

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC127429.3 - phase: 0 /pseudo
(423 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
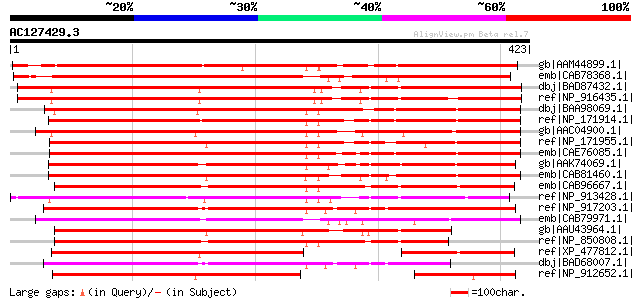
Score E
Sequences producing significant alignments: (bits) Value
gb|AAM44899.1| unknown protein [Arabidopsis thaliana] gi|1752899... 428 e-118
emb|CAB78368.1| putative protein [Arabidopsis thaliana] gi|47536... 425 e-117
dbj|BAD87432.1| putative flavin monoxygenase-like protein floozy... 424 e-117
ref|NP_916435.1| putative dimethylaniline monooxygenase [Oryza s... 409 e-113
dbj|BAA98069.1| dimethylaniline monooxygenase-like [Arabidopsis ... 376 e-103
ref|NP_171914.1| flavin-containing monooxygenase family protein ... 373 e-102
gb|AAC04900.1| putative flavin-containing monooxygenase [Arabido... 373 e-102
ref|NP_171955.1| flavin-containing monooxygenase / FMO (YUCCA3) ... 370 e-101
emb|CAE76085.1| B1340F09.23 [Oryza sativa (japonica cultivar-gro... 368 e-100
gb|AAK74069.1| flavin monoxygenase-like protein floozy [Petunia ... 365 2e-99
emb|CAB81460.1| putative protein [Arabidopsis thaliana] gi|42181... 360 6e-98
emb|CAB96667.1| putative protein [Arabidopsis thaliana] gi|15239... 348 2e-94
ref|NP_913428.1| dimethylaniline monooxygenase-like protein [Ory... 341 2e-92
ref|NP_917203.1| P0707D10.20 [Oryza sativa (japonica cultivar-gr... 339 8e-92
emb|CAB79971.1| dimethylaniline monooxygenase-like protein [Arab... 331 2e-89
gb|AAU43964.1| putative dimethylaniline monooxygenase [Oryza sat... 318 2e-85
ref|NP_850808.1| flavin-containing monooxygenase family protein ... 281 4e-74
ref|XP_477812.1| putative dimethylaniline monooxygenase [Oryza s... 279 1e-73
dbj|BAD68007.1| flavin-containing monooxygenase-like [Oryza sati... 275 1e-72
ref|NP_912652.1| Putative flavin-containing monooxygenase [Oryza... 237 4e-61
>gb|AAM44899.1| unknown protein [Arabidopsis thaliana] gi|17528990|gb|AAL38705.1|
unknown protein [Arabidopsis thaliana]
gi|22327064|ref|NP_197944.2| flavin-containing
monooxygenase, putative / FMO, putative [Arabidopsis
thaliana]
Length = 417
Score = 428 bits (1101), Expect = e-118
Identities = 234/429 (54%), Positives = 284/429 (65%), Gaps = 38/429 (8%)
Query: 3 CYLRELEGKQAHDPLFIEKMINNNKNSTSSSDRCLWIPGPLIVGAGPSGLAVAAYLKQKG 62
C+ RE+EGK AHD ++ TS C+ + GP+IVGAGPSGLA AA LK++G
Sbjct: 4 CWKREMEGKLAHD----------HRGMTSPRRICV-VTGPVIVGAGPSGLATAACLKERG 52
Query: 63 VPSLILERSNCIASLWKLKTYDRLRLHLPKQVCELPLMEFPSGFPTYPTKQQFIEYLESY 122
+ S++LERSNCIASLW+LKTYDRL LHLPKQ CELP++ FP FPTYPTKQQFIEYLE Y
Sbjct: 53 ITSVLLERSNCIASLWQLKTYDRLHLHLPKQFCELPIIPFPGDFPTYPTKQQFIEYLEDY 112
Query: 123 SKNFDIRPWFNETVMHAEFDATLGFWRVRSEGKAGMVTEFVCRWLIVATGENAEAVVPEI 182
++ FDI+P FN+TV A FD LG WRV S G+ G TE+VCRWL+ ATGENAE VVP
Sbjct: 113 ARRFDIKPEFNQTVESAAFDENLGMWRVTSVGEEG-TTEYVCRWLVAATGENAEPVVPRF 171
Query: 183 EGVDEF--VGSIRHTSLYKSGEEFRGKKVLVVGCGNSGMEVCLDLCNHDAAPSIVVRDSP 240
EG+D+F G ++HT YK+G +F GK+VLVVGCGNSGMEVCLDLCN A PS+VVRD+
Sbjct: 172 EGMDKFAAAGVVKHTCHYKTGGDFAGKRVLVVGCGNSGMEVCLDLCNFGAQPSLVVRDAV 231
Query: 241 ---TKRYAGKINF--------W----VVHVVAQVATRATGGSHPSHCVMVNARQHRTVRF 285
+ G F W +V V +R G ++ + R
Sbjct: 232 HVLPREMLGTSTFGLSMFLLKWLPIRLVDRFLLVVSRFILGD----TTLLGLNRPRLGPL 287
Query: 286 GSASVGSP*TQKTVWKDTGPRCGCPCQD*KRRHQGIKRLKRYTVEFADGSTENFDAIILA 345
++ KT D G D K GI+RLKR+ VEF +G TE FDAIILA
Sbjct: 288 ELKNISG----KTPVLDVGTLAKIKTGDIK-VCSGIRRLKRHEVEFDNGKTERFDAIILA 342
Query: 346 TGYKSNVPYWLKDKGMFSKEDGYPRKPFPNGWKGENGLYAVGFTKRGLLGASMDAKNIAE 405
TGYKSNVP WLK+ MFSK+DG+P + FP GW+GE GLYAVGFTKRG+ GASMDAK IAE
Sbjct: 343 TGYKSNVPSWLKENKMFSKKDGFPIQEFPEGWRGECGLYAVGFTKRGISGASMDAKRIAE 402
Query: 406 DIERCWKAE 414
DI +CWK +
Sbjct: 403 DIHKCWKQD 411
>emb|CAB78368.1| putative protein [Arabidopsis thaliana] gi|4753660|emb|CAB41936.1|
putative protein [Arabidopsis thaliana]
gi|22136612|gb|AAM91625.1| unknown protein [Arabidopsis
thaliana] gi|16555354|gb|AAL23751.1| flavin-containing
monooxygenase YUCCA2 [Arabidopsis thaliana]
gi|15235652|ref|NP_193062.1| flavin-containing
monooxygenase / FMO (YUCCA2) [Arabidopsis thaliana]
gi|7485670|pir||T07706 hypothetical protein F17N18.150 -
Arabidopsis thaliana
Length = 415
Score = 425 bits (1092), Expect = e-117
Identities = 234/430 (54%), Positives = 283/430 (65%), Gaps = 56/430 (13%)
Query: 4 YLRELEGKQAHDPLFIEKMINNNKNSTSSSDRCLWIPGPLIVGAGPSGLAVAAYLKQKGV 63
++ E GK+ HDP ++E+ RCL IPGP+IVG+GPSGLA AA LK + +
Sbjct: 3 FVTETLGKRIHDP-YVEET------------RCLMIPGPIIVGSGPSGLATAACLKSRDI 49
Query: 64 PSLILERSNCIASLWKLKTYDRLRLHLPKQVCELPLMEFPSGFPTYPTKQQFIEYLESYS 123
PSLILERS CIASLW+ KTYDRLRLHLPK CELPLM FPS +PTYPTKQQF++YLESY+
Sbjct: 50 PSLILERSTCIASLWQHKTYDRLRLHLPKDFCELPLMPFPSSYPTYPTKQQFVQYLESYA 109
Query: 124 KNFDIRPWFNETVMHAEFDATLGFWRVRSE-GKAGMVTEFVCRWLIVATGENAEAVVPEI 182
++FD++P FN+TV A+FD G WRVR+ GK E+V RWL+VATGENAE V+PEI
Sbjct: 110 EHFDLKPVFNQTVEEAKFDRRCGLWRVRTTGGKKDETMEYVSRWLVVATGENAEEVMPEI 169
Query: 183 EGVDEFVGSIRHTSLYKSGEEFRGKKVLVVGCGNSGMEVCLDLCNHDAAPSIVVRDSPTK 242
