

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130801.12 + phase: 0
(124 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
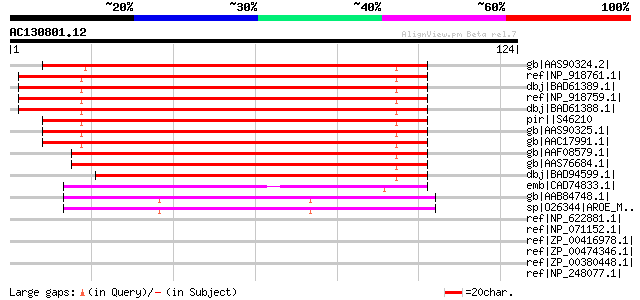
Score E
Sequences producing significant alignments: (bits) Value
gb|AAS90324.2| 3-dehydroquinate dehydratase / shikimate dehydrog... 128 3e-29
ref|NP_918761.1| putative 3-dehydroquinate dehydratase / shikima... 94 1e-18
dbj|BAD61389.1| putative dehydroquinate dehydratase [Oryza sativ... 94 1e-18
ref|NP_918759.1| putative 3-dehydroquinate dehydratase [Oryza sa... 92 3e-18
dbj|BAD61388.1| putative dehydroquinate dehydratase [Oryza sativ... 92 3e-18
pir||S46210 3-dehydroquinate dehydratase (EC 4.2.1.10) / shikima... 91 6e-18
gb|AAS90325.1| 3-dehydroquinate dehydratase / shikimate dehydrog... 91 6e-18
gb|AAC17991.1| dehydroquinate dehydratase/shikimate:NADP oxidore... 91 6e-18
gb|AAF08579.1| putative dehydroquinase shikimate dehydrogenase [... 82 3e-15
gb|AAS76684.1| At3g06350 [Arabidopsis thaliana] 82 3e-15
dbj|BAD94599.1| putative dehydroquinase shikimate dehydrogenase ... 74 8e-13
emb|CAD74833.1| 3-dehydroquinate dehydratase / shikimate 5-dehyd... 50 1e-05
gb|AAB84748.1| shikimate 5-dehydrogenase [Methanothermobacter th... 46 2e-04
sp|O26344|AROE_METTH Shikimate dehydrogenase 46 2e-04
ref|NP_622881.1| Shikimate 5-dehydrogenase [Thermoanaerobacter t... 42 0.003
ref|NP_071152.1| shikimate 5-dehydrogenase (aroE) [Archaeoglobus... 42 0.004
ref|ZP_00416978.1| Shikimate/quinate 5-dehydrogenase [Azotobacte... 40 0.010
ref|ZP_00474346.1| Shikimate 5-dehydrogenase [Chromohalobacter s... 40 0.017
ref|ZP_00380448.1| COG0169: Shikimate 5-dehydrogenase [Brevibact... 40 0.017
ref|NP_248077.1| shikimate 5-dehydrogenase (aroE) [Methanocaldoc... 40 0.017
>gb|AAS90324.2| 3-dehydroquinate dehydratase / shikimate dehydrogenase isoform 2
[Nicotiana tabacum]
Length = 521
Score = 128 bits (322), Expect = 3e-29
Identities = 68/101 (67%), Positives = 81/101 (79%), Gaps = 7/101 (6%)
Query: 9 PLDGRLFVR--AGGAGKALAFGAKTRGARIVIFDIDFDKSRSLACAVFGEVQSYKNLVNF 66
PL G+LFV AGGAG+ALAFGAK+R A IVIFDIDFD++++LA AV GE ++NL +F
Sbjct: 367 PLAGKLFVLVGAGGAGRALAFGAKSRRAEIVIFDIDFDRAKALAAAVSGEALPFENLASF 426
Query: 67 QPVKGAILANATPIGMHPDTDRIPVAE-----YQLVFDAVY 102
QP KGAILANATPIGMHP+ DRIPV+E Y +VFDAVY
Sbjct: 427 QPEKGAILANATPIGMHPNKDRIPVSEASLKDYVVVFDAVY 467
>ref|NP_918761.1| putative 3-dehydroquinate dehydratase / shikimate 5-dehydrogenase
[Oryza sativa (japonica cultivar-group)]
Length = 500
Score = 93.6 bits (231), Expect = 1e-18
