

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126792.7 - phase: 0
(253 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
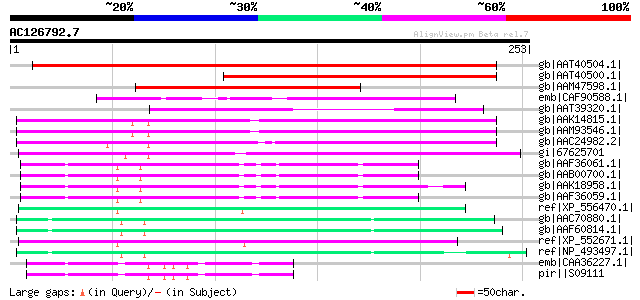
Score E
Sequences producing significant alignments: (bits) Value
gb|AAT40504.1| putative polyprotein [Solanum demissum] 275 7e-73
gb|AAT40500.1| putative reverse transcriptase [Solanum demissum] 166 6e-40
gb|AAM47598.1| NBS/LRR resistance protein-like protein [Capsicum... 123 6e-27
emb|CAF90588.1| unnamed protein product [Tetraodon nigroviridis] 94 4e-18
gb|AAT39320.1| hypothetical protein PGEC400L14.17 [Solanum demis... 87 6e-16
gb|AAK14815.1| polyprotein [Schistosoma japonicum] 84 5e-15
gb|AAM93546.1| polyprotein [Schistosoma japonicum] 84 5e-15
gb|AAC24982.2| reverse transcriptase [synthetic construct] 84 5e-15
gi|67625701 TPA: endonuclease-reverse transcriptase [Schistosoma... 75 1e-12
gb|AAF36061.1| Hypothetical protein Y76B12C.5 [Caenorhabditis el... 74 5e-12
gb|AAB00700.1| Hypothetical protein C34D4.5 [Caenorhabditis eleg... 72 1e-11
gb|AAK18958.1| Hypothetical protein F56C9.2 [Caenorhabditis eleg... 70 7e-11
gb|AAF36059.1| Hypothetical protein Y75D11A.4 [Caenorhabditis el... 69 1e-10
ref|XP_556470.1| ENSANGP00000028171 [Anopheles gambiae str. PEST... 65 2e-09
gb|AAC70880.1| Hypothetical protein F21E9.5 [Caenorhabditis eleg... 63 7e-09
gb|AAF60814.1| Hypothetical protein Y58G8A.2 [Caenorhabditis ele... 63 7e-09
ref|XP_552671.1| ENSANGP00000005174 [Anopheles gambiae str. PEST... 63 7e-09
ref|NP_493497.1| predicted CDS, reverse transcriptase family mem... 63 9e-09
emb|CAA36227.1| unnamed protein product [Drosophila melanogaster... 59 2e-07
pir||S09111 hypothetical protein 2 - fruit fly (Drosophila melan... 58 3e-07
>gb|AAT40504.1| putative polyprotein [Solanum demissum]
Length = 832
Score = 275 bits (704), Expect = 7e-73
Identities = 129/226 (57%), Positives = 166/226 (73%)
Query: 12 YEGASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVLTEHIQELAPRCMLFADDV 71
Y A T VRT G +E FP+ IGLHQGS LSP++F LV+D LT IQE P CMLFADD+
Sbjct: 370 YYTAKTRVRTVGGDSEHFPVEIGLHQGSVLSPFLFALVMDELTRSIQETVPWCMLFADDI 429
Query: 72 VLVGDSREEVNGRLETWRQALEAYGFCLSRSKTEYMECNFSGRRSKSTLEVKVGDHIIPQ 131
VL+ ++R+ VN RLE WRQ LE+ GF LSR+KTEY+ C FS ++ +EV++ +IP+
Sbjct: 430 VLIDETRDRVNARLEVWRQTLESKGFRLSRTKTEYLGCKFSDGLDETDVEVRLAAQVIPK 489
Query: 132 VTRFKYLESFVQNDGEIEADVSHRIQVGWLKWRRASCVLCDKKVPFKLKGKFYRTAVRSA 191
F+YL + +Q G+I+ DV+HR+ W+KWR AS VLCDKK+ KLKGKFYR VR A
Sbjct: 490 KESFRYLGAVIQGSGDIDDDVTHRVGAAWMKWRLASGVLCDKKISPKLKGKFYRVVVRPA 549
Query: 192 LLYGTECWAVKSQHENQVSVAEMRMLRWMSGKTRHDRIRNDIIRER 237
LLYG ECW VK+ H +++ VAEMRMLRWM G TR D+IRN++IRE+
Sbjct: 550 LLYGAECWPVKNAHVHKMHVAEMRMLRWMCGHTRSDKIRNEVIREK 595
