

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126784.6 - phase: 0
(135 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
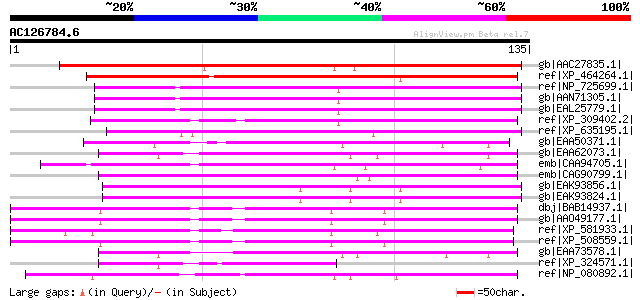
Score E
Sequences producing significant alignments: (bits) Value
gb|AAC27835.1| hypothetical protein [Arabidopsis thaliana] gi|74... 133 1e-30
ref|XP_464264.1| unknown protein [Oryza sativa (japonica cultiva... 108 3e-23
ref|NP_725699.1| CG30105-PA [Drosophila melanogaster] gi|2162705... 64 8e-10
gb|AAN71305.1| RE11282p [Drosophila melanogaster] 64 8e-10
gb|EAL25779.1| GA15653-PA [Drosophila pseudoobscura] 64 1e-09
ref|XP_309402.2| ENSANGP00000015645 [Anopheles gambiae str. PEST... 63 1e-09
ref|XP_635195.1| hypothetical protein DDB0183878 [Dictyostelium ... 59 3e-08
gb|EAA50371.1| hypothetical protein MG04130.4 [Magnaporthe grise... 58 6e-08
gb|EAA62073.1| hypothetical protein AN7493.2 [Aspergillus nidula... 57 1e-07
emb|CAA94705.1| SPAC12B10.15c [Schizosaccharomyces pombe] gi|191... 56 2e-07
emb|CAG90799.1| unnamed protein product [Debaryomyces hansenii C... 55 5e-07
gb|EAK93856.1| hypothetical protein CaO19.1309 [Candida albicans... 54 6e-07
gb|EAK93824.1| hypothetical protein CaO19.8889 [Candida albicans... 54 6e-07
dbj|BAB14937.1| unnamed protein product [Homo sapiens] 51 7e-06
gb|AAO49177.1| AYP1 [Homo sapiens] gi|28628363|gb|AAO49176.1| AY... 50 9e-06
ref|XP_581933.1| PREDICTED: similar to AYP1 protein [Bos taurus] 50 1e-05
ref|XP_508559.1| PREDICTED: similar to AYP1 protein; RNase H1 sm... 49 2e-05
gb|EAA73578.1| hypothetical protein FG04252.1 [Gibberella zeae P... 49 3e-05
ref|XP_324571.1| predicted protein [Neurospora crassa] gi|289238... 47 8e-05
ref|NP_080892.1| AYP1 [Mus musculus] gi|28628361|gb|AAO49175.1| ... 47 1e-04
>gb|AAC27835.1| hypothetical protein [Arabidopsis thaliana] gi|7485405|pir||T00554
hypothetical protein At2g39440 [imported] - Arabidopsis
thaliana gi|15225467|ref|NP_181476.1| expressed protein
[Arabidopsis thaliana]
Length = 773
Score = 133 bits (334), Expect = 1e-30
Identities = 67/142 (47%), Positives = 87/142 (61%), Gaps = 22/142 (15%)
Query: 14 DGIEDLSGRVHLLPCCIKHDGPTEVSQYFKPKPTGV----------VGEDGLPLQQSHFR 63
+ I DLS +VH LPCCI+ DGP EVS YFKPK + V DG+ +++HFR
Sbjct: 83 NSIVDLSSQVHQLPCCIRFDGPVEVSHYFKPKSSVRLMWKAVEFVDVEIDGVKTEEAHFR 142
Query: 64 GRLLEGTTLPLPHNYSGFVL---------GKKNS---VESSNSWETSVTFNDITYWNHDS 111
