

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC129091.5 - phase: 0 /pseudo
(980 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
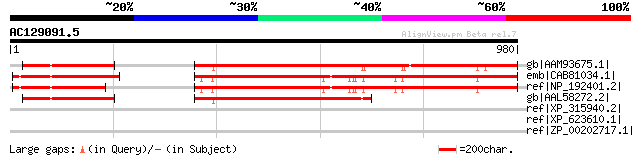
Score E
Sequences producing significant alignments: (bits) Value
gb|AAM93675.1| putative glycine-rich protein [Oryza sativa (japo... 716 0.0
emb|CAB81034.1| hypothetical protein [Arabidopsis thaliana] gi|2... 654 0.0
ref|NP_192401.2| expressed protein [Arabidopsis thaliana] 654 0.0
gb|AAL58272.2| hypothetical protein, 3'-partial [Oryza sativa (j... 407 e-111
ref|XP_315940.2| ENSANGP00000013473 [Anopheles gambiae str. PEST... 38 1.9
ref|XP_623610.1| PREDICTED: similar to Thyroid hormone receptor ... 37 2.5
ref|ZP_00202717.1| COG3181: Uncharacterized protein conserved in... 37 2.5
>gb|AAM93675.1| putative glycine-rich protein [Oryza sativa (japonica
cultivar-group)] gi|31432881|gb|AAP54457.1| putative
glycine-rich protein [Oryza sativa (japonica
cultivar-group)] gi|37535736|ref|NP_922170.1| putative
glycine-rich protein [Oryza sativa (japonica
cultivar-group)]
Length = 1291
Score = 716 bits (1849), Expect = 0.0
Identities = 393/736 (53%), Positives = 485/736 (65%), Gaps = 115/736 (15%)
Query: 357 LRPVVLNPIFGS---MDGNPPMRTVWESKVDISIPATDD--------------------- 392
++PVVL+PIFGS G PP +TVW ++V+ SIP ++D
Sbjct: 511 VQPVVLHPIFGSPANFGGQPPTQTVWSTRVNKSIPPSEDLKNPQSYVPMPTTSDERSSSE 570
Query: 393 ----KSKGIHFNPFDLPKYLGTLARVVFSAQGGEIAVAFFQGRVRIFSGSNFEPVTNYEI 448
++ + F+P+DLP + LA++V+SA GGE+AVAF +G V IFSG NFE V +Y +
Sbjct: 571 CSVDRANRLSFDPYDLPNDVRQLAQIVYSAHGGEVAVAFLRGGVHIFSGPNFEQVDSYHV 630
Query: 449 NVGSSISVPAFSATSCCSASVWHDTSKGEAMLKIIRVLPPPFPIGQEKATSSTWEHAISE 508
NVGS+I+ PAFS++ CC ASVWHDT K +LKIIRVLPP Q K +S+ WE AI++
Sbjct: 631 NVGSAIAPPAFSSSGCCLASVWHDTLKDRTILKIIRVLPPAILNAQTKVSSAVWERAIAD 690
Query: 509 -RFWFSLLVGVDWWDVVGCTQRAAEEGIVSVNGVIAVLDADFHSLPTAQHRQQYCLGLDM 567
RFW+SLL GVDWWD VGCTQ AAE+GIVS+N VIA+LDADFH LPT Q RQQ+C LD
Sbjct: 691 SRFWWSLLAGVDWWDAVGCTQSAAEDGIVSLNSVIALLDADFHCLPTIQQRQQHCPNLDR 750
Query: 568 IKCRLLVGSNAQEVRATVLDMQARVLLDMLGKGIESALINPSALLPDPWQASEEILSNFD 627