>gb|AAT40500.1| putative reverse transcriptase [Solanum demissum]
Length = 213
Score = 166 bits (420), Expect = 6e-40
Identities = 74/133 (55%), Positives = 98/133 (73%)
Query: 105 EYMECNFSGRRSKSTLEVKVGDHIIPQVTRFKYLESFVQNDGEIEADVSHRIQVGWLKWR 164
EY+ C FS ++ +EV++ +IP+ FKYL + +Q G+I+ DV+HR+ W+KWR
Sbjct: 7 EYLGCKFSDVLDETDVEVRLAAQVIPKKESFKYLGAVIQGSGDIDDDVTHRVGAAWMKWR 66
Query: 165 RASCVLCDKKVPFKLKGKFYRTAVRSALLYGTECWAVKSQHENQVSVAEMRMLRWMSGKT 224
AS VLCDKK+P KLKGKFYR VR ALLYG ECW VK+ H +++ VAEMRMLRWM G T
Sbjct: 67 LASGVLCDKKIPLKLKGKFYRVVVRPALLYGAECWPVKNAHVHKMHVAEMRMLRWMCGHT 126
Query: 225 RHDRIRNDIIRER 237
R D+IRN++IRE+
Sbjct: 127 RSDKIRNEVIREK 139
>gb|AAM47598.1| NBS/LRR resistance protein-like protein [Capsicum annuum]
Length = 122
Score = 123 bits (308), Expect = 6e-27
Identities = 57/110 (51%), Positives = 81/110 (72%)
Query: 62 PRCMLFADDVVLVGDSREEVNGRLETWRQALEAYGFCLSRSKTEYMECNFSGRRSKSTLE 121
P CMLFADDVVL+ ++R VN +LE WRQ LE+ GF +SR+KTEY+EC F+ R ++ +
Sbjct: 10 PWCMLFADDVVLIDETRGGVNDKLELWRQTLESKGFRVSRTKTEYVECKFNDVRRENEVV 69
Query: 122 VKVGDHIIPQVTRFKYLESFVQNDGEIEADVSHRIQVGWLKWRRASCVLC 171
V++ + + +FKYL S +Q++GEI+ DVSHRI GW+KW+ AS V+C
Sbjct: 70 VRLEAQEVKKRDKFKYLGSVIQSNGEIDEDVSHRIGAGWMKWKLASGVMC 119
>emb|CAF90588.1| unnamed protein product [Tetraodon nigroviridis]
Length = 183
Score = 94.0 bits (232), Expect = 4e-18
Identities = 61/175 (34%), Positives = 96/175 (54%), Gaps = 17/175 (9%)
Query: 43 PYIFTLVLDVLTEHIQELAPRCMLFADDVVLVGDSREEVNGRLETWRQALEAYGFCLSRS 102
P + +V+D LT+ +++ +P +FA D+V+ SRE+V +LE R ALE+ S
Sbjct: 19 PLLVAMVMDRLTDEVRQESPWTTMFAGDIVMC--SREQVEEKLEERRFALES-------S 69
Query: 103 KTEYMECNFSGRRSKSTLEVKVGDHIIPQVTRFKYLESFVQNDGEIEADVSHRIQVGWLK 162
KTE E + SG E+K G+ + K L S VQ GE +V R+Q GW
Sbjct: 70 KTEN-ERDLSGSVRLQGEEIKKGEDL-------KNLGSTVQTSGECGKEVKKRVQAGWNW 121
Query: 163 WRRASCVLCDKKVPFKLKGKFYRTAVRSALLYGTECWAVKSQHENQVSVAEMRML 217
W + S V+CD+ V K+K K +T R A+++G E ++ + E ++ +AEM+ L
Sbjct: 122 WGKVSGVMCDRGVSAKIKRKVDKTVARPAIIFGLETVPLRKRQEAELELAEMKAL 176
>gb|AAT39320.1| hypothetical protein PGEC400L14.17 [Solanum demissum]
Length = 139
Score = 86.7 bits (213), Expect = 6e-16
Identities = 50/163 (30%), Positives = 75/163 (45%), Gaps = 49/163 (30%)
Query: 69 DDVVLVGDSREEVNGRLETWRQALEAYGFCLSRSKTEYMECNFSGRRSKSTLEVKVGDHI 128
+D+VL+ ++R VN R E WR L++ G LS++KTEYMEC FS K+ +V++ +
Sbjct: 4 NDIVLINETRGGVNDRQEIWRSTLDSKGLTLSKTKTEYMECKFSVASEKANRKVRIDTQL 63
Query: 129 IPQVTRFKYLESFVQNDGEIEADVSHRIQVGWLKWRRASCVLCDKKVPFKLKGKFYRTAV 188
IP+ FK + +
Sbjct: 64 IPKKGSFKVI-------------------------------------------------I 74
Query: 189 RSALLYGTECWAVKSQHENQVSVAEMRMLRWMSGKTRHDRIRN 231
R +LY EC +V++ + Q+ VAEMRM RWM +TR D+I N
Sbjct: 75 RQTMLYEVECLSVQNSYVQQMKVAEMRMFRWMCRQTRKDKIGN 117