+G+ +F G I HTS YKSGE F KK+LVVGCGNSGMEVCLDLCN +A PS+VVRDS
Sbjct: 170 DGIPDFGGPILHTSSYKSGEIFSEKKILVVGCGNSGMEVCLDLCNFNALPSLVVRDS--- 226
Query: 243 RYAGKINFWVVHVVAQ----VATRATGGS----HPSHCVMVNARQHRTVRFGSASVGSP* 294
VHV+ Q ++T S P H V +R +G
Sbjct: 227 ----------VHVLPQEMLGISTFGISTSLLKWFPVHVV-----DRFLLRMSRLVLGDTD 271
Query: 295 TQKTVWKDTGP-----RCG-CPCQD*K----------RRHQGIKRLKRYTVEFADGSTEN 338
V GP +CG P D + + +KR+ Y+ EF DG +N
Sbjct: 272 RLGLVRPKLGPLERKIKCGKTPVLDVGTLAKIRSGHIKVYPELKRVMHYSAEFVDGRVDN 331
Query: 339 FDAIILATGYKSNVPYWLKDKGMFSKEDGYPRKPFPNGWKGENGLYAVGFTKRGLLGASM 398
FDAIILATGYKSNVP WLK MFS++DG+P KPFPNGWKGE+GLYAVGFTK GLLGA++
Sbjct: 332 FDAIILATGYKSNVPMWLKGVNMFSEKDGFPHKPFPNGWKGESGLYAVGFTKLGLLGAAI 391
Query: 399 DAKNIAEDIE 408
DAK IAEDIE
Sbjct: 392 DAKKIAEDIE 401
>dbj|BAD87432.1| putative flavin monoxygenase-like protein floozy [Oryza sativa
(japonica cultivar-group)]
Length = 438
Score = 424 bits (1091), Expect = e-117
Identities = 230/436 (52%), Positives = 288/436 (65%), Gaps = 33/436 (7%)
Query: 7 ELEGKQAHDPLFIEKMINNNKNSTSS-------------SDRCLWIPGPLIVGAGPSGLA 53
E+EGK+AHDP+F N + S ++RC+ + GP+IVGAGPSGLA
Sbjct: 6 EIEGKRAHDPIFQNYFSQNCRQSVDGFCKKRSADAAVARAERCIRVLGPIIVGAGPSGLA 65
Query: 54 VAAYLKQKGVPSLILERSNCIASLWKLKTYDRLRLHLPKQVCELPLMEFPSGFPTYPTKQ 113
VAA LK+KGV SL+LERSNCIASLW+LKTYDRL LHLP+Q CELPLM FP+ +P YP+KQ
Sbjct: 66 VAACLKEKGVDSLVLERSNCIASLWQLKTYDRLSLHLPRQFCELPLMPFPAYYPIYPSKQ 125
Query: 114 QFIEYLESYSKNFDIRPWFNETVMHAEFDATLGFWRVRSE--GKAGMVTEFVCRWLIVAT 171
QF+ YLESY+ F I P +N TV+ AE+D L WRVR+ G G E+V RWL+VAT
Sbjct: 126 QFVAYLESYAARFGICPTYNRTVVCAEYDEQLQLWRVRTRATGIMGEEVEYVSRWLVVAT 185
Query: 172 GENAEAVVPEIEGVDEFVGSIRHTSLYKSGEEFRGKKVLVVGCGNSGMEVCLDLCNHDAA 231
GENAE V+PEI+G+D+F G++ HTS YKSG F GK+VLVVG GNSGMEVCLDLCNH+A
Sbjct: 186 GENAEVVLPEIDGLDDFKGTVMHTSSYKSGGAFAGKRVLVVGSGNSGMEVCLDLCNHNAN 245
Query: 232 PSIVVRDSPTKRYAGK---INFWV-----VHVVAQVATRATGGSHPSHCVMVNARQHRTV 283
P IVV P + ++ W+ VHVV ++ + ++ + Q
Sbjct: 246 PHIVVHILPREMLGQSTFGLSMWLLKWLPVHVVDRILLLI------AQTMLGDTAQLGLK 299
Query: 284 R--FGSASVGSP*TQKTVWKDTGPRCGCPCQD*KRRHQGIKRLKRYTVEFADGSTENFDA 341
R G + S + KT D G D K R IK++ VEF D E FD
Sbjct: 300 RPTIGPLELKSL-SGKTPVLDVGTFAKIKSGDIKVR-PAIKQISGRQVEFMDTRLEEFDV 357
Query: 342 IILATGYKSNVPYWLKDKGMFSKEDGYPRKPFPNGWKGENGLYAVGFTKRGLLGASMDAK 401
I+LATGYKSNVP+WLKD+ +FS++DG PRK FPNGWKGENGLY+VGFT+RGL+G S+DA+
Sbjct: 358 IVLATGYKSNVPFWLKDRELFSEKDGLPRKAFPNGWKGENGLYSVGFTRRGLMGTSVDAR 417
Query: 402 NIAEDIERCWKAEAKH 417
IA DIE+ WKA KH
Sbjct: 418 RIAHDIEQQWKARGKH 433
>ref|NP_916435.1| putative dimethylaniline monooxygenase [Oryza sativa (japonica
cultivar-group)]
Length = 430
Score = 409 bits (1052), Expect = e-113
Identities = 227/436 (52%), Positives = 281/436 (64%), Gaps = 41/436 (9%)
Query: 7 ELEGKQAHDPLFIEKMINNNKNSTSS-------------SDRCLWIPGPLIVGAGPSGLA 53
E+EGK+AHDP+F N + S ++RC+ + GP+IVGAGPSGLA
Sbjct: 6 EIEGKRAHDPIFQNYFSQNCRQSVDGFCKKRSADAAVARAERCIRVLGPIIVGAGPSGLA 65
Query: 54 VAAYLKQKGVPSLILERSNCIASLWKLKTYDRLRLHLPKQVCELPLMEFPSGFPTYPTKQ 113
VAA LK+KGV SL+LERSNCIASLW+LKTYDRL LHLP+Q CELPLM FP+ +P YP+KQ
Sbjct: 66 VAACLKEKGVDSLVLERSNCIASLWQLKTYDRLSLHLPRQFCELPLMPFPAYYPIYPSKQ 125
Query: 114 QFIEYLESYSKNFDIRPWFNETVMHAEFDATLGFWRVRSE--GKAGMVTEFVCRWLIVAT 171
QF+ YLESY+ F I P +N TV+ AE+D L WRVR+ G G E+V RWL+VAT
Sbjct: 126 QFVAYLESYAARFGICPTYNRTVVCAEYDEQLQLWRVRTRATGIMGEEVEYVSRWLVVAT 185
Query: 172 GENAEAVVPEIEGVDEFVGSIRHTSLYKSGEEFRGKKVLVVGCGNSGMEVCLDLCNHDAA 231
GENAE V+PEI+G+D+F G++ HTS YKSG F GK+VLVVG GNSGMEVCLDLCNH+A
Sbjct: 186 GENAEVVLPEIDGLDDFKGTVMHTSSYKSGGAFAGKRVLVVGSGNSGMEVCLDLCNHNAN 245
Query: 232 PSIVVRDSPTKRYAGK---INFWV-----VHVVAQVATRATGGSHPSHCVMVNARQHRTV 283
P IVV P + ++ W+ VHVV ++ + ++ + Q
Sbjct: 246 PHIVVHILPREMLGQSTFGLSMWLLKWLPVHVVDRILLLI------AQTMLGDTAQLGLK 299
Query: 284 R--FGSASVGSP*TQKTVWKDTGPRCGCPCQD*KRRHQGIKRLKRYTVEFADGSTENFDA 341
R G + S + KT D G D K R IK++ VEF D E FD
Sbjct: 300 RPTIGPLELKSL-SGKTPVLDVGTFAKIKSGDIKVR-PAIKQISGRQVEFMDTRLEEFDV 357
Query: 342 IILATGYKSNVPYWLKDKGMFSKEDGYPRKPFPNGWKGENGLYAVGFTKRGLLGASMDAK 401
I+LATGYKSNVP+WLK DG PRK FPNGWKGENGLY+VGFT+RGL+G S+DA+
Sbjct: 358 IVLATGYKSNVPFWLK--------DGLPRKAFPNGWKGENGLYSVGFTRRGLMGTSVDAR 409
Query: 402 NIAEDIERCWKAEAKH 417
IA DIE+ WKA KH
Sbjct: 410 RIAHDIEQQWKARGKH 425
>dbj|BAA98069.1| dimethylaniline monooxygenase-like [Arabidopsis thaliana]
gi|15240026|ref|NP_199202.1| flavin-containing
monooxygenase family protein / FMO family protein
[Arabidopsis thaliana]
Length = 424
Score = 376 bits (966), Expect = e-103
Identities = 201/408 (49%), Positives = 263/408 (64%), Gaps = 30/408 (7%)
Query: 29 STSSSDR--CLWIPGPLIVGAGPSGLAVAAYLKQKGVPSLILERSNCIASLWKLKTYDRL 86
S SSDR C+W+ GP+IVGAGPSGLA AA L+++GVP ++LER++CIASLW+ +TYDR+
Sbjct: 10 SEDSSDRRRCIWVNGPVIVGAGPSGLATAACLREEGVPFVVLERADCIASLWQKRTYDRI 69
Query: 87 RLHLPKQVCELPLMEFPSGFPTYPTKQQFIEYLESYSKNFDIRPWFNETVMHAEFDATLG 146
+LHLPK+VC+LP M FP +P YPTK+QFIEYLESY+ F+I P FNE V A +D T G
Sbjct: 70 KLHLPKKVCQLPKMPFPEDYPEYPTKRQFIEYLESYANKFEITPQFNECVQSARYDETSG 129