IKCRLL G+NAQ+VRA VLDMQAR+LLDMLGKGIESALINPS LLP+PWQAS ++LS+
Sbjct: 751 IKCRLLEGTNAQDVRALVLDMQARLLLDMLGKGIESALINPSTLLPEPWQASSDMLSSIG 810
Query: 628 MEEMAVEPELTPCIQAYVDSVIDLASHLITRLRHYAKIFRTLANQAVTVASG-------- 679
++M V+P L IQ YVD+V+DLASH ITRLR YA RTLA+ AV +SG
Sbjct: 811 PDKMTVDPALLLSIQGYVDAVLDLASHFITRLRRYASFCRTLASHAVGASSGSGNSRNMV 870
Query: 680 ---------------------STSG--------------------GVPNLTLNPSISGPS 698
ST+G G N NP ISG S
Sbjct: 871 TSPTNSSPSPSTNQGNQGGVASTTGSSQMQEWVQGAIAKISNNTDGAANAAPNP-ISGRS 929
Query: 699 SLMLISINTGTFPGTPAVRLIGDCHFLHRLCQLLFFCFFFKRSQLARYKSGLRRTAETSL 758
S M ISINTGTFPGTPAVRLIGDCHFLHRLCQLL FC F+R Q R + ++++++S+
Sbjct: 930 SFMPISINTGTFPGTPAVRLIGDCHFLHRLCQLLLFCLLFRRRQSPRIPANAQKSSDSSM 989
Query: 759 ----------------VRS-------NDGQTGRDRQIVPGSKGGEEPSPG--PVRLGNGN 793
VRS DG T R + + G+KG EE G R+G+GN
Sbjct: 990 QKQHLMNSKTEDNTLAVRSGLGAAKLEDGTTSRGQMV--GAKGAEENPVGNKSARIGSGN 1047
Query: 794 AGQGYSVEEVKVIFQVLLDLCRRTSGLQHPLPVSQVGSSNIQVQLHYIEGSYTVLPEVVE 853
AGQGY+ +EVKV+F +L+DLC+RT+ LQHPLP SQVGSSNI ++LHYI+G+YTVLPEVVE
Sbjct: 1048 AGQGYTSDEVKVLFLILVDLCKRTATLQHPLPSSQVGSSNIIIRLHYIDGNYTVLPEVVE 1107
Query: 854 ASLGPYMQNMPRLGDADDTGLLLRELQLHPPAEEWHQLNMFVRPCTDSN-----DTPKPF 908
ASLGP+MQNMPR AD GLLLREL+L PPAEEWH+ NMF P ++ + D +
Sbjct: 1108 ASLGPHMQNMPRPRGADAAGLLLRELELQPPAEEWHRRNMFGGPWSEPDDLGPLDNTRQL 1167
Query: 909 RSNPLDSRSL----ESNDIDYGTNGLRPKKRRMIERDAAFGLNTSLGLGAYLGIMGSRRD 964
+ N +R L E D +G L P+KRR+ ERDAAFGL TS+GLG++LG+MGSRRD
Sbjct: 1168 KINGSTNRHLSDMEEDGDSSFGIQNLWPRKRRLSERDAAFGLKTSVGLGSFLGVMGSRRD 1227
Query: 965 VITTSWKTGLEGVWYK 980
VIT WKTGLEG WYK
Sbjct: 1228 VITAVWKTGLEGEWYK 1243
Score = 210 bits (534), Expect = 2e-52
Identities = 108/181 (59%), Positives = 129/181 (70%), Gaps = 5/181 (2%)
Query: 25 EEVEAGTVFDISLEQSPSTSLHKITVHESMRSNFSAVAWCAKLNVIACATETCVNGIPRS 84
++ TVF I L+Q PS+ HK+ V E R NFSAVAWC KLN IACA+ETC IP S
Sbjct: 109 QQASPATVFRIRLKQPPSSLRHKMRVPELCR-NFSAVAWCGKLNAIACASETCAR-IPSS 166
Query: 85 SVNPWFWIPIHIVIPERPTEIAAFNVVADSSLDSVQSIQWSPICCPRALLIANFQGRVTI 144
