

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134322.24 + phase: 0
(456 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
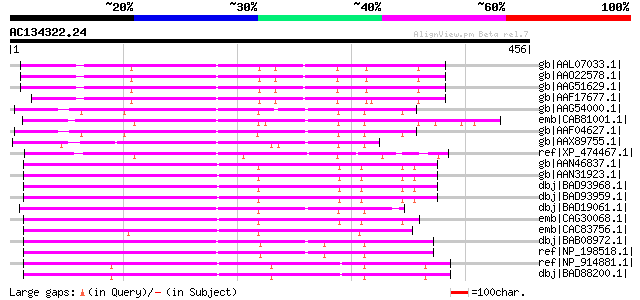
Score E
Sequences producing significant alignments: (bits) Value
gb|AAL07033.1| auxin response factor ARF17 [Arabidopsis thaliana] 263 8e-69
gb|AAO22578.1| auxin response factor ARF17 [Arabidopsis thaliana... 262 1e-68
gb|AAG51629.1| putative auxin response factor; 79762-82020 [Arab... 262 1e-68
gb|AAF17677.1| F28K19.6 [Arabidopsis thaliana] 243 9e-63
gb|AAG54000.1| auxin response factor 10 [Arabidopsis thaliana] g... 239 2e-61
emb|CAB81001.1| transcription factor-like protein [Arabidopsis t... 235 2e-60
gb|AAF04627.1| auxin response factor 10 [Arabidopsis thaliana] 226 9e-58
gb|AAX89755.1| putative auxin response factor 10 [Gossypium raim... 201 3e-50
ref|XP_474467.1| OSJNBa0039K24.27 [Oryza sativa (japonica cultiv... 186 2e-45
gb|AAN46837.1| At5g62000/mtg10_20 [Arabidopsis thaliana] gi|1419... 181 3e-44
gb|AAN31923.1| auxin response factor [Arabidopsis thaliana] gi|6... 181 3e-44
dbj|BAD93968.1| ARF1-binding protein [Arabidopsis thaliana] 181 3e-44
dbj|BAD93959.1| ARF1-binding protein [Arabidopsis thaliana] gi|6... 181 3e-44
dbj|BAD19061.1| auxin response factor 1 [Cucumis sativus] 180 7e-44
emb|CAG30068.1| putative auxin response factor [Brassica napus] 179 1e-43
emb|CAC83756.1| auxin response factor 1 [Oryza sativa (japonica ... 178 4e-43
dbj|BAB08972.1| auxin responsive transcription factor [Arabidops... 177 5e-43
ref|NP_198518.1| auxin-responsive factor (ARF8) [Arabidopsis tha... 177 5e-43
ref|NP_914881.1| auxin response factor 2 [Oryza sativa (japonica... 177 6e-43
dbj|BAD88200.1| putative auxin response factor [Oryza sativa (ja... 177 6e-43
>gb|AAL07033.1| auxin response factor ARF17 [Arabidopsis thaliana]
Length = 585
Score = 263 bits (672), Expect = 8e-69
Identities = 159/439 (36%), Positives = 232/439 (52%), Gaps = 74/439 (16%)
Query: 10 VRPEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVS 69
V P IW CA A+V+IP LHSRVYYFPQGH+E+ P S++ + S + CI++
Sbjct: 16 VDPTIWRACAGASVQIPVLHSRVYYFPQGHVEHCCPLLSTLPSSTSPVP-------CIIT 68
Query: 70 AVDLLADPHTDEVFVKLLLTPITNDVHLENPKEEVA----NLNDRNEVVSFVKTLTRSDV 125
++ LLADP TDEVF L+L P+T + +++D N+V +F K LT SD
Sbjct: 69 SIQLLADPVTDEVFAHLILQPMTQQQFTPTNYSQFGRFDGDVDDNNKVTTFAKILTPSDA 128
Query: 126 NNARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISE 185
NN F +PRFCAD+VFP L+ + + Q L+VTD+HG V F H+ RG P+R++L +
Sbjct: 129 NNGGGFSVPRFCADSVFPLLNFQIDPPVQKLYVTDIHGAVWDFRHIYRGTPRRHLL-TTG 187
Query: 186 WNSFVKRKKLVAGDSVIFMKDSTGKIFVGIRRN-----------------TQFVAAAAEQ 228
W+ FV KKL+AGDSV+FM+ S ++F+G+RR + + ++
Sbjct: 188 WSKFVNSKKLIAGDSVVFMRKSADEMFIGVRRTPISSSDGGSSYYGGDEYNGYYSQSSVA 247
Query: 229 KKDE---------------LEKAVMEALKLAEENKAFEIVYYPQGDDWCDFVVDGNVVDE 273
K+D+ +AV +A+ A + FE+V+YP W +FVV V+
Sbjct: 248 KEDDGSPKKTFRRSGNGKLTAEAVTDAINRASQGLPFEVVFYPAA-GWSEFVVRAEDVES 306
Query: 274 SMKIQWNPRMRVK--MKTDKSSRIP-YQGTISIVSRTSNLWR-----MLQVNWDEFQVSQ 325
SM + W P RVK M+T+ SSRI +QG +S + + WR LQ+ WDE ++ Q
Sbjct: 307 SMSMYWTPGTRVKMAMETEDSSRITWFQGIVSSTYQETGPWRGSPWKQLQITWDEPEILQ 366
Query: 326 IPRRVNPWWVELISH-KPAPTPFPQTKKFRTTQ--------------------SSAQLSD 364
+RVNPW VE+ +H TPFP K+ + Q SSA D
Sbjct: 367 NVKRVNPWQVEIAAHATQLHTPFPPAKRLKYPQPGGGFLSGDDGEILYPQSGLSSAAAPD 426
Query: 365 KKETLLNGDGFPVDIQRVR 383
++ + FP +Q R
Sbjct: 427 PSPSMFSYSTFPAGMQGAR 445
>gb|AAO22578.1| auxin response factor ARF17 [Arabidopsis thaliana]
gi|18411720|ref|NP_565161.1| transcriptional factor B3
family protein [Arabidopsis thaliana]
gi|46576532|sp|Q84WU6|ARFQ_ARATH Auxin response factor
17
Length = 585
Score = 262 bits (670), Expect = 1e-68
Identities = 159/439 (36%), Positives = 231/439 (52%), Gaps = 74/439 (16%)
Query: 10 VRPEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVS 69
V P IW CA A+V+IP LHSRVYYFPQGH+E+ P S++ + S + CI++
Sbjct: 16 VDPTIWRACAGASVQIPVLHSRVYYFPQGHVEHCCPLLSTLPSSTSPVP-------CIIT 68
Query: 70 AVDLLADPHTDEVFVKLLLTPITNDVHLENPKEEVA----NLNDRNEVVSFVKTLTRSDV 125
++ LLADP TDEVF L+L P+T +++D N+V +F K LT SD
Sbjct: 69 SIQLLADPVTDEVFAHLILQPMTQQQFTPTNYSRFGRFDGDVDDNNKVTTFAKILTPSDA 128
Query: 126 NNARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISE 185
NN F +PRFCAD+VFP L+ + + Q L+VTD+HG V F H+ RG P+R++L +
Sbjct: 129 NNGGGFSVPRFCADSVFPLLNFQIDPPVQKLYVTDIHGAVWDFRHIYRGTPRRHLL-TTG 187
Query: 186 WNSFVKRKKLVAGDSVIFMKDSTGKIFVGIRRN-----------------TQFVAAAAEQ 228
