

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130802.9 + phase: 0 /pseudo
(937 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
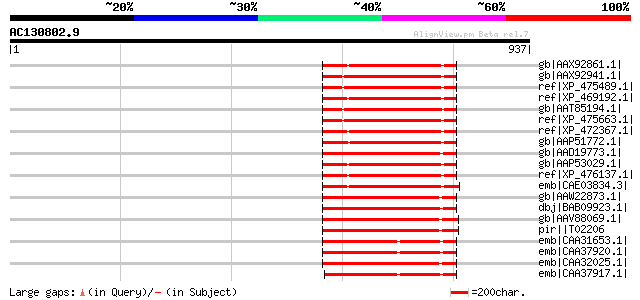
Score E
Sequences producing significant alignments: (bits) Value
gb|AAX92861.1| retrotransposon protein, putative, Ty1-copia sub-... 273 1e-71
gb|AAX92941.1| retrotransposon protein, putative, Ty1-copia sub-... 271 6e-71
ref|XP_475489.1| putative polyprotein [Oryza sativa (japonica cu... 270 2e-70
ref|XP_469192.1| putative polyprotein [Oryza sativa (japonica cu... 268 6e-70
gb|AAT85194.1| putative polyprotein [Oryza sativa (japonica cult... 267 1e-69
ref|XP_475663.1| putative polyprotein [Oryza sativa (japonica cu... 267 1e-69
ref|XP_472367.1| OSJNBb0012E08.6 [Oryza sativa (japonica cultiva... 266 3e-69
gb|AAP51772.1| putative pol polyprotein [Oryza sativa (japonica ... 261 8e-68
gb|AAD19773.1| putative retroelement pol polyprotein [Arabidopsi... 258 8e-67
gb|AAP53029.1| putative retrotransposon-related protein [Oryza s... 257 1e-66
ref|XP_476137.1| putative polyprotein [Oryza sativa (japonica cu... 257 1e-66
emb|CAE03834.3| OSJNBb0013J13.11 [Oryza sativa (japonica cultiva... 256 2e-66
gb|AAW22873.1| putative polyprotein [Lycopersicon esculentum] 249 2e-64
dbj|BAB09923.1| copia-like retrotransposable element [Arabidopsi... 247 1e-63
gb|AAV88069.1| hypothetical retrotransposon [Ipomoea batatas] 245 4e-63
pir||T02206 hypothetical protein - common tobacco retrotransposo... 245 6e-63
emb|CAA31653.1| polyprotein [Arabidopsis thaliana] gi|99721|pir|... 244 1e-62
emb|CAA37920.1| unnamed protein product [Arabidopsis thaliana] g... 243 3e-62
emb|CAA32025.1| unnamed protein product [Nicotiana tabacum] gi|1... 242 4e-62
emb|CAA37917.1| reverse transcriptase [Arabidopsis thaliana] gi|... 242 5e-62
>gb|AAX92861.1| retrotransposon protein, putative, Ty1-copia sub-class [Oryza sativa
(japonica cultivar-group)]
Length = 1373
Score = 273 bits (699), Expect = 1e-71
Identities = 136/243 (55%), Positives = 177/243 (71%), Gaps = 4/243 (1%)
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
+ SLYGLKQSPR+W +RFD F+L F RS YDSCVY+ N I YLLLYVDD+L+A+
Sbjct: 964 KRSLYGLKQSPRQWNKRFDSFMLSHSFKRSKYDSCVYIKHVNGSPI-YLLLYVDDMLIAA 1022
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMS 684
SK EI KLK+ L+ EF+MKDLG AK++LG++I R+R G LFLSQ Y+KK +RF M
Sbjct: 1023 KSKIEITKLKKLLSSEFDMKDLGSAKKILGMEISRDRKSGLLFLSQHNYIKKVLQRFNMQ 1082
Query: 685 NSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIV 744
N+K VSTP+ H KLS QCP + E + M PY+S VGS+MY MVCSRPDL+YA+S+V
Sbjct: 1083 NAKAVSTPIAPHFKLSAAQCPSIDAEIEYMSRVPYSSAVGSLMYAMVCSRPDLSYAMSLV 1142
Query: 745 SRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSL 804
SR+M+NPG HW+A++W+ RYL G+ LK+ R + L GYVD+DYA ++D R+SL
Sbjct: 1143 SRYMSNPGKEHWRAVQWIFRYLRGTTYSCLKFGRT---DKGLIGYVDSDYAADLDRRRSL 1199
Query: 805 VGF 807
G+
Sbjct: 1200 TGY 1202
>gb|AAX92941.1| retrotransposon protein, putative, Ty1-copia sub-class [Oryza sativa
(japonica cultivar-group)]
Length = 2340
Score = 271 bits (694), Expect = 6e-71
Identities = 133/243 (54%), Positives = 180/243 (73%), Gaps = 4/243 (1%)
