

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133779.11 - phase: 0
(1067 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
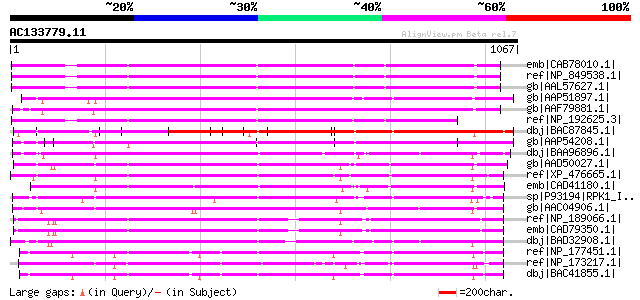
Score E
Sequences producing significant alignments: (bits) Value
emb|CAB78010.1| receptor protein kinase-like protein [Arabidopsi... 776 0.0
ref|NP_849538.1| leucine-rich repeat family protein / protein ki... 776 0.0
gb|AAL57627.1| AT4g08850/T32A17_160 [Arabidopsis thaliana] 775 0.0
gb|AAP51897.1| putative protein kinase [Oryza sativa (japonica c... 768 0.0
gb|AAF79881.1| Contains similarity to receptor protein kinase-li... 747 0.0
ref|NP_192625.3| leucine-rich repeat family protein / protein ki... 699 0.0
dbj|BAC87845.1| leucine-rich repeat receptor-like protein kinase... 632 e-179
gb|AAP54208.1| putative protein kinase [Oryza sativa (japonica c... 612 e-173
dbj|BAA96896.1| receptor-like protein kinase [Arabidopsis thalia... 576 e-162
gb|AAD50027.1| Similar to leucine-rich receptor-like protein kin... 569 e-160
ref|XP_476665.1| putative LRR receptor-like kinase [Oryza sativa... 562 e-158
emb|CAD41180.1| OSJNBb0002J11.4 [Oryza sativa (japonica cultivar... 550 e-155
sp|P93194|RPK1_IPONI Receptor-like protein kinase precursor gi|1... 545 e-153
gb|AAC04906.1| putative receptor-like protein kinase [Arabidopsi... 533 e-149
ref|NP_189066.1| leucine-rich repeat transmembrane protein kinas... 529 e-148
emb|CAD79350.1| LRR receptor-like kinase 2 [Arabidopsis thaliana] 528 e-148
dbj|BAD32908.1| putative receptor-like protein kinase 2 [Oryza s... 526 e-147
ref|NP_177451.1| leucine-rich repeat transmembrane protein kinas... 525 e-147
ref|NP_173217.1| leucine-rich repeat transmembrane protein kinas... 524 e-147
dbj|BAC41855.1| unknown protein [Arabidopsis thaliana] 523 e-146
>emb|CAB78010.1| receptor protein kinase-like protein [Arabidopsis thaliana]
gi|7321074|emb|CAB82121.1| receptor protein kinase-like
protein [Arabidopsis thaliana] gi|25407456|pir||B85089
receptor protein kinase-like protein [imported] -
Arabidopsis thaliana
Length = 1027
Score = 776 bits (2005), Expect = 0.0
Identities = 451/1044 (43%), Positives = 616/1044 (58%), Gaps = 49/1044 (4%)
Query: 4 LPTLIMILCVLP-TLSVAEDSEAKLALLKWKDSFDDQ-SQTLLSTWKN-NTNPCKPKWRG 60
L L++I VL + +V+ E ALLKWK +F +Q S + LS+W N NT+ W G
Sbjct: 10 LQVLLIISIVLSCSFAVSATVEEANALLKWKSTFTNQTSSSKLSSWVNPNTSSFCTSWYG 69
Query: 61 IKCDKSNFISTIGLANLGLKGTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISI 120
+ C + I + L N G++GT FSS PNL +D+ N F GTI G S
Sbjct: 70 VACSLGSIIR-LNLTNTGIEGTFEDFPFSSLPNLTFVDLSMNRFSGTISPLWGRFSK--- 125
Query: 121 LTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSIGNLTNLSYLILGGNNWSGGP 180
L++ D+S +L G IP +G+L+NL L L N +G
Sbjct: 126 ---------------------LEYFDLSINQLVGEIPPELGDLSNLDTLHLVENKLNGS- 163
Query: 181 IPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKLD 240
IP EIG+L + +AI + L G IP G LT L + L NSLSG IP IGNL L
Sbjct: 164 IPSEIGRLTKVTEIAIYDNLLTGPIPSSFGNLTKLVNLYLFINSLSGSIPSEIGNLPNLR 223
Query: 241 TLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSG 300
L L N ++G IP S N+ ++T+L LSG IP I N+ L L+L N L+G
Sbjct: 224 ELCLDRNN-LTGKIPSSFGNLKNVTLLNMFENQLSGEIPPEIGNMTALDTLSLHTNKLTG 282
Query: 301 SIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWL 360
IPST+G++K L L+L N L+G IP +G + ++ L + EN LTG +P S G L L
Sbjct: 283 PIPSTLGNIKTLAVLHLYLNQLNGSIPPELGEMESMIDLEISENKLTGPVPDSFGKLTAL 342
Query: 361 TVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTG 420
+ N+L G IP G+ N T V N+F G LP IC GG L L D N F G
Sbjct: 343 EWLFLRDNQLSGPIPPGIANSTELTVLQVDTNNFTGFLPDTICRGGKLENLTLDDNHFEG 402
Query: 421 PIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNL 480
P+P SL+ C S+ R+ + N GDI++ FGVYP L ++DLS+N FHGQ+S NW +S L
Sbjct: 403 PVPKSLRDCKSLIRVRFKGNSFSGDISEAFGVYPTLNFIDLSNNNFHGQLSANWEQSQKL 462
Query: 481 QTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFS 540
FI+SNN+I+G IP + +T+L L LSSN++TG+LP E + + + L+++ N S
Sbjct: 463 VAFILSNNSITGAIPPEIWNMTQLSQLDLSSNRITGELP-ESISNINRISKLQLNGNRLS 521
Query: 541 DNIPSEIGLLQRLQELDLGGNELSGKIPKELVELPNLRMLNLSRNKIEGIIPIKFD--SG 598
IPS I LL L+ LDL N S +IP L LP L +NLSRN ++ IP S
Sbjct: 522 GKIPSGIRLLTNLEYLDLSSNRFSSEIPPTLNNLPRLYYMNLSRNDLDQTIPEGLTKLSQ 581
Query: 599 LESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSGTIPQNFGRNLVF--VNISDNQLE 656
L+ LDLS N L G I + L L +L+LSHN LSG IP +F L V++S N L+
Sbjct: 582 LQMLDLSYNQLDGEISSQFRSLQNLERLDLSHNNLSGQIPPSFKDMLALTHVDVSHNNLQ 641
Query: 657 GPLPKIPAFLSASFESLKNNNHLCGNI---RGLDPCATSHSRKR---KNVLRPVFIALGA 710
GP+P AF +A ++ + N LCG++ +GL PC+ + S+K +N++ + + +
Sbjct: 642 GPIPDNAAFRNAPPDAFEGNKDLCGSVNTTQGLKPCSITSSKKSHKDRNLIIYILVPIIG 701
Query: 711 VILVLCVVGALMYIMCGRKKPNE-ESQTEEVQRGVLFSIWSHDGKMMFENIIEATANFDD 769
I++L V + +C RK+ + E T+ G SI+S DGK+ ++ II+AT FD
Sbjct: 702 AIIILSVCAGIF--ICFRKRTKQIEEHTDSESGGETLSIFSFDGKVRYQEIIKATGEFDP 759
Query: 770 KYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKS-FMSEIETLTGIKHRNI 828
KYL+G G G VYKA+L ++AVKKL+ TD +S S+K F++EI LT I+HRN+
Sbjct: 760 KYLIGTGGHGKVYKAKLPNA-IMAVKKLNETTDSSISNPSTKQEFLNEIRALTEIRHRNV 818
Query: 829 IKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQAVAFDWEKRVNVVKGVANALSYLHHD 888
+KL GFCSH + +FLVY+++E GSL ++L ND +A DW KR+NVVKGVA+ALSY+HHD
Sbjct: 819 VKLFGFCSHRRNTFLVYEYMERGSLRKVLENDDEAKKLDWGKRINVVKGVAHALSYMHHD 878
Query: 889 CSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLKPGLHSWTQFAGTFGYAAPELAQTME 948
SP I+HRDISS N+LL DYEA +SDFGTAK LKP +W+ AGT+GY APELA M+
Sbjct: 879 RSPAIVHRDISSGNILLGEDYEAKISDFGTAKLLKPDSSNWSAVAGTYGYVAPELAYAMK 938
Query: 949 VNEKCDVYSFGVLALETIMGKHPGDLISLFLSPSTRPMANNMLLTDVLDQRPQQVMEPID 1008
V EKCDVYSFGVL LE I G+HPGDL+S S+ P + L + D R + I
Sbjct: 939 VTEKCDVYSFGVLTLEVIKGEHPGDLVSTL---SSSPPDATLSLKSISDHRLPEPTPEIK 995
Query: 1009 EEVILIARLAFACLSQNPRLRPSM 1032
EEV+ I ++A CL +P+ RP+M
Sbjct: 996 EEVLEILKVALLCLHSDPQARPTM 1019
>ref|NP_849538.1| leucine-rich repeat family protein / protein kinase family protein
[Arabidopsis thaliana]
Length = 1045
Score = 776 bits (2005), Expect = 0.0
Identities = 451/1044 (43%), Positives = 616/1044 (58%), Gaps = 49/1044 (4%)
Query: 4 LPTLIMILCVLP-TLSVAEDSEAKLALLKWKDSFDDQ-SQTLLSTWKN-NTNPCKPKWRG 60
L L++I VL + +V+ E ALLKWK +F +Q S + LS+W N NT+ W G
Sbjct: 28 LQVLLIISIVLSCSFAVSATVEEANALLKWKSTFTNQTSSSKLSSWVNPNTSSFCTSWYG 87
Query: 61 IKCDKSNFISTIGLANLGLKGTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISI 120
+ C + I + L N G++GT FSS PNL +D+ N F GTI G S
Sbjct: 88 VACSLGSIIR-LNLTNTGIEGTFEDFPFSSLPNLTFVDLSMNRFSGTISPLWGRFSK--- 143
Query: 121 LTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSIGNLTNLSYLILGGNNWSGGP 180
L++ D+S +L G IP +G+L+NL L L N +G
Sbjct: 144 ---------------------LEYFDLSINQLVGEIPPELGDLSNLDTLHLVENKLNGS- 181
Query: 181 IPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKLD 240
IP EIG+L + +AI + L G IP G LT L + L NSLSG IP IGNL L
Sbjct: 182 IPSEIGRLTKVTEIAIYDNLLTGPIPSSFGNLTKLVNLYLFINSLSGSIPSEIGNLPNLR 241
Query: 241 TLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSG 300
L L N ++G IP S N+ ++T+L LSG IP I N+ L L+L N L+G
Sbjct: 242 ELCLDRNN-LTGKIPSSFGNLKNVTLLNMFENQLSGEIPPEIGNMTALDTLSLHTNKLTG 300
Query: 301 SIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWL 360
IPST+G++K L L+L N L+G IP +G + ++ L + EN LTG +P S G L L
Sbjct: 301 PIPSTLGNIKTLAVLHLYLNQLNGSIPPELGEMESMIDLEISENKLTGPVPDSFGKLTAL 360
Query: 361 TVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTG 420
+ N+L G IP G+ N T V N+F G LP IC GG L L D N F G
Sbjct: 361 EWLFLRDNQLSGPIPPGIANSTELTVLQVDTNNFTGFLPDTICRGGKLENLTLDDNHFEG 420
Query: 421 PIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNL 480
P+P SL+ C S+ R+ + N GDI++ FGVYP L ++DLS+N FHGQ+S NW +S L
Sbjct: 421 PVPKSLRDCKSLIRVRFKGNSFSGDISEAFGVYPTLNFIDLSNNNFHGQLSANWEQSQKL 480
Query: 481 QTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFS 540
FI+SNN+I+G IP + +T+L L LSSN++TG+LP E + + + L+++ N S
Sbjct: 481 VAFILSNNSITGAIPPEIWNMTQLSQLDLSSNRITGELP-ESISNINRISKLQLNGNRLS 539
Query: 541 DNIPSEIGLLQRLQELDLGGNELSGKIPKELVELPNLRMLNLSRNKIEGIIPIKFD--SG 598
IPS I LL L+ LDL N S +IP L LP L +NLSRN ++ IP S
Sbjct: 540 GKIPSGIRLLTNLEYLDLSSNRFSSEIPPTLNNLPRLYYMNLSRNDLDQTIPEGLTKLSQ 599
Query: 599 LESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSGTIPQNFGRNLVF--VNISDNQLE 656
L+ LDLS N L G I + L L +L+LSHN LSG IP +F L V++S N L+
Sbjct: 600 LQMLDLSYNQLDGEISSQFRSLQNLERLDLSHNNLSGQIPPSFKDMLALTHVDVSHNNLQ 659
Query: 657 GPLPKIPAFLSASFESLKNNNHLCGNI---RGLDPCATSHSRKR---KNVLRPVFIALGA 710
GP+P AF +A ++ + N LCG++ +GL PC+ + S+K +N++ + + +
Sbjct: 660 GPIPDNAAFRNAPPDAFEGNKDLCGSVNTTQGLKPCSITSSKKSHKDRNLIIYILVPIIG 719
Query: 711 VILVLCVVGALMYIMCGRKKPNE-ESQTEEVQRGVLFSIWSHDGKMMFENIIEATANFDD 769
I++L V + +C RK+ + E T+ G SI+S DGK+ ++ II+AT FD
Sbjct: 720 AIIILSVCAGIF--ICFRKRTKQIEEHTDSESGGETLSIFSFDGKVRYQEIIKATGEFDP 777
Query: 770 KYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKS-FMSEIETLTGIKHRNI 828
KYL+G G G VYKA+L ++AVKKL+ TD +S S+K F++EI LT I+HRN+
Sbjct: 778 KYLIGTGGHGKVYKAKLPNA-IMAVKKLNETTDSSISNPSTKQEFLNEIRALTEIRHRNV 836
Query: 829 IKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQAVAFDWEKRVNVVKGVANALSYLHHD 888
+KL GFCSH + +FLVY+++E GSL ++L ND +A DW KR+NVVKGVA+ALSY+HHD
Sbjct: 837 VKLFGFCSHRRNTFLVYEYMERGSLRKVLENDDEAKKLDWGKRINVVKGVAHALSYMHHD 896
Query: 889 CSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLKPGLHSWTQFAGTFGYAAPELAQTME 948
SP I+HRDISS N+LL DYEA +SDFGTAK LKP +W+ AGT+GY APELA M+
Sbjct: 897 RSPAIVHRDISSGNILLGEDYEAKISDFGTAKLLKPDSSNWSAVAGTYGYVAPELAYAMK 956
Query: 949 VNEKCDVYSFGVLALETIMGKHPGDLISLFLSPSTRPMANNMLLTDVLDQRPQQVMEPID 1008
V EKCDVYSFGVL LE I G+HPGDL+S S+ P + L + D R + I
Sbjct: 957 VTEKCDVYSFGVLTLEVIKGEHPGDLVSTL---SSSPPDATLSLKSISDHRLPEPTPEIK 1013
Query: 1009 EEVILIARLAFACLSQNPRLRPSM 1032
EEV+ I ++A CL +P+ RP+M
Sbjct: 1014 EEVLEILKVALLCLHSDPQARPTM 1037
>gb|AAL57627.1| AT4g08850/T32A17_160 [Arabidopsis thaliana]
Length = 1045
Score = 775 bits (2002), Expect = 0.0
Identities = 450/1044 (43%), Positives = 616/1044 (58%), Gaps = 49/1044 (4%)
Query: 4 LPTLIMILCVLP-TLSVAEDSEAKLALLKWKDSFDDQ-SQTLLSTWKN-NTNPCKPKWRG 60
L L++I VL + +V+ E ALLKWK +F +Q S + LS+W N NT+ W G
Sbjct: 28 LQVLLIISIVLSCSFAVSATVEEANALLKWKSTFTNQTSSSKLSSWVNPNTSSFCTSWYG 87
Query: 61 IKCDKSNFISTIGLANLGLKGTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISI 120
+ C + I + L N G++GT FSS PNL +D+ N F GTI G S
Sbjct: 88 VACSLGSIIR-LNLTNTGIEGTFEDFPFSSLPNLTFVDLSMNRFSGTISPLWGRFSK--- 143
Query: 121 LTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSIGNLTNLSYLILGGNNWSGGP 180
L++ D+S +L G IP +G+L+NL L L N +G
Sbjct: 144 ---------------------LEYFDLSINQLVGEIPPELGDLSNLDTLHLVENKLNGS- 181
Query: 181 IPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKLD 240
IP EIG+L + +AI + L G IP G LT L + L NSLSG IP IGNL L
Sbjct: 182 IPSEIGRLTKVTEIAIYDNLLTGPIPSSFGNLTKLVNLYLFINSLSGSIPSEIGNLPNLR 241
Query: 241 TLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSG 300
L L N ++G IP S N+ ++T+L LSG IP I N+ L L+L N L+G
Sbjct: 242 ELCLDRNN-LTGKIPSSFGNLKNVTLLNMFENQLSGEIPPEIGNMTALDTLSLHTNKLTG 300
Query: 301 SIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWL 360
IPST+G++K L L+L N L+G IP +G + ++ L + EN LTG +P S G L L
Sbjct: 301 PIPSTLGNIKTLAVLHLYLNQLNGSIPPELGEMESMIDLEISENKLTGPVPDSFGKLTAL 360
Query: 361 TVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTG 420
+ N+L G IP G+ N T + N+F G LP IC GG L L D N F G
Sbjct: 361 EWLFLRDNQLSGPIPPGIANSTELTVLQLDTNNFTGFLPDTICRGGKLENLTLDDNHFEG 420
Query: 421 PIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNL 480