W+ FV KKL+AGDSV+FM+ S ++F+G+RR + + ++
Sbjct: 188 WSKFVNSKKLIAGDSVVFMRKSADEMFIGVRRTPISSSDGGSSYYGGDEYNGYYSQSSVA 247
Query: 229 KKDE---------------LEKAVMEALKLAEENKAFEIVYYPQGDDWCDFVVDGNVVDE 273
K+D+ +AV +A+ A + FE+V+YP W +FVV V+
Sbjct: 248 KEDDGSPKKTFRRSGNGKLTAEAVTDAINRASQGLPFEVVFYPAA-GWSEFVVRAEDVES 306
Query: 274 SMKIQWNPRMRVK--MKTDKSSRIP-YQGTISIVSRTSNLWR-----MLQVNWDEFQVSQ 325
SM + W P RVK M+T+ SSRI +QG +S + + WR LQ+ WDE ++ Q
Sbjct: 307 SMSMYWTPGTRVKMAMETEDSSRITWFQGIVSSTYQETGPWRGSPWKQLQITWDEPEILQ 366
Query: 326 IPRRVNPWWVELISH-KPAPTPFPQTKKFRTTQ--------------------SSAQLSD 364
+RVNPW VE+ +H TPFP K+ + Q SSA D
Sbjct: 367 NVKRVNPWQVEIAAHATQLHTPFPPAKRLKYPQPGGGFLSGDDGEILYPQSGLSSAAAPD 426
Query: 365 KKETLLNGDGFPVDIQRVR 383
++ + FP +Q R
Sbjct: 427 PSPSMFSYSTFPAGMQGAR 445
>gb|AAG51629.1| putative auxin response factor; 79762-82020 [Arabidopsis thaliana]
Length = 596
Score = 262 bits (670), Expect = 1e-68
Identities = 159/439 (36%), Positives = 231/439 (52%), Gaps = 74/439 (16%)
Query: 10 VRPEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVS 69
V P IW CA A+V+IP LHSRVYYFPQGH+E+ P S++ + S + CI++
Sbjct: 16 VDPTIWRACAGASVQIPVLHSRVYYFPQGHVEHCCPLLSTLPSSTSPVP-------CIIT 68
Query: 70 AVDLLADPHTDEVFVKLLLTPITNDVHLENPKEEVA----NLNDRNEVVSFVKTLTRSDV 125
++ LLADP TDEVF L+L P+T +++D N+V +F K LT SD
Sbjct: 69 SIQLLADPVTDEVFAHLILQPMTQQQFTPTNYSRFGRFDGDVDDNNKVTTFAKILTPSDA 128
Query: 126 NNARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISE 185
NN F +PRFCAD+VFP L+ + + Q L+VTD+HG V F H+ RG P+R++L +
Sbjct: 129 NNGGGFSVPRFCADSVFPLLNFQIDPPVQKLYVTDIHGAVWDFRHIYRGTPRRHLL-TTG 187
Query: 186 WNSFVKRKKLVAGDSVIFMKDSTGKIFVGIRRN-----------------TQFVAAAAEQ 228
W+ FV KKL+AGDSV+FM+ S ++F+G+RR + + ++
Sbjct: 188 WSKFVNSKKLIAGDSVVFMRKSADEMFIGVRRTPISSSDGGSSYYGGDEYNGYYSQSSVA 247
Query: 229 KKDE---------------LEKAVMEALKLAEENKAFEIVYYPQGDDWCDFVVDGNVVDE 273
K+D+ +AV +A+ A + FE+V+YP W +FVV V+
Sbjct: 248 KEDDGSPKKTFRRSGNGKLTAEAVTDAINRASQGLPFEVVFYPAA-GWSEFVVRAEDVES 306
Query: 274 SMKIQWNPRMRVK--MKTDKSSRIP-YQGTISIVSRTSNLWR-----MLQVNWDEFQVSQ 325
SM + W P RVK M+T+ SSRI +QG +S + + WR LQ+ WDE ++ Q
Sbjct: 307 SMSMYWTPGTRVKMAMETEDSSRITWFQGIVSSTYQETGPWRGSPWKQLQITWDEPEILQ 366
Query: 326 IPRRVNPWWVELISH-KPAPTPFPQTKKFRTTQ--------------------SSAQLSD 364
+RVNPW VE+ +H TPFP K+ + Q SSA D
Sbjct: 367 NVKRVNPWQVEIAAHATQLHTPFPPAKRLKYPQPGGGFLSGDDGEILYPQSGLSSAAAPD 426
Query: 365 KKETLLNGDGFPVDIQRVR 383
++ + FP +Q R
Sbjct: 427 PSPSMFSYSTFPAGMQGAR 445
>gb|AAF17677.1| F28K19.6 [Arabidopsis thaliana]
Length = 652
Score = 243 bits (620), Expect = 9e-63
Identities = 154/436 (35%), Positives = 226/436 (51%), Gaps = 81/436 (18%)
Query: 20 TAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVSAVDLLADPHT 79
+A+V+IP LHSRVYYFPQGH+E+ P S++ + S + CI++++ LLADP T
Sbjct: 23 SASVQIPVLHSRVYYFPQGHVEHCCPLLSTLPSSTSPVP-------CIITSIQLLADPVT 75
Query: 80 DEVFVKLLLTPITNDVHLENPKEEVA----NLNDRNEVVSFVKTLTRSDVNNARSFHIPR 135
DEVF L+L P+T +++D N+V +F K LT SD NN F +PR
Sbjct: 76 DEVFAHLILQPMTQQQFTPTNYSRFGRFDGDVDDNNKVTTFAKILTPSDANNGGGFSVPR 135
Query: 136 FCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFVKRKKL 195
FCAD+VFP L+ + + Q L+VTD+HG V F H+ RG P+R++L + W+ FV KKL
Sbjct: 136 FCADSVFPLLNFQIDPPVQKLYVTDIHGAVWDFRHIYRGTPRRHLL-TTGWSKFVNSKKL 194
Query: 196 VAGDSVIFMKDSTGKIFVGIRRN-----------------TQFVAAAAEQKKDE------ 232
+AGDSV+FM+ S ++F+G+RR + + ++ K+D+
Sbjct: 195 IAGDSVVFMRKSADEMFIGVRRTPISSSDGGSSYYGGDEYNGYYSQSSVAKEDDGSPKKT 254
Query: 233 ---------LEKAVMEALKLAEENKAFEIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRM 283
+AV +A+ A + FE+V+YP W +FVV V+ SM + W P
Sbjct: 255 FRRSGNGKLTAEAVTDAINRASQGLPFEVVFYPAA-GWSEFVVRAEDVESSMSMYWTPGT 313
Query: 284 RVK--MKTDKSSRIP-YQGTISIVSRTSNLWR-----MLQV-------NWDEFQVSQIPR 328
RVK M+T+ SSRI +QG +S + + WR LQV WDE ++ Q +
Sbjct: 314 RVKMAMETEDSSRITWFQGIVSSTYQETGPWRGSPWKQLQVYDVFEMITWDEPEILQNVK 373
Query: 329 RVNPWWVELISH-KPAPTPFPQTKKFRTTQ--------------------SSAQLSDKKE 367
RVNPW VE+ +H TPFP K+ + Q SSA D
Sbjct: 374 RVNPWQVEIAAHATQLHTPFPPAKRLKYPQPGGGFLSGDDGEILYPQSGLSSAAAPDPSP 433
Query: 368 TLLNGDGFPVDIQRVR 383
++ + FP +Q R
Sbjct: 434 SMFSYSTFPAGMQGAR 449
>gb|AAG54000.1| auxin response factor 10 [Arabidopsis thaliana]
gi|4432846|gb|AAD20695.1| unknown protein [Arabidopsis
thaliana] gi|46576666|sp|Q9SKN5|ARFJ_ARATH Auxin
response factor 10 gi|13272405|gb|AAK17141.1| unknown
protein [Arabidopsis thaliana]