Query: 147 FWRVR--SEGKAGMVTEFVCRWLIVATGENAEAVVPEIEGV-DEFVGSIRHTSLYKSGEE 203
WR++ S +G E++CRWL+VATGENAE VVPEI+G+ EF G + H+ YKSGE+
Sbjct: 130 LWRIKTTSSSSSGSEMEYICRWLVVATGENAEKVVPEIDGLTTEFEGEVIHSCEYKSGEK 189
Query: 204 FRGKKVLVVGCGNSGMEVCLDLCNHDAAPSIVVRDSP---TKRYAGKINF---------- 250
+RGK VLVVGCGNSGMEV LDL NH+A S+VVR S + GK +F
Sbjct: 190 YRGKSVLVVGCGNSGMEVSLDLANHNANASMVVRSSVHVLPREILGKSSFEISMMLMKWF 249
Query: 251 --WVVHVVAQVATRATGGSHPSHCVMVNARQHRTVRFGSASVGSP*TQKTVWKDTGPRCG 308
W+V + + G+ + + ++ S KT D G
Sbjct: 250 PLWLVDKILLILAWLILGNLTKYGLKRPTMGPMELKIVSG--------KTPVLDIGAMEK 301
Query: 309 CPCQD*KRRHQGIKRLKRYTVEFADGSTENFDAIILATGYKSNVPYWLKDKGMFSKEDGY 368
+ GIKR R VE DG + DA++LATGY+SNVP WL++ +FSK +G+
Sbjct: 302 IKSGE-VEIVPGIKRFSRSHVELVDGQRLDLDAVVLATGYRSNVPSWLQENDLFSK-NGF 359
Query: 369 PRKPFPNGWKGENGLYAVGFTKRGLLGASMDAKNIAEDIERCWKAEAK 416
P+ PFPN WKG++GLYA GFT++GL GAS DA NIA+DI W+ E K
Sbjct: 360 PKSPFPNAWKGKSGLYAAGFTRKGLAGASADAVNIAQDIGNVWREETK 407
>ref|NP_171914.1| flavin-containing monooxygenase family protein / FMO family protein
[Arabidopsis thaliana] gi|3142293|gb|AAC16744.1|
Contains similarity to myosin IB heavy chain gb|X70400
from Gallus gallus. [Arabidopsis thaliana]
gi|7485875|pir||T00955 hypothetical protein F20D22.5 -
Arabidopsis thaliana
Length = 421
Score = 373 bits (958), Expect = e-102
Identities = 202/401 (50%), Positives = 260/401 (64%), Gaps = 27/401 (6%)
Query: 32 SSDRCLWIPGPLIVGAGPSGLAVAAYLKQKGVPSLILERSNCIASLWKLKTYDRLRLHLP 91
S RC+W+ GP+IVGAGPSGLA AA L +GVP +++ERS+CIASLW+ +TYDRL+LHLP
Sbjct: 15 SERRCVWVNGPVIVGAGPSGLATAACLHDQGVPFVVVERSDCIASLWQKRTYDRLKLHLP 74
Query: 92 KQVCELPLMEFPSGFPTYPTKQQFIEYLESYSKNFDIRPWFNETVMHAEFDATLGFWRVR 151
K+ C+LP M FP +P YPTK+QFI+YLESY+ FDI+P FN++V A FD T G WRVR
Sbjct: 75 KKFCQLPKMPFPDHYPEYPTKRQFIDYLESYANRFDIKPEFNKSVESARFDETSGLWRVR 134
Query: 152 SEGKAGMVTEFVCRWLIVATGENAEAVVPEIEG-VDEFVGSIRHTSLYKSGEEFRGKKVL 210
+ G E++CRWL+VATGENAE VVPEI G + EF G + H YKSGE+FRGK+VL
Sbjct: 135 TTSD-GEEMEYICRWLVVATGENAERVVPEINGLMTEFDGEVIHACEYKSGEKFRGKRVL 193
Query: 211 VVGCGNSGMEVCLDLCNHDAAPSIVVRDSP---TKRYAGKINF------------WVVHV 255
VVGCGNSGMEV LDL NH+A S+VVR S + GK F W+V
Sbjct: 194 VVGCGNSGMEVSLDLANHNAITSMVVRSSVHVLPREIMGKSTFGISVMMMKWLPLWLVDK 253
Query: 256 VAQVATRATGGSHPSHCVMVNARQHRTVRFGSASVGSP*TQKTVWKDTGPRCGCPCQD*K 315
+ + + GS ++ + + G + S T KT D G D
Sbjct: 254 LLLILSWLVLGSLSNYGL-------KRPDIGPMELKSM-TGKTPVLDIGALEKIKSGD-V 304
Query: 316 RRHQGIKRLKRYTVEFADGSTENFDAIILATGYKSNVPYWLKDKGMFSKEDGYPRKPFPN 375
IK+ R+ VE DG + DA++LATGY+SNVP WL++ FSK +G+P+ PFPN
Sbjct: 305 EIVPAIKQFSRHHVELVDGQKLDIDAVVLATGYRSNVPSWLQESEFFSK-NGFPKSPFPN 363
Query: 376 GWKGENGLYAVGFTKRGLLGASMDAKNIAEDIERCWKAEAK 416
WKG++GLYA GFT++GL GAS+DA NIA+DI W+ E K
Sbjct: 364 AWKGKSGLYAAGFTRKGLAGASVDAVNIAQDIGNVWREETK 404
>gb|AAC04900.1| putative flavin-containing monooxygenase [Arabidopsis thaliana]
gi|25408251|pir||H84742 probable flavin-containing
monooxygenase [imported] - Arabidopsis thaliana
gi|15225816|ref|NP_180881.1| flavin-containing
monooxygenase, putative / FMO, putative [Arabidopsis
thaliana]
Length = 431
Score = 373 bits (957), Expect = e-102
Identities = 211/430 (49%), Positives = 259/430 (60%), Gaps = 51/430 (11%)
Query: 22 MINNNKNSTSS-----------SDRCLWIPGPLIVGAGPSGLAVAAYLKQKGVPSLILER 70
M NNN S + S RC+W+ GP+IVGAGPSGLAVAA LK++ VP +ILER
Sbjct: 1 MCNNNNTSCVNISSMLQPEDIFSRRCIWVNGPVIVGAGPSGLAVAADLKRQEVPFVILER 60
Query: 71 SNCIASLWKLKTYDRLRLHLPKQVCELPLMEFPSGFPTYPTKQQFIEYLESYSKNFDIRP 130
+NCIASLW+ +TYDRL+LHLPKQ C+LP + FP P YPTK QFIEYLESY+ +FD+RP
Sbjct: 61 ANCIASLWQNRTYDRLKLHLPKQFCQLPNLPFPEDIPEYPTKYQFIEYLESYATHFDLRP 120
Query: 131 WFNETVMHAEFDATLGFWRV----RSEGKAGMVTEFVCRWLIVATGENAEAVVPEIEGVD 186
FNETV A++D G WRV RSE E++CRWL+VATGENAE VVPE EG++
Sbjct: 121 KFNETVQSAKYDKRFGLWRVQTVLRSELLGYCEFEYICRWLVVATGENAEKVVPEFEGLE 180
Query: 187 EFVGSIRHTSLYKSGEEFRGKKVLVVGCGNSGMEVCLDLCNHDAAPSIVVRDSP---TKR 243
+F G + H YKSGE +RGK+VLVVGCGNSGMEV LDLCNHDA+PS+VVR S +
Sbjct: 181 DFGGDVLHAGDYKSGERYRGKRVLVVGCGNSGMEVSLDLCNHDASPSMVVRSSVHVLPRE 240
Query: 244 YAGKINF------------WVVHVVAQVATRATGGSHPSHCVMVNARQHRTVRFG--SAS 289
GK F W+V V TR G+ T ++G
Sbjct: 241 VLGKSTFELSVTMMKWMPVWLVDKTLLVLTRLLLGN--------------TDKYGLKRPE 286
Query: 290 VGSP*TQKTVWKDTGPRCGCPCQD*KRRHQ---GIKRLKRYTVEFADGSTENFDAIILAT 346
+G + T K G + + GI + VE DG D++ILAT
Sbjct: 287 IGPLELKNTAGKTPVLDIGAISMIKSGKIKIVAGIAKFGPGKVELVDGRVLQIDSVILAT 346
Query: 347 GYKSNVPYWLKDKGMFSKEDGYPRKPFPNGWKGENGLYAVGFTKRGLLGASMDAKNIAED 406
GY+SNVP WLK+ + E G + PFP GWKG+ GLYAVGFT RGL GAS DA ++A D
Sbjct: 347 GYRSNVPSWLKENDL--GEIGIEKNPFPKGWKGKAGLYAVGFTGRGLSGASFDAMSVAHD 404
Query: 407 IERCWKAEAK 416
I WK E K
Sbjct: 405 IANSWKEETK 414
>ref|NP_171955.1| flavin-containing monooxygenase / FMO (YUCCA3) [Arabidopsis
thaliana] gi|16555356|gb|AAL23752.1| flavin-containing
monooxygenase YUCCA3 [Arabidopsis thaliana]
gi|46518417|gb|AAS99690.1| At1g04610 [Arabidopsis
thaliana] gi|2494132|gb|AAB80641.1| Contains similarity
to human dimethylaniline monooxygenase (gb|M64082).