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
++SLYGLKQSPR+WY+RFD F+L F RS YDSCVY LK + +YLLLYVDD+L+A+
Sbjct: 1170 KKSLYGLKQSPRQWYKRFDSFMLSQKFRRSNYDSCVY-LKVVDGSAIYLLLYVDDMLIAA 1228
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMS 684
K EI KLK +L+ EFEMKDLG AK++LG++I R R +L+LSQ GY++K RF M
Sbjct: 1229 KDKSEIAKLKAQLSSEFEMKDLGAAKKILGMEITRKRHSFKLYLSQKGYIEKVLRRFNMH 1288
Query: 685 NSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIV 744
++K VSTPL H +LS CPQS+ + + M PY+S VGS+MY MVCSRPDL++A+S+V
Sbjct: 1289 DAKPVSTPLAAHFRLSSDLCPQSDYDIEYMSRVPYSSAVGSLMYAMVCSRPDLSHALSVV 1348
Query: 745 SRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSL 804
SR+MANPG HW+A++W+ RYL G+ L++ R+ D L GYVD+D+AG++D +SL
Sbjct: 1349 SRYMANPGKEHWKAVQWIFRYLRGTSSACLQFGRS---RDGLVGYVDSDFAGDLDRGRSL 1405
Query: 805 VGF 807
G+
Sbjct: 1406 AGY 1408
>ref|XP_475489.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|48475213|gb|AAT44282.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1243
Score = 270 bits (689), Expect = 2e-70
Identities = 130/243 (53%), Positives = 180/243 (73%), Gaps = 4/243 (1%)
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
++SLYGLKQSPR+WY+RFD F+L F S YDSCVY LK + ++YLLLYVDD+L+A+
Sbjct: 935 KKSLYGLKQSPRQWYKRFDSFMLSQKFRISNYDSCVY-LKVVDGSVIYLLLYVDDMLIAA 993
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMS 684
K EI KLK +L+ EFEMKDLG AK++LG++I R R G+L+LSQ GY++K RF M
Sbjct: 994 KDKSEIEKLKAQLSSEFEMKDLGAAKKILGMEITRERHSGKLYLSQKGYIEKVLRRFNMH 1053
Query: 685 NSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIV 744
++K VSTPL H +LS CP S+ + + M PY+S VGS+MY MVC RPDL++A+S+V
Sbjct: 1054 DAKPVSTPLAAHFRLSSDLCPLSDYDIEYMSRVPYSSAVGSLMYAMVCCRPDLSHALSVV 1113
Query: 745 SRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSL 804
+R+MANPG HW+A++W+ RYL G+ L++ R+ D L GYVD+D+AG++D R+S+
Sbjct: 1114 NRYMANPGKEHWKAVQWIFRYLRGTSSACLQFERS---RDGLVGYVDSDFAGDLDRRRSI 1170
Query: 805 VGF 807
G+
Sbjct: 1171 TGY 1173
>ref|XP_469192.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|53370655|gb|AAU89150.1| integrase core domain
containing protein [Oryza sativa (japonica
cultivar-group)] gi|40538906|gb|AAR87163.1| putative
polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1322
Score = 268 bits (685), Expect = 6e-70
Identities = 130/243 (53%), Positives = 181/243 (73%), Gaps = 4/243 (1%)
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
+ SLYGLKQSPR+WY+RFD F+L GF RS +DSCVY+ N I YLLLYVDD+L+A+
Sbjct: 953 KRSLYGLKQSPRQWYKRFDSFMLSHGFKRSEFDSCVYIKFVNGSPI-YLLLYVDDMLIAA 1011
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMS 684
SK++I LK++L+ EF+MKDLG AK++LG++I R+R+ G LFLSQ Y+KK +RF M
Sbjct: 1012 KSKEQITTLKKQLSSEFDMKDLGAAKKILGMEITRDRNSGLLFLSQQSYIKKVLQRFNMH 1071
Query: 685 NSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIV 744
++K VSTP+ H KLS QC ++++ + M PY+S VGS+MY MVCS PDL++A+S+V
Sbjct: 1072 DAKPVSTPIAPHFKLSALQCASTDEDVEYMSRVPYSSAVGSLMYAMVCSWPDLSHAMSLV 1131
Query: 745 SRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSL 804
SR+MANPG HW+A++W+ RYL G+ LK+ R + L GYVD+D+A ++D R+SL
Sbjct: 1132 SRYMANPGKEHWKAVQWIFRYLRGTADACLKFGRI---DKGLVGYVDSDFAADLDKRRSL 1188
Query: 805 VGF 807
G+
Sbjct: 1189 TGY 1191
