

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
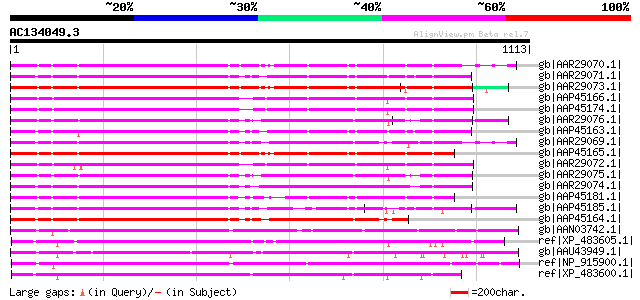
Query= AC134049.3 - phase: 0
(1113 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
gb|AAR29070.1| blight resistance protein RGA1 [Solanum bulbocast... 715 0.0
gb|AAR29071.1| blight resistance protein RGA3 [Solanum bulbocast... 711 0.0
gb|AAR29073.1| blight resistance protein B149 [Solanum bulbocast... 706 0.0
gb|AAP45166.1| putative disease resistant protein RGA4 [Solanum ... 705 0.0
gb|AAP45174.1| putative disease resistant protein rga4 [Solanum ... 704 0.0
gb|AAR29076.1| blight resistance protein T118 [Solanum tarijense] 691 0.0
gb|AAP45163.1| putative disease resistant protein RGA1 [Solanum ... 691 0.0
gb|AAR29069.1| blight resistance protein RPI [Solanum bulbocasta... 689 0.0
gb|AAP45165.1| putative disease resistant protein RGA3 [Solanum ... 688 0.0
gb|AAR29072.1| blight resistance protein RGA4 [Solanum bulbocast... 680 0.0
gb|AAR29075.1| blight resistance protein SH20 [Solanum tuberosum] 677 0.0
gb|AAR29074.1| blight resistance protein SH10 [Solanum tuberosum] 669 0.0
gb|AAP45181.1| putative disease resistant protein rga3 [Solanum ... 659 0.0
gb|AAP45185.1| putative disease resistant protein rga1 [Solanum ... 658 0.0
gb|AAP45164.1| putative disease resistant protein RGA2 [Solanum ... 630 e-179
gb|AAN03742.1| NBS-LRR-like protein [Oryza sativa (japonica cult... 548 e-154
ref|XP_483605.1| putative NBS-LRR resistance protein RGH1 [Oryza... 548 e-154
gb|AAU43949.1| putative NBS-LRR protein [Oryza sativa (japonica ... 547 e-154
ref|NP_915900.1| putative NBS-LRR type resistance protein [Oryza... 540 e-152
ref|XP_483600.1| putative NBS-LRR resistance protein RGH1 [Oryza... 539 e-151
>gb|AAR29070.1| blight resistance protein RGA1 [Solanum bulbocastanum]
Length = 992
Score = 715 bits (1846), Expect = 0.0
Identities = 453/1093 (41%), Positives = 630/1093 (57%), Gaps = 109/1093 (9%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAEA L+++L +L+ I+ E+ L GF++EF +L+S+ + I+A LEDA+EKQ I
Sbjct: 1 MAEAFLQVLLDNLTFFIQGELGLVFGFEKEFKKLSSMFSMIQAVLEDAQEKQLKYKAI-- 58
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSH-KHIAFRYKLAKKMKRIGV 119
K+WL KL AAY +DDI+D+C TEA +K + G H + I F YK+ K+MK +
Sbjct: 59 --KNWLQKLNVAAYEVDDILDDCKTEAAR--FKQAVLGRYHPRTITFCYKVGKRMKEMME 114
Query: 120 WLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVGDASE 179
LD IA E+ FHL E + ER RQT ++T+P VYGR +++D+IV L+ + S
Sbjct: 115 KLDAIAEERRNFHLDERIIERQAAR---RQTGFVLTEPKVYGREKEEDEIVKILINNVSY 171
Query: 180 QEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGAT 239
E++ V PI+G+GGLGKTTLAQ+VFN +I HF LKIWVCVS+DF KR+ KAI+E
Sbjct: 172 SEEVPVLPILGMGGLGKTTLAQMVFNDQRITEHFNLKIWVCVSDDFDEKRLIKAIVESIE 231
Query: 240 KKSCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGASILVTT 299
KS D+DL LQ+KLQ+LL KRY LVLDDVWN+ QE W L++VL G GASIL+TT
Sbjct: 232 GKSLGDMDLAPLQKKLQELLNGKRYFLVLDDVWNEDQEKWDNLRAVLKIGASGASILITT 291
Query: 300 RLPKVAKIMGTIPHHELSRLSDEDCWELFKQRAFGPNEVQQKELVIVGKEIIKKCGGFPL 359
RL K+ IMGT+ ++LS LS EDCW LFKQRAF +L+ +GKEI+KKCGG PL
Sbjct: 292 RLEKIGSIMGTLQLYQLSNLSQEDCWLLFKQRAFCHQTETSPKLMEIGKEIVKKCGGVPL 351
Query: 360 AAIALGSLLRFKREEKEWLYVKESKLWNL-QGEAYVMPALRLSYLHLPVKLRQCFSFCAL 418
AA LG LLRFKREE EW +V++S++WNL Q E V+PALRLSY HLP+ LRQCF++CA+
Sbjct: 352 AAKTLGGLLRFKREESEWEHVRDSEIWNLPQDENSVLPALRLSYHHLPLDLRQCFAYCAV 411
Query: 419 FPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWNELYWRSFFENTENVGFGQITI 478
FPKD I K+ LI LW A+ F+ S +E +D+GNEVWNELY RSFF+ E V G+ T
Sbjct: 412 FPKDTKIEKEYLIALWMAHSFLLSKGNMELEDVGNEVWNELYLRSFFQEIE-VKSGK-TY 469
Query: 479 FKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHLLIYNRNSFAEANSIQLHHVKSLK 538
FKMHDL+HDLA S+ S+R ++ + +++ ++ + SI V S
Sbjct: 470 FKMHDLIHDLATSM---FSASASSRSIRQINVKDDEDMMFIVTNYKDMMSIGFSEVVS-- 524
Query: 539 TYMEFNFDVYEAGQLSPQVLNCYSLRVLLSHRLNNLSSSIGRLKYLRYLDISEGRFKNLP 598
+Y F + +S +VLN L + L SS+G L +LRYLD+S + +LP
Sbjct: 525 SYSPSLFKRF----VSLRVLN------LSNSEFEQLPSSVGDLVHLRYLDLSGNKICSLP 574
Query: 599 NSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSLNTL 658
LCKL NL+ L L C SL LP ++L L+NL L C LTS+P +IG LT L TL
Sbjct: 575 KRLCKLQNLQTLDLYNCQSLSCLPKQTSKLCSLRNLVLDHC-PLTSMPPRIGLLTCLKTL 633
Query: 659 SKYIVGEERGFLLEELGQLNLKGQLHIKNLERLKSVTDAKKANMSRK-KLNQLWLSWERN 717
++VGE +G+ L EL LNL+G + I +LER+K+ +AK+AN+S K L+ L +SW+R
Sbjct: 634 GYFVVGERKGYQLGELRNLNLRGAISITHLERVKNDMEAKEANLSAKANLHSLSMSWDRP 693
Query: 718 EVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDCKSCL 777
+ +E ++LEAL+P+ Y + + G P W++ L ++ S+ + C++C
Sbjct: 694 NRYESEE--VKVLEALKPHPNLKY-LEIIDFCGFCLPDWMNHSVLKNVVSILISGCENCS 750
Query: 778 NLPELWKLPSLKYLKLSN-MIHVIYLFHESY-DGEGLMALKTLFLEKLPNLIGLSREERV 835
LP +LP L+ L+L + + V Y+ + +L+ L + NL GL R +
Sbjct: 751 CLPPFGELPCLESLELQDGSVEVEYVEDSGFLTRRRFPSLRKLHIGGFCNLKGLQRMKGA 810
Query: 836 -MFPRLKALEITECPNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLHFSDNEEL 894
FP L+ ++I++CP + P L S+ L I G+ + SSI L +L SL N +
Sbjct: 811 EQFPVLEEMKISDCP-MFVFPTLSSVKKLEIWGEADAGGLSSISNLSTLTSLKIFSNHTV 869
Query: 895 IYFPDGILRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCRNIEELPNEVMQR 954
+ + +NL + L L LK LPT + ++ L+ L I C +E LP E ++
Sbjct: 870 TSLLEEMFKNLEN-LIYLSVSFLENLKELPTSLASLNNLKCLDIRYCYALESLPEEGLEG 928
Query: 955 LHSLKELDIVGCDKLKLSSDFQYLTCLETLAIGSCSEVEGFHEALQHMTTLKSLTLSDLP 1014
L SL EL + C+ LK CL E LQH+TTL SL
Sbjct: 929 LSSLTELFVEHCNMLK---------CLP--------------EGLQHLTTLTSL------ 959
Query: 1015 NLEYLPECIGNLTLLHEINIYSCPKLACLPTSIQQISGLEILSIHDCSKLEKRCQKEIGE 1074
I CP+L KRC+K IGE
Sbjct: 960 ------------------KIRGCPQLI------------------------KRCEKGIGE 977
Query: 1075 DWPKIVHVQYIEI 1087
DW KI H+ + I
Sbjct: 978 DWHKISHIPNVNI 990
>gb|AAR29071.1| blight resistance protein RGA3 [Solanum bulbocastanum]
Length = 979
Score = 711 bits (1834), Expect = 0.0
Identities = 436/999 (43%), Positives = 603/999 (59%), Gaps = 58/999 (5%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAEA +++VL +L+ ++ E+ L GF EF RL+S+ +TI+A LEDA+EKQ +D +
Sbjct: 1 MAEAFIQVVLDNLTSFLKGELVLLFGFQDEFQRLSSMFSTIQAVLEDAQEKQLND----K 56
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSH-KHIAFRYKLAKKMKRIGV 119
+++WL KL A Y +DDI+DE T+A + S+ G H K I FR+K+ K+M ++
Sbjct: 57 PLENWLQKLNAATYEVDDILDEYKTKATR--FLQSEYGRYHPKVIPFRHKVGKRMDQVMK 114
Query: 120 WLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVGDASE 179
L+ IA E+ FHL E + ER R+T S++T+P VYGR+++KD+IV L+ + S+
Sbjct: 115 KLNAIAEERKNFHLQEKIIERQAAT---RETGSVLTEPQVYGRDKEKDEIVKILINNVSD 171
Query: 180 QEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGAT 239
+ LSV PI+G+GGLGKTTL+Q+VFN ++ F KIW+CVS+DF KR+ KAI+E
Sbjct: 172 AQKLSVLPILGMGGLGKTTLSQMVFNDQRVTERFYPKIWICVSDDFDEKRLIKAIVESIE 231
Query: 240 KKSCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGASILVTT 299
KS D+DL LQ+KLQ+LL KRY LVLDDVWN+ Q W L++VL G GA +L TT
Sbjct: 232 GKSLSDMDLAPLQKKLQELLNGKRYFLVLDDVWNEDQHKWANLRAVLKVGASGAFVLTTT 291
Query: 300 RLPKVAKIMGTIPHHELSRLSDEDCWELFKQRAFGPNEVQQKELVIVGKEIIKKCGGFPL 359
RL KV IMGT+ +ELS LS EDCW LF QRAFG E LV +GKEI+KKCGG PL
Sbjct: 292 RLEKVGSIMGTLQPYELSNLSPEDCWFLFMQRAFGHQEEINPNLVAIGKEIVKKCGGVPL 351
Query: 360 AAIALGSLLRFKREEKEWLYVKESKLWNL-QGEAYVMPALRLSYLHLPVKLRQCFSFCAL 418
AA LG +LRFKREE+EW +V++S +WNL Q E+ ++PALRLSY HLP+ LRQCF +CA+
Sbjct: 352 AAKTLGGILRFKREEREWEHVRDSPIWNLPQDESSILPALRLSYHHLPLDLRQCFVYCAV 411
Query: 419 FPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWNELYWRSFFENTENVGFGQITI 478
FPKD ++K+ LI W A+GF+ S LE +D+GNEVWNELY RSFF+ E V G+ T
Sbjct: 412 FPKDTKMAKENLIAFWMAHGFLLSKGNLELEDVGNEVWNELYLRSFFQEIE-VESGK-TY 469
Query: 479 FKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHLLIYNRNSFAEANSIQLHHVKSLK 538
FKMHDL+HDLA S+ S T S R + N N SI V S
Sbjct: 470 FKMHDLIHDLATSLF----------SANTSSSNIREI---NANYDGYMMSIGFAEVVS-- 514
Query: 539 TYMEFNFDVYEAGQLSPQVLNCY-SLRV--LLSHRLNNLSSSIGRLKYLRYLDISEG-RF 594
SP +L + SLRV L + LN L SSIG L +LRYLD+S R
Sbjct: 515 -------------SYSPSLLQKFVSLRVLNLRNSNLNQLPSSIGDLVHLRYLDLSGNFRI 561
Query: 595 KNLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTS 654
+NLP LC+L NL+ L L C SL LP ++L L+NL L C SLTS P +IG LT
Sbjct: 562 RNLPKRLCRLQNLQTLDLHYCDSLSCLPKQTSKLGSLRNLLLDGC-SLTSTPPRIGLLTC 620
Query: 655 LNTLSKYIVGEERGFLLEELGQLNLKGQLHIKNLERLKSVTDAKKANMSRK-KLNQLWLS 713
L +LS +++G+ +G+ L EL LNL G + I L+R+K +DAK+AN+S K L+ L LS
Sbjct: 621 LKSLSCFVIGKRKGYQLGELKNLNLYGSISITKLDRVKKDSDAKEANLSAKANLHSLCLS 680
Query: 714 WERNEVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDC 773
W+ + + ++LEAL+P++ Y + G+ G P W++ L ++ S+ + C
Sbjct: 681 WDLDGKHRYD---SEVLEALKPHSNLKY-LEINGFGGIRLPDWMNQSVLKNVVSIRIRGC 736
Query: 774 KSCLNLPELWKLPSLKYLKL-SNMIHVIYLFHESYDGEGLMALKTLFLEKLPNLIGLSRE 832
++C LP +LP L+ L+L + V Y+ + G +L+ L + NL GL ++
Sbjct: 737 ENCSCLPPFGELPCLESLELHTGSADVEYVEDNVHPGR-FPSLRKLVIWDFSNLKGLLKK 795
Query: 833 E-RVMFPRLKALEITECPNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLHFSDN 891
E FP L+ + CP + +P L S+ L + + + SI L +L SL S+N
Sbjct: 796 EGEKQFPVLEEMTFYWCP-MFVIPTLSSVKTLKVIAT-DATVLRSISNLRALTSLDISNN 853
Query: 892 EELIYFPDGILRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCRNIEELPNEV 951
E P+ + ++LA+ LK L LK LPT + ++AL+ L C +E LP E
Sbjct: 854 VEATSLPEEMFKSLAN-LKYLNISFFRNLKELPTSLASLNALKSLKFEFCNALESLPEEG 912
Query: 952 MQRLHSLKELDIVGCDKLK-LSSDFQYLTCLETLAIGSC 989
++ L SL EL + C LK L Q+LT L TL I C
Sbjct: 913 VKGLTSLTELSVSNCMMLKCLPEGLQHLTALTTLTITQC 951
Score = 39.3 bits (90), Expect = 0.75
Identities = 35/145 (24%), Positives = 67/145 (46%), Gaps = 5/145 (3%)
Query: 874 PSSIHKLGSLESLHFSDNEELIYFPDGILRNLASPLKTLGFHRHSKLKMLPTEMIHIHAL 933
PS + K SL L+ N L P I L+ L + +++ LP + + L
Sbjct: 518 PSLLQKFVSLRVLNLR-NSNLNQLPSSI--GDLVHLRYLDLSGNFRIRNLPKRLCRLQNL 574
Query: 934 QQLYINDCRNIEELPNEVMQRLHSLKELDIVGCDKLKLSSDFQYLTCLETLAIGSCSEVE 993
Q L ++ C ++ LP + +L SL+ L + GC LTCL++L+ + +
Sbjct: 575 QTLDLHYCDSLSCLPKQT-SKLGSLRNLLLDGCSLTSTPPRIGLLTCLKSLSCFVIGKRK 633
Query: 994 GFH-EALQHMTTLKSLTLSDLPNLE 1017
G+ L+++ S++++ L ++
Sbjct: 634 GYQLGELKNLNLYGSISITKLDRVK 658
>gb|AAR29073.1| blight resistance protein B149 [Solanum bulbocastanum]
Length = 971
Score = 706 bits (1822), Expect = 0.0
Identities = 431/999 (43%), Positives = 608/999 (60%), Gaps = 39/999 (3%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAEA ++++L +L+ I+ E+ L GF++EF +L+S+ + I+A LEDA+EKQ I
Sbjct: 1 MAEAFIQVLLDNLTFFIQGELGLVFGFEKEFKKLSSMFSMIQAVLEDAQEKQLKYKAI-- 58
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSH-KHIAFRYKLAKKMKRIGV 119
K+WL KL AAY +DDI+D+C TEA +K + G H + I F YK+ K+MK +
Sbjct: 59 --KNWLQKLNVAAYEVDDILDDCKTEAAR--FKQAVLGRYHPRTITFCYKVGKRMKEMME 114
Query: 120 WLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVGDASE 179
LD IA E+ FHL E + ER RQT ++T+P VYGR +++D+IV L+ + S
Sbjct: 115 KLDAIAEERRNFHLDERIIERQAAR---RQTGFVLTEPKVYGREKEEDEIVKILINNVSY 171
Query: 180 QEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGAT 239
E++ V PI+G+GGLGKTTLAQ+VFN +I HF LKIWVCVS+DF KR+ KAI+E
Sbjct: 172 SEEVPVLPILGMGGLGKTTLAQMVFNDQRITEHFNLKIWVCVSDDFDEKRLIKAIVESIE 231
Query: 240 KKSCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGASILVTT 299
KS D+DL LQ+KLQ+LL KRY LVLDDVWN+ QE W L++VL G GASIL+TT
Sbjct: 232 GKSLGDMDLAPLQKKLQELLNGKRYFLVLDDVWNEDQEKWDNLRAVLKIGASGASILITT 291
Query: 300 RLPKVAKIMGTIPHHELSRLSDEDCWELFKQRAFGPNEVQQKELVIVGKEIIKKCGGFPL 359
RL K+ IMGT+ ++LS LS EDCW LFKQRAF +L+ +GKEI+KKCGG PL
Sbjct: 292 RLEKIGSIMGTLQLYQLSNLSQEDCWLLFKQRAFCHQTETSPKLMEIGKEIVKKCGGVPL 351
Query: 360 AAIALGSLLRFKREEKEWLYVKESKLWNL-QGEAYVMPALRLSYLHLPVKLRQCFSFCAL 418
AA LG LLRFKREE EW +V++S++W L Q E V+PALRLSY HLP+ LRQCF++CA+