+ +P FWIPIHI+ PERPTE + FNV ADS D VQ I+WSP CPRALL+ANF GR+TI
Sbjct: 167 NSSPPFWIPIHILNPERPTECSVFNVKADSPRDFVQFIEWSPRSCPRALLVANFHGRITI 226
Query: 145 WTQPSRGPPNLVIDTNCWQREHEWRQETAVVTKWLSGASP---LDGPSFSVSVLYHNQER 201
WTQP++GP NLV D + WQ EHEWRQ+ +VVTKWLSG SP L S + S L +E+
Sbjct: 227 WTQPTKGPTNLVRDASSWQCEHEWRQDLSVVTKWLSGISPYRWLPANSSTSSNLKTFEEK 286
Query: 202 F 202
F
Sbjct: 287 F 287
>emb|CAB81034.1| hypothetical protein [Arabidopsis thaliana] gi|25372623|pir||H85061
hypothetical protein AT4g04920 [imported] - Arabidopsis
thaliana
Length = 1196
Score = 654 bits (1686), Expect = 0.0
Identities = 384/741 (51%), Positives = 456/741 (60%), Gaps = 120/741 (16%)
Query: 357 LRPVVLNPIFGS----MDGNPPMRTVWESKVDISIPAT---------------------- 390
++PVVL+ IFG+ G P +TVW S+VD+SIP T
Sbjct: 439 VQPVVLHQIFGNPTSNFGGQVPTQTVWVSRVDMSIPPTKDFKNHQVAAAGPSVDAPKEPD 498
Query: 391 --DDKSKGIHFNPFDLPKYLGTLARVVFSAQGGEIAVAFFQGRVRIFSGSNFEPVTNYEI 448
D+K+ + F+PFDLP + TLAR+V+SA GGEIA+AF +G V IFSG F PV NY+I
Sbjct: 499 SGDEKANKVVFDPFDLPSDIRTLARIVYSAHGGEIAIAFLRGGVHIFSGPTFSPVENYQI 558
Query: 449 NVGSSISVPAFSATSCCSASVWHDTSKGEAMLKIIRVLPPPFPIGQEKATSSTWEHAISE 508
NVGS+I+ PAFS TSCCSASVWHD +K AMLKIIRVLPP P Q K STWE AI+E
Sbjct: 559 NVGSAIAAPAFSPTSCCSASVWHDAAKDCAMLKIIRVLPPALPRNQSKVDQSTWERAIAE 618
Query: 509 RFWFSLLVGVDWWDVVGCTQRAAEEGIVSVNGVIAVLDADFHSLPTAQHRQQYCLGLDMI 568
RFW+SLLVGVDWWD VGCTQ AAE+GIVS+N VIAV+DADFHSLP+ QHRQQY LD I
Sbjct: 619 RFWWSLLVGVDWWDAVGCTQSAAEDGIVSLNSVIAVMDADFHSLPSTQHRQQYGPNLDRI 678
Query: 569 KCRLLVGSNAQEVRATVLDMQARVLLDMLGKGIESAL--------------------INP 608
KCRLL G+NAQEVRA VLDMQAR+LLDMLGKGIESAL INP
Sbjct: 679 KCRLLEGTNAQEVRAMVLDMQARLLLDMLGKGIESALVNPSALVFEPWRVDGETITGINP 738
Query: 609 SALLPDPWQASEEILSNFDMEEMAVEPELTPCIQAYVDSVIDLASHL-----------IT 657
A+ DP S + + + + ++ Y LASH +T
Sbjct: 739 EAMAVDPALVSS--IQAYVDAVLDLASHFITRLRRYASFCRTLASHAASAGTGSNRNNVT 796
Query: 658 RLRHYA------KIF-------------RTLANQAVTVASGST---------------SG 683
A ++F T A T +SGS+ S
Sbjct: 797 SPTQNASSPATPQVFPDKSLYLAVGQPTTTTTTTATTNSSGSSHVQAWMQGAIAKISSSN 856
Query: 684 GVPNLTLNPSISGPSSLMLISINTGTFPGTPAVRLIGDCHFLHRLCQLLFFCFFFKRSQL 743