[Arabidopsis thaliana] gi|25406796|pir||G86178
hypothetical protein [imported] - Arabidopsis thaliana
gi|40823347|gb|AAR92277.1| At1g04610 [Arabidopsis
thaliana]
Length = 437
Score = 370 bits (950), Expect = e-101
Identities = 200/410 (48%), Positives = 256/410 (61%), Gaps = 43/410 (10%)
Query: 33 SDRCLWIPGPLIVGAGPSGLAVAAYLKQKGVPSLILERSNCIASLWKLKTYDRLRLHLPK 92
S RC+W+ GP+IVGAGPSGLAVAA LK++GVP +ILER+NCIASLW+ +TYDRL+LHLPK
Sbjct: 28 SRRCIWVNGPVIVGAGPSGLAVAAGLKREGVPFIILERANCIASLWQNRTYDRLKLHLPK 87
Query: 93 QVCELPLMEFPSGFPTYPTKQQFIEYLESYSKNFDIRPWFNETVMHAEFDATLGFWRVRS 152
Q C+LP FP FP YPTK QFI+YLESY+ NFDI P FNETV A++D T G WRV++
Sbjct: 88 QFCQLPNYPFPDEFPEYPTKFQFIQYLESYAANFDINPKFNETVQSAKYDETFGLWRVKT 147
Query: 153 EGKAGMV----TEFVCRWLIVATGENAEAVVPEIEGVDEFVGSIRHTSLYKSGEEFRGKK 208
G + E++CRW++VATGENAE VVP+ EG+++F G + H YKSG ++GKK
Sbjct: 148 ISNMGQLGSCEFEYICRWIVVATGENAEKVVPDFEGLEDFGGDVLHAGDYKSGGRYQGKK 207
Query: 209 VLVVGCGNSGMEVCLDLCNHDAAPSIVVRDSP---TKRYAGKINF------------WVV 253
VLVVGCGNSGMEV LDL NH A PS+VVR + + GK F W+
Sbjct: 208 VLVVGCGNSGMEVSLDLYNHGANPSMVVRSAVHVLPREIFGKSTFELGVTMMKYMPVWLA 267
Query: 254 HVVAQVATRATGGSHPSHCVMVNARQHRTVRFGSASVGSP*TQKTVWKDTGPRCGCPCQD 313
R G+ + + + + G + + KT D G
Sbjct: 268 DKTILFLARIILGNTDKYGL-------KRPKIGPLELKNK-EGKTPVLDIGAL------- 312
Query: 314 *KRRHQGIKRLKRYTVEFADGSTE-------NFDAIILATGYKSNVPYWLKDKGMFSKED 366
+ G ++ ++F G E D++ILATGY+SNVP WLKD FS +D
Sbjct: 313 -PKIRSGKIKIVPGIIKFGKGKVELIDGRVLEIDSVILATGYRSNVPSWLKDNDFFS-DD 370
Query: 367 GYPRKPFPNGWKGENGLYAVGFTKRGLLGASMDAKNIAEDIERCWKAEAK 416
G P+ PFPNGWKGE GLYAVGFT++GL GAS+DA ++A DI WK E+K
Sbjct: 371 GIPKNPFPNGWKGEAGLYAVGFTRKGLFGASLDAMSVAHDIANRWKEESK 420
>emb|CAE76085.1| B1340F09.23 [Oryza sativa (japonica cultivar-group)]
gi|50921567|ref|XP_471144.1| B1340F09.23 [Oryza sativa
(japonica cultivar-group)] gi|38346518|emb|CAE03813.2|
OSJNBa0027H09.13 [Oryza sativa (japonica
cultivar-group)]
Length = 419
Score = 368 bits (944), Expect = e-100
Identities = 197/399 (49%), Positives = 254/399 (63%), Gaps = 25/399 (6%)
Query: 33 SDRCLWIPGPLIVGAGPSGLAVAAYLKQKGVPSLILERSNCIASLWKLKTYDRLRLHLPK 92
S + W+ GP+IVGAGPSGLAVAA L+++GVP +LER++CIASLW+ +TYDRL+LHLPK
Sbjct: 14 SPQTSWVSGPIIVGAGPSGLAVAASLREQGVPFTMLERADCIASLWQKRTYDRLKLHLPK 73
Query: 93 QVCELPLMEFPSGFPTYPTKQQFIEYLESYSKNFDIRPWFNETVMHAEFDATLGFWRVRS 152
Q CELP M FP+ +P YPT++QFI+YLE Y+ FDI P F TV+ A +D T G WRVR+
Sbjct: 74 QFCELPRMAFPAHYPEYPTRRQFIDYLEDYAAAFDINPLFGHTVLSARYDETSGLWRVRA 133
Query: 153 EGKAGMVTEFVCRWLIVATGENAEAVVPEIEGVDEFVGSIRHTSLYKSGEEFRGKKVLVV 212
AG E++ WL+VATGENAE+VVP+I G+D F G + H + YKSGE +RGK+VLVV
Sbjct: 134 SSSAGAEMEYIGSWLVVATGENAESVVPDIPGIDGFGGEVVHVADYKSGEAYRGKRVLVV 193
Query: 213 GCGNSGMEVCLDLCNHDAAPSIVVRDSP---TKRYAGKINF------------WVVHVVA 257
GCGNSGMEV LDLC+H A P++VVRD+ + GK F W+V +
Sbjct: 194 GCGNSGMEVSLDLCDHGARPAMVVRDAVHVLPREVLGKSTFELAVLLMAWLPLWLVDKIL 253
Query: 258 QVATRATGGSHPSHCVMVNARQHRTVRFGSASVGSP*TQKTVWKDTGPRCGCPCQD*KRR 317
+ G + + R+ T G + + T +T D G +
Sbjct: 254 VLLAWLVLG----NLAKLGIRRPAT---GPLELKNT-TGRTPVLDYGALARIRSGE-ITV 304
Query: 318 HQGIKRLKRYTVEFADGSTENFDAIILATGYKSNVPYWLKDKGMFSKEDGYPRKPFPNGW 377
G+ R R E ADG DA++LATGY+SNVP WL+ F+K DGYP+ FPNGW
Sbjct: 305 VPGVARFGRGFAELADGRVIALDAVVLATGYRSNVPQWLQGNDFFNK-DGYPKTAFPNGW 363
Query: 378 KGENGLYAVGFTKRGLLGASMDAKNIAEDIERCWKAEAK 416
KGE+GLYAVGFT+RGL GAS DA A+D+ R WK K
Sbjct: 364 KGESGLYAVGFTRRGLSGASADAMRAAKDLARVWKEATK 402
>gb|AAK74069.1| flavin monoxygenase-like protein floozy [Petunia x hybrida]
Length = 412
Score = 365 bits (936), Expect = 2e-99
Identities = 197/397 (49%), Positives = 252/397 (62%), Gaps = 32/397 (8%)
Query: 32 SSDRCLWIPGPLIVGAGPSGLAVAAYLKQKGVPSLILERSNCIASLWKLKTYDRLRLHLP 91
+ + LW+ GP+I+GAGPSGLAV+A LK+ GVPSLILERS+CIASLW+ KTYDRL+LHLP
Sbjct: 9 TQQKWLWVNGPIIIGAGPSGLAVSACLKENGVPSLILERSDCIASLWQHKTYDRLKLHLP 68
Query: 92 KQVCELPLMEFPSGFPTYPTKQQFIEYLESYSKNFDIRPWFNETVMHAEFDATLGFWRVR 151
KQ C+LPL FP FP YPTK+QFI YLESY+K+F I P + + V AEFD GFW+V+
Sbjct: 69 KQFCQLPLFGFPDNFPKYPTKRQFISYLESYAKHFSINPKYKQAVQVAEFDHVSGFWKVQ 128
Query: 152 SEGKAGMVTEFVCRWLIVATGENAEAVVPEIEGVDEFVGSIRHTSLYKSGEEFRGKKVLV 211
++ ++ +WLIVATGENAE V+P I+G+D+F G + HTSLYKSG EF ++VLV
Sbjct: 129 TQN-----FQYFSKWLIVATGENAEPVIPNIQGMDKFKGPVMHTSLYKSGTEFNNQRVLV 183
Query: 212 VGCGNSGMEVCLDLCNHDAAPSIVVRDSP---TKRYAGKINFWVVHVVAQ---------- 258
+GCGN GMEV LDLC H+A P +V R+S + G F + + +