Sbjct: 352 AAKTLGGLLRFKREESEWEHVRDSEIWXLPQDENSVLPALRLSYHHLPLDLRQCFAYCAV 411
Query: 419 FPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWNELYWRSFFENTENVGFGQITI 478
FPKD I K+ LI LW A+ F+ S +E +D+GNEVWNELY RSFF+ E V G+ T
Sbjct: 412 FPKDTKIEKEYLIALWMAHSFLLSKGNMELEDVGNEVWNELYLRSFFQGIE-VKSGK-TY 469
Query: 479 FKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHLLIYNRNSFAEANSIQLHHVKSLK 538
FKMHDL+HDLA S+ S+R ++ + +++ ++ + SI V S
Sbjct: 470 FKMHDLIHDLATSM---FSASASSRSIRQINVKDDEDMMFIVTNYKDMMSIGFSEVVS-- 524
Query: 539 TYMEFNFDVYEAGQLSPQVLNCYSLRVLLSHRLNNLSSSIGRLKYLRYLDISEGRFKNLP 598
+Y F + +S +VLN L + L SS+G L +LRYLD+S + +LP
Sbjct: 525 SYSPSLFKRF----VSLRVLN------LSNSEFEQLPSSVGDLVHLRYLDLSGNKICSLP 574
Query: 599 NSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSLNTL 658
LCKL NL+ L L C SL LP ++L L+NL L C LTS+P +IG LT L TL
Sbjct: 575 KRLCKLRNLQTLDLYNCQSLSCLPKQTSKLCSLRNLVLDHC-PLTSMPPRIGLLTCLKTL 633
Query: 659 SKYIVGEERGFLLEELGQLNLKGQLHIKNLERLKSVTDAKKANMSRK-KLNQLWLSWERN 717
++VGE +G+ L EL LNL+G + I +LER+K+ +AK+AN+S K L+ L +SW+R
Sbjct: 634 GYFVVGERKGYQLGELRNLNLRGAISITHLERVKNDMEAKEANLSAKANLHSLSMSWDRP 693
Query: 718 EVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDCKSCL 777
+ +E ++LEAL+P+ Y + + G P W++ L ++ S+ + C++C
Sbjct: 694 NRYESEE--VKVLEALKPHPNLKY-LEIIDFCGFCLPDWMNHSVLKNVVSILISGCENCS 750
Query: 778 NLPELWKLPSLKYLKLSN-MIHVIYLFHESY-DGEGLMALKTLFLEKLPNLIGLSREERV 835
LP +LP L+ L+L + + V Y+ + +L+ L + NL GL R +
Sbjct: 751 CLPPFGELPCLESLELQDGSVEVEYVEDSGFLTRRRFPSLRKLHIGGFCNLKGLQRMKGA 810
Query: 836 -MFPRLKALEITECPNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLHFSDNEEL 894
FP L+ ++I++CP + P L S+ L I G+ + SSI L +L SL N +
Sbjct: 811 EQFPVLEEMKISDCP-MFVFPTLSSVKKLEIWGEADAGGLSSISNLSTLTSLKIFSNHTV 869
Query: 895 IYFPDGILRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCRNIEELPNEVMQR 954
+ + +NL + L L LK LPT + ++ L+ L I C +E LP E ++
Sbjct: 870 TSLLEEMFKNLEN-LIYLSVSFLENLKELPTSLASLNNLKCLDIRYCYALESLPEEGLEG 928
Query: 955 LHSLKELDIVGCDKLK-LSSDFQYLTCLETLAIGSCSEV 992
L SL EL + C+ LK L Q+LT L +L I C ++
Sbjct: 929 LSSLTELFVEHCNMLKCLPEGLQHLTTLTSLKIRGCPQL 967
Score = 56.6 bits (135), Expect = 5e-06
Identities = 70/265 (26%), Positives = 104/265 (38%), Gaps = 38/265 (14%)
Query: 838 PRLKALEITE-----CPNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLHFSDNE 892
P LK LEI + P+ + L ++ + I G N +L LESL D
Sbjct: 711 PNLKYLEIIDFCGFCLPDWMNHSVLKNVVSILISGCENCSCLPPFGELPCLESLELQDGS 770
Query: 893 -ELIYFPDG--ILRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCRNIEELPN 949
E+ Y D + R L+ L LK L + +Q + + I + P
Sbjct: 771 VEVEYVEDSGFLTRRRFPSLRKLHIGGFCNLKGLQ----RMKGAEQFPVLEEMKISDCPM 826
Query: 950 EVMQRLHSLKELDIVGCDKLKLSSDFQYLTCLETLAIGSCSEVEGF-HEALQHMTTLKSL 1008
V L S+K+L+I G S L+ L +L I S V E +++ L L
Sbjct: 827 FVFPTLSSVKKLEIWGEADAGGLSSISNLSTLTSLKIFSNHTVTSLLEEMFKNLENLIYL 886
Query: 1009 TLSDLPNLEYLP-------------------------ECIGNLTLLHEINIYSCPKLACL 1043
++S L NL+ LP E + L+ L E+ + C L CL
Sbjct: 887 SVSFLENLKELPTSLASLNNLKCLDIRYCYALESLPEEGLEGLSSLTELFVEHCNMLKCL 946
Query: 1044 PTSIQQISGLEILSIHDCSKLEKRC 1068
P +Q ++ L L I C +L KRC
Sbjct: 947 PEGLQHLTTLTSLKIRGCPQLIKRC 971
>gb|AAP45166.1| putative disease resistant protein RGA4 [Solanum bulbocastanum]
gi|46576965|sp|Q7XA39|RGA4_SOLBU Putative disease
resistance protein RGA4 (RGA4-blb)
Length = 988
Score = 705 bits (1820), Expect = 0.0
Identities = 427/1007 (42%), Positives = 598/1007 (58%), Gaps = 56/1007 (5%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAEA L+++L +L+ I ++ L GF++E +L+S+ +TI+A L+DA+EKQ D I
Sbjct: 1 MAEAFLQVLLENLTSFIGDKLVLIFGFEKECEKLSSVFSTIQAVLQDAQEKQLKDKAI-- 58
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSHKHIA-FRYKLAKKMKRIGV 119
++WL KL AAY +DDI+ EC EA+ E S+ G H I FR+K+ ++MK I
Sbjct: 59 --ENWLQKLNSAAYEVDDILGECKNEAIRFEQ--SRLGFYHPGIINFRHKIGRRMKEIME 114
Query: 120 WLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVGDASE 179
LD I+ E+ KFH E + ER R+T ++T+P VYGR++++D+IV L+ + +
Sbjct: 115 KLDAISEERRKFHFLEKITERQAAAAT-RETGFVLTEPKVYGRDKEEDEIVKILINNVNV 173
Query: 180 QEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGAT 239
E+L V+PI+G+GGLGKTTLAQ++FN +++ HF KIWVCVS+DF KR+ K II
Sbjct: 174 AEELPVFPIIGMGGLGKTTLAQMIFNDERVTKHFNPKIWVCVSDDFDEKRLIKTIIGNIE 233
Query: 240 KKSCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGASILVTT 299
+ S DL Q+KLQ+LL KRYLLVLDDVWND E W +L++VL G +GASIL TT
Sbjct: 234 RSSPHVEDLASFQKKLQELLNGKRYLLVLDDVWNDDLEKWAKLRAVLTVGARGASILATT 293
Query: 300 RLPKVAKIMGTIPHHELSRLSDEDCWELFKQRAFGPNEVQQKELVIVGKEIIKKCGGFPL 359
RL KV IMGT+ + LS LS D LF QRAFG + LV +GKEI+KKCGG PL
Sbjct: 294 RLEKVGSIMGTLQPYHLSNLSPHDSLLLFMQRAFGQQKEANPNLVAIGKEIVKKCGGVPL 353
Query: 360 AAIALGSLLRFKREEKEWLYVKESKLWNL-QGEAYVMPALRLSYLHLPVKLRQCFSFCAL 418
AA LG LLRFKREE EW +V+++++W+L Q E+ ++PALRLSY HLP+ LRQCF++CA+
Sbjct: 354 AAKTLGGLLRFKREESEWEHVRDNEIWSLPQDESSILPALRLSYHHLPLDLRQCFAYCAV 413
Query: 419 FPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWNELYWRSFFENTENVGFGQITI 478
FPKD + K+ LI LW A+GF+ S LE +D+GNEVWNELY RSFF+ E T
Sbjct: 414 FPKDTKMIKENLITLWMAHGFLLSKGNLELEDVGNEVWNELYLRSFFQEIEAKSGN--TY 471
Query: 479 FKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHLLIYNRNSFAEANSIQLHHVKSLK 538
FK+HDL+HDLA S+ ++ A +I+ +VK K
Sbjct: 472 FKIHDLIHDLATSLF---------------------------SASASCGNIREINVKDYK 504
Query: 539 TYMEFNFDVYEAGQLSPQVLNCY-SLRVL-LSH-RLNNLSSSIGRLKYLRYLDISEGRFK 595
+ F SP +L + SLRVL LS+ +L L SSIG L +LRYLD+S F+
Sbjct: 505 HTVSIGFAAV-VSSYSPSLLKKFVSLRVLNLSYSKLEQLPSSIGDLLHLRYLDLSCNNFR 563
Query: 596 NLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSL 655
+LP LCKL NL+ L + C SL LP ++L L++L + C LTS P +IG LT L
Sbjct: 564 SLPERLCKLQNLQTLDVHNCYSLNCLPKQTSKLSSLRHLVVDGC-PLTSTPPRIGLLTCL 622
Query: 656 NTLSKYIVGEERGFLLEELGQLNLKGQLHIKNLERLKSVTDAKKANMSRK-KLNQLWLSW 714
TL +IVG ++G+ L EL LNL G + I +LER+K+ TDA +AN+S K L L +SW
Sbjct: 623 KTLGFFIVGSKKGYQLGELKNLNLCGSISITHLERVKNDTDA-EANLSAKANLQSLSMSW 681
Query: 715 ERNEVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDCK 774
+ + ++ + ++LEAL+P+ Y + + G FP WI+ L + S+ + CK
Sbjct: 682 DNDGPNRYESKEVKVLEALKPHPNLKY-LEIIAFGGFRFPSWINHSVLEKVISVRIKSCK 740
Query: 775 SCLNLPELWKLPSLKYLKLSN-MIHVIYLFHESYDG-----EGLMALKTLFLEKLPNLIG 828
+CL LP +LP L+ L+L N V Y+ + +LK L + +L G
Sbjct: 741 NCLCLPPFGELPCLENLELQNGSAEVEYVEEDDVHSRFSTRRSFPSLKKLRIWFFRSLKG 800
Query: 829 LSREE-RVMFPRLKALEITECPNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLH 887
L +EE FP L+ + I CP L P L S+ L + G N + SSI L +L SL
Sbjct: 801 LMKEEGEEKFPMLEEMAILYCP-LFVFPTLSSVKKLEVHGNTNTRGLSSISNLSTLTSLR 859
Query: 888 FSDNEELIYFPDGILRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCRNIEEL 947
N P+ + +L + L+ L F LK LPT + ++AL++L I C ++E
Sbjct: 860 IGANYRATSLPEEMFTSLTN-LEFLSFFDFKNLKDLPTSLTSLNALKRLQIESCDSLESF 918
Query: 948 PNEVMQRLHSLKELDIVGCDKLK-LSSDFQYLTCLETLAIGSCSEVE 993
P + ++ L SL +L + C LK L Q+LT L L + C EVE
Sbjct: 919 PEQGLEGLTSLTQLFVKYCKMLKCLPEGLQHLTALTNLGVSGCPEVE 965
Score = 40.8 bits (94), Expect = 0.26
Identities = 39/160 (24%), Positives = 71/160 (44%), Gaps = 16/160 (10%)
Query: 874 PSSIHKLGSLESLHFSDNEELIYFPDGILRNLASPLKTLGFHRHSKL-----KMLPTEMI 928
PS + K SL L+ S ++ L L S + L R+ L + LP +
Sbjct: 520 PSLLKKFVSLRVLNLSYSK---------LEQLPSSIGDLLHLRYLDLSCNNFRSLPERLC 570
Query: 929 HIHALQQLYINDCRNIEELPNEVMQRLHSLKELDIVGCDKLKLSSDFQYLTCLETLAIGS 988
+ LQ L +++C ++ LP + +L SL+ L + GC LTCL+TL
Sbjct: 571 KLQNLQTLDVHNCYSLNCLPKQT-SKLSSLRHLVVDGCPLTSTPPRIGLLTCLKTLGFFI 629
Query: 989 CSEVEGFH-EALQHMTTLKSLTLSDLPNLEYLPECIGNLT 1027
+G+ L+++ S++++ L ++ + NL+
Sbjct: 630 VGSKKGYQLGELKNLNLCGSISITHLERVKNDTDAEANLS 669
Score = 39.7 bits (91), Expect = 0.57
Identities = 26/83 (31%), Positives = 44/83 (52%), Gaps = 3/83 (3%)
Query: 983 TLAIGSCSEVEGFHEAL-QHMTTLKSLTLSDLPNLEYLPECIGNLTLLHEINIYSCPKLA 1041
T++IG + V + +L + +L+ L LS LE LP IG+L L +++ SC
Sbjct: 506 TVSIGFAAVVSSYSPSLLKKFVSLRVLNLS-YSKLEQLPSSIGDLLHLRYLDL-SCNNFR 563
Query: 1042 CLPTSIQQISGLEILSIHDCSKL 1064
LP + ++ L+ L +H+C L
Sbjct: 564 SLPERLCKLQNLQTLDVHNCYSL 586
Score = 37.7 bits (86), Expect = 2.2
Identities = 32/121 (26%), Positives = 52/121 (42%), Gaps = 5/121 (4%)
Query: 941 CRNIEELPNEVMQRLHSLKELDIVGCDKLKLSSDFQYLTCLETLAIGSCSEVEGFHEALQ 1000
C NI E+ + + S+ +V L F L L S S++E ++
Sbjct: 492 CGNIREINVKDYKHTVSIGFAAVVSSYSPSLLKKFVSLRVLNL----SYSKLEQLPSSIG 547
Query: 1001 HMTTLKSLTLSDLPNLEYLPECIGNLTLLHEINIYSCPKLACLPTSIQQISGLEILSIHD 1060
+ L+ L LS N LPE + L L +++++C L CLP ++S L L +
Sbjct: 548 DLLHLRYLDLS-CNNFRSLPERLCKLQNLQTLDVHNCYSLNCLPKQTSKLSSLRHLVVDG 606
Query: 1061 C 1061
C
Sbjct: 607 C 607
>gb|AAP45174.1| putative disease resistant protein rga4 [Solanum bulbocastanum]
Length = 988
Score = 704 bits (1818), Expect = 0.0
Identities = 427/1007 (42%), Positives = 597/1007 (58%), Gaps = 56/1007 (5%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAEA L+++L +L+ I ++ L GF++E +L+S+ +TI+A ++DA+EKQ D I
Sbjct: 1 MAEAFLQVLLENLTSFIGDKLVLIFGFEKECEKLSSVFSTIQAVVQDAQEKQLKDKAI-- 58
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSHKHIA-FRYKLAKKMKRIGV 119
++WL KL AAY +DDI+ EC EA+ E S+ G H I FR+K+ ++MK I
Sbjct: 59 --ENWLQKLNSAAYEVDDILGECKNEAIRFEQ--SRLGFYHPGIINFRHKIGRRMKEIME 114
Query: 120 WLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVGDASE 179
LD IA E+ KFH E + ER R+T ++T+P VYGR++++D+IV L+ + +
Sbjct: 115 KLDAIAEERRKFHFLEKITERQAAAAT-RETGFVLTEPKVYGRDKEEDEIVKILINNVNV 173
Query: 180 QEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGAT 239
E+L V+PI+G+GGLGKTTLAQ++FN +++ HF KIWVCVS+DF KR+ K II
Sbjct: 174 AEELPVFPIIGMGGLGKTTLAQMIFNDERVTKHFNPKIWVCVSDDFDEKRLIKTIIGNIE 233
Query: 240 KKSCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGASILVTT 299
+ S DL Q+KLQ+LL KRYLLVLDDVWND E W +L++VL G +GASIL TT
Sbjct: 234 RSSPHVEDLASFQKKLQELLNGKRYLLVLDDVWNDDLEKWAKLRAVLTVGARGASILATT 293
Query: 300 RLPKVAKIMGTIPHHELSRLSDEDCWELFKQRAFGPNEVQQKELVIVGKEIIKKCGGFPL 359
RL KV IMGT + LS LS D LF QRAFG + LV +GKEI+KKCGG PL
Sbjct: 294 RLEKVGSIMGTSQPYHLSNLSPHDSLLLFMQRAFGQQKEANPNLVAIGKEIVKKCGGVPL 353
Query: 360 AAIALGSLLRFKREEKEWLYVKESKLWNL-QGEAYVMPALRLSYLHLPVKLRQCFSFCAL 418
AA LG LLRFKREE EW +V+++++W+L Q E+ ++PALRLSY HLP+ LRQCF++CA+
Sbjct: 354 AAKTLGGLLRFKREESEWEHVRDNEIWSLPQDESSILPALRLSYHHLPLDLRQCFAYCAV 413
Query: 419 FPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWNELYWRSFFENTENVGFGQITI 478
FPKD + K+ LI LW A+GF+ S LE +D+GNEVWNELY RSFF+ E T
Sbjct: 414 FPKDTKMIKENLITLWMAHGFLLSKGNLELEDVGNEVWNELYLRSFFQEIEAKSGN--TY 471
Query: 479 FKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHLLIYNRNSFAEANSIQLHHVKSLK 538
FK+HDL+HDLA S+ ++ A +I+ +VK K
Sbjct: 472 FKIHDLIHDLATSLF---------------------------SASASCGNIREINVKDYK 504
Query: 539 TYMEFNFDVYEAGQLSPQVLNCY-SLRVL-LSH-RLNNLSSSIGRLKYLRYLDISEGRFK 595
+ F SP +L + SLRVL LS+ +L L SSIG L +LRYLD+S F+
Sbjct: 505 HTVSIGFSAV-VSSYSPSLLKKFVSLRVLNLSYSKLEQLPSSIGDLLHLRYLDLSCNNFR 563
Query: 596 NLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSL 655
+LP LCKL NL+ L + C SL LP ++L L++L + C LTS P +IG LT L
Sbjct: 564 SLPERLCKLQNLQTLDVHNCYSLNCLPKQTSKLSSLRHLVVDGC-PLTSTPPRIGLLTCL 622
Query: 656 NTLSKYIVGEERGFLLEELGQLNLKGQLHIKNLERLKSVTDAKKANMSRK-KLNQLWLSW 714
TL +IVG ++G+ L EL LNL G + I +LER+K+ TDA +AN+S K L L +SW
Sbjct: 623 KTLGFFIVGSKKGYQLGELKNLNLCGSISITHLERVKNDTDA-EANLSAKANLQSLSMSW 681
Query: 715 ERNEVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDCK 774
+ + ++ + ++LEAL+P+ Y + + G FP WI+ L + S+ + CK
Sbjct: 682 DNDGPNRYESEEVKVLEALKPHPNLKY-LEIIAFGGFRFPSWINHSVLEKVISVRIKSCK 740
Query: 775 SCLNLPELWKLPSLKYLKLSN-MIHVIYLFHESYDG-----EGLMALKTLFLEKLPNLIG 828