N T +P ISG + M ISINTGTFPGTPAVRLIGDCHFLHRLCQLL FCF + S+
Sbjct: 857 DGSNSTASP-ISGSPTFMPISINTGTFPGTPAVRLIGDCHFLHRLCQLLLFCFLQRSSRF 915
Query: 744 ----ARYKSGLRRTAETS-------------LVRSNDGQTGRDRQIVPGSKGGEEPSPGP 786
A S +T TS L R D Q R Q+ G KG +E S
Sbjct: 916 PQRNADVSSQKLQTGATSKLEEVNSAKPTPALNRIEDAQGFRGAQLGTGVKGIDENSART 975
Query: 787 VRLGNGNAGQGYSVEEVKVIFQVLLDLCRRTSGLQHPLPVSQVGSSNIQVQLHYIEGSYT 846
++G+GNAGQGY+ EEV+V+F +L+DLC+RTSGL HPLP SQVGS NIQV+LHYI+G+YT
Sbjct: 976 TKMGSGNAGQGYTYEEVRVLFHILMDLCKRTSGLAHPLPGSQVGSGNIQVRLHYIDGNYT 1035
Query: 847 VLPEVVEASLGPYMQNMPRLGDADDTGLLLRELQLHPPAEEWHQLNMFVRPCTDSND--- 903
VLPEVVEA+LGP+MQNMPR AD GLLLREL+LHPP+EEWH+ N+F P ++ D
Sbjct: 1036 VLPEVVEAALGPHMQNMPRPRGADAAGLLLRELELHPPSEEWHRRNLFGGPGSEPEDMIL 1095
Query: 904 -TPKPFRSNPLDSRSLESNDIDYGTN---GLRPKKRRMIERDAAFGLNTSLGLGAYLGIM 959
SN LD + I G N L P+KRRM ERDAAFG NTS+GLGAYLGIM
Sbjct: 1096 TDDVSKLSNSLDLPDTNFSGICDGYNRVHSLWPRKRRMSERDAAFGSNTSVGLGAYLGIM 1155
Query: 960 GSRRDVITTSWKTGLEGVWYK 980
GSRRDV+T +WKTGLEGVWYK
Sbjct: 1156 GSRRDVVTATWKTGLEGVWYK 1176
Score = 219 bits (559), Expect = 3e-55
Identities = 116/207 (56%), Positives = 141/207 (68%), Gaps = 6/207 (2%)
Query: 6 ESEAEAEAAATVNDPMMEEEEVEAGTVFDISLEQSPSTSLHKITVHESMRSNFSAVAWCA 65
+ E E + +++ N ME + V TVF + L+Q S LHK++V E R NFSAVAWC
Sbjct: 64 DGEKEDDNSSSSN---MEIDPVSPATVFCVKLKQPNSNLLHKMSVPELCR-NFSAVAWCG 119
Query: 66 KLNVIACATETCVNGIPRSSVNPWFWIPIHIVIPERPTEIAAFNVVADSSLDSVQSIQWS 125
KLN IACA+ETC IP S N FWIPIHI+IPERPTE A FNVVADS DSVQ I+WS
Sbjct: 120 KLNAIACASETCAR-IPSSKANTPFWIPIHILIPERPTECAVFNVVADSPRDSVQFIEWS 178
Query: 126 PICCPRALLIANFQGRVTIWTQPSRGPPNLVIDTNCWQREHEWRQETAVVTKWLSGASPL 185
P CPRALLIANF GR+TIWTQP++G NLV D WQ EHEWRQ+ AVVTKWL+GASP
Sbjct: 179 PTSCPRALLIANFHGRITIWTQPTQGSANLVHDATSWQCEHEWRQDIAVVTKWLTGASPW 238
Query: 186 DGPSFSVSVLYHNQER-FNFIGPKGLL 211
S + + + ++ GP G++
Sbjct: 239 PSNQGSTAPKWFSTKKGLLGAGPSGIM 265
>ref|NP_192401.2| expressed protein [Arabidopsis thaliana]
Length = 1250
Score = 654 bits (1686), Expect = 0.0
Identities = 384/741 (51%), Positives = 456/741 (60%), Gaps = 120/741 (16%)