Identities = 53/107 (49%), Positives = 70/107 (64%), Gaps = 7/107 (6%)
Query: 3 KHLQVHPLDGRLFV--RAGGAGKALAFGAKTRGARIVIFDIDFDKSRSLACAVFGEVQSY 60
K+ V PL GRL V AGGAGKA+A+GAK +GARIV+ + ++K+ SLA AV G
Sbjct: 349 KNAAVTPLAGRLLVVVGAGGAGKAIAYGAKEKGARIVVANRTYEKAVSLAAAVGGHALRL 408
Query: 61 KNLVNFQPVKGAILANATPIGMHPDTDRIPVAE-----YQLVFDAVY 102
L F+P +G ILANAT +GM+P+ D P+ + Y +VFDAVY
Sbjct: 409 AELETFRPEEGMILANATSLGMYPNVDGTPIPKQALSFYDVVFDAVY 455
>dbj|BAD61389.1| putative dehydroquinate dehydratase [Oryza sativa (japonica
cultivar-group)]
Length = 657
Score = 93.6 bits (231), Expect = 1e-18
Identities = 53/107 (49%), Positives = 70/107 (64%), Gaps = 7/107 (6%)
Query: 3 KHLQVHPLDGRLFV--RAGGAGKALAFGAKTRGARIVIFDIDFDKSRSLACAVFGEVQSY 60
K+ V PL GRL V AGGAGKA+A+GAK +GARIV+ + ++K+ SLA AV G
Sbjct: 432 KNAAVTPLAGRLLVVVGAGGAGKAIAYGAKEKGARIVVANRTYEKAVSLAAAVGGHALRL 491
Query: 61 KNLVNFQPVKGAILANATPIGMHPDTDRIPVAE-----YQLVFDAVY 102
L F+P +G ILANAT +GM+P+ D P+ + Y +VFDAVY
Sbjct: 492 AELETFRPEEGMILANATSLGMYPNVDGTPIPKQALSFYDVVFDAVY 538
>ref|NP_918759.1| putative 3-dehydroquinate dehydratase [Oryza sativa (japonica
cultivar-group)]
Length = 531
Score = 92.0 bits (227), Expect = 3e-18
Identities = 51/107 (47%), Positives = 69/107 (63%), Gaps = 7/107 (6%)
Query: 3 KHLQVHPLDGRLFV--RAGGAGKALAFGAKTRGARIVIFDIDFDKSRSLACAVFGEVQSY 60
K + PL GRL V AGGAGKA+A+GAK +GAR+V+ + ++K+ SLA AV G
Sbjct: 370 KDAAISPLAGRLVVVVGAGGAGKAIAYGAKEKGARVVVANRTYEKAVSLAAAVGGHALRL 429
Query: 61 KNLVNFQPVKGAILANATPIGMHPDTDRIPVAE-----YQLVFDAVY 102
L F+P +G ILANAT +GM+P+ D P+ + Y +VFDAVY
Sbjct: 430 AELETFRPEEGMILANATSLGMYPNVDGTPIPKKALSFYDVVFDAVY 476
>dbj|BAD61388.1| putative dehydroquinate dehydratase [Oryza sativa (japonica
cultivar-group)]
Length = 509
Score = 92.0 bits (227), Expect = 3e-18
Identities = 51/107 (47%), Positives = 69/107 (63%), Gaps = 7/107 (6%)
Query: 3 KHLQVHPLDGRLFV--RAGGAGKALAFGAKTRGARIVIFDIDFDKSRSLACAVFGEVQSY 60
K + PL GRL V AGGAGKA+A+GAK +GAR+V+ + ++K+ SLA AV G
Sbjct: 348 KDAAISPLAGRLVVVVGAGGAGKAIAYGAKEKGARVVVANRTYEKAVSLAAAVGGHALRL 407
Query: 61 KNLVNFQPVKGAILANATPIGMHPDTDRIPVAE-----YQLVFDAVY 102
L F+P +G ILANAT +GM+P+ D P+ + Y +VFDAVY
Sbjct: 408 AELETFRPEEGMILANATSLGMYPNVDGTPIPKKALSFYDVVFDAVY 454
>pir||S46210 3-dehydroquinate dehydratase (EC 4.2.1.10) / shikimate
5-dehydrogenase (EC 1.1.1.25) precursor - common tobacco
(fragment) gi|535771|gb|AAA34069.1| dehydroquinate
dehydratase/shikimate dehydrogenase
Length = 579
Score = 90.9 bits (224), Expect = 6e-18
