

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC129091.14 + phase: 0
(439 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
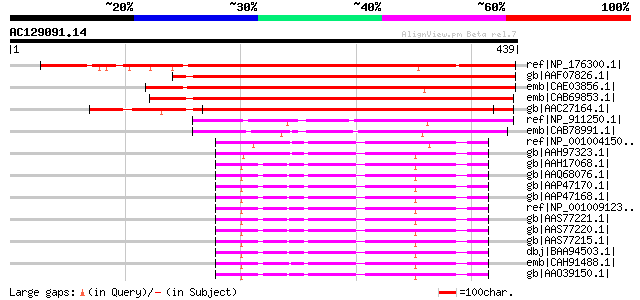
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_176300.1| alpha 1,4-glycosyltransferase family protein / ... 396 e-109
gb|AAF07826.1| unknown protein [Arabidopsis thaliana] gi|5923667... 339 9e-92
emb|CAE03856.1| OSJNBa0081C01.2 [Oryza sativa (japonica cultivar... 333 8e-90
emb|CAB69853.1| putative protein [Arabidopsis thaliana] gi|51315... 332 1e-89
gb|AAC27164.1| hypothetical protein [Arabidopsis thaliana] gi|15... 315 2e-84
ref|NP_911250.1| unknown protein [Oryza sativa (japonica cultiva... 133 1e-29
emb|CAB78991.1| putative protein [Arabidopsis thaliana] gi|57383... 131 4e-29
ref|NP_001004150.1| alpha 1,4-galactosyltransferase [Mus musculu... 121 4e-26
gb|AAH97323.1| Unknown (protein for MGC:114326) [Rattus norvegic... 118 3e-25
gb|AAH17068.1| Alpha 1,4-galactosyltransferase [Homo sapiens] gi... 112 2e-23
gb|AAQ68076.1| globotriaosylceramide/CD77 synthase [Homo sapiens... 112 2e-23
gb|AAP47170.1| alpha-1,4-galactosyltransferase [Homo sapiens] 112 2e-23
gb|AAP47168.1| alpha-1,4-galactosyltransferase [Homo sapiens] gi... 112 2e-23
ref|NP_001009123.1| alpha 1,4-galactosyltransferase [Pan troglod... 112 2e-23
gb|AAS77221.1| alpha-1,4-galactosyltransferase [Homo sapiens] 112 2e-23
gb|AAS77220.1| alpha-1,4-galactosyltransferase [Homo sapiens] 112 2e-23
gb|AAS77215.1| alpha-1,4-galactosyltransferase [Homo sapiens] 112 2e-23
dbj|BAA94503.1| alpha-1,4-N-acetylglucosaminyltransferase/alpha-... 112 2e-23
emb|CAH91488.1| hypothetical protein [Pongo pygmaeus] 112 2e-23
gb|AAO39150.1| alpha-1,4-galactosyltransferase [Homo sapiens] gi... 111 5e-23
>ref|NP_176300.1| alpha 1,4-glycosyltransferase family protein / glycosyltransferase
sugar-binding DXD motif-containing protein [Arabidopsis
thaliana] gi|12323337|gb|AAG51645.1| hypothetical
protein; 81821-83128 [Arabidopsis thaliana]
gi|25404304|pir||B96636 hypothetical protein T7P1.18
[imported] - Arabidopsis thaliana
Length = 435
Score = 396 bits (1018), Expect = e-109
Identities = 210/431 (48%), Positives = 283/431 (64%), Gaps = 33/431 (7%)
Query: 27 LHNFKKKILSILFKLPTSFFALLVLFLLAYNAFTVFCIHIPHHYSPHKPL--LLPPPS-- 82
+ K+ I+S +F LP S LL++ LL YN+F+VF +H+ P++P+ L P
Sbjct: 13 VQRLKRLIVSFVFCLPMSLLGLLLMLLLIYNSFSVFSLHLV----PNQPIQSTLSPTHLQ 68
Query: 83 --HTKTKLSSSVFFAHKEDNSP--IINKTHFPLLHKISIHKF---KRVKKHKRGLKTLHS 135
H +T SSSV D+S ++ +T + K ++ K+ ++ KR + +
Sbjct: 69 ILHHQTSTSSSV-----SDSSLLLVVKETSLGFIQKQNVSSTRIEKKTRRFKRSTELTPA 123
Query: 136 DTKF------PLFQKRLGAFFNGNSSSCSCKLRFFMTWISPLKAFGDRELLSVESLFKSH 189
T+ FQ R+ + S SC+ FFMTWIS +++FGDRE ++ESLFK H
Sbjct: 124 ITQRLQVKSRQRFQTRVKSLL----SKSSCESLFFMTWISSIESFGDRERFTIESLFKFH 179
Query: 190 PKACLVIVSKSMDSDKGTQILRPFVKNGFRVIAIEPDFNYIFKNTHAESWFNRLIQGNVN 249
P CL++VS S D D+GT IL+PF G +V+ I+PDF YIFK+T AE WF RL +G ++
Sbjct: 180 PNGCLILVSNSFDCDRGTLILKPFTDKGLKVLPIKPDFAYIFKDTSAEKWFERLKKGTLS 239
Query: 250 PGEISLGQNLSNLLRLSLLYKFGGIYIDADIIIMKSFSKFRNTIGAQNIDVKTKKWSRLN 309
PG I L QNLSNLLRL LLYK+GGIY+D D+II+KS S N IGAQ +D TKKWSRLN
Sbjct: 240 PGVIPLEQNLSNLLRLVLLYKYGGIYLDTDVIILKSLSNLHNVIGAQTVDPVTKKWSRLN 299
Query: 310 NAVLIFDKKHPLLLKFIEEFALTFDGNKWGHNGPYLISRVVSR--VSGREGYNFSVVPPS 367
NAVLIFDK HPLL +FI+EF+ TF+GNKWGHNGPYL+SRV++R +S FSV+PPS
Sbjct: 300 NAVLIFDKNHPLLKRFIDEFSRTFNGNKWGHNGPYLVSRVITRIKISSSSDLGFSVLPPS 359
Query: 368 AFYPVDWRGIKSLFRGPGDEIHSKWLVKKMVQIRKESYAVHLWNRQSGKLEVVKGSIIDS 427
AFYPVDW IK +R P +E WL K++ +RK ++AVHLWNR+S KL + +GSII
Sbjct: 360 AFYPVDWTRIKGFYRAPTNE-SDAWLRKRLTHLRKNTFAVHLWNRESKKLRIEEGSIIHQ 418
Query: 428 IISSCCIFCNT 438
++S CIFCN+
Sbjct: 419 LMSHSCIFCNS 429