P+P SL+ C S+ R+ + N GDI++ FGVYP L ++DLS+N FHGQ+S NW +S L
Sbjct: 421 PVPKSLRDCKSLIRVRFKGNSFSGDISEAFGVYPTLNFIDLSNNNFHGQLSANWEQSQKL 480
Query: 481 QTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFS 540
FI+SNN+I+G IP + +T+L L LSSN++TG+LP E + + + L+++ N S
Sbjct: 481 VAFILSNNSITGAIPPEIWNMTQLSQLDLSSNRITGELP-ESISNINRISKLQLNGNRLS 539
Query: 541 DNIPSEIGLLQRLQELDLGGNELSGKIPKELVELPNLRMLNLSRNKIEGIIPIKFD--SG 598
IPS I LL L+ LDL N S +IP L LP L +NLSRN ++ IP S
Sbjct: 540 GKIPSGIRLLTNLEYLDLSSNRFSSEIPPTLNNLPRLYYMNLSRNDLDQTIPEGLTKLSQ 599
Query: 599 LESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSGTIPQNFGRNLVF--VNISDNQLE 656
L+ LDLS N L G I + L L +L+LSHN LSG IP +F L V++S N L+
Sbjct: 600 LQMLDLSYNQLDGEISSQFRSLQNLERLDLSHNNLSGQIPPSFKDMLALTHVDVSHNNLQ 659
Query: 657 GPLPKIPAFLSASFESLKNNNHLCGNI---RGLDPCATSHSRKR---KNVLRPVFIALGA 710
GP+P AF +A ++ + N LCG++ +GL PC+ + S+K +N++ + + +
Sbjct: 660 GPIPDNAAFRNAPPDAFEGNKDLCGSVNTTQGLKPCSITSSKKSHKDRNLIIYILVPIIG 719
Query: 711 VILVLCVVGALMYIMCGRKKPNE-ESQTEEVQRGVLFSIWSHDGKMMFENIIEATANFDD 769
I++L V + +C RK+ + E T+ G SI+S DGK+ ++ II+AT FD
Sbjct: 720 AIIILSVCAGIF--ICFRKRTKQIEEHTDSESGGETLSIFSFDGKVRYQEIIKATGEFDP 777
Query: 770 KYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKS-FMSEIETLTGIKHRNI 828
KYL+G G G VYKA+L ++AVKKL+ TD +S S+K F++EI LT I+HRN+
Sbjct: 778 KYLIGTGGHGKVYKAKLPNA-IMAVKKLNETTDSSISNPSTKQEFLNEIRALTEIRHRNV 836
Query: 829 IKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQAVAFDWEKRVNVVKGVANALSYLHHD 888
+KL GFCSH + +FLVY+++E GSL ++L ND +A DW KR+NVVKGVA+ALSY+HHD
Sbjct: 837 VKLFGFCSHRRNTFLVYEYMERGSLRKVLENDDEAKKLDWGKRINVVKGVAHALSYMHHD 896
Query: 889 CSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLKPGLHSWTQFAGTFGYAAPELAQTME 948
SP I+HRDISS N+LL DYEA +SDFGTAK LKP +W+ AGT+GY APELA M+
Sbjct: 897 RSPAIVHRDISSGNILLGEDYEAKISDFGTAKLLKPDSSNWSAVAGTYGYVAPELAYAMK 956
Query: 949 VNEKCDVYSFGVLALETIMGKHPGDLISLFLSPSTRPMANNMLLTDVLDQRPQQVMEPID 1008
V EKCDVYSFGVL LE I G+HPGDL+S S+ P + L + D R + I
Sbjct: 957 VTEKCDVYSFGVLTLEVIKGEHPGDLVSTL---SSSPPDATLSLKSISDHRLPEPTPEIK 1013
Query: 1009 EEVILIARLAFACLSQNPRLRPSM 1032
EEV+ I ++A CL +P+ RP+M
Sbjct: 1014 EEVLEILKVALLCLHSDPQARPTM 1037
>gb|AAP51897.1| putative protein kinase [Oryza sativa (japonica cultivar-group)]
gi|37530616|ref|NP_919610.1| putative protein kinase
[Oryza sativa (japonica cultivar-group)]
gi|20042880|gb|AAM08708.1| Putative protein kinase [Oryza
sativa] gi|16924042|gb|AAL31654.1| Putative protein
kinase [Oryza sativa]
Length = 1098
Score = 768 bits (1983), Expect = 0.0
Identities = 436/1095 (39%), Positives = 631/1095 (56%), Gaps = 77/1095 (7%)
Query: 26 KLALLKWKDSFDDQSQTLLSTWKNNTNPCKPKWRGIKCDKSN-----FISTIGLANLGLK 80
++ALL WK + + S+W+ +T+PC W GI C ++ I+ I L + G+
Sbjct: 17 QMALLHWKSTLQSTGPQMRSSWQASTSPCN--WTGITCRAAHQAMSWVITNISLPDAGIH 74
Query: 81 GTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCTLT 140
G L L FSS P L ID+ +NS YG IP+ I +LS ++ L + N G +P E+ L
Sbjct: 75 GQLGELNFSSLPFLTYIDLSSNSVYGPIPSSISSLSALTYLDLQLNQLTGRMPDEISELQ 134
Query: 141 GLQFLDISFCKLNGAIPKSIGNLT------------------------NLSYLILGGNNW 176
L LD+S+ L G IP S+GNLT NL L L N
Sbjct: 135 RLTMLDLSYNNLTGHIPASVGNLTMITELSIHRNMVSGPIPKEIGMLANLQLLQLSNNTL 194
Query: 177 SG-----------------------GPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLT 213
SG GP+PP++ KL NL +LA+ + L G IP IG LT
Sbjct: 195 SGEIPTTLANLTNLDTFYLDGNELSGPVPPKLCKLTNLQYLALGDNKLTGEIPTCIGNLT 254
Query: 214 NLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIG 273
+ + L +N + G IP IGNL+ L LVL+ N K+ G +P L N++ L L+
Sbjct: 255 KMIKLYLFRNQIIGSIPPEIGNLAMLTDLVLNEN-KLKGSLPTELGNLTMLNNLFLHENQ 313
Query: 274 LSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNL 333
++GSIP + + NL+ L L N +SGSIP T+ +L LI L L N ++G IP GNL
Sbjct: 314 ITGSIPPGLGIISNLQNLILHSNQISGSIPGTLANLTKLIALDLSKNQINGSIPQEFGNL 373
Query: 334 INLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSEND 393
+NLQ+LS++EN ++G+IP S+GN + + +N+L +P NITN + ++ N
Sbjct: 374 VNLQLLSLEENQISGSIPKSLGNFQNMQNLNFRSNQLSNSLPQEFGNITNMVELDLASNS 433
Query: 394 FVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVY 453
G LP+ IC+G SL+LL N F GP+P SLKTC+S+ R+ L+ NQ+ GDI++ FGVY
Sbjct: 434 LSGQLPANICAGTSLKLLFLSLNMFNGPVPRSLKTCTSLVRLFLDGNQLTGDISKHFGVY 493
Query: 454 PKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQ 513
PKL+ + L N+ GQISP WG L I+ N I+G IP L L L LSSN
Sbjct: 494 PKLKKMSLMSNRLSGQISPKWGACPELAILNIAENMITGTIPPALSKLPNLVELKLSSNH 553
Query: 514 LTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKIPKELVE 573
+ G +P E+ G + +L+ L +S N S +IPS++G L+ L+ LD+ N LSG IP+EL
Sbjct: 554 VNGVIPPEI-GNLINLYSLNLSFNKLSGSIPSQLGNLRDLEYLDVSRNSLSGPIPEELGR 612
Query: 574 LPNLRMLNLSRNKIEGIIPIKFDSGLE---SLDLSGNFLKGNIPTGLADLVRLSKLNLSH 630
L++L ++ N G +P + LD+S N L G +P + L LNLSH
Sbjct: 613 CTKLQLLRINNNHFSGNLPATIGNLASIQIMLDVSNNKLDGLLPQDFGRMQMLVFLNLSH 672
Query: 631 NMLSGTIPQNFGR--NLVFVNISDNQLEGPLPKIPAFLSASFESLKNNNHLCGNIRGLDP 688
N +G IP +F +L ++ S N LEGPLP F +AS NN LCGN+ GL
Sbjct: 673 NQFTGRIPTSFASMVSLSTLDASYNNLEGPLPAGRLFQNASASWFLNNKGLCGNLSGLPS 732
Query: 689 CATSHSRKRKNVLR---PVFIALGAVILVLCVVGALMYIMCGRKKPNEESQTEEVQRGVL 745
C ++ ++ + R PV + LG IL V+G + + ++KP E + + +
Sbjct: 733 CYSAPGHNKRKLFRFLLPVVLVLGFAILATVVLGTV--FIHNKRKPQESTTAKGRD---M 787
Query: 746 FSIWSHDGKMMFENIIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEM 805
FS+W+ DG++ FE+I+ AT +FDDKY++G G G VY+A+L +G VVAVKKLH T+E +
Sbjct: 788 FSVWNFDGRLAFEDIVRATEDFDDKYIIGAGGYGKVYRAQLQDGQVVAVKKLH-TTEEGL 846
Query: 806 SCFSSKSFMSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQAVA 865
K F E+E LT I+ R+I+KL+GFCSH ++ FLVY+++E GSL L +D A A
Sbjct: 847 G--DEKRFSCEMEILTQIRQRSIVKLYGFCSHPEYRFLVYEYIEQGSLHMTLADDELAKA 904
Query: 866 FDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLKPG 925
DW+KR ++K VA AL YLHHDC+PPIIHRDI+S N+LL+ +A+VSDFGTA+ L+P
Sbjct: 905 LDWQKRNILIKDVAQALCYLHHDCNPPIIHRDITSNNILLDTTLKAYVSDFGTARILRPD 964
Query: 926 LHSWTQFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGKHPGDLISLFLSPSTRP 985
+W+ AGT+GY APEL+ T V EKCDVYSFG++ LE ++GKHP DL L T
Sbjct: 965 SSNWSALAGTYGYIAPELSYTSLVTEKCDVYSFGMVMLEVVIGKHPRDL----LQHLTSS 1020
Query: 986 MANNMLLTDVLDQRPQQVMEPIDEEVILIARLAFACLSQNPRLRPSMGQVCK-MLAIGKS 1044
+N+ + ++LD RP +E ++ + ++ F+CL +P+ RP+M +V + ++ S
Sbjct: 1021 RDHNITIKEILDSRPLAPTTTEEENIVSLIKVVFSCLKASPQARPTMQEVYQTLIDYQTS 1080
Query: 1045 PLVGKQLHMIRLEQL 1059
+ K + L++L
Sbjct: 1081 SFLSKNCSRVILDEL 1095
>gb|AAF79881.1| Contains similarity to receptor protein kinase-like protein from
Arabidopsis thaliana gb|AL161513. It contains a
eukaryotic protein kinase domain PF|00069. EST
gb|AI997574 comes from this gene
gi|15219699|ref|NP_174809.1| leucine-rich repeat
transmembrane protein kinase, putative [Arabidopsis
thaliana] gi|25518391|pir||B86479 hypothetical protein
F14D7.1 - Arabidopsis thaliana
Length = 1120
Score = 747 bits (1928), Expect = 0.0
Identities = 436/1109 (39%), Positives = 630/1109 (56%), Gaps = 96/1109 (8%)
Query: 6 TLIMILCVLPTLSVAEDSEAKLALLKWKDSFDDQSQTLLSTW----KNNTNPCKPKWRGI 61
++I+ + + ++AE + ALLKWK +F + S+ LS+W NT+ W G+
Sbjct: 18 SIILSCSISASATIAEAN----ALLKWKSTFTNSSK--LSSWVHDANTNTSFSCTSWYGV 71
Query: 62 KCDKSN--------------------FISTIGLANLGLKGTLHSLT----FSSFPNLLMI 97
C+ FIS LA + L L S T F + L+
Sbjct: 72 SCNSRGSIEELNLTNTGIEGTFQDFPFISLSNLAYVDLSMNLLSGTIPPQFGNLSKLIYF 131
Query: 98 DIRNNSFYGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAIP 157
D+ N G I +GNL N+++L NY IP E+ + + L +S KL G+IP
Sbjct: 132 DLSTNHLTGEISPSLGNLKNLTVLYLHQNYLTSVIPSELGNMESMTDLALSQNKLTGSIP 191
Query: 158 KSIGNLTNLSYLILGGN------------------------------------------- 174
S+GNL NL L L N
Sbjct: 192 SSLGNLKNLMVLYLYENYLTGVIPPELGNMESMTDLALSQNKLTGSIPSTLGNLKNLMVL 251
Query: 175 ----NWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIP 230
N+ G IPPEIG + ++ +LA+ ++ L GSIP +G L NL + L +N L+GGIP
Sbjct: 252 YLYENYLTGVIPPEIGNMESMTNLALSQNKLTGSIPSSLGNLKNLTLLSLFQNYLTGGIP 311
Query: 231 ETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKE 290
+GN+ + L LSNN K++G IP SL N+ +LT+LY L+G IP + N+ ++ +
Sbjct: 312 PKLGNIESMIDLELSNN-KLTGSIPSSLGNLKNLTILYLYENYLTGVIPPELGNMESMID 370
Query: 291 LALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTI 350
L L+ N L+GSIPS+ G+LKNL LYL N L+G IP +GN+ ++ L + +N LTG++
Sbjct: 371 LQLNNNKLTGSIPSSFGNLKNLTYLYLYLNYLTGVIPQELGNMESMINLDLSQNKLTGSV 430
Query: 351 PASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRL 410
P S GN L + N L G IP G+ N ++ + ++ N+F G P +C G L+
Sbjct: 431 PDSFGNFTKLESLYLRVNHLSGAIPPGVANSSHLTTLILDTNNFTGFFPETVCKGRKLQN 490
Query: 411 LNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQI 470
++ D+N GPIP SL+ C S+ R N+ GDI + FG+YP L ++D S NKFHG+I
Sbjct: 491 ISLDYNHLEGPIPKSLRDCKSLIRARFLGNKFTGDIFEAFGIYPDLNFIDFSHNKFHGEI 550
Query: 471 SPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLF 530
S NW KS L I+SNNNI+G IP + +T+L L LS+N L G+LP E +G + +L
Sbjct: 551 SSNWEKSPKLGALIMSNNNITGAIPTEIWNMTQLVELDLSTNNLFGELP-EAIGNLTNLS 609
Query: 531 DLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKIPKELVELPNLRMLNLSRNKIEGI 590
L+++ N S +P+ + L L+ LDL N S +IP+ L +NLSRNK +G
Sbjct: 610 RLRLNGNQLSGRVPAGLSFLTNLESLDLSSNNFSSEIPQTFDSFLKLHDMNLSRNKFDGS 669
Query: 591 IP-IKFDSGLESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSGTIPQNFGRNLVFVN 649
IP + + L LDLS N L G IP+ L+ L L KL+LSHN LSG IP F + N
Sbjct: 670 IPRLSKLTQLTQLDLSHNQLDGEIPSQLSSLQSLDKLDLSHNNLSGLIPTTFEGMIALTN 729
Query: 650 --ISDNQLEGPLPKIPAFLSASFESLKNNNHLCGNI--RGLDPCAT-SHSRKRKNVLRPV 704
IS+N+LEGPLP P F A+ ++L+ N LC NI + L PC +K N++ +
Sbjct: 730 VDISNNKLEGPLPDTPTFRKATADALEENIGLCSNIPKQRLKPCRELKKPKKNGNLVVWI 789
Query: 705 FIALGAVILVLCVVGALMYIMCGRKKPNEESQTEEVQRGVLFSIWSHDGKMMFENIIEAT 764
+ + V+++L + A + C RK+ + + + + G SI+S DGK +++IIE+T
Sbjct: 790 LVPILGVLVILSIC-ANTFTYCIRKRKLQNGRNTDPETGENMSIFSVDGKFKYQDIIEST 848
Query: 765 ANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSC-FSSKSFMSEIETLTGI 823
FD +L+G G VY+A L + ++AVK+LH DEE+S + F++E++ LT I
Sbjct: 849 NEFDPTHLIGTGGYSKVYRANLQD-TIIAVKRLHDTIDEEISKPVVKQEFLNEVKALTEI 907
Query: 824 KHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQAVAFDWEKRVNVVKGVANALS 883
+HRN++KL GFCSH + +FL+Y+++E GSL+++L ND +A W KR+NVVKGVA+ALS
Sbjct: 908 RHRNVVKLFGFCSHRRHTFLIYEYMEKGSLNKLLANDEEAKRLTWTKRINVVKGVAHALS 967
Query: 884 YLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLKPGLHSWTQFAGTFGYAAPEL 943
Y+HHD PI+HRDISS N+LL+ DY A +SDFGTAK LK +W+ AGT+GY APE
Sbjct: 968 YMHHDRITPIVHRDISSGNILLDNDYTAKISDFGTAKLLKTDSSNWSAVAGTYGYVAPEF 1027
Query: 944 AQTMEVNEKCDVYSFGVLALETIMGKHPGDLISLFLSPSTRPMANNMLLTDVLDQRPQQV 1003
A TM+V EKCDVYSFGVL LE I+GKHPGDL+S S S+ P + L + D+R +
Sbjct: 1028 AYTMKVTEKCDVYSFGVLILELIIGKHPGDLVS---SLSSSP-GEALSLRSISDERVLEP 1083
Query: 1004 MEPIDEEVILIARLAFACLSQNPRLRPSM 1032
E+++ + +A CL NP RP+M
Sbjct: 1084 RGQNREKLLKMVEMALLCLQANPESRPTM 1112
>ref|NP_192625.3| leucine-rich repeat family protein / protein kinase family protein
[Arabidopsis thaliana]
Length = 1009
Score = 699 bits (1805), Expect = 0.0
Identities = 408/953 (42%), Positives = 561/953 (58%), Gaps = 46/953 (4%)
Query: 4 LPTLIMILCVLP-TLSVAEDSEAKLALLKWKDSFDDQ-SQTLLSTWKN-NTNPCKPKWRG 60
L L++I VL + +V+ E ALLKWK +F +Q S + LS+W N NT+ W G
Sbjct: 28 LQVLLIISIVLSCSFAVSATVEEANALLKWKSTFTNQTSSSKLSSWVNPNTSSFCTSWYG 87
Query: 61 IKCDKSNFISTIGLANLGLKGTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISI 120
+ C + I + L N G++GT FSS PNL +D+ N F GTI G S
Sbjct: 88 VACSLGSIIR-LNLTNTGIEGTFEDFPFSSLPNLTFVDLSMNRFSGTISPLWGRFSK--- 143
Query: 121 LTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSIGNLTNLSYLILGGNNWSGGP 180