Sbjct: 184 IGCGNFGMEVSLDLCRHNAIPHMVARNSVHILPREMLGISTFSMAMALLKCLPLRIVDKF 243
Query: 259 ---VATRATGGSHPSHCVMVNARQHRTVRFGSASVGSP*TQKTVWKDTGPRCGCPCQD*K 315
VA G + + R+ +T G + + T KT D G +
Sbjct: 244 LLLVANLTLGNTD-----KLGLRRPKT---GPIELKNA-TGKTPVLDVGALSQIKAGKIQ 294
Query: 316 RRHQGIKRLKRYTVEFADGSTENFDAIILATGYKSNVPYWLKDKGMFSKEDGYPRKPFPN 375
H +K + + +F DG FD+IILATGYKSNVP WLK F+ E G P+ PFPN
Sbjct: 295 IVH-AVKEITKIGAKFVDGKEGEFDSIILATGYKSNVPSWLKGTEFFT-EQGMPKTPFPN 352
Query: 376 GWKGENGLYAVGFTKRGLLGASMDAKNIAEDIERCWK 412
GWKGENGLY VGFT+RGLLG + DAKNIA DI W+
Sbjct: 353 GWKGENGLYTVGFTRRGLLGTACDAKNIARDIGDEWR 389
>emb|CAB81460.1| putative protein [Arabidopsis thaliana] gi|4218126|emb|CAA22980.1|
putative protein [Arabidopsis thaliana]
gi|15235409|ref|NP_194601.1| flavin-containing
monooxygenase family protein / FMO family protein
[Arabidopsis thaliana] gi|7485532|pir||T04527
hypothetical protein F16A16.170 - Arabidopsis thaliana
Length = 426
Score = 360 bits (923), Expect = 6e-98
Identities = 202/410 (49%), Positives = 257/410 (62%), Gaps = 41/410 (10%)
Query: 32 SSDRCLWIPGPLIVGAGPSGLAVAAYLKQKGVPSLILERSNCIASLWKLKTYDRLRLHLP 91
+++RC+W+ GP+IVGAGPSGLA AA L ++ VP ++LER++CIASLW+ +TYDRL+LHLP
Sbjct: 15 TNNRCIWVNGPVIVGAGPSGLATAACLHEQNVPFVVLERADCIASLWQKRTYDRLKLHLP 74
Query: 92 KQVCELPLMEFPSGFPTYPTKQQFIEYLESYSKNFDIRPWFNETVMHAEFDATLGFWRVR 151
KQ C+LP M FP FP YPTK+QFI+YLESY+ F+I P FNE V A FD T G WRV+
Sbjct: 75 KQFCQLPKMPFPEDFPEYPTKRQFIDYLESYATRFEINPKFNECVQTARFDETSGLWRVK 134
Query: 152 SEGKAGMV---TEFVCRWLIVATGENAEAVVPEIEGVDEFVGSIRHTSLYKSGEEFRGKK 208
+ K+ E++CRWL+VATGENAE V+PEI+G+ EF G + H YKSGE+F GKK
Sbjct: 135 TVSKSESTQTEVEYICRWLVVATGENAERVMPEIDGLSEFSGEVIHACDYKSGEKFAGKK 194
Query: 209 VLVVGCGNSGMEVCLDLCNHDAAPSIVVRDS---PTKRYAGKINF------------WVV 253
VLVVGCGNSGMEV LDL NH A PS+VVR S + GK F W+V
Sbjct: 195 VLVVGCGNSGMEVSLDLANHFAKPSMVVRSSLHVMPREVMGKSTFELAMKMLRWFPLWLV 254
Query: 254 HVVAQVATRATGGSHPSHCVMVNARQHRTVR--FGSASVGSP*TQKTVWKDTGP----RC 307
+ V S V+ N ++ R G + S KT D G R
Sbjct: 255 DKILLVL---------SWMVLGNIEKYGLKRPEMGPMELKSV-KGKTPVLDIGAIEKIRL 304
Query: 308 GCPCQD*KRRHQGIKRLKRYTVEFADGSTENFDAIILATGYKSNVPYWLKDKGMFSKEDG 367
G GIKR VE +G + D+++LATGY+SNVPYWL++ F+K +G
Sbjct: 305 GK-----INVVPGIKRFNGNKVELVNGEQLDVDSVVLATGYRSNVPYWLQENEFFAK-NG 358
Query: 368 YPRKPFP-NGWKGENGLYAVGFTKRGLLGASMDAKNIAEDIERCWKAEAK 416
+P+ NGWKG GLYAVGFT++GL GASMDA IA+DI W+ E K
Sbjct: 359 FPKTVADNNGWKGRTGLYAVGFTRKGLSGASMDAVKIAQDIGSVWQLETK 408
>emb|CAB96667.1| putative protein [Arabidopsis thaliana]
gi|15239020|ref|NP_196693.1| flavin-containing
monooxygenase family protein / FMO family protein
[Arabidopsis thaliana]
Length = 411
Score = 348 bits (892), Expect = 2e-94
Identities = 190/386 (49%), Positives = 240/386 (61%), Gaps = 22/386 (5%)
Query: 37 LWIPGPLIVGAGPSGLAVAAYLKQKGVPSLILERSNCIASLWKLKTYDRLRLHLPKQVCE 96
+++PGP+IVGAGPSGLAVAA L +GVPS+ILER++C+ASLW+ +TYDRL+LHLPK CE
Sbjct: 12 IFVPGPIIVGAGPSGLAVAACLSNRGVPSVILERTDCLASLWQKRTYDRLKLHLPKHFCE 71
Query: 97 LPLMEFPSGFPTYPTKQQFIEYLESYSKNFDIRPWFNETVMHAEFDATLGFWRVRSEGKA 156
LPLM FP FP YP+KQ FI Y+ESY+ F+I+P FN+TV AEFD G W V+++
Sbjct: 72 LPLMPFPKNFPKYPSKQLFISYVESYAARFNIKPVFNQTVEKAEFDDASGLWNVKTQDGV 131
Query: 157 GMVTEFVCRWLIVATGENAEAVVPEIEGVDEFVGSIRHTSLYKSGEEFRGKKVLVVGCGN 216
+ WL+VATGENAE V P I G+ +F G + HTS YKSG F +KVLVVGCGN
Sbjct: 132 -----YTSTWLVVATGENAEPVFPNIPGLKKFTGPVVHTSAYKSGSAFANRKVLVVGCGN 186
Query: 217 SGMEVCLDLCNHDAAPSIVVRDSP---TKRYAGKINF--------WVVHVVAQVATRATG 265
SGMEV LDLC ++A P +VVR+S + + G F W +
Sbjct: 187 SGMEVSLDLCRYNALPHMVVRNSVHVLPRDFFGLSTFGIAMTLLKWFPLKLVDKFLLLLA 246
Query: 266 GSHPSHCVMVNARQHRTVRFGSASVGSP*TQKTVWKDTGPRCGCPCQD*KRRHQGIKRLK 325
S + ++ R+ +T +V T KT D G K Q +K +
Sbjct: 247 NSTLGNTDLLGLRRPKTGPIELKNV----TGKTPVLDVGAISLIRSGQIKVT-QAVKEIT 301
Query: 326 RYTVEFADGSTENFDAIILATGYKSNVPYWLKDKGMFSKEDGYPRKPFPNGWKGENGLYA 385
R +F +G FD+IILATGYKSNVP WLK+ F+KE G P+ PFPNGWKGE GLY
Sbjct: 302 RNGAKFLNGKEIEFDSIILATGYKSNVPDWLKENSFFTKE-GMPKTPFPNGWKGEKGLYT 360
Query: 386 VGFTKRGLLGASMDAKNIAEDIERCW 411
VGFT+RGL G + DA IAEDI W
Sbjct: 361 VGFTRRGLSGTAYDAVKIAEDITDQW 386
>ref|NP_913428.1| dimethylaniline monooxygenase-like protein [Oryza sativa (japonica
cultivar-group)] gi|12698319|dbj|BAB07916.2| putative
flavin-containing monooxygenase YUCCA3 [Oryza sativa
(japonica cultivar-group)] gi|13027342|dbj|BAB32703.1|
putative flavin-containing monooxygenase YUCCA3 [Oryza
sativa (japonica cultivar-group)]