>gb|AAF07826.1| unknown protein [Arabidopsis thaliana] gi|5923667|gb|AAD56318.1|
hypothetical protein [Arabidopsis thaliana]
gi|15232004|ref|NP_187514.1| alpha
1,4-glycosyltransferase family protein /
glycosyltransferase sugar-binding DXD motif-containing
protein [Arabidopsis thaliana]
Length = 411
Score = 339 bits (870), Expect = 9e-92
Identities = 156/297 (52%), Positives = 219/297 (73%), Gaps = 5/297 (1%)
Query: 142 FQKRLGAFFNGNSSSCSCKLRFFMTWISPLKAFGDRELLSVESLFKSHPKACLVIVSKSM 201
FQ+R F + C+++F MTWISP + FG RE+LSVES+FKSH + CL+I+S +M
Sbjct: 115 FQQRATEFLRDD-----CEVKFMMTWISPAELFGKREILSVESVFKSHARGCLMILSSTM 169
Query: 202 DSDKGTQILRPFVKNGFRVIAIEPDFNYIFKNTHAESWFNRLIQGNVNPGEISLGQNLSN 261
DS +G +IL+PF+ G+RV+A+ PD ++ K+T ESW + G +PG+ISL QNLSN
Sbjct: 170 DSLQGFRILKPFLDRGYRVMAVTPDLPFLLKDTAGESWLEEIQTGKRDPGKISLAQNLSN 229
Query: 262 LLRLSLLYKFGGIYIDADIIIMKSFSKFRNTIGAQNIDVKTKKWSRLNNAVLIFDKKHPL 321
L+RL+ L+KFGG+Y+D D+I++KSF RN IGAQ ++ ++ W+RLNNAVLIFDK HP
Sbjct: 230 LMRLAYLFKFGGVYLDTDMIVLKSFKTLRNVIGAQTLEPVSRNWTRLNNAVLIFDKNHPF 289
Query: 322 LLKFIEEFALTFDGNKWGHNGPYLISRVVSRVSGREGYNFSVVPPSAFYPVDWRGIKSLF 381
LLK IEEFALTF+GN WGHNGPYL+SRV V G +GYNF+++ P AFYPV+W I+ LF
Sbjct: 290 LLKSIEEFALTFNGNVWGHNGPYLVSRVARAVEGTDGYNFTILTPPAFYPVNWVEIEKLF 349
Query: 382 RGPGDEIHSKWLVKKMVQIRKESYAVHLWNRQSGKLEVVKGSIIDSIISSCCIFCNT 438
+ P E SK + K+++++K SY +HLWN+ S K E+ +GS +D ++S+ CI C++
Sbjct: 350 KVPRTEKDSKRVQVKVLEMQKRSYGLHLWNKFSRKFEIEQGSAMDKLVSNQCIICDS 406
>emb|CAE03856.1| OSJNBa0081C01.2 [Oryza sativa (japonica cultivar-group)]
gi|21742092|emb|CAD41203.1| OSJNBa0074L08.14 [Oryza
sativa (japonica cultivar-group)]
gi|50926638|ref|XP_473266.1| OSJNBa0074L08.14 [Oryza
sativa (japonica cultivar-group)]
Length = 445
Score = 333 bits (853), Expect = 8e-90
Identities = 166/326 (50%), Positives = 218/326 (65%), Gaps = 8/326 (2%)
Query: 118 HKFKRVKKHKRGLKTLHSDT-KFPLFQKRLGAFFNGNSSSCSCKLRFFMTWISPLKAFGD 176
+K K++K LK L +T + F R G F +S C RFFMTW+SPL FG
Sbjct: 116 NKKASAKRNKSLLKLLLRETPRTRRFAARAGELF---ASPRPCTRRFFMTWLSPLARFGR 172
Query: 177 RELLSVESLFKSHPKACLVIVSKSMDSDKGTQILRPFVKNGFRVIAIEPDFNYIFKNTHA 236
RELL VESLF+SH ACL+I S +MDSD G L PF+ G RV A PD Y+ T A
Sbjct: 173 RELLVVESLFRSHRDACLLIASDTMDSDGGGDRLGPFLDRGLRVAAASPDMAYLLNGTPA 232
Query: 237 ESWFNRLIQGNVNPGEISLGQNLSNLLRLSLLYKFGGIYIDADIIIMKSFSKFRNTIGAQ 296
E+W + +G+V+PG I LGQNLSNLLRL+LLYK+GG+Y+DAD+++++ FS RN IGAQ
Sbjct: 233 EAWLGAVQRGDVSPGSIPLGQNLSNLLRLALLYKYGGVYLDADVVVLRPFSDLRNAIGAQ 292
Query: 297 NIDVKTKKWSRLNNAVLIFDKKHPLLLKFIEEFALTFDGNKWGHNGPYLISRVVSRVSGR 356
+D T W RLNNAV++FD+ HPLL +FI EFA FDG+KWGHNGPYL+SRV +R R
Sbjct: 293 AVDASTGDWMRLNNAVMVFDRGHPLLREFIAEFAAKFDGSKWGHNGPYLVSRVAARWRRR 352
Query: 357 E----GYNFSVVPPSAFYPVDWRGIKSLFRGPGDEIHSKWLVKKMVQIRKESYAVHLWNR 412
+ +V+PP+AFYPVDW I LF P D +W+ K+ I+ ES+ +HLWNR
Sbjct: 353 RRPEAEADLTVLPPAAFYPVDWNKIGGLFVAPKDRKGERWVKAKVESIKGESFGIHLWNR 412
Query: 413 QSGKLEVVKGSIIDSIISSCCIFCNT 438
+S LE+ +GS+I ++S C+FCN+
Sbjct: 413 ESRSLEMEEGSVIGRLLSDSCLFCNS 438
>emb|CAB69853.1| putative protein [Arabidopsis thaliana] gi|51315396|gb|AAT99803.1|
At5g01250 [Arabidopsis thaliana]
gi|15240929|ref|NP_195745.1| alpha
1,4-glycosyltransferase family protein /
glycosyltransferase sugar-binding DXD motif-containing
protein [Arabidopsis thaliana] gi|11357836|pir||T45965
hypothetical protein F7J8.230 - Arabidopsis thaliana
Length = 407
Score = 332 bits (851), Expect = 1e-89
Identities = 154/315 (48%), Positives = 215/315 (67%), Gaps = 5/315 (1%)
Query: 122 RVKKHKRGLKTLHSDTKFPLFQKRLGAFFNGNSSSCSCKLRFFMTWISPLKAFGDRELLS 181
+V + + ++ D FQKR+ F C++ F MTWISP FG+RE+L+
Sbjct: 91 QVNEKLQVIEVFSGDNLSDKFQKRVNEFVGDG-----CEVNFVMTWISPADFFGNREVLA 145
Query: 182 VESLFKSHPKACLVIVSKSMDSDKGTQILRPFVKNGFRVIAIEPDFNYIFKNTHAESWFN 241
+ES+FKSHP CL+I+S +MDS +G L+PF+ G++V+A+ PD ++ K T E W +