GR L+G T+ LP YSGFVL GK+ + E + WE FN++TYWNHDS
Sbjct: 143 GRKLQGATISLPSGYSGFVLRQASNLNANGKRKASMPTEDNQCWEVKAKFNNLTYWNHDS 202
Query: 112 VPSNNDDFSRAFHWIPVAEAVS 133
+PS +D F R+FHW +AEAV+
Sbjct: 203 LPSKDDTFFRSFHWFSIAEAVT 224
>ref|XP_464264.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|49388604|dbj|BAD25719.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 161
Score = 108 bits (270), Expect = 3e-23
Identities = 56/119 (47%), Positives = 73/119 (61%), Gaps = 8/119 (6%)
Query: 21 GRVHLLPCCIKHDGPTEVSQYFKPKPTGVVGEDGLPLQQSHFRGRLLEGTTLPLPHNYSG 80
GRVHLLPC IK +G VS YFKPK TGV E G+ ++++ FRGR L+G T+ LP Y G
Sbjct: 24 GRVHLLPCGIKQNGAAAVSDYFKPKDTGVEVE-GIRVEEAFFRGRKLQGATISLPDGYRG 82
Query: 81 FVLGKKNSVESSNSWETSVT-------FNDITYWNHDSVPSNNDDFSRAFHWIPVAEAV 132
+VL K++ + E V+ F +ITYWNHD+ PS D R FH + VA A+
Sbjct: 83 YVLEKRSGGKDMKKLEGEVSNFKSRAEFQNITYWNHDTTPSAEDPLPRCFHLLTVANAM 141
>ref|NP_725699.1| CG30105-PA [Drosophila melanogaster] gi|21627059|gb|AAM68481.1|
CG30105-PA [Drosophila melanogaster]
Length = 155
Score = 63.9 bits (154), Expect = 8e-10
Identities = 43/114 (37%), Positives = 52/114 (44%), Gaps = 4/114 (3%)
Query: 23 VHLLPCCIKHDGPTEVSQYFKPKPTGVVGEDGLPLQQSHFRGRLLEGTTLPLPHNYSGFV 82
VH LP I DG V YF T E G + + RG L G L +P Y G V
Sbjct: 20 VHYLPAKIDGDGEANVKNYFN-NYTREATEFGTGILTNALRGFPLMGEKLKVPEGYCGLV 78
Query: 83 LG---KKNSVESSNSWETSVTFNDITYWNHDSVPSNNDDFSRAFHWIPVAEAVS 133
L K S S + F D TYWN+D VPSN D + +A VA+A+S
Sbjct: 79 LQETEKPISDTSDRKLRLTGVFQDFTYWNYDKVPSNGDPYRQALPLPDVAQALS 132
>gb|AAN71305.1| RE11282p [Drosophila melanogaster]
Length = 165
Score = 63.9 bits (154), Expect = 8e-10
Identities = 43/114 (37%), Positives = 52/114 (44%), Gaps = 4/114 (3%)
Query: 23 VHLLPCCIKHDGPTEVSQYFKPKPTGVVGEDGLPLQQSHFRGRLLEGTTLPLPHNYSGFV 82
VH LP I DG V YF T E G + + RG L G L +P Y G V
Sbjct: 30 VHYLPAKIDGDGEANVKNYFN-NYTREATEFGTGILTNALRGFPLMGEKLKVPEGYCGLV 88
Query: 83 LG---KKNSVESSNSWETSVTFNDITYWNHDSVPSNNDDFSRAFHWIPVAEAVS 133
L K S S + F D TYWN+D VPSN D + +A VA+A+S
Sbjct: 89 LQETEKPISDTSDRKLRLTGVFQDFTYWNYDKVPSNGDPYRQALPLPDVAQALS 142
>gb|EAL25779.1| GA15653-PA [Drosophila pseudoobscura]
Length = 154
Score = 63.5 bits (153), Expect = 1e-09
Identities = 40/114 (35%), Positives = 53/114 (46%), Gaps = 4/114 (3%)
Query: 23 VHLLPCCIKHDGPTEVSQYFKPKPTGVVGEDGLPLQQSHFRGRLLEGTTLPLPHNYSGFV 82
+H LP I DG V YF T E G + + RG L G + +P Y G V