>gb|AAK14815.1| polyprotein [Schistosoma japonicum]
Length = 1091
Score = 83.6 bits (205), Expect = 5e-15
Identities = 62/246 (25%), Positives = 110/246 (44%), Gaps = 16/246 (6%)
Query: 4 YIRAIKDMYEGASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVLTEHIQ----- 58
YI I+ +Y + VR + + + G+ Q LSP++F ++D+L E
Sbjct: 737 YINLIQALYSNTTCRVRAYGRLSSELTTSSGVRQACPLSPFLFNFIIDILLELTLSSSDF 796
Query: 59 ---ELAPRCML----FADDVVLVGDSREEVNGRLETWRQALEAYGFCLSRSKTEYMECNF 111
+L P L +ADD+VL+ + +++ L T + G S SK + + ++
Sbjct: 797 PGVDLFPGDKLTDLEYADDIVLLSEDADKMQDFLTTLNMNVSMLGMRFSPSKCKMLLQDW 856
Query: 112 SGRRSKSTLEVKVGDHIIPQVTRFKYLESFVQNDGEIEADVSHRIQVGWLKWRRASCVLC 171
K + +G I V RF YL S + +G + ++S RI + +
Sbjct: 857 LNSAPK----LVIGRETIECVNRFTYLGSLISPNGLVSDEISARIHKARSAFANLRHLWR 912
Query: 172 DKKVPFKLKGKFYRTAVRSALLYGTECWAVKSQHENQVSVAEMRMLRWMSGKTRHDRIRN 231
+ + KG+ Y AVRS L YG E W ++ + ++ V + R LR ++ +R+ N
Sbjct: 913 RRDIRLMTKGRVYCAAVRSVLPYGCETWPLRVEDIRRILVFDHRCLRNIARVCWDNRVSN 972
Query: 232 DIIRER 237
+R R
Sbjct: 973 AWVRNR 978
>gb|AAM93546.1| polyprotein [Schistosoma japonicum]
Length = 976
Score = 83.6 bits (205), Expect = 5e-15
Identities = 62/246 (25%), Positives = 110/246 (44%), Gaps = 16/246 (6%)
Query: 4 YIRAIKDMYEGASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVLTEHIQ----- 58
YI I+ +Y + VR + + + G+ Q LSP++F ++D+L E
Sbjct: 622 YINLIQALYSNTTCRVRAYGRLSSELTTSSGVRQACPLSPFLFNFIIDILLELTLSSSDF 681
Query: 59 ---ELAPRCML----FADDVVLVGDSREEVNGRLETWRQALEAYGFCLSRSKTEYMECNF 111
+L P L +ADD+VL+ + +++ L T + G S SK + + ++
Sbjct: 682 PGVDLFPGDKLTDLEYADDIVLLSEDADKMQDFLTTLNMNVSMLGMRFSPSKCKMLLQDW 741
Query: 112 SGRRSKSTLEVKVGDHIIPQVTRFKYLESFVQNDGEIEADVSHRIQVGWLKWRRASCVLC 171
K + +G I V RF YL S + +G + ++S RI + +
Sbjct: 742 LNSAPK----LVIGRETIECVNRFTYLGSLISPNGLVSDEISARIHKARSAFANLRHLWR 797
Query: 172 DKKVPFKLKGKFYRTAVRSALLYGTECWAVKSQHENQVSVAEMRMLRWMSGKTRHDRIRN 231
+ + KG+ Y AVRS L YG E W ++ + ++ V + R LR ++ +R+ N
Sbjct: 798 RRDIRLMTKGRVYCAAVRSVLPYGCETWPLRVEDIRRILVFDHRCLRNIARVCWDNRVSN 857
Query: 232 DIIRER 237
+R R
Sbjct: 858 AWVRNR 863
>gb|AAC24982.2| reverse transcriptase [synthetic construct]
Length = 1016
Score = 83.6 bits (205), Expect = 5e-15
Identities = 61/246 (24%), Positives = 113/246 (45%), Gaps = 16/246 (6%)
Query: 4 YIRAIKDMYEGASTSVRTQDGTTEDFPITIGLHQGSTLSPYIF---------TLVLDVLT 54
+I +K +Y S VR + + F + G+ QG +SP++F T ++DV
Sbjct: 662 FINILKALYTNTSGRVRAYNHLSPLFHSSSGVRQGCPISPFLFNFAIDDILETALMDVSN 721
Query: 55 EHIQELAPRCML---FADDVVLVGDSREEVNGRLETWRQALEAYGFCLSRSKTEYMECNF 111
+ L +L +ADD+VL+ D+ + + L ++ YG C + SK + + ++
Sbjct: 722 GGVDMLPGERLLDLEYADDIVLLCDNAQGMQSALNQLAISVRRYGMCFAPSKCKVLLQDW 781
Query: 112 SGRRSKSTLEVKVGDHIIPQVTRFKYLESFVQNDGEIEADVSHRIQVGWLKWRRASCVLC 171
TL+ G+ I V +F YL S++ G + ++ RI + +