Sbjct: 146 IESVFKSHPYGCLMILSATMDSPQGYATLKPFIDRGYKVLAVTPDLPFLLKGTAGELWLD 205
Query: 242 RLIQGNVNPGEISLGQNLSNLLRLSLLYKFGGIYIDADIIIMKSFSKFRNTIGAQNIDVK 301
+ G +PG+ISL QNLSNL+RL+ LYK+GG+Y+D D+I++KSF RN IGAQ +D
Sbjct: 206 EIKSGKRDPGKISLAQNLSNLMRLAYLYKYGGVYLDTDMIVLKSFKGLRNVIGAQTLDPS 265
Query: 302 TKKWSRLNNAVLIFDKKHPLLLKFIEEFALTFDGNKWGHNGPYLISRVVSRVSGREGYNF 361
+ W+RLNNAVLIFDK HPLLLKF+EEFA TF+GN WG+NGPYL+SRV V G GYNF
Sbjct: 266 STNWTRLNNAVLIFDKNHPLLLKFMEEFAKTFNGNIWGYNGPYLVSRVARAVEGSSGYNF 325
Query: 362 SVVPPSAFYPVDWRGIKSLFRGPGDEIHSKWLVKKMVQIRKESYAVHLWNRQSGKLEVVK 421
+V+ PS FY V+W IK LF+ P E SKW+ K++ +++ Y +HLWN+ S K E+ +
Sbjct: 326 TVMRPSVFYSVNWLEIKKLFKVPKTEKDSKWVKTKLLHMQRNGYGLHLWNKFSRKYEIEQ 385
Query: 422 GSIIDSIISSCCIFC 436
GS + ++S CI C
Sbjct: 386 GSAMWKLVSEHCIIC 400
>gb|AAC27164.1| hypothetical protein [Arabidopsis thaliana]
gi|15224453|ref|NP_181350.1| alpha
1,4-glycosyltransferase family protein /
glycosyltransferase sugar-binding DXD motif-containing
protein [Arabidopsis thaliana] gi|7452433|pir||T01247
hypothetical protein At2g38150 [imported] - Arabidopsis
thaliana
Length = 736
Score = 315 bits (806), Expect = 2e-84
Identities = 158/372 (42%), Positives = 234/372 (62%), Gaps = 17/372 (4%)
Query: 70 YSPHKPLLLP-PPSHTKTKLSSSVFFAHKEDNSPIINKTHFPLLHKISIHKFKRVKKHKR 128
+ P+ PL P P + ++S V HKE K PLL K +R+ +R
Sbjct: 369 FKPNVPLPRPYGPMSSSCNINSVVDSEHKE-------KELDPLLPPRKASKNQRIDWFRR 421
Query: 129 GL---KTLHSDTKFPLFQKRLGAFFNGNSSSCSCKLRFFMTWISPLKAFGDRELLSVESL 185
L + L S TK F R+ +N N C +FFM W+SP +FG RE+L++++L
Sbjct: 422 KLPELEILKSTTKSKSFHTRVLDLYNKN-----CSAQFFMIWLSPANSFGPREMLAIDTL 476
Query: 186 FKSHPKACLVIVSKSMDSDKGTQILRPFVKNGFRVIAIEPDFNYIFKNTHAESWFNRLIQ 245
F ++P ACL I+S S+DS G IL+P GF +IA+ D ++ KNT AE+W RL
Sbjct: 477 FTTNPGACLAILSNSLDSPNGYTILKPLFDQGFNLIAVTIDIPFLVKNTPAEAWLKRLKS 536
Query: 246 GNVNPGEISLGQNLSNLLRLSLLYKFGGIYIDADIIIMKSFSKFRNTIGAQNIDVKTKKW 305
GN++PG I L NLS+L RL++LYK+GG+Y+D DII + + RN IGAQ+ D TK+W
Sbjct: 537 GNMDPGSIPLFMNLSDLTRLAVLYKYGGVYLDTDIIFLNDMTGLRNAIGAQSSDPATKRW 596
Query: 306 SRLNNAVLIFDKKHPLLLKFIEEFALTFDGNKWGHNGPYLISRVVSRVSGREGY-NFSVV 364
+RLNNAV++FD HPL+ +F++E+A TFDGNKWG+N PYL+SRV+ R+ + GY N ++
Sbjct: 597 TRLNNAVMVFDIYHPLMREFLQEYATTFDGNKWGYNSPYLVSRVIKRLGNKPGYNNLTIF 656
Query: 365 PPSAFYPVDWRGIKSLFRGPGDEIHSKWLVKKMVQIRKESYAVHLWNRQSGKLEVVKGSI 424
P AFYPV+W I+ LF+ P +KW+ K + + K SY +HLWN+ + K+++ +GS+
Sbjct: 657 SPDAFYPVNWIKIQKLFKKPATTREAKWVEKTVQDMNKGSYMIHLWNKVTRKIKIEEGSV 716
Query: 425 IDSIISSCCIFC 436
+ +++S+ C C
Sbjct: 717 MHTLVSTHCTVC 728
Score = 270 bits (691), Expect = 5e-71
Identities = 129/253 (50%), Positives = 177/253 (68%), Gaps = 1/253 (0%)
Query: 168 ISPLKAFGDRELLSVESLFKSHPKACLVIVSKSMDSDKGTQILRPFVKNGFRVIAIEPDF 227
+S + FG RE+L+VES+FK+HP+ CL+IVS S+DS +G IL+P G++V A PD
Sbjct: 103 VSSDEYFGKREMLAVESVFKAHPQGCLMIVSGSLDSLQGDSILKPLNDRGYKVFAATPDM 162
Query: 228 NYIFKNTHAESWFNRLIQGNVNPGEISLGQNLSNLLRLSLLYKFGGIYIDADIIIMKSFS 287
+ + +NT A+SWF + +PG I L QNLSNL RL+ LYK+GG+Y+D D I+ +SF
Sbjct: 163 SLLLENTPAKSWFQEMKSCKRDPGRIPLHQNLSNLARLAFLYKYGGVYLDTDFIVTRSFK 222
Query: 288 KFRNTIGAQNI-DVKTKKWSRLNNAVLIFDKKHPLLLKFIEEFALTFDGNKWGHNGPYLI 346
+N+IGAQ + + +K W+RLNNAVLIF+K HPL+ FIEEFA TFDGNKWGHNGPYL+
Sbjct: 223 GLKNSIGAQTVVEGDSKNWTRLNNAVLIFEKDHPLVYSFIEEFASTFDGNKWGHNGPYLV 282
Query: 347 SRVVSRVSGREGYNFSVVPPSAFYPVDWRGIKSLFRGPGDEIHSKWLVKKMVQIRKESYA 406
+RV R G NF+V+PP AFYP +W I LF+ P S L +V++ +ESY
Sbjct: 283 TRVAQRARETIGDNFTVLPPVAFYPFNWLDIPRLFQTPRGSNDSTLLKTDLVKLNRESYG 342
Query: 407 VHLWNRQSGKLEV 419
+HLWN+ + KL++
Sbjct: 343 LHLWNKITRKLKI 355
>ref|NP_911250.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|27817893|dbj|BAC55659.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 615
Score = 133 bits (334), Expect = 1e-29