Sbjct: 20 LHYLPAKISGDGEANVETYFN-NYTREAPEYGSGMMNNALRGYPLVGERMKVPEGYKGLV 78
Query: 83 LG---KKNSVESSNSWETSVTFNDITYWNHDSVPSNNDDFSRAFHWIPVAEAVS 133
L K S + + F++ TYWN+D VPSN D F +A VA+A+S
Sbjct: 79 LQETEKPLSESTDRQLRLTGVFDEFTYWNYDKVPSNGDPFRQAMLMADVAKALS 132
>ref|XP_309402.2| ENSANGP00000015645 [Anopheles gambiae str. PEST]
gi|55244718|gb|EAA05041.2| ENSANGP00000015645 [Anopheles
gambiae str. PEST]
Length = 129
Score = 63.2 bits (152), Expect = 1e-09
Identities = 38/114 (33%), Positives = 53/114 (46%), Gaps = 7/114 (6%)
Query: 22 RVHLLPCCIKHDGPTEVSQYFKPKPTGVVGEDGLPLQQSHFRGRLLEGTTLPLPHNYSGF 81
++ +P IK DGP + Q+F P DG RG L G LP Y+G
Sbjct: 3 KLQYIPATIKGDGPANLEQFFTPYTE--TQRDGTLCNA--LRGYPLRGKATSLPAGYTGV 58
Query: 82 VLG---KKNSVESSNSWETSVTFNDITYWNHDSVPSNNDDFSRAFHWIPVAEAV 132
+L K S E + + F D TYWN+D PS ND ++A W+ +AE +
Sbjct: 59 MLQETKKPLSDEDDRTLTFAGAFRDFTYWNYDRQPSRNDPMAKALGWLQLAEVL 112
>ref|XP_635195.1| hypothetical protein DDB0183878 [Dictyostelium discoideum]
gi|60463507|gb|EAL61692.1| hypothetical protein
DDB0183878 [Dictyostelium discoideum]
Length = 233
Score = 58.9 bits (141), Expect = 3e-08
Identities = 37/119 (31%), Positives = 60/119 (50%), Gaps = 11/119 (9%)
Query: 26 LPCCIKHDGPTEVSQYFK--PKP--------TGVVGEDGLPLQQSHFRGRLLEGTTLPLP 75
LP I ++G +VS YF+ KP T D S FRG L G + +P
Sbjct: 34 LPFSIGYNGFAKVSNYFQVTDKPATTTTTTTTTTPSPDDKKYLYSSFRGIQLIGEKIKIP 93
Query: 76 HNYSGFVLGKKNSVESSN-SWETSVTFNDITYWNHDSVPSNNDDFSRAFHWIPVAEAVS 133
+ + G+V ++ +++N WE FN++TYWN ++VPS+ D +AF + + V+
Sbjct: 94 NGFDGYVFRDEDEQDNNNRKWEPISKFNELTYWNRETVPSDFDKQIQAFKSLNIQSMVN 152
>gb|EAA50371.1| hypothetical protein MG04130.4 [Magnaporthe grisea 70-15]
gi|39944238|ref|XP_361656.1| hypothetical protein
MG04130.4 [Magnaporthe grisea 70-15]
Length = 331
Score = 57.8 bits (138), Expect = 6e-08
Identities = 39/134 (29%), Positives = 61/134 (45%), Gaps = 29/134 (21%)
Query: 20 SGRVHLLPCCIKHDGPT-EVSQYFKPKPTGVVGEDGLPLQQSHFRGRLLEGTTLPLPHNY 78
S VHLLPC I HDGP +V Y+KP + +DG + + FRGR L G + +P Y
Sbjct: 17 SAAVHLLPCTIHHDGPVDQVQSYWKPTES----QDG--KKTAFFRGRKLHGKAIRVPEGY 70
Query: 79 SGFVLGK--------------------KNSVESSNSWETSVTFNDITYWNHDS-VPSNND 117
G V+ K ++ + +T F ++ W H+S +++D
Sbjct: 71 KGIVVEKGPEDGPAETRADEPVTVDVDAEEKVATTALDTKAEFEEMVIWGHESTADASSD 130
Query: 118 DFSRAF-HWIPVAE 130
+ R W+ +AE
Sbjct: 131 PYVRGVEEWVSLAE 144
>gb|EAA62073.1| hypothetical protein AN7493.2 [Aspergillus nidulans FGSC A4]