>gb|AAT85194.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1241
Score = 267 bits (683), Expect = 1e-69
Identities = 132/243 (54%), Positives = 179/243 (73%), Gaps = 4/243 (1%)
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
++SLYGLKQSPR+WY+RFD F+L F RS YDSCVY LK + +YLLLYVDD+L+A+
Sbjct: 872 KKSLYGLKQSPRQWYKRFDSFMLSQKFRRSNYDSCVY-LKVVDGSAIYLLLYVDDMLIAA 930
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMS 684
K EI KLK +L+ EFEMKDLG AK++LG++I R R G+L+LSQ Y++K RF M
Sbjct: 931 KDKSEIAKLKAQLSSEFEMKDLGAAKKILGMEITRERHSGKLYLSQKCYIEKVLHRFNMH 990
Query: 685 NSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIV 744
++K VST L H +LS CPQS + + M PY+S V S+MY MVCSRPDL++A+S+V
Sbjct: 991 DAKLVSTLLAAHFRLSSDLCPQSAYDIEYMSRVPYSSAVSSLMYAMVCSRPDLSHALSVV 1050
Query: 745 SRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSL 804
SR+MANPG HW+A++W+ RYL G+ L++ R++ D L GYVD+D+AG++D R+SL
Sbjct: 1051 SRYMANPGKEHWKAVQWIFRYLRGTSSACLQFGRSS---DGLVGYVDSDFAGDLDRRRSL 1107
Query: 805 VGF 807
G+
Sbjct: 1108 TGY 1110
>ref|XP_475663.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|48475188|gb|AAT44257.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1211
Score = 267 bits (682), Expect = 1e-69
Identities = 131/243 (53%), Positives = 178/243 (72%), Gaps = 4/243 (1%)
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
++SLYGLKQSPR+WY+RFD F+ F RS YDSCVY LK + +YLLLYVDD+L+A+
Sbjct: 842 KKSLYGLKQSPRQWYKRFDSFMFSQKFRRSNYDSCVY-LKVVDGSSIYLLLYVDDMLIAA 900
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMS 684
K EI KLK +L+ EFEMKDLG AK++LG++I R R G+L+LSQ GY++K RF M
Sbjct: 901 KDKSEIEKLKAQLSSEFEMKDLGAAKKILGMEITRERHSGKLYLSQKGYIEKVLRRFNMH 960
Query: 685 NSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIV 744
++K VSTPL H +LS CPQS+ + + M PY S VGS+MY MVCSR DL++A+S+V
Sbjct: 961 DAKPVSTPLAAHFRLSSDLCPQSDYDIEYMSRVPYLSVVGSLMYAMVCSRLDLSHALSVV 1020
Query: 745 SRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSL 804
SR+MANPG HW+ ++W+ +YL G+ L++ R+ D L GYVD+D+AG++D R+SL
Sbjct: 1021 SRYMANPGKEHWKVVQWIFKYLRGTSSACLQFGRS---RDGLVGYVDSDFAGDLDRRRSL 1077
Query: 805 VGF 807
G+
Sbjct: 1078 TGY 1080
>ref|XP_472367.1| OSJNBb0012E08.6 [Oryza sativa (japonica cultivar-group)]
gi|38345209|emb|CAD40782.2| OSJNBb0012E08.6 [Oryza
sativa (japonica cultivar-group)]
Length = 375
Score = 266 bits (679), Expect = 3e-69
Identities = 129/243 (53%), Positives = 181/243 (74%), Gaps = 4/243 (1%)
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
+ SLYGLKQSPR+WY+RFD F+L GF RS +DSCVY+ N I YLLLYVDD+L+A+
Sbjct: 20 KRSLYGLKQSPRQWYKRFDSFMLSHGFKRSEFDSCVYIKFVNGSPI-YLLLYVDDMLIAA 78
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMS 684
SK++I LK++L+ EF+MK+LG AK++LG++I R+R+ G LFLSQ Y+KK +RF M
Sbjct: 79 KSKEQITTLKKQLSSEFDMKNLGAAKKILGMEITRDRNSGLLFLSQQSYIKKVLQRFNMH 138
Query: 685 NSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIV 744
++K VSTP+ H KLS QC ++++ + M PY+S VGS+MY MVCSRPDL++A+S+V
Sbjct: 139 DAKPVSTPIVSHFKLSALQCASTDEDVEYMSRVPYSSDVGSLMYAMVCSRPDLSHAMSLV 198
Query: 745 SRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSL 804
SR+MANPG HW+A++W+ RYL G+ LK+ R + L GYVD+D+A ++D R+SL
Sbjct: 199 SRYMANPGKEHWKAVQWIFRYLRGTADACLKFGRT---DKGLVGYVDSDFAADLDKRRSL 255