Identities = 82/286 (28%), Positives = 142/286 (48%), Gaps = 21/286 (7%)
Query: 159 CKLRFFMTWISPLKAFGDRELLSVESLFKSHPKACLVIVSKSMDSDKGTQILRPFVKNGF 218
C ++ FM W SP A+G R +ESL + HP+AC+V++S++++ + + FVK G+
Sbjct: 339 CSVKVFMVWNSPQWAYGVRHQRGLESLLRQHPEACVVMLSETLE----LEFFQEFVKEGY 394
Query: 219 RVIAIEPDFNYIFKNTHAESW---FNRLIQGNVNPGEISLGQNLSNLLRLSLLYKFGGIY 275
+V P+ + + + T + +N + P + S L+RL+ LYK+GGIY
Sbjct: 395 KVAVALPNLDELLEGTLTHDFVSVWNEWRKTKYYP------LHYSELVRLAALYKYGGIY 448
Query: 276 IDADIIIMKSFSKFRNTIGAQNIDVKTKKWSRLNNAVLIFDKKHPLLLKFIEEFALTFDG 335
+D+D++++K + RN+IG + + + S + AVL F+K P L + ++EF T+D
Sbjct: 449 LDSDVVVLKPLNALRNSIG---VVKQVSENSSFSGAVLAFEKNSPFLAECLKEFHSTYDD 505
Query: 336 NKWGHNGPYLISRVVSRVSGREGYN-----FSVVPPSAFYPVDWRGIKSLFRGPGDEIHS 390
NG L++RV+ +S + N P AFYP+ I F
Sbjct: 506 ELLQWNGAELMTRVIRNMSDKADDNSGHLDIKFEPSVAFYPISSTDITRYFSEADSTDER 565
Query: 391 KWLVKKMVQIRKESYAVHLWNRQSGKLEVVKGSIIDSIISSCCIFC 436
+I +S HLWN + L S+++ I++ C+ C
Sbjct: 566 AQHDALFSRIVNDSTTFHLWNSITSSLVPEPNSLVERILNRYCLHC 611
>emb|CAB78991.1| putative protein [Arabidopsis thaliana] gi|5738362|emb|CAB52870.1|
putative protein [Arabidopsis thaliana]
gi|25407572|pir||F85225 hypothetical protein AT4g19900
[imported] - Arabidopsis thaliana
gi|15235222|ref|NP_193724.1| glycosyl
transferase-related [Arabidopsis thaliana]
Length = 1302
Score = 131 bits (330), Expect = 4e-29
Identities = 88/282 (31%), Positives = 142/282 (50%), Gaps = 23/282 (8%)
Query: 159 CKLRFFMTWISPLKAFGDRELLSVESLFKSHPKACLVIVSKSMDSDKGTQILRPFVKNGF 218
C +R FM W SP F R +ESL H AC+V+ S++++ D FVK+ +
Sbjct: 368 CSMRVFMVWNSPGWMFSVRHQRGLESLLSQHRDACVVVFSETVELDF---FRNSFVKDSY 424
Query: 219 RVIAIEPDFNYIFKNT----HAESWFNRLIQGNVNPGEISLGQNLSNLLRLSLLYKFGGI 274
+V P+ + + ++T A WF+ + P + S L+RL+ LYK+GG+
Sbjct: 425 KVAVAMPNLDELLQDTPTHVFASVWFDWR-KTKFYP------THYSELVRLAALYKYGGV 477
Query: 275 YIDADIIIMKSFSKFRNTIGAQNIDVKTKKWSRLNNAVLIFDKKHPLLLKFIEEFALTFD 334
Y+D+D+I++ S S RNTIG ++ LN AV+ F+KK P LL+ + E+ LT+D
Sbjct: 478 YLDSDVIVLGSLSSLRNTIGMED----QVAGESLNGAVMSFEKKSPFLLECLNEYYLTYD 533
Query: 335 GNKWGHNGPYLISRVVSR-VSGR----EGYNFSVVPPSAFYPVDWRGIKSLFRGPGDEIH 389
NG L++RV R ++G+ ++ P S F+P++ + I + F P E
Sbjct: 534 DKCLRCNGADLLTRVAKRFLNGKNRRMNQQELNIRPSSVFFPINSQQITNYFAYPAIEDE 593
Query: 390 SKWLVKKMVQIRKESYAVHLWNRQSGKLEVVKGSIIDSIISS 431
+ +I ES H WN + L S++ +ISS
Sbjct: 594 RSQQDESFKKILNESLTFHFWNSVTSSLIPEPESLVAKLISS 635
>ref|NP_001004150.1| alpha 1,4-galactosyltransferase [Mus musculus]
gi|38350359|gb|AAR18365.1| Gb3 synthase [Mus musculus]
gi|59797925|sp|Q67BJ4|A4GAT_MOUSE Lactosylceramide
4-alpha-galactosyltransferase
(Alpha-1,4-galactosyltransferase)
(UDP-galactose:beta-D-galactosyl-beta1-R
4-alpha-D-galactosyltransferase)
(Alpha-1,4-N-acetylglucosaminyltransferase)
(Alpha4Gal-T1) (Globotriaosylceramide synthase) (Gb3
synthase)
Length = 359
Score = 121 bits (304), Expect = 4e-26
Identities = 76/246 (30%), Positives = 128/246 (51%), Gaps = 29/246 (11%)
Query: 179 LLSVESLFKSHPKACLVIVSKSMDSDKGTQI--LRPFVKNGFRVIAIEP-DFNYIFKNTH 235
+ SVES ++HP++ +V++ K + D Q L + + F + I P D +F++T
Sbjct: 101 MCSVESAARAHPESQVVVLMKGLPRDTTAQPRNLGISLLSCFPNVWIRPLDLQELFEDTP 160
Query: 236 AESWFNRLIQGNVNPGEISLGQNLSNLLRLSLLYKFGGIYIDADIIIMKSFSKFRNTIGA 295
+W++ + P ++ + LS+ R++LL+KFGGIY+D D I++K+ NT+G
Sbjct: 161 LAAWYSEA-RHRWEPYQLPV---LSDASRIALLWKFGGIYLDTDFIVLKNLLNLTNTLGI 216
Query: 296 QNIDVKTKKWSRLNNAVLIFDKKHPLLLKFIEEFALTFDGNKWGHNGPYLISRVVSRVSG 355
Q+ V LN A L F++KH L + +F ++G WGH GP L++RV +
Sbjct: 217 QSRYV-------LNGAFLAFERKHEFLALCLHDFVANYNGWIWGHQGPQLLTRVFKKWCS 269
Query: 356 REGYNFS-------VVPPSAFYPVDWRGIKSLFRGPGDEIHSKWLVKKMVQIRKESYAVH 408
+ S +PP AFYP+ W+ K F E ++ Q+ +YAVH
Sbjct: 270 IQSLEKSHACRGVTALPPEAFYPIPWQNWKKYFEDISPE--------ELTQLLNATYAVH 321
Query: 409 LWNRQS 414
+WN++S
Sbjct: 322 VWNKKS 327