Identities = 52/101 (51%), Positives = 64/101 (62%), Gaps = 7/101 (6%)
Query: 9 PLDGRLFV--RAGGAGKALAFGAKTRGARIVIFDIDFDKSRSLACAVFGEVQSYKNLVNF 66
PL G+LFV AGGAGKALA+GAK +GAR+VI + ++++R LA V G+ S L NF
Sbjct: 383 PLAGKLFVVIGAGGAGKALAYGAKEKGARVVIANRTYERARELADVVGGQALSLDELSNF 442
Query: 67 QPVKGAILANATPIGMHPDTDRIPVAE-----YQLVFDAVY 102
P ILAN T IGM P D P+ + Y LVFDAVY
Sbjct: 443 HPENDMILANTTSIGMQPKVDDTPIFKEALRYYSLVFDAVY 483
>gb|AAS90325.1| 3-dehydroquinate dehydratase / shikimate dehydrogenase isoform 1
[Nicotiana tabacum]
Length = 569
Score = 90.9 bits (224), Expect = 6e-18
Identities = 52/101 (51%), Positives = 64/101 (62%), Gaps = 7/101 (6%)
Query: 9 PLDGRLFV--RAGGAGKALAFGAKTRGARIVIFDIDFDKSRSLACAVFGEVQSYKNLVNF 66
PL G+LFV AGGAGKALA+GAK +GAR+VI + ++++R LA V G+ S L NF
Sbjct: 373 PLAGKLFVVIGAGGAGKALAYGAKEKGARVVIANRTYERARELADVVGGQALSLDELSNF 432
Query: 67 QPVKGAILANATPIGMHPDTDRIPVAE-----YQLVFDAVY 102
P ILAN T IGM P D P+ + Y LVFDAVY
Sbjct: 433 HPENDMILANTTSIGMQPKVDDTPIFKEALRYYSLVFDAVY 473
>gb|AAC17991.1| dehydroquinate dehydratase/shikimate:NADP oxidoreductase
[Lycopersicon esculentum] gi|3169888|gb|AAC17993.1|
dehydroquinate dehydratase/shikimate:NADP oxidoreductase
[Lycopersicon esculentum] gi|7436862|pir||T06264
3-dehydroquinate dehydratase (EC 4.2.1.10) / shikimate
5-dehydrogenase (EC 1.1.1.25) - tomato
Length = 545
Score = 90.9 bits (224), Expect = 6e-18
Identities = 51/101 (50%), Positives = 64/101 (62%), Gaps = 7/101 (6%)
Query: 9 PLDGRLFV--RAGGAGKALAFGAKTRGARIVIFDIDFDKSRSLACAVFGEVQSYKNLVNF 66
PL G+LFV AGGAGKA+A+GAK +GAR+VI + ++++R LA V E S L NF
Sbjct: 392 PLAGKLFVVIGAGGAGKAIAYGAKEKGARVVIANRTYERARELAIVVGAEALSLDELSNF 451
Query: 67 QPVKGAILANATPIGMHPDTDRIPVAE-----YQLVFDAVY 102
P ILAN T IGM P D P+++ Y LVFDAVY
Sbjct: 452 HPENDMILANTTSIGMQPKVDDTPISKEALKHYSLVFDAVY 492
>gb|AAF08579.1| putative dehydroquinase shikimate dehydrogenase [Arabidopsis
thaliana] gi|15230703|ref|NP_187286.1| dehydroquinate
dehydratase, putative / shikimate dehydrogenase,
putative [Arabidopsis thaliana]
Length = 603
Score = 82.0 bits (201), Expect = 3e-15
Identities = 41/92 (44%), Positives = 62/92 (66%), Gaps = 5/92 (5%)
Query: 16 VRAGGAGKALAFGAKTRGARIVIFDIDFDKSRSLACAVFGEVQSYKNLVNFQPVKGAILA 75
+ AGGAGKALA+GAK +GA++VI + ++++ LA A+ G+ S +L N+ P G +LA
Sbjct: 459 IGAGGAGKALAYGAKEKGAKVVIANRTYERALELAEAIGGKALSLTDLDNYHPEDGMVLA 518
Query: 76 NATPIGMHPDTDRIPVAE-----YQLVFDAVY 102
N T +GM P+ + P+++ Y LVFDAVY
Sbjct: 519 NTTSMGMQPNVEETPISKDALKHYALVFDAVY 550
>gb|AAS76684.1| At3g06350 [Arabidopsis thaliana]
Length = 603