gi|67901012|ref|XP_680762.1| hypothetical protein
AN7493_2 [Aspergillus nidulans FGSC A4]
gi|49108701|ref|XP_411630.1| hypothetical protein
AN7493.2 [Aspergillus nidulans FGSC A4]
Length = 159
Score = 56.6 bits (135), Expect = 1e-07
Identities = 38/128 (29%), Positives = 57/128 (43%), Gaps = 23/128 (17%)
Query: 24 HLLPCCIKHDGPTE-VSQYFKPKPTGVVGEDGLPLQQSHFRGRLLEGTTLPLPHNYSGFV 82
++LPC I +DG E +++Y+ P E LQ ++FRGR L G + LP Y G V
Sbjct: 25 NILPCRIHYDGSVESLNRYWTPS----TDEKDKDLQTAYFRGRRLRGRRVQLPEGYEGVV 80
Query: 83 LGKKN------------SVESSNS-----WETSVTFNDITYWNHDSVPSNNDDFSRAF-H 124
+ S E N+ E TF D W H+ +P+ +D F +
Sbjct: 81 VTHTEREIPNATDNVTVSKEGENNKAVKLLEKQATFGDYVVWGHEVIPAADDSFVKGVEE 140
Query: 125 WIPVAEAV 132
W+ AE +
Sbjct: 141 WVKFAEVM 148
>emb|CAA94705.1| SPAC12B10.15c [Schizosaccharomyces pombe]
gi|19115559|ref|NP_594647.1| hypothetical protein
SPAC12B10.15c [Schizosaccharomyces pombe 972h-]
gi|7490784|pir||T37582 hypothetical protein
SPAC12B10.15c - fission yeast (Schizosaccharomyces
pombe) gi|1723557|sp|Q10448|YDEF_SCHPO Hypothetical
protein C12B10.15c in chromosome I
Length = 147
Score = 55.8 bits (133), Expect = 2e-07
Identities = 42/135 (31%), Positives = 62/135 (45%), Gaps = 14/135 (10%)
Query: 9 IDLKRDGIEDLSGRVHLLPCCIKHDGPTEVSQYFKPKPTGVVGEDGLPLQQSHFRGRLLE 68
+ L++ G +L + LLPC I +DGP V +YF K + + RGR L
Sbjct: 8 VKLEQAGSSNLE-KASLLPCHISYDGPAPVFEYFHDKIQ--ISNNTHSKTTVCLRGRELN 64
Query: 69 GTTLPLPHNYSGFVL----GKKNSVES-----SNSWETSVTFNDITYWNHDSVPSNNDDF 119
G L LP NY+G V+ G +N S ++W + TF + WN D++ S+ D
Sbjct: 65 GEELDLPENYTGQVVLCDDGLENDETSEKEIPESTWCINSTFEKVMLWNRDNMRSSEKDQ 124
Query: 120 SR--AFHWIPVAEAV 132
+ WI A V
Sbjct: 125 WKRGVLEWIQFASKV 139
>emb|CAG90799.1| unnamed protein product [Debaryomyces hansenii CBS767]
gi|50427361|ref|XP_462293.1| unnamed protein product
[Debaryomyces hansenii]
Length = 138
Score = 54.7 bits (130), Expect = 5e-07
Identities = 29/120 (24%), Positives = 58/120 (48%), Gaps = 11/120 (9%)
Query: 24 HLLPCCIKHDGPTEVSQYFKPKPTGVVGEDGLPLQQSHFRGRLLEGTTLPLPHNYSGFVL 83
+++PC I+++G YF P G ++ +++RG L G + + ++YSG+++
Sbjct: 17 NIVPCHIQYNGAANTKDYFTPSKKQDTLPTGEQVEVAYYRGCKLVGKNMNISNDYSGYLI 76
Query: 84 GKKNSV-----ESS------NSWETSVTFNDITYWNHDSVPSNNDDFSRAFHWIPVAEAV 132
K S+ E+S N++ F +IT + HD+VP + W ++E +
Sbjct: 77 NKSESLARVEDETSEDYLTVNTYTPVAKFEEITVYGHDTVPELTSQWGLIQEWKDISEVI 136
