRPETHTFHLPCGEVAPTLQDVSYLLGLPLAGD----AVGPVDM----- 1006
Query: 113 KALHLEVADIKDKFTTK----DGLPCLPYNFLHQKATT 146
D KD+ T + LP LP L Q T
Sbjct: 1007 ------GVDWKDELTARFAPVQRLPGLPLEPLQQHRNT 1038
>emb|CAE02346.1| OSJNBb0072M01.7 [Oryza sativa (japonica cultivar-group)]
gi|38345695|emb|CAD41115.2| OSJNBb0070J16.11 [Oryza
sativa (japonica cultivar-group)]
gi|50926446|ref|XP_473170.1| OSJNBb0070J16.11 [Oryza
sativa (japonica cultivar-group)]
Length = 1416
Score = 34.3 bits (77), Expect = 5.1
Identities = 20/54 (37%), Positives = 30/54 (55%), Gaps = 4/54 (7%)
Query: 52 VDVMLLNTLVQFYDPIYHGFTFPDFQLFPTLEEYSHWVGLPVLDEVPFHAFGPI 105
VD LL LV + P H F P ++ PTL++ S+ +GLP+ + A GP+
Sbjct: 746 VDRSLLAALVDRWRPETHTFHLPCGEVAPTLQDVSYLLGLPLAGD----AVGPV 795
>ref|YP_078945.1| carbamoyl-phosphate synthetase (catalytic subunit) [Bacillus
licheniformis ATCC 14580] gi|52003365|gb|AAU23307.1|
carbamoyl-phosphate synthetase (catalytic subunit)
[Bacillus licheniformis ATCC 14580]
gi|52785531|ref|YP_091360.1| PyrAB [Bacillus
licheniformis ATCC 14580] gi|52348033|gb|AAU40667.1|
PyrAB [Bacillus licheniformis DSM 13]
Length = 1070
Score = 33.9 bits (76), Expect = 6.7
Identities = 21/70 (30%), Positives = 33/70 (47%), Gaps = 1/70 (1%)
Query: 3 VVGRRTHGYKFKAPDTESLEG-LRGLTKMLKNPGHFQERYGNLLDILKTKVDVMLLNTLV 61
++ +R H +K TE G LR + +K G E NLLD+++ ++NTL
Sbjct: 952 LIAKRFHAIGYKILATEGTAGYLRDASVPVKTVGKIGEEGTNLLDVIRNGEAQFVVNTLT 1011
Query: 62 QFYDPIYHGF 71
+ P GF
Sbjct: 1012 KGKQPARDGF 1021
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.324 0.144 0.458
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 613,028,211
Number of Sequences: 2540612
Number of extensions: 28580886
Number of successful extensions: 47227
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 47206
Number of HSP's gapped (non-prelim): 22
length of query: 324
length of database: 863,360,394
effective HSP length: 128
effective length of query: 196
effective length of database: 538,162,058
effective search space: 105479763368
effective search space used: 105479763368
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 75 (33.5 bits)
Medicago: description of AC134521.1