Score = 82.0 bits (201), Expect = 3e-15
Identities = 41/92 (44%), Positives = 62/92 (66%), Gaps = 5/92 (5%)
Query: 16 VRAGGAGKALAFGAKTRGARIVIFDIDFDKSRSLACAVFGEVQSYKNLVNFQPVKGAILA 75
+ AGGAGKALA+GAK +GA++VI + ++++ LA A+ G+ S +L N+ P G +LA
Sbjct: 459 IGAGGAGKALAYGAKEKGAKVVIANRTYERALELAEAIGGKALSLTDLDNYHPEDGMVLA 518
Query: 76 NATPIGMHPDTDRIPVAE-----YQLVFDAVY 102
N T +GM P+ + P+++ Y LVFDAVY
Sbjct: 519 NTTSMGMQPNVEETPISKDALKHYALVFDAVY 550
>dbj|BAD94599.1| putative dehydroquinase shikimate dehydrogenase [Arabidopsis
thaliana]
Length = 139
Score = 73.9 bits (180), Expect = 8e-13
Identities = 37/86 (43%), Positives = 57/86 (66%), Gaps = 5/86 (5%)
Query: 22 GKALAFGAKTRGARIVIFDIDFDKSRSLACAVFGEVQSYKNLVNFQPVKGAILANATPIG 81
GKALA+GAK +GA++VI + ++++ LA A+ G+ S +L N+ P G +LAN T +G
Sbjct: 1 GKALAYGAKEKGAKVVIANRTYERALKLAEAIGGKALSLTDLDNYHPEDGMVLANTTSMG 60
Query: 82 MHPDTDRIPVAE-----YQLVFDAVY 102
M P+ + P+++ Y LVFDAVY
Sbjct: 61 MQPNVEETPISKDALKHYALVFDAVY 86
>emb|CAD74833.1| 3-dehydroquinate dehydratase / shikimate 5-dehydrogenase precursor
[Rhodopirellula baltica SH 1]
gi|32474293|ref|NP_867287.1| 3-dehydroquinate
dehydratase / shikimate 5-dehydrogenase precursor
[Rhodopirellula baltica SH 1]
Length = 496
Score = 50.1 bits (118), Expect = 1e-05
Identities = 30/94 (31%), Positives = 47/94 (49%), Gaps = 8/94 (8%)
Query: 14 LFVRAGGAGKALAFGAKTRGARIVIFDIDFDKSRSLACAVFGEVQSYKNLVNFQPVKGAI 73
L + AGG +A+A+G + R + I +++ LA + V ++ K +
Sbjct: 346 LVLGAGGVSRAIAWGLRQRKCDVTISSRTRERAEMLAADIGCRVIDWEER---HEAKVQL 402
Query: 74 LANATPIGMHPDTDRIP-----VAEYQLVFDAVY 102
L N TPIGMHPD D P + ++ +VFD VY
Sbjct: 403 LINGTPIGMHPDVDNTPFNQSALNQFMVVFDTVY 436
>gb|AAB84748.1| shikimate 5-dehydrogenase [Methanothermobacter thermautotrophicus
str. Delta H] gi|15678270|ref|NP_275385.1| shikimate
5-dehydrogenase [Methanothermobacter thermautotrophicus
str. Delta H] gi|7431134|pir||C69130 shikimate
5-dehydrogenase - Methanobacterium thermoautotrophicum
(strain Delta H)
Length = 260
Score = 46.2 bits (108), Expect = 2e-04
Identities = 30/97 (30%), Positives = 53/97 (53%), Gaps = 6/97 (6%)
Query: 14 LFVRAGGAGKALAFGAKTRGAR-IVIFDIDFDKSRSLACAVFGEVQSYKNLVNFQPVKGA 72
L + AGGA +A AF +GA I I + +K+R LA + ++ + ++ ++G+
Sbjct: 103 LILGAGGAARACAFQLAEKGASDITILNRTPEKARLLAEDMADKLGFEASYGGYELIQGS 162
Query: 73 -----ILANATPIGMHPDTDRIPVAEYQLVFDAVYLH 104
IL + TP+GMHP TD P+ +L+ + + +H
Sbjct: 163 VKSADILIDTTPVGMHPHTDDRPLVGAELMHEGLVVH 199
>sp|O26344|AROE_METTH Shikimate dehydrogenase
Length = 283
Score = 46.2 bits (108), Expect = 2e-04