+CL LP +LP L+ L+L N V Y+ + +LK L + +L G
Sbjct: 741 NCLCLPPFGELPCLENLELQNGSAEVEYVEEDDVHSRFSTRRSFPSLKKLRIWFFRSLKG 800
Query: 829 LSREE-RVMFPRLKALEITECPNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLH 887
L +EE FP L+ + I CP L P L S+ L + G N + SSI L +L SL
Sbjct: 801 LMKEEGEEKFPMLEEMAILYCP-LFVFPTLSSVKKLEVHGNTNTRGLSSISNLSTLTSLR 859
Query: 888 FSDNEELIYFPDGILRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCRNIEEL 947
N P+ + +L + L+ L F LK LPT + ++AL++L I C ++E
Sbjct: 860 IGANYRATSLPEEMFTSLTN-LEFLSFFDFKNLKDLPTSLTSLNALKRLQIESCDSLESF 918
Query: 948 PNEVMQRLHSLKELDIVGCDKLK-LSSDFQYLTCLETLAIGSCSEVE 993
P + ++ L SL +L + C LK L Q+LT L L + C EVE
Sbjct: 919 PEQGLEGLTSLTQLFVKYCKMLKCLPEGLQHLTALTNLGVSGCPEVE 965
Score = 40.8 bits (94), Expect = 0.26
Identities = 39/160 (24%), Positives = 71/160 (44%), Gaps = 16/160 (10%)
Query: 874 PSSIHKLGSLESLHFSDNEELIYFPDGILRNLASPLKTLGFHRHSKL-----KMLPTEMI 928
PS + K SL L+ S ++ L L S + L R+ L + LP +
Sbjct: 520 PSLLKKFVSLRVLNLSYSK---------LEQLPSSIGDLLHLRYLDLSCNNFRSLPERLC 570
Query: 929 HIHALQQLYINDCRNIEELPNEVMQRLHSLKELDIVGCDKLKLSSDFQYLTCLETLAIGS 988
+ LQ L +++C ++ LP + +L SL+ L + GC LTCL+TL
Sbjct: 571 KLQNLQTLDVHNCYSLNCLPKQT-SKLSSLRHLVVDGCPLTSTPPRIGLLTCLKTLGFFI 629
Query: 989 CSEVEGFH-EALQHMTTLKSLTLSDLPNLEYLPECIGNLT 1027
+G+ L+++ S++++ L ++ + NL+
Sbjct: 630 VGSKKGYQLGELKNLNLCGSISITHLERVKNDTDAEANLS 669
Score = 39.3 bits (90), Expect = 0.75
Identities = 26/83 (31%), Positives = 44/83 (52%), Gaps = 3/83 (3%)
Query: 983 TLAIGSCSEVEGFHEAL-QHMTTLKSLTLSDLPNLEYLPECIGNLTLLHEINIYSCPKLA 1041
T++IG + V + +L + +L+ L LS LE LP IG+L L +++ SC
Sbjct: 506 TVSIGFSAVVSSYSPSLLKKFVSLRVLNLS-YSKLEQLPSSIGDLLHLRYLDL-SCNNFR 563
Query: 1042 CLPTSIQQISGLEILSIHDCSKL 1064
LP + ++ L+ L +H+C L
Sbjct: 564 SLPERLCKLQNLQTLDVHNCYSL 586
Score = 37.4 bits (85), Expect = 2.8
Identities = 32/121 (26%), Positives = 52/121 (42%), Gaps = 5/121 (4%)
Query: 941 CRNIEELPNEVMQRLHSLKELDIVGCDKLKLSSDFQYLTCLETLAIGSCSEVEGFHEALQ 1000
C NI E+ + + S+ +V L F L L S S++E ++
Sbjct: 492 CGNIREINVKDYKHTVSIGFSAVVSSYSPSLLKKFVSLRVLNL----SYSKLEQLPSSIG 547
Query: 1001 HMTTLKSLTLSDLPNLEYLPECIGNLTLLHEINIYSCPKLACLPTSIQQISGLEILSIHD 1060
+ L+ L LS N LPE + L L +++++C L CLP ++S L L +
Sbjct: 548 DLLHLRYLDLS-CNNFRSLPERLCKLQNLQTLDVHNCYSLNCLPKQTSKLSSLRHLVVDG 606
Query: 1061 C 1061
C
Sbjct: 607 C 607
>gb|AAR29076.1| blight resistance protein T118 [Solanum tarijense]
Length = 948
Score = 691 bits (1782), Expect = 0.0
Identities = 425/1011 (42%), Positives = 597/1011 (59%), Gaps = 86/1011 (8%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAEA ++++L +++ I+ E+ L LGF+ EF ++S +TI+A LEDA+EKQ D I
Sbjct: 1 MAEAFIQVLLENITSFIQGELGLLLGFENEFENISSRFSTIQAVLEDAQEKQLKDKAI-- 58
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSH-KHIAFRYKLAKKMKRIGV 119
K+WL KL AAY +DD++DEC LE S+ G H K I FR+K+ K++K +
Sbjct: 59 --KNWLQKLNAAAYKVDDLLDECKAARLEQ----SRLGRHHPKAIVFRHKIGKRIKEMME 112
Query: 120 WLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVGDASE 179
LD IA E+ FHL E + ER P+ T ++T+P VYGR++++D+IV L+ + S
Sbjct: 113 KLDAIAKERTDFHLHEKIIERQVARPE---TGPVLTEPQVYGRDKEEDEIVKILINNVSN 169
Query: 180 QEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGAT 239
+LSV PI+G+GGLGKTTLAQ+VFN ++ HF KIW+CVS+DF KR+ + II
Sbjct: 170 ALELSVLPILGMGGLGKTTLAQMVFNDQRVTEHFYPKIWICVSDDFDEKRLIETIIGNIE 229
Query: 240 KKSCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGASILVTT 299
+ S + DL Q+KLQ LL KRYLLVLDDVWN+ Q+ W L++VL G GAS+L TT
Sbjct: 230 RSSLDVKDLASFQKKLQQLLNGKRYLLVLDDVWNEDQQKWDNLRAVLKVGASGASVLTTT 289
Query: 300 RLPKVAKIMGTIPHHELSRLSDEDCWELFKQRAFGPNEVQQKELVIVGKEIIKKCGGFPL 359
RL KV IMGT+ ++LS LS +DCW LF QRA+ E LV +GKEI+KK GG PL
Sbjct: 290 RLEKVGSIMGTLQPYQLSNLSQDDCWLLFIQRAYRHQEEISPNLVAIGKEIVKKSGGVPL 349
Query: 360 AAIALGSLLRFKREEKEWLYVKESKLWNL-QGEAYVMPALRLSYLHLPVKLRQCFSFCAL 418
AA LG LLRFKRE++EW +V++ ++WNL Q E ++P LRLSY HLP+ LRQCF++CA+
Sbjct: 350 AAKTLGGLLRFKREKREWEHVRDREIWNLPQDEMSILPVLRLSYHHLPLDLRQCFAYCAV 409
Query: 419 FPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWNELYWRSFFENTENVGFGQITI 478
FPKD + K+ +I LW A+GF+ S + LE +D+GNEVWNELY RSFF+ E V +G T
Sbjct: 410 FPKDTKMEKKKVISLWMAHGFLLSRRNLELEDVGNEVWNELYLRSFFQEIE-VRYGN-TY 467
Query: 479 FKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHLLIYNRNSFAEANSIQLHHVKSLK 538
FKMHDL+HDLA S+ S T S R + N S+
Sbjct: 468 FKMHDLIHDLATSLF----------SANTSSSNIREI---NVESY--------------- 499
Query: 539 TYMEFNFDVYE-AGQLSPQVLNCY-SLRVL-LSH-RLNNLSSSIGRLKYLRYLDISEG-R 593
T+M + E SP +L + SLRVL LS+ + L SSIG L +LRY+D+S
Sbjct: 500 THMMMSIGFSEVVSSYSPSLLQKFVSLRVLNLSYSKFEELPSSIGDLVHLRYMDLSNNIE 559
Query: 594 FKNLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLT 653
++LP LCKL NL+ L L C L LP ++L L+NL L C LT P +IG LT
Sbjct: 560 IRSLPKQLCKLQNLQTLDLQYCTRLCCLPKQTSKLGSLRNLLLHGCHRLTRTPPRIGSLT 619
Query: 654 SLNTLSKYIVGEERGFLLEELGQLNLKGQLHIKNLERLKSVTDAKKANMSRKK-LNQLWL 712
L TL +++V ++G+ L ELG LNL G + I +LER+K+ +AK+AN+S K+ L+ L +
Sbjct: 620 CLKTLGQFVVKRKKGYQLGELGSLNLYGSIKISHLERVKNDKEAKEANLSAKENLHSLSM 679
Query: 713 SWERNEVSQLQENVE-QILEALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELV 771
W+ +E E+ E ++LEAL+P++ L + G+ G P W++ L ++ +E+
Sbjct: 680 KWDDDERPHRYESEEVEVLEALKPHS-NLTCLTISGFRGIRLPDWMNHSVLKNIVLIEIS 738
Query: 772 DCKSCLNLPELWKLPSLKYLKLSNMIHVIYLFHESYDGEG--------LMALKTLFLEKL 823
CK+C LP LP L+ L+L Y+ D E +L+ L + K
Sbjct: 739 GCKNCSCLPPFGDLPCLESLQLYRG-SAEYVEEVDIDVEDSGFPTRIRFPSLRKLCICKF 797
Query: 824 PNLIGLSREE-RVMFPRLKALEITECPNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGS 882
NL GL ++E FP L+ +EI CP +P+LS L +
Sbjct: 798 DNLKGLVKKEGGEQFPVLEEMEIRYCP-------IPTLSS----------------NLKA 834
Query: 883 LESLHFSDNEELIYFPDGILRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCR 942
L SL+ SDN+E FP+ + ++LA+ LK L LK LPT + ++AL+ L I C
Sbjct: 835 LTSLNISDNKEATSFPEEMFKSLAN-LKYLNISHFKNLKELPTSLASLNALKSLKIQWCC 893
Query: 943 NIEELPNEVMQRLHSLKELDIVGCDKLK-LSSDFQYLTCLETLAIGSCSEV 992
+E +P E ++ L SL EL + C LK L Q+LT L + I C ++
Sbjct: 894 ALESIPEEGVKGLTSLTELIVKFCKMLKCLPEGLQHLTALTRVKIWGCPQL 944
Score = 60.1 bits (144), Expect = 4e-07
Identities = 63/253 (24%), Positives = 105/253 (40%), Gaps = 29/253 (11%)
Query: 822 KLPNLIGLSREERVMFPRLKALEITEC-PNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKL 880
+LP+ + S + ++ + + C P LPCL SL +Y +++ +
Sbjct: 719 RLPDWMNHSVLKNIVLIEISGCKNCSCLPPFGDLPCLESLQLYRGSAEYVEEVDIDVEDS 778
Query: 881 GSLESLHFSDNEELIYFPDGILRNLASPLKTLGFHRHSKLKM----LPTEMIHIHALQQL 936
G + F +L L+ L F ++++ +PT ++ AL L
Sbjct: 779 GFPTRIRFPSLRKLCICKFDNLKGLVKKEGGEQFPVLEEMEIRYCPIPTLSSNLKALTSL 838
Query: 937 YINDCRNIEELPNEVMQRLHSLKELDIVGCDKLKLSSDFQYLTCLETLAIGSCSEVEGFH 996
I+D + P E+ + L +LK L+I S F+ L L T
Sbjct: 839 NISDNKEATSFPEEMFKSLANLKYLNI---------SHFKNLKELPT------------- 876
Query: 997 EALQHMTTLKSLTLSDLPNLEYLPE-CIGNLTLLHEINIYSCPKLACLPTSIQQISGLEI 1055
+L + LKSL + LE +PE + LT L E+ + C L CLP +Q ++ L
Sbjct: 877 -SLASLNALKSLKIQWCCALESIPEEGVKGLTSLTELIVKFCKMLKCLPEGLQHLTALTR 935
Query: 1056 LSIHDCSKLEKRC 1068
+ I C +L KRC
Sbjct: 936 VKIWGCPQLIKRC 948
Score = 39.7 bits (91), Expect = 0.57
Identities = 23/95 (24%), Positives = 44/95 (46%), Gaps = 8/95 (8%)
Query: 988 SCSEVEGFHEALQHMTTLKSLTLSDLPNLEYLPECIGNLTLLHEINIYSCPKLACLPTSI 1047
S S+ E ++ + L+ + LS+ + LP+ + L L +++ C +L CLP
Sbjct: 532 SYSKFEELPSSIGDLVHLRYMDLSNNIEIRSLPKQLCKLQNLQTLDLQYCTRLCCLPKQT 591
Query: 1048 QQISGLEILSIHDCSKLEK--------RCQKEIGE 1074
++ L L +H C +L + C K +G+
Sbjct: 592 SKLGSLRNLLLHGCHRLTRTPPRIGSLTCLKTLGQ 626
>gb|AAP45163.1| putative disease resistant protein RGA1 [Solanum bulbocastanum]
gi|46576967|sp|Q7XA42|RGA1_SOLBU Putative disease
resistance protein RGA1 (RGA3-blb)
Length = 1025
Score = 691 bits (1782), Expect = 0.0
Identities = 438/1042 (42%), Positives = 601/1042 (57%), Gaps = 98/1042 (9%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAEA +++VL +L+ ++ E+ L GF EF RL+S+ +TI+A LEDA+EKQ +D +
Sbjct: 1 MAEAFIQVVLDNLTSFLKGELVLLFGFQDEFQRLSSMFSTIQAVLEDAQEKQLND----K 56
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSH-KHIAFRYKLAKKMKRIGV 119
+++WL KL A Y +DDI+DE T+A + S+ G H K I FR+K+ K+M ++
Sbjct: 57 PLENWLQKLNAATYEVDDILDEYKTKATR--FLQSEYGRYHPKVIPFRHKVGKRMDQVMK 114
Query: 120 WLDDIAAEKNKFHLTEIVRERSGVVPD--------------------------------- 146
L+ IA E+ KFHL E + ER +
Sbjct: 115 KLNAIAEERKKFHLQEKIIERQAATRETVSMFSTCSSSHPYLFIVQNPSLFNYFLPTPKE 174
Query: 147 --------WRQTT--SIVTQPLVYGRNEDKDKIVDFLVGDASEQEDLSVYPIVGLGGLGK 196
W S++T+P VYGR+++KD+IV L+ AS+ + LSV PI+G+GGLGK
Sbjct: 175 LECNNILIWTLLAPGSVLTEPQVYGRDKEKDEIVKILINTASDAQKLSVLPILGMGGLGK 234
Query: 197 TTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGATKKSCEDLDLELLQRKLQ 256
TTL+Q+VFN ++ F KIW+C+S+DF KR+ KAI+E KS D+DL LQ+KLQ
Sbjct: 235 TTLSQMVFNDQRVTERFYPKIWICISDDFNEKRLIKAIVESIEGKSLSDMDLAPLQKKLQ 294
Query: 257 DLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGASILVTTRLPKVAKIMGTIPHHEL 316
+LL KRY LVLDDVWN+ Q W L++VL G GA +L TTRL KV IMGT+ +EL
Sbjct: 295 ELLNGKRYFLVLDDVWNEDQHKWANLRAVLKVGASGAFVLTTTRLEKVGSIMGTLQPYEL 354
Query: 317 SRLSDEDCWELFKQRAFGPNEVQQKELVIVGKEIIKKCGGFPLAAIALGSLLRFKREEKE 376
S LS EDCW LF QRAFG E L+ +GKEI+KKCGG PLAA LG +LRFKREE+E
Sbjct: 355 SNLSPEDCWFLFMQRAFGHQEEINPNLMAIGKEIVKKCGGVPLAAKTLGGILRFKREERE 414
Query: 377 WLYVKESKLWNL-QGEAYVMPALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWT 435
W +V++S +WNL Q E+ ++PALRLSY HLP+ LRQCF +CA+FPKD ++K+ LI W
Sbjct: 415 WEHVRDSPIWNLPQDESSILPALRLSYHHLPLDLRQCFVYCAVFPKDTKMAKENLIAFWM 474
Query: 436 ANGFISSNQMLEADDIGNEVWNELYWRSFFENTENVGFGQITIFKMHDLVHDLAGSVTQD 495
A+GF+ S LE +D+GNEVWNELY RSFF+ E V G+ T FKMHDL+HDLA S+
Sbjct: 475 AHGFLLSKGNLELEDVGNEVWNELYLRSFFQEIE-VESGK-TYFKMHDLIHDLATSLF-- 530
Query: 496 VCCITDDNSMRTMSEETRHLLIYNRNSFAEANSIQLHHVKSLKTYMEFNFDVYEAGQLSP 555
S T S R + N N SI V S SP
Sbjct: 531 --------SANTSSSNIREI---NANYDGYMMSIGFAEVVS---------------SYSP 564
Query: 556 QVLNCY-SLRV--LLSHRLNNLSSSIGRLKYLRYLDISEG-RFKNLPNSLCKLCNLEVLK 611
+L + SLRV L + LN L SSIG L +LRYLD+S R +NLP LCKL NL+ L
Sbjct: 565 SLLQKFVSLRVLNLRNSNLNQLPSSIGDLVHLRYLDLSGNFRIRNLPKRLCKLQNLQTLD 624
Query: 612 LDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSLNTLSKYIVGEERGFLL 671
L C SL LP ++L L+NL L C SLTS P +IG LT L +LS +++G+ +G L
Sbjct: 625 LHYCDSLSCLPKQTSKLGSLRNLLLDGC-SLTSTPPRIGLLTCLKSLSCFVIGKRKGHQL 683
Query: 672 EELGQLNLKGQLHIKNLERLKSVTDAKKANMSRK-KLNQLWLSWERNEVSQLQENVEQIL 730
EL LNL G + I L+R+K TDAK+AN+S K L+ L LSW+ + + ++L
Sbjct: 684 GELKNLNLYGSISITKLDRVKKDTDAKEANLSAKANLHSLCLSWDLDGKHRYD---SEVL 740
Query: 731 EALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDCKSCLNLPELWKLPSLKY 790
EAL+P++ Y + G+ G P W++ L ++ S+ + C++C LP +LP L+
Sbjct: 741 EALKPHSNLKY-LEINGFGGIRLPDWMNQSVLKNVVSIRIRGCENCSCLPPFGELPCLES 799
Query: 791 LKL-SNMIHVIYLFHESYDGEGLMALKTLFLEKLPNLIGLSR-EERVMFPRLKALEITEC 848
L+L + V Y+ + G +L+ L + NL GL + E FP L+ + C
Sbjct: 800 LELHTGSADVEYVEDNVHPGR-FPSLRKLVIWDFSNLKGLLKMEGEKQFPVLEEMTFYWC 858
Query: 849 PNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLHFSDNEELIYFPDGILRNLASP 908
P + +P L S+ L + + + SI L +L SL SDN E P+ + ++LA+
Sbjct: 859 P-MFVIPTLSSVKTLKVI-VTDATVLRSISNLRALTSLDISDNVEATSLPEEMFKSLAN- 915
Query: 909 LKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCRNIEELPNEVMQRLHSLKELDIVGCDK 968
LK L LK LPT + ++AL+ L C +E LP E ++ L SL EL + C
Sbjct: 916 LKYLKISFFRNLKELPTSLASLNALKSLKFEFCDALESLPEEGVKGLTSLTELSVSNCMM 975
Query: 969 LK-LSSDFQYLTCLETLAIGSC 989
LK L Q+LT L TL I C
Sbjct: 976 LKCLPEGLQHLTALTTLTITQC 997
>gb|AAR29069.1| blight resistance protein RPI [Solanum bulbocastanum]
gi|32693281|gb|AAP86601.1| putative disease resistant