>gb|AAH97323.1| Unknown (protein for MGC:114326) [Rattus norvegicus]
gi|11560038|ref|NP_071576.1| alpha
1,4-galactosyltransferase [Rattus norvegicus]
gi|59797638|sp|Q9JI93|A4GAT_RAT Lactosylceramide
4-alpha-galactosyltransferase
(Alpha-1,4-galactosyltransferase)
(UDP-galactose:beta-D-galactosyl-beta1-R
4-alpha-D-galactosyltransferase)
(Alpha-1,4-N-acetylglucosaminyltransferase)
(Alpha4Gal-T1) (Globotriaosylceramide synthase) (Gb3
synthase) gi|9082162|gb|AAF82758.1| Gb3 synthase [Rattus
norvegicus]
Length = 360
Score = 118 bits (296), Expect = 3e-25
Identities = 77/246 (31%), Positives = 126/246 (50%), Gaps = 29/246 (11%)
Query: 179 LLSVESLFKSHPKACLVIVSKSM--DSDKGTQILRPFVKNGFRVIAIEP-DFNYIFKNTH 235
+ SVES ++HP++ +V++ K + D+ + L + + F + I P D +F++T
Sbjct: 102 MCSVESAARAHPESQVVVLMKGLPRDTTAWPRNLGISLLSCFPNVQIRPLDLQELFEDTP 161
Query: 236 AESWFNRLIQGNVNPGEISLGQNLSNLLRLSLLYKFGGIYIDADIIIMKSFSKFRNTIGA 295
+W+ Q P + + LS+ R++LL+KFGGIY+D D I++K+ N +G
Sbjct: 162 LAAWYLEA-QHRWEPYLLPV---LSDASRIALLWKFGGIYLDTDFIVLKNLRNLTNMLGI 217
Query: 296 QNIDVKTKKWSRLNNAVLIFDKKHPLLLKFIEEFALTFDGNKWGHNGPYLISRVV----- 350
Q+ V LN A L F++KH L I +F ++G WGH GP L++RV
Sbjct: 218 QSRYV-------LNGAFLAFERKHEFLALCIRDFVAHYNGWIWGHQGPQLLTRVFKKWCS 270
Query: 351 --SRVSGREGYNFSVVPPSAFYPVDWRGIKSLFRGPGDEIHSKWLVKKMVQIRKESYAVH 408
S R + +PP AFYP+ W+ K F E ++ Q+ +YAVH
Sbjct: 271 IHSLKESRACRGVTALPPEAFYPIPWQNWKKYFEDVSPE--------ELAQLLNATYAVH 322
Query: 409 LWNRQS 414
+WN++S
Sbjct: 323 VWNKKS 328
>gb|AAH17068.1| Alpha 1,4-galactosyltransferase [Homo sapiens]
gi|56208108|emb|CAI18757.1| OTTHUMP00000028708 [Homo
sapiens] gi|8250233|emb|CAB93532.1|
alpha-4-galactosyltransferase [Homo sapiens]
gi|60459546|gb|AAX20109.1| alpha
1,4-galactosyltransferase [Homo sapiens]
gi|25452796|sp|Q9NPC4|A4GAT_HUMAN Lactosylceramide
4-alpha-galactosyltransferase
(Alpha-1,4-galactosyltransferase)
(UDP-galactose:beta-D-galactosyl-beta1-R
4-alpha-D-galactosyltransferase)
(Alpha-1,4-N-acetylglucosaminyltransferase)
(Alpha4Gal-T1) (Globotriaosylceramide synthase) (Gb3
synthase) (CD77 synthase) (P1/Pk synthase)
gi|8392830|ref|NP_059132.1| alpha
1,4-galactosyltransferase [Homo sapiens]
gi|7959011|dbj|BAA95915.1| Gb3/CD77 synthase [Homo
sapiens]
Length = 353
Score = 112 bits (281), Expect = 2e-23
Identities = 72/250 (28%), Positives = 124/250 (48%), Gaps = 37/250 (14%)
Query: 179 LLSVESLFKSHPKACLVIVSK-------SMDSDKGTQILRPFVKNGFRVIAIEPDFNYIF 231
+ SVES ++HP++ ++++ K S+ G +L F V + D +F
Sbjct: 95 MCSVESAARTHPESHVLVLMKGLPGGNASLPRHLGISLLSCFPN----VQMLPLDLRELF 150
Query: 232 KNTHAESWFNRLIQGNVNPGEISLGQNLSNLLRLSLLYKFGGIYIDADIIIMKSFSKFRN 291
++T W+ +QG P + + LS+ R++L++KFGGIY+D D I++K+ N
Sbjct: 151 RDTPLADWY-AAVQGRWEPYLLPV---LSDASRIALMWKFGGIYLDTDFIVLKNLRNLTN 206
Query: 292 TIGAQNIDVKTKKWSRLNNAVLIFDKKHPLLLKFIEEFALTFDGNKWGHNGPYLISRVV- 350
+G Q+ V LN A L F+++H + + +F ++G WGH GP L++RV
Sbjct: 207 VLGTQSRYV-------LNGAFLAFERRHEFMALCMRDFVDHYNGWIWGHQGPQLLTRVFK 259
Query: 351 ------SRVSGREGYNFSVVPPSAFYPVDWRGIKSLFRGPGDEIHSKWLVKKMVQIRKES 404
S R + +PP AFYP+ W+ K F E ++ ++ +
Sbjct: 260 KWCSIRSLAESRACRGVTTLPPEAFYPIPWQDWKKYFEDINPE--------ELPRLLSAT 311
Query: 405 YAVHLWNRQS 414
YAVH+WN++S
Sbjct: 312 YAVHVWNKKS 321
>gb|AAQ68076.1| globotriaosylceramide/CD77 synthase [Homo sapiens]
gi|33115183|gb|AAH55286.1| A4GALT protein [Homo sapiens]
Length = 353
Score = 112 bits (281), Expect = 2e-23
Identities = 72/250 (28%), Positives = 124/250 (48%), Gaps = 37/250 (14%)
Query: 179 LLSVESLFKSHPKACLVIVSK-------SMDSDKGTQILRPFVKNGFRVIAIEPDFNYIF 231
+ SVES ++HP++ ++++ K S+ G +L F V + D +F
Sbjct: 95 MCSVESAARTHPESHVLVLMKGLPGGNASLPRHLGISLLSCFPN----VQMLPLDLRELF 150
Query: 232 KNTHAESWFNRLIQGNVNPGEISLGQNLSNLLRLSLLYKFGGIYIDADIIIMKSFSKFRN 291
++T W+ +QG P + + LS+ R++L++KFGGIY+D D I++K+ N