L++ D+S +L G IP +G+L+NL L L N +G
Sbjct: 144 ---------------------LEYFDLSINQLVGEIPPELGDLSNLDTLHLVENKLNGS- 181
Query: 181 IPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKLD 240
IP EIG+L + +AI + L G IP G LT L + L NSLSG IP IGNL L
Sbjct: 182 IPSEIGRLTKVTEIAIYDNLLTGPIPSSFGNLTKLVNLYLFINSLSGSIPSEIGNLPNLR 241
Query: 241 TLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSG 300
L L N ++G IP S N+ ++T+L LSG IP I N+ L L+L N L+G
Sbjct: 242 ELCLDRNN-LTGKIPSSFGNLKNVTLLNMFENQLSGEIPPEIGNMTALDTLSLHTNKLTG 300
Query: 301 SIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWL 360
IPST+G++K L L+L N L+G IP +G + ++ L + EN LTG +P S G L L
Sbjct: 301 PIPSTLGNIKTLAVLHLYLNQLNGSIPPELGEMESMIDLEISENKLTGPVPDSFGKLTAL 360
Query: 361 TVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTG 420
+ N+L G IP G+ N T V N+F G LP IC GG L L D N F G
Sbjct: 361 EWLFLRDNQLSGPIPPGIANSTELTVLQVDTNNFTGFLPDTICRGGKLENLTLDDNHFEG 420
Query: 421 PIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNL 480
P+P SL+ C S+ R+ + N GDI++ FGVYP L ++DLS+N FHGQ+S NW +S L
Sbjct: 421 PVPKSLRDCKSLIRVRFKGNSFSGDISEAFGVYPTLNFIDLSNNNFHGQLSANWEQSQKL 480
Query: 481 QTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFS 540
FI+SNN+I+G IP + +T+L L LSSN++TG+LP E + + + L+++ N S
Sbjct: 481 VAFILSNNSITGAIPPEIWNMTQLSQLDLSSNRITGELP-ESISNINRISKLQLNGNRLS 539
Query: 541 DNIPSEIGLLQRLQELDLGGNELSGKIPKELVELPNLRMLNLSRNKIEGIIP--IKFDSG 598
IPS I LL L+ LDL N S +IP L LP L +NLSRN ++ IP + S
Sbjct: 540 GKIPSGIRLLTNLEYLDLSSNRFSSEIPPTLNNLPRLYYMNLSRNDLDQTIPEGLTKLSQ 599
Query: 599 LESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSGTIPQNFGRNLVF--VNISDNQLE 656
L+ LDLS N L G I + L L +L+LSHN LSG IP +F L V++S N L+
Sbjct: 600 LQMLDLSYNQLDGEISSQFRSLQNLERLDLSHNNLSGQIPPSFKDMLALTHVDVSHNNLQ 659
Query: 657 GPLPKIPAFLSASFESLKNNNHLCGNI---RGLDPCATSHSRKR---KNVLRPVFIALGA 710
GP+P AF +A ++ + N LCG++ +GL PC+ + S+K +N++ + + +
Sbjct: 660 GPIPDNAAFRNAPPDAFEGNKDLCGSVNTTQGLKPCSITSSKKSHKDRNLIIYILVPIIG 719
Query: 711 VILVLCVVGALMYIMCGRKKPNE-ESQTEEVQRGVLFSIWSHDGKMMFENIIEATANFDD 769
I++L V + +C RK+ + E T+ G SI+S DGK+ ++ II+AT FD
Sbjct: 720 AIIILSVCAGI--FICFRKRTKQIEEHTDSESGGETLSIFSFDGKVRYQEIIKATGEFDP 777
Query: 770 KYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSK-SFMSEIETLTGIKHRNI 828
KYL+G G G VYKA+L ++AVKKL+ TD +S S+K F++EI LT I+HRN+
Sbjct: 778 KYLIGTGGHGKVYKAKLPNA-IMAVKKLNETTDSSISNPSTKQEFLNEIRALTEIRHRNV 836
Query: 829 IKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQAVAFDWEKRVNVVKGVANALSYLHHD 888
+KL GFCSH + +FLVY+++E GSL ++L ND +A DW KR+NVVKGVA+ALSY+HHD
Sbjct: 837 VKLFGFCSHRRNTFLVYEYMERGSLRKVLENDDEAKKLDWGKRINVVKGVAHALSYMHHD 896
Query: 889 CSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLKPGLHSWTQFAGTFGYAAP 941
SP I+HRDISS N+LL DYEA +SDFGTAK LKP +W+ AGT+GY AP
Sbjct: 897 RSPAIVHRDISSGNILLGEDYEAKISDFGTAKLLKPDSSNWSAVAGTYGYVAP 949
>dbj|BAC87845.1| leucine-rich repeat receptor-like protein kinase 1 [Populus nigra]
Length = 856
Score = 632 bits (1630), Expect = e-179
Identities = 350/736 (47%), Positives = 467/736 (62%), Gaps = 17/736 (2%)
Query: 335 NLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDF 394
NL +++ N+L GTIP+ I NL +T + N +G +P + N+T+ + + N+F
Sbjct: 119 NLLTPNLRNNSLYGTIPSHISNLTKITNLNLCHNHFNGSLPPEMNNLTHLMVLHLFSNNF 178
Query: 395 VGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYP 454
GHLP +C GG L A +N F+GPIP SL+ C+S+ R+ L+ NQ+ G+I++DFG+YP
Sbjct: 179 TGHLPRDLCLGGLLVNFTASYNHFSGPIPKSLRNCTSLFRVRLDWNQLTGNISEDFGLYP 238
Query: 455 KLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQL 514
L Y+DLS N +G+++ WG NL + +SNNNI+G IP + T L ++ LSSN L
Sbjct: 239 NLNYVDLSHNNLYGELTWKWGGFNNLTSLKLSNNNITGEIPSEIAKATGLQMIDLSSNLL 298
Query: 515 TGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKIPKELVEL 574
G +P E LG +K+L++L + NNH +P EI +L +L+ L+L N L G IPK+L E
Sbjct: 299 KGTIPKE-LGKLKALYNLTLHNNHLFGVVPFEIQMLSQLRALNLASNNLGGSIPKQLGEC 357
Query: 575 PNLRMLNLSRNKIEGIIP--IKFDSGLESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNM 632
NL LNLS NK G IP I F L LDLSGN L G IP+ + L +L +NLSHN
Sbjct: 358 SNLLQLNLSHNKFIGSIPSEIGFLHFLGDLDLSGNLLAGEIPSEIGQLKQLETMNLSHNK 417
Query: 633 LSGTIPQNFGR--NLVFVNISDNQLEGPLPKIPAFLSASFESLKNNNHLCGNIRGLDPCA 690
LSG IP F +L V+IS N+LEGP+PKI F+ A E+ NN+ LCGN GL PC
Sbjct: 418 LSGLIPTAFVDLVSLTTVDISYNELEGPIPKIKGFIEAPLEAFMNNSGLCGNANGLKPCT 477
Query: 691 TSHSRKRKN--VLRPVFIALGAVILVLCVVGALMYIMCGRKKPNEESQTEEVQRGVLFSI 748
SRK+ N V+ +F LG+++L+L +VG L + + S E Q + F +
Sbjct: 478 LLTSRKKSNKIVILILFPLLGSLLLLLIMVGCLYFHH--QTSRERISCLGERQSPLSFVV 535
Query: 749 WSHDGKMMFENIIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCF 808
W H+ +++ E II+A NF+ +G G G VY+A L G VVAVKK H D E+
Sbjct: 536 WGHEEEILHETIIQAANNFNFNNCIGKGGYGIVYRAMLPTGQVVAVKKFHPSRDGEL--M 593
Query: 809 SSKSFMSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQAVAFDW 868
+ ++F +EI L I+HRNI+KLHGFCS + SFLVY+F+E GSL L+++ Q + DW
Sbjct: 594 NLRTFRNEIRMLIDIRHRNIVKLHGFCSLIEHSFLVYEFIERGSLKMNLSSEEQVMDLDW 653
Query: 869 EKRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLKPGLHS 928
+R+NVVKGVA+ALSYLHHDCSPPIIHRDISS NVLL+ +YEAHVSDFGTA+ L P +
Sbjct: 654 NRRLNVVKGVASALSYLHHDCSPPIIHRDISSSNVLLDSEYEAHVSDFGTARLLMPDSTN 713
Query: 929 WTQFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGKHPGDLISL-----FLSPST 983
WT FAGT GY APELA TM VNEKCDVYSFGV+ +E IMG HPGDLIS F S S
Sbjct: 714 WTSFAGTLGYTAPELAYTMRVNEKCDVYSFGVVTMEVIMGMHPGDLISFLYASAFSSSSC 773
Query: 984 RPMANNMLLTDVLDQRPQQVMEPIDEEVILIARLAFACLSQNPRLRPSMGQVCKMLAIGK 1043
+ + LL DV+DQR + E V+ I ++AFACL NP+ RP+M QV L I +
Sbjct: 774 SQINQHALLKDVIDQRIPLPENRVAEGVVSIIKIAFACLLANPQSRPTMRQVASEL-IAR 832
Query: 1044 SPLVGKQLHMIRLEQL 1059
P + K I +E L
Sbjct: 833 WPPLPKSFSAITVEDL 848
Score = 254 bits (649), Expect = 1e-65
Identities = 156/437 (35%), Positives = 230/437 (51%), Gaps = 29/437 (6%)
Query: 8 IMILCVLPTL-----------------SVAEDSEAKLALLKWKDSFDDQ-SQTLLSTWKN 49
++++C++P+ AE +E ALLKW+ S DD SQ++LS+W
Sbjct: 19 LLLMCIIPSFFAFPSNSSATSFGAAKYEAAEGNEEAEALLKWRASLDDSHSQSVLSSWVG 78
Query: 50 NTNPCKPKWRGIKCDKSNFISTIGLANLGLKGTLHSLTFSSFPNLLMIDIRNNSFYGTIP 109
++ PCK W GI CD S ++ L + GL+GTLHS FSSFPNLL ++RNNS YGTIP
Sbjct: 79 SS-PCK--WLGITCDNSGSVANFSLPHFGLRGTLHSFNFSSFPNLLTPNLRNNSLYGTIP 135
Query: 110 AQIGNLSNISILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSI---GNLTNL 166
+ I NL+ I+ L +N+F+GS+P EM LT L L + G +P+ + G L N
Sbjct: 136 SHISNLTKITNLNLCHNHFNGSLPPEMNNLTHLMVLHLFSNNFTGHLPRDLCLGGLLVNF 195
Query: 167 SYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLS 226
+ N GPIP + +L + + + L G+I ++ G NL Y+DLS N+L
Sbjct: 196 T----ASYNHFSGPIPKSLRNCTSLFRVRLDWNQLTGNISEDFGLYPNLNYVDLSHNNLY 251
Query: 227 GGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLV 286
G + G + L +L LSNN ++G IP + + L ++ + L G+IP + L
Sbjct: 252 GELTWKWGGFNNLTSLKLSNN-NITGEIPSEIAKATGLQMIDLSSNLLKGTIPKELGKLK 310
Query: 287 NLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNL 346
L L L NHL G +P I L L L L SNNL G IP +G NL L++ N
Sbjct: 311 ALYNLTLHNNHLFGVVPFEIQMLSQLRALNLASNNLGGSIPKQLGECSNLLQLNLSHNKF 370
Query: 347 TGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGG 406
G+IP+ IG L +L +++ N L G IP+ + + + +S N G +P+
Sbjct: 371 IGSIPSEIGFLHFLGDLDLSGNLLAGEIPSEIGQLKQLETMNLSHNKLSGLIPTAFVDLV 430
Query: 407 SLRLLNADHNRFTGPIP 423
SL ++ +N GPIP
Sbjct: 431 SLTTVDISYNELEGPIP 447
Score = 168 bits (425), Expect = 1e-39
Identities = 111/353 (31%), Positives = 174/353 (48%), Gaps = 2/353 (0%)
Query: 190 NLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTK 249
NLL ++ ++L G+IP I LT + ++L N +G +P + NL+ L L L +N
Sbjct: 119 NLLTPNLRNNSLYGTIPSHISNLTKITNLNLCHNHFNGSLPPEMNNLTHLMVLHLFSNN- 177
Query: 250 MSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDL 309
+G +P L L SG IP S++N +L + LD N L+G+I G
Sbjct: 178 FTGHLPRDLCLGGLLVNFTASYNHFSGPIPKSLRNCTSLFRVRLDWNQLTGNISEDFGLY 237
Query: 310 KNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNK 369
NL + L NNL G + G NL L + NN+TG IP+ I L + ++++N
Sbjct: 238 PNLNYVDLSHNNLYGELTWKWGGFNNLTSLKLSNNNITGEIPSEIAKATGLQMIDLSSNL 297
Query: 370 LHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTC 429
L G IP L + + + N G +P +I LR LN N G IP L C
Sbjct: 298 LKGTIPKELGKLKALYNLTLHNNHLFGVVPFEIQMLSQLRALNLASNNLGGSIPKQLGEC 357
Query: 430 SSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNN 489
S++ ++ L N+ G I + G L LDLS N G+I G+ L+T +S+N
Sbjct: 358 SNLLQLNLSHNKFIGSIPSEIGFLHFLGDLDLSGNLLAGEIPSEIGQLKQLETMNLSHNK 417
Query: 490 ISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDN 542
+SG+IP F+ L L + +S N+L G +P ++ G +++ + ++N+ N
Sbjct: 418 LSGLIPTAFVDLVSLTTVDISYNELEGPIP-KIKGFIEAPLEAFMNNSGLCGN 469
Score = 112 bits (280), Expect = 7e-23
Identities = 79/240 (32%), Positives = 123/240 (50%), Gaps = 7/240 (2%)
Query: 449 DFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVLH 508
+F +P L +L +N +G I + + + +N+ +G +P + LT L VLH
Sbjct: 113 NFSSFPNLLTPNLRNNSLYGTIPSHISNLTKITNLNLCHNHFNGSLPPEMNNLTHLMVLH 172
Query: 509 LSSNQLTGKLPMEV-LGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKI 567
L SN TG LP ++ LGG+ L + S NHFS IP + L + L N+L+G I
Sbjct: 173 LFSNNFTGHLPRDLCLGGL--LVNFTASYNHFSGPIPKSLRNCTSLFRVRLDWNQLTGNI 230
Query: 568 PKELVELPNLRMLNLSRNKIEGIIPIKFD--SGLESLDLSGNFLKGNIPTGLADLVRLSK 625
++ PNL ++LS N + G + K+ + L SL LS N + G IP+ +A L
Sbjct: 231 SEDFGLYPNLNYVDLSHNNLYGELTWKWGGFNNLTSLKLSNNNITGEIPSEIAKATGLQM 290
Query: 626 LNLSHNMLSGTIPQNFGRNLVFVNIS--DNQLEGPLPKIPAFLSASFESLKNNNHLCGNI 683
++LS N+L GTIP+ G+ N++ +N L G +P LS +N+L G+I
Sbjct: 291 IDLSSNLLKGTIPKELGKLKALYNLTLHNNHLFGVVPFEIQMLSQLRALNLASNNLGGSI 350
Score = 89.7 bits (221), Expect = 5e-16
Identities = 63/201 (31%), Positives = 96/201 (47%), Gaps = 29/201 (14%)
Query: 57 KWRGIKCDKSNFISTIGLANLGLKGTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLS 116
KW G N ++++ L+N + G + S + L MID+ +N GTIP ++G L
Sbjct: 257 KWGGF-----NNLTSLKLSNNNITGEIPS-EIAKATGLQMIDLSSNLLKGTIPKELGKLK 310
Query: 117 NISILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSIGNLTNLSYLILGGNNW 176
+ LT NN+ G +P E+ L+ L+ L+++ L G+IPK +G +NL L L N +
Sbjct: 311 ALYNLTLHNNHLFGVVPFEIQMLSQLRALNLASNNLGGSIPKQLGECSNLLQLNLSHNKF 370
Query: 177 SG-----------------------GPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLT 213
G G IP EIG+L L + + + L G IP L
Sbjct: 371 IGSIPSEIGFLHFLGDLDLSGNLLAGEIPSEIGQLKQLETMNLSHNKLSGLIPTAFVDLV 430
Query: 214 NLAYIDLSKNSLSGGIPETIG 234
+L +D+S N L G IP+ G
Sbjct: 431 SLTTVDISYNELEGPIPKIKG 451
Score = 71.6 bits (174), Expect = 1e-10
Identities = 59/196 (30%), Positives = 87/196 (44%), Gaps = 10/196 (5%)
Query: 492 GVIPLDFIGLT-----KLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSE 546
G P ++G+T + L L G L +L + NN IPS
Sbjct: 78 GSSPCKWLGITCDNSGSVANFSLPHFGLRGTLHSFNFSSFPNLLTPNLRNNSLYGTIPSH 137
Query: 547 IGLLQRLQELDLGGNELSGKIPKELVELPNLRMLNLSRNKIEGIIPIKFDSG--LESLDL 604
I L ++ L+L N +G +P E+ L +L +L+L N G +P G L +
Sbjct: 138 ISNLTKITNLNLCHNHFNGSLPPEMNNLTHLMVLHLFSNNFTGHLPRDLCLGGLLVNFTA 197
Query: 605 SGNFLKGNIPTGLADLVRLSKLNLSHNMLSGTIPQNFG--RNLVFVNISDNQLEGPLP-K 661
S N G IP L + L ++ L N L+G I ++FG NL +V++S N L G L K
Sbjct: 198 SYNHFSGPIPKSLRNCTSLFRVRLDWNQLTGNISEDFGLYPNLNYVDLSHNNLYGELTWK 257
Query: 662 IPAFLSASFESLKNNN 677
F + + L NNN
Sbjct: 258 WGGFNNLTSLKLSNNN 273
>gb|AAP54208.1| putative protein kinase [Oryza sativa (japonica cultivar-group)]
gi|37535238|ref|NP_921921.1| putative protein kinase
[Oryza sativa (japonica cultivar-group)]
gi|13489172|gb|AAK27806.1| putative protein kinase [Oryza
sativa (japonica cultivar-group)]
Length = 1278
Score = 612 bits (1579), Expect = e-173