protein RGA2 [Solanum bulbocastanum]
gi|46576968|sp|Q7XBQ9|RGA2_SOLBU Disease resistance
protein RGA2 (RGA2-blb) (Blight resistance protein RPI)
Length = 970
Score = 689 bits (1777), Expect = 0.0
Identities = 441/1099 (40%), Positives = 618/1099 (56%), Gaps = 143/1099 (13%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAEA ++++L +L+ ++ E+ L GF EF RL+S+ +TI+A LEDA+EKQ ++ +
Sbjct: 1 MAEAFIQVLLDNLTSFLKGELVLLFGFQDEFQRLSSMFSTIQAVLEDAQEKQLNN----K 56
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSH-KHIAFRYKLAKKMKRIGV 119
+++WL KL A Y +DDI+DE T+A + S+ G H K I FR+K+ K+M ++
Sbjct: 57 PLENWLQKLNAATYEVDDILDEYKTKATR--FSQSEYGRYHPKVIPFRHKVGKRMDQVMK 114
Query: 120 WLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVGDASE 179
L IA E+ FHL E + ER V R+T S++T+P VYGR+++KD+IV L+ + S+
Sbjct: 115 KLKAIAEERKNFHLHEKIVERQAVR---RETGSVLTEPQVYGRDKEKDEIVKILINNVSD 171
Query: 180 QEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGAT 239
+ LSV PI+G+GGLGKTTLAQ+VFN ++ HF KIW+CVSEDF KR+ KAI+E
Sbjct: 172 AQHLSVLPILGMGGLGKTTLAQMVFNDQRVTEHFHSKIWICVSEDFDEKRLIKAIVESIE 231
Query: 240 KKSC-EDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGASILVT 298
+ ++DL LQ+KLQ+LL KRYLLVLDDVWN+ Q+ W L++VL G GAS+L T
Sbjct: 232 GRPLLGEMDLAPLQKKLQELLNGKRYLLVLDDVWNEDQQKWANLRAVLKVGASGASVLTT 291
Query: 299 TRLPKVAKIMGTIPHHELSRLSDEDCWELFKQRAFGPNEVQQKELVIVGKEIIKKCGGFP 358
TRL KV IMGT+ +ELS LS EDCW LF QRAFG E LV +GKEI+KK GG P
Sbjct: 292 TRLEKVGSIMGTLQPYELSNLSQEDCWLLFMQRAFGHQEEINPNLVAIGKEIVKKSGGVP 351
Query: 359 LAAIALGSLLRFKREEKEWLYVKESKLWNL-QGEAYVMPALRLSYLHLPVKLRQCFSFCA 417
LAA LG +L FKREE+ W +V++S +WNL Q E+ ++PALRLSY LP+ L+QCF++CA
Sbjct: 352 LAAKTLGGILCFKREERAWEHVRDSPIWNLPQDESSILPALRLSYHQLPLDLKQCFAYCA 411
Query: 418 LFPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWNELYWRSFFENTENVGFGQIT 477
+FPKD + K+ LI LW A+GF+ S +E +D+G+EVW ELY RSFF+ E V G+ T
Sbjct: 412 VFPKDAKMEKEKLISLWMAHGFLLSKGNMELEDVGDEVWKELYLRSFFQEIE-VKDGK-T 469
Query: 478 IFKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHLLIYNRNSFAEANSIQLHHVKSL 537
FKMHDL+HDLA S+ S T S R + N++S+ SI V
Sbjct: 470 YFKMHDLIHDLATSLF----------SANTSSSNIREI---NKHSYTHMMSIGFAEVVFF 516
Query: 538 KTYMEFNFDVYEAGQLSPQVLNCYSLRVLL--SHRLNNLSSSIGRLKYLRYLDISEGRFK 595
T P + SLRVL N L SSIG L +LRYL++ +
Sbjct: 517 YTL--------------PPLEKFISLRVLNLGDSTFNKLPSSIGDLVHLRYLNLYGSGMR 562
Query: 596 NLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSL 655
+LP LCKL NL+ L L C L LP ++L L+NL L SLT +P +IG LT L
Sbjct: 563 SLPKQLCKLQNLQTLDLQYCTKLCCLPKETSKLGSLRNLLLDGSQSLTCMPPRIGSLTCL 622
Query: 656 NTLSKYIVGEERGFLLEELGQLNLKGQLHIKNLERLKSVTDAKKANMSRK-KLNQLWLSW 714
TL +++VG ++G+ L ELG LNL G + I +LER+K+ DAK+AN+S K L+ L +SW
Sbjct: 623 KTLGQFVVGRKKGYQLGELGNLNLYGSIKISHLERVKNDKDAKEANLSAKGNLHSLSMSW 682
Query: 715 ERNEVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDCK 774
+ ++LEAL+P++ L S + G+ G + P+W++ L ++ S+ + + +
Sbjct: 683 NNFGPHIYESEEVKVLEALKPHS-NLTSLKIYGFRGIHLPEWMNHSVLKNIVSILISNFR 741
Query: 775 SCLNLPELWKLPSLKYLKLSNMIHVIYLFHESYDGEGLMALKTLFLEKLPNLIGLSREER 834
+C LP LP L+ L+ L S D E ++E++ + R
Sbjct: 742 NCSCLPPFGDLPCLESLE---------LHWGSADVE--------YVEEVDIDVHSGFPTR 784
Query: 835 VMFPRLKALEITECPNLLGL------PCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLHF 888
+ FP L+ L+I + +L GL P L ++ I L S+ L +L SL
Sbjct: 785 IRFPSLRKLDIWDFGSLKGLLKKEGEEQFPVLEEMIIHECPFLTLSSN---LRALTSLRI 841
Query: 889 SDNEELIYFPDGILRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCRNIEELP 948
N+ FP+ + +NLA+ LK L R + LK LPT + ++AL+ L I C +E LP
Sbjct: 842 CYNKVATSFPEEMFKNLAN-LKYLTISRCNNLKELPTSLASLNALKSLKIQLCCALESLP 900
Query: 949 NEVMQRLHSLKELDIVGCDKLKLSSDFQYLTCLETLAIGSCSEVEGFHEALQHMTTLKSL 1008
E ++ L SL EL + C+ LK CL E LQH+TTL SL
Sbjct: 901 EEGLEGLSSLTELFVEHCNMLK---------CLP--------------EGLQHLTTLTSL 937
Query: 1009 TLSDLPNLEYLPECIGNLTLLHEINIYSCPKLACLPTSIQQISGLEILSIHDCSKLEKRC 1068
I CP+L KRC
Sbjct: 938 ------------------------KIRGCPQLI------------------------KRC 949
Query: 1069 QKEIGEDWPKIVHVQYIEI 1087
+K IGEDW KI H+ + I
Sbjct: 950 EKGIGEDWHKISHIPNVNI 968
>gb|AAP45165.1| putative disease resistant protein RGA3 [Solanum bulbocastanum]
gi|46576966|sp|Q7XA40|RGA3_SOLBU Putative disease
resistance protein RGA3 (RGA1-blb) (Blight resistance
protein B149)
Length = 947
Score = 688 bits (1775), Expect = 0.0
Identities = 417/959 (43%), Positives = 587/959 (60%), Gaps = 38/959 (3%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAEA L+++L +L+ I+ E+ L GF++EF +L+S+ + I+A LEDA+EKQ I
Sbjct: 1 MAEAFLQVLLDNLTFFIQGELGLVFGFEKEFKKLSSMFSMIQAVLEDAQEKQLKYKAI-- 58
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSH-KHIAFRYKLAKKMKRIGV 119
K+WL KL AAY +DDI+D+C TEA +K + G H + I F YK+ K+MK +
Sbjct: 59 --KNWLQKLNVAAYEVDDILDDCKTEAAR--FKQAVLGRYHPRTITFCYKVGKRMKEMME 114
Query: 120 WLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVGDASE 179
LD IA E+ FHL E + ER RQT ++T+P VYGR +++D+IV L+ + S
Sbjct: 115 KLDAIAEERRNFHLDERIIERQAAR---RQTGFVLTEPKVYGREKEEDEIVKILINNVSY 171
Query: 180 QEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGAT 239
E++ V PI+G+GGLGKTTLAQ+VFN +I HF LKIWVCVS+DF KR+ KAI+E
Sbjct: 172 SEEVPVLPILGMGGLGKTTLAQMVFNDQRITEHFNLKIWVCVSDDFDEKRLIKAIVESIE 231
Query: 240 KKSCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGASILVTT 299
KS D+DL LQ+KLQ+LL KRY LVLDDVWN+ QE W L++VL G GASIL+TT
Sbjct: 232 GKSLGDMDLAPLQKKLQELLNGKRYFLVLDDVWNEDQEKWDNLRAVLKIGASGASILITT 291
Query: 300 RLPKVAKIMGTIPHHELSRLSDEDCWELFKQRAFGPNEVQQKELVIVGKEIIKKCGGFPL 359
RL K+ IMGT+ ++LS LS EDCW LFKQRAF +L+ +GKEI+KKCGG PL
Sbjct: 292 RLEKIGSIMGTLQLYQLSNLSQEDCWLLFKQRAFCHQTETSPKLMEIGKEIVKKCGGVPL 351
Query: 360 AAIALGSLLRFKREEKEWLYVKESKLWNL-QGEAYVMPALRLSYLHLPVKLRQCFSFCAL 418
AA LG LLRFKREE EW +V++S++WNL Q E V+PALRLSY HLP+ LRQCF++CA+
Sbjct: 352 AAKTLGGLLRFKREESEWEHVRDSEIWNLPQDENSVLPALRLSYHHLPLDLRQCFAYCAV 411
Query: 419 FPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWNELYWRSFFENTENVGFGQITI 478
FPKD I K+ LI LW A+ F+ S +E +D+GNEVWNELY RSFF+ E V G+ T
Sbjct: 412 FPKDTKIEKEYLIALWMAHSFLLSKGNMELEDVGNEVWNELYLRSFFQEIE-VKSGK-TY 469
Query: 479 FKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHLLIYNRNSFAEANSIQLHHVKSLK 538
FKMHDL+HDLA S+ S+R ++ + +++ ++ + SI V S
Sbjct: 470 FKMHDLIHDLATSM---FSASASSRSIRQINVKDDEDMMFIVTNYKDMMSIGFSEVVS-- 524
Query: 539 TYMEFNFDVYEAGQLSPQVLNCYSLRVLLSHRLNNLSSSIGRLKYLRYLDISEGRFKNLP 598
+Y F + +S +VLN L + L SS+G L +LRYLD+S + +LP
Sbjct: 525 SYSPSLFKRF----VSLRVLN------LSNSEFEQLPSSVGDLVHLRYLDLSGNKICSLP 574
Query: 599 NSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSLNTL 658
LCKL NL+ L L C SL LP ++L L+NL L C LTS+P +IG LT L TL
Sbjct: 575 KRLCKLQNLQTLDLYNCQSLSCLPKQTSKLCSLRNLVLDHC-PLTSMPPRIGLLTCLKTL 633
Query: 659 SKYIVGEERGFLLEELGQLNLKGQLHIKNLERLKSVTDAKKANMSRK-KLNQLWLSWERN 717
++VGE +G+ L EL LNL+G + I +LER+K+ +AK+AN+S K L+ L +SW+R
Sbjct: 634 GYFVVGERKGYQLGELRNLNLRGAISITHLERVKNDMEAKEANLSAKANLHSLSMSWDRP 693
Query: 718 EVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDCKSCL 777
+ +E ++LEAL+P+ Y + + G P W++ L ++ S+ + C++C
Sbjct: 694 NRYESEE--VKVLEALKPHPNLKY-LEIIDFCGFCLPDWMNHSVLKNVVSILISGCENCS 750
Query: 778 NLPELWKLPSLKYLKLSN-MIHVIYLFHESY-DGEGLMALKTLFLEKLPNLIGLSREERV 835
LP +LP L+ L+L + + V Y+ + +L+ L + NL GL R +
Sbjct: 751 CLPPFGELPCLESLELQDGSVEVEYVEDSGFLTRRRFPSLRKLHIGGFCNLKGLQRMKGA 810
Query: 836 -MFPRLKALEITECPNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLHFSDNEEL 894
FP L+ ++I++CP + P L S+ L I G+ + SSI L +L SL N +
Sbjct: 811 EQFPVLEEMKISDCP-MFVFPTLSSVKKLEIWGEADAGGLSSISNLSTLTSLKIFSNHTV 869
Query: 895 IYFPDGILRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCRNIEELPNEVMQ 953
+ + +NL + L L LK LPT + ++ L+ L I C +E LP E ++
Sbjct: 870 TSLLEEMFKNLEN-LIYLSVSFLENLKELPTSLASLNNLKCLDIRYCYALESLPEEGLE 927
>gb|AAR29072.1| blight resistance protein RGA4 [Solanum bulbocastanum]
Length = 1040
Score = 680 bits (1755), Expect = 0.0
Identities = 429/1058 (40%), Positives = 600/1058 (56%), Gaps = 106/1058 (10%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAEA L+++L +L+ I ++ L GF++E +L+S+ +TI+A L+DA+EKQ D I
Sbjct: 1 MAEAFLQVLLENLTSFIGDKLVLIFGFEKECEKLSSVFSTIQAVLQDAQEKQLKDKAI-- 58
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSHKHIA-FRYKLAKKMKRIGV 119
++WL KL AAY +DDI+ EC EA+ E S+ G H I FR+K+ ++MK I
Sbjct: 59 --ENWLQKLNSAAYEVDDILGECKNEAIRFEQ--SRLGFYHPGIINFRHKIGRRMKEIME 114
Query: 120 WLDDIAAEKNKFHLTEIV---------RERSGVVPDWRQ--------------------- 149
LD I+ E+ KFH E + RE G W +
Sbjct: 115 KLDAISEERRKFHFLEKITERQAAAATRETVGWQWGWARLEYKRLLLGVLMRIMSLRMHV 174
Query: 150 -TTS--------------------IVTQPLVYGRNEDKDKIVDFLVGDASEQEDLSVYPI 188
T S ++T+P VYGR++++D+IV L+ + + E+L V+PI
Sbjct: 175 STCSTLYEFKFYLCTPKVGARRCFVLTEPKVYGRDKEEDEIVKILINNVNVAEELPVFPI 234
Query: 189 VGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGATKKSCEDLDL 248
+G+GGLGKTTLAQ++FN +++ HF KIWVCVS+DF KR+ K II + S DL
Sbjct: 235 IGMGGLGKTTLAQMIFNDERVTKHFNPKIWVCVSDDFDEKRLIKTIIGNIERSSPHVEDL 294
Query: 249 ELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGASILVTTRLPKVAKIM 308
Q+KLQ+LL KRYLLVLDDVWND E W +L++VL G +GASIL TTRL KV IM
Sbjct: 295 ASFQKKLQELLNGKRYLLVLDDVWNDDLEKWAKLRAVLTVGARGASILATTRLEKVGSIM 354
Query: 309 GTIPHHELSRLSDEDCWELFKQRAFGPNEVQQKELVIVGKEIIKKCGGFPLAAIALGSLL 368
GT+ + LS LS D LF QRAFG + LV +GKEI+KKCGG PLAA LG LL
Sbjct: 355 GTLQPYHLSNLSPHDSLLLFMQRAFGQQKEANPNLVAIGKEIVKKCGGVPLAAKTLGGLL 414
Query: 369 RFKREEKEWLYVKESKLWNL-QGEAYVMPALRLSYLHLPVKLRQCFSFCALFPKDEIISK 427
RFKREE EW +V+++++W+L Q E+ ++PALRLSY HLP+ LRQCF++CA+FPKD + K
Sbjct: 415 RFKREESEWEHVRDNEIWSLPQDESSILPALRLSYHHLPLDLRQCFAYCAVFPKDTKMIK 474
Query: 428 QLLIDLWTANGFISSNQMLEADDIGNEVWNELYWRSFFENTENVGFGQITIFKMHDLVHD 487
+ LI LW A+GF+ S LE +D+GNEVWNELY RSFF+ E T FK+HDL+HD
Sbjct: 475 ENLITLWMAHGFLLSKGNLELEDVGNEVWNELYLRSFFQEIEAKSGN--TYFKIHDLIHD 532
Query: 488 LAGSVTQDVCCITDDNSMRTMSEETRHLLIYNRNSFAEANSIQLHHVKSLKTYMEFNFDV 547
LA S+ ++ A +I+ +VK K + F
Sbjct: 533 LATSLF---------------------------SASASCGNIREINVKDYKHTVSIGFAA 565
Query: 548 YEAGQLSPQVLNCY-SLRVL-LSH-RLNNLSSSIGRLKYLRYLDISEGRFKNLPNSLCKL 604
SP +L + SLRVL LS+ +L L SSIG L +LRYLD+S F++LP LCKL
Sbjct: 566 V-VSSYSPSLLKKFVSLRVLNLSYSKLEQLPSSIGDLLHLRYLDLSCNNFRSLPERLCKL 624
Query: 605 CNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSLNTLSKYIVG 664
NL+ L + C SL LP ++L L++L + C LTS P +IG LT L TL +IVG
Sbjct: 625 QNLQTLDVHNCYSLNCLPKQTSKLSSLRHLVVDGC-PLTSTPPRIGLLTCLKTLGFFIVG 683
Query: 665 EERGFLLEELGQLNLKGQLHIKNLERLKSVTDAKKANMSRK-KLNQLWLSWERNEVSQLQ 723
++G+ L EL LNL G + I +LER+K+ TDA +AN+S K L L +SW+ + ++ +
Sbjct: 684 SKKGYQLGELKNLNLCGSISITHLERVKNDTDA-EANLSAKANLQSLSMSWDNDGPNRYE 742
Query: 724 ENVEQILEALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDCKSCLNLPELW 783
++LEAL+P+ Y + + G FP WI+ L + S+ + CK+CL LP
Sbjct: 743 SKEVKVLEALKPHPNLKY-LEIIAFGGFRFPSWINHSVLEKVISVRIKSCKNCLCLPPFG 801
Query: 784 KLPSLKYLKLSN-MIHVIYLFHESYDG-----EGLMALKTLFLEKLPNLIGLSREE-RVM 836
+LP L+ L+L N V Y+ + +LK L + +L GL +EE
Sbjct: 802 ELPCLENLELQNGSAEVEYVEEDDVHSRFSTRRSFPSLKKLRIWFFRSLKGLMKEEGEEK 861
Query: 837 FPRLKALEITECPNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLHFSDNEELIY 896
FP L+ + I CP L P L S+ L + G N + SSI L +L SL N
Sbjct: 862 FPMLEEMAILYCP-LFVFPTLSSVKKLEVHGNTNTRGLSSISNLSTLTSLRIGANYRATS 920
Query: 897 FPDGILRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCRNIEELPNEVMQRLH 956
P+ + +L + L+ L F LK LPT + ++AL++L I C ++E P + ++ L
Sbjct: 921 LPEEMFTSLTN-LEFLSFFDFKNLKDLPTSLTSLNALKRLQIESCDSLESFPEQGLEGLT 979
Query: 957 SLKELDIVGCDKLK-LSSDFQYLTCLETLAIGSCSEVE 993
SL +L + C LK L Q+LT L L + C EVE