gi|15226389|ref|NP_180402.1| auxin-responsive factor
(ARF10) [Arabidopsis thaliana]
Length = 693
Score = 239 bits (609), Expect = 2e-61
Identities = 156/416 (37%), Positives = 217/416 (51%), Gaps = 76/416 (18%)
Query: 5 KQPSHVRPEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFR--P 62
+Q + P++WH CA + V+IP L+S V+YF QGH E+A H + R P
Sbjct: 2 EQEKSLDPQLWHACAGSMVQIPSLNSTVFYFAQGHTEHA--------HAPPDFHAPRVPP 53
Query: 63 FTLCIVSAVDLLADPHTDEVFVKLLLTPIT-NDVHLEN-------PKEEVANLNDRNEVV 114
LC V +V LAD TDEVF K+ L P+ ND+ LEN P N N + +
Sbjct: 54 LILCRVVSVKFLADAETDEVFAKITLLPLPGNDLDLENDAVLGLTPPSSDGNGNGKEKPA 113
Query: 115 SFVKTLTRSDVNNARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRG 174
SF KTLT+SD NN F +PR+CA+ +FP+LD AE Q + D+HGE KF H+ RG
Sbjct: 114 SFAKTLTQSDANNGGGFSVPRYCAETIFPRLDYSAEPPVQTVIAKDIHGETWKFRHIYRG 173
Query: 175 FPKRNMLYISEWNSFVKRKKLVAGDSVIFMKDSTGKIFVGIRR----------------- 217
P+R++L + W++FV +KKL+AGDS++F++ +G + VGIRR
Sbjct: 174 TPRRHLL-TTGWSTFVNQKKLIAGDSIVFLRSESGDLCVGIRRAKRGGLGSNAGSDNPYP 232
Query: 218 -------------------------NTQFVAAAAEQKKDELEKAVMEALKLAEENKAFEI 252
N AAA + + E AV EA+ A +AFE+
Sbjct: 233 GFSGFLRDDESTTTTSKLMMMKRNGNNDGNAAATGRVRVE---AVAEAVARAACGQAFEV 289
Query: 253 VYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKM--KTDKSSRIP-YQGTISIVSRTSN 309
VYYP+ +F V V +M+I+W MR KM +T+ SSRI + GT+S V
Sbjct: 290 VYYPRAST-PEFCVKAADVRSAMRIRWCSGMRFKMAFETEDSSRISWFMGTVSAVQVADP 348
Query: 310 L------WRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA--PTPFPQTKKFRTTQ 357
+ WR+LQV WDE + Q +RV+PW VEL+S+ P +PF KK R Q
Sbjct: 349 IRWPNSPWRLLQVAWDEPDLLQNVKRVSPWLVELVSNMPTIHLSPFSPRKKIRIPQ 404
>emb|CAB81001.1| transcription factor-like protein [Arabidopsis thaliana]
gi|4938484|emb|CAB43843.1| transcription factor-like
protein [Arabidopsis thaliana] gi|7484815|pir||T08984
auxin response factor 7 homolog F6G3.110 - Arabidopsis
thaliana
Length = 653
Score = 235 bits (600), Expect = 2e-60
Identities = 173/519 (33%), Positives = 245/519 (46%), Gaps = 107/519 (20%)
Query: 12 PEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVSAV 71
P++WH CA V++P ++S+V+YFPQGH ENA P LC V A+
Sbjct: 18 PQLWHACAGGMVRMPPMNSKVFYFPQGHAENAYDCVDFGN------LPIPPMVLCRVLAI 71
Query: 72 DLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLN----DRNEVVSFVKTLTRSDVNN 127
+AD +DEVF KL L P+ +D ++++ + + N + + SF KTLT+SD NN
Sbjct: 72 KYMADAESDEVFAKLRLIPLKDDEYVDHEYGDGEDSNGFESNSEKTPSFAKTLTQSDANN 131
Query: 128 ARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWN 187
F +PR+CA+ +FP+LD AE Q + DVHG+V KF H+ RG P+R++L + W+
Sbjct: 132 GGGFSVPRYCAETIFPRLDYNAEPPVQTILAKDVHGDVWKFRHIYRGTPRRHLL-TTGWS 190
Query: 188 SFVKRKKLVAGDSVIFMKDSTGKIFVGIRR--------NTQFVA---------------- 223
+FV +KKLVAGDS++FM+ G + VGIRR ++ A
Sbjct: 191 NFVNQKKLVAGDSIVFMRAENGDLCVGIRRAKRGGIGNGPEYSAGWNPIGGSCGYSSLLR 250
Query: 224 ------------AAAEQKKDELEKAVMEALKLAEENKAFEIVYYPQGDDWCDFVVDGNVV 271
+ A++K ++V+EA LA + FE+VYYP+ +F V
Sbjct: 251 EDESNSLRRSNCSLADRKGKVTAESVIEAATLAISGRPFEVVYYPRAST-SEFCVKALDA 309
Query: 272 DESMKIQWNPRMRVKM--KTDKSSRIP-YQGTISIVSRTSNL------WRMLQVNWDEFQ 322
+M+I W MR KM +T+ SSRI + GT+S V+ + + WR+LQV WDE
Sbjct: 310 RAAMRIPWCSGMRFKMAFETEDSSRISWFMGTVSAVNVSDPIRWPNSPWRLLQVAWDEPD 369
Query: 323 VSQIPRRVNPWWVELISH-KPAP-TPF-PQTKKFRTTQ---------SSAQLSDKKETLL 370
+ Q +RVNPW VEL+S+ P P T F P KK R Q S S L+
Sbjct: 370 LLQNVKRVNPWLVELVSNVHPIPLTSFSPPRKKMRLPQHPDYNNLINSIPVPSFPSNPLI 429
Query: 371 NG-------DGFPVDIQRVRHDLVSISGPIHS---HIILNSSETKFP------------- 407
D PV +Q RH+ G S H LN P
Sbjct: 430 RSSPLSSVLDNVPVGLQGARHNAHQYYGLSSSDLHHYYLNRPPPPPPPSSLQLSPSLGLR 489
Query: 408 ---------------ATHNCNTKNDSDGSIMLFGKIIQP 431
T CN I+LFGK+I P
Sbjct: 490 NIDTKNEKGFCFLTMGTTPCNDTKSKKSHIVLFGKLILP 528
>gb|AAF04627.1| auxin response factor 10 [Arabidopsis thaliana]
Length = 701
Score = 226 bits (577), Expect = 9e-58
Identities = 149/414 (35%), Positives = 208/414 (49%), Gaps = 71/414 (17%)
Query: 5 KQPSHVRPEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFR--P 62
+Q + P++WH CA + V+IP L+S V+YF QGH E+A H + R P
Sbjct: 2 EQEKSLDPQLWHACAGSMVQIPSLNSTVFYFAQGHTEHA--------HAPPDFHAPRVPP 53
Query: 63 FTLCIVSAVDLLADPHTDEVFVKLLLTPIT-NDVHLEN-------PKEEVANLNDRNEVV 114
LC V +V LAD TDEVF K+ L P+ ND+ LEN P N N + +
Sbjct: 54 LILCRVVSVKFLADAETDEVFAKITLLPLPGNDLDLENDAVLGLTPPSSDGNGNGKEKPA 113
Query: 115 SFVKTLTRSDVNNARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRG 174
SF KTLT+SD NN F +PR+CA+ +FP+LD AE Q + D+HGE KF H+ RG