>gb|EAK93856.1| hypothetical protein CaO19.1309 [Candida albicans SC5314]
Length = 137
Score = 54.3 bits (129), Expect = 6e-07
Identities = 34/119 (28%), Positives = 59/119 (49%), Gaps = 10/119 (8%)
Query: 25 LLPCCIKHDGPTEVSQYFKPKPTGVVGEDGLPLQQSHFRGRLLEGTTLPL-PHNYSGFVL 83
++P I++ GP YF P T DG L ++FRG L G TL L ++ G+V+
Sbjct: 18 VVPMNIQYSGPANTEDYFAPSKTQETQPDGTLLDVAYFRGCKLVGKTLELDKYDVKGYVI 77
Query: 84 GKKN---SVESSNSWETSVT------FNDITYWNHDSVPSNNDDFSRAFHWIPVAEAVS 133
K S + + +T+VT F +T + HD++PS + +S ++ ++ V+
Sbjct: 78 NKSEHLVSDKETGEVKTAVTYIPVGNFQKLTVYGHDTLPSRTNQWSLIPEYLSISNIVN 136
>gb|EAK93824.1| hypothetical protein CaO19.8889 [Candida albicans SC5314]
Length = 137
Score = 54.3 bits (129), Expect = 6e-07
Identities = 34/119 (28%), Positives = 59/119 (49%), Gaps = 10/119 (8%)
Query: 25 LLPCCIKHDGPTEVSQYFKPKPTGVVGEDGLPLQQSHFRGRLLEGTTLPL-PHNYSGFVL 83
++P I++ GP YF P T DG L ++FRG L G TL L ++ G+V+
Sbjct: 18 VVPMNIQYSGPANTEDYFAPSKTQETQPDGTLLDVAYFRGCKLVGKTLELDKYDVKGYVI 77
Query: 84 GKKN---SVESSNSWETSVT------FNDITYWNHDSVPSNNDDFSRAFHWIPVAEAVS 133
K S + + +T+VT F +T + HD++PS + +S ++ ++ V+
Sbjct: 78 NKSEHLVSDKETGEVKTAVTYIPVGNFQKLTVYGHDTLPSRTNQWSLIPEYLSISNIVN 136
>dbj|BAB14937.1| unnamed protein product [Homo sapiens]
Length = 163
Score = 50.8 bits (120), Expect = 7e-06
Identities = 42/161 (26%), Positives = 67/161 (41%), Gaps = 34/161 (21%)
Query: 1 MEKGGEAKIDLKRDGIEDLSGR------VHLLPCCIKHDGPTEVSQYFKPKPTGVVGEDG 54
ME G EA I+ R + + R +HLLPC + DGP V ++F P G +G
Sbjct: 1 MESGDEAAIERHRVHLRSATLRDAVPATLHLLPCEVAVDGPAPVGRFFTPAIR--QGPEG 58
Query: 55 LPLQQSHFRGRLLEGTTLPLPHNYSGFV--------LGKKNSVESSNSWE---------- 96
L + FRGR L G + +P G+V +GK + + S + +
Sbjct: 59 LEVS---FRGRCLRGEEVAVPPGLVGYVMVTEEKVSMGKPDPLRDSGTDDQEEEPLERDF 115
Query: 97 -----TSVTFNDITYWNHDSVPSNNDDFSRAFHWIPVAEAV 132
+ F+ T W +++P + A W +A A+
Sbjct: 116 DRFIGATANFSRFTLWGLETIPGPDAKVRGALTWPSLAAAI 156
>gb|AAO49177.1| AYP1 [Homo sapiens] gi|28628363|gb|AAO49176.1| AYP1 [Homo sapiens]
gi|19421812|gb|AAL87739.1| RNase H1 small subunit [Homo
sapiens] gi|23270834|gb|AAH23588.1| AYP1 protein [Homo
sapiens] gi|38176285|ref|NP_115569.2| AYP1 protein [Homo
sapiens]
Length = 164
Score = 50.4 bits (119), Expect = 9e-06
Identities = 42/162 (25%), Positives = 67/162 (40%), Gaps = 35/162 (21%)
Query: 1 MEKGGEAKIDLKRDGIEDLSGR------VHLLPCCIKHDGPTEVSQYFKPKPTGVVGEDG 54
ME G EA I+ R + + R +HLLPC + DGP V ++F P G +G