Sbjct: 782 QDSHPVLTLD---GEQI-EVVEKFVYLGSYISAGGGVSDEIDARIMKARAAYANLGHLWR 837
Query: 172 DKKVPFKLKGKFYRTAVRSALLYGTECWAVKSQHENQVSVAEMRMLRWMSGKTRHDRIRN 231
+ V +KG+ Y +VR+ LLY E W ++ + ++SV + R LR ++ + N
Sbjct: 838 LRDVSLAVKGRIYNASVRAVLLYACETWPLRVEDVRRLSVFDHRCLRRIADIQWQHHVSN 897
Query: 232 DIIRER 237
+R R
Sbjct: 898 AEVRHR 903
>gi|67625701 TPA: endonuclease-reverse transcriptase [Schistosoma mansoni]
Length = 992
Score = 75.5 bits (184), Expect = 1e-12
Identities = 59/256 (23%), Positives = 110/256 (42%), Gaps = 16/256 (6%)
Query: 5 IRAIKDMYEGASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVLTE-------HI 57
+ I++ Y+G + + T+ F + G+ QG LSP++F LV+D + + H
Sbjct: 659 VNIIRNSYDGLNCQIVHGGQLTDSFEVKTGVRQGCLLSPFLFLLVIDWIMKTSTSGGMHG 718
Query: 58 QELAPRCML----FADDVVLVGDSREEVNGRLETWRQALEAYGFCLSRSKTEYMECNFSG 113
+ R L FADD+ L+ +++++ + + A A G +++ K++ + N
Sbjct: 719 IQWTGRMQLDDLDFADDLALLSQTQQQMQEKTTSVAAASAAVGLNINKGKSKTLRYN--- 775
Query: 114 RRSKSTLEVKVGDHIIPQVTRFKYLESFVQNDGEIEADVSHRIQVGWLKWRRASCVLCDK 173
+ T + + + V F YL S + G +ADV RI + + + K
Sbjct: 776 --TICTNPITLDGEALEDVEIFTYLGSIIDEHGGSDADVRARIGKARAAYLQLKNIWSSK 833
Query: 174 KVPFKLKGKFYRTAVRSALLYGTECWAVKSQHENQVSVAEMRMLRWMSGKTRHDRIRNDI 233
++ K + + T V++ LLYG E W ++ V LR + D I N +
Sbjct: 834 QLSTNTKVRIFNTNVKTVLLYGAETWRTTKAIIQKIQVFINSCLRKILRIRWPDTISNKL 893
Query: 234 IRERERESGGSTYSRK 249
+ E + RK
Sbjct: 894 LWETTNQIPAEEEIRK 909
>gb|AAF36061.1| Hypothetical protein Y76B12C.5 [Caenorhabditis elegans]
gi|17544228|ref|NP_500154.1| predicted CDS, reverse
transcriptase family member (4C744) [Caenorhabditis
elegans]
Length = 938
Score = 73.6 bits (179), Expect = 5e-12
Identities = 64/205 (31%), Positives = 100/205 (48%), Gaps = 19/205 (9%)
Query: 6 RAIKDMYEGASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLD-VLTEHIQELAP-- 62
+ I+++ EG + D + + G+ QG + SP +F+ L +LT+ ELA
Sbjct: 691 KMIQELMEGGQAEITVHDKKLK-VNLCTGIRQGDSASPALFSAALQAILTDCDNELAGVG 749
Query: 63 --------RCMLFADDVVLVGDSREEVNGRLETWRQALEAYGFCLSRSKTEYMECNFSGR 114
R + FADDVVL+ + EEV RLE + YG +++SKT ++ F
Sbjct: 750 ISVEGRHIRRLEFADDVVLICSTPEEVQERLEILDRISSNYGLKINQSKTVLLKNKFC-- 807
Query: 115 RSKSTLEVKVGDHIIPQVTRFKYLESFVQNDGEIEADVSHRIQVGWLKWRRASCVLCDKK 174
RS+ L G IIP V +YL ++ G I+ ++S RI+ GW VL +
Sbjct: 808 RSQDIL--FNGSPIIP-VPGCRYLGRWIDISGSIDEEISRRIRAGWGALVGIKEVL--RI 862
Query: 175 VPFKLKGKFYRTAVRSALLYGTECW 199
+P K + ++ V ALLY +E W
Sbjct: 863 MPNKERIILFKQNVLPALLYASETW 887
>gb|AAB00700.1| Hypothetical protein C34D4.5 [Caenorhabditis elegans]
gi|17539028|ref|NP_501121.1| predicted CDS, reverse
transcriptase family member (4H911) [Caenorhabditis
elegans] gi|7497012|pir||T29286 hypothetical protein
C34D4.5 - Caenorhabditis elegans
Length = 624