Sbjct: 114 SFAKTLTQSDANNGGGFSVPRYCAETIFPRLDYSAEPPVQTVNAKDIHGETWKFRHIYRG 173
Query: 175 FPKRNMLYISEWNSFVKRKKLVAGDSVIFMKDSTGKIFVGIR------------------ 216
P+R++L + W++FV +KKL+AGDS++F++ +G + VGIR
Sbjct: 174 TPRRHLL-TTGWSTFVNQKKLIAGDSIVFLRSESGDLCVGIRRAKRGGLGSNAGSDNPYP 232
Query: 217 ----------------------RNTQFVAAAAEQKKDELEKAVMEALKLAEENKAFEIVY 254
RN AA + +E + +KAFE+VY
Sbjct: 233 GFSGFLRDDESTTTTSKLMMMKRNGNNDGNAAATGRVRVEAVAGSGGACSXVDKAFEVVY 292
Query: 255 YPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKM--KTDKSSRIP-YQGTISIVSRTSNL- 310
YP+ +F V V +M+ W MR KM +T+ SSRI + GT S V +
Sbjct: 293 YPRAST-PEFCVKAADVRSAMRXXWCXGMRXKMAFETEDSSRISWFMGTXSAVQVADPIR 351
Query: 311 -----WRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA--PTPFPQTKKFRTTQ 357
WR+LQV WDE + Q +RV+PW V L+S+ P +PF KK R Q
Sbjct: 352 WPNSPWRLLQVAWDEPDLXQNVKRVSPWLVXLVSNMPTIHLSPFSXWKKIRIPQ 405
>gb|AAX89755.1| putative auxin response factor 10 [Gossypium raimondii]
Length = 417
Score = 201 bits (512), Expect = 3e-50
Identities = 126/363 (34%), Positives = 191/363 (51%), Gaps = 53/363 (14%)
Query: 3 LQKQPSHVRPEIWHTCATAAVKIPKLHSRVYYFPQGHLENAS-----PSSSSITHTHSFL 57
+++ + P++WH CA V+IP L+S+V+YFPQGH E++ PSS +
Sbjct: 1 MKESDKSLDPQLWHACAGPMVQIPPLNSKVFYFPQGHAEHSLAAVDFPSSPPVP------ 54
Query: 58 QSFRPFTLCIVSAVDLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFV 117
LC V+++ +AD TDEV+ K+LL P+ N L+ V ++ + SF
Sbjct: 55 ----ALVLCRVASLKFMADTETDEVYAKILLMPLPN-TELDLEHVAVFGSDNAEKPASFA 109
Query: 118 KTLTRSDVNNARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPK 177
KTLT+SD NN F +PR+CA+ +FP LD + Q + DVHGE KF H+ RG P+
Sbjct: 110 KTLTQSDANNGGGFSVPRYCAETIFPPLDYTEDPPVQTVVAVDVHGETWKFRHIYRGTPR 169
Query: 178 RNMLYISEWNSFVKRKKLVAGDSVIFMKDSTGKIFVGIRRNTQFVAAAAEQ--------K 229
R++L + W++FV KKLVAGDS++F++ G + VGIRR + E +
Sbjct: 170 RHLL-TTGWSTFVNHKKLVAGDSIVFLRSENGGLCVGIRRAKRGTGNGPEAGSPFLSFLR 228
Query: 230 KDELE------------------KAVMEALKLAEENKAFEIVYYPQGDDWCDFVVDGNVV 271
+DE + +AV++A LA + FE+VYYP+ +F V + V
Sbjct: 229 EDESKMMMMNRNGDWRGKGKLKAEAVLQAATLAASGQPFEVVYYPRAST-PEFCVKASSV 287
Query: 272 DESMKIQWNPRMRVKM--KTDKSSRIP-YQGTISIVSRTSNL------WRMLQVNWDEFQ 322
+M++ W MR KM +T+ SSRI + GT+S V + WR+ Q+ Q
Sbjct: 288 KAAMRVPWCCGMRFKMAFETEDSSRISWFMGTVSSVQVVDPIRWPNSPWRLFQLEELSTQ 347
Query: 323 VSQ 325
V Q
Sbjct: 348 VPQ 350
>ref|XP_474467.1| OSJNBa0039K24.27 [Oryza sativa (japonica cultivar-group)]
gi|38345524|emb|CAE01808.2| OSJNBa0039K24.27 [Oryza
sativa (japonica cultivar-group)]
Length = 529
Score = 186 bits (471), Expect = 2e-45
Identities = 129/401 (32%), Positives = 197/401 (48%), Gaps = 50/401 (12%)
Query: 14 IWHTCATAAVKIPKLHSRVYYFPQGHLENA-SPSSSSITHTHSFLQSFRPFTLCIVSAVD 72
+W CA +IP + ++V YFP+GH E +P + H F LC ++AVD
Sbjct: 28 VWLACAAPLSRIPVVGTQVSYFPEGHAEQCPAPLPDPLPSAHRFF-------LCTITAVD 80
Query: 73 LLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLN----DRNEVVSFVKTLTRSDVNNA 128
L AD T E + + L P+ +D P A + E + K LT+SD NN
Sbjct: 81 LSADTTTGEPYATISLLPLRHDAPAPAPAPAPAAAELAEAESQEFRYYAKQLTQSDANNG 140
Query: 129 RSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNS 188
F +PR CAD++FP L+L+ + Q L + D+ G+ +F H+ RG P+R++L + W+
Sbjct: 141 GGFSVPRLCADHIFPALNLDDDPPVQSLTMGDLQGDSWEFRHIYRGTPRRHLL-TTGWSK 199
Query: 189 FVKRKKLVAGDSVIFM----KDSTGKIFVGIRRNTQFVAAAAEQKKDELE-KAVMEALKL 243
FV K+LVAGD+V+FM K+ VG+RR ++ +A + ++ + VMEA++L
Sbjct: 200 FVNAKQLVAGDTVVFMWCGAPAPERKLLVGVRRAARYSGESACNARGRVQPQEVMEAVRL 259
Query: 244 AEENKAFEIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVK---MKTDKSSRIPY-QG 299
A E AF + YYP+ +FVV VD+ + W M+V+ M+ + + R+ + G
Sbjct: 260 AAEQAAFRVTYYPRHGAG-EFVVPRVEVDKGLTTPWRCGMQVRAQVMEAEDTRRLAWLNG 318
Query: 300 TISIVSRTSNLWRMLQVNWDEFQVSQI--PRRVNPWWVELISHKPAPTPFPQTKKFRTTQ 357
T++ + R +WR L+V WD S R VNPW V+ + P P K
Sbjct: 319 TLTNL-RHQQIWRTLEVEWDASAASSSMKNRFVNPWQVQPVDF----PPLPMGLKISNNN 373
Query: 358 SSAQLSDKKETLLNGDGF-------------PVDIQRVRHD 385
SA + NGD P DIQ RH+
Sbjct: 374 ISA-------PVCNGDSLLVPPILMHPQPQPPADIQGARHN 407
>gb|AAN46837.1| At5g62000/mtg10_20 [Arabidopsis thaliana]
gi|14190369|gb|AAK55665.1| AT5g62000/mtg10_20
[Arabidopsis thaliana]
Length = 678
Score = 181 bits (460), Expect = 3e-44
Identities = 124/381 (32%), Positives = 182/381 (47%), Gaps = 18/381 (4%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVSAVD 72
E+WH CA V +P+ RV+YFPQGH+E S++ L LC V VD
Sbjct: 61 ELWHACAGPLVTVPRQDDRVFYFPQGHIEQVEASTNQAAEQQMPLYDLPSKLLCRVINVD 120