Sbjct: 1 MESGDEAAIERHRVHLRSATLRDAVPATLHLLPCEVAVDGPAPVGRFFTPAIR--QGPEG 58
Query: 55 LPLQQSHFRGRLLEGTTLPLPHNYSGFV---------LGKKNSVESSNSWE--------- 96
L + FRGR L G + +P G+V +GK + + S + +
Sbjct: 59 LEVS---FRGRCLRGEEVAVPPGLVGYVMVTEEKKVSMGKPDPLRDSGTDDQEEEPLERD 115
Query: 97 ------TSVTFNDITYWNHDSVPSNNDDFSRAFHWIPVAEAV 132
+ F+ T W +++P + A W +A A+
Sbjct: 116 FDRFIGATANFSRFTLWGLETIPGPDAKVRGALTWPSLAAAI 157
>ref|XP_581933.1| PREDICTED: similar to AYP1 protein [Bos taurus]
Length = 160
Score = 50.1 bits (118), Expect = 1e-05
Identities = 43/162 (26%), Positives = 68/162 (41%), Gaps = 36/162 (22%)
Query: 1 MEKGGEAKIDLKR-----DGIEDLS-GRVHLLPCCIKHDGPTEVSQYFKPKPTGVVGEDG 54
ME E ID +R D + D + +HLLPC + + PT V ++F P +G DG
Sbjct: 1 MENSYEETIDKRRVHLRPDTLRDPAPASLHLLPCEVPVNRPTPVGRFFTPAIR--MGRDG 58
Query: 55 LPLQQSHFRGRLLEGTTLPLPHNYSGFV---------LGKKNSVESSNS----------- 94
L ++ FRGR L G + +P + G+V +GK++ E
Sbjct: 59 L---EASFRGRSLRGEEVVVPPGFVGYVVTEEKAEVLMGKQDDHERQEQELLEPPEALER 115
Query: 95 -----WETSVTFNDITYWNHDSVPSNNDDFSRAFHWIPVAEA 131
+ +F+ T W +S+P + A W +A A
Sbjct: 116 DCDRFMGATASFSSFTVWGLESIPGPDAKLRGALSWPSLAAA 157
>ref|XP_508559.1| PREDICTED: similar to AYP1 protein; RNase H1 small subunit [Pan
troglodytes]
Length = 246
Score = 49.3 bits (116), Expect = 2e-05
Identities = 42/161 (26%), Positives = 66/161 (40%), Gaps = 35/161 (21%)
Query: 1 MEKGGEAKIDLKRDGIEDLSGR------VHLLPCCIKHDGPTEVSQYFKPKPTGVVGEDG 54
ME G EA I+ R + + R +HLLPC + DGP V ++F P G +G
Sbjct: 68 MESGDEAAIERHRVHLRSATLRDAVPATLHLLPCEVAVDGPAPVGRFFTPAIR--QGPEG 125
Query: 55 LPLQQSHFRGRLLEGTTLPLPHNYSGFV---------LGKKNSVESSNSWE--------- 96
L + FRGR L G + +P G+V +GK + + S + +
Sbjct: 126 LEVS---FRGRCLRGEEVAVPPGLVGYVMVTEEKEVSMGKPDPLRDSGTDDQEEEPLERD 182
Query: 97 ------TSVTFNDITYWNHDSVPSNNDDFSRAFHWIPVAEA 131
+ F+ T W +++P + A W +A A
Sbjct: 183 FDRFIGATANFSRFTLWGLETIPGPDAKVRGALTWPSLAAA 223
>gb|EAA73578.1| hypothetical protein FG04252.1 [Gibberella zeae PH-1]
gi|46116820|ref|XP_384428.1| hypothetical protein
FG04252.1 [Gibberella zeae PH-1]
Length = 149
Score = 48.9 bits (115), Expect = 3e-05
Identities = 35/129 (27%), Positives = 61/129 (47%), Gaps = 31/129 (24%)
Query: 24 HLLPCCIKHDGPTE-VSQYFKPKPTGVVGEDGLPLQQSHFRGRLLEGTTLPLPHNYSGFV 82
++LPC I HDGP E S Y+ P T ++FRGR L+G + LP + G V
Sbjct: 20 NILPCRIHHDGPIEPTSTYWTPAAT-----------DAYFRGRKLQGKIVKLPKDCRGIV 68
Query: 83 LGK------KNSV-----------ESSNSWETSVTFNDITYWNHDSV-PSNNDDFSRAF- 123