Score = 72.0 bits (175), Expect = 1e-11
Identities = 63/205 (30%), Positives = 98/205 (47%), Gaps = 19/205 (9%)
Query: 6 RAIKDMYEGASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLD-VLTEHIQELAP-- 62
+ I++M EG + D + + G+ QG + SP +F+ L +LT+ E A
Sbjct: 291 KMIQEMMEGGQAEISVHDKKLK-VNLRTGVRQGDSASPALFSAALQAILTDCDNEFAGVG 349
Query: 63 --------RCMLFADDVVLVGDSREEVNGRLETWRQALEAYGFCLSRSKTEYMECNFSGR 114
R + FADDVVL+ + EEV RLE + YG +++SKT ++ F
Sbjct: 350 IKVEGRHIRRLEFADDVVLICSTPEEVQERLEILDRISSIYGLKINQSKTVLLKNKFC-- 407
Query: 115 RSKSTLEVKVGDHIIPQVTRFKYLESFVQNDGEIEADVSHRIQVGWLKWRRASCVLCDKK 174
RS+ G IIP V +YL ++ G I+ ++S RI+ GW VL +
Sbjct: 408 RSQDVF--FNGSPIIP-VPGCRYLGRWIDISGSIDEEISRRIRAGWGALVGIKEVL--RI 462
Query: 175 VPFKLKGKFYRTAVRSALLYGTECW 199
+P K + ++ V ALLY +E W
Sbjct: 463 MPNKERIILFKQNVLPALLYASETW 487
>gb|AAK18958.1| Hypothetical protein F56C9.2 [Caenorhabditis elegans]
gi|17553662|ref|NP_498615.1| predicted CDS, reverse
transcriptase family member (3I419) [Caenorhabditis
elegans] gi|7504398|pir||T16474 hypothetical protein
F56C9.2 - Caenorhabditis elegans
Length = 772
Score = 69.7 bits (169), Expect = 7e-11
Identities = 66/228 (28%), Positives = 107/228 (45%), Gaps = 26/228 (11%)
Query: 6 RAIKDMYEGASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLD-VLTEHIQELAP-- 62
+ I++M +G + D + + G+ QG + SP +F+ L +LT+ E A
Sbjct: 439 KMIQEMMDGGQAEITVHDKKLK-VNLCTGVRQGDSASPALFSAALQAILTDCDNEFAGVG 497
Query: 63 --------RCMLFADDVVLVGDSREEVNGRLETWRQALEAYGFCLSRSKTEYMECNFSGR 114
R + FADDVVL+ + EV RLE + YG +++SKT ++ F
Sbjct: 498 INVEGRHIRRLEFADDVVLICSTPGEVQERLEILDRISSNYGLKINQSKTVLLKNKFC-- 555
Query: 115 RSKSTLEVKVGDHIIPQVTRFKYLESFVQNDGEIEADVSHRIQVGWLKWRRASCVLCDKK 174
RS+ L G IIP V +YL ++ G I+ ++S RI+ GW VL +
Sbjct: 556 RSQDVL--FNGSPIIP-VPGCRYLGRWIDISGSIDEEISRRIRAGWGALVGIKEVL--RI 610
Query: 175 VPFKLKGKFYRTAVRSALLYGTECWAVKSQHENQVSVAEMRMLRWMSG 222
+P K + ++ V ALLY +E W + + +R+ R +SG
Sbjct: 611 MPNKERIILFKQNVLPALLYASETWTCNAG-------STLRLKRTVSG 651
>gb|AAF36059.1| Hypothetical protein Y75D11A.4 [Caenorhabditis elegans]
gi|17570529|ref|NP_508323.1| predicted CDS, reverse
transcriptase family member (XC378) [Caenorhabditis
elegans]
Length = 480
Score = 68.9 bits (167), Expect = 1e-10
Identities = 63/205 (30%), Positives = 97/205 (46%), Gaps = 19/205 (9%)
Query: 6 RAIKDMYEGASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLD-VLTEHIQELAP-- 62
+ I++M E + D + + G+ QG + SP +F+ L +LT+ ELA
Sbjct: 147 KMIQEMMERGQAEITVHDKKLK-VNLCTGVRQGDSASPALFSAALQAILTDCDNELAGVG 205
Query: 63 --------RCMLFADDVVLVGDSREEVNGRLETWRQALEAYGFCLSRSKTEYMECNFSGR 114
R + FADDVVL+ + EEV RLE + YG + +SKT ++ F
Sbjct: 206 ISVEGRHIRRLEFADDVVLICSTPEEVQERLEILDRISSYYGLKIDQSKTVLLKNKFC-- 263
Query: 115 RSKSTLEVKVGDHIIPQVTRFKYLESFVQNDGEIEADVSHRIQVGWLKWRRASCVLCDKK 174