Sbjct: 151 RDTPLADWY-AAVQGRWEPYLLPV---LSDASRIALMWKFGGIYLDTDFIVLKNLRNLTN 206
Query: 292 TIGAQNIDVKTKKWSRLNNAVLIFDKKHPLLLKFIEEFALTFDGNKWGHNGPYLISRVV- 350
+G Q+ V LN A L F+++H + + +F ++G WGH GP L++RV
Sbjct: 207 VLGTQSRYV-------LNGAFLAFERRHEFMALCMRDFVDHYNGWIWGHQGPQLLTRVFK 259
Query: 351 ------SRVSGREGYNFSVVPPSAFYPVDWRGIKSLFRGPGDEIHSKWLVKKMVQIRKES 404
S R + +PP AFYP+ W+ K F E ++ ++ +
Sbjct: 260 KWCSIRSLAESRACRGVTTLPPEAFYPIPWQDWKKYFEDINPE--------ELPRLLSAT 311
Query: 405 YAVHLWNRQS 414
YAVH+WN++S
Sbjct: 312 YAVHVWNKKS 321
>gb|AAP47170.1| alpha-1,4-galactosyltransferase [Homo sapiens]
Length = 354
Score = 112 bits (281), Expect = 2e-23
Identities = 72/250 (28%), Positives = 124/250 (48%), Gaps = 37/250 (14%)
Query: 179 LLSVESLFKSHPKACLVIVSK-------SMDSDKGTQILRPFVKNGFRVIAIEPDFNYIF 231
+ SVES ++HP++ ++++ K S+ G +L F V + D +F
Sbjct: 95 MCSVESAARTHPESHVLVLMKGLPGGNASLPRHLGISLLSCFPN----VQMLPLDLRELF 150
Query: 232 KNTHAESWFNRLIQGNVNPGEISLGQNLSNLLRLSLLYKFGGIYIDADIIIMKSFSKFRN 291
++T W+ +QG P + + LS+ R++L++KFGGIY+D D I++K+ N
Sbjct: 151 RDTPLADWY-AAVQGRWEPYLLPV---LSDASRIALMWKFGGIYLDTDFIVLKNLRNLTN 206
Query: 292 TIGAQNIDVKTKKWSRLNNAVLIFDKKHPLLLKFIEEFALTFDGNKWGHNGPYLISRVV- 350
+G Q+ V LN A L F+++H + + +F ++G WGH GP L++RV
Sbjct: 207 VLGTQSRYV-------LNGAFLAFERRHEFMALCMRDFVDHYNGWIWGHQGPQLLTRVFK 259
Query: 351 ------SRVSGREGYNFSVVPPSAFYPVDWRGIKSLFRGPGDEIHSKWLVKKMVQIRKES 404
S R + +PP AFYP+ W+ K F E ++ ++ +
Sbjct: 260 KWCSIRSLAESRACRGVTTLPPEAFYPIPWQDWKKYFEDINPE--------ELPRLLSAT 311
Query: 405 YAVHLWNRQS 414
YAVH+WN++S
Sbjct: 312 YAVHVWNKKS 321
>gb|AAP47168.1| alpha-1,4-galactosyltransferase [Homo sapiens]
gi|31324070|gb|AAP47167.1|
alpha-1,4-galactosyltransferase [Homo sapiens]
Length = 352
Score = 112 bits (281), Expect = 2e-23
Identities = 72/250 (28%), Positives = 124/250 (48%), Gaps = 37/250 (14%)
Query: 179 LLSVESLFKSHPKACLVIVSK-------SMDSDKGTQILRPFVKNGFRVIAIEPDFNYIF 231
+ SVES ++HP++ ++++ K S+ G +L F V + D +F
Sbjct: 94 MCSVESAARTHPESHVLVLMKGLPGGNASLPRHLGISLLSCFPN----VQMLPLDLRELF 149
Query: 232 KNTHAESWFNRLIQGNVNPGEISLGQNLSNLLRLSLLYKFGGIYIDADIIIMKSFSKFRN 291
++T W+ +QG P + + LS+ R++L++KFGGIY+D D I++K+ N
Sbjct: 150 RDTPLADWY-AAVQGRWEPYLLPV---LSDASRIALMWKFGGIYLDTDFIVLKNLRNLTN 205
Query: 292 TIGAQNIDVKTKKWSRLNNAVLIFDKKHPLLLKFIEEFALTFDGNKWGHNGPYLISRVV- 350
+G Q+ V LN A L F+++H + + +F ++G WGH GP L++RV
Sbjct: 206 VLGTQSRYV-------LNGAFLAFERRHEFMALCMRDFVDHYNGWIWGHQGPQLLTRVFK 258
Query: 351 ------SRVSGREGYNFSVVPPSAFYPVDWRGIKSLFRGPGDEIHSKWLVKKMVQIRKES 404
S R + +PP AFYP+ W+ K F E ++ ++ +
Sbjct: 259 KWCSIRSLAESRACRGVTTLPPEAFYPIPWQDWKKYFEDINPE--------ELPRLLSAT 310
Query: 405 YAVHLWNRQS 414
YAVH+WN++S
Sbjct: 311 YAVHVWNKKS 320
>ref|NP_001009123.1| alpha 1,4-galactosyltransferase [Pan troglodytes]
gi|25452795|sp|Q9N291|A4GAT_PANTR Lactosylceramide
4-alpha-galactosyltransferase
(Alpha-1,4-galactosyltransferase)
(UDP-galactose:beta-D-galactosyl-beta1-R
4-alpha-D-galactosyltransferase)
(Alpha-1,4-N-acetylglucosaminyltransferase)
(Globotriaosylceramide synthase) (Gb3 synthase)
gi|7593034|dbj|BAA94504.1|
alpha-1,4-N-acetylglucosaminyltransferase/alpha-
1,4-galactosyltransferase [Pan troglodytes]
Length = 353
Score = 112 bits (281), Expect = 2e-23
Identities = 72/250 (28%), Positives = 124/250 (48%), Gaps = 37/250 (14%)
Query: 179 LLSVESLFKSHPKACLVIVSK-------SMDSDKGTQILRPFVKNGFRVIAIEPDFNYIF 231
+ SVES ++HP++ ++++ K S+ G +L F V + D +F
Sbjct: 95 MCSVESAARTHPESHVLVLMKGLPGGNASLPRHLGISLLSCFPN----VQMLPLDLRELF 150
Query: 232 KNTHAESWFNRLIQGNVNPGEISLGQNLSNLLRLSLLYKFGGIYIDADIIIMKSFSKFRN 291
++T W+ +QG P + + LS+ R++L++KFGGIY+D D I++K+ N
Sbjct: 151 QDTPLADWY-AAVQGRWEPYLLPV---LSDASRIALMWKFGGIYLDTDFIVLKNLRNLTN 206
Query: 292 TIGAQNIDVKTKKWSRLNNAVLIFDKKHPLLLKFIEEFALTFDGNKWGHNGPYLISRVV- 350