Length = 439
Score = 341 bits (875), Expect = 2e-92
Identities = 200/444 (45%), Positives = 262/444 (58%), Gaps = 57/444 (12%)
Query: 1 MDCYLRELEGKQAHDPLFIEKMINNNKNSTS----SSDRCLWIPGPLIVGAGPSGLAVAA 56
MDC+ E EGK+AHDPL+ + +T D+ + +PG +IVGAGP+G+AV A
Sbjct: 1 MDCFA-ETEGKRAHDPLYQRRAAAAATPATGVPVDDVDKVVDVPGAVIVGAGPAGVAVGA 59
Query: 57 YLKQKGVPSLILERSNCIASLWKLKTYDRLRLHLPKQVCELPLMEFPSGFPTYPTKQQFI 116
L +GV ++LER CIASLW+ +TYDRL LHLPK+ CELPL FP+ FP YPT+ QF+
Sbjct: 60 LLGLRGVAYVVLERCGCIASLWRHRTYDRLCLHLPKRFCELPLRPFPASFPEYPTRDQFL 119
Query: 117 EYLESYSKNFDIRPWFNETVMHAEFDATLGFWRVRSEGKAG-----------MVTEFVCR 165
YL++Y++ F + P F V+ AE+D +W E A +T + R
Sbjct: 120 GYLDAYAREFGVEPVFRRAVISAEYDGE-SWWVYTREVVAAAAGGEQAVLGCTMTVYRSR 178
Query: 166 WLIVATGENAEAVVPEIEGVDEFVGSIRHTSLYKSGEEFRGKKVLVVGCGNSGMEVCLDL 225
WL+VATGENAE VVPE++G F G + H+S Y++G+ + GKKVLVVGCGNSGMEV LDL
Sbjct: 179 WLVVATGENAEPVVPEMDGAGRFKGQMMHSSEYRNGDGYAGKKVLVVGCGNSGMEVSLDL 238
Query: 226 CNHDAAPSIVVRDSP---TKRYAGKINF----------------WVVHVVAQVA---TRA 263
CNH+A S+VVRD+ + G F W+V +++ + T
Sbjct: 239 CNHNARASMVVRDTVHVLPREILGFSTFGLSMWLLRWLSVQTVDWLVLLLSFLVFGDTAR 298
Query: 264 TGGSHPSHCVMVNARQHRTVRFGSASVGSP*TQKTVWKDTGPRCGCPCQD*KRRHQGIKR 323
G PS G + S + KT D G D K I+
Sbjct: 299 LGIPRPS--------------LGPFELKSV-SGKTPVLDVGTLAKIKSGDIKVT-PAIQC 342
Query: 324 LKRYTVEFADGSTENFDAIILATGYKSNVPYWLKDKGMFSKEDGYPRKPFPNGWKGENGL 383
+ + VEF DGSTE FD +ILATGYKSNVPYWLK+K FS++DG+PRK N WKG+NGL
Sbjct: 343 FQEHGVEFVDGSTEEFDVVILATGYKSNVPYWLKEKEFFSEKDGFPRK--GNAWKGQNGL 400
Query: 384 YAVGFTKRGLLGASMDAKNIAEDI 407
YAVGF++RGL G SMDA NI +DI
Sbjct: 401 YAVGFSRRGLSGVSMDANNIVQDI 424
>ref|NP_917203.1| P0707D10.20 [Oryza sativa (japonica cultivar-group)]
Length = 406
Score = 339 bits (870), Expect = 8e-92
Identities = 190/396 (47%), Positives = 243/396 (60%), Gaps = 20/396 (5%)
Query: 28 NSTSSSDRCLWIPGPLIVGAGPSGLAVAAYLKQKGVPSLILERSNCIASLWKLKTYDRLR 87
N + R W+PG +IVGAGPSGLA AA L +GVP+ +LERS+ +AS W+ + YDRL
Sbjct: 3 NKPAQERRETWVPGAVIVGAGPSGLAAAACLAARGVPATVLERSDSLASTWRHRMYDRLA 62
Query: 88 LHLPKQVCELPLMEFPSGFPTYPTKQQFIEYLESYSKNFDIRPWFNETVMHAEFDATLGF 147
LHLPK+ CELPL+ FP +PTYP+K QF+ Y+E+Y+ + P F TV A FDA +G
Sbjct: 63 LHLPKRFCELPLLPFPEEYPTYPSKDQFVAYMEAYAAAAGVAPRFGATVEEAAFDAAVGA 122
Query: 148 WRVRSEGKAGMVTEFVCRWLIVATGENAEAVVPEIEGVDEFVGSIRHTSLYKSGEEFRGK 207
WRVR +G G V + RWL+VATGENAE VP+ G+ +F G HTS YKSGE+F GK
Sbjct: 123 WRVRLDG--GEV--LMARWLVVATGENAEPRVPDFPGMQKFAGCAMHTSEYKSGEQFAGK 178
Query: 208 KVLVVGCGNSGMEVCLDLCNHDAAPSIVVRDSP---TKRYAGKINFWVVHVV-----AQV 259
KVLVVGCGNSGMEV LDLC H A PS+VVR++ + G F + + Q+
Sbjct: 179 KVLVVGCGNSGMEVSLDLCRHGAKPSMVVRNTVHVLPREMFGLSTFGIAMALLRWLPVQL 238
Query: 260 ATRATGGSHPSHCVMVNARQH--RTVRFGSASVGSP*TQKTVWKDTGPRCGCPCQD*KRR 317
R +H ++ N Q R + G + + T +T D G + K +
Sbjct: 239 VDRFL--LTAAHLILGNTGQFGLRRPKTGPIELKNL-TGRTPVLDVGTL--DHIKSGKIK 293
Query: 318 HQG-IKRLKRYTVEFADGSTENFDAIILATGYKSNVPYWLKDKGMFSKEDGYPRKPFPNG 376
G +K + R V F DG E FD IILATGY+SNVP WLKD G +G + PFPN
Sbjct: 294 VVGAVKEMTRQGVRFTDGKEEQFDTIILATGYRSNVPSWLKDAGDLFTREGISKVPFPNS 353
Query: 377 WKGENGLYAVGFTKRGLLGASMDAKNIAEDIERCWK 412
W+G NGLY VGFT+RGLLG S DA N+A+DI W+
Sbjct: 354 WRGRNGLYTVGFTQRGLLGTSSDALNVAKDIHCQWR 389
>emb|CAB79971.1| dimethylaniline monooxygenase-like protein [Arabidopsis thaliana]
gi|4914451|emb|CAB43691.1| dimethylaniline
monooxygenase-like protein [Arabidopsis thaliana]
gi|15236840|ref|NP_194980.1| flavin-containing
monooxygenase / FMO (YUCCA) [Arabidopsis thaliana]
gi|16555352|gb|AAL23750.1| flavin-containing
monooxygenase YUCCA [Arabidopsis thaliana]
gi|7486767|pir||T08587 hypothetical protein L23H3.20 -
Arabidopsis thaliana
Length = 414
Score = 331 bits (849), Expect = 2e-89
Identities = 193/415 (46%), Positives = 250/415 (59%), Gaps = 38/415 (9%)
Query: 22 MINNNKNSTSSSDRCLWIPGPLIVGAGPSGLAVAAYLKQKGVPSLILERSNCIASLWKLK 81
M ++ N T + + + GP+I+GAGPSGLA +A L +GVPSLILERS+ IASLWK K
Sbjct: 1 MESHPHNKTDQTQHIILVHGPIIIGAGPSGLATSACLSSRGVPSLILERSDSIASLWKSK 60
Query: 82 TYDRLRLHLPKQVCELPLMEFPSGFPTYPTKQQFIEYLESYSKNFDIRPWFNETVMHAEF 141
TYDRLRLHLPK C LPL++FP +P YP+K +F+ YLESY+ +F I P FN+ V +A +
Sbjct: 61 TYDRLRLHLPKHFCRLPLLDFPEYYPKYPSKNEFLAYLESYASHFRIAPRFNKNVQNAAY 120
Query: 142 DATLGFWRVRSEGKAGMVTEFVCRWLIVATGENAEAVVPEIEGVDEFV-GSIRHTSLYKS 200
D++ GFWRV++ TE++ +WLIVATGENA+ PEI G +F G I H S YKS
Sbjct: 121 DSSSGFWRVKTHDN----TEYLSKWLIVATGENADPYFPEIPGRKKFSGGKIVHASEYKS 176