RS+ L G IIP V +YL ++ G I+ ++S RI+ GW VL +
Sbjct: 264 RSQDVL--FNGSPIIP-VPGCRYLGRWIDISGSIDEEISRRIRAGWGALVGIKEVL--RI 318
Query: 175 VPFKLKGKFYRTAVRSALLYGTECW 199
+P K ++ V ALLY ++ W
Sbjct: 319 MPNKENIILFKQNVLPALLYASKTW 343
>ref|XP_556470.1| ENSANGP00000028171 [Anopheles gambiae str. PEST]
gi|55238617|gb|EAL39934.1| ENSANGP00000028171 [Anopheles
gambiae str. PEST]
Length = 777
Score = 64.7 bits (156), Expect = 2e-09
Identities = 54/230 (23%), Positives = 93/230 (39%), Gaps = 12/230 (5%)
Query: 5 IRAIKDMYEGASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLD--------VLTEH 56
IR ++ + VR + F T GL QG L+ +F L L+ T
Sbjct: 505 IRLVRMTMTNVTCQVRVDGKLSGPFATTKGLRQGDGLACLLFNLALERAIRDSRVETTGT 564
Query: 57 IQELAPRCMLFADDVVLVGDSREEVNGRLETWRQALEAYGFCLSRSKTEYMECNFS---- 112
I + + + +ADD+ ++G V + QA E G ++ +KT+ M +
Sbjct: 565 IFYKSTQILAYADDIDIIGLRLSYVAEAYQGIEQAAENLGLQINEAKTKLMVATSADLPI 624
Query: 113 GRRSKSTLEVKVGDHIIPQVTRFKYLESFVQNDGEIEADVSHRIQVGWLKWRRASCVLCD 172
+ +V++G+ V F YL S V ND +E ++ R+ +
Sbjct: 625 NNPNLRRRDVQIGERTFEVVPEFTYLGSKVSNDNSMEVELRARMLAANRSFYSLKKQFTS 684
Query: 173 KKVPFKLKGKFYRTAVRSALLYGTECWAVKSQHENQVSVAEMRMLRWMSG 222
K + + K Y T + L Y +E W + E ++ E +MLR + G
Sbjct: 685 KNLSRRTKLGLYSTYIVPVLTYASETWTLSKSDEALLAAFERKMLRRILG 734
>gb|AAC70880.1| Hypothetical protein F21E9.5 [Caenorhabditis elegans]
gi|17567237|ref|NP_508248.1| predicted CDS, reverse
transcriptase family member (XB968) [Caenorhabditis
elegans] gi|7499579|pir||T31973 hypothetical protein
F21E9.5 - Caenorhabditis elegans
Length = 864
Score = 63.2 bits (152), Expect = 7e-09
Identities = 55/251 (21%), Positives = 101/251 (39%), Gaps = 20/251 (7%)
Query: 4 YIRAIKDMYEGASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVL---------- 53
YI +++ Y ST+ T P+T G+ QG +SP +F+ L+ +
Sbjct: 492 YINLLQECYTDCSTTF-TPFHKNVTVPVTRGVRQGDPISPNLFSACLEHVFRQLNWKHFK 550
Query: 54 ------TEHIQELAPRC--MLFADDVVLVGDSREEVNGRLETWRQALEAYGFCLSRSKTE 105
TE I+ + FADD+VLV + + L + + G ++ KT+
Sbjct: 551 GDERYETEGIRVNGQNLTNLRFADDIVLVAHNPRTASQMLTELVEKCSSVGLKINTGKTK 610
Query: 106 YMECNFSGRRSKSTLEVKVGDHIIPQVTRFKYLESFVQNDGEIEADVSHRIQVGWLKWRR 165
+ F+ +R + II V + YL + + + ++ R + W +
Sbjct: 611 VLRNRFAYKRKVEIRCPNTTNIIIDDVNEYIYLGRQINDSNNLLPELHRRRRAAWAAFTN 670
Query: 166 ASCVLCDKKVPFKLKGKFYRTAVRSALLYGTECWAVKSQHENQVSVAEMRMLRWMSGKTR 225
P +L+ + + V AL YG+E W + +V V + R + G T
Sbjct: 671 IKSTTDQITCP-RLRANLFDSTVLPALTYGSEAWTFTKELAERVRVTHAALERKLVGLTL 729
Query: 226 HDRIRNDIIRE 236
++ + +I RE
Sbjct: 730 TEQRKRNIHRE 740
>gb|AAF60814.1| Hypothetical protein Y58G8A.2 [Caenorhabditis elegans]
gi|17566042|ref|NP_503166.1| predicted CDS, reverse
transcriptase family member (5A739) [Caenorhabditis
elegans]
Length = 769