Identities = 354/884 (40%), Positives = 508/884 (57%), Gaps = 40/884 (4%)
Query: 93 NLLMIDIRNNSFYGTIPAQIGN-LSNISILTFKNNYFDGSIPQEMCTLTGLQFLDISFCK 151
N+ +D+ N+ +G IP + L N+ L N F G IP + LT LQ L ++
Sbjct: 213 NVTYLDLSQNTLFGKIPDTLPEKLPNLRYLNLSINAFSGPIPASLGKLTKLQDLRMAANN 272
Query: 152 LNGAIPKSIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGF 211
L G +P+ +G++ L L LG N GGPIPP +G+L L L I+ S L ++P ++G
Sbjct: 273 LTGGVPEFLGSMPQLRILELGDNQL-GGPIPPVLGQLQMLQRLDIKNSGLSSTLPSQLGN 331
Query: 212 LTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNT----------------------- 248
L NL + +LS N LSGG+P + + +S N
Sbjct: 332 LKNLIFFELSLNQLSGGLPPEFAGMRAMRYFGISTNNLTGEIPPVLFTSWPELISFQVQN 391
Query: 249 -KMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIG 307
++G IP L S L +LY +GSIP + L NL EL L +N L+G IPS+ G
Sbjct: 392 NSLTGKIPPELGKASKLNILYLFTNKFTGSIPAELGELENLTELDLSVNSLTGPIPSSFG 451
Query: 308 DLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVAT 367
+LK L KL L NNL+G IP IGN+ LQ L V N+L G +PA+I L+ L V
Sbjct: 452 NLKQLTKLALFFNNLTGVIPPEIGNMTALQSLDVNTNSLHGELPATITALRSLQYLAVFD 511
Query: 368 NKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLK 427
N + G IP L + N F G LP IC G +L L A++N FTG +P LK
Sbjct: 512 NHMSGTIPADLGKGLALQHVSFTNNSFSGELPRHICDGFALDHLTANYNNFTGALPPCLK 571
Query: 428 TCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISN 487
C+++ R+ LE N GDI++ FGV+PKL YLD+S NK G++S WG+ +NL +
Sbjct: 572 NCTALVRVRLEENHFTGDISEAFGVHPKLVYLDVSGNKLTGELSSAWGQCINLTLLHLDG 631
Query: 488 NNISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEI 547
N ISG IP F +T L L+L+ N LTG +P VLG ++ +F+L +S+N FS IP+ +
Sbjct: 632 NRISGGIPAAFGSMTSLKDLNLAGNNLTGGIP-PVLGNIR-VFNLNLSHNSFSGPIPASL 689
Query: 548 GLLQRLQELDLGGNELSGKIPKELVELPNLRMLNLSRNKIEGIIPIKFDSGLES---LDL 604
+LQ++D GN L G IP + +L L +L+LS+N++ G IP + + + LDL
Sbjct: 690 SNNSKLQKVDFSGNMLDGTIPVAISKLDALILLDLSKNRLSGEIPSELGNLAQLQILLDL 749
Query: 605 SGNFLKGNIPTGLADLVRLSKLNLSHNMLSGTIPQNFGR--NLVFVNISDNQLEGPLPKI 662
S N L G IP L L+ L +LNLSHN LSG+IP F R +L V+ S N+L G +P
Sbjct: 750 SSNSLSGAIPPNLEKLITLQRLNLSHNELSGSIPAGFSRMSSLESVDFSYNRLTGSIPSG 809
Query: 663 PAFLSASFESLKNNNHLCGNIRGLDPC----ATSHSRKRKNVLRPVFIALGAVILVLCVV 718
F +AS + N+ LCG+++GL PC S S K V+ +++ V+L+L VV
Sbjct: 810 NVFQNASASAYVGNSGLCGDVQGLTPCDISSTGSSSGHHKRVVIATVVSVVGVVLLLAVV 869
Query: 719 GALMYIMCGRKKPNEESQTEE-VQRGVLFSIWSHDGKMMFENIIEATANFDDKYLVGVGS 777
++ ++C R++P E+ + E +IW +GK F +I+ AT NF++ + +G G
Sbjct: 870 TCII-LLC-RRRPREKKEVESNTNYSYESTIWEKEGKFTFFDIVNATDNFNETFCIGKGG 927
Query: 778 QGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSEIETLTGIKHRNIIKLHGFCSH 837
G+VY+AELS G VVAVK+ H+ ++ + KSF +EI+ LT ++HRNI+KLHGFC+
Sbjct: 928 FGSVYRAELSSGQVVAVKRFHVADTGDIPDVNKKSFENEIKALTEVRHRNIVKLHGFCTS 987
Query: 838 SKFSFLVYKFLEGGSLDQILNNDTQAVAFDWEKRVNVVKGVANALSYLHHDCSPPIIHRD 897
+ +LVY++LE GSL + L + DW RV VV+G+A+AL+YLHHDC+P I+HRD
Sbjct: 988 GDYMYLVYEYLERGSLGKTLYGEEGKKKMDWGMRVKVVQGLAHALAYLHHDCNPAIVHRD 1047
Query: 898 ISSKNVLLNLDYEAHVSDFGTAKFLKPGLHSWTQFAGTFGYAAP 941
I+ N+LL D+E + DFGTAK L +WT AG++GY AP
Sbjct: 1048 ITVNNILLESDFEPRLCDFGTAKLLGGASTNWTSVAGSYGYMAP 1091
Score = 318 bits (816), Expect = 5e-85
Identities = 233/735 (31%), Positives = 347/735 (46%), Gaps = 69/735 (9%)
Query: 7 LIMILCVLPTLSVAEDSEAKLALLKWKDSFDDQSQTLLSTWKNNTNPCKPKWRGIKCDKS 66
L++++ V+ A +++A LL WK D + L S W C WRG+ CD +
Sbjct: 10 LLLLVVVVAAADAATEADA---LLAWKAGLQDGAAAL-SGWSRAAPVCA--WRGVACDAA 63
Query: 67 NF---ISTIGLANLGLKGTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISILTF 123
++++ L GL G L +L F++ P L +D+ N+F G IPA I L +++ L
Sbjct: 64 AGGARVTSLRLRGAGLGGGLDALDFAALPALAELDLNGNNFTGAIPASISRLRSLASLDL 123
Query: 124 KNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSIGNLTNLSYLILGGN--------- 174
NN F SIP ++ L+GL L + L GAIP + L +++ LG N
Sbjct: 124 GNNGFSDSIPPQLGDLSGLVDLRLYNNNLVGAIPHQLSRLPKVAHFDLGANYLTDEDFAK 183
Query: 175 --------------NWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEI-GFLTNLAYID 219
N G P I K N+ +L + ++ L G IP + L NL Y++
Sbjct: 184 FSPMPTVTFMSLYLNSFNGSFPEFILKSGNVTYLDLSQNTLFGKIPDTLPEKLPNLRYLN 243
Query: 220 LSKNSLSGGIPETIGNLSKLDTLVLSNN-----------------------TKMSGPIPH 256
LS N+ SG IP ++G L+KL L ++ N ++ GPIP
Sbjct: 244 LSINAFSGPIPASLGKLTKLQDLRMAANNLTGGVPEFLGSMPQLRILELGDNQLGGPIPP 303
Query: 257 SLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLY 316
L + L L N GLS ++P + NL NL L +N LSG +P ++ +
Sbjct: 304 VLGQLQMLQRLDIKNSGLSSTLPSQLGNLKNLIFFELSLNQLSGGLPPEFAGMRAMRYFG 363
Query: 317 LGSNNLSGPIP----ASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHG 372
+ +NNL+G IP S LI+ Q VQ N+LTG IP +G L + + TNK G
Sbjct: 364 ISTNNLTGEIPPVLFTSWPELISFQ---VQNNSLTGKIPPELGKASKLNILYLFTNKFTG 420
Query: 373 RIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSI 432
IP L + N +S N G +PS + L L N TG IP + +++
Sbjct: 421 SIPAELGELENLTELDLSVNSLTGPIPSSFGNLKQLTKLALFFNNLTGVIPPEIGNMTAL 480
Query: 433 ERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISG 492
+ + + N + G++ LQYL + DN G I + GK L LQ +NN+ SG
Sbjct: 481 QSLDVNTNSLHGELPATITALRSLQYLAVFDNHMSGTIPADLGKGLALQHVSFTNNSFSG 540
Query: 493 VIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQR 552
+P L L + N TG LP L +L +++ NHF+ +I G+ +
Sbjct: 541 ELPRHICDGFALDHLTANYNNFTGALP-PCLKNCTALVRVRLEENHFTGDISEAFGVHPK 599
Query: 553 LQELDLGGNELSGKIPKELVELPNLRMLNLSRNKIEGIIPIKFDS--GLESLDLSGNFLK 610
L LD+ GN+L+G++ + NL +L+L N+I G IP F S L+ L+L+GN L
Sbjct: 600 LVYLDVSGNKLTGELSSAWGQCINLTLLHLDGNRISGGIPAAFGSMTSLKDLNLAGNNLT 659
Query: 611 GNIPTGLADLVRLSKLNLSHNMLSGTIPQNFGRN--LVFVNISDNQLEGPLPKIPAFLSA 668
G IP L + +R+ LNLSHN SG IP + N L V+ S N L+G +P + L A
Sbjct: 660 GGIPPVLGN-IRVFNLNLSHNSFSGPIPASLSNNSKLQKVDFSGNMLDGTIPVAISKLDA 718
Query: 669 SFESLKNNNHLCGNI 683
+ N L G I
Sbjct: 719 LILLDLSKNRLSGEI 733
Score = 244 bits (622), Expect = 1e-62
Identities = 151/448 (33%), Positives = 236/448 (51%), Gaps = 4/448 (0%)
Query: 73 GLANLGLKGTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISILTFKNNYFDGSI 132
G++ L G + + F+S+P L+ ++NNS G IP ++G S ++IL N F GSI
Sbjct: 363 GISTNNLTGEIPPVLFTSWPELISFQVQNNSLTGKIPPELGKASKLNILYLFTNKFTGSI 422
Query: 133 PQEMCTLTGLQFLDISFCKLNGAIPKSIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLL 192
P E+ L L LD+S L G IP S GNL L+ L L NN + G IPPEIG + L
Sbjct: 423 PAELGELENLTELDLSVNSLTGPIPSSFGNLKQLTKLALFFNNLT-GVIPPEIGNMTALQ 481
Query: 193 HLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSG 252
L + ++L G +P I L +L Y+ + N +SG IP +G L + +NN+ SG
Sbjct: 482 SLDVNTNSLHGELPATITALRSLQYLAVFDNHMSGTIPADLGKGLALQHVSFTNNS-FSG 540
Query: 253 PIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNL 312
+P + + +L L + +G++P ++N L + L+ NH +G I G L
Sbjct: 541 ELPRHICDGFALDHLTANYNNFTGALPPCLKNCTALVRVRLEENHFTGDISEAFGVHPKL 600
Query: 313 IKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHG 372
+ L + N L+G + ++ G INL +L + N ++G IPA+ G++ L +A N L G
Sbjct: 601 VYLDVSGNKLTGELSSAWGQCINLTLLHLDGNRISGGIPAAFGSMTSLKDLNLAGNNLTG 660
Query: 373 RIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSI 432
IP L NI + +S N F G +P+ + + L+ ++ N G IP ++ ++
Sbjct: 661 GIPPVLGNI-RVFNLNLSHNSFSGPIPASLSNNSKLQKVDFSGNMLDGTIPVAISKLDAL 719
Query: 433 ERITLEVNQIEGDIAQDFGVYPKLQ-YLDLSDNKFHGQISPNWGKSLNLQTFIISNNNIS 491
+ L N++ G+I + G +LQ LDLS N G I PN K + LQ +S+N +S
Sbjct: 720 ILLDLSKNRLSGEIPSELGNLAQLQILLDLSSNSLSGAIPPNLEKLITLQRLNLSHNELS 779
Query: 492 GVIPLDFIGLTKLGVLHLSSNQLTGKLP 519
G IP F ++ L + S N+LTG +P
Sbjct: 780 GSIPAGFSRMSSLESVDFSYNRLTGSIP 807
Score = 95.1 bits (235), Expect = 1e-17
Identities = 50/118 (42%), Positives = 71/118 (59%), Gaps = 1/118 (0%)
Query: 942 ELAQTMEVNEKCDVYSFGVLALETIMGKHPGDLISLFLSPSTRPMANNMLLTDVLDQRPQ 1001
E A TM V EKCDVYSFGV+ALE +MGKHPGDL++ + S+ +++LL D+LDQR
Sbjct: 1157 EFAYTMRVTEKCDVYSFGVVALEVMMGKHPGDLLTSLPAISSSE-EDDLLLKDILDQRLD 1215
Query: 1002 QVMEPIDEEVILIARLAFACLSQNPRLRPSMGQVCKMLAIGKSPLVGKQLHMIRLEQL 1059
+ EEV+ I R+A C NP RPSM V + ++ + + +I + +L
Sbjct: 1216 APTGQLAEEVVFIVRIALGCTRVNPESRPSMRSVAQEISAHTQAYLSEPFKLITISKL 1273
>dbj|BAA96896.1| receptor-like protein kinase [Arabidopsis thaliana]
gi|15237562|ref|NP_201198.1| leucine-rich repeat
transmembrane protein kinase, putative [Arabidopsis
thaliana]
Length = 1102
Score = 576 bits (1484), Expect = e-162
Identities = 395/1104 (35%), Positives = 583/1104 (52%), Gaps = 82/1104 (7%)
Query: 6 TLIMILCVLPTLSVAEDSEAKLALLKWKDSFDDQSQTLLSTWKNNTNPCKPKWRGIKCDK 65
+L++IL + T + + + LL+ K F D Q L + N++ PC W G+ C
Sbjct: 14 SLLLILLISETTGLNLEGQY---LLEIKSKFVDAKQNLRNWNSNDSVPCG--WTGVMC-- 66
Query: 66 SNFIS-----TIGLANLGLKGTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISI 120
SN+ S ++ L+++ L G L S + +L +D+ N G IP +IGN S++ I
Sbjct: 67 SNYSSDPEVLSLNLSSMVLSGKL-SPSIGGLVHLKQLDLSYNGLSGKIPKEIGNCSSLEI 125
Query: 121 LTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSIGNLTNLSYLILGGNNWSG-- 178
L NN FDG IP E+ L L+ L I +++G++P IGNL +LS L+ NN SG
Sbjct: 126 LKLNNNQFDGEIPVEIGKLVSLENLIIYNNRISGSLPVEIGNLLSLSQLVTYSNNISGQL 185
Query: 179 ---------------------GPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAY 217
G +P EIG +L+ L + ++ L G +P+EIG L L+
Sbjct: 186 PRSIGNLKRLTSFRAGQNMISGSLPSEIGGCESLVMLGLAQNQLSGELPKEIGMLKKLSQ 245
Query: 218 IDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGS 277
+ L +N SG IP I N + L+TL L N ++ GPIP L ++ SL LY GL+G+
Sbjct: 246 VILWENEFSGFIPREISNCTSLETLALYKN-QLVGPIPKELGDLQSLEFLYLYRNGLNGT 304
Query: 278 IPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQ 337
IP I NL E+ N L+G IP +G+++ L LYL N L+G IP + L NL
Sbjct: 305 IPREIGNLSYAIEIDFSENALTGEIPLELGNIEGLELLYLFENQLTGTIPVELSTLKNLS 364
Query: 338 VLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGH 397
L + N LTG IP L+ L + ++ N L G IP L ++ +S+N G
Sbjct: 365 KLDLSINALTGPIPLGFQYLRGLFMLQLFQNSLSGTIPPKLGWYSDLWVLDMSDNHLSGR 424
Query: 398 LPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQ 457
+PS +C ++ +LN N +G IPT + TC ++ ++ L N + G + +
Sbjct: 425 IPSYLCLHSNMIILNLGTNNLSGNIPTGITTCKTLVQLRLARNNLVGRFPSNLCKQVNVT 484
Query: 458 YLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLTGK 517
++L N+F G I G LQ +++N +G +P + L++LG L++SSN+LTG+
Sbjct: 485 AIELGQNRFRGSIPREVGNCSALQRLQLADNGFTGELPREIGMLSQLGTLNISSNKLTGE 544
Query: 518 LPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKIPKELVELPNL 577
+P E+ K L L + N+FS +PSE+G L +L+ L L N LSG IP L L L
Sbjct: 545 VPSEIFN-CKMLQRLDMCCNNFSGTLPSEVGSLYQLELLKLSNNNLSGTIPVALGNLSRL 603
Query: 578 RMLNLSRNKIEGIIPIKFDS--GLE-SLDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLS 634
L + N G IP + S GL+ +L+LS N L G IP L++LV L L L++N LS
Sbjct: 604 TELQMGGNLFNGSIPRELGSLTGLQIALNLSYNKLTGEIPPELSNLVMLEFLLLNNNNLS 663
Query: 635 GTIPQNFGR--NLVFVNISDNQLEGPLPKIPAFLSASFESLKNNNHLCG----NIRGLDP 688
G IP +F +L+ N S N L GP IP + S S N LCG P
Sbjct: 664 GEIPSSFANLSSLLGYNFSYNSLTGP---IPLLRNISMSSFIGNEGLCGPPLNQCIQTQP 720
Query: 689 CATSHSRKRKNVLRPV-FIALGAVIL---VLCVVGALMYIMCGRKKP-------NEESQT 737
A S S + +R IA+ A ++ L ++ ++Y+M ++P ++ Q
Sbjct: 721 FAPSQSTGKPGGMRSSKIIAITAAVIGGVSLMLIALIVYLM---RRPVRTVASSAQDGQP 777
Query: 738 EEVQRGVLFSIWSHDGKMMFENIIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKL 797
E+ + F +G F++++ AT NFD+ ++VG G+ G VYKA L G +AVKKL
Sbjct: 778 SEMSLDIYFP--PKEG-FTFQDLVAATDNFDESFVVGRGACGTVYKAVLPAGYTLAVKKL 834
Query: 798 HLVTDEEMSCFSSKSFMSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQIL 857
+ + SF +EI TL I+HRNI+KLHGFC+H + L+Y+++ GSL +IL
Sbjct: 835 ASNHEGGNNNNVDNSFRAEILTLGNIRHRNIVKLHGFCNHQGSNLLLYEYMPKGSLGEIL 894
Query: 858 NNDTQAVAFDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFG 917