Query: 73 LLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNARSFH 132
L A+ TDEV+ ++ L P N KE R +V SF KTLT SD + F
Sbjct: 121 LKAEADTDEVYAQITLLPEANQDENAIEKEAPLPPPPRFQVHSFCKTLTASDTSTHGGFS 180
Query: 133 IPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFVKR 192
+ R AD P LD+ + +Q L D+H +F H+ RG P+R++L S W+ FV
Sbjct: 181 VLRRHADECLPPLDMSRQPPTQELVAKDLHANEWRFRHIFRGQPRRHLLQ-SGWSVFVSS 239
Query: 193 KKLVAGDSVIFMKDSTGKIFVGIRR--NTQFVAAAAEQKKDELEKAVMEALKLAEENKAF 250
K+LVAGD+ IF++ G++ VG+RR Q ++ + V+ A
Sbjct: 240 KRLVAGDAFIFLRGENGELRVGVRRAMRQQGNVPSSVISSHSMHLGVLATAWHAISTGTM 299
Query: 251 EIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKMK--TDKSSRIPYQGTISIVSRT- 307
VYY +F+V + ES+K ++ MR KM+ +++ + GTI + +
Sbjct: 300 FTVYYKPRTSPSEFIVPFDQYMESVKNNYSIGMRFKMRFEGEEAPEQRFTGTIVGIEESD 359
Query: 308 -----SNLWRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA--PTPFPQTKKFR-----T 355
+ WR L+V WDE P RV+PW VE PA P P P+ K+ R +
Sbjct: 360 PTRWPKSKWRSLKVRWDETSSIPRPDRVSPWKVEPALAPPALSPVPMPRPKRPRSNIAPS 419
Query: 356 TQSSAQLSDKKETLLNGDGFP 376
+ S+ L+ + T N D P
Sbjct: 420 SPDSSMLTREGTTKANMDPLP 440
>gb|AAN31923.1| auxin response factor [Arabidopsis thaliana]
gi|62319959|dbj|BAD94058.1| ARF1-binding protein
[Arabidopsis thaliana] gi|62319913|dbj|BAD93985.1|
ARF1-binding protein [Arabidopsis thaliana]
gi|10176918|dbj|BAB10162.1| auxin response factor-like
protein [Arabidopsis thaliana]
gi|30697612|ref|NP_201006.2| transcriptional factor B3
family protein / auxin-responsive factor, putative
(ARF1) [Arabidopsis thaliana]
gi|42573768|ref|NP_974980.1| transcriptional factor B3
family protein / auxin-responsive factor, putative
(ARF1) [Arabidopsis thaliana]
gi|30697610|ref|NP_851244.1| transcriptional factor B3
family protein / auxin-responsive factor, putative
(ARF1) [Arabidopsis thaliana] gi|49616349|gb|AAT67071.1|
ARF2 [Arabidopsis thaliana]
gi|46395940|sp|Q94JM3|ARFB_ARATH Auxin response factor 2
(ARF1-binding protein) (ARF1-BP)
Length = 859
Score = 181 bits (460), Expect = 3e-44
Identities = 124/381 (32%), Positives = 182/381 (47%), Gaps = 18/381 (4%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVSAVD 72
E+WH CA V +P+ RV+YFPQGH+E S++ L LC V VD
Sbjct: 61 ELWHACAGPLVTVPRQDDRVFYFPQGHIEQVEASTNQAAEQQMPLYDLPSKLLCRVINVD 120
Query: 73 LLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNARSFH 132
L A+ TDEV+ ++ L P N KE R +V SF KTLT SD + F
Sbjct: 121 LKAEADTDEVYAQITLLPEANQDENAIEKEAPLPPPPRFQVHSFCKTLTASDTSTHGGFS 180
Query: 133 IPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFVKR 192
+ R AD P LD+ + +Q L D+H +F H+ RG P+R++L S W+ FV
Sbjct: 181 VLRRHADECLPPLDMSRQPPTQELVAKDLHANEWRFRHIFRGQPRRHLLQ-SGWSVFVSS 239
Query: 193 KKLVAGDSVIFMKDSTGKIFVGIRR--NTQFVAAAAEQKKDELEKAVMEALKLAEENKAF 250
K+LVAGD+ IF++ G++ VG+RR Q ++ + V+ A
Sbjct: 240 KRLVAGDAFIFLRGENGELRVGVRRAMRQQGNVPSSVISSHSMHLGVLATAWHAISTGTM 299
Query: 251 EIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKMK--TDKSSRIPYQGTISIVSRT- 307
VYY +F+V + ES+K ++ MR KM+ +++ + GTI + +
Sbjct: 300 FTVYYKPRTSPSEFIVPFDQYMESVKNNYSIGMRFKMRFEGEEAPEQRFTGTIVGIEESD 359
Query: 308 -----SNLWRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA--PTPFPQTKKFR-----T 355
+ WR L+V WDE P RV+PW VE PA P P P+ K+ R +
Sbjct: 360 PTRWPKSKWRSLKVRWDETSSIPRPDRVSPWKVEPALAPPALSPVPMPRPKRPRSNIAPS 419
Query: 356 TQSSAQLSDKKETLLNGDGFP 376
+ S+ L+ + T N D P
Sbjct: 420 SPDSSMLTREGTTKANMDPLP 440
>dbj|BAD93968.1| ARF1-binding protein [Arabidopsis thaliana]
Length = 859
Score = 181 bits (460), Expect = 3e-44
Identities = 124/381 (32%), Positives = 182/381 (47%), Gaps = 18/381 (4%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVSAVD 72
E+WH CA V +P+ RV+YFPQGH+E S++ L LC V VD
Sbjct: 61 ELWHACAGPLVTVPRQDDRVFYFPQGHIEQVEASTNQAAEQQMPLYDLPSKLLCRVINVD 120
Query: 73 LLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNARSFH 132
L A+ TDEV+ ++ L P N KE R +V SF KTLT SD + F
Sbjct: 121 LKAEADTDEVYAQITLLPEANQDENAIEKEAPLPPPPRFQVHSFCKTLTASDTSTHGGFS 180
Query: 133 IPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFVKR 192
+ R AD P LD+ + +Q L D+H +F H+ RG P+R++L S W+ FV
Sbjct: 181 VLRRHADECLPPLDMSRQPPTQELVAKDLHANEWRFRHIFRGQPRRHLLQ-SGWSVFVSS 239
Query: 193 KKLVAGDSVIFMKDSTGKIFVGIRR--NTQFVAAAAEQKKDELEKAVMEALKLAEENKAF 250
K+LVAGD+ IF++ G++ VG+RR Q ++ + V+ A
Sbjct: 240 KRLVAGDAFIFLRGENGELRVGVRRAMRQQGNVPSSVISSHSMHLGVLATAWHAISTGTM 299
Query: 251 EIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKMK--TDKSSRIPYQGTISIVSRT- 307