Query: 357 LRPVVLNPIFGS----MDGNPPMRTVWESKVDISIPAT---------------------- 390
++PVVL+ IFG+ G P +TVW S+VD+SIP T
Sbjct: 493 VQPVVLHQIFGNPTSNFGGQVPTQTVWVSRVDMSIPPTKDFKNHQVAAAGPSVDAPKEPD 552
Query: 391 --DDKSKGIHFNPFDLPKYLGTLARVVFSAQGGEIAVAFFQGRVRIFSGSNFEPVTNYEI 448
D+K+ + F+PFDLP + TLAR+V+SA GGEIA+AF +G V IFSG F PV NY+I
Sbjct: 553 SGDEKANKVVFDPFDLPSDIRTLARIVYSAHGGEIAIAFLRGGVHIFSGPTFSPVENYQI 612
Query: 449 NVGSSISVPAFSATSCCSASVWHDTSKGEAMLKIIRVLPPPFPIGQEKATSSTWEHAISE 508
NVGS+I+ PAFS TSCCSASVWHD +K AMLKIIRVLPP P Q K STWE AI+E
Sbjct: 613 NVGSAIAAPAFSPTSCCSASVWHDAAKDCAMLKIIRVLPPALPRNQSKVDQSTWERAIAE 672
Query: 509 RFWFSLLVGVDWWDVVGCTQRAAEEGIVSVNGVIAVLDADFHSLPTAQHRQQYCLGLDMI 568
RFW+SLLVGVDWWD VGCTQ AAE+GIVS+N VIAV+DADFHSLP+ QHRQQY LD I
Sbjct: 673 RFWWSLLVGVDWWDAVGCTQSAAEDGIVSLNSVIAVMDADFHSLPSTQHRQQYGPNLDRI 732
Query: 569 KCRLLVGSNAQEVRATVLDMQARVLLDMLGKGIESAL--------------------INP 608
KCRLL G+NAQEVRA VLDMQAR+LLDMLGKGIESAL INP
Sbjct: 733 KCRLLEGTNAQEVRAMVLDMQARLLLDMLGKGIESALVNPSALVFEPWRVDGETITGINP 792
Query: 609 SALLPDPWQASEEILSNFDMEEMAVEPELTPCIQAYVDSVIDLASHL-----------IT 657
A+ DP S + + + + ++ Y LASH +T
Sbjct: 793 EAMAVDPALVSS--IQAYVDAVLDLASHFITRLRRYASFCRTLASHAASAGTGSNRNNVT 850
Query: 658 RLRHYA------KIF-------------RTLANQAVTVASGST---------------SG 683
A ++F T A T +SGS+ S
Sbjct: 851 SPTQNASSPATPQVFPDKSLYLAVGQPTTTTTTTATTNSSGSSHVQAWMQGAIAKISSSN 910
Query: 684 GVPNLTLNPSISGPSSLMLISINTGTFPGTPAVRLIGDCHFLHRLCQLLFFCFFFKRSQL 743
N T +P ISG + M ISINTGTFPGTPAVRLIGDCHFLHRLCQLL FCF + S+
Sbjct: 911 DGSNSTASP-ISGSPTFMPISINTGTFPGTPAVRLIGDCHFLHRLCQLLLFCFLQRSSRF 969
Query: 744 ----ARYKSGLRRTAETS-------------LVRSNDGQTGRDRQIVPGSKGGEEPSPGP 786
A S +T TS L R D Q R Q+ G KG +E S
Sbjct: 970 PQRNADVSSQKLQTGATSKLEEVNSAKPTPALNRIEDAQGFRGAQLGTGVKGIDENSART 1029
Query: 787 VRLGNGNAGQGYSVEEVKVIFQVLLDLCRRTSGLQHPLPVSQVGSSNIQVQLHYIEGSYT 846
++G+GNAGQGY+ EEV+V+F +L+DLC+RTSGL HPLP SQVGS NIQV+LHYI+G+YT
Sbjct: 1030 TKMGSGNAGQGYTYEEVRVLFHILMDLCKRTSGLAHPLPGSQVGSGNIQVRLHYIDGNYT 1089
Query: 847 VLPEVVEASLGPYMQNMPRLGDADDTGLLLRELQLHPPAEEWHQLNMFVRPCTDSND--- 903