Score = 63.2 bits (152), Expect = 7e-09
Identities = 56/255 (21%), Positives = 101/255 (38%), Gaps = 20/255 (7%)
Query: 4 YIRAIKDMYEGASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVL---------- 53
YI +++ Y ST+ T P+ G+ QG +SP +F+ L+ +
Sbjct: 439 YINLLQECYTDCSTTF-TPFHKNVTVPVIRGVRQGDPISPNLFSACLEHVFRQLNWKHFK 497
Query: 54 ------TEHIQELAPRC--MLFADDVVLVGDSREEVNGRLETWRQALEAYGFCLSRSKTE 105
TE I+ + FADD+VLV + + L + + G ++ KT+
Sbjct: 498 GDERYETEGIRVNGQNLTNLRFADDIVLVAHNPRTASQMLTELVEKCSSVGLKINTGKTK 557
Query: 106 YMECNFSGRRSKSTLEVKVGDHIIPQVTRFKYLESFVQNDGEIEADVSHRIQVGWLKWRR 165
+ F+ +R + II V + YL + + + ++ R + W +
Sbjct: 558 VLRNRFAYKRKVEIRCPNTTNIIIDDVNEYIYLGRQINDSNNLLPELHRRRRAAWAAFTN 617
Query: 166 ASCVLCDKKVPFKLKGKFYRTAVRSALLYGTECWAVKSQHENQVSVAEMRMLRWMSGKTR 225
P +L+ + + V AL YG+E W + +V V + R + G T
Sbjct: 618 IKSTTDQITCP-RLRANLFDSTVLPALTYGSEAWTFTKELAERVRVTHAALERKLVGLTL 676
Query: 226 HDRIRNDIIRERERE 240
++ +I RE RE
Sbjct: 677 TEQRERNIHREEVRE 691
>ref|XP_552671.1| ENSANGP00000005174 [Anopheles gambiae str. PEST]
gi|55235101|gb|EAL38938.1| ENSANGP00000005174 [Anopheles
gambiae str. PEST]
Length = 329
Score = 63.2 bits (152), Expect = 7e-09
Identities = 52/226 (23%), Positives = 92/226 (40%), Gaps = 12/226 (5%)
Query: 5 IRAIKDMYEGASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLD--------VLTEH 56
IR ++ + VR + F T GL QG L+ +F L L+ T
Sbjct: 8 IRLVRMTMTNVTCQVRVDGKLSGPFATTKGLRQGDGLACLLFNLALERAIRDSRVETTGT 67
Query: 57 IQELAPRCMLFADDVVLVGDSREEVNGRLETWRQALEAYGFCLSRSKTEYMECNFSG--- 113
I + + + +ADD+ ++ V + QA E+ G ++ +KT+ M +G
Sbjct: 68 IFYKSTQILAYADDIDIIDLRLSYVAEAYQGIEQAAESLGLQINEAKTKLMVATSAGLPI 127
Query: 114 -RRSKSTLEVKVGDHIIPQVTRFKYLESFVQNDGEIEADVSHRIQVGWLKWRRASCVLCD 172
++ +V++G+ V F L S V ND +EA++ R+ +
Sbjct: 128 NNQNLRRRDVQIGERTFEVVPEFTCLGSKVSNDNSMEAELRARMLAANRSFYSLKKQFTS 187
Query: 173 KKVPFKLKGKFYRTAVRSALLYGTECWAVKSQHENQVSVAEMRMLR 218
K + + K Y T + Y +E W + E ++ E +MLR
Sbjct: 188 KNLSRRTKLGLYSTYIVPVFTYASETWTLSKSDETLLAAFERKMLR 233
>ref|NP_493497.1| predicted CDS, reverse transcriptase family member (1O881)
[Caenorhabditis elegans]
Length = 835
Score = 62.8 bits (151), Expect = 9e-09
Identities = 60/269 (22%), Positives = 108/269 (39%), Gaps = 32/269 (11%)
Query: 4 YIRAIKDMYEGASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVL---------- 53
YI +++ Y ST+ T P+T G+ QG +SP +F+ L+ +
Sbjct: 558 YINLLQECYTDCSTTF-TPFHKNVTVPVTRGVRQGDPISPNLFSACLEHVFRQLNWKHFK 616
Query: 54 ------TEHIQELAPRC--MLFADDVVLVGDSREEVNGRLETWRQALEAYGFCLSRSKTE 105
TE I+ + FADD+VLV + V+ L + + G ++ KT+
Sbjct: 617 GDERYETEGIRVNGQNLTNLRFADDIVLVAHNPRTVSQMLTELVEKCSSVGLKINTGKTK 676
Query: 106 YMECNFSGRRSKSTLEVKVGDHIIPQVTRFKYLESFVQNDGEIEADVSHRIQVGWLKWRR 165
+ FS +R + II V + YL + + + A++ R + W +