++ + DW KR + G A L+YLHHDC P I HRDI S N+LL+ +EAHV DFG
Sbjct: 895 HDPS--CNLDWSKRFKIALGAAQGLAYLHHDCKPRIFHRDIKSNNILLDDKFEAHVGDFG 952
Query: 918 TAKFL-KPGLHSWTQFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGKHP----- 971
AK + P S + AG++GY APE A TM+V EK D+YS+GV+ LE + GK P
Sbjct: 953 LAKVIDMPHSKSMSAIAGSYGYIAPEYAYTMKVTEKSDIYSYGVVLLELLTGKAPVQPID 1012
Query: 972 --GDLISLFLSPSTRPMANNMLLTDVLDQRPQQVMEPIDEEVILIARLAFACLSQNPRLR 1029
GD+++ S R + L + VLD R E I ++ + ++A C S +P R
Sbjct: 1013 QGGDVVNWVRSYIRR----DALSSGVLDARLTLEDERIVSHMLTVLKIALLCTSVSPVAR 1068
Query: 1030 PSMGQVCKMLAIGKSPLVGKQLHM 1053
PSM QV ML I G+Q H+
Sbjct: 1069 PSMRQVVLML-IESERSEGEQEHL 1091
>gb|AAD50027.1| Similar to leucine-rich receptor-like protein kinase [Arabidopsis
thaliana] gi|15220056|ref|NP_173166.1| leucine-rich
repeat family protein / protein kinase family protein
[Arabidopsis thaliana] gi|25518557|pir||E86308
hypothetical protein F20D23.7 - Arabidopsis thaliana
Length = 1133
Score = 569 bits (1467), Expect = e-160
Identities = 383/1088 (35%), Positives = 563/1088 (51%), Gaps = 65/1088 (5%)
Query: 8 IMILCVLPTLSVAEDSEAKLALLKWKDSFDDQSQTLLSTWKNNTNPCKPKWRGIKCDKSN 67
I+ILC + V +E LL++K +D + L S + ++NPC W GI C
Sbjct: 10 IVILCSFSFILVRSLNEEGRVLLEFKAFLNDSNGYLASWNQLDSNPCN--WTGIACTHLR 67
Query: 68 FISTIGLANLGLKGTLHSL--------------TFSSFP---------NLLMIDIRNNSF 104
++++ L + L GTL L F S P +L ++D+ N F
Sbjct: 68 TVTSVDLNGMNLSGTLSPLICKLHGLRKLNVSTNFISGPIPQDLSLCRSLEVLDLCTNRF 127
Query: 105 YGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSIGNLT 164
+G IP Q+ + + L NY GSIP+++ L+ LQ L I L G IP S+ L
Sbjct: 128 HGVIPIQLTMIITLKKLYLCENYLFGSIPRQIGNLSSLQELVIYSNNLTGVIPPSMAKLR 187
Query: 165 NLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNS 224
L + G N +SG IP EI +L L + ++ L GS+P+++ L NL + L +N
Sbjct: 188 QLRIIRAGRNGFSG-VIPSEISGCESLKVLGLAENLLEGSLPKQLEKLQNLTDLILWQNR 246
Query: 225 LSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQN 284
LSG IP ++GN+S+L+ L L N +G IP + ++ + LY L+G IP I N
Sbjct: 247 LSGEIPPSVGNISRLEVLALHENY-FTGSIPREIGKLTKMKRLYLYTNQLTGEIPREIGN 305
Query: 285 LVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQEN 344
L++ E+ N L+G IP G + NL L+L N L GPIP +G L L+ L + N
Sbjct: 306 LIDAAEIDFSENQLTGFIPKEFGHILNLKLLHLFENILLGPIPRELGELTLLEKLDLSIN 365
Query: 345 NLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICS 404
L GTIP + L +L ++ N+L G+IP + +N+ +S N G +P+ C
Sbjct: 366 RLNGTIPQELQFLPYLVDLQLFDNQLEGKIPPLIGFYSNFSVLDMSANSLSGPIPAHFCR 425
Query: 405 GGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDN 464
+L LL+ N+ +G IP LKTC S+ ++ L NQ+ G + + L L+L N
Sbjct: 426 FQTLILLSLGSNKLSGNIPRDLKTCKSLTKLMLGDNQLTGSLPIELFNLQNLTALELHQN 485
Query: 465 KFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLG 524
G IS + GK NL+ ++NNN +G IP + LTK+ ++SSNQLTG +P E LG
Sbjct: 486 WLSGNISADLGKLKNLERLRLANNNFTGEIPPEIGNLTKIVGFNISSNQLTGHIPKE-LG 544
Query: 525 GMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKIPKELVELPNLRMLNLSR 584
++ L +S N FS I E+G L L+ L L N L+G+IP +L L L L
Sbjct: 545 SCVTIQRLDLSGNKFSGYIAQELGQLVYLEILRLSDNRLTGEIPHSFGDLTRLMELQLGG 604
Query: 585 NKIEGIIPI---KFDSGLESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSGTIPQNF 641
N + IP+ K S SL++S N L G IP L +L L L L+ N LSG IP +
Sbjct: 605 NLLSENIPVELGKLTSLQISLNISHNNLSGTIPDSLGNLQMLEILYLNDNKLSGEIPASI 664
Query: 642 GR--NLVFVNISDNQLEGPLPKIPAFLSASFESLKNNNHLCGNIRGLDPCATSHS----- 694
G +L+ NIS+N L G +P F + N+ LC + R HS
Sbjct: 665 GNLMSLLICNISNNNLVGTVPDTAVFQRMDSSNFAGNHGLCNSQRSHCQPLVPHSDSKLN 724
Query: 695 -----RKRKNVLRPVFIALGAVILVLCVVGALMYIMCGRKKPNEESQTEEVQRGVLFSIW 749
+R+ +L I +G+V L+ +G I R++P + ++ + V+ S +
Sbjct: 725 WLINGSQRQKILTITCIVIGSVFLIT-FLGLCWTIK--RREPAFVALEDQTKPDVMDSYY 781
Query: 750 SHDGKMMFENIIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFS 809
++ +++AT NF + ++G G+ G VYKAE+S G V+AVKKL+ S
Sbjct: 782 FPKKGFTYQGLVDATRNFSEDVVLGRGACGTVYKAEMSGGEVIAVKKLN---SRGEGASS 838
Query: 810 SKSFMSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQAVAFDWE 869
SF +EI TL I+HRNI+KL+GFC H + L+Y+++ GSL + L + DW
Sbjct: 839 DNSFRAEISTLGKIRHRNIVKLYGFCYHQNSNLLLYEYMSKGSLGEQLQRGEKNCLLDWN 898
Query: 870 KRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLKPGL-HS 928
R + G A L YLHHDC P I+HRDI S N+LL+ ++AHV DFG AK + S
Sbjct: 899 ARYRIALGAAEGLCYLHHDCRPQIVHRDIKSNNILLDERFQAHVGDFGLAKLIDLSYSKS 958
Query: 929 WTQFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGKHP-------GDLISLFLSP 981
+ AG++GY APE A TM+V EKCD+YSFGV+ LE I GK P GDL++
Sbjct: 959 MSAVAGSYGYIAPEYAYTMKVTEKCDIYSFGVVLLELITGKPPVQPLEQGGDLVNW---- 1014
Query: 982 STRPMANNMLLT-DVLDQRPQQVMEPIDEEVILIARLAFACLSQNPRLRPSMGQVCKML- 1039
R NM+ T ++ D R + E+ L+ ++A C S +P RP+M +V M+
Sbjct: 1015 -VRRSIRNMIPTIEMFDARLDTNDKRTVHEMSLVLKIALFCTSNSPASRPTMREVVAMIT 1073
Query: 1040 -AIGKSPL 1046
A G S L
Sbjct: 1074 EARGSSSL 1081
>ref|XP_476665.1| putative LRR receptor-like kinase [Oryza sativa (japonica
cultivar-group)] gi|34395052|dbj|BAC84715.1| putative LRR
receptor-like kinase [Oryza sativa (japonica
cultivar-group)]
Length = 1109
Score = 562 bits (1449), Expect = e-158
Identities = 361/1089 (33%), Positives = 559/1089 (51%), Gaps = 66/1089 (6%)
Query: 2 MVLPTLIMILCVLPTLSVAEDSEAKLALLKWKDSFDDQSQTLLSTW-----KNNTNPCKP 56
++L + V + + + A AL+++K DD L S+W +PC
Sbjct: 8 VLLAAAVFFAAVAAAAAASSSAAAVAALMEFKTKLDDVDGRL-SSWDAAGGSGGGDPCG- 65
Query: 57 KWRGIKCDKSNFISTIGLANLGLKGTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLS 116
W GI C + ++ + L L L G L S + P L ++++ N+ G +P +
Sbjct: 66 -WPGIACSAAMEVTAVTLHGLNLHGEL-SAAVCALPRLAVLNVSKNALAGALPPGLAACR 123
Query: 117 NISILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSIGNLTNLSYLILGGNNW 176
+ +L N G IP +C+L L+ L +S L+G IP +IGNLT L L + NN
Sbjct: 124 ALEVLDLSTNSLHGGIPPSLCSLPSLRQLFLSENFLSGEIPAAIGNLTALEELEIYSNNL 183
Query: 177 SGG-----------------------PIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLT 213
+GG PIP EI +L L + ++NL G +P E+ L
Sbjct: 184 TGGIPTTIAALQRLRIIRAGLNDLSGPIPVEISACASLAVLGLAQNNLAGELPGELSRLK 243
Query: 214 NLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIG 273
NL + L +N+LSG IP +G++ L+ L L++N +G +P L + SL LY
Sbjct: 244 NLTTLILWQNALSGEIPPELGDIPSLEMLALNDNA-FTGGVPRELGALPSLAKLYIYRNQ 302
Query: 274 LSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNL 333
L G+IP + +L + E+ L N L+G IP +G + L LYL N L G IP +G L
Sbjct: 303 LDGTIPRELGDLQSAVEIDLSENKLTGVIPGELGRIPTLRLLYLFENRLQGSIPPELGEL 362
Query: 334 INLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSEND 393
++ + + NNLTGTIP NL L ++ N++HG IP L +N +S+N
Sbjct: 363 TVIRRIDLSINNLTGTIPMEFQNLTDLEYLQLFDNQIHGVIPPMLGAGSNLSVLDLSDNR 422
Query: 394 FVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVY 453
G +P +C L L+ NR G IP +K C ++ ++ L N + G + + +
Sbjct: 423 LTGSIPPHLCKFQKLIFLSLGSNRLIGNIPPGVKACRTLTQLQLGGNMLTGSLPVELSLL 482
Query: 454 PKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQ 513
L LD++ N+F G I P GK +++ I+S N G IP LTKL ++SSNQ
Sbjct: 483 RNLSSLDMNRNRFSGPIPPEIGKFRSIERLILSENYFVGQIPPGIGNLTKLVAFNISSNQ 542
Query: 514 LTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKIPKELVE 573
LTG +P E L L L +S N + IP E+G L L++L L N L+G +P
Sbjct: 543 LTGPIPRE-LARCTKLQRLDLSKNSLTGVIPQELGTLVNLEQLKLSDNSLNGTVPSSFGG 601
Query: 574 LPNLRMLNLSRNKIEGIIPIKFD--SGLE-SLDLSGNFLKGNIPTGLADLVRLSKLNLSH 630
L L L + N++ G +P++ + L+ +L++S N L G IPT L +L L L L++
Sbjct: 602 LSRLTELQMGGNRLSGQLPVELGQLTALQIALNVSYNMLSGEIPTQLGNLHMLEFLYLNN 661
Query: 631 NMLSGTIPQNFGR--NLVFVNISDNQLEGPLPKIPAFLSASFESLKNNNHLCGNIRGLDP 688
N L G +P +FG +L+ N+S N L GPLP F + NN LCG I+G
Sbjct: 662 NELEGEVPSSFGELSSLLECNLSYNNLAGPLPSTTLFQHMDSSNFLGNNGLCG-IKGKSC 720
Query: 689 CATSHSR--------KRKNVLRPVFIALGAVILVLCVVGALMYIMCG--RKKPNEESQTE 738
S S ++K +LR I++ ++++ V L+ ++C + K + E
Sbjct: 721 SGLSGSAYASREAAVQKKRLLREKIISISSIVIAF-VSLVLIAVVCWSLKSKIPDLVSNE 779
Query: 739 EVQRGVLFSIWSHDGKMMFENIIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLH 798
E + G + ++ F+ +++ T +F + ++G G+ G VYKA + +G VAVKKL
Sbjct: 780 ERKTGFSGPHYFLKERITFQELMKVTDSFSESAVIGRGACGTVYKAIMPDGRRVAVKKLK 839
Query: 799 LVTDEEMSCFSSKSFMSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILN 858
+ +SF +EI TL ++HRNI+KL+GFCS+ + ++Y+++ GSL ++L+
Sbjct: 840 C---QGEGSNVDRSFRAEITTLGNVRHRNIVKLYGFCSNQDCNLILYEYMANGSLGELLH 896
Query: 859 NDTQAVAFDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGT 918
DW+ R + G A L YLH DC P +IHRDI S N+LL+ EAHV DFG
Sbjct: 897 GSKDVCLLDWDTRYRIALGAAEGLRYLHSDCKPKVIHRDIKSNNILLDEMMEAHVGDFGL 956
Query: 919 AKFLK-PGLHSWTQFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGKHP------ 971
AK + + + AG++GY APE A TM+V EKCD+YSFGV+ LE + G+ P
Sbjct: 957 AKLIDISNSRTMSAIAGSYGYIAPEYAFTMKVTEKCDIYSFGVVLLELVTGQSPIQPLEQ 1016
Query: 972 -GDLISLFLSPSTRPMANNMLLTDVLDQRPQQVMEPIDEEVILIARLAFACLSQNPRLRP 1030
GDL++L + N+ + L+ ++V+ EE+ L+ ++A C S++P RP
Sbjct: 1017 GGDLVNLVRRMTNSSTTNSEIFDSRLNLNSRRVL----EEISLVLKIALFCTSESPLDRP 1072
Query: 1031 SMGQVCKML 1039
SM +V ML
Sbjct: 1073 SMREVISML 1081
>emb|CAD41180.1| OSJNBb0002J11.4 [Oryza sativa (japonica cultivar-group)]
gi|32490277|emb|CAE05566.1| OSJNBb0116K07.19 [Oryza
sativa (japonica cultivar-group)]
gi|50926296|ref|XP_473095.1| OSJNBb0116K07.19 [Oryza
sativa (japonica cultivar-group)]
Length = 1104
Score = 550 bits (1417), Expect = e-155
Identities = 370/1050 (35%), Positives = 548/1050 (51%), Gaps = 74/1050 (7%)
Query: 44 LSTWKNNTNPCKPKWRGIKCDKSNF--ISTIGLANLGLKGTLHSLTFSSFPNLLMIDIRN 101
L W N +P W+G+ C + + ++ L+N+ L GT+ + L +D+
Sbjct: 51 LDDW-NPEDPSPCGWKGVNCSSGSTPAVVSLNLSNMNLSGTVDP-SIGGLAELTNLDLSF 108
Query: 102 NSFYGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSIG 161
N F GTIPA+IGN S ++ L NN F G+IP E+ L + ++ KL GAIP IG
Sbjct: 109 NGFSGTIPAEIGNCSKLTGLNLNNNQFQGTIPAELGKLAMMITFNLCNNKLFGAIPDEIG 168
Query: 162 NLTNLSYLILGGNNWSG-----------------------GPIPPEIGKLNNLLHLAIQK 198
N+ +L L+ NN SG G IP EIG+ NL+ + +
Sbjct: 169 NMASLEDLVGYSNNLSGSIPHTIGRLKNLKTVRLGQNAISGNIPVEIGECLNLVVFGLAQ 228
Query: 199 SNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSL 258
+ L G +P+EIG LTN+ + L N LS IP IGN L T+ L +N + GPIP ++
Sbjct: 229 NKLGGPLPKEIGKLTNMTDLILWGNQLSSVIPPEIGNCINLRTIALYDNN-LVGPIPATI 287
Query: 259 WNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLG 318
N+ +L LY L+G+IP I NL +E+ N L+G +P G + L LYL
Sbjct: 288 GNIQNLQRLYLYRNLLNGTIPLEIGNLSLAEEIDFSENVLTGGVPKEFGKIPRLYLLYLF 347
Query: 319 SNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPN-- 376
N L+GPIP + L NL L + N L+G IPA + L ++ N L G IP
Sbjct: 348 QNQLTGPIPTELCVLRNLSKLDLSINTLSGPIPACFQYMSRLIQLQLFNNMLSGDIPPRF 407
Query: 377 GLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERIT 436
G+Y+ + F S N+ G +P +C +L LLN N+ G IP + +C S+ ++
Sbjct: 408 GIYSRLWVVDF--SNNNITGQIPRDLCRQSNLILLNLGANKLIGNIPHGITSCKSLVQLR 465
Query: 437 LEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPL 496
L N + G D L ++L NKF+G I P G +LQ ++NN + +P
Sbjct: 466 LADNSLTGSFPTDLCNLVNLTTIELGRNKFNGPIPPQIGNCKSLQRLDLTNNYFTSELPQ 525
Query: 497 DFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQEL 556
+ L+KL V ++SSN+L G +P+E+ L L +S N F ++P+E+G L +L+ L
Sbjct: 526 EIGNLSKLVVFNISSNRLGGSIPLEIFN-CTMLQRLDLSQNSFEGSLPNEVGSLPQLELL 584
Query: 557 DLGGNELSGKIPKELVELPNLRMLNLSRNKIEGIIPIKFD--SGLE-SLDLSGNFLKGNI 613
N LSG+IP L +L +L L + N+ G IP + S L+ +++LS N L GNI
Sbjct: 585 SFADNRLSGEIPPILGKLSHLTALQIGGNQFSGGIPKELGLLSSLQIAMNLSYNNLSGNI 644
Query: 614 PTGLADLVRLSKLNLSHNMLSGTIPQNFGR--NLVFVNISDNQLEGPLPKIPAFLSASFE 671
P+ L +L L L L++N L+G IP F +L+ N+S N L G LP IP F + +
Sbjct: 645 PSELGNLALLENLFLNNNKLTGEIPDTFANLSSLLEFNVSYNNLTGALPTIPLFDNMAST 704