VYY +F+V + ES+K ++ MR KM+ +++ + GTI + +
Sbjct: 300 FTVYYKPRTSPSEFIVPFDQYMESVKNNYSIGMRFKMRFEGEEAPEQRFTGTIVGIEESD 359
Query: 308 -----SNLWRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA--PTPFPQTKKFR-----T 355
+ WR L+V WDE P RV+PW VE PA P P P+ K+ R +
Sbjct: 360 PTRWPKSKWRSLKVRWDETSSIPRPDRVSPWKVEPALAPPALSPVPMPRPKRPRSNIAPS 419
Query: 356 TQSSAQLSDKKETLLNGDGFP 376
+ S+ L+ + T N D P
Sbjct: 420 SPDSSMLTREGTTKANMDPLP 440
>dbj|BAD93959.1| ARF1-binding protein [Arabidopsis thaliana]
gi|62319857|dbj|BAD93897.1| ARF1-binding protein
[Arabidopsis thaliana] gi|62319853|dbj|BAD93891.1|
ARF1-binding protein [Arabidopsis thaliana]
Length = 859
Score = 181 bits (460), Expect = 3e-44
Identities = 124/381 (32%), Positives = 182/381 (47%), Gaps = 18/381 (4%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVSAVD 72
E+WH CA V +P+ RV+YFPQGH+E S++ L LC V VD
Sbjct: 61 ELWHACAGPLVTVPRQDDRVFYFPQGHIEQVEASTNQAAEQQMPLYDLPSKLLCRVINVD 120
Query: 73 LLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNARSFH 132
L A+ TDEV+ ++ L P N KE R +V SF KTLT SD + F
Sbjct: 121 LKAEADTDEVYAQITLLPEANQDENAIEKEAPLPPPPRFQVHSFCKTLTASDTSTHGGFS 180
Query: 133 IPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFVKR 192
+ R AD P LD+ + +Q L D+H +F H+ RG P+R++L S W+ FV
Sbjct: 181 VLRRHADECLPPLDMSRQPPTQELVAKDLHANEWRFRHIFRGQPRRHLLQ-SGWSVFVSS 239
Query: 193 KKLVAGDSVIFMKDSTGKIFVGIRR--NTQFVAAAAEQKKDELEKAVMEALKLAEENKAF 250
K+LVAGD+ IF++ G++ VG+RR Q ++ + V+ A
Sbjct: 240 KRLVAGDAFIFLRGENGELRVGVRRAMRQQGNVPSSVISSHSMHLGVLATAWHAISTGTM 299
Query: 251 EIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKMK--TDKSSRIPYQGTISIVSRT- 307
VYY +F+V + ES+K ++ MR KM+ +++ + GTI + +
Sbjct: 300 FTVYYKPRTSPSEFIVPFDQYMESVKNNYSIGMRFKMRFEGEEAPEQRFTGTIVGIEESD 359
Query: 308 -----SNLWRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA--PTPFPQTKKFR-----T 355
+ WR L+V WDE P RV+PW VE PA P P P+ K+ R +
Sbjct: 360 PTRWPKSKWRSLKVRWDETSSIPRPDRVSPWKVEPALAPPALSPVPMPRPKRPRSNIAPS 419
Query: 356 TQSSAQLSDKKETLLNGDGFP 376
+ S+ L+ + T N D P
Sbjct: 420 SPDSSMLTREGTTKANMDPLP 440
>dbj|BAD19061.1| auxin response factor 1 [Cucumis sativus]
Length = 1081
Score = 180 bits (457), Expect = 7e-44
Identities = 115/350 (32%), Positives = 174/350 (48%), Gaps = 18/350 (5%)
Query: 9 HVRPEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIV 68
++ E+WH CA V +P + S V YFPQGH E + S + T + +C++
Sbjct: 19 NINSELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMNKETDFIPNYPNLPSKLICML 78
Query: 69 SAVDLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNA 128
V L ADP TDEV+ ++ L P+ ++ R F KTLT SD +
Sbjct: 79 HNVTLHADPETDEVYAQMTLQPVNKYEKEALLASDIGLKQSRQPAEFFCKTLTASDTSTH 138
Query: 129 RSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNS 188
F +PR A+ +FP LD + +Q L D+H F H+ RG PKR++L + W+
Sbjct: 139 GGFSVPRRAAEKIFPPLDYSMQPPAQELVARDLHDNSWTFRHIYRGQPKRHLL-TTGWSV 197
Query: 189 FVKRKKLVAGDSVIFMKDSTGKIFVGIRR--NTQFVAAAAEQKKDELEKAVM-EALKLAE 245
FV K+L AGDSV+F++D ++ +GIRR Q +++ D + ++ A A
Sbjct: 198 FVSTKRLFAGDSVLFIRDEKSQLLLGIRRANRQQPALSSSVISSDSMHIGILASAAHAAA 257
Query: 246 ENKAFEIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKM--KTDKSSRIPYQGTISI 303
N F I Y P+ +FV+ +++M Q + MR +M +T++S Y GTI+
Sbjct: 258 NNSPFTIFYNPRASP-SEFVIPLAKYNKAMYTQVSLGMRFRMMFETEESGVRRYMGTITG 316
Query: 304 VSRTSNL------WRMLQVNWDEFQVSQIPRRVNPWWVELISHKPAPTPF 347
+S ++ WR LQV WDE + P RV+ W VE P TPF
Sbjct: 317 ISDMDSVRWKNSQWRNLQVGWDESAAGERPNRVSIWEVE-----PVVTPF 361
>emb|CAG30068.1| putative auxin response factor [Brassica napus]
Length = 848
Score = 179 bits (455), Expect = 1e-43
Identities = 118/360 (32%), Positives = 175/360 (47%), Gaps = 13/360 (3%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVSAVD 72
E+WH CA V +P+ RV+YFPQGH+E S++ L LC V VD
Sbjct: 56 ELWHACAGPLVTVPRQDDRVFYFPQGHIEQVEASTNQAAEQQMPLYDLPSKILCRVINVD 115
Query: 73 LLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNARSFH 132
L A+ TDEV+ ++ L P KE R +V SF KTLT SD + F
Sbjct: 116 LKAEADTDEVYAQITLLPEPVQDENSIEKEAPPPPPPRFQVHSFCKTLTASDTSTHGGFS 175
Query: 133 IPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFVKR 192
+ R AD P LD+ + +Q L D+H +F H+ RG P+R++L S W+ FV
Sbjct: 176 VLRRHADECLPPLDMSRQPPTQELVAKDLHASEWRFRHIFRGQPRRHLLQ-SGWSVFVSS 234
Query: 193 KKLVAGDSVIFMKDSTGKIFVGIRR--NTQFVAAAAEQKKDELEKAVMEALKLAEENKAF 250
K+LVAGD+ IF++ G++ VG+RR Q ++ + V+ A
Sbjct: 235 KRLVAGDAFIFLRGENGELRVGVRRAMRQQGNVPSSVISSHSMHLGVLATAWHAISTGTM 294