VLPEVVEA+LGP+MQNMPR AD GLLLREL+LHPP+EEWH+ N+F P ++ D
Sbjct: 1090 VLPEVVEAALGPHMQNMPRPRGADAAGLLLRELELHPPSEEWHRRNLFGGPGSEPEDMIL 1149
Query: 904 -TPKPFRSNPLDSRSLESNDIDYGTN---GLRPKKRRMIERDAAFGLNTSLGLGAYLGIM 959
SN LD + I G N L P+KRRM ERDAAFG NTS+GLGAYLGIM
Sbjct: 1150 TDDVSKLSNSLDLPDTNFSGICDGYNRVHSLWPRKRRMSERDAAFGSNTSVGLGAYLGIM 1209
Query: 960 GSRRDVITTSWKTGLEGVWYK 980
GSRRDV+T +WKTGLEGVWYK
Sbjct: 1210 GSRRDVVTATWKTGLEGVWYK 1230
Score = 219 bits (558), Expect = 4e-55
Identities = 112/179 (62%), Positives = 130/179 (72%), Gaps = 5/179 (2%)
Query: 6 ESEAEAEAAATVNDPMMEEEEVEAGTVFDISLEQSPSTSLHKITVHESMRSNFSAVAWCA 65
+ E E + +++ N ME + V TVF + L+Q S LHK++V E R NFSAVAWC
Sbjct: 64 DGEKEDDNSSSSN---MEIDPVSPATVFCVKLKQPNSNLLHKMSVPELCR-NFSAVAWCG 119
Query: 66 KLNVIACATETCVNGIPRSSVNPWFWIPIHIVIPERPTEIAAFNVVADSSLDSVQSIQWS 125
KLN IACA+ETC IP S N FWIPIHI+IPERPTE A FNVVADS DSVQ I+WS
Sbjct: 120 KLNAIACASETCAR-IPSSKANTPFWIPIHILIPERPTECAVFNVVADSPRDSVQFIEWS 178
Query: 126 PICCPRALLIANFQGRVTIWTQPSRGPPNLVIDTNCWQREHEWRQETAVVTKWLSGASP 184
P CPRALLIANF GR+TIWTQP++G NLV D WQ EHEWRQ+ AVVTKWL+GASP
Sbjct: 179 PTSCPRALLIANFHGRITIWTQPTQGSANLVHDATSWQCEHEWRQDIAVVTKWLTGASP 237
>gb|AAL58272.2| hypothetical protein, 3'-partial [Oryza sativa (japonica
cultivar-group)]
Length = 812
Score = 407 bits (1045), Expect = e-111
Identities = 211/371 (56%), Positives = 264/371 (70%), Gaps = 31/371 (8%)
Query: 357 LRPVVLNPIFGS---MDGNPPMRTVWESKVDISIPATDD--------------------- 392
++PVVL+PIFGS G PP +TVW ++V+ SIP ++D
Sbjct: 439 VQPVVLHPIFGSPANFGGQPPTQTVWSTRVNKSIPPSEDLKNPQSYVPMPTTSDERSSSE 498
Query: 393 ----KSKGIHFNPFDLPKYLGTLARVVFSAQGGEIAVAFFQGRVRIFSGSNFEPVTNYEI 448
++ + F+P+DLP + LA++V+SA GGE+AVAF +G V IFSG NFE V +Y +
Sbjct: 499 CSVDRANRLSFDPYDLPNDVRQLAQIVYSAHGGEVAVAFLRGGVHIFSGPNFEQVDSYHV 558
Query: 449 NVGSSISVPAFSATSCCSASVWHDTSKGEAMLKIIRVLPPPFPIGQEKATSSTWEHAISE 508
NVGS+I+ PAFS++ CC ASVWHDT K +LKIIRVLPP Q K +S+ WE AI++
Sbjct: 559 NVGSAIAPPAFSSSGCCLASVWHDTLKDRTILKIIRVLPPAILNAQTKVSSAVWERAIAD 618
Query: 509 -RFWFSLLVGVDWWDVVGCTQRAAEEGIVSVNGVIAVLDADFHSLPTAQHRQQYCLGLDM 567