Sbjct: 980 SLTQLFVKYCKMLKCLPEGLQHLTALTNLGVSGCPEVE 1017
Score = 40.8 bits (94), Expect = 0.26
Identities = 39/160 (24%), Positives = 71/160 (44%), Gaps = 16/160 (10%)
Query: 874 PSSIHKLGSLESLHFSDNEELIYFPDGILRNLASPLKTLGFHRHSKL-----KMLPTEMI 928
PS + K SL L+ S ++ L L S + L R+ L + LP +
Sbjct: 572 PSLLKKFVSLRVLNLSYSK---------LEQLPSSIGDLLHLRYLDLSCNNFRSLPERLC 622
Query: 929 HIHALQQLYINDCRNIEELPNEVMQRLHSLKELDIVGCDKLKLSSDFQYLTCLETLAIGS 988
+ LQ L +++C ++ LP + +L SL+ L + GC LTCL+TL
Sbjct: 623 KLQNLQTLDVHNCYSLNCLPKQT-SKLSSLRHLVVDGCPLTSTPPRIGLLTCLKTLGFFI 681
Query: 989 CSEVEGFH-EALQHMTTLKSLTLSDLPNLEYLPECIGNLT 1027
+G+ L+++ S++++ L ++ + NL+
Sbjct: 682 VGSKKGYQLGELKNLNLCGSISITHLERVKNDTDAEANLS 721
Score = 39.7 bits (91), Expect = 0.57
Identities = 26/83 (31%), Positives = 44/83 (52%), Gaps = 3/83 (3%)
Query: 983 TLAIGSCSEVEGFHEAL-QHMTTLKSLTLSDLPNLEYLPECIGNLTLLHEINIYSCPKLA 1041
T++IG + V + +L + +L+ L LS LE LP IG+L L +++ SC
Sbjct: 558 TVSIGFAAVVSSYSPSLLKKFVSLRVLNLS-YSKLEQLPSSIGDLLHLRYLDL-SCNNFR 615
Query: 1042 CLPTSIQQISGLEILSIHDCSKL 1064
LP + ++ L+ L +H+C L
Sbjct: 616 SLPERLCKLQNLQTLDVHNCYSL 638
Score = 37.7 bits (86), Expect = 2.2
Identities = 32/121 (26%), Positives = 52/121 (42%), Gaps = 5/121 (4%)
Query: 941 CRNIEELPNEVMQRLHSLKELDIVGCDKLKLSSDFQYLTCLETLAIGSCSEVEGFHEALQ 1000
C NI E+ + + S+ +V L F L L S S++E ++
Sbjct: 544 CGNIREINVKDYKHTVSIGFAAVVSSYSPSLLKKFVSLRVLNL----SYSKLEQLPSSIG 599
Query: 1001 HMTTLKSLTLSDLPNLEYLPECIGNLTLLHEINIYSCPKLACLPTSIQQISGLEILSIHD 1060
+ L+ L LS N LPE + L L +++++C L CLP ++S L L +
Sbjct: 600 DLLHLRYLDLS-CNNFRSLPERLCKLQNLQTLDVHNCYSLNCLPKQTSKLSSLRHLVVDG 658
Query: 1061 C 1061
C
Sbjct: 659 C 659
>gb|AAR29075.1| blight resistance protein SH20 [Solanum tuberosum]
Length = 947
Score = 677 bits (1748), Expect = 0.0
Identities = 421/1010 (41%), Positives = 593/1010 (58%), Gaps = 85/1010 (8%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAEA ++++L +++ I+ E+ L LGF+ +F ++S +TI+A LEDA+EKQ D I
Sbjct: 1 MAEAFIQVLLENITSFIQGELGLLLGFENDFENISSRFSTIQAVLEDAQEKQLKDKAI-- 58
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSH-KHIAFRYKLAKKMKRIGV 119
K+WL KL A Y +DD++DEC LE S+ G H K I FR+K+ K++K +
Sbjct: 59 --KNWLQKLNAAVYKVDDLLDECKAARLEQ----SRLGCHHPKAIVFRHKIGKRIKEMME 112
Query: 120 WLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVGDASE 179
LD IA E+ FHL E + ER P+ T ++T+P VYGR++++D+IV L+ + S
Sbjct: 113 KLDAIAKERTDFHLHEKIIERQVARPE---TGFVLTEPQVYGRDKEEDEIVKILINNVSN 169
Query: 180 QEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGAT 239
++LSV PI+G+GGLGKTTLAQ+VFN ++ HF KIW+CVS+DF KR+ + II
Sbjct: 170 AQELSVLPILGMGGLGKTTLAQMVFNDQRVTEHFYPKIWICVSDDFDEKRLIENIIGNIE 229
Query: 240 KKSCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGASILVTT 299
+ S + DL Q+KLQ LL KRYLLVLDDVWN+ Q+ W L+ VL G GAS+L TT
Sbjct: 230 RSSLDVKDLASFQKKLQQLLNGKRYLLVLDDVWNEDQQKWDNLRVVLKVGASGASVLTTT 289
Query: 300 RLPKVAKIMGTIPHHELSRLSDEDCWELFKQRAFGPNEVQQKELVIVGKEIIKKCGGFPL 359
RL KV +MGT+ ++LS LS +DCW LF QRAF E LV +GKEI+KK GG PL
Sbjct: 290 RLEKVGSVMGTLQPYQLSNLSQDDCWLLFIQRAFRHQEEISPNLVAIGKEIVKKSGGVPL 349
Query: 360 AAIALGSLLRFKREEKEWLYVKESKLWNL-QGEAYVMPALRLSYLHLPVKLRQCFSFCAL 418
AA LG LLRFKRE++EW +V++S++WNL Q E ++PALRLSY HLP+ LRQCF++CA+
Sbjct: 350 AAKTLGGLLRFKREKREWEHVRDSEIWNLPQDEMSILPALRLSYHHLPLALRQCFAYCAV 409
Query: 419 FPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWNELYWRSFFENTENVGFGQITI 478
FPKD + K+ +I LW A+GF+ S + LE +D+ NE WNELY RSFF+ E V +G T
Sbjct: 410 FPKDTKMEKKKVISLWMAHGFLLSRRNLELEDVRNEGWNELYLRSFFQEIE-VRYGN-TY 467
Query: 479 FKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHLLIYNRNSFAEANSIQLHHVKSLK 538
FKM DL+HDLA S+ S T S R + N S+
Sbjct: 468 FKMXDLIHDLAXSLL----------SANTSSSNIREI---NVESY--------------- 499
Query: 539 TYMEFNFDVYE-AGQLSPQVLNCY-SLRVL-LSH-RLNNLSSSIGRLKYLRYLDISEG-R 593
T+M + E SP +L + SLRVL LS+ + L SSIG L +LRY+D+S
Sbjct: 500 THMMMSIGFSEVVSSYSPSLLQKFVSLRVLNLSYSKFEELPSSIGDLVHLRYMDLSNNIE 559
Query: 594 FKNLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLT 653
++LP LCKL NL+ L L C L LP ++L L+NL L C LT P +IG LT
Sbjct: 560 IRSLPKQLCKLQNLQTLDLQYCTRLCCLPKQTSKLGSLRNLLLHGCHRLTRTPPRIGSLT 619
Query: 654 SLNTLSKYIVGEERGFLLEELGQLNLKGQLHIKNLERLKSVTDAKKANMSRKK-LNQLWL 712
L TL + +V ++G+ L ELG LNL G + I +LER+K+ +AK+AN+S K+ L+ L +
Sbjct: 620 CLKTLGQSVVKRKKGYQLGELGSLNLYGSIKISHLERVKNDKEAKEANLSAKENLHSLSM 679
Query: 713 SWERNEVSQLQENVE-QILEALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELV 771
W+ +E E+ E ++LEAL+P++ L + G+ G P W++ L ++ +E+
Sbjct: 680 KWDDDEHPHRYESEEVEVLEALKPHS-NLTCLKISGFRGIRLPDWMNHSVLKNIVLIEIS 738
Query: 772 DCKSCLNLPELWKLPSLKYLKLSNMIHVIYLFHESYDGEG-------LMALKTLFLEKLP 824
CK+C LP LP L+ L+L Y+ D + L +L+ L + K
Sbjct: 739 GCKNCSCLPPFGDLPCLESLELYRG-SAEYVEEVDIDVDSGFPTRIRLPSLRKLCICKFD 797
Query: 825 NLIGLSREE-RVMFPRLKALEITECPNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSL 883
NL GL ++E FP L+ +EI CP +P+LS L +L
Sbjct: 798 NLKGLLKKEGGEQFPVLEEMEIRYCP-------IPTLSP----------------NLKAL 834
Query: 884 ESLHFSDNEELIYFPDGILRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCRN 943
SL+ SDN+E FP+ + ++LA+ LK L LK LPT + ++AL+ L I C
Sbjct: 835 TSLNISDNKEATSFPEEMFKSLAN-LKYLNISHFKNLKELPTSLASLNALKSLKIQWCCA 893
Query: 944 IEELPNEVMQRLHSLKELDIVGCDKLK-LSSDFQYLTCLETLAIGSCSEV 992
+E +P E ++ L SL EL + LK L +LT L L I C ++
Sbjct: 894 LENIPKEGVKGLTSLTELIVKFSKVLKCLPEGLHHLTALTRLKIWGCPQL 943
Score = 39.7 bits (91), Expect = 0.57
Identities = 23/95 (24%), Positives = 44/95 (46%), Gaps = 8/95 (8%)
Query: 988 SCSEVEGFHEALQHMTTLKSLTLSDLPNLEYLPECIGNLTLLHEINIYSCPKLACLPTSI 1047
S S+ E ++ + L+ + LS+ + LP+ + L L +++ C +L CLP
Sbjct: 532 SYSKFEELPSSIGDLVHLRYMDLSNNIEIRSLPKQLCKLQNLQTLDLQYCTRLCCLPKQT 591
Query: 1048 QQISGLEILSIHDCSKLEK--------RCQKEIGE 1074
++ L L +H C +L + C K +G+
Sbjct: 592 SKLGSLRNLLLHGCHRLTRTPPRIGSLTCLKTLGQ 626
>gb|AAR29074.1| blight resistance protein SH10 [Solanum tuberosum]
Length = 948
Score = 669 bits (1726), Expect = 0.0
Identities = 418/1004 (41%), Positives = 587/1004 (57%), Gaps = 72/1004 (7%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAEA +++++ +L+ ++ E+ L GF EF RL+S+ +TI+A LEDA+EKQ +D +
Sbjct: 1 MAEAFIQVLIDNLTSFLKGELVLLFGFQNEFQRLSSIFSTIQAVLEDAQEKQLND----K 56
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSH-KHIAFRYKLAKKMKRIGV 119
+++WL KL A Y +DDI+DE T+A + S G H K I FR+K+ K+M ++
Sbjct: 57 PLENWLQKLNAATYEVDDILDEYKTKATR--FSQSAYGRYHPKVIPFRHKVGKRMDQVMK 114
Query: 120 WLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVGDASE 179
L+ IA E+ FHL E + ER V R+T S++T+P VYGR++++D+IV L+ + S+
Sbjct: 115 KLNAIAEERKNFHLHEKIIERQAVR---RETGSVLTEPQVYGRDKEEDEIVKILINNVSD 171
Query: 180 QEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGAT 239
+ LSV PI+G+GGLGKTTLAQ+VFN +I HF KIW+CVSEDF KR+ KAIIE
Sbjct: 172 AQHLSVLPILGMGGLGKTTLAQMVFNDQRITEHFHSKIWICVSEDFDEKRLLKAIIESIE 231
Query: 240 KKSC-EDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGASILVT 298
+ ++DL LQ+KLQ+LL KRY LVLDDVWN+ Q+ W L++VL G GA +L T
Sbjct: 232 GRPLLGEMDLAPLQKKLQELLNGKRYFLVLDDVWNEDQQKWANLRAVLKVGASGAFVLAT 291
Query: 299 TRLPKVAKIMGTIPHHELSRLSDEDCWELFKQRAFGPNEVQQKELVIVGKEIIKKCGGFP 358
TRL KV IMGT+ +ELS LS EDCW LF Q AFG E LV +GKEI+KK GG P
Sbjct: 292 TRLEKVGSIMGTLQPYELSNLSQEDCWLLFIQCAFGHQEEINPNLVAIGKEIVKKSGGVP 351
Query: 359 LAAIALGSLLRFKREEKEWLYVKESKLWNL-QGEAYVMPALRLSYLHLPVKLRQCFSFCA 417
LAA LG +LRFKREE+EW +V++S++WNL Q E ++PALRLSY HLP+ LRQCF++CA
Sbjct: 352 LAAKTLGGILRFKREEREWEHVRDSEIWNLPQEERSILPALRLSYHHLPLDLRQCFAYCA 411
Query: 418 LFPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWNELYWRSFFENTENVGFGQIT 477
+FPKD + K+ LI LW A+GF+ L+ +D+GNEV EL RSFF+ E G+ T
Sbjct: 412 VFPKDTKMEKEKLISLWMAHGFLLLEGKLQPEDVGNEVSKELCLRSFFQEIE-AKCGK-T 469
Query: 478 IFKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHLLIYNRNSFAEANSIQLHHVKSL 537
FKMHDL HDLA S+ S T S R + +VK
Sbjct: 470 YFKMHDLHHDLATSLF----------SASTSSSNIREI-----------------NVKGY 502
Query: 538 KTYMEFNFDVYEAGQLSPQVLNCY-SLRVLLSHRLN--NLSSSIGRLKYLRYLDISEGR- 593
M SP + + SLRVL L+ LSSSIG L ++R LD+SE
Sbjct: 503 PHKMMSIGFTEVVSSYSPSLSQKFVSLRVLNLSNLHFEELSSSIGDLVHMRCLDLSENSG 562
Query: 594 FKNLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLT 653
++LP LCKL NL+ L L C SL LP ++L L+NL CD L S+P +IG LT
Sbjct: 563 IRSLPKQLCKLQNLQTLDLHNCYSLSCLPKEPSKLGSLRNLFFHGCDELNSMPPRIGSLT 622
Query: 654 SLNTLSKYIVG-EERGFLLEELGQLNLKGQLHIKNLERLKSVTDAKKANMSRK-KLNQLW 711
L TL G +++G+ L +L +NL G + I +LER+K+V DAK+AN+S K L+ L
Sbjct: 623 FLKTLKWICCGIQKKGYQLGKLRDVNLYGSIEITHLERVKNVMDAKEANLSAKGNLHSLI 682
Query: 712 LSWERNEVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELV 771
++W R + +++EAL+P+ L + G+ G FP+W++ L ++ S+E+
Sbjct: 683 MNWSRKGPHIYESEEVRVIEALKPH-PNLTCLTISGFRGFRFPEWMNHSVLKNVVSIEIS 741
Query: 772 DCKSCLNLPELWKLPSLKYLKL-SNMIHVIYLFHESYDGEGLMALKTLFLEKLPNLIGLS 830
CK+C LP +LP LK L+L V Y+ +L+ LF+ + PNL GL
Sbjct: 742 GCKNCSCLPPFGELPCLKRLELQKGSAEVEYVDSGFPTRRRFPSLRKLFIGEFPNLKGLL 801
Query: 831 REE-RVMFPRLKALEITECPNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLHFS 889
++E FP L+ + I C ++ L S +L SLH S
Sbjct: 802 KKEGEEKFPVLERMTIFYC-HMFVYTTLSS-------------------NFRALTSLHIS 841
Query: 890 DNEELIYFPDGILRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCRNIEELPN 949
N E P+ I ++ A+ LK L LK LP+ + ++AL+ L I+ C +E LP
Sbjct: 842 HNNEATSLPEEIFKSFAN-LKYLKISLFYNLKELPSSLACLNALKTLEIHSCSALESLPE 900
Query: 950 EVMQRLHSLKELDIVGCDKLK-LSSDFQYLTCLETLAIGSCSEV 992
E ++ L SL EL + C+ LK L Q+LT L +L + C ++
Sbjct: 901 EGVKGLTSLTELFVYDCEMLKFLPEGLQHLTALTSLKLRRCPQL 944
Score = 40.4 bits (93), Expect = 0.34
Identities = 29/122 (23%), Positives = 55/122 (44%), Gaps = 8/122 (6%)
Query: 943 NIEELPNEVMQRLHSLKELDIVGCDKLKLSSDFQYLTCLETLAIGSCSEVEGFHEALQHM 1002
N++ P+++M S+ ++V LS F L L S E ++ +
Sbjct: 498 NVKGYPHKMM----SIGFTEVVSSYSPSLSQKFVSLRVLNL----SNLHFEELSSSIGDL 549
Query: 1003 TTLKSLTLSDLPNLEYLPECIGNLTLLHEINIYSCPKLACLPTSIQQISGLEILSIHDCS 1062
++ L LS+ + LP+ + L L +++++C L+CLP ++ L L H C
Sbjct: 550 VHMRCLDLSENSGIRSLPKQLCKLQNLQTLDLHNCYSLSCLPKEPSKLGSLRNLFFHGCD 609
Query: 1063 KL 1064
+L
Sbjct: 610 EL 611
>gb|AAP45181.1| putative disease resistant protein rga3 [Solanum bulbocastanum]
Length = 908
Score = 659 bits (1699), Expect = 0.0
Identities = 404/959 (42%), Positives = 568/959 (59%), Gaps = 77/959 (8%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAEA L+++L +L+ I+ E+ L GF++EF +L+S+ + I+A LEDA+EKQ I
Sbjct: 1 MAEAFLQVLLDNLTFFIQGELGLVFGFEKEFKKLSSMFSMIQAVLEDAQEKQLKYKAI-- 58
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSH-KHIAFRYKLAKKMKRIGV 119
K+WL KL AAY +DDI+D+C TEA +K + G H + I F YK+ K+MK +
Sbjct: 59 --KNWLQKLNVAAYEVDDILDDCKTEAAR--FKQAVLGRYHPRTITFCYKVGKRMKEMME 114
Query: 120 WLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVGDASE 179
LD IA E+ FHL E + ER RQT ++T+P VYG+ +++D+IV L+ + S
Sbjct: 115 KLDAIAEERRNFHLDERIIERQAAR---RQTGFVLTEPKVYGKEKEEDEIVKILINNVSY 171
Query: 180 QEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGAT 239
+++ V PI+G+GGLGKTTLAQ+VFN +I HF LKIWVCVS+DF KR+ KAI+E
Sbjct: 172 SKEVPVLPILGMGGLGKTTLAQMVFNDQRITEHFNLKIWVCVSDDFDEKRLIKAIVESIE 231
Query: 240 KKSCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGASILVTT 299
KS D+DL LQ+KLQ+LL KRY LVLDDVWN+ QE W L++VL G GASIL+TT
Sbjct: 232 GKSLGDMDLAPLQKKLQELLNGKRYFLVLDDVWNEDQEKWDNLRAVLKIGASGASILITT 291
Query: 300 RLPKVAKIMGTIPHHELSRLSDEDCWELFKQRAFGPNEVQQKELVIVGKEIIKKCGGFPL 359
RL K+ IMGT+ ++LS LS EDCW LFKQRAF +L+ +GKEI+KKCGG PL
Sbjct: 292 RLEKIGSIMGTLQLYQLSNLSQEDCWLLFKQRAFCHQTETSPKLMEIGKEIVKKCGGVPL 351
Query: 360 AAIALGSLLRFKREEKEWLYVKESKLWNL-QGEAYVMPALRLSYLHLPVKLRQCFSFCAL 418
AA LG LLRFKREE EW +V++S++WNL Q E V+PALRLSY HLP+ LRQCF++CA+