Identities = 30/97 (30%), Positives = 53/97 (53%), Gaps = 6/97 (6%)
Query: 14 LFVRAGGAGKALAFGAKTRGAR-IVIFDIDFDKSRSLACAVFGEVQSYKNLVNFQPVKGA 72
L + AGGA +A AF +GA I I + +K+R LA + ++ + ++ ++G+
Sbjct: 126 LILGAGGAARACAFQLAEKGASDITILNRTPEKARLLAEDMADKLGFEASYGGYELIQGS 185
Query: 73 -----ILANATPIGMHPDTDRIPVAEYQLVFDAVYLH 104
IL + TP+GMHP TD P+ +L+ + + +H
Sbjct: 186 VKSADILIDTTPVGMHPHTDDRPLVGAELMHEGLVVH 222
>ref|NP_622881.1| Shikimate 5-dehydrogenase [Thermoanaerobacter tengcongensis MB4]
gi|20516261|gb|AAM24485.1| Shikimate 5-dehydrogenase
[Thermoanaerobacter tengcongensis MB4]
gi|24211499|sp|Q8RAG2|AROE_THETN Shikimate dehydrogenase
Length = 280
Score = 42.0 bits (97), Expect = 0.003
Identities = 30/95 (31%), Positives = 47/95 (48%), Gaps = 10/95 (10%)
Query: 18 AGGAGKALAFGAKTRGAR-IVIFDIDFDKSRSLACAVFGEVQSYKNLVNFQPVKGA---- 72
AGGA KA+ F G IVI + +K+++LA + E + + + + V+
Sbjct: 132 AGGASKAICFALAREGVESIVIANRTLNKAKALAEYIREEFKMKCDYCSIEEVEKFNEID 191
Query: 73 ILANATPIGMHPDTDRIPVAE-----YQLVFDAVY 102
IL N T +GMHP+ PV+E V+D +Y
Sbjct: 192 ILINTTSVGMHPEVGNSPVSEEVVAKANFVYDLIY 226
>ref|NP_071152.1| shikimate 5-dehydrogenase (aroE) [Archaeoglobus fulgidus DSM 4304]
gi|2648193|gb|AAB88930.1| shikimate 5-dehydrogenase
(aroE) [Archaeoglobus fulgidus DSM 4304]
gi|9087116|sp|O27957|AROE_ARCFU Shikimate dehydrogenase
Length = 269
Score = 41.6 bits (96), Expect = 0.004
Identities = 33/98 (33%), Positives = 48/98 (48%), Gaps = 10/98 (10%)
Query: 14 LFVRAGGAGKALAFGAKTRGARIVIFDIDFDKSRSLACAV--FGEVQSYKNLVNFQPVKG 71
L V AGGAGKA A G+ +++ + +K R + +GE + L + +KG
Sbjct: 117 LVVGAGGAGKAAALALLDMGSTVIVANRTEEKGREAVEMLRRYGEC-IFWPLSRVEELKG 175
Query: 72 A--ILANATPIGMHPDTDRIPVAE-----YQLVFDAVY 102
++ NATP+GM IPV +LVFD VY
Sbjct: 176 KVDVVVNATPLGMRGFKAEIPVPPSMLDGVELVFDTVY 213
>ref|ZP_00416978.1| Shikimate/quinate 5-dehydrogenase [Azotobacter vinelandii AvOP]
gi|67087151|gb|EAM06618.1| Shikimate/quinate
5-dehydrogenase [Azotobacter vinelandii AvOP]
Length = 372
Score = 40.4 bits (93), Expect = 0.010
Identities = 27/91 (29%), Positives = 51/91 (55%), Gaps = 5/91 (5%)
Query: 16 VRAGGAGKALAFGAKTRGARIVIFDIDFDKSRSLACAVFGEVQSYKNLVNFQPVKGAILA 75
+ AGGA KA+A+G + GAR+ +F+ ++ R+LA ++ EV ++ +F ++
Sbjct: 226 IGAGGAAKAVAYGCRQAGARVEVFNRSAERGRTLAESL--EVGFGGSIDDFAADAFDVVI 283
Query: 76 NATPIGM-HPDTDRIP--VAEYQLVFDAVYL 103
NAT G P T+ + +A + +V D ++
Sbjct: 284 NATSAGFRQPGTNPLDGRLASHLVVMDVAFI 314
>ref|ZP_00474346.1| Shikimate 5-dehydrogenase [Chromohalobacter salexigens DSM 3043]
gi|67518347|gb|EAM22322.1| Shikimate 5-dehydrogenase