Query: 805 VGF 807
+
Sbjct: 256 TDY 258
>gb|AAP51772.1| putative pol polyprotein [Oryza sativa (japonica cultivar-group)]
gi|37530366|ref|NP_919485.1| putative pol polyprotein
[Oryza sativa (japonica cultivar-group)]
gi|19551102|gb|AAL91607.1| Putative pol polyprotein
[Oryza sativa (japonica cultivar-group)]
gi|18542925|gb|AAL75759.1| Putative pol polyprotein
[Oryza sativa]
Length = 1005
Score = 261 bits (667), Expect = 8e-68
Identities = 126/243 (51%), Positives = 178/243 (72%), Gaps = 5/243 (2%)
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
+ S YGLKQSPR+WY+RFD F+L GF RS +DSCVY+ N I LLYVDD+L+A+
Sbjct: 637 KRSFYGLKQSPRQWYKRFDLFMLSHGFKRSEFDSCVYIKFVNGSPIY--LLYVDDMLIAA 694
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMS 684
SK++I LK++L+ EF+MKDLG AK++LG++I R+R+ G LFLSQ Y+K +RF M
Sbjct: 695 KSKEQITTLKKQLSSEFDMKDLGAAKKILGMEITRDRNSGLLFLSQQSYIKNVLQRFNMH 754
Query: 685 NSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIV 744
++K VS P+ H KLS+ QC ++++ + M PY+S VGS+MY MVCSRPDL++A+S+V
Sbjct: 755 DAKLVSIPIAPHFKLSVLQCASTDEDVEYMSRVPYSSAVGSLMYAMVCSRPDLSHAMSLV 814
Query: 745 SRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSL 804
SR+MANPG HW+A++W+ RYL G+ LK+ R + L GYVD+D+A ++D R+SL
Sbjct: 815 SRYMANPGKEHWKAVQWIFRYLRGTADACLKFGRT---DKGLVGYVDSDFAADLDKRRSL 871
Query: 805 VGF 807
G+
Sbjct: 872 TGY 874
>gb|AAD19773.1| putative retroelement pol polyprotein [Arabidopsis thaliana]
gi|25301697|pir||B84512 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana
Length = 1335
Score = 258 bits (658), Expect = 8e-67
Identities = 128/243 (52%), Positives = 174/243 (70%), Gaps = 2/243 (0%)
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
+ SLYGLKQSPR+W RFDEF+ + RS YDSCVY K N +YLLLYVDD+L+AS
Sbjct: 963 KRSLYGLKQSPRQWNLRFDEFMRGIKYTRSAYDSCVYFKKCNGDTYIYLLLYVDDMLIAS 1022
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMS 684
++K E+ +LK+ L+ EFEMKDLG AK++LG++I R+RD G L LSQ GY+KK F+M
Sbjct: 1023 ANKSEVNELKQLLSREFEMKDLGDAKKILGMEISRDRDAGLLTLSQEGYVKKVLRSFQMD 1082
Query: 685 NSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIV 744
N+K VSTPLG H KL + E++ + M+ PYA+ +GSIMY M+ +RPDLAY++ ++
Sbjct: 1083 NAKPVSTPLGIHFKLKAATDKEYEEQFERMKIVPYANTIGSIMYSMIGTRPDLAYSLGVI 1142
Query: 745 SRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSL 804
SRFM+ P HWQA+KWVLRY+ G+ K L + + Q++ L GY D+DY N DTR+S+
Sbjct: 1143 SRFMSKPLKDHWQAVKWVLRYMRGTEKKKLCFRK--QEDFLLRGYCDSDYGSNFDTRRSI 1200
Query: 805 VGF 807
G+
Sbjct: 1201 TGY 1203
>gb|AAP53029.1| putative retrotransposon-related protein [Oryza sativa (japonica
cultivar-group)] gi|37532880|ref|NP_920742.1| putative
retrotransposon-related protein [Oryza sativa (japonica
cultivar-group)] gi|22655747|gb|AAN04164.1| Putative
retrotransposon protein [Oryza sativa (japonica
cultivar-group)] gi|16905223|gb|AAL31093.1| putative
retrotransposon-related protein [Oryza sativa]
Length = 1229
Score = 257 bits (657), Expect = 1e-66
Identities = 126/243 (51%), Positives = 179/243 (72%), Gaps = 4/243 (1%)
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
+ SLYGLKQSPR+WY+RFD F+L GF RS +DSCVY+ N I YLLLYVDD+L+A+