+G Q+ V LN A L F+++H + + +F ++G WGH GP L++RV
Sbjct: 207 VLGTQSRYV-------LNGAFLAFERRHEFMALCMRDFVDHYNGWIWGHQGPQLLTRVFK 259
Query: 351 ------SRVSGREGYNFSVVPPSAFYPVDWRGIKSLFRGPGDEIHSKWLVKKMVQIRKES 404
S R + +PP AFYP+ W+ K F E ++ ++ +
Sbjct: 260 KWCSIRSLAESRSCRGVTTLPPEAFYPIPWQDWKKYFEDINPE--------ELPRLLSAT 311
Query: 405 YAVHLWNRQS 414
YAVH+WN++S
Sbjct: 312 YAVHVWNKKS 321
>gb|AAS77221.1| alpha-1,4-galactosyltransferase [Homo sapiens]
Length = 436
Score = 112 bits (281), Expect = 2e-23
Identities = 72/250 (28%), Positives = 124/250 (48%), Gaps = 37/250 (14%)
Query: 179 LLSVESLFKSHPKACLVIVSK-------SMDSDKGTQILRPFVKNGFRVIAIEPDFNYIF 231
+ SVES ++HP++ ++++ K S+ G +L F V + D +F
Sbjct: 95 MCSVESAARTHPESHVLVLMKGLPGGNASLPRHLGISLLSCFPN----VQMLPLDLRELF 150
Query: 232 KNTHAESWFNRLIQGNVNPGEISLGQNLSNLLRLSLLYKFGGIYIDADIIIMKSFSKFRN 291
++T W+ +QG P + + LS+ R++L++KFGGIY+D D I++K+ N
Sbjct: 151 RDTPLADWY-AAVQGRWEPYLLPV---LSDASRIALMWKFGGIYLDTDFIVLKNLRNLTN 206
Query: 292 TIGAQNIDVKTKKWSRLNNAVLIFDKKHPLLLKFIEEFALTFDGNKWGHNGPYLISRVV- 350
+G Q+ V LN A L F+++H + + +F ++G WGH GP L++RV
Sbjct: 207 VLGTQSRYV-------LNGAFLAFERRHEFMALCMRDFVDHYNGWIWGHQGPQLLTRVFK 259
Query: 351 ------SRVSGREGYNFSVVPPSAFYPVDWRGIKSLFRGPGDEIHSKWLVKKMVQIRKES 404
S R + +PP AFYP+ W+ K F E ++ ++ +
Sbjct: 260 KWCSIRSLAESRACRGVTTLPPEAFYPIPWQDWKKYFEDINPE--------ELPRLLSAT 311
Query: 405 YAVHLWNRQS 414
YAVH+WN++S
Sbjct: 312 YAVHVWNKKS 321
>gb|AAS77220.1| alpha-1,4-galactosyltransferase [Homo sapiens]
Length = 436
Score = 112 bits (281), Expect = 2e-23
Identities = 72/250 (28%), Positives = 124/250 (48%), Gaps = 37/250 (14%)
Query: 179 LLSVESLFKSHPKACLVIVSK-------SMDSDKGTQILRPFVKNGFRVIAIEPDFNYIF 231
+ SVES ++HP++ ++++ K S+ G +L F V + D +F
Sbjct: 95 MCSVESAARTHPESHVLVLMKGLPGGNASLPRHLGISLLSCFPN----VQMLPLDLRELF 150
Query: 232 KNTHAESWFNRLIQGNVNPGEISLGQNLSNLLRLSLLYKFGGIYIDADIIIMKSFSKFRN 291
++T W+ +QG P + + LS+ R++L++KFGGIY+D D I++K+ N
Sbjct: 151 RDTPLADWY-AAVQGRWEPYLLPV---LSDASRIALMWKFGGIYLDTDFIVLKNLRNLTN 206
Query: 292 TIGAQNIDVKTKKWSRLNNAVLIFDKKHPLLLKFIEEFALTFDGNKWGHNGPYLISRVV- 350
+G Q+ V LN A L F+++H + + +F ++G WGH GP L++RV
Sbjct: 207 VLGTQSRYV-------LNGAFLAFERRHEFMALCMRDFVDHYNGWIWGHQGPQLLTRVFK 259
Query: 351 ------SRVSGREGYNFSVVPPSAFYPVDWRGIKSLFRGPGDEIHSKWLVKKMVQIRKES 404
S R + +PP AFYP+ W+ K F E ++ ++ +
Sbjct: 260 KWCSIRSLAESRACRGVTTLPPEAFYPIPWQDWKKYFEDINPE--------ELPRLLSAT 311
Query: 405 YAVHLWNRQS 414
YAVH+WN++S
Sbjct: 312 YAVHVWNKKS 321
>gb|AAS77215.1| alpha-1,4-galactosyltransferase [Homo sapiens]
Length = 353
Score = 112 bits (281), Expect = 2e-23
Identities = 72/250 (28%), Positives = 124/250 (48%), Gaps = 37/250 (14%)
Query: 179 LLSVESLFKSHPKACLVIVSK-------SMDSDKGTQILRPFVKNGFRVIAIEPDFNYIF 231
+ SVES ++HP++ ++++ K S+ G +L F V + D +F
Sbjct: 95 MYSVESAARTHPESHVLVLMKGLPGGNASLPRHLGISLLSCFPN----VQMLPLDLRELF 150
Query: 232 KNTHAESWFNRLIQGNVNPGEISLGQNLSNLLRLSLLYKFGGIYIDADIIIMKSFSKFRN 291
++T W+ +QG P + + LS+ R++L++KFGGIY+D D I++K+ N
Sbjct: 151 RDTPLADWY-AAVQGRWEPYLLPV---LSDASRIALMWKFGGIYLDTDFIVLKNLRNLTN 206
Query: 292 TIGAQNIDVKTKKWSRLNNAVLIFDKKHPLLLKFIEEFALTFDGNKWGHNGPYLISRVV- 350
+G Q+ V LN A L F+++H + + +F ++G WGH GP L++RV
Sbjct: 207 VLGTQSRYV-------LNGAFLAFERRHEFMALCMRDFVDHYNGWIWGHQGPQLLTRVFK 259
Query: 351 ------SRVSGREGYNFSVVPPSAFYPVDWRGIKSLFRGPGDEIHSKWLVKKMVQIRKES 404
S R + +PP AFYP+ W+ K F E ++ ++ +
Sbjct: 260 KWCSIRSLAESRACRGVTTLPPEAFYPIPWQDWKKYFEDINPE--------ELPRLLSAT 311
Query: 405 YAVHLWNRQS 414
YAVH+WN++S
Sbjct: 312 YAVHVWNKKS 321
>dbj|BAA94503.1| alpha-1,4-N-acetylglucosaminyltransferase/alpha-
1,4-galactosyltransferase [Homo sapiens]
Length = 353
Score = 112 bits (281), Expect = 2e-23
Identities = 72/250 (28%), Positives = 124/250 (48%), Gaps = 37/250 (14%)
Query: 179 LLSVESLFKSHPKACLVIVSK-------SMDSDKGTQILRPFVKNGFRVIAIEPDFNYIF 231