Sbjct: 677 VLRNRFSYKRKVEIRCPNTTNIIIDDVNEYIYLGRQINDSNNLLAELHRRRRAAWAAFTN 736
Query: 166 ASCVLCDKKVPFKLKGKFYRTAVRSALLYGTECWAVKSQHENQVSVAEMRMLRWMSGKTR 225
+ P +L+ + + V AL YG+E W + +V V +
Sbjct: 737 IKSTMDQITCP-RLRANLFDSTVLPALTYGSEAWTFTKELAERVRVK----------VRK 785
Query: 226 HDRIRNDIIRERERESG--GSTYSRKVGR 252
++R+ +I ++R+ G G R GR
Sbjct: 786 KSKLRDPLIHIKKRKLGWAGHVARRTDGR 814
>emb|CAA36227.1| unnamed protein product [Drosophila melanogaster]
gi|140023|sp|P16425|Y2R2_DROME Hypothetical 115 kDa
protein in type I retrotransposable element R1DM (ORF 2)
Length = 1021
Score = 58.5 bits (140), Expect = 2e-07
Identities = 49/141 (34%), Positives = 73/141 (51%), Gaps = 21/141 (14%)
Query: 9 KDMYEGASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVLTEHIQELAPRCML-- 66
+ + G +R+ GT E P+T G QGS P+I+ +++DVL +Q L P C L
Sbjct: 607 QSFFSGRRAVIRSSSGTVE-VPVTRGCPQGSISGPFIWDILMDVL---LQRLQPYCQLSA 662
Query: 67 FADDVVLV--GDSR---EEVNGRL----ETWRQALEAYGFCLSRSKTEYMECNFSGRRSK 117
+ADD++L+ G+SR EE +L ETW + G CLS SKT M + RR+
Sbjct: 663 YADDLLLLVEGNSRAVLEEKGAQLMSIVETWGAEV---GDCLSTSKTVIMLLKGALRRAP 719
Query: 118 STLEVKVGDHIIPQVTRFKYL 138
+ V+ +P V +YL
Sbjct: 720 T---VRFAGRNLPYVRSCRYL 737
>pir||S09111 hypothetical protein 2 - fruit fly (Drosophila melanogaster)
transposable element R1
Length = 1021
Score = 57.8 bits (138), Expect = 3e-07
Identities = 49/141 (34%), Positives = 73/141 (51%), Gaps = 21/141 (14%)
Query: 9 KDMYEGASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVLTEHIQELAPRCML-- 66
+ + G +R+ GT E P+T G QGS P+I+ +++DVL +Q L P C L
Sbjct: 607 QSFFSGRRAVIRSSSGTVE-VPVTRGCPQGSISGPFIWDILMDVL---LQRLQPYCQLSA 662
Query: 67 FADDVVLV--GDSR---EEVNGRL----ETWRQALEAYGFCLSRSKTEYMECNFSGRRSK 117
+ADD++L+ G+SR EE +L ETW + G CLS SKT M + RR+
Sbjct: 663 YADDLLLLVEGNSRAVLEEKGAQLMSIVETWGAEV---GDCLSTSKTVIMLLKGALRRAP 719
Query: 118 STLEVKVGDHIIPQVTRFKYL 138
+ V+ +P V +YL
Sbjct: 720 T---VRFAGANLPYVRSCRYL 737
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.321 0.135 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 410,904,199
Number of Sequences: 2540612
Number of extensions: 15701319
Number of successful extensions: 34038
Number of sequences better than 10.0: 567
Number of HSP's better than 10.0 without gapping: 271
Number of HSP's successfully gapped in prelim test: 296
Number of HSP's that attempted gapping in prelim test: 33612
Number of HSP's gapped (non-prelim): 607
length of query: 253
length of database: 863,360,394
effective HSP length: 125
effective length of query: 128
effective length of database: 545,783,894
effective search space: 69860338432
effective search space used: 69860338432
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 73 (32.7 bits)
Medicago: description of AC126792.7