Sbjct: 899 KRSLYGLKQSPRQWYKRFDSFMLSHGFKRSEFDSCVYIKFVNGSHI-YLLLYVDDMLIAA 957
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMS 684
SK++I LK++L+ EF+MKDL +K++LG++I R+ + G LFLSQ Y+KK +RF +
Sbjct: 958 KSKEQITTLKKQLSSEFDMKDLVASKKILGMEITRDINSGLLFLSQQSYIKKVLQRFNIH 1017
Query: 685 NSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIV 744
++K VSTP+ H KLS QC ++++ + M PY+S VGS+MY MVCSRP L++A+S+V
Sbjct: 1018 DAKPVSTPIAPHFKLSALQCTSTDEDVEYMSRVPYSSVVGSLMYAMVCSRPVLSHAMSLV 1077
Query: 745 SRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSL 804
SR+MANPG HW+A++W+ RYL G+ LK+ R + L GYVD+D+A ++D R+SL
Sbjct: 1078 SRYMANPGKEHWKAVQWIFRYLRGTADACLKFGRT---DKGLVGYVDSDFAADLDKRRSL 1134
Query: 805 VGF 807
G+
Sbjct: 1135 TGY 1137
>ref|XP_476137.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|48475101|gb|AAT44170.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
gi|46576026|gb|AAT01387.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1175
Score = 257 bits (657), Expect = 1e-66
Identities = 127/243 (52%), Positives = 177/243 (72%), Gaps = 4/243 (1%)
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
+ SLYGLKQSPR+WY+RFD F+L GF RS +DSCVY+ N I YLLLYVDD+L+A+
Sbjct: 806 KRSLYGLKQSPRQWYKRFDSFMLSHGFKRSEFDSCVYIKFVNVSPI-YLLLYVDDMLIAA 864
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMS 684
SK++I LK++L+ EF+MKDL AK++LG++I R+R+ G LFLSQ Y+KK +RF M
Sbjct: 865 KSKEQITTLKKQLSSEFDMKDLDAAKKILGMEITRDRNSGWLFLSQQSYIKKVLQRFNMH 924
Query: 685 NSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIV 744
++K VST + H KLS QC ++++ + M PY+S VGS+MY MVCSR DL++A+S+V
Sbjct: 925 DTKPVSTHIAPHFKLSALQCASTDEDVEYMSRVPYSSVVGSLMYAMVCSRLDLSHAMSLV 984
Query: 745 SRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSL 804
SR+MANPG HW+A++W+ RYL + LK+ R L GYVD+D+A ++D R+SL
Sbjct: 985 SRYMANPGKEHWKAIQWIFRYLRDTANACLKFGRT---NKGLIGYVDSDFAADLDKRRSL 1041
Query: 805 VGF 807
G+
Sbjct: 1042 TGY 1044
>emb|CAE03834.3| OSJNBb0013J13.11 [Oryza sativa (japonica cultivar-group)]
gi|50930401|ref|XP_474728.1| OSJNBb0013J13.11 [Oryza
sativa (japonica cultivar-group)]
Length = 576
Score = 256 bits (655), Expect = 2e-66
Identities = 127/248 (51%), Positives = 177/248 (71%), Gaps = 4/248 (1%)
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
+ SLYGLKQSPR+WY+RFD F+L GF S +DSCVY+ N I YLLLYVDD+L+A+
Sbjct: 286 KRSLYGLKQSPRQWYKRFDSFMLSHGFKWSEFDSCVYIKFVNGSPI-YLLLYVDDMLIAA 344
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMS 684
SK++I LK++L+ EF+MKDLG AK++LG++I R+ + G LFLSQ Y+KK E F M
Sbjct: 345 KSKEQITTLKKQLSSEFDMKDLGAAKKILGMEITRDINFGLLFLSQQSYIKKVLEHFNMH 404
Query: 685 NSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIV 744
++K +STP+ H KLS QC ++++ M PY+S VGS+MY M+CSRPDL++A+S+V
Sbjct: 405 DAKPISTPIAAHFKLSALQCASTDEDVGYMSRVPYSSAVGSLMYAMICSRPDLSHAMSLV 464
Query: 745 SRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSL 804
SR+MANPG HW+A++W RYL G+ K+ R + L GYVD+D+A N+D R+S
Sbjct: 465 SRYMANPGKEHWKAVQWFFRYLRGTTDACSKFGRT---DKGLVGYVDSDFAANLDKRRSF 521
Query: 805 VGFCVYSL 812
+Y+L
Sbjct: 522 TVKVLYAL 529
>gb|AAW22873.1| putative polyprotein [Lycopersicon esculentum]
Length = 687
Score = 249 bits (637), Expect = 2e-64
Identities = 122/243 (50%), Positives = 170/243 (69%), Gaps = 3/243 (1%)