Sbjct: 352 AAKTLGGLLRFKREESEWEHVRDSEIWNLPQDENSVLPALRLSYHHLPLDLRQCFAYCAV 411
Query: 419 FPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWNELYWRSFFENTENVGFGQITI 478
FPKD I K+ LI LW A+ F+ S +E +D+GNEVWNELY RSFF+ E V G+ T
Sbjct: 412 FPKDTKIEKEYLIALWMAHSFLLSKGNMELEDVGNEVWNELYLRSFFQEIE-VKSGK-TY 469
Query: 479 FKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHLLIYNRNSFAEANSIQLHHVKSLK 538
FKMHDL+HDLA SM + S +R + R + + + V + K
Sbjct: 470 FKMHDLIHDLA-------------TSMFSASASSRSI----RQINVKDDEDMMFIVTNYK 512
Query: 539 TYMEFNFDVYEAGQLSPQVLNCYSLRVLLSHRLNNLSSSIGRLKYLRYLDISEGRFKNLP 598
M F +V++ Y S FK+LP
Sbjct: 513 DMMSIGFS---------EVVSSY----------------------------SPSLFKSLP 535
Query: 599 NSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSLNTL 658
LCKL NL+ L L C SL LP ++L L+NL L C LTS+P +IG LT L TL
Sbjct: 536 KRLCKLQNLQTLDLYNCQSLSCLPKQTSKLCSLRNLVLDHC-PLTSMPPRIGLLTCLKTL 594
Query: 659 SKYIVGEERGFLLEELGQLNLKGQLHIKNLERLKSVTDAKKANMSRK-KLNQLWLSWERN 717
++VGE +G+ L EL LNL+G + I +LER+K+ +AK+AN+S K L+ L +SW+R
Sbjct: 595 GYFVVGERKGYQLGELRNLNLRGAISITHLERVKNDMEAKEANLSAKANLHSLSMSWDRP 654
Query: 718 EVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDCKSCL 777
+ +E ++LEAL+P+ Y + + G P W++ L ++ S+ + C++C
Sbjct: 655 NRYESEE--VKVLEALKPHPNLKY-LEIIDFCGFCLPDWMNHSVLKNVVSILISGCENCS 711
Query: 778 NLPELWKLPSLKYLKLSN-MIHVIYLFHESY-DGEGLMALKTLFLEKLPNLIGLSR-EER 834
LP +LP L+ L+L + + V ++ + +L+ L + NL GL R E
Sbjct: 712 CLPPFGELPCLESLELQDGSVEVEFVEDSGFPTRRRFPSLRKLHIGGFCNLKGLQRMEGE 771
Query: 835 VMFPRLKALEITECPNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLHFSDNEEL 894
FP L+ ++I++CP + P L S+ L I G+ + + SSI L +L SL N +
Sbjct: 772 EQFPVLEEMKISDCP-MFVFPTLSSVKKLEIWGEADARGLSSISNLSTLTSLKIFSNHTV 830
Query: 895 IYFPDGILRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCRNIEELPNEVMQ 953
+ + ++L + LK L LK LPT + ++ L+ L I C +E LP E ++
Sbjct: 831 TSLLEEMFKSLEN-LKYLSVSYLENLKELPTSLASLNNLKCLDIRYCYALESLPEEGLE 888
>gb|AAP45185.1| putative disease resistant protein rga1 [Solanum bulbocastanum]
Length = 923
Score = 658 bits (1697), Expect = 0.0
Identities = 420/999 (42%), Positives = 574/999 (57%), Gaps = 114/999 (11%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAEA +++VL +L+ ++ E+ L GF EF RL+S+ +TI+A LEDA+EKQ +D +
Sbjct: 1 MAEAFIQVVLDNLTSFLKGELVLLFGFQDEFQRLSSMFSTIQAVLEDAQEKQLND----K 56
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSH-KHIAFRYKLAKKMKRIGV 119
+++WL KL A Y +DDI+DE T+A + S+ G H K I FR+K+ K+M ++
Sbjct: 57 PLENWLQKLNAATYEVDDILDEYKTKATR--FLLSEYGRYHPKVIPFRHKVGKRMDQVMK 114
Query: 120 WLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVGDASE 179
L+ IA E+ FHL E + ER R+T S++T+ VYGR+++KD+IV L AS+
Sbjct: 115 KLNAIAEERKNFHLQEKIIERQAAT---RETGSVLTESQVYGRDKEKDEIVKILTNTASD 171
Query: 180 QEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGAT 239
+ LSV PI+G+GGLGKTTL+Q+VFN ++ F KIW+CVS+DF KR+ KAI+E
Sbjct: 172 AQKLSVLPILGMGGLGKTTLSQMVFNDQRVTERFYPKIWICVSDDFNEKRLIKAIVESIE 231
Query: 240 KKSCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGASILVTT 299
KS D+DL LQ+KLQ+LL KRY LVLDDVWN+ Q W L++VL G GA +L TT
Sbjct: 232 GKSLSDMDLAPLQKKLQELLNGKRYFLVLDDVWNEDQHKWANLRAVLKVGASGAFVLTTT 291
Query: 300 RLPKVAKIMGTIPHHELSRLSDEDCWELFKQRAFGPNEVQQKELVIVGKEIIKKCGGFPL 359
RL KV IMGT+ +ELS LS EDCW LF QRAFG E LV +GKEI+KKCGG PL
Sbjct: 292 RLEKVGSIMGTLQPYELSNLSPEDCWFLFMQRAFGHQEEINPNLVAIGKEIVKKCGGVPL 351
Query: 360 AAIALGSLLRFKREEKEWLYVKESKLWNL-QGEAYVMPALRLSYLHLPVKLRQCFSFCAL 418
AA LG +LRFKREE+EW +V++S +WNL Q E+ ++PALRLSY HLP+ LRQCF +CA+
Sbjct: 352 AAKTLGGILRFKREEREWEHVRDSPIWNLPQDESSILPALRLSYHHLPLDLRQCFVYCAV 411
Query: 419 FPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWNELYWRSFFENTENVGFGQITI 478
FPKD ++K+ LI W A+GF+ S LE +D+GNEVWNELY RSFF+ E V G+ T
Sbjct: 412 FPKDTKMAKENLIAFWMAHGFLLSKGNLELEDVGNEVWNELYLRSFFQEIE-VESGK-TY 469
Query: 479 FKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHLLIYNRNSFAEANSIQLHHVKSLK 538
FKMHDL+HDLA S+ S T S R + N N SI V S
Sbjct: 470 FKMHDLIHDLATSLF----------SANTSSSNIREI---NANYDGYMMSIGFAEVVS-- 514
Query: 539 TYMEFNFDVYEAGQLSPQVLNCY-SLRV--LLSHRLNNLSSSIGRLKYLRYLDISEG-RF 594
SP +L + SLRV L + LN L SSIG L +LRYLD+S R
Sbjct: 515 -------------SYSPSLLQKFVSLRVLNLRNSNLNQLPSSIGDLVHLRYLDLSGNVRI 561
Query: 595 KNLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTS 654
++LP LCKL NL Q L L CDSL+ LP+Q
Sbjct: 562 RSLPRRLCKLQNL------------------------QTLDLHYCDSLSCLPKQ------ 591
Query: 655 LNTLSKYI-VGEERGFLLEELGQLNLKGQLHIKNLERLKSVTDAKKANMSRK-KLNQLWL 712
T+SK + +E+G+ L EL LNL G + I L+R+K TDAK+AN+S K L+ L L
Sbjct: 592 --TISKLLCYWQEKGYQLGELKNLNLYGSISITKLDRVKKDTDAKEANLSAKANLHSLCL 649
Query: 713 SWERNEVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELVD 772
SW+ + + ++LEAL+P++ Y + G+ G P W++ L ++ S+ +
Sbjct: 650 SWDLDGKHRYD---SEVLEALKPHSNLKY-LEINGFGGILLPDWMNQSVLKNVVSIRIRG 705
Query: 773 CKSCLNLPELWKLPSLKYLKL-SNMIHVIYLFHESYDGEGLMALKTLFLEKLPNLIGLSR 831
C++C LP +LP L+ L+L + V Y+ E +GE
Sbjct: 706 CENCSCLPPFGELPCLESLELHTGSAEVEYV--EDNEGE--------------------- 742
Query: 832 EERVMFPRLKALEITECPNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLHFSDN 891
FP L+ + CP + +P L S+ L + + + SI L +L SL S+N
Sbjct: 743 ---KQFPVLEEMTFYWCP-MFVIPTLSSVKTLKVIAT-DATVLRSISNLRALTSLDISNN 797
Query: 892 EELIYFPDGILRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCRNIEELPNEV 951
E P+ + ++LA+ LK L LK LPT + ++AL+ L C +E LP E
Sbjct: 798 VEATSLPEEMFKSLAN-LKYLNISFFRNLKELPTSLASLNALKSLKFEFCDALESLPEEG 856
Query: 952 MQRLHSLKELDIVGCDKLK-LSSDFQYLTCLETLAIGSC 989
++ L SL EL + C LK L Q+LT L TL I C
Sbjct: 857 VKGLTSLTELSVSNCMMLKCLPEGLQHLTALTTLTITQC 895
Score = 85.9 bits (211), Expect = 7e-15
Identities = 104/356 (29%), Positives = 153/356 (42%), Gaps = 71/356 (19%)
Query: 762 LNDLKSLELVDCKSCLNLPELWKLPSLKYLKLSNM--IHVIYLFHE-----SYDGEGLMA 814
L +LK+L L S L + K K LS +H + L + YD E L A
Sbjct: 607 LGELKNLNLYGSISITKLDRVKKDTDAKEANLSAKANLHSLCLSWDLDGKHRYDSEVLEA 666
Query: 815 LKT----LFLE-------KLPNLIGLSREERVMFPRLKALEITEC-PNLLGLPCLPSLSD 862
LK +LE LP+ + S + V+ R++ E C P LPCL SL
Sbjct: 667 LKPHSNLKYLEINGFGGILLPDWMNQSVLKNVVSIRIRGCENCSCLPPFGELPCLESLE- 725
Query: 863 LYIQGKYNQQLPSSIHKLGSLESLHFSDNEELIYFPDGILRNLA---SPLKTLGFHRHSK 919
+H GS E + DNE FP +L + P+ + K
Sbjct: 726 --------------LHT-GSAEVEYVEDNEGEKQFP--VLEEMTFYWCPMFVIPTLSSVK 768
Query: 920 -LKMLPTE------MIHIHALQQLYINDCRNIEELPNEVMQRLHSLKELDIVGCDKLKLS 972
LK++ T+ + ++ AL L I++ LP E+ + L +LK L+I
Sbjct: 769 TLKVIATDATVLRSISNLRALTSLDISNNVEATSLPEEMFKSLANLKYLNI--------- 819
Query: 973 SDFQYLTCLETLAIGSCSEVEGFHEALQHMTTLKSLTLSDLPNLEYLPE-CIGNLTLLHE 1031
S F+ L L T +L + LKSL LE LPE + LT L E
Sbjct: 820 SFFRNLKELPT--------------SLASLNALKSLKFEFCDALESLPEEGVKGLTSLTE 865
Query: 1032 INIYSCPKLACLPTSIQQISGLEILSIHDCSKLEKRCQKEIGEDWPKIVHVQYIEI 1087
+++ +C L CLP +Q ++ L L+I C + KRC++ IGEDW KI H+ Y+ +
Sbjct: 866 LSVSNCMMLKCLPEGLQHLTALTTLTITQCPIVFKRCERGIGEDWHKISHIPYLTL 921
Score = 38.9 bits (89), Expect = 0.97
Identities = 58/225 (25%), Positives = 91/225 (39%), Gaps = 17/225 (7%)
Query: 859 SLSDLYIQGKYNQQLPSSIHKLGSLESLHFSDNEELIYFPDGILRNLASPLKTLGFHRHS 918
SL L ++ QLPSSI L L L S N + P + + L+TL H
Sbjct: 526 SLRVLNLRNSNLNQLPSSIGDLVHLRYLDLSGNVRIRSLPRRLCK--LQNLQTLDLHYCD 583
Query: 919 KLKMLPTEMIHIHALQQLYINDCRNIEELPNEVMQRLHSLKELDIVGCDKLKLSSDFQYL 978
L LP + I L + + EL N + S+ +LD V D ++
Sbjct: 584 SLSCLPKQT--ISKLLCYWQEKGYQLGELKNLNLYGSISITKLDRVKKDTDAKEANLSAK 641
Query: 979 TCLETLAIGSCSEVEGFH-------EALQHMTTLKSLTLSDLPNLEYLPECIGNLTLLH- 1030
L +L + +++G H EAL+ + LK L ++ + LP+ + L +
Sbjct: 642 ANLHSLCLS--WDLDGKHRYDSEVLEALKPHSNLKYLEINGFGGI-LLPDWMNQSVLKNV 698
Query: 1031 -EINIYSCPKLACLPTSIQQISGLEILSIHDCSKLEKRCQKEIGE 1074
I I C +CLP ++ LE L +H S + + GE
Sbjct: 699 VSIRIRGCENCSCLP-PFGELPCLESLELHTGSAEVEYVEDNEGE 742
Score = 37.0 bits (84), Expect = 3.7
Identities = 23/82 (28%), Positives = 41/82 (49%), Gaps = 2/82 (2%)
Query: 984 LAIGSCSEVEGFHEAL-QHMTTLKSLTLSDLPNLEYLPECIGNLTLLHEINIYSCPKLAC 1042
++IG V + +L Q +L+ L L + NL LP IG+L L +++ ++
Sbjct: 505 MSIGFAEVVSSYSPSLLQKFVSLRVLNLRN-SNLNQLPSSIGDLVHLRYLDLSGNVRIRS 563
Query: 1043 LPTSIQQISGLEILSIHDCSKL 1064
LP + ++ L+ L +H C L
Sbjct: 564 LPRRLCKLQNLQTLDLHYCDSL 585
>gb|AAP45164.1| putative disease resistant protein RGA2 [Solanum bulbocastanum]
Length = 819
Score = 630 bits (1625), Expect = e-179
Identities = 372/860 (43%), Positives = 525/860 (60%), Gaps = 62/860 (7%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAEA ++++L +L+ ++ E+ L GF EF RL+S+ +TI+A LEDA+EKQ ++ +
Sbjct: 1 MAEAFIQVLLDNLTSFLKGELVLLFGFQDEFQRLSSMFSTIQAVLEDAQEKQLNN----K 56
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSH-KHIAFRYKLAKKMKRIGV 119
+++WL KL A Y +DDI+DE T+A + S+ G H K I FR+K+ K+M ++
Sbjct: 57 PLENWLQKLNAATYEVDDILDEYKTKATR--FSQSEYGRYHPKVIPFRHKVGKRMDQVMK 114
Query: 120 WLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVGDASE 179
L IA E+ FHL E + ER V R+T S++T+P VYGR+++KD+IV L+ + S+
Sbjct: 115 KLKAIAEERKNFHLHEKIVERQAVR---RETGSVLTEPQVYGRDKEKDEIVKILINNVSD 171
Query: 180 QEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGAT 239
+ LSV PI+G+GGLGKTTLAQ+VFN ++ HF KIW+CVSEDF KR+ KAI+E
Sbjct: 172 AQHLSVLPILGMGGLGKTTLAQMVFNDQRVTEHFHSKIWICVSEDFDEKRLIKAIVESIE 231
Query: 240 KKSC-EDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGASILVT 298
+ ++DL LQ+KLQ+LL KRYLLVLDDVWN+ Q+ W L++VL G GAS+L T
Sbjct: 232 GRPLLGEMDLAPLQKKLQELLNGKRYLLVLDDVWNEDQQKWANLRAVLKVGASGASVLTT 291
Query: 299 TRLPKVAKIMGTIPHHELSRLSDEDCWELFKQRAFGPNEVQQKELVIVGKEIIKKCGGFP 358
TRL KV IMGT+ +ELS LS EDCW LF QRAFG E LV +GKEI+KK GG P
Sbjct: 292 TRLEKVGSIMGTLQPYELSNLSQEDCWLLFMQRAFGHQEEINPNLVAIGKEIVKKSGGVP 351
Query: 359 LAAIALGSLLRFKREEKEWLYVKESKLWNL-QGEAYVMPALRLSYLHLPVKLRQCFSFCA 417
LAA LG +L FKREE+ W +V++S +WNL Q E+ ++PALRLSY LP+ L+QCF++CA
Sbjct: 352 LAAKTLGGILCFKREERAWEHVRDSPIWNLPQDESSILPALRLSYHQLPLDLKQCFAYCA 411
Query: 418 LFPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWNELYWRSFFENTENVGFGQIT 477
+FPKD + K+ LI LW A+GF+ S +E +D+G+EVW ELY RSFF+ E V G+ T
Sbjct: 412 VFPKDAKMEKEKLISLWMAHGFLLSKGNMELEDVGDEVWKELYLRSFFQEIE-VKDGK-T 469
Query: 478 IFKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHLLIYNRNSFAEANSIQLHHVKSL 537
FKMHDL+HDLA S+ S T S R + N++S+ SI V
Sbjct: 470 YFKMHDLIHDLATSLF----------SANTSSSNIREI---NKHSYTHMMSIGFAEVVFF 516
Query: 538 KTYMEFNFDVYEAGQLSPQVLNCYSLRVLL--SHRLNNLSSSIGRLKYLRYLDISEGRFK 595
T P + SLRVL N L SSIG L +LRYL++ +
Sbjct: 517 YTL--------------PPLEKFISLRVLNLGDSTFNKLPSSIGDLVHLRYLNLYGSGMR 562
Query: 596 NLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSL 655
+LP LCKL NL+ L L C L LP ++L L+NL L SLT +P +IG LT L
Sbjct: 563 SLPKQLCKLQNLQTLDLQYCTKLCCLPKETSKLGSLRNLLLDGSQSLTCMPPRIGSLTCL 622
Query: 656 NTLSKYIVGEERGFLLEELGQLNLKGQLHIKNLERLKSVTDAKKANMSRK-KLNQLWLSW 714
TL +++VG ++G+ L ELG LNL G + I +LER+K+ DAK+AN+S K L+ L +SW
Sbjct: 623 KTLGQFVVGRKKGYQLGELGNLNLYGSIKISHLERVKNDKDAKEANLSAKGNLHSLSMSW 682
Query: 715 ERNEVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDCK 774
+ ++LEAL+P++ L S + G+ G + P+W++ L ++ S+ + + +
Sbjct: 683 NNFGPHIYESEEVKVLEALKPHS-NLTSLKIYGFRGIHLPEWMNHSVLKNIVSILISNFR 741
Query: 775 SCLNLPELWKLPSLKYLKLSNMIHVIYLFHESYDGEGLMALKTLFLEKLPNLIGLSREER 834
+C LP LP L+ L+ L S D E ++E++ + R
Sbjct: 742 NCSCLPPFGDLPCLESLE---------LHWGSADVE--------YVEEVDIDVHSGFPTR 784
Query: 835 VMFPRLKALEITECPNLLGL 854
+ FP L+ L+I + +L GL
Sbjct: 785 IRFPSLRKLDIWDFGSLKGL 804
>gb|AAN03742.1| NBS-LRR-like protein [Oryza sativa (japonica cultivar-group)]
Length = 1108
Score = 548 bits (1413), Expect = e-154