RFW+SLL GVDWWD VGCTQ AAE+GIVS+N VIA+LDADFH LPT Q RQQ+C LD
Sbjct: 619 SRFWWSLLAGVDWWDAVGCTQSAAEDGIVSLNSVIALLDADFHCLPTIQQRQQHCPNLDR 678
Query: 568 IKCRLLVGSNAQEVRATVLDMQARVLLDMLGKGIESALINPSALLPDPWQASEEILSNFD 627
IKCRLL G+NAQ+VRA VLDMQAR+LLDMLGKGIESALINPS LLP+PWQAS ++LS+
Sbjct: 679 IKCRLLEGTNAQDVRALVLDMQARLLLDMLGKGIESALINPSTLLPEPWQASSDMLSSIG 738
Query: 628 MEEMAVEPELTPCIQAYVDSVIDLASHLITRLRHYAKIFRTLANQAVTVASGSTSGGVPN 687
++M V+P L IQ YVD+V+DLASH ITRLR YA RTLA+ AV +SG SG N
Sbjct: 739 PDKMTVDPALLLSIQGYVDAVLDLASHFITRLRRYASFCRTLASHAVGASSG--SGNSRN 796
Query: 688 LTLNPSISGPS 698
+ +P+ S PS
Sbjct: 797 MVTSPTNSSPS 807
Score = 210 bits (534), Expect = 2e-52
Identities = 108/181 (59%), Positives = 129/181 (70%), Gaps = 5/181 (2%)
Query: 25 EEVEAGTVFDISLEQSPSTSLHKITVHESMRSNFSAVAWCAKLNVIACATETCVNGIPRS 84
++ TVF I L+Q PS+ HK+ V E R NFSAVAWC KLN IACA+ETC IP S
Sbjct: 37 QQASPATVFRIRLKQPPSSLRHKMRVPELCR-NFSAVAWCGKLNAIACASETCAR-IPSS 94
Query: 85 SVNPWFWIPIHIVIPERPTEIAAFNVVADSSLDSVQSIQWSPICCPRALLIANFQGRVTI 144
+ +P FWIPIHI+ PERPTE + FNV ADS D VQ I+WSP CPRALL+ANF GR+TI
Sbjct: 95 NSSPPFWIPIHILNPERPTECSVFNVKADSPRDFVQFIEWSPRSCPRALLVANFHGRITI 154
Query: 145 WTQPSRGPPNLVIDTNCWQREHEWRQETAVVTKWLSGASP---LDGPSFSVSVLYHNQER 201
WTQP++GP NLV D + WQ EHEWRQ+ +VVTKWLSG SP L S + S L +E+
Sbjct: 155 WTQPTKGPTNLVRDASSWQCEHEWRQDLSVVTKWLSGISPYRWLPANSSTSSNLKTFEEK 214
Query: 202 F 202
F
Sbjct: 215 F 215
>ref|XP_315940.2| ENSANGP00000013473 [Anopheles gambiae str. PEST]
gi|55238722|gb|EAA11790.2| ENSANGP00000013473 [Anopheles
gambiae str. PEST]
Length = 357
Score = 37.7 bits (86), Expect = 1.9
Identities = 28/108 (25%), Positives = 48/108 (43%), Gaps = 4/108 (3%)
Query: 404 LPKYLGTLARVVFSAQGGEIAVAFFQGRV-RIFSGSNFEPVTNYEINVGSSISVPAFSAT 462
+P + LA + FS G EIA A +G V R+FS S+ + + V +S+ + + +
Sbjct: 178 IPAHDSPLAAIAFSQIGTEIATASEKGTVIRVFSVSDGSKLFEFRRGVKRCVSIASLAFS 237
Query: 463 SCCSASVWHDTSKGEAMLKIIRVLPPPFPIGQEKATSSTWEHAISERF 510
+C ++ + K+ RV P E+ W IS+ F
Sbjct: 238 TCSKYLCCSSNTETVHVFKLERVSP---ETNDEQGAGEAWYGKISKAF 282