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
++SLYGLKQ+PR+WY +FD F+ + R+ D CVY K +E + L LYVDD+L+
Sbjct: 317 KKSLYGLKQAPRQWYHKFDSFMSNNEYKRTTADPCVYFRKFSEGNFIILCLYVDDMLIVG 376
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMS 684
+ I +LKE L+ F+MKDLGPAK++LG++I R+R G+L+LSQ Y+++ ERF M
Sbjct: 377 QDVEMICRLKEDLSKSFDMKDLGPAKQILGMEIARDRKAGKLWLSQENYIERVLERFNMK 436
Query: 685 NSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIV 744
N+K V+TPL H KLS + CP +E EK+ M PY+S VGS+MY MVC+RPD+A+AV +V
Sbjct: 437 NAKPVNTPLAAHFKLSKRCCPTTEKEKESMSHIPYSSVVGSLMYAMVCTRPDIAHAVGLV 496
Query: 745 SRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSL 804
SR++ANP VHW+A+KW+LRYL G+ L + E LEG+ DAD AG++D RKS
Sbjct: 497 SRYLANPSKVHWEAVKWILRYLRGTSNLSLCF---GGGEPILEGFTDADMAGDLDNRKST 553
Query: 805 VGF 807
G+
Sbjct: 554 SGY 556
>dbj|BAB09923.1| copia-like retrotransposable element [Arabidopsis thaliana]
Length = 1342
Score = 247 bits (631), Expect = 1e-63
Identities = 122/243 (50%), Positives = 168/243 (68%), Gaps = 2/243 (0%)
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
++SLYGLKQSPR+W +RFD F++ +G+ RS Y+ CVY + N+ +YLLLYVDD+L+AS
Sbjct: 970 KKSLYGLKQSPRQWNQRFDSFMINSGYQRSKYNPCVYTQQLNDGSYIYLLLYVDDMLIAS 1029
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMS 684
+KD+I KLKE LN EFEMKDLGPA+++LG++I RNR++G L LSQ Y+ F M
Sbjct: 1030 QNKDQIQKLKESLNREFEMKDLGPARKILGMEITRNREQGILDLSQSEYVAGVLRAFGMD 1089
Query: 685 NSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIV 744
SK TPLG H KL + + M+ PY + +GSIMY M+ SRPDLAY V +V
Sbjct: 1090 QSKVSQTPLGAHFKLRAANEKTLARDAEYMKLVPYPNAIGSIMYSMIGSRPDLAYPVGVV 1149
Query: 745 SRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSL 804
SRFM+ P HWQA+KWV+RY+ G+ L++ + D+ + GY D+DYA ++D R+S+
Sbjct: 1150 SRFMSKPSKEHWQAVKWVMRYMKGTQDTCLRFKK--DDKFEIRGYCDSDYATDLDRRRSI 1207
Query: 805 VGF 807
GF
Sbjct: 1208 TGF 1210
>gb|AAV88069.1| hypothetical retrotransposon [Ipomoea batatas]
Length = 1415
Score = 245 bits (626), Expect = 4e-63
Identities = 117/245 (47%), Positives = 171/245 (69%), Gaps = 3/245 (1%)
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
++SLYGLKQ+PR+WY++F + K G+ ++ D CV++ + ++ + LLLYVDD+L+
Sbjct: 955 KKSLYGLKQAPRQWYKKFTSVMSKHGYKKTSSDHCVFVNRYSDDDFVILLLYVDDMLIVG 1014
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMS 684
+ I +LK+ L+ F MKD+GPAK++LG+ I R+R +L+LSQ Y++K ERF M+
Sbjct: 1015 RNASRIQELKQELSKSFSMKDMGPAKQILGMKIIRDRQNKKLWLSQEKYIEKVLERFHMN 1074
Query: 685 NSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIV 744
+K VSTPL H KL +QCP SE EK+ M+ PY+S VGS+MY MVC+RPD+A+AV +V
Sbjct: 1075 EAKPVSTPLDMHFKLCKKQCPSSEKEKEEMQRVPYSSAVGSLMYAMVCTRPDIAHAVGVV 1134
Query: 745 SRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSL 804
SRF++NPG HW A+KW+LRYL G+ L + + L GY D+D AG++DTRKS
Sbjct: 1135 SRFLSNPGREHWDAVKWILRYLRGTSSLSLCF---GTGKPILTGYTDSDMAGDIDTRKST 1191
Query: 805 VGFCV 809
G+ +
Sbjct: 1192 SGYLI 1196
>pir||T02206 hypothetical protein - common tobacco retrotransposon Tto1
gi|1167523|dbj|BAA11674.1| ORF(AA 1-1338) [Nicotiana
tabacum]
Length = 1338
Score = 245 bits (625), Expect = 6e-63
Identities = 119/244 (48%), Positives = 173/244 (70%), Gaps = 3/244 (1%)