Identities = 389/1130 (34%), Positives = 594/1130 (52%), Gaps = 75/1130 (6%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQ----EFNRLASLLTTIKATLEDAEEKQFSDS 56
+ EAVL + +L E S L F Q E L+S L+TI A +EDAEE+Q D
Sbjct: 3 IGEAVLSAFMQALFEKAVAAASSELKFPQNIAVELQNLSSSLSTILAHVEDAEERQLKDQ 62
Query: 57 EIGRDVKDWLLKLKDAAYTLDDIMDECATEALE-----------MEYKASKCGLSHKHIA 105
+ WL +LKD AY +DD++DE A E L ++ + C + K+
Sbjct: 63 A----ARSWLSRLKDVAYEMDDLLDEHAAEVLRSKLAGPSNYHHLKVRICFCCIWLKNGL 118
Query: 106 FRYKLAKKMKRIGVWLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNED 165
F L K++ RI +D + K++ + I+R + + +T+S++ VYGR ED
Sbjct: 119 FNRDLVKQIMRIEGKIDRLI--KDRHIVDPIMRFNREEIRERPKTSSLIDDSSVYGREED 176
Query: 166 KDKIVDFLVG-DASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSED 224
K+ IV+ L+ + S +LS+ PIVG+GG+GKTTL QLV+N ++ HF+L++W+CVSE+
Sbjct: 177 KEVIVNMLLTTNNSNHVNLSILPIVGMGGVGKTTLTQLVYNDVRVKKHFQLRMWLCVSEN 236
Query: 225 FTLKRMTKAIIEG-ATKKSCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLK 283
F ++TK IE A+ S ++ LLQ L + L+ KR+LLVLDDVWN+ + W R +
Sbjct: 237 FDEAKLTKETIESVASGLSSATTNMNLLQEDLSNKLKGKRFLLVLDDVWNEDPDRWDRYR 296
Query: 284 SVLACGGKGASILVTTRLPKVAKIMGTIPHHELSRLSDEDCWELFKQRAFGPNEVQ-QKE 342
L G KG+ I+VTTR V K++G + + L +LS DCW LF+ AF +
Sbjct: 297 CALVAGAKGSKIMVTTRNENVGKLVGGLTPYYLKQLSYNDCWHLFRSYAFADGDSSAHPN 356
Query: 343 LVIVGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWNLQGEA-YVMPALRLS 401
L ++GKEI+ K G PLAA ALGSLL K E +W + ES++W L + ++PALRLS
Sbjct: 357 LEMIGKEIVHKLKGLPLAARALGSLLCAKDNEDDWKNILESEIWELPSDKNNILPALRLS 416
Query: 402 YLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWNELYW 461
Y HLP L++CF+FC++F KD + K +L+ +W A G+I ++IGN ++EL
Sbjct: 417 YNHLPPILKRCFAFCSVFHKDYVFEKDILVQIWMAVGYIQPQGRRRMEEIGNNYFDELLS 476
Query: 462 RSFFENTENVGFGQITIFKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHLLIYNRN 521
RSFF+ ++ + MHD +HDLA SV+ D C D+ + +E L ++ +
Sbjct: 477 RSFFQKHKDG-------YVMHDAMHDLAQSVSIDECMRLDNLPNNSTTERNARHLSFSCD 529
Query: 522 SFAEANSIQLHHVKSLKTYMEFNFDVYEAGQL-SPQVLNCYSLRVLLSHR--LNNLSSSI 578
+ ++ ++ + N + + S LN L VL +R + L S+
Sbjct: 530 NKSQTTFEAFRGFNRARSLLLLNGYKSKTSSIPSDLFLNLRYLHVLDLNRQEITELPESV 589
Query: 579 GRLKYLRYLDISEGRFKNLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRD 638
G+LK LRYL++S + LP+S+ KL L+ LKL C L L +L R
Sbjct: 590 GKLKMLRYLNLSGTVVRKLPSSIGKLYCLQTLKLRNCSH---------NLVNLLSLEAR- 639
Query: 639 CDSLTSLPRQIGKLTSLNTLSKYIVGEERGFLLEELGQLN-LKGQLHIKNLERLKSVTDA 697
+ +T + R IGKLT L L +++V +++G+ + EL +N + G + IKNLE + S +A
Sbjct: 640 TELITGIAR-IGKLTCLQKLEEFVVHKDKGYKVSELKAMNKIGGHICIKNLESVSSAEEA 698
Query: 698 KKANMSRKK----LNQLWLSWERNEVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAYF 753
+A +S K L+ +W S R+ S+ + L +L+P+ +L V + G F
Sbjct: 699 DEALLSEKAHISILDLIWSS-SRDFTSEEANQDIETLTSLEPH-DELKELTVKAFAGFEF 756
Query: 754 PQWISIPSLNDLKSLELVDCKSCLNLPELWKLPSLKYLKLSNMIHVIYL---FHESYDGE 810
P WI L+ L+++ L DC +C LP L +LP LK + + +I + F S + +
Sbjct: 757 PHWI----LSHLQTIHLSDCTNCSILPALGQLPLLKVIIIGGFPTIIKIGDEFSGSSEVK 812
Query: 811 GLMALKTLFLEKLPNL-IGLSREERVMFPRLKALEITECPNLLGLPCLPS-LSDLYIQGK 868
G +LK L E PNL S ++ P L+ L++ +CP + LP LPS L +L I
Sbjct: 813 GFPSLKELVFEDTPNLERWTSTQDGEFLPFLRELQVLDCPKVTELPLLPSTLVELKISEA 872
Query: 869 YNQQLPSSIHK---LGSLESLHFSDNEELIYFPDGILRNLASPLKTLGFHRHSKLKMLPT 925
LP +H L SL L L G+L S L+ L +L PT
Sbjct: 873 GFSVLP-EVHAPRFLPSLTRLQIHKCPNLTSLQQGLLSQQLSALQQLTITNCPELIHPPT 931
Query: 926 EMIH-IHALQQLYINDCRNIEELPNE-VMQRLHSLKELDIVGCDKL--KLSSDFQYLTCL 981
E + + ALQ L+I DC + + ++ R+ +++L I C + L + L L
Sbjct: 932 EGLRTLTALQSLHIYDCPRLATAEHRGLLPRM--IEDLRITSCSNIINPLLDELNELFAL 989
Query: 982 ETLAIGSCSEVEGFHEALQHMTTLKSLTLSDLPNLEYLPECIGNLTLLHEINIYSCPKLA 1041
+ L I C + F E L TLK L + + NL LP C+ + L + I +C +
Sbjct: 990 KNLVIADCVSLNTFPEKLP--ATLKKLEIFNCSNLASLPACLQEASCLKTMTILNCVSIK 1047
Query: 1042 CLPTSIQQISGLEILSIHDCSKLEKRCQKEIGEDWPKIVHVQYIEIENDN 1091
CLP +S LE L I +C L +RCQ+ GEDWPKI H+ IEI++D+
Sbjct: 1048 CLPAHGLPLS-LEELYIKECPFLAERCQENSGEDWPKISHIAIIEIDDDS 1096
>ref|XP_483605.1| putative NBS-LRR resistance protein RGH1 [Oryza sativa (japonica
cultivar-group)] gi|42407847|dbj|BAD08990.1| putative
NBS-LRR resistance protein RGH1 [Oryza sativa (japonica
cultivar-group)] gi|42408544|dbj|BAD09722.1| putative
NBS-LRR resistance protein RGH1 [Oryza sativa (japonica
cultivar-group)]
Length = 1124
Score = 548 bits (1411), Expect = e-154
Identities = 383/1135 (33%), Positives = 601/1135 (52%), Gaps = 99/1135 (8%)
Query: 5 VLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGRDVKD 64
V+ V G ++ + + ++ G D + +L L ++ L DAE K SE VK
Sbjct: 9 VVRGVAGKAADALVQSVTRMCGIDGDRRKLERQLLAVQCKLADAEAK----SETNPAVKR 64
Query: 65 WLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLS--------HKHIAFRYKLAKKMKR 116
W+ LK AY DD++D+ EAL E K H + FR +++K+
Sbjct: 65 WMKDLKAVAYEADDVLDDFEYEALRREVKIGDSTTRKVLGFFTPHSPLLFRVTMSRKLGD 124
Query: 117 IGVWLDDIAAEKNKFHLTEIVRERSGVVPD--WRQTTSIVTQPL-VYGRNEDKDKIVDFL 173
+ ++++ E NKF L E V VP +R T S + + ++GR DK+ +V
Sbjct: 125 VLKKINELVEEMNKFGLMEHVE-----VPQLPYRLTHSGLDESADIFGREHDKEVLVKLT 179
Query: 174 VGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKA 233
+ D +Q++L V PIVG+GGLGKTTLA+L++N + HF+LK+W CVSE+F + + K+
Sbjct: 180 L-DQHDQQNLQVLPIVGMGGLGKTTLAKLIYNDPSVQEHFQLKMWHCVSENFEVGSLLKS 238
Query: 234 IIEGATKKSCEDLD-LELLQRKLQDLLRRKRYLLVLDDVWNDKQENW-QRLKSVL-ACGG 290
I+E AT + C+ ++ +ELL+R+L++ R+R+LLVLDDVWND++ W LK +L + GG
Sbjct: 239 IVELATNRRCQLINTIELLRRQLEEAFGRRRFLLVLDDVWNDEENKWADDLKPLLNSVGG 298
Query: 291 KGASILVTTRLPKVAKIMGTIPHHELSRLSDEDCWELFKQRAFGPNEVQQKELVIVGKEI 350
G+ I+VTTR +VA IMGT+ +EL L+++D WE+F +RAFG +Q +LV +G I
Sbjct: 299 AGSVIVVTTRSQRVASIMGTLEPYELRCLNEDDSWEVFSKRAFGKQVQEQAKLVSIGTRI 358
Query: 351 IKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLW-NLQGEAYVMPALRLSYLHLPVKL 409
+KKC G PLA +G L+ K+ EW + ES + +QG+ VM L+LSY HL ++
Sbjct: 359 VKKCRGVPLALKTMGGLMSSKQSVSEWEVIAESNIGARVQGKNDVMDILKLSYRHLSPEM 418
Query: 410 RQCFSFCALFPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWNELYWRSFFENTE 469
+QCF+FCA+FP+D + K LI LW ANGFI + ++ G ++++L WRSF ++ +
Sbjct: 419 KQCFAFCAIFPQDYEMVKDELIQLWMANGFIQEEENMDLTHKGEMIFHDLVWRSFLQDVK 478
Query: 470 N---VGFG-QITIFKMHDLVHDLAGSVTQDVCCITDD-NSMRTMSEETRHLLIYNRNSFA 524
+G+ + KMHDL+HDLA VT + T + + ++ ++ RHL I
Sbjct: 479 EEFIIGYHCDSIVCKMHDLMHDLAKDVTDECASTTKELDQLKGSIKDVRHLRI--PEEME 536
Query: 525 EANSIQLHHVKSLKTYMEFNFDVYEAGQLSPQVLNCYSLRVLLSHRLNNLSSSIGRLKYL 584
E + SL T ++ ++ +S + N S+R L R + ++S+I K++
Sbjct: 537 ETMTELFKGTSSLHTLIDRSWR-STLWNVSVE-FNLASVRAL---RCSVINSAITNAKHI 591
Query: 585 RYLDISEGRFKNLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTS 644
R+LD+SE LP+S+C L NL+ L+L+ C L+ LP G+ +++L ++ L CDSL
Sbjct: 592 RFLDLSETSIVRLPDSICMLYNLQSLRLNSCDELEYLPKGMRTMRKLIHIYLYWCDSLRR 651
Query: 645 LPRQIGKLTSLNTLSKYIVGEERGFLLEELGQL-NLKGQLHIKNLERLKSVTDAKKANMS 703
+P IG L +L TL+ Y+V E G +EEL L +L +L + NL ++KS AK+ANM
Sbjct: 652 MPPNIGLLNNLRTLTTYVVDTEAGCGIEELKDLQHLTNRLELYNLHKVKSEEKAKQANMY 711
Query: 704 RKK-LNQLWLSWERNEVSQLQENV---EQILEALQPYAQKLYSFGVGGYTGAYFPQWISI 759
+KK L+++ W R + +N E++LE+L PY L + GY G P+W+
Sbjct: 712 QKKNLSEVLFFWGRQKRCMPNDNAYNEERVLESLAPYCSNLKVLELHGYGGVEIPEWMRD 771
Query: 760 P-SLNDLKSLELVDCKSCLNLPELWKLPSLKYLKLSNMIHVIYL-FHESYDGEG------ 811
P + + L + +C C +LP +W L SL+ L LS M ++ L ++ + EG
Sbjct: 772 PHTFQRISKLNISNCPRCKDLPPVWLLVSLEELSLSCMDNLTTLCTNDDVEAEGCGTSLQ 831
Query: 812 -LMALKTLFLEKLPNL----IGLSREER--VMFPRLKALEITECPNLLGLPCLPSLSDLY 864
LK +FL LPNL + +S + + P+L+ L I++CP L G+P P L DL
Sbjct: 832 IFPKLKKMFLRNLPNLERWAVNISGDPSSFITLPQLEILRISDCPKLAGIPDCPVLRDLN 891
Query: 865 IQGKYNQQLPSSIHKLGSLESLHF-SDNEELIYFPDGI--------LRNLASPLKTL--- 912
I N + S H + SL L + ++ + + P G +R+LA+ + +L
Sbjct: 892 IDRCSNIAVSSLAH-VTSLSYLSYDAEGFDSMTMPLGSWSSLMRLKVRSLANMVISLEDQ 950
Query: 913 ---GFHRHSKLKMLPTE------------------MIHIHALQQLYINDCRNIEELPNEV 951
G L+ L +H ++ L I DC +I P E
Sbjct: 951 QNQGESNLVNLRRLNLHGPKCFTTVSGFSELHHGIWVHFAFVEHLVIGDCHDIVRWPTEE 1010
Query: 952 MQRLHSLKELDIVGCDKL----KLSSDFQYLTCLETLAIGSCSEVEGFHEALQHMTTLKS 1007
++ L L+ L I L LS + YL+CLE L I SCS G E + +L+
Sbjct: 1011 LRCLIRLRSLHIFKFTSLGINFSLSEEILYLSCLEELNITSCS---GIVEIPKLPASLEE 1067
Query: 1008 LTLSDLPNLEY-LPECIGNLTLLHEINIYSCPKLACLPTSIQQISGLEILSIHDC 1061
L + NL LP +GNL L + C L LP + ++ L L + C
Sbjct: 1068 LFIQSCQNLVVPLPPNLGNLASLRNFIVIKCESLKLLPDGMDGLTSLRKLHLDGC 1122
Score = 40.4 bits (93), Expect = 0.34
Identities = 78/350 (22%), Positives = 144/350 (40%), Gaps = 89/350 (25%)
Query: 598 PNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSLNT 657
P+S L LE+L++ C L +P L++L++ C ++ + L + +
Sbjct: 858 PSSFITLPQLEILRISDCPKLAGIPD----CPVLRDLNIDRCSNIA-----VSSLAHVTS 908
Query: 658 LSKYIVGEERGF--LLEELGQLNLKGQLHIKNLERLKSVTDAKKANMSRKKLNQLWLSWE 715
LS Y+ + GF + LG + +L +++L ANM +S E
Sbjct: 909 LS-YLSYDAEGFDSMTMPLGSWSSLMRLKVRSL-----------ANMV--------ISLE 948
Query: 716 RNEVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDCKS 775
++ +Q + N+ L L + K ++ V G++ + W+ + + L + DC
Sbjct: 949 -DQQNQGESNLVN-LRRLNLHGPKCFTT-VSGFSELHHGIWVHFAFV---EHLVIGDCHD 1002
Query: 776 CLNLPELWKLPSLKYLKLSNMIHVIYLFHESYDGEGLMALKTLFLEKLPNL-IGLSREER 834
+ W L+ L + L++L + K +L I S E
Sbjct: 1003 IVR----WPTEELRCL---------------------IRLRSLHIFKFTSLGINFSLSEE 1037
Query: 835 VMFPR-LKALEITECPNLLGLPCLP-SLSDLYIQGKYNQQLPSSIHKLGSLESLHFSDNE 892
+++ L+ L IT C ++ +P LP SL +L+IQ N +P +
Sbjct: 1038 ILYLSCLEELNITSCSGIVEIPKLPASLEELFIQSCQNLVVPLPPN-------------- 1083
Query: 893 ELIYFPDGILRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCR 942
L NLAS L+ + LK+LP M + +L++L+++ CR
Sbjct: 1084 ---------LGNLAS-LRNFIVIKCESLKLLPDGMDGLTSLRKLHLDGCR 1123
>gb|AAU43949.1| putative NBS-LRR protein [Oryza sativa (japonica cultivar-group)]
gi|47777415|gb|AAT38049.1| putative NBS-LRR resistance
protein [Oryza sativa (japonica cultivar-group)]
Length = 1222
Score = 547 bits (1410), Expect = e-154
Identities = 404/1239 (32%), Positives = 624/1239 (49%), Gaps = 169/1239 (13%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
+ A+L +L E + E S G ++ + L +LL + + DAE++
Sbjct: 4 LLSALLPALLKKAGESLGTEFSFIGGIERRRSELYTLLLAVNQVINDAEDQASKKPA--- 60
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEY--KASKCGLS--------HKHIAFRYKL 110
VK W+ KLK AA DD +DE E L E + K + + F+Y++
Sbjct: 61 -VKSWIAKLKLAACDADDALDELHYEELRCEALRRGHKINTGVRAFFSSHYNPLLFKYRI 119
Query: 111 AKKMKRIGVWLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIV 170
K++++I +D + ++ N+F S V + QT S V + V GR++++D+IV
Sbjct: 120 GKRLQQIVERIDQLVSQMNRFGFLNC----SMPVDERMQTYSYVDEQEVIGRDKERDEIV 175
Query: 171 DFLVGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRM 230
L+ ++E ++L + PIVG+GGLGKTTLAQLVFN K+ HF+ +WVCVSE+F++ +
Sbjct: 176 HMLL--SAETDELLILPIVGIGGLGKTTLAQLVFNDVKVKAHFQKHMWVCVSENFSVPVI 233
Query: 231 TKAIIEGATKKSC--EDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLAC 288
K II+ A C + +LELLQ++L++ L +KRYLLVLDDVWN+ ++ W L+++L
Sbjct: 234 VKGIIDTAIGNDCGLKFDNLELLQQRLREELGQKRYLLVLDDVWNEDKQKWGALRTLLGS 293
Query: 289 GGKGASILVTTRLPKVAKIMGTIPHHELSRLSDEDCWELFKQRAFGPNEVQQKELVIVGK 348
G G++++VTTR KVA IM +I L L+ ED W +F +RAFG V+ ELV VGK
Sbjct: 294 CGMGSAVVVTTRNVKVASIMESISPLCLENLNPEDSWIVFSRRAFGTGVVETPELVEVGK 353
Query: 349 EIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWNLQGEAYVMPALRLSYLHLPVK 408
I++KC G PLA ++G+L+ K+E ++WL + ES W+ E+ ++PAL L Y +LP
Sbjct: 354 RIVEKCCGLPLAIKSMGALMSTKQETRDWLSILESNTWD--EESQILPALSLGYKNLPSH 411
Query: 409 LRQCFSFCALFPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWNELYWRSFFENT 468
++QCF+FCA+FPKD I K LI LW +NGFI S +M + ++ GN V+ EL WRSFF+N
Sbjct: 412 MKQCFAFCAVFPKDYEIDKDDLIHLWVSNGFIPSKKMSDIEENGNHVFWELVWRSFFQNV 471