Query: 201 GEEFRGKKVLVVGCGNSGMEVCLDLCNHDAAPSIVVRDSPTKRYAGKINFWVVHVVAQ-- 258
GEEFR +KVLVVGCGNSGME+ LDL H+A+P +VVR++ VHV+ +
Sbjct: 177 GEEFRRQKVLVVGCGNSGMEISLDLVRHNASPHLVVRNT-------------VHVLPREI 223
Query: 259 --VATRATGGS----HPSHCV------MVNARQHRTVRFG--SASVGSP*TQKTVWKDTG 304
V+T G + P V M N T R G G + K
Sbjct: 224 LGVSTFGVGMTLLKCLPLRLVDKFLLLMANLSFGNTDRLGLRRPKTGPLELKNVTGKSPV 283
Query: 305 PRCGCPCQD*KRRHQGIKRLKRYT---VEFADGSTENFDAIILATGYKSNVPYWLKDKGM 361
G Q ++ +K T +F DG ++FD+II ATGYKSNVP WL+ G
Sbjct: 284 LDVGAMSLIRSGMIQIMEGVKEITKKGAKFMDGQEKDFDSIIFATGYKSNVPTWLQG-GD 342
Query: 362 FSKEDGYPRKPFPNGWKGENGLYAVGFTKRGLLGASMDAKNIAEDIERCWKAEAK 416
F +DG P+ PFPNGW+G GLY VGFT+RGLLG + DA IA +I W+ E K
Sbjct: 343 FFTDDGMPKTPFPNGWRGGKGLYTVGFTRRGLLGTASDAVKIAGEIGDQWRDEIK 397
>gb|AAU43964.1| putative dimethylaniline monooxygenase [Oryza sativa (japonica
cultivar-group)]
Length = 348
Score = 318 bits (815), Expect = 2e-85
Identities = 175/341 (51%), Positives = 221/341 (64%), Gaps = 29/341 (8%)
Query: 37 LWIPGPLIVGAGPSGLAVAAYLKQKGVPSLILERSNCIASLWKLKTYDRLRLHLPKQVCE 96
+W+ GP++VGAGPSGLA AA LK+KG+ SL+LERS+C+A LW+LK YDRL LHLP+Q CE
Sbjct: 3 VWVQGPIVVGAGPSGLAAAACLKEKGIDSLVLERSSCLAPLWQLKMYDRLSLHLPRQFCE 62
Query: 97 LPLMEFPSGFPTYPTKQQFIEYLESYSKNFDIRPWFNETVMHAEFDATLGFWRVRSEGKA 156
LPL FP+ +P YPTKQQF+ YLESY+ F I P +N TV+ AEFD L WRVR+
Sbjct: 63 LPLFPFPASYPDYPTKQQFVAYLESYAAKFGINPMYNHTVVCAEFDERLMLWRVRTTQAT 122
Query: 157 GMV---TEFVCRWLIVATGENAEAVVPEIEGVDEFVGSIRHTSLYKSGEEFRGKKVLVVG 213
GM+ E+V +WL+VATGEN+EAV+P I+G++EF GS+ HTS YKSG +F GK VLVVG
Sbjct: 123 GMMEDDVEYVSQWLVVATGENSEAVLPVIDGLEEFRGSVIHTSAYKSGSKFAGKTVLVVG 182
Query: 214 CGNSGMEVCLDLCNHDAAPSIVVR-------DSPTKRYAGKINFWV-VHVVAQVATRATG 265
CGNSGMEVCLDLCNH+ P IVV PT R A + W+ +H+V ++
Sbjct: 183 CGNSGMEVCLDLCNHNGYPRIVVHILPREMLGQPTFRLAMWLLKWLPIHIVDRIL----- 237
Query: 266 GSHPSHCVMVNARQHRTVRFG--SASVG----SP*TQKTVWKDTGPRCGCPCQD*KRRHQ 319
++ A T +FG S+G + KT D G D K R
Sbjct: 238 ------LLVARAILGDTSQFGLKRPSLGPLELKSLSGKTPILDIGTLAKIKSGDIKVR-P 290
Query: 320 GIKRLKRYTVEFADGSTENFDAIILATGYKSNVPYWLKDKG 360
I+R+ V+F DG +E FDAI+LATGYKSNVP WLK G
Sbjct: 291 AIRRIAGQQVKFVDGRSEQFDAIVLATGYKSNVPCWLKVYG 331
>ref|NP_850808.1| flavin-containing monooxygenase family protein / FMO family protein
[Arabidopsis thaliana]
Length = 357
Score = 281 bits (718), Expect = 4e-74
Identities = 157/332 (47%), Positives = 202/332 (60%), Gaps = 21/332 (6%)
Query: 37 LWIPGPLIVGAGPSGLAVAAYLKQKGVPSLILERSNCIASLWKLKTYDRLRLHLPKQVCE 96
+++PGP+IVGAGPSGLAVAA L +GVPS+ILER++C+ASLW+ +TYDRL+LHLPK CE
Sbjct: 12 IFVPGPIIVGAGPSGLAVAACLSNRGVPSVILERTDCLASLWQKRTYDRLKLHLPKHFCE 71
Query: 97 LPLMEFPSGFPTYPTKQQFIEYLESYSKNFDIRPWFNETVMHAEFDATLGFWRVRSEGKA 156
LPLM FP FP YP+KQ FI Y+ESY+ F+I+P FN+TV AEFD G W V+++
Sbjct: 72 LPLMPFPKNFPKYPSKQLFISYVESYAARFNIKPVFNQTVEKAEFDDASGLWNVKTQDGV 131
Query: 157 GMVTEFVCRWLIVATGENAEAVVPEIEGVDEFVGSIRHTSLYKSGEEFRGKKVLVVGCGN 216
+ WL+VATGENAE V P I G+ +F G + HTS YKSG F +KVLVVGCGN
Sbjct: 132 -----YTSTWLVVATGENAEPVFPNIPGLKKFTGPVVHTSAYKSGSAFANRKVLVVGCGN 186
Query: 217 SGMEVCLDLCNHDAAPSIVVRDSP---TKRYAGKINF--------WVVHVVAQVATRATG 265
SGMEV LDLC ++A P +VVR+S + + G F W +
Sbjct: 187 SGMEVSLDLCRYNALPHMVVRNSVHVLPRDFFGLSTFGIAMTLLKWFPLKLVDKFLLLLA 246
Query: 266 GSHPSHCVMVNARQHRTVRFGSASVGSP*TQKTVWKDTGPRCGCPCQD*KRRHQGIKRLK 325
S + ++ R+ +T +V T KT D G K Q +K +
Sbjct: 247 NSTLGNTDLLGLRRPKTGPIELKNV----TGKTPVLDVGAISLIRSGQIKVT-QAVKEIT 301
Query: 326 RYTVEFADGSTENFDAIILATGYKSNVPYWLK 357
R +F +G FD+IILATGYKSNVP WLK
Sbjct: 302 RNGAKFLNGKEIEFDSIILATGYKSNVPDWLK 333
>ref|XP_477812.1| putative dimethylaniline monooxygenase [Oryza sativa (japonica
cultivar-group)] gi|33147034|dbj|BAC80117.1| putative
dimethylaniline monooxygenase [Oryza sativa (japonica
cultivar-group)]
Length = 398
Score = 279 bits (714), Expect = 1e-73
Identities = 126/209 (60%), Positives = 167/209 (79%), Gaps = 4/209 (1%)
Query: 35 RCLWIPGPLIVGAGPSGLAVAAYLKQKGVPSLILERSNCIASLWKLKTYDRLRLHLPKQV 94
R +W+ GP++VGAGP+GL+VAA L+++GVPS++LER++CIASLW+ +TYDRLRLHLPK
Sbjct: 4 RVVWVNGPIVVGAGPAGLSVAACLRERGVPSVLLERADCIASLWQRRTYDRLRLHLPKHF 63
Query: 95 CELPLMEFPSGFPTYPTKQQFIEYLESYSKNFDIRPWFNETVMHAEFDATLGFWRVRSE- 153
CELP M FP G+P YP ++QF++YL++Y+ + P FN++V A +D G WRVR+E
Sbjct: 64 CELPGMPFPDGYPEYPDRRQFVDYLQAYAARAGVEPRFNQSVTSARYDDAAGLWRVRAED 123
Query: 154 ---GKAGMVTEFVCRWLIVATGENAEAVVPEIEGVDEFVGSIRHTSLYKSGEEFRGKKVL 210