Query: 672 SLKNNNHLCGNIRGLDPCATSHSRKRKN--------VLRPVFIALGAVILVLCVVGALMY 723
S N LCG G + S + N V+ V +G + L+L V+ ++Y
Sbjct: 705 SFLGNKGLCGGQLGKCGSESISSSQSSNSGSPPLGKVIAIVAAVIGGISLILIVI--IVY 762
Query: 724 IMCGRKKPNEESQTEEVQRGVLFSIWSH-----DGKMMFENIIEATANFDDKYLVGVGSQ 778
M +KP E +Q +FS S+ F+ ++ AT NFD+ ++G G+
Sbjct: 763 HM---RKPLET--VAPLQDKQIFSAGSNMQVSTKDAYTFQELVSATNNFDESCVIGRGAC 817
Query: 779 GNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSEIETLTGIKHRNIIKLHGFCSHS 838
G VY+A L G +AVKKL + SF +EI TL I+HRNI+KL+GF H
Sbjct: 818 GTVYRAILKAGQTIAVKKL---ASNREGSNTDNSFRAEILTLGKIRHRNIVKLYGFIYHQ 874
Query: 839 KFSFLVYKFLEGGSLDQILNNDTQAVAFDWEKRVNVVKGVANALSYLHHDCSPPIIHRDI 898
+ L+Y+++ GSL ++L+ + + + DWE R + G A LSYLHHDC P IIHRDI
Sbjct: 875 GSNLLLYEYMPRGSLGELLHGQSSS-SLDWETRFMIALGSAEGLSYLHHDCKPRIIHRDI 933
Query: 899 SSKNVLLNLDYEAHVSDFGTAKFL-KPGLHSWTQFAGTFGYAAPELAQTMEVNEKCDVYS 957
S N+LL+ ++EAHV DFG AK + P S + AG++GY APE A TM+V EK D+YS
Sbjct: 934 KSNNILLDENFEAHVGDFGLAKVIDMPYSKSMSAIAGSYGYIAPEYAYTMKVTEKSDIYS 993
Query: 958 FGVLALETIMGKHP-------GDLISLFLSPSTRPMANNMLLTDVLDQRPQQVMEPIDEE 1010
+GV+ LE + G+ P GDL++ + +N L +LD+ + +
Sbjct: 994 YGVVLLELLTGRAPVQPLELGGDLVTWV----KNYIRDNSLGPGILDKNLNLEDKTSVDH 1049
Query: 1011 VILIARLAFACLSQNPRLRPSMGQVCKMLA 1040
+I + ++A C S +P RP M V ML+
Sbjct: 1050 MIEVLKIALLCTSMSPYDRPPMRNVVVMLS 1079
>sp|P93194|RPK1_IPONI Receptor-like protein kinase precursor gi|14495542|gb|AAB36558.2|
receptor-like protein kinase INRPK1 [Ipomoea nil]
Length = 1109
Score = 545 bits (1403), Expect = e-153
Identities = 379/1120 (33%), Positives = 556/1120 (48%), Gaps = 120/1120 (10%)
Query: 6 TLIMILCVLPTLSVA----EDSEAKLALLK-WKDSFDDQSQTLLSTWK-NNTNPCKPKWR 59
T ++ LC ++ A D A L+L + W D +Q+ W +++ PC W
Sbjct: 7 TFLLFLCSTSSIYAAFALNSDGAALLSLTRHWTSIPSDITQS----WNASDSTPCS--WL 60
Query: 60 GIKCDKSNFISTIGLANLGLKGTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNIS 119
G++CD+ F+ T+ L++ G+ G S +L + + N F+G+IP+Q+GN S +
Sbjct: 61 GVECDRRQFVDTLNLSSYGISGEFGP-EISHLKHLKKVVLSGNGFFGSIPSQLGNCSLLE 119
Query: 120 ILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKL------------------------NGA 155
+ +N F G+IP + L L+ L + F L NG+
Sbjct: 120 HIDLSSNSFTGNIPDTLGALQNLRNLSLFFNSLIGPFPESLLSIPHLETVYFTGNGLNGS 179
Query: 156 IPKSIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNL 215
IP +IGN++ L+ L L N +SG P+P +G + L L + +NLVG++P + L NL
Sbjct: 180 IPSNIGNMSELTTLWLDDNQFSG-PVPSSLGNITTLQELYLNDNNLVGTLPVTLNNLENL 238
Query: 216 AYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLS 275
Y+D+ NSL G IP + ++DT+ LSNN + +G +P L N +SL + LS
Sbjct: 239 VYLDVRNNSLVGAIPLDFVSCKQIDTISLSNN-QFTGGLPPGLGNCTSLREFGAFSCALS 297
Query: 276 GSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLIN 335
G IP L L L L NH SG IP +G K++I L L N L G IP +G L
Sbjct: 298 GPIPSCFGQLTKLDTLYLAGNHFSGRIPPELGKCKSMIDLQLQQNQLEGEIPGELGMLSQ 357
Query: 336 LQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFV 395
LQ L + NNL+G +P SI ++ L ++ N L G +P + + +S + EN F
Sbjct: 358 LQYLHLYTNNLSGEVPLSIWKIQSLQSLQLYQNNLSGELPVDMTELKQLVSLALYENHFT 417
Query: 396 GHLPSQICSGGSLRLLNADHNRFTGPIPTSLKT------------------------CSS 431
G +P + + SL +L+ N FTG IP +L + CS+
Sbjct: 418 GVIPQDLGANSSLEVLDLTRNMFTGHIPPNLCSQKKLKRLLLGYNYLEGSVPSDLGGCST 477
Query: 432 IERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNIS 491
+ER+ LE N + G + DF L + DLS N F G I P+ G N+ +S+N +S
Sbjct: 478 LERLILEENNLRGGLP-DFVEKQNLLFFDLSGNNFTGPIPPSLGNLKNVTAIYLSSNQLS 536
Query: 492 GVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQ 551
G IP + L KL L+LS N L G LP E L L +L S+N + +IPS +G L
Sbjct: 537 GSIPPELGSLVKLEHLNLSHNILKGILPSE-LSNCHKLSELDASHNLLNGSIPSTLGSLT 595
Query: 552 RLQELDLGGNELSGKIPKELVELPNLRMLNLSRNKIEGIIP-IKFDSGLESLDLSGNFLK 610
L +L LG N SG IP L + L L L N + G IP + L SL+LS N L
Sbjct: 596 ELTKLSLGENSFSGGIPTSLFQSNKLLNLQLGGNLLAGDIPPVGALQALRSLNLSSNKLN 655
Query: 611 GNIPTGLADLVRLSKLNLSHNMLSGTIPQ-NFGRNLVFVNISDNQLEGPLP-KIPAFLSA 668
G +P L L L +L++SHN LSGT+ + ++L F+NIS N GP+P + FL++
Sbjct: 656 GQLPIDLGKLKMLEELDVSHNNLSGTLRVLSTIQSLTFINISHNLFSGPVPPSLTKFLNS 715
Query: 669 SFESLKNNNHLCGNIRG----------LDPC--ATSHSRKRKNVLRPVFIALGAVILVLC 716
S S N+ LC N L PC ++ + + L I LGA++ ++C
Sbjct: 716 SPTSFSGNSDLCINCPADGLACPESSILRPCNMQSNTGKGGLSTLGIAMIVLGALLFIIC 775
Query: 717 VV--GALMYIMCGRKKPNEESQTEEVQRGVLFSIWSHDGKMMFENIIEATANFDDKYLVG 774
+ A +++ C KK +E + S DG ++ ++EAT N +DKY++G
Sbjct: 776 LFLFSAFLFLHC--KKSVQE---------IAISAQEGDGSLL-NKVLEATENLNDKYVIG 823
Query: 775 VGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSEIETLTGIKHRNIIKLHGF 834
G+ G +YKA LS V AVKKL + S S + EIET+ ++HRN+IKL F
Sbjct: 824 KGAHGTIYKATLSPDKVYAVKKLVFTGIKN----GSVSMVREIETIGKVRHRNLIKLEEF 879
Query: 835 CSHSKFSFLVYKFLEGGSLDQILNNDTQAVAFDWEKRVNVVKGVANALSYLHHDCSPPII 894
++ ++Y ++E GSL IL+ DW R N+ G A+ L+YLH DC P I+
Sbjct: 880 WLRKEYGLILYTYMENGSLHDILHETNPPKPLDWSTRHNIAVGTAHGLAYLHFDCDPAIV 939
Query: 895 HRDISSKNVLLNLDYEAHVSDFGTAKFLKPGLHS--WTQFAGTFGYAAPELAQTMEVNEK 952
HRDI N+LL+ D E H+SDFG AK L S GT GY APE A T + +
Sbjct: 940 HRDIKPMNILLDSDLEPHISDFGIAKLLDQSATSIPSNTVQGTIGYMAPENAFTTVKSRE 999
Query: 953 CDVYSFGVLALETIMGKH--------PGDLISLFLSPST-----RPMANNMLLTDVLDQR 999
DVYS+GV+ LE I K D++ S T + + + LL +++D
Sbjct: 1000 SDVYSYGVVLLELITRKKALDPSFNGETDIVGWVRSVWTQTGEIQKIVDPSLLDELID-- 1057
Query: 1000 PQQVMEPIDEEVILIARLAFACLSQNPRLRPSMGQVCKML 1039
VME + E + LA C + RP+M V K L
Sbjct: 1058 -SSVMEQVTEAL----SLALRCAEKEVDKRPTMRDVVKQL 1092
>gb|AAC04906.1| putative receptor-like protein kinase [Arabidopsis thaliana]
gi|25408249|pir||B84742 probable receptor-like protein
kinase [imported] - Arabidopsis thaliana
gi|15225805|ref|NP_180875.1| leucine-rich repeat
transmembrane protein kinase, putative [Arabidopsis
thaliana]
Length = 1124
Score = 533 bits (1372), Expect = e-149
Identities = 365/1095 (33%), Positives = 567/1095 (51%), Gaps = 76/1095 (6%)
Query: 7 LIMILCVLPTLSVAEDSEAKLALLKWKDSFDDQSQTLLSTWKN-NTNPCKPKWRGIKCDK 65
++ +L +L S + +S+ + LL+ K+ S L W + PC W G+ C
Sbjct: 19 VLFLLTLLVWTSESLNSDGQF-LLELKNRGFQDSLNRLHNWNGIDETPCN--WIGVNCSS 75
Query: 66 --------SNFISTIGLANLGLKGTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSN 117
S ++++ L+++ L G + S + NL+ +++ N+ G IP +IGN S
Sbjct: 76 QGSSSSSNSLVVTSLDLSSMNLSGIV-SPSIGGLVNLVYLNLAYNALTGDIPREIGNCSK 134
Query: 118 ISILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSIGNLTNLSYLILGGNNWS 177
+ ++ NN F GSIP E+ L+ L+ +I KL+G +P+ IG+L NL L+ NN +
Sbjct: 135 LEVMFLNNNQFGGSIPVEINKLSQLRSFNICNNKLSGPLPEEIGDLYNLEELVAYTNNLT 194
Query: 178 GGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLS 237
G P+P +G LN L +++ G+IP EIG NL + L++N +SG +P+ IG L
Sbjct: 195 G-PLPRSLGNLNKLTTFRAGQNDFSGNIPTEIGKCLNLKLLGLAQNFISGELPKEIGMLV 253
Query: 238 KLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINH 297
KL ++L N K SG IP + N++SL L L G IP I N+ +LK+L L N
Sbjct: 254 KLQEVILWQN-KFSGFIPKDIGNLTSLETLALYGNSLVGPIPSEIGNMKSLKKLYLYQNQ 312
Query: 298 LSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNL 357
L+G+IP +G L ++++ N LSG IP + + L++L + +N LTG IP + L
Sbjct: 313 LNGTIPKELGKLSKVMEIDFSENLLSGEIPVELSKISELRLLYLFQNKLTGIIPNELSKL 372
Query: 358 KWLTVFEVATNKLHGRIPNGLYNIT---------NWISFVV---------------SEND 393
+ L +++ N L G IP G N+T N +S V+ SEN
Sbjct: 373 RNLAKLDLSINSLTGPIPPGFQNLTSMRQLQLFHNSLSGVIPQGLGLYSPLWVVDFSENQ 432
Query: 394 FVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVY 453
G +P IC +L LLN NR G IP + C S+ ++ + N++ G +
Sbjct: 433 LSGKIPPFICQQSNLILLNLGSNRIFGNIPPGVLRCKSLLQLRVVGNRLTGQFPTELCKL 492
Query: 454 PKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQ 513
L ++L N+F G + P G LQ ++ N S +P + L+ L ++SSN
Sbjct: 493 VNLSAIELDQNRFSGPLPPEIGTCQKLQRLHLAANQFSSNLPNEISKLSNLVTFNVSSNS 552
Query: 514 LTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKIPKELVE 573
LTG +P E+ K L L +S N F ++P E+G L +L+ L L N SG IP +
Sbjct: 553 LTGPIPSEI-ANCKMLQRLDLSRNSFIGSLPPELGSLHQLEILRLSENRFSGNIPFTIGN 611
Query: 574 LPNLRMLNLSRNKIEGIIPIKFD--SGLE-SLDLSGNFLKGNIPTGLADLVRLSKLNLSH 630
L +L L + N G IP + S L+ +++LS N G IP + +L L L+L++
Sbjct: 612 LTHLTELQMGGNLFSGSIPPQLGLLSSLQIAMNLSYNDFSGEIPPEIGNLHLLMYLSLNN 671
Query: 631 NMLSGTIPQNFGR--NLVFVNISDNQLEGPLPKIPAFLSASFESLKNNNHLCG-NIRGLD 687
N LSG IP F +L+ N S N L G LP F + + S N LCG ++R D
Sbjct: 672 NHLSGEIPTTFENLSSLLGCNFSYNNLTGQLPHTQIFQNMTLTSFLGNKGLCGGHLRSCD 731
Query: 688 PCATSH---------SRKRKNVLRPVFIALGAV-ILVLCVVGALMYIMCGRKKPNEESQT 737
P +S S +R ++ V +G + +L++ +V + P +
Sbjct: 732 PSHSSWPHISSLKAGSARRGRIIIIVSSVIGGISLLLIAIVVHFLRNPVEPTAPYVHDKE 791
Query: 738 EEVQRGVLFSIWSHDGKMMFENIIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKL 797
Q ++ + + ++I+EAT F D Y+VG G+ G VYKA + G +AVKKL
Sbjct: 792 PFFQESDIYFVPKE--RFTVKDILEATKGFHDSYIVGRGACGTVYKAVMPSGKTIAVKKL 849
Query: 798 HLVTD--EEMSCFSSKSFMSEIETLTGIKHRNIIKLHGFCSH--SKFSFLVYKFLEGGSL 853
+ S + SF +EI TL I+HRNI++L+ FC H S + L+Y+++ GSL
Sbjct: 850 ESNREGNNNNSNNTDNSFRAEILTLGKIRHRNIVRLYSFCYHQGSNSNLLLYEYMSRGSL 909
Query: 854 DQILNNDTQAVAFDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHV 913
++L+ ++ + DW R + G A L+YLHHDC P IIHRDI S N+L++ ++EAHV
Sbjct: 910 GELLHGG-KSHSMDWPTRFAIALGAAEGLAYLHHDCKPRIIHRDIKSNNILIDENFEAHV 968
Query: 914 SDFGTAKFL-KPGLHSWTQFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGKHP- 971
DFG AK + P S + AG++GY APE A TM+V EKCD+YSFGV+ LE + GK P
Sbjct: 969 GDFGLAKVIDMPLSKSVSAVAGSYGYIAPEYAYTMKVTEKCDIYSFGVVLLELLTGKAPV 1028
Query: 972 ------GDLISLFLSPSTRPMANNMLLTDVLDQRPQQVMEP-IDEEVILIARLAFACLSQ 1024
GDL + + + ++ L +++LD +V + I +I + ++A C
Sbjct: 1029 QPLEQGGDLATW----TRNHIRDHSLTSEILDPYLTKVEDDVILNHMITVTKIAVLCTKS 1084
Query: 1025 NPRLRPSMGQVCKML 1039
+P RP+M +V ML
Sbjct: 1085 SPSDRPTMREVVLML 1099
>ref|NP_189066.1| leucine-rich repeat transmembrane protein kinase, putative
[Arabidopsis thaliana]
Length = 1141
Score = 529 bits (1363), Expect = e-148
Identities = 363/1081 (33%), Positives = 550/1081 (50%), Gaps = 89/1081 (8%)
Query: 8 IMILCVLPTLSVAEDSEAKLALLKWKDSFDDQSQTL-LSTWKNNTN-PCKPKWRGIKCDK 65
I I C +LS AE + L W S +L L W + N PC W I C
Sbjct: 23 IFIFCF--SLSDAEQNPEASILYSWLHSSSPTPSSLSLFNWNSIDNTPCN-NWTFITCSS 79
Query: 66 SNFISTIGLANLGLK----------GTLHSLTFS------SFPNLL-------MIDIRNN 102
FI+ I + ++ L+ +L LT S + P L ++D+ +N
Sbjct: 80 QGFITDIDIESVPLQLSLPKNLPAFRSLQKLTISGANLTGTLPESLGDCLGLKVLDLSSN 139
Query: 103 SFYGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSIGN 162
G IP + L N+ L +N G IP ++ + L+ L + L G+IP +G
Sbjct: 140 GLVGDIPWSLSKLRNLETLILNSNQLTGKIPPDISKCSKLKSLILFDNLLTGSIPTELGK 199
Query: 163 LTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSK 222
L+ L + +GGN G IP EIG +NL L + ++++ G++P +G L L + +
Sbjct: 200 LSGLEVIRIGGNKEISGQIPSEIGDCSNLTVLGLAETSVSGNLPSSLGKLKKLETLSIYT 259
Query: 223 NSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSI 282
+SG IP +GN S+L L L N+ +SG IP + ++ L L+ L G IP+ I
Sbjct: 260 TMISGEIPSDLGNCSELVDLFLYENS-LSGSIPREIGQLTKLEQLFLWQNSLVGGIPEEI 318
Query: 283 QNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQ 342
N NLK + L +N LSGSIPS+IG L L + + N SG IP +I N +L L +
Sbjct: 319 GNCSNLKMIDLSLNLLSGSIPSSIGRLSFLEEFMISDNKFSGSIPTTISNCSSLVQLQLD 378
Query: 343 ENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQI 402
+N ++G IP+ +G L LT+F +N+L G IP GL + T+ + +S N G +PS +
Sbjct: 379 KNQISGLIPSELGTLTKLTLFFAWSNQLEGSIPPGLADCTDLQALDLSRNSLTGTIPSGL 438
Query: 403 CSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLS 462
+L L N +G IP + CSS+ R+ L N+I G+I G K+ +LD S
Sbjct: 439 FMLRNLTKLLLISNSLSGFIPQEIGNCSSLVRLRLGFNRITGEIPSGIGSLKKINFLDFS 498
Query: 463 DNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEV 522