>ref|XP_623610.1| PREDICTED: similar to Thyroid hormone receptor associated protein 5
[Apis mellifera]
Length = 924
Score = 37.4 bits (85), Expect = 2.5
Identities = 40/163 (24%), Positives = 71/163 (43%), Gaps = 28/163 (17%)
Query: 512 FSLLVGVDWWDVVGCTQRAAEEGIVSVNGVIAVLDADFHSLPTAQHRQQY-CLGLDMIKC 570
+ L+ G+DW DV+ C + + E + LD F+ Q QQY + IK
Sbjct: 444 YCLVTGLDWLDVLLCLRSSMIEPLCDR------LDVSFNR--QLQPIQQYHYIQFLCIKI 495
Query: 571 RLLVGSNAQEVRATVLDMQARVLLDMLGKGIESALINPSALLPDPWQASE---------- 620
L++ S ++ D+ + +++ + +S L+ PS L+ +E
Sbjct: 496 SLMLVSG----QSKAADLSSLLMIHSVATAFKS-LLRPSDLISHDKGPAESLATVINEAI 550
Query: 621 ----EILSNFDMEEMAVEPELTPCIQAYVDSVIDLASHLITRL 659
++L + D +E VEP +Q + V DLA +L+ RL
Sbjct: 551 TDVDKVLLHLDPKEFTVEPSTLQSLQQLIQWVADLALNLLARL 593
>ref|ZP_00202717.1| COG3181: Uncharacterized protein conserved in bacteria [Ralstonia
eutropha JMP134]
Length = 269
Score = 37.4 bits (85), Expect = 2.5
Identities = 33/123 (26%), Positives = 60/123 (47%), Gaps = 13/123 (10%)
Query: 536 VSVNGVIAVLDAD--FHS----LPTAQHRQQ------YCLGLDMIKCRLLVGSNAQEVRA 583
V+VN + V+ AD + S L A+H Q+ Y G ++ L + AQ V
Sbjct: 69 VAVNPAVFVVKADSPYRSIRDLLAAAKHEQRPISIGNYSTGYQLMATWLGTATGAQ-VNH 127
Query: 584 TVLDMQARVLLDMLGKGIESALINPSALLPDPWQASEEILSNFDMEEMAVEPELTPCIQA 643
A+++ D++G +E+A+ +PS++LP + I++ + M PE+ I++
Sbjct: 128 ISYKGGAQMVTDVIGGQLETAVTDPSSILPMHREGRVRIIATTGTKRMEALPEVPTLIES 187
Query: 644 YVD 646
VD
Sbjct: 188 GVD 190
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.327 0.141 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,695,746,373
Number of Sequences: 2540612
Number of extensions: 73534051
Number of successful extensions: 195368
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 195329
Number of HSP's gapped (non-prelim): 17
length of query: 980
length of database: 863,360,394
effective HSP length: 138
effective length of query: 842
effective length of database: 512,755,938
effective search space: 431740499796
effective search space used: 431740499796
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 80 (35.4 bits)
Medicago: description of AC129091.5