Query: 251 EIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKMK--TDKSSRIPYQGTISIVSRT- 307
VYY +F+V + ES+KI ++ MR KM+ +++ + GTI + +
Sbjct: 295 FTVYYKPRTSPSEFIVPFDQYTESVKINYSIGMRFKMRFEGEEAPEQRFTGTIVGIEDSD 354
Query: 308 -----SNLWRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA--PTPFPQTKKFRTTQSSA 360
+ WR L+V WDE P RV+PW +E PA P P P+ K+ R+ +S+
Sbjct: 355 PTRWAKSKWRSLKVRWDETTSIPRPDRVSPWKIEPALSPPALSPVPMPRPKRPRSNLASS 414
>emb|CAC83756.1| auxin response factor 1 [Oryza sativa (japonica cultivar-group)]
Length = 836
Score = 178 bits (451), Expect = 4e-43
Identities = 119/359 (33%), Positives = 175/359 (48%), Gaps = 18/359 (5%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVSAVD 72
E+W CA V +P++ +V+YFPQGH+E S++ + L + LC V V+
Sbjct: 24 ELWSACAGPLVTVPRVGEKVFYFPQGHIEQVEASTNQVGEQRMQLYNLPWKILCEVMNVE 83
Query: 73 LLADPHTDEVFVKLLLTPITNDVHLENPKEE-----VANLNDRNEVVSFVKTLTRSDVNN 127
L A+P TDEV+ +L L P + EE A + R V SF KTLT SD +
Sbjct: 84 LKAEPDTDEVYAQLTLLPESKQQEDNGSTEEEVPSAPAAGHVRPRVHSFCKTLTASDTST 143
Query: 128 ARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWN 187
F + R AD P LD+ + +Q L D+HG +F H+ RG P+R++L S W+
Sbjct: 144 HGGFSVLRRHADECLPPLDMSRQPPTQELVAKDLHGVEWRFRHIFRGQPRRHLLQ-SGWS 202
Query: 188 SFVKRKKLVAGDSVIFMKDSTGKIFVGIRR--NTQFVAAAAEQKKDELEKAVMEALKLAE 245
FV K+LVAGD+ IF++ G++ VG+RR Q ++ + V+ A
Sbjct: 203 VFVSAKRLVAGDAFIFLRGENGELRVGVRRAMRQQTNVPSSVISSHSMHLGVLATAWHAV 262
Query: 246 ENKAFEIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKMKTDKSSRIPYQGTISIVS 305
VYY +FVV + ES+K ++ MR KM+ + + T +IV
Sbjct: 263 NTGTMFTVYYKPRTSPAEFVVPYDRYMESLKQNYSIGMRFKMRFEGEEAPEQRFTGTIVG 322
Query: 306 R--------TSNLWRMLQVNWDEFQVSQIPRRVNPWWVE-LISHKPA-PTPFPQTKKFR 354
+ WR L+V WDE P RV+PW +E +S P P P P+TK+ R
Sbjct: 323 MGDSDPAGWPESKWRSLKVRWDEASSIPRPERVSPWQIEPAVSPPPVNPLPVPRTKRLR 381
>dbj|BAB08972.1| auxin responsive transcription factor [Arabidopsis thaliana]
Length = 821
Score = 177 bits (450), Expect = 5e-43
Identities = 118/374 (31%), Positives = 189/374 (49%), Gaps = 16/374 (4%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTH-SFLQSFRPFTLCIVSAV 71
E+WH CA V +P SRV YFPQGH E + +++ H S P +C + V
Sbjct: 22 ELWHACAGPLVSLPSSGSRVVYFPQGHSEQVAATTNKEVDGHIPNYPSLPPQLICQLHNV 81
Query: 72 DLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNARSF 131
+ AD TDEV+ ++ L P+T + E + + F KTLT SD + F
Sbjct: 82 TMHADVETDEVYAQMTLQPLTPEEQKETFVPIELGIPSKQPSNYFCKTLTASDTSTHGGF 141
Query: 132 HIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFVK 191
+PR A+ VFP LD + +Q L D+H KF H+ RG PKR++L + W+ FV
Sbjct: 142 SVPRRAAEKVFPPLDYTLQPPAQELIARDLHDVEWKFRHIFRGQPKRHLL-TTGWSVFVS 200
Query: 192 RKKLVAGDSVIFMKDSTGKIFVGIRRNT--QFVAAAAEQKKDELEKAVMEALKLAE-ENK 248
K+LVAGDSVIF+++ ++F+GIR T Q + ++ D + ++ A A N
Sbjct: 201 AKRLVAGDSVIFIRNEKNQLFLGIRHATRPQTIVPSSVLSSDSMHIGLLAAAAHASATNS 260
Query: 249 AFEIVYYPQGDDWCDFVVDGNVVDESM---KIQWNPRMRVKMKTDKSSRIPYQGTISIVS 305
F + ++P+ +FV+ + +++ +I R R+ +T++SS Y GTI+ +S
Sbjct: 261 CFTVFFHPRASQ-SEFVIQLSKYIKAVFHTRISVGMRFRMLFETEESSVRRYMGTITGIS 319
Query: 306 RTSNL------WRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA-PTPFPQTKKFRTTQS 358
++ WR ++V WDE + RV+ W +E ++ P P+ FP K
Sbjct: 320 DLDSVRWPNSHWRSVKVGWDESTAGERQPRVSLWEIEPLTTFPMYPSLFPLRLKRPWHAG 379
Query: 359 SAQLSDKKETLLNG 372
++ L D + L +G
Sbjct: 380 TSSLPDGRGDLGSG 393
>ref|NP_198518.1| auxin-responsive factor (ARF8) [Arabidopsis thaliana]
gi|49616355|gb|AAT67074.1| ARF8 [Arabidopsis thaliana]
gi|46576647|sp|Q9FGV1|ARFH_ARATH Auxin response factor 8
gi|4104931|gb|AAD02219.1| auxin response factor 8
[Arabidopsis thaliana]
Length = 811
Score = 177 bits (450), Expect = 5e-43
Identities = 118/374 (31%), Positives = 189/374 (49%), Gaps = 16/374 (4%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTH-SFLQSFRPFTLCIVSAV 71
E+WH CA V +P SRV YFPQGH E + +++ H S P +C + V
Sbjct: 22 ELWHACAGPLVSLPSSGSRVVYFPQGHSEQVAATTNKEVDGHIPNYPSLPPQLICQLHNV 81
Query: 72 DLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNARSF 131
+ AD TDEV+ ++ L P+T + E + + F KTLT SD + F
Sbjct: 82 TMHADVETDEVYAQMTLQPLTPEEQKETFVPIELGIPSKQPSNYFCKTLTASDTSTHGGF 141
Query: 132 HIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFVK 191
+PR A+ VFP LD + +Q L D+H KF H+ RG PKR++L + W+ FV
Sbjct: 142 SVPRRAAEKVFPPLDYTLQPPAQELIARDLHDVEWKFRHIFRGQPKRHLL-TTGWSVFVS 200
Query: 192 RKKLVAGDSVIFMKDSTGKIFVGIRRNT--QFVAAAAEQKKDELEKAVMEALKLAE-ENK 248
K+LVAGDSVIF+++ ++F+GIR T Q + ++ D + ++ A A N
Sbjct: 201 AKRLVAGDSVIFIRNEKNQLFLGIRHATRPQTIVPSSVLSSDSMHIGLLAAAAHASATNS 260