+ + K +V E + + + F+++ W H++V ++ D + R+
Sbjct: 69 VERVSQQDPKTAVEEIVEDEDAEQEQTGRMQVTAEFDEMVVWGHEAVADASADPYVRSME 128
Query: 124 HWIPVAEAV 132
W+ VA+ +
Sbjct: 129 EWLQVADKI 137
>ref|XP_324571.1| predicted protein [Neurospora crassa] gi|28923812|gb|EAA32977.1|
predicted protein [Neurospora crassa]
Length = 169
Score = 47.4 bits (111), Expect = 8e-05
Identities = 28/63 (44%), Positives = 35/63 (55%), Gaps = 6/63 (9%)
Query: 24 HLLPCCIKHDGPTE-VSQYFKPKPTGVVGEDGLPLQQSHFRGRLLEGTTLPLPHNYSGFV 82
+LLPC I H G E ++ P+ +DG L+ S+FRGR L G TLPLP Y G V
Sbjct: 26 NLLPCRIHHSGSVEPTDSFWNPR----CEQDG-NLKVSYFRGRKLHGKTLPLPEGYKGVV 80
Query: 83 LGK 85
K
Sbjct: 81 ASK 83
>ref|NP_080892.1| AYP1 [Mus musculus] gi|28628361|gb|AAO49175.1| AYP1 [Mus musculus]
gi|12858665|dbj|BAB31401.1| unnamed protein product [Mus
musculus] gi|12847755|dbj|BAB27694.1| unnamed protein
product [Mus musculus]
Length = 166
Score = 47.0 bits (110), Expect = 1e-04
Identities = 42/155 (27%), Positives = 66/155 (42%), Gaps = 32/155 (20%)
Query: 5 GEAKIDLKRDGIEDLS-GRVHLLPCCIKHDGPTEVSQYFKPKPTGVVGEDGLPLQQSHFR 63
G+ +I L+ + + ++HLLPC + P V ++F P V D LQ S FR
Sbjct: 10 GKQRIHLRPGSLRGAAPAKLHLLPCDVLVSRPAPVDRFFTP----AVRHDADGLQAS-FR 64
Query: 64 GRLLEGTTLPLPHNYSGFVL----------GKKN--------SVESSNSWETSV------ 99
GR L G + +P ++GFV+ GK N + E+ E
Sbjct: 65 GRGLRGEEVAVPPGFAGFVMVTEEKGEGLIGKLNFSGDAEDKADEAQEPLERDFDRLIGA 124
Query: 100 --TFNDITYWNHDSVPSNNDDFSRAFHWIPVAEAV 132
+F+ T W ++VP + RA W +A A+
Sbjct: 125 TGSFSHFTLWGLETVPGPDAKVHRALGWPSLAAAI 159
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.316 0.136 0.429
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 270,156,728
Number of Sequences: 2540612
Number of extensions: 11629616
Number of successful extensions: 22149
Number of sequences better than 10.0: 77
Number of HSP's better than 10.0 without gapping: 25
Number of HSP's successfully gapped in prelim test: 57
Number of HSP's that attempted gapping in prelim test: 22084
Number of HSP's gapped (non-prelim): 89
length of query: 135
length of database: 863,360,394
effective HSP length: 111
effective length of query: 24
effective length of database: 581,352,462
effective search space: 13952459088
effective search space used: 13952459088
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.5 bits)
S2: 67 (30.4 bits)
Medicago: description of AC126784.6