[Chromohalobacter salexigens DSM 3043]
Length = 288
Score = 39.7 bits (91), Expect = 0.017
Identities = 31/100 (31%), Positives = 46/100 (46%), Gaps = 11/100 (11%)
Query: 14 LFVRAGGAGKALAFGAKTRGA-RIVIFDIDFDK----SRSLACAVFGEVQSYKNLVNFQP 68
L + GG GKA+AFG GA +++ D+D +K +R L A F +
Sbjct: 124 LLIGTGGVGKAVAFGLAKLGADEVLLMDLDLEKAERLARELDHAGFSAAAVAPEALEATA 183
Query: 69 VKGAILANATPIGMHPDTDRIP-----VAEYQLVFDAVYL 103
+ + N TPIG H + P + Q +FDAVY+
Sbjct: 184 ARCDGVVNCTPIG-HEKSPGYPLPPALIRADQWIFDAVYV 222
>ref|ZP_00380448.1| COG0169: Shikimate 5-dehydrogenase [Brevibacterium linens BL2]
Length = 280
Score = 39.7 bits (91), Expect = 0.017
Identities = 24/73 (32%), Positives = 43/73 (58%), Gaps = 10/73 (13%)
Query: 16 VRAGGAGKALAFGAKTRGAR-IVIFDIDFDKSRSLACAVFGEVQSYKNLVNFQPVKGAI- 73
+ AGGAG+A+AF G + +VI DI ++++ LA + +V++ ++ ++ AI
Sbjct: 133 IGAGGAGRAVAFALSESGVKDLVIADISAERAQDLAAEIGDDVRA----ISMSELEPAIT 188
Query: 74 ----LANATPIGM 82
+ NATP+GM
Sbjct: 189 SADGIVNATPVGM 201
>ref|NP_248077.1| shikimate 5-dehydrogenase (aroE) [Methanocaldococcus jannaschii DSM
2661] gi|1591730|gb|AAB99086.1| shikimate
5-dehydrogenase (aroE) [Methanocaldococcus jannaschii
DSM 2661] gi|2492956|sp|Q58484|AROE_METJA Shikimate
dehydrogenase
Length = 282
Score = 39.7 bits (91), Expect = 0.017
Identities = 33/99 (33%), Positives = 50/99 (50%), Gaps = 17/99 (17%)
Query: 18 AGGAGKALAFGAKTRGARIVIFDIDFDKSRSLACAV-------FGEVQSYKNL-VNFQPV 69
AGGA +A+AF + I+I + +K+ +LA + FGE + L V+ V
Sbjct: 131 AGGAARAVAFEL-AKDNNIIIANRTVEKAEALAKEIAEKLNKKFGEEVKFSGLDVDLDGV 189
Query: 70 KGAILANATPIGMHPDTDRIPVA------EYQLVFDAVY 102
I+ NATPIGM+P+ D P+ E +V D +Y
Sbjct: 190 D--IIINATPIGMYPNIDVEPIVKAEKLREDMVVMDLIY 226
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.327 0.143 0.438
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 211,832,006
Number of Sequences: 2540612
Number of extensions: 7709201
Number of successful extensions: 31628
Number of sequences better than 10.0: 174
Number of HSP's better than 10.0 without gapping: 40
Number of HSP's successfully gapped in prelim test: 134
Number of HSP's that attempted gapping in prelim test: 31564
Number of HSP's gapped (non-prelim): 174
length of query: 124
length of database: 863,360,394
effective HSP length: 100
effective length of query: 24
effective length of database: 609,299,194
effective search space: 14623180656
effective search space used: 14623180656
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 68 (30.8 bits)
Medicago: description of AC130801.12