Query: 566 ESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMASS 625
+SLYGLKQ+PR+WY++F+ + + G+ ++ D CV+ K ++ + LLLYVDD+L+
Sbjct: 959 KSLYGLKQAPRQWYKKFESVMGQHGYKKTTSDHCVFAQKFSDDDFIILLLYVDDMLIVGR 1018
Query: 626 SKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMSN 685
+ I LKE+L+ F MKDLGPAK++LG+ I R+R+ +L+LSQ Y++K +RF M
Sbjct: 1019 NVSRINSLKEQLSKFFAMKDLGPAKQILGMRIMRDREAKKLWLSQEKYIEKVLQRFNMEK 1078
Query: 686 SKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIVS 745
+K VS PL +H +LS +Q P ++DE++ ME PYAS VGS+MY MVC+RPD+A+AV +VS
Sbjct: 1079 TKAVSCPLANHFRLSTKQSPSTDDERRKMERIPYASAVGSLMYAMVCTRPDIAHAVGVVS 1138
Query: 746 RFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSLV 805
RF++NPG HW A+KW+LRYL G+ K L + +D L GY DAD AG+VD+RKS
Sbjct: 1139 RFLSNPGKEHWDAVKWILRYLRGTSKLCLCF---GEDNPVLVGYTDADMAGDVDSRKSTS 1195
Query: 806 GFCV 809
G+ +
Sbjct: 1196 GYLI 1199
>emb|CAA31653.1| polyprotein [Arabidopsis thaliana] gi|99721|pir||S05465
retrovirus-related polyprotein - Arabidopsis thaliana
retrotransposon Ta1-3
Length = 1291
Score = 244 bits (623), Expect = 1e-62
Identities = 120/243 (49%), Positives = 171/243 (69%), Gaps = 5/243 (2%)
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
++SLYGLKQSPR+W +RF+ F++ F+RS +D+CVY+ + +E+ LYLLLYVDD+L+A
Sbjct: 993 KKSLYGLKQSPRQWNKRFNRFMIDQNFIRSEHDACVYVKQVSEQEHLYLLLYVDDMLIAG 1052
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMS 684
SK EI K+KE+L+ EFEMKD+GPA R+LGIDI R+ G L +SQ Y+ +RF M+
Sbjct: 1053 KSKSEINKVKEQLSMEFEMKDMGPASRILGIDIIRDMKNGVLRMSQASYIHNVVQRFNMA 1112
Query: 685 NSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIV 744
+K +P+G H KL+ + +DE PYAS VGSIMY M+ RPDLAY + +V
Sbjct: 1113 EAKVTRSPIGAHFKLA---AVRDDDECIDNNAVPYASAVGSIMYAMIGIRPDLAYVICLV 1169
Query: 745 SRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSL 804
SR+MA PG +HW+A+KW+LRY+ GS L +T+ + E + GY D+DYA ++D R+S+
Sbjct: 1170 SRYMARPGSIHWEAVKWILRYMRGSQDLNLVFTK--EKEFRVTGYCDSDYAADLDRRRSV 1227
Query: 805 VGF 807
G+
Sbjct: 1228 SGY 1230
>emb|CAA37920.1| unnamed protein product [Arabidopsis thaliana] gi|99730|pir||S23315
hypothetical protein 3 - Arabidopsis thaliana
retrotransposon Ta1-2 (strain Kashmir) (fragment)
Length = 494
Score = 243 bits (619), Expect = 3e-62
Identities = 120/243 (49%), Positives = 171/243 (69%), Gaps = 5/243 (2%)
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
++SLYGLKQSPR+W +RF F++ F+RS +D+CVY+ + E+ LYLLLYVDD+L+A
Sbjct: 218 KKSLYGLKQSPRQWNKRFSRFMIDQNFIRSEHDACVYVKQAGEQDHLYLLLYVDDMLIAG 277
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMS 684
SK EI K+KE+L+ EFEMKD+GPA R+LGIDI R+R G L +SQ Y+ +RF M+
Sbjct: 278 KSKSEINKVKEQLSMEFEMKDMGPASRILGIDIIRDRKNGVLRMSQARYIHNVVQRFNMA 337
Query: 685 NSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIV 744
+K +P+G H KL+ + +DE PYAS VG+IMY M+ +RPDLAYA+ +V
Sbjct: 338 EAKVTRSPIGAHFKLA---AVRDDDECIDNNALPYASAVGNIMYAMIGTRPDLAYAICLV 394
Query: 745 SRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSL 804
SR+MA PG +HW+A+KW+LRY+ GS L +T+ + E + Y D+DYA ++D R+S+
Sbjct: 395 SRYMARPGNIHWEAVKWILRYMRGSQDLNLVFTK--EKEFRVTRYCDSDYAADLDRRRSV 452
Query: 805 VGF 807
G+
Sbjct: 453 SGY 455