Query: 469 ENVG---------FGQ--ITIFKMHDLVHDLAGSVTQDVCCITDD-NSMRTMSEETRHLL 516
+ +G +GQ +T FK+HDL+HDLA ++ D C ++ ++ + + H+
Sbjct: 472 KQIGSIFQRKVYRYGQSDVTTFKIHDLMHDLAVHISGDECLALENLAKIKKIPKNVHHMA 531
Query: 517 IYNRNSFAEANSIQLHHVKSLKTYMEFNFDVYEAGQLSPQVLNCYSLRVLLSH--RLNNL 574
+ + H + +++ F D + N LRV+ H +
Sbjct: 532 FEGQQKI----GFLMQHCRVIRSV--FALDKNDMHIAQDIKFNESPLRVVGLHIFGIEKF 585
Query: 575 SSSIGRLKYLRYLDISEGRFKNLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNL 634
+K+LRYLD+S LP + L NL+VL L+ C L LP G+ + L+++
Sbjct: 586 PVEPAFMKHLRYLDLSGSYINTLPEAASALYNLQVLILNRCRRLTHLPDGMKFMISLRHV 645
Query: 635 SLRDCDSLTSLPRQIGKLTSLNTLSKYIVGEERGFLLEELGQLNLKGQLHIKNLERLKSV 694
L DC LTS+P +G+L +L TL+K++ G E G+ + EL L L G+L I NL ++ +
Sbjct: 646 YLDDCARLTSMPAGLGQLINLRTLTKFVPGNESGYRINELNDLKLGGKLQIFNLIKVTNP 705
Query: 695 TDAKKANMSRK-KLNQLWLSWERNEVSQLQE------NVEQILEALQPYAQKLYSFGVGG 747
+AK+AN+ K L QL L W ++ ++LQ E++L+AL+P L +
Sbjct: 706 IEAKEANLECKTNLQQLALCWGTSKSAELQAEDLHLYRHEEVLDALKP-PNGLTVLKLRQ 764
Query: 748 YTGAYFPQWISIP-SLNDLKSLELVDCKSCLNLPELWKLPSLKYLKLSNMIHVIYLFH-- 804
Y G FP W+ +L ++ L++ D +C+ LP +WKLP L+ L+L +M + YL +
Sbjct: 765 YMGTTFPIWMENGITLRNIVKLKVTDSINCMKLPSVWKLPFLEVLRLKDMKKLKYLCNGF 824
Query: 805 --ESYDGEGLMA---LKTLFLEKLPNL-----IGLSREERVMFPRLKALEITECPNLLGL 854
+ L+A LK L LE++ +L + + FP L A+EI +CP L +
Sbjct: 825 CSDKECDHQLVAFPKLKLLSLERMESLENWQEYDVEQVTPANFPVLDAMEIIDCPKLTAM 884
Query: 855 PCLPSLSDLYIQG-KYNQQLPSSIHKL---------GSLES-----LHFSDN-EELIYFP 898
P P L L + G K L SS+ L GSLE H+ +N E
Sbjct: 885 PNAPVLKSLSVIGNKILIGLSSSVSNLSYLYLGASQGSLERKKTLIYHYKENLEGTTDSK 944
Query: 899 DGILRNLASPLKTL-GFHRHSKLKMLPTEMIHIH-------------------------- 931
D +L + S +L H + P ++ +I
Sbjct: 945 DHVLAHHFSSWGSLTKLHLQGFSALAPEDIQNISGHVMSVQNLDLISCDCFIQYDTLQSP 1004
Query: 932 --------ALQQLYINDCRNIEELPNEVMQRLHSLKELDIVGCDKL-------------- 969
LQ L I C ++ P E Q L SLK LDI C+
Sbjct: 1005 LWFWKSFACLQHLTIEYCNSLTFWPGEEFQSLTSLKRLDIRYCNNFTGMPPAQVSVKSFE 1064
Query: 970 --------KLSSDFQY-----LTCLETLAIGSCSEVEGFHEALQHMTTLKSLT------L 1010
++ +F Y T L L I SC+ +E E L + L+SL+ L
Sbjct: 1065 DEGMHNLERIEIEFCYNLVAFPTSLSYLRICSCNVLEDLPEGLGCLGALRSLSIDYNPRL 1124
Query: 1011 SDLP------------------NLEYLPECIGNLTLLHEINIYSCPKLACLPTSIQQ-IS 1051
LP +L LPE + NLT L+++ I++CP L LP +QQ +
Sbjct: 1125 KSLPPSIQRLSNLTRLYLGTNDSLTTLPEGMHNLTALNDLAIWNCPSLKALPEGLQQRLH 1184
Query: 1052 GLEILSIHDCSKLEKRCQKEIGEDWPKIVHVQYIEIEND 1090
LE L I C L +RC K G+ W K+ + + + D
Sbjct: 1185 SLEKLFIRQCPTLVRRC-KRGGDYWSKVKDIPDLRVTGD 1222
>ref|NP_915900.1| putative NBS-LRR type resistance protein [Oryza sativa (japonica
cultivar-group)] gi|13872974|dbj|BAB44079.1| putative
NBS-LRR type resistance protein [Oryza sativa (japonica
cultivar-group)]
Length = 1110
Score = 540 bits (1392), Expect = e-152
Identities = 380/1122 (33%), Positives = 602/1122 (52%), Gaps = 56/1122 (4%)
Query: 4 AVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGRDVK 63
A ++ + LSE + G + L+S L+ ++A L+DAEEKQ +D+ V+
Sbjct: 9 AFMQTLFQKLSEATLDHFISWRGIHGKLESLSSTLSQLQAFLDDAEEKQLTDAS----VR 64
Query: 64 DWLLKLKDAAYTLDDIMDECATEALEME-----YKASKCGLSHKHIA---FRYKLAKKMK 115
WL KLKD AY LDD++D + +++ M+ + LS ++ +++++ K+
Sbjct: 65 GWLAKLKDIAYDLDDLLDSYSAKSMRMKQRQVIFPTKASFLSSSFLSRNLYQHRIKHKIN 124
Query: 116 RIGVWLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVG 175
I LD IA E++ L I R + Q++S+V V+GR D++++V ++
Sbjct: 125 IILERLDKIAQERDTIGLQMICEMRRYDTSERPQSSSLVDSSAVFGRERDREEMVRLVLS 184
Query: 176 DASEQE-DLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAI 234
D +L V P+VG+GGLGKTTL Q+V++ D++ HF+L+IW+ VSE F +++T+
Sbjct: 185 DNGHNSCNLCVIPVVGMGGLGKTTLMQMVYHDDRVREHFDLRIWIYVSESFDERKLTQET 244
Query: 235 IEGAT-KKSCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGA 293
+E + +S ++ +LQ L +LR KRYLLVLDDVWN+ + W ++ L GG G+
Sbjct: 245 LEASDYDQSVASTNMNMLQETLSRVLRGKRYLLVLDDVWNEDLDKWHSYRAALISGGFGS 304
Query: 294 SILVTTRLPKVAKIMGTIPHHELSRLSDEDCWELFKQRAFGPNEVQ-QKELVIVGKEIIK 352
I+VT+R V +IMG I ++L +LSD+D W +FK AF + EL +G EI+K
Sbjct: 305 KIVVTSRNENVGRIMGGIEPYKLQKLSDDDSWSVFKSHAFRDGDCSAHPELEAIGMEIVK 364
Query: 353 KCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWNLQGEA-YVMPALRLSYLHLPVKLRQ 411
K G PLA+ ALGSLL K +E+EW + ++ +W L + ++PALRLSY HLP L+Q
Sbjct: 365 KLKGLPLASKALGSLLFCKTDEEEWKDILQNDIWELPADKNNILPALRLSYNHLPPHLKQ 424
Query: 412 CFSFCALFPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWNELYWRSFFENTENV 471
CF+FC+++PKD + ++ L+ +W A GFI ++ +D GN +NEL RSFF+ EN
Sbjct: 425 CFAFCSVYPKDYMFRREKLVKIWLALGFIRQSRKKRMEDTGNAYFNELLSRSFFQPYEN- 483
Query: 472 GFGQITIFKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHLLIYNRNSFAEANSIQL 531
+ MHD +HDLA S++ + C D + +TRHL +++ + L
Sbjct: 484 ------NYVMHDAMHDLAKSISMEDCDHLDYGRRHDNAIKTRHLSFPCKDAKC-MHFNPL 536
Query: 532 HHVKSLKTYMEFNFDVYEAGQLSPQV-LNCYSLRVLLSH--RLNNLSSSIGRLKYLRYLD 588
+ + L+T + QL + + LRVL H L L SIG LK LR+LD
Sbjct: 537 YGFRKLRTLTIIHGYKSRMSQLPHGLFMKLEYLRVLDMHGQGLKELPESIGNLKQLRFLD 596
Query: 589 ISEGRFKNLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQ 648
+S + LP SL KL NL++LKL C L+++P G+TRL L++L L S
Sbjct: 597 LSSTEIETLPASLVKLYNLQILKLSDCNFLREVPQGITRLINLRHLEA--STRLLSRIHG 654
Query: 649 IGKLTSLNTLSKYIVGEERGFLLEELGQLN-LKGQLHIKNLERLKSVTDAKKANMSRKK- 706
IG L L L +++V + G + EL ++ L+GQL I+ L + + DA A + K+
Sbjct: 655 IGSLVCLQELEEFVVQKRSGHNVTELNNMDELQGQLSIRGLNNVPNGQDAVCAKLRNKEH 714
Query: 707 LNQLWLSWERNEVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLK 766
L L L W+ + S E +++LE LQP+ L + G+ G FP W++ L L+
Sbjct: 715 LRTLHLIWDEDCESNPSEQ-QEVLEGLQPHLD-LKELVIKGFPGVRFPSWLASSFLPKLQ 772
Query: 767 SLELVDCKSCLNLPELWKLPSLKYLKLSNMIHVIYLFHESYDG----EGLMALKTLFLEK 822
++ + +C+S LP L +LP LKYL ++ + V L E + G +G AL+ L LE
Sbjct: 773 TIHICNCRS-TRLPALGQLPFLKYLVIAGVTEVTQLSSE-FTGFGQPKGFPALEDLLLED 830
Query: 823 LPNLI-GLSREERVMFPRLKALEITECPNLLGLPCLPS-LSDLYIQGKYNQQLPSSIHKL 880
+PNL + +FP+L L + +CP L LP +PS L L+I + LP +
Sbjct: 831 MPNLSEWIFDVADQLFPQLTELGLIKCPQLKKLPPIPSTLRTLWISESGLESLPELQNNS 890
Query: 881 --GSLESLHFSDNEELIYFPDGILRNLASPLKTLGFHRHSKLKMLPTEMIH-IHALQQLY 937
S SL+ +D L G+L + LK+L L LP E + +L+ L+
Sbjct: 891 CPSSPTSLYINDCPNLTSLRVGLLAYRPTALKSLTIAHCEGLVSLPEECFRPLISLRSLH 950
Query: 938 INDCRNIEELPNEVMQ---RLHSLKELDIVGCDKLK--LSSDFQYLTCLETLAIGSCSEV 992
I +C + +P ++ S++++ + C L L + YL L I C ++
Sbjct: 951 IYECPCL--VPWTALEGGLLPTSIEDIRLNSCTPLASVLLNGLSYLPHLRHFEIADCPDI 1008
Query: 993 EGF-HEALQHMTTLKSLTLSDLPNLEYLPECIGNLTLLHEINIYSCPKLACLPTSIQQIS 1051
F E L H TL+ L +S +L+ LP + N++ L + I +CP + LP +
Sbjct: 1009 NNFPAEGLPH--TLQFLEISCCDDLQCLPPGLHNISSLETLRISNCPGVESLPKEGLPM- 1065
Query: 1052 GLEILSIHDCSKLEKRCQKEIGEDWPKIVHVQYIEIENDNLI 1093
GL L I C +++++CQ E GE KI H++ IEI+ D ++
Sbjct: 1066 GLNELYIKGCPQIKQQCQ-EGGEYHAKIAHIRDIEIDGDVIV 1106
>ref|XP_483600.1| putative NBS-LRR resistance protein RGH1 [Oryza sativa (japonica
cultivar-group)] gi|42407842|dbj|BAD08985.1| putative
NBS-LRR resistance protein RGH1 [Oryza sativa (japonica
cultivar-group)]
Length = 1048
Score = 539 bits (1388), Expect = e-151
Identities = 349/1022 (34%), Positives = 562/1022 (54%), Gaps = 69/1022 (6%)
Query: 5 VLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGRDVKD 64
V+ V+G + + + ++ G D + ++L L ++ L DAE K SE VK
Sbjct: 9 VVRGVVGKAAGALVQSVTRMCGVDGDRHKLERQLLAVQCKLSDAEAK----SETSPAVKR 64
Query: 65 WLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLS--------HKHIAFRYKLAKKMKR 116
W+ LK AY DD++D+ EAL + + H + FR ++KK+
Sbjct: 65 WMKDLKAVAYEADDVLDDFHYEALRRDAQIGDSTTDKVLGYFTPHSPLLFRVAMSKKLNS 124
Query: 117 IGVWLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVGD 176
+ ++++ E NKF L E + + V + + + + + GR++DK+ +V+ L+
Sbjct: 125 VLKKINELVEEMNKFGLVERADQATVHVIHPQTHSGLDSLMEIVGRDDDKEMVVNLLLEQ 184
Query: 177 ASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIE 236
S++ + V IVG+GGLGKTTLA++V+N ++ FEL +W+CVS+DF + + ++IIE
Sbjct: 185 RSKRM-VEVLSIVGMGGLGKTTLAKMVYNDTRVQQRFELPMWLCVSDDFNVVSLVRSIIE 243
Query: 237 GATKKSCEDLD-LELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVL-ACGGKGAS 294
AT+ +C D +ELL+ +L +++ RKRYLLVLDDVWN+++ W+ L+ +L + G G+
Sbjct: 244 LATRGNCTLPDRIELLRSRLHEVVGRKRYLLVLDDVWNEEEHKWEELRPLLHSAGAPGSV 303
Query: 295 ILVTTRLPKVAKIMGTIPHHELSRLSDEDCWELFKQRAFGPNEVQQKELVIVGKEIIKKC 354
+LVTTR +VA IMGT+P H LS L+ +D WELF+++AF E QQ E +G I+KKC
Sbjct: 304 VLVTTRSQRVASIMGTVPAHTLSYLNHDDSWELFRKKAFSKEEEQQPEFAEIGNRIVKKC 363
Query: 355 GGFPLAAIALGSLLRFKREEKEWLYVKESKLWNLQGEA-YVMPALRLSYLHLPVKLRQCF 413
G PLA +G L+ K+ +EW + SK W G ++ L+LSY HLP++++QCF
Sbjct: 364 KGLPLALKTMGGLMSSKKRIQEWEAIAGSKSWEDVGTTNEILSILKLSYRHLPLEMKQCF 423
Query: 414 SFCALFPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWNELYWRSFFENTE---- 469
+FCA+FPKD + + L+ LW AN FI M++ ++ G V+NEL WRSFF++ +
Sbjct: 424 AFCAIFPKDYQMERDKLVQLWIANNFIQEEGMMDLEERGQFVFNELVWRSFFQDVKVESF 483
Query: 470 NVGFGQ----ITIFKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHLLIYNRNSFAE 525
+VG Q IT + MHDL+HDLA SVT++ D N + ++ RHL+ ++ +
Sbjct: 484 HVGIKQTYKSITCY-MHDLMHDLAKSVTEECVDAQDLNQQKASMKDVRHLM---SSAKLQ 539
Query: 526 ANSIQLHHVKSLKTYMEFNFDVYEAGQLSPQVLNCYSLRVLLSHRLNNLSSSIGRLKYLR 585
NS HV L T + + + + LN SLR L + +LN ++ + +LR
Sbjct: 540 ENSELFKHVGPLHTLLSPYWSKSSPLPRNIKRLNLTSLRALHNDKLNVSPKALASITHLR 599
Query: 586 YLDIS-EGRFKNLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTS 644
YLD+S + ++LP+S+C L +L+ L+L+GC+ LQ LP G+ + +L++L L C SL
Sbjct: 600 YLDLSHSSKLEHLPDSICMLYSLQALRLNGCLKLQHLPEGMRFMSKLRHLYLIGCHSLKR 659
Query: 645 LPRQIGKLTSLNTLSKYIVGEERGFLLEELGQL-NLKGQLHIKNLERLKSVTDAKKANMS 703
+P +IG+L +L TL+ ++V + G LEEL L +L G+L + NL+ ++S ++A++AN+
Sbjct: 660 MPPRIGQLKNLRTLTTFVVDTKDGCGLEELKDLHHLGGRLELFNLKAIQSGSNAREANLH 719
Query: 704 -RKKLNQLWLSW--------ERNEVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAYFP 754
++ + +L L W + + + +N ++I+E P +L + V G
Sbjct: 720 IQENVTELLLHWCHDIFEYSDHDFDLDVVDNKKEIVEFSLP-PSRLETLQVWGSGHIEMS 778
Query: 755 QWISIPSL-NDLKSLELVDCKSCLNLPELWKLPSLKYLKLSNMIHVIYL---FHESYDG- 809
W+ P++ LK L + +C C +LP LW+ SL+ L LS + ++ L + G
Sbjct: 779 SWMKNPAIFLCLKELHMSECWRCKDLPPLWQSVSLESLSLSRLDNLTTLSSGIDMAVPGC 838
Query: 810 ----EGLMALKTLFLEKLPNLIGLSREE--RVMFPRLKALEITECPNLLGLPCLPSL-SD 862
E LK + L LPNL E VMFP LK L+I CP L+ +P P L +
Sbjct: 839 NGSLEIFPKLKKMHLHYLPNLEKWMDNEVTSVMFPELKELKIYNCPKLVNIPKAPILCKN 898
Query: 863 LYIQGKYNQQLPSSIHKL---------------GSLESLHFSDNEELIYFPDGILRNLAS 907
L PS + KL SLE+L ++ L+ P + R +
Sbjct: 899 LTSSSSEESLFPSGLEKLYIEFCNNLLEIPKLPASLETLRINECTSLVSLPPNLAR--LA 956
Query: 908 PLKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCRNIEELPNEVMQRLHSLKELDIVGCD 967
L+ L S L+ LP M + LQ+L + C +E LP ++QRL +L++L +G
Sbjct: 957 KLRDLTLFSCSSLRNLPDVMDGLTGLQELCVRQCPGVETLPQSLLQRLPNLRKLMTLGSH 1016
Query: 968 KL 969
KL
Sbjct: 1017 KL 1018
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.320 0.138 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,882,171,729
Number of Sequences: 2540612
Number of extensions: 80600773
Number of successful extensions: 240061
Number of sequences better than 10.0: 4253
Number of HSP's better than 10.0 without gapping: 1719
Number of HSP's successfully gapped in prelim test: 2582
Number of HSP's that attempted gapping in prelim test: 210271
Number of HSP's gapped (non-prelim): 15472
length of query: 1113
length of database: 863,360,394
effective HSP length: 139
effective length of query: 974
effective length of database: 510,215,326
effective search space: 496949727524
effective search space used: 496949727524
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 81 (35.8 bits)
Medicago: description of AC134049.3