N+ HG++ G LQ +SNN++ G +P L+ L VL +S+NQ +GK+P
Sbjct: 499 SNRLHGKVPDEIGSCSELQMIDLSNNSLEGSLPNPVSSLSGLQVLDVSANQFSGKIPAS- 557
Query: 523 LGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKIPKELVELPNLRM-LN 581
LG + SL L +S N FS +IP+ +G+ LQ LDLG NELSG+IP EL ++ NL + LN
Sbjct: 558 LGRLVSLNKLILSKNLFSGSIPTSLGMCSGLQLLDLGSNELSGEIPSELGDIENLEIALN 617
Query: 582 LSRNKIEGIIPIKFDSGLESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSGTI-PQN 640
LS N+ L G IP+ +A L +LS L+LSHNML G + P
Sbjct: 618 LSSNR----------------------LTGKIPSKIASLNKLSILDLSHNMLEGDLAPLA 655
Query: 641 FGRNLVFVNISDNQLEGPLPKIPAFLSASFESLKNNNHLCGNIRGLDPCATSHSR----- 695
NLV +NIS N G LP F S + L+ N LC + + D C ++ +
Sbjct: 656 NIENLVSLNISYNSFSGYLPDNKLFRQLSPQDLEGNKKLCSSTQ--DSCFLTYRKGNGLG 713
Query: 696 -----KRKNVLRPVFIALGAVILVLCVVGALMYIMCGRKKPNE-ESQTEEVQRGVLFSIW 749
R LR L + +VL ++GA+ I R NE +S+ E + W
Sbjct: 714 DDGDASRTRKLRLTLALLITLTVVLMILGAVAVIRARRNIDNERDSELGETYK------W 767
Query: 750 SHD--GKMMFENIIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVT----DE 803
K+ F ++ + + ++G G G VY+A++ G V+AVKKL +
Sbjct: 768 QFTPFQKLNF-SVDQIIRCLVEPNVIGKGCSGVVYRADVDNGEVIAVKKLWPAMVNGGHD 826
Query: 804 EMSCFSSKSFMSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQA 863
E + SF +E++TL I+H+NI++ G C + L+Y ++ GSL +L ++ +
Sbjct: 827 EKTKNVRDSFSAEVKTLGTIRHKNIVRFLGCCWNRNTRLLMYDYMPNGSLGSLL-HERRG 885
Query: 864 VAFDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLK 923
+ DW+ R ++ G A L+YLHHDC PPI+HRDI + N+L+ LD+E +++DFG AK +
Sbjct: 886 SSLDWDLRYRILLGAAQGLAYLHHDCLPPIVHRDIKANNILIGLDFEPYIADFGLAKLVD 945
Query: 924 PG--LHSWTQFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGKHPGDLI---SLF 978
G AG++GY APE +M++ EK DVYS+GV+ LE + GK P D +
Sbjct: 946 EGDIGRCSNTVAGSYGYIAPEYGYSMKITEKSDVYSYGVVVLEVLTGKQPIDPTVPEGIH 1005
Query: 979 LSPSTRPMANNMLLTDVLDQRPQQVMEPIDEEVILIARLAFACLSQNPRLRPSMGQVCKM 1038
L R ++ +VLD + E +E++ + A C++ +P RP+M V M
Sbjct: 1006 LVDWVRQNRGSL---EVLDSTLRSRTEAEADEMMQVLGTALLCVNSSPDERPTMKDVAAM 1062
Query: 1039 L 1039
L
Sbjct: 1063 L 1063
>emb|CAD79350.1| LRR receptor-like kinase 2 [Arabidopsis thaliana]
Length = 1120
Score = 528 bits (1361), Expect = e-148
Identities = 363/1081 (33%), Positives = 550/1081 (50%), Gaps = 89/1081 (8%)
Query: 8 IMILCVLPTLSVAEDSEAKLALLKWKDSFDDQSQTL-LSTWKNNTN-PCKPKWRGIKCDK 65
I I C +LS AE + L W S +L L W + N PC W I C
Sbjct: 23 IFIFCF--SLSDAEQNPEASILYSWLHSSSPTPSSLSLFNWNSIDNTPCN-NWTFITCSS 79
Query: 66 SNFISTIGLANLGLK----------GTLHSLTFS------SFPNLL-------MIDIRNN 102
FI+ I + ++ L+ +L LT S + P L ++D+ +N
Sbjct: 80 QGFITDIDIESVPLQLSLPKNLPAFRSLQKLTISGANLTGTLPESLGDCLGLKVLDLSSN 139
Query: 103 SFYGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSIGN 162
G IP + L N+ L +N G IP ++ + L+ L + L G+IP +G
Sbjct: 140 GLVGDIPWSLSKLRNLETLILNSNQLTGKIPPDISKCSKLKSLILFDNLLTGSIPTELGK 199
Query: 163 LTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSK 222
L+ L + +GGN G IP EIG +NL L + ++++ G++P +G L L + +
Sbjct: 200 LSGLEVIRIGGNKEISGQIPLEIGDCSNLTVLGLAETSVSGNLPSSLGKLKKLETLSIYT 259
Query: 223 NSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSI 282
+SG IP +GN S+L L L N+ +SG IP + ++ L L+ L G IP+ I
Sbjct: 260 TMISGEIPSDLGNCSELVDLFLYENS-LSGSIPREIGQLTKLEQLFLWQNSLVGGIPEEI 318
Query: 283 QNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQ 342
N NLK + L +N LSGSIPS+IG L L + + N SG IP +I N +L L +
Sbjct: 319 GNCSNLKMIDLSLNLLSGSIPSSIGRLSFLEEFMISDNKFSGSIPTTISNCSSLVQLQLD 378
Query: 343 ENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQI 402
+N ++G IP+ +G L LT+F +N+L G IP GL + T+ + +S N G +PS +
Sbjct: 379 KNQISGLIPSELGTLTKLTLFFAWSNQLEGSIPPGLADCTDLQALDLSRNSLTGTIPSGL 438
Query: 403 CSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLS 462
+L L N +G IP + CSS+ R+ L N+I G+I G K+ +LD S
Sbjct: 439 FMLRNLTKLLLISNSLSGFIPQEIGNCSSLVRLRLGFNRITGEIPSGIGSLKKINFLDFS 498
Query: 463 DNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEV 522
N+ HG++ G LQ +SNN++ G +P L+ L VL +S+NQ +GK+P
Sbjct: 499 SNRLHGKVPDEIGSCSELQMIDLSNNSLEGSLPNPVSSLSGLQVLDVSANQFSGKIPAS- 557
Query: 523 LGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKIPKELVELPNLRM-LN 581
LG + SL L +S N FS +IP+ +G+ LQ LDLG NELSG+IP EL ++ NL + LN
Sbjct: 558 LGRLVSLNKLILSKNLFSGSIPTSLGMCSGLQLLDLGSNELSGEIPSELGDIENLEIALN 617
Query: 582 LSRNKIEGIIPIKFDSGLESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSGTI-PQN 640
LS N+ L G IP+ +A L +LS L+LSHNML G + P
Sbjct: 618 LSSNR----------------------LTGKIPSKIASLNKLSILDLSHNMLEGDLAPLA 655
Query: 641 FGRNLVFVNISDNQLEGPLPKIPAFLSASFESLKNNNHLCGNIRGLDPCATSHSR----- 695
NLV +NIS N G LP F S + L+ N LC + + D C ++ +
Sbjct: 656 NIENLVSLNISYNSFSGYLPDNKLFRQLSPQDLEGNKKLCSSTQ--DSCFLTYRKGNGLG 713
Query: 696 -----KRKNVLRPVFIALGAVILVLCVVGALMYIMCGRKKPNE-ESQTEEVQRGVLFSIW 749
R LR L + +VL ++GA+ I R NE +S+ E + W
Sbjct: 714 DDGDASRTRKLRLTLALLITLTVVLMILGAVAVIRARRNIDNERDSELGETYK------W 767
Query: 750 SHD--GKMMFENIIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVT----DE 803
K+ F ++ + + ++G G G VY+A++ G V+AVKKL +
Sbjct: 768 QFTPFQKLNF-SVDQIIRCLVEPNVIGKGCSGVVYRADVDNGEVIAVKKLWPAMVNGGHD 826
Query: 804 EMSCFSSKSFMSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQA 863
E + SF +E++TL I+H+NI++ G C + L+Y ++ GSL +L ++ +
Sbjct: 827 EKTKNVRDSFSAEVKTLGTIRHKNIVRFLGCCWNRNTRLLMYDYMPNGSLGSLL-HERRG 885
Query: 864 VAFDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLK 923
+ DW+ R ++ G A L+YLHHDC PPI+HRDI + N+L+ LD+E +++DFG AK +
Sbjct: 886 SSLDWDLRYRILLGAAQGLAYLHHDCLPPIVHRDIKANNILIGLDFEPYIADFGLAKLVD 945
Query: 924 PG--LHSWTQFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGKHPGDLI---SLF 978
G AG++GY APE +M++ EK DVYS+GV+ LE + GK P D +
Sbjct: 946 EGDIGRCSNTVAGSYGYIAPEYGYSMKITEKSDVYSYGVVVLEVLTGKQPIDPTVPEGIH 1005
Query: 979 LSPSTRPMANNMLLTDVLDQRPQQVMEPIDEEVILIARLAFACLSQNPRLRPSMGQVCKM 1038
L R ++ +VLD + E +E++ + A C++ +P RP+M V M
Sbjct: 1006 LVDWVRQNRGSL---EVLDSTLRSRTEAEADEMMQVLGTALLCVNSSPDERPTMKDVAAM 1062
Query: 1039 L 1039
L
Sbjct: 1063 L 1063
>dbj|BAD32908.1| putative receptor-like protein kinase 2 [Oryza sativa (japonica
cultivar-group)]
Length = 1072
Score = 526 bits (1354), Expect = e-147
Identities = 359/1081 (33%), Positives = 550/1081 (50%), Gaps = 86/1081 (7%)
Query: 1 MMVLPTLIMILCVLPTLSVAEDSEAKLALLKWKDSFDDQSQTLLSTWKNNT-NPCKPKWR 59
++V+ +++ + V P +++ D +A L+LL +L +W PC W+
Sbjct: 10 VVVVVVVVLGVVVRPAAALSADGKALLSLLPAA-----APSPVLPSWDPTAATPCS--WQ 62
Query: 60 GIKCDKSNFISTIGLANLGLK--------GTLHSL----------------TFSSFPNLL 95
G+ C + + ++ L N L +L SL ++S L
Sbjct: 63 GVTCSPQSRVVSLSLPNTFLNLSSLPPQLASLSSLQLLNLSTCNISGAIPPAYASLAALR 122
Query: 96 MIDIRNNSFYGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGA 155
++D+ +N+ YG IPA +G LS + L +N G+IP+ + +L LQ L + LNG
Sbjct: 123 VLDLSSNALYGDIPASLGALSGLQYLLLNSNRLTGAIPRSLASLAALQVLCVQDNLLNGT 182
Query: 156 IPKSIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNL 215
IP S+G LT L +GGN GPIP +G L+NL + L G+IP+E+G L NL
Sbjct: 183 IPASLGALTALQQFRVGGNPGLSGPIPASLGALSNLTVFGAAATALSGAIPEELGNLANL 242
Query: 216 AYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLS 275
+ L +SG IP +G ++L L L N K++GPIP L + LT L LS
Sbjct: 243 QTLALYDTGVSGPIPAALGGCAELRNLYLHMN-KLTGPIPPELGRLQKLTSLLLWGNALS 301
Query: 276 GSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLIN 335
G IP + N L L L N L+G +P +G L L +L+L N L+G IPA + N +
Sbjct: 302 GRIPPELSNCSALVVLDLSGNRLAGEVPGALGRLAALEQLHLSDNQLAGRIPAELSNCSS 361
Query: 336 LQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFV 395
L L + +N LTG IP +G L+ L V + N L G IP L N T + +S N
Sbjct: 362 LTALQLDKNGLTGAIPPQLGELRALQVLFLWGNALSGAIPPSLGNCTELYALDLSRNRLA 421
Query: 396 GHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPK 455
G +P ++ + L L N +G +P S+ CSS+ R+ L NQ+ G+I ++ G P
Sbjct: 422 GGIPDEVFALQKLSKLLLLGNALSGRLPPSVADCSSLVRLRLGENQLAGEIPREIGKLPN 481
Query: 456 LQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLT 515
L +LDL NKF G + L+ + NN+ +G IP F L L L LS N+LT
Sbjct: 482 LVFLDLYSNKFTGALPGELANITVLELLDVHNNSFTGAIPPQFGELMNLEQLDLSMNKLT 541
Query: 516 GKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKIPKELVELP 575
G++P G L L +S N S +P I LQ+L L+L N SG IP E+ L
Sbjct: 542 GEIPAS-FGNFSYLNKLILSGNMLSGTLPKSIRNLQKLTMLELSNNSFSGPIPPEIGALS 600
Query: 576 NLRMLNLSRNKIEGIIPIKFDSGLESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSG 635
+L + SLDLS N G +P ++ L +L L+LS N L G
Sbjct: 601 SLSI---------------------SLDLSSNRFTGELPDEMSSLTQLQSLDLSSNGLYG 639
Query: 636 TIPQNFG-RNLVFVNISDNQLEGPLPKIPAFLSASFESLKNNNHLCGNIRGLDPCATSHS 694
+I G +L +NIS N G +P P F + S S NN +LC + G CA+
Sbjct: 640 SISVLSGLTSLTSLNISYNNFSGAIPVTPFFKTLSSSSYINNPNLCESYDG-HTCASDMV 698
Query: 695 RKR--KNVLRPVFI--ALGAVILVLCVVGALMYIMCGRKKPNEESQTEEVQRGVLFS--- 747
R+ K V + + LG++ L+L VV L I R +++ + V G FS
Sbjct: 699 RRTALKTVKTVILVCAVLGSITLLLVVVWIL--INRSRTLAGKKAMSMSVAGGDDFSHPW 756
Query: 748 IWSHDGKMMF--ENIIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEM 805
++ K+ F +NI+E D+ ++G G G VY+AE+ G ++AVKKL + EE
Sbjct: 757 TFTPFQKLNFCVDNILEC---LRDENVIGKGCSGVVYRAEMPNGEIIAVKKLWKTSKEE- 812
Query: 806 SCFSSKSFMSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQAVA 865
+F +EI+ L I+HRNI+KL G+CS+ L+Y ++ G+L Q+L ++ +
Sbjct: 813 ---PIDAFAAEIQILGHIRHRNIVKLLGYCSNKYVKLLLYNYIPNGNLQQLLKDNR---S 866
Query: 866 FDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFL-KP 924
DW+ R + G A L+YLHHDC P I+HRD+ N+LL+ YEA+++DFG AK + P
Sbjct: 867 LDWDTRYKIAVGAAQGLAYLHHDCVPAILHRDVKCNNILLDTKYEAYLADFGLAKLMNSP 926
Query: 925 GL-HSWTQFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGKHP-----GDLISLF 978
H+ ++ AG++GY APE T ++ EK DVYS+GV+ LE + G+ GD + +
Sbjct: 927 NYHHAMSRIAGSYGYIAPEYGYTTKITEKSDVYSYGVVLLEILSGRSAVEAVVGDSLHI- 985
Query: 979 LSPSTRPMANNMLLTDVLDQRPQQVMEPIDEEVILIARLAFACLSQNPRLRPSMGQVCKM 1038
+ + + M + ++LD + + + + + +E++ +A C++ P RP+M +V
Sbjct: 986 VEWAKKKMGSYEPAVNILDPKLRGMPDQLVQEMLQTLGIAIFCVNPAPAERPTMKEVVAF 1045
Query: 1039 L 1039
L
Sbjct: 1046 L 1046
>ref|NP_177451.1| leucine-rich repeat transmembrane protein kinase, putative
[Arabidopsis thaliana] gi|5903097|gb|AAD55655.1| Highly
similar to receptor-like protein kinase [Arabidopsis
thaliana] gi|25406249|pir||D96756 receptor-like protein
kinase homolog [imported] - Arabidopsis thaliana
Length = 1123
Score = 525 bits (1352), Expect = e-147
Identities = 366/1103 (33%), Positives = 542/1103 (48%), Gaps = 112/1103 (10%)
Query: 22 DSEAKLALLKWKDSFDDQSQTLLSTWKNNTNPCKP-KWRGIKCDKSNFISTIGLANLGLK 80
D L+LLK D Q + STWK N + P W GI CD S ++++ +
Sbjct: 32 DGLTLLSLLKHLDRVPPQ---VTSTWKINASEATPCNWFGITCDDSKNVASLNFTRSRVS 88
Query: 81 GTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISILTFKNNYFD----------- 129
G L +L ++D+ N+F GTIP+ +GN + ++ L N F
Sbjct: 89 GQLGP-EIGELKSLQILDLSTNNFSGTIPSTLGNCTKLATLDLSENGFSDKIPDTLDSLK 147
Query: 130 -------------GSIPQEMCTLTGLQFLDISFCKLNGAIPKSIGNLTNLSYLILGGNNW 176
G +P+ + + LQ L + + L G IP+SIG+ L L + N +
Sbjct: 148 RLEVLYLYINFLTGELPESLFRIPKLQVLYLDYNNLTGPIPQSIGDAKELVELSMYANQF 207
Query: 177 SGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYI------------------ 218
SG IP IG ++L L + ++ LVGS+P+ + L NL +
Sbjct: 208 SGN-IPESIGNSSSLQILYLHRNKLVGSLPESLNLLGNLTTLFVGNNSLQGPVRFGSPNC 266
Query: 219 ------DLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNI 272
DLS N GG+P +GN S LD LV+ + +SG IP SL + +LT+L
Sbjct: 267 KNLLTLDLSYNEFEGGVPPALGNCSSLDALVIVSGN-LSGTIPSSLGMLKNLTILNLSEN 325
Query: 273 GLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGN 332
LSGSIP + N +L L L+ N L G IPS +G L+ L L L N SG IP I
Sbjct: 326 RLSGSIPAELGNCSSLNLLKLNDNQLVGGIPSALGKLRKLESLELFENRFSGEIPIEIWK 385