>emb|CAA32025.1| unnamed protein product [Nicotiana tabacum]
gi|130582|sp|P10978|POLX_TOBAC Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
; Reverse transcriptase ; Endonuclease]
Length = 1328
Score = 242 bits (618), Expect = 4e-62
Identities = 119/242 (49%), Positives = 165/242 (68%), Gaps = 3/242 (1%)
Query: 566 ESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMASS 625
+SLYGLKQ+PR+WY +FD F+ ++++ D CVY + +E + LLLYVDD+L+
Sbjct: 958 KSLYGLKQAPRQWYMKFDSFMKSQTYLKTYSDPCVYFKRFSENNFIILLLYVDDMLIVGK 1017
Query: 626 SKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMSN 685
K I KLK L+ F+MKDLGPA+++LG+ I R R +L+LSQ Y+++ ERF M N
Sbjct: 1018 DKGLIAKLKGDLSKSFDMKDLGPAQQILGMKIVRERTSRKLWLSQEKYIERVLERFNMKN 1077
Query: 686 SKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIVS 745
+K VSTPL H KLS + CP + +EK M PY+S VGS+MY MVC+RPD+A+AV +VS
Sbjct: 1078 AKPVSTPLAGHLKLSKKMCPTTVEEKGNMAKVPYSSAVGSLMYAMVCTRPDIAHAVGVVS 1137
Query: 746 RFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSLV 805
RF+ NPG HW+A+KW+LRYL G+ L + + L+GY DAD AG++D RKS
Sbjct: 1138 RFLENPGKEHWEAVKWILRYLRGTTGDCLCF---GGSDPILKGYTDADMAGDIDNRKSST 1194
Query: 806 GF 807
G+
Sbjct: 1195 GY 1196
>emb|CAA37917.1| reverse transcriptase [Arabidopsis thaliana] gi|99755|pir||S23312
retrovirus-related polyprotein KAS-1 - Arabidopsis
thaliana retrotransposon Ta1 (fragment)
Length = 587
Score = 242 bits (617), Expect = 5e-62
Identities = 119/240 (49%), Positives = 169/240 (69%), Gaps = 5/240 (2%)
Query: 568 LYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMASSSK 627
LYGLKQSPR+W +RF+ F++ F+RS +D+CVY+ + E+ LYLLLYVDD+L+A +K
Sbjct: 220 LYGLKQSPRQWNKRFNRFMIDQNFIRSEHDACVYVKQVTEQEHLYLLLYVDDMLIAGKNK 279
Query: 628 DEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMSNSK 687
EI K+KE+L+ EFEMKD+GPA R+LGIDI R+R G L +SQ Y+ +RF M+ +K
Sbjct: 280 SEINKVKEQLSMEFEMKDMGPASRILGIDIIRDRKNGVLRMSQASYIHNVVQRFNMAEAK 339
Query: 688 TVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIVSRF 747
+P+G H KL+ + +DE PYAS VGSIMY M+ +RPDLAY + +VSR+
Sbjct: 340 VTRSPIGAHFKLA---AVRDDDECIDNNVVPYASAVGSIMYAMIGTRPDLAYDICLVSRY 396
Query: 748 MANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSLVGF 807
MA PG +HW+A+KW+LRY+ GS L +T+ + E + GY D+DYA ++D R+S+ G+
Sbjct: 397 MARPGSIHWEAVKWILRYMRGSQDLNLVFTK--EKEFRVTGYCDSDYAADLDRRRSVSGY 454
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.356 0.161 0.595
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,397,459,948
Number of Sequences: 2540612
Number of extensions: 55858894
Number of successful extensions: 198370
Number of sequences better than 10.0: 1519
Number of HSP's better than 10.0 without gapping: 1362
Number of HSP's successfully gapped in prelim test: 157
Number of HSP's that attempted gapping in prelim test: 194389
Number of HSP's gapped (non-prelim): 1870
length of query: 937
length of database: 863,360,394
effective HSP length: 138
effective length of query: 799
effective length of database: 512,755,938
effective search space: 409691994462
effective search space used: 409691994462
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 80 (35.4 bits)
Medicago: description of AC130802.9