+ SVES ++HP++ ++++ K S+ G +L F V + D +F
Sbjct: 95 MCSVESAARTHPESHVLVLMKGLPGGNASLPRHLGISLLSCFPN----VQMLPLDLRELF 150
Query: 232 KNTHAESWFNRLIQGNVNPGEISLGQNLSNLLRLSLLYKFGGIYIDADIIIMKSFSKFRN 291
++T W+ +QG P + + LS+ R++L++KFGGIY+D D I++K+ N
Sbjct: 151 RDTPLADWY-AAVQGRWEPYLLPV---LSDASRIALMWKFGGIYLDTDFIVLKNLRNLTN 206
Query: 292 TIGAQNIDVKTKKWSRLNNAVLIFDKKHPLLLKFIEEFALTFDGNKWGHNGPYLISRVV- 350
+G Q+ V LN A L F+++H + + +F ++G WGH GP L++RV
Sbjct: 207 VLGTQSRYV-------LNGAFLAFERRHEFMALCMRDFVDHYNGWIWGHQGPQLLTRVFK 259
Query: 351 ------SRVSGREGYNFSVVPPSAFYPVDWRGIKSLFRGPGDEIHSKWLVKKMVQIRKES 404
S R + +PP AFYP+ W+ K F E ++ ++ +
Sbjct: 260 KWCSIRSLAESRACRGVTTLPPEAFYPIPWQDWKKYFEDINPE--------ELPRLLSAT 311
Query: 405 YAVHLWNRQS 414
YAVH+WN++S
Sbjct: 312 YAVHVWNKKS 321
>emb|CAH91488.1| hypothetical protein [Pongo pygmaeus]
Length = 353
Score = 112 bits (280), Expect = 2e-23
Identities = 72/250 (28%), Positives = 123/250 (48%), Gaps = 37/250 (14%)
Query: 179 LLSVESLFKSHPKACLVIVSK-------SMDSDKGTQILRPFVKNGFRVIAIEPDFNYIF 231
+ SVES ++HP++ ++++ K S+ G +L F V + D +F
Sbjct: 95 MCSVESAARTHPESHVLVLMKGLPGGNASLPRHLGISLLSCFPN----VQMLPLDLRELF 150
Query: 232 KNTHAESWFNRLIQGNVNPGEISLGQNLSNLLRLSLLYKFGGIYIDADIIIMKSFSKFRN 291
++T W+ +QG P + + LS+ R++L++KFGGIY+D D I++K+ N
Sbjct: 151 RDTPLADWYTA-VQGRWEPYLLPV---LSDASRIALMWKFGGIYLDTDFIVLKNLRNLTN 206
Query: 292 TIGAQNIDVKTKKWSRLNNAVLIFDKKHPLLLKFIEEFALTFDGNKWGHNGPYLISRVV- 350
+G Q+ V LN A L F ++H + + +F ++G WGH GP L++RV
Sbjct: 207 VLGTQSRYV-------LNGAFLAFQRRHEFMALCMRDFVDHYNGWIWGHQGPQLLTRVFK 259
Query: 351 ------SRVSGREGYNFSVVPPSAFYPVDWRGIKSLFRGPGDEIHSKWLVKKMVQIRKES 404
S R + +PP AFYP+ W+ K F E ++ ++ +
Sbjct: 260 KWCSIRSLAESRACRGVTTLPPEAFYPIPWQDWKKYFEDISPE--------ELPRLLNAT 311
Query: 405 YAVHLWNRQS 414
YAVH+WN++S
Sbjct: 312 YAVHVWNKKS 321
>gb|AAO39150.1| alpha-1,4-galactosyltransferase [Homo sapiens]
gi|37726539|gb|AAO39149.1|
alpha-1,4-galactosyltransferase [Homo sapiens]
Length = 353
Score = 111 bits (277), Expect = 5e-23
Identities = 72/250 (28%), Positives = 123/250 (48%), Gaps = 37/250 (14%)
Query: 179 LLSVESLFKSHPKACLVIVSK-------SMDSDKGTQILRPFVKNGFRVIAIEPDFNYIF 231
+ SVES ++HP++ ++++ K S+ G +L F V + D +F
Sbjct: 95 MCSVESAARTHPESHVLVLMKGLPGGNASLPRHLGISLLSCFPN----VQMLPLDLRELF 150
Query: 232 KNTHAESWFNRLIQGNVNPGEISLGQNLSNLLRLSLLYKFGGIYIDADIIIMKSFSKFRN 291
++T W+ +QG P + + LS+ R++L +KFGGIY+D D I++K+ N
Sbjct: 151 RDTPLADWY-AAVQGRWEPYLLPV---LSDASRIALKWKFGGIYLDTDFIVLKNLRNLTN 206
Query: 292 TIGAQNIDVKTKKWSRLNNAVLIFDKKHPLLLKFIEEFALTFDGNKWGHNGPYLISRVV- 350
+G Q+ V LN A L F+++H + + +F ++G WGH GP L++RV
Sbjct: 207 VLGTQSRYV-------LNGAFLAFERRHEFMALCMRDFVDHYNGWIWGHQGPQLLTRVFK 259
Query: 351 ------SRVSGREGYNFSVVPPSAFYPVDWRGIKSLFRGPGDEIHSKWLVKKMVQIRKES 404
S R + +PP AFYP+ W+ K F E ++ ++ +
Sbjct: 260 KWCSIRSLAESRACRGVTTLPPEAFYPIPWQDWKKYFEDINPE--------ELPRLLSAT 311
Query: 405 YAVHLWNRQS 414
YAVH+WN++S
Sbjct: 312 YAVHVWNKKS 321
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.326 0.141 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 783,148,506
Number of Sequences: 2540612
Number of extensions: 32992264
Number of successful extensions: 116445
Number of sequences better than 10.0: 98
Number of HSP's better than 10.0 without gapping: 51
Number of HSP's successfully gapped in prelim test: 48
Number of HSP's that attempted gapping in prelim test: 116221
Number of HSP's gapped (non-prelim): 123
length of query: 439
length of database: 863,360,394
effective HSP length: 131
effective length of query: 308
effective length of database: 530,540,222
effective search space: 163406388376
effective search space used: 163406388376
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 77 (34.3 bits)
Medicago: description of AC129091.14