AG VTE++ RWL+VATGENAE VVPEI+G D+F G + H + YKSG +RGK+VL
Sbjct: 124 VSVDAAGDVTEYIGRWLVVATGENAERVVPEIDGADDFEGPVSHVAEYKSGAAYRGKRVL 183
Query: 211 VVGCGNSGMEVCLDLCNHDAAPSIVVRDS 239
VVGCGNSGMEVCLDLC+H+A P++VVRDS
Sbjct: 184 VVGCGNSGMEVCLDLCHHNALPAMVVRDS 212
Score = 115 bits (288), Expect = 3e-24
Identities = 55/92 (59%), Positives = 66/92 (70%), Gaps = 1/92 (1%)
Query: 320 GIKRLKRYTVEFADGSTENFDAIILATGYKSNVPYWLKDKGMFSKEDGYPRKPFPNGWKG 379
GI+RL R E DG DA+ILATGY+SNVP WLK F++E GYPR PFP+GWKG
Sbjct: 300 GIRRLLRGGAELVDGRRVPADAVILATGYQSNVPQWLKGSDFFTQE-GYPRVPFPDGWKG 358
Query: 380 ENGLYAVGFTKRGLLGASMDAKNIAEDIERCW 411
E+GLY+VGFT+RGL G S DA +A+DI W
Sbjct: 359 ESGLYSVGFTRRGLSGVSSDAVKVAQDIAMAW 390
>dbj|BAD68007.1| flavin-containing monooxygenase-like [Oryza sativa (japonica
cultivar-group)]
Length = 364
Score = 275 bits (704), Expect = 1e-72
Identities = 161/343 (46%), Positives = 207/343 (59%), Gaps = 20/343 (5%)
Query: 28 NSTSSSDRCLWIPGPLIVGAGPSGLAVAAYLKQKGVPSLILERSNCIASLWKLKTYDRLR 87
N + R W+PG +IVGAGPSGLA AA L +GVP+ +LERS+ +AS W+ + YDRL
Sbjct: 3 NKPAQERRETWVPGAVIVGAGPSGLAAAACLAARGVPATVLERSDSLASTWRHRMYDRLA 62
Query: 88 LHLPKQVCELPLMEFPSGFPTYPTKQQFIEYLESYSKNFDIRPWFNETVMHAEFDATLGF 147
LHLPK+ CELPL+ FP +PTYP+K QF+ Y+E+Y+ + P F TV A FDA +G
Sbjct: 63 LHLPKRFCELPLLPFPEEYPTYPSKDQFVAYMEAYAAAAGVAPRFGATVEEAAFDAAVGA 122
Query: 148 WRVRSEGKAGMVTEFVCRWLIVATGENAEAVVPEIEGVDEFVGSIRHTSLYKSGEEFRGK 207
WRVR +G G V + RWL+VATGENAE VP+ G+ +F G HTS YKSGE+F GK
Sbjct: 123 WRVRLDG--GEV--LMARWLVVATGENAEPRVPDFPGMQKFAGCAMHTSEYKSGEQFAGK 178
Query: 208 KVLVVGCGNSGMEVCLDLCNHDAAPSIVVRDSP---TKRYAGKINFWVVHVV-----AQV 259
KVLVVGCGNSGMEV LDLC H A PS+VVR++ + G F + + Q+
Sbjct: 179 KVLVVGCGNSGMEVSLDLCRHGAKPSMVVRNTVHVLPREMFGLSTFGIAMALLRWLPVQL 238
Query: 260 ATRATGGSHPSHCVMVNARQH--RTVRFGSASVGSP*TQKTVWKDTGPRCGCPCQD*KRR 317
R +H ++ N Q R + G + + T +T D G + K +
Sbjct: 239 VDRFL--LTAAHLILGNTGQFGLRRPKTGPIELKNL-TGRTPVLDVGTL--DHIKSGKIK 293
Query: 318 HQG-IKRLKRYTVEFADGSTENFDAIILATGYKSNVPYWLKDK 359
G +K + R V F DG E FD IILATGY+SNVP WLK K
Sbjct: 294 VVGAVKEMTRQGVRFTDGKEEQFDTIILATGYRSNVPSWLKVK 336
>ref|NP_912652.1| Putative flavin-containing monooxygenase [Oryza sativa (japonica
cultivar-group)] gi|22773255|gb|AAN06861.1| Putative
flavin-containing monooxygenase [Oryza sativa (japonica
cultivar-group)]
Length = 444
Score = 237 bits (605), Expect = 4e-61
Identities = 114/206 (55%), Positives = 140/206 (67%), Gaps = 4/206 (1%)
Query: 36 CLWIPGPLIVGAGPSGLAVAAYLKQKGVPSLILERSNCIASLWKLKTYDRLRLHLPKQVC 95
C+ + GP+IVGAGPSGLAVAA L+Q G P ++ERS +A LW +TYDRLRLHLPK C
Sbjct: 20 CIVLDGPIIVGAGPSGLAVAATLRQHGAPFTVVERSGGVADLWTNRTYDRLRLHLPKVFC 79
Query: 96 ELPLMEFPSGFPTYPTKQQFIEYLESYSKNFDIRPWFNETVMHAEFDATLGFWRV----R 151
ELP + FP FPTYPTK F+ YL SY+ F I P TV A +D WRV
Sbjct: 80 ELPHVAFPPDFPTYPTKHDFLRYLHSYAARFAIAPLLRRTVTRAWYDHPASLWRVTTTTT 139
Query: 152 SEGKAGMVTEFVCRWLIVATGENAEAVVPEIEGVDEFVGSIRHTSLYKSGEEFRGKKVLV 211
S ++TE+ WL+VA+GENAE VVP+++G + F G H+S Y+SGE FRG +VLV
Sbjct: 140 SSSATSVITEYASPWLVVASGENAEVVVPKVKGRERFAGEALHSSEYRSGERFRGMRVLV 199
Query: 212 VGCGNSGMEVCLDLCNHDAAPSIVVR 237
VGCGNSGME+CLDLC H A P + VR
Sbjct: 200 VGCGNSGMEMCLDLCEHGAMPFMSVR 225
Score = 82.8 bits (203), Expect = 2e-14
Identities = 42/85 (49%), Positives = 52/85 (60%), Gaps = 3/85 (3%)
Query: 331 FADGSTENFDAIILATGYKSNVPYWLKDKGMFSKEDGYPRKPFPNG---WKGENGLYAVG 387
F DG+ FDA+I ATGY+SNVP WL++ G E+G R + W+G NGLY VG
Sbjct: 348 FVDGNEMAFDAVIFATGYRSNVPSWLQEDGELFTEEGKLRSSGSSSEWRWRGPNGLYCVG 407
Query: 388 FTKRGLLGASMDAKNIAEDIERCWK 412
F+ RGLLGA DA A DI W+
Sbjct: 408 FSGRGLLGAGADALRAAADIAGRWQ 432
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.322 0.138 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 781,676,263
Number of Sequences: 2540612
Number of extensions: 34648984
Number of successful extensions: 117393
Number of sequences better than 10.0: 1128
Number of HSP's better than 10.0 without gapping: 662
Number of HSP's successfully gapped in prelim test: 466
Number of HSP's that attempted gapping in prelim test: 114651
Number of HSP's gapped (non-prelim): 2058
length of query: 423
length of database: 863,360,394
effective HSP length: 131
effective length of query: 292
effective length of database: 530,540,222
effective search space: 154917744824
effective search space used: 154917744824
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 76 (33.9 bits)
Medicago: description of AC127429.3