Query: 333 LINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSEN 392
+L L V +NNLTG +P + +K L + + N +G IP GL ++ + E
Sbjct: 386 SQSLTQLLVYQNNLTGELPVEMTEMKKLKIATLFNNSFYGAIPPGL-----GVNSSLEEV 440
Query: 393 DFVGH-----LPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIA 447
DF+G+ +P +C G LR+LN N G IP S+ C +I R L N + G +
Sbjct: 441 DFIGNKLTGEIPPNLCHGRKLRILNLGSNLLHGTIPASIGHCKTIRRFILRENNLSG-LL 499
Query: 448 QDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVL 507
+F L +LD + N F G I + G NL + +S N +G IP L LG +
Sbjct: 500 PEFSQDHSLSFLDFNSNNFEGPIPGSLGSCKNLSSINLSRNRFTGQIPPQLGNLQNLGYM 559
Query: 508 HLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKI 567
+LS N L G LP + L SL + N + ++PS + L L L N SG I
Sbjct: 560 NLSRNLLEGSLPAQ-LSNCVSLERFDVGFNSLNGSVPSNFSNWKGLTTLVLSENRFSGGI 618
Query: 568 PKELVELPNLRMLNLSRNKIEGIIPIKF---DSGLESLDLSGNFLKGNIPTGLADLVRLS 624
P+ L EL L L ++RN G IP + + LDLSGN L G IP L DL++L+
Sbjct: 619 PQFLPELKKLSTLQIARNAFGGEIPSSIGLIEDLIYDLDLSGNGLTGEIPAKLGDLIKLT 678
Query: 625 KLNLSHNMLSGTIPQNFG-RNLVFVNISDNQLEGPLP-KIPAFLSASFESLKNNNHLC-- 680
+LN+S+N L+G++ G +L+ V++S+NQ GP+P + L + S N +LC
Sbjct: 679 RLNISNNNLTGSLSVLKGLTSLLHVDVSNNQFTGPIPDNLEGQLLSEPSSFSGNPNLCIP 738
Query: 681 ------GNIRGLDPCATSHSRKRKNVLRP---VFIALGAVILVLCVVGALMYIMCGRKKP 731
N R S+ RK+ L V IA+ + +LVL VV AL++I R+K
Sbjct: 739 HSFSASNNSRSALKYCKDQSKSRKSGLSTWQIVLIAVLSSLLVLVVVLALVFICLRRRKG 798
Query: 732 NEESQTEEVQRGVLFSIWSHDG-KMMFENIIEATANFDDKYLVGVGSQGNVYKAELSEGL 790
E + +G ++ ++ AT N ++KY +G G+ G VY+A L G
Sbjct: 799 RPEKDA--------YVFTQEEGPSLLLNKVLAATDNLNEKYTIGRGAHGIVYRASLGSGK 850
Query: 791 VVAVKKLHLVTDEEMSCFSSKSFMSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEG 850
V AVK+L + +++S M EI+T+ ++HRN+IKL GF ++Y+++
Sbjct: 851 VYAVKRLVFASHIR----ANQSMMREIDTIGKVRHRNLIKLEGFWLRKDDGLMLYRYMPK 906
Query: 851 GSLDQILNN-DTQAVAFDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDY 909
GSL +L+ + DW R NV GVA+ L+YLH+DC PPI+HRDI +N+L++ D
Sbjct: 907 GSLYDVLHGVSPKENVLDWSARYNVALGVAHGLAYLHYDCHPPIVHRDIKPENILMDSDL 966
Query: 910 EAHVSDFGTAKFLKPGLHSWTQFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGK 969
E H+ DFG A+ L S GT GY APE A + DVYS+GV+ LE + K
Sbjct: 967 EPHIGDFGLARLLDDSTVSTATVTGTTGYIAPENAFKTVRGRESDVYSYGVVLLELVTRK 1026
Query: 970 HPGDL-------ISLFLSPSTRPMANNM--LLTDVLDQRPQQVMEPID----EEVILIAR 1016
D I ++ + NN+ ++T ++D P V E +D E+V+ +
Sbjct: 1027 RAVDKSFPESTDIVSWVRSALSSSNNNVEDMVTTIVD--PILVDELLDSSLREQVMQVTE 1084
Query: 1017 LAFACLSQNPRLRPSMGQVCKML 1039
LA +C Q+P +RP+M K+L
Sbjct: 1085 LALSCTQQDPAMRPTMRDAVKLL 1107
>ref|NP_173217.1| leucine-rich repeat transmembrane protein kinase, putative
[Arabidopsis thaliana] gi|25513592|pir||E86312 F11A6.9
protein - Arabidopsis thaliana gi|9802748|gb|AAF99817.1|
Unknown protein [Arabidopsis thaliana]
Length = 1088
Score = 524 bits (1349), Expect = e-147
Identities = 362/1082 (33%), Positives = 543/1082 (49%), Gaps = 91/1082 (8%)
Query: 18 SVAEDSEAKLALLKWKDSFDDQSQTLLSTWKNNTN---PCKPKWRGIKCDKS-NFISTIG 73
SV+ + LALL FD + STWK NT+ PC W G+ CD S N + T+
Sbjct: 23 SVSSLNSDGLALLSLLKHFDKVPLEVASTWKENTSETTPCNNNWFGVICDLSGNVVETLN 82
Query: 74 LANLGLKGTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISILTFKNNYFDGSIP 133
L+ GL G L S +L+ +D+ NSF G +P+ +GN +++ L NN F G +P
Sbjct: 83 LSASGLSGQLGS-EIGELKSLVTLDLSLNSFSGLLPSTLGNCTSLEYLDLSNNDFSGEVP 141
Query: 134 QEMCTLTGLQFLDISFCKLNGAIPKSIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLH 193
+L L FL + L+G IP S+G L L L + NN SG IP +G + L +
Sbjct: 142 DIFGSLQNLTFLYLDRNNLSGLIPASVGGLIELVDLRMSYNNLSG-TIPELLGNCSKLEY 200
Query: 194 LAIQKSNLVGSIPQEIGFLTNLAYI------------------------DLSKNSLSGGI 229
LA+ + L GS+P + L NL + DLS N GG+
Sbjct: 201 LALNNNKLNGSLPASLYLLENLGELFVSNNSLGGRLHFGSSNCKKLVSLDLSFNDFQGGV 260
Query: 230 PETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLK 289
P IGN S L +LV+ ++G IP S+ + ++V+ + LSG+IP + N +L+
Sbjct: 261 PPEIGNCSSLHSLVMVK-CNLTGTIPSSMGMLRKVSVIDLSDNRLSGNIPQELGNCSSLE 319
Query: 290 ELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGT 349
L L+ N L G IP + LK L L L N LSG IP I + +L + V N LTG
Sbjct: 320 TLKLNDNQLQGEIPPALSKLKKLQSLELFFNKLSGEIPIGIWKIQSLTQMLVYNNTLTGE 379
Query: 350 IPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLR 409
+P + LK L + N +G IP L + + N F G +P +C G LR
Sbjct: 380 LPVEVTQLKHLKKLTLFNNGFYGDIPMSLGLNRSLEEVDLLGNRFTGEIPPHLCHGQKLR 439
Query: 410 LLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQ 469
L N+ G IP S++ C ++ER+ LE N++ G + +F L Y++L N F G
Sbjct: 440 LFILGSNQLHGKIPASIRQCKTLERVRLEDNKLSG-VLPEFPESLSLSYVNLGSNSFEGS 498
Query: 470 ISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSL 529
I + G NL T +S N ++G+IP + L LG+L+LS N L G LP ++ G + L
Sbjct: 499 IPRSLGSCKNLLTIDLSQNKLTGLIPPELGNLQSLGLLNLSHNYLEGPLPSQLSGCARLL 558
Query: 530 FDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKIPKELVELPNLRMLNLSRNKIEG 589
+ + +N + +IPS + L L L N G IP+ L EL L L ++RN G
Sbjct: 559 Y-FDVGSNSLNGSIPSSFRSWKSLSTLVLSDNNFLGAIPQFLAELDRLSDLRIARNAFGG 617
Query: 590 IIPIK---FDSGLESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSG--TIPQNFGRN 644
IP S LDLS N G IPT L L+ L +LN+S+N L+G ++ Q+ ++
Sbjct: 618 KIPSSVGLLKSLRYGLDLSANVFTGEIPTTLGALINLERLNISNNKLTGPLSVLQSL-KS 676
Query: 645 LVFVNISDNQLEGPLPKIPAFLSASFESLKNNNHLCGNIRGLDPCATSHSRKRKNVLRPV 704
L V++S NQ GP IP L ++ N LC I+ + ++ K+ V
Sbjct: 677 LNQVDVSYNQFTGP---IPVNLLSNSSKFSGNPDLC--IQASYSVSAIIRKEFKSCKGQV 731
Query: 705 --------FIALGAVILVLCVVGALMYIMCGRKKPNEESQTEEVQRGVLFSIWSHDG-KM 755
IA G+ + VL ++ AL ++C K+ ++TE+ +I + +G +
Sbjct: 732 KLSTWKIALIAAGSSLSVLALLFALFLVLCRCKRG---TKTEDA------NILAEEGLSL 782
Query: 756 MFENIIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMS 815
+ ++ AT N DDKY++G G+ G VY+A L G AVKKL + E + ++++
Sbjct: 783 LLNKVLAATDNLDDKYIIGRGAHGVVYRASLGSGEEYAVKKL--IFAEHIR--ANQNMKR 838
Query: 816 EIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQAVA-FDWEKRVNV 874
EIET+ ++HRN+I+L F + ++Y+++ GSL +L+ Q A DW R N+
Sbjct: 839 EIETIGLVRHRNLIRLERFWMRKEDGLMLYQYMPNGSLHDVLHRGNQGEAVLDWSARFNI 898
Query: 875 VKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLKPGLHSWTQFAG 934
G+++ L+YLHHDC PPIIHRDI +N+L++ D E H+ DFG A+ L S G
Sbjct: 899 ALGISHGLAYLHHDCHPPIIHRDIKPENILMDSDMEPHIGDFGLARILDDSTVSTATVTG 958
Query: 935 TFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGKHPGD--------LISLFLS------ 980
T GY APE A +++ DVYS+GV+ LE + GK D ++S S
Sbjct: 959 TTGYIAPENAYKTVRSKESDVYSYGVVLLELVTGKRALDRSFPEDINIVSWVRSVLSSYE 1018
Query: 981 ---PSTRPMANNMLLTDVLDQRPQQVMEPIDEEVILIARLAFACLSQNPRLRPSMGQVCK 1037
+ P+ + L+ ++LD + + E+ I + LA C + P RPSM V K
Sbjct: 1019 DEDDTAGPIVDPKLVDELLDTK-------LREQAIQVTDLALRCTDKRPENRPSMRDVVK 1071
Query: 1038 ML 1039
L
Sbjct: 1072 DL 1073
>dbj|BAC41855.1| unknown protein [Arabidopsis thaliana]
Length = 1123
Score = 523 bits (1347), Expect = e-146
Identities = 365/1103 (33%), Positives = 542/1103 (49%), Gaps = 112/1103 (10%)
Query: 22 DSEAKLALLKWKDSFDDQSQTLLSTWKNNTNPCKP-KWRGIKCDKSNFISTIGLANLGLK 80
D L+LLK D Q + STWK N + P W GI CD S ++++ +
Sbjct: 32 DGLTLLSLLKHLDRVPPQ---VTSTWKINASEATPCNWFGITCDDSKNVASLNFTRSRVS 88
Query: 81 GTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISILTFKNNYFD----------- 129
G L +L ++D+ N+F GTIP+ +GN + ++ L N F
Sbjct: 89 GQLGP-EIGELKSLQILDLSTNNFSGTIPSTLGNCTKLATLDLSENGFSDKIPDTLDSLK 147
Query: 130 -------------GSIPQEMCTLTGLQFLDISFCKLNGAIPKSIGNLTNLSYLILGGNNW 176
G +P+ + + LQ L + + L G IP+SIG+ L L + N +
Sbjct: 148 RLEVLYLYINFLTGELPESLFRIPKLQVLYLDYNNLTGPIPQSIGDAKELVELSMYANQF 207
Query: 177 SGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYI------------------ 218
SG IP IG ++L L + ++ LVGS+P+ + L NL +
Sbjct: 208 SGN-IPESIGNSSSLQILYLHRNKLVGSLPESLNLLGNLTTLFVGNNSLQGPVRFGSPNC 266
Query: 219 ------DLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNI 272
DLS N GG+P +GN S LD LV+ + +SG IP SL + +LT+L
Sbjct: 267 KNLLTLDLSYNEFEGGVPPALGNCSSLDALVIVSGN-LSGTIPSSLGMLKNLTILNLSEN 325
Query: 273 GLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGN 332
LSGSIP + N +L L L+ N L G IPS +G L+ L L L N SG IP I
Sbjct: 326 RLSGSIPAELGNCSSLNLLKLNDNQLVGGIPSALGKLRKLESLELFENRFSGEIPIEIWK 385
Query: 333 LINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSEN 392
+L L V +NNLTG +P + +K L + + N +G IP GL ++ + E
Sbjct: 386 SQSLTQLLVYQNNLTGELPVEMTEMKKLKIATLFNNSFYGAIPPGL-----GVNSSLEEV 440
Query: 393 DFVGH-----LPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIA 447
DF+G+ +P +C G LR+LN N G IP S+ C +I R L N + G +
Sbjct: 441 DFIGNKLTGEIPPNLCHGRKLRILNLGSNLLHGTIPASIGHCKTIRRFILRENNLSG-LL 499
Query: 448 QDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVL 507
+F L +LD + N F G I + G NL + +S N +G IP L LG +
Sbjct: 500 PEFSQDHSLSFLDFNSNNFEGPIPGSLGSCKNLSSINLSRNRFTGQIPPQLGNLQNLGYM 559
Query: 508 HLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKI 567
+LS N L G LP + L SL + N + ++PS + L L L N SG I
Sbjct: 560 NLSRNLLEGSLPAQ-LSNCVSLERFDVGFNSLNGSVPSNFSNWKGLTTLVLSENRFSGGI 618
Query: 568 PKELVELPNLRMLNLSRNKIEGIIPIKF---DSGLESLDLSGNFLKGNIPTGLADLVRLS 624
P+ L EL L L ++RN G IP + + LDLSGN L G IP L DL++L+
Sbjct: 619 PQFLPELKKLSTLQIARNAFGGEIPSSIGLIEDLIYDLDLSGNGLTGEIPAKLGDLIKLT 678
Query: 625 KLNLSHNMLSGTIPQNFG-RNLVFVNISDNQLEGPLP-KIPAFLSASFESLKNNNHLC-- 680
+LN+S+N L+G++ G +L+ V++S+NQ GP+P + L + S N +LC
Sbjct: 679 RLNISNNNLTGSLSVLKGLTSLLHVDVSNNQFTGPIPDNLEGQLLSEPSSFSGNPNLCIP 738
Query: 681 ------GNIRGLDPCATSHSRKRKNVLRP---VFIALGAVILVLCVVGALMYIMCGRKKP 731
+ R S+ RK+ L V IA+ + +LVL VV AL++I R+K
Sbjct: 739 HSFSASNDSRSALKYCKDQSKSRKSGLSTWQIVLIAVLSSLLVLVVVLALVFICLRRRKG 798
Query: 732 NEESQTEEVQRGVLFSIWSHDG-KMMFENIIEATANFDDKYLVGVGSQGNVYKAELSEGL 790
E + +G ++ ++ AT N ++KY +G G+ G VY+A L G
Sbjct: 799 RPEKDA--------YVFTQEEGPSLLLNKVLAATDNLNEKYTIGRGAHGIVYRASLGSGK 850
Query: 791 VVAVKKLHLVTDEEMSCFSSKSFMSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEG 850
V AVK+L + +++S M EI+T+ ++HRN+IKL GF ++Y+++
Sbjct: 851 VYAVKRLVFASHIR----ANQSMMREIDTIGKVRHRNLIKLEGFWLRKDDGLMLYRYMPK 906
Query: 851 GSLDQILNN-DTQAVAFDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDY 909
GSL +L+ + DW R NV GVA+ L+YLH+DC PPI+HRDI +N+L++ D
Sbjct: 907 GSLYDVLHGVSPKENVLDWSARYNVALGVAHGLAYLHYDCHPPIVHRDIKPENILMDSDL 966
Query: 910 EAHVSDFGTAKFLKPGLHSWTQFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGK 969
E H+ DFG A+ L S GT GY APE A + DVYS+GV+ LE + K
Sbjct: 967 EPHIGDFGLARLLDDSTVSTATVTGTTGYIAPENAFKTVRGRESDVYSYGVVLLELVTRK 1026
Query: 970 HPGDL-------ISLFLSPSTRPMANNM--LLTDVLDQRPQQVMEPID----EEVILIAR 1016
D I ++ + NN+ ++T ++D P V E +D E+V+ +
Sbjct: 1027 RAVDKSFPESTDIVSWVRSALSSSNNNVEDMVTTIVD--PILVDELLDSSLREQVMQVTE 1084
Query: 1017 LAFACLSQNPRLRPSMGQVCKML 1039
LA +C Q+P +RP+M K+L
Sbjct: 1085 LALSCTQQDPAMRPTMRDAVKLL 1107
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.319 0.138 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,847,078,011
Number of Sequences: 2540612
Number of extensions: 81255181
Number of successful extensions: 389265
Number of sequences better than 10.0: 23035
Number of HSP's better than 10.0 without gapping: 7091
Number of HSP's successfully gapped in prelim test: 16014
Number of HSP's that attempted gapping in prelim test: 191452
Number of HSP's gapped (non-prelim): 60307
length of query: 1067
length of database: 863,360,394
effective HSP length: 139
effective length of query: 928
effective length of database: 510,215,326
effective search space: 473479822528
effective search space used: 473479822528
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 81 (35.8 bits)
Medicago: description of AC133779.11