Query: 249 AFEIVYYPQGDDWCDFVVDGNVVDESM---KIQWNPRMRVKMKTDKSSRIPYQGTISIVS 305
F + ++P+ +FV+ + +++ +I R R+ +T++SS Y GTI+ +S
Sbjct: 261 CFTVFFHPRASQ-SEFVIQLSKYIKAVFHTRISVGMRFRMLFETEESSVRRYMGTITGIS 319
Query: 306 RTSNL------WRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA-PTPFPQTKKFRTTQS 358
++ WR ++V WDE + RV+ W +E ++ P P+ FP K
Sbjct: 320 DLDSVRWPNSHWRSVKVGWDESTAGERQPRVSLWEIEPLTTFPMYPSLFPLRLKRPWHAG 379
Query: 359 SAQLSDKKETLLNG 372
++ L D + L +G
Sbjct: 380 TSSLPDGRGDLGSG 393
>ref|NP_914881.1| auxin response factor 2 [Oryza sativa (japonica cultivar-group)]
Length = 826
Score = 177 bits (449), Expect = 6e-43
Identities = 120/391 (30%), Positives = 182/391 (45%), Gaps = 18/391 (4%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVSAVD 72
E+WH CA V +P++ V+YFPQGH+E S + + + L LC V V+
Sbjct: 19 ELWHACAGPLVTVPRVGDLVFYFPQGHIEQVEASMNQVADSQMRLYDLPSKLLCRVLNVE 78
Query: 73 LLADPHTDEVFVKLLL--TPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNARS 130
L A+ TDEV+ +++L P N++ +E + R V SF KTLT SD +
Sbjct: 79 LKAEQDTDEVYAQVMLMPEPEQNEMAVEKTTPTSGPVQARPPVRSFCKTLTASDTSTHGG 138
Query: 131 FHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFV 190
F + R AD P LD+ +Q L D+H +F H+ RG P+R++L S W+ FV
Sbjct: 139 FSVLRRHADECLPPLDMTQSPPTQELVAKDLHSMDWRFRHIFRGQPRRHLLQ-SGWSVFV 197
Query: 191 KRKKLVAGDSVIFMKDSTGKIFVGIRRNTQFVAAAAEQ--KKDELEKAVMEALKLAEENK 248
K+LVAGD+ IF++ G++ VG+RR + ++ + V+ A K
Sbjct: 198 SSKRLVAGDAFIFLRGENGELRVGVRRAMRQLSNVPSSVISSQSMHLGVLATAWHAINTK 257
Query: 249 AFEIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKMKTDKSSRIPYQGTISIVSRTS 308
+ VYY +F++ + ES+K ++ MR +M+ + P Q + +
Sbjct: 258 SMFTVYYKPRTSPSEFIIPYDQYMESVKNNYSVGMRFRMRFE-GEEAPEQRFTGTIIGSE 316
Query: 309 NL--------WRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA-PTPFPQTKKFRTTQSS 359
NL WR L+V WDE P RV+PW +E S P P P + K+ R
Sbjct: 317 NLDPVWPESSWRSLKVRWDEPSTIPRPDRVSPWKIEPASSPPVNPLPLSRVKRPRPNAPP 376
Query: 360 AQLSD---KKETLLNGDGFPVDIQRVRHDLV 387
A KE D P QR ++ V
Sbjct: 377 ASPESPILTKEAATKVDTDPAQAQRSQNSTV 407
>dbj|BAD88200.1| putative auxin response factor [Oryza sativa (japonica
cultivar-group)]
Length = 808
Score = 177 bits (449), Expect = 6e-43
Identities = 120/391 (30%), Positives = 182/391 (45%), Gaps = 18/391 (4%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVSAVD 72
E+WH CA V +P++ V+YFPQGH+E S + + + L LC V V+
Sbjct: 24 ELWHACAGPLVTVPRVGDLVFYFPQGHIEQVEASMNQVADSQMRLYDLPSKLLCRVLNVE 83
Query: 73 LLADPHTDEVFVKLLL--TPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNARS 130
L A+ TDEV+ +++L P N++ +E + R V SF KTLT SD +
Sbjct: 84 LKAEQDTDEVYAQVMLMPEPEQNEMAVEKTTPTSGPVQARPPVRSFCKTLTASDTSTHGG 143
Query: 131 FHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFV 190
F + R AD P LD+ +Q L D+H +F H+ RG P+R++L S W+ FV
Sbjct: 144 FSVLRRHADECLPPLDMTQSPPTQELVAKDLHSMDWRFRHIFRGQPRRHLLQ-SGWSVFV 202
Query: 191 KRKKLVAGDSVIFMKDSTGKIFVGIRRNTQFVAAAAEQ--KKDELEKAVMEALKLAEENK 248
K+LVAGD+ IF++ G++ VG+RR + ++ + V+ A K
Sbjct: 203 SSKRLVAGDAFIFLRGENGELRVGVRRAMRQLSNVPSSVISSQSMHLGVLATAWHAINTK 262
Query: 249 AFEIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKMKTDKSSRIPYQGTISIVSRTS 308
+ VYY +F++ + ES+K ++ MR +M+ + P Q + +
Sbjct: 263 SMFTVYYKPRTSPSEFIIPYDQYMESVKNNYSVGMRFRMRFE-GEEAPEQRFTGTIIGSE 321
Query: 309 NL--------WRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA-PTPFPQTKKFRTTQSS 359
NL WR L+V WDE P RV+PW +E S P P P + K+ R
Sbjct: 322 NLDPVWPESSWRSLKVRWDEPSTIPRPDRVSPWKIEPASSPPVNPLPLSRVKRPRPNAPP 381
Query: 360 AQLSD---KKETLLNGDGFPVDIQRVRHDLV 387
A KE D P QR ++ V
Sbjct: 382 ASPESPILTKEAATKVDTDPAQAQRSQNSTV 412
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.319 0.133 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 773,879,104
Number of Sequences: 2540612
Number of extensions: 31682697
Number of successful extensions: 83149
Number of sequences better than 10.0: 165
Number of HSP's better than 10.0 without gapping: 103
Number of HSP's successfully gapped in prelim test: 62
Number of HSP's that attempted gapping in prelim test: 82607
Number of HSP's gapped (non-prelim): 193
length of query: 456
length of database: 863,360,394
effective HSP length: 131
effective length of query: 325
effective length of database: 530,540,222
effective search space: 172425572150
effective search space used: 172425572150
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 77 (34.3 bits)
Medicago: description of AC134322.24


