

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126015.11 + phase: 0
(371 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
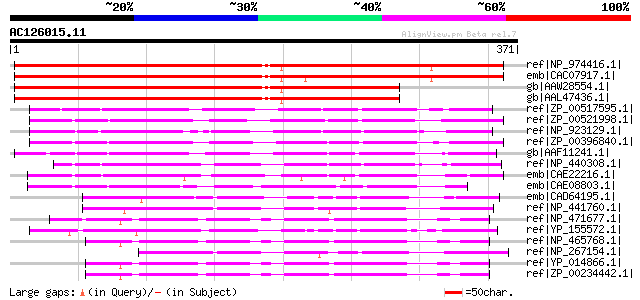
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_974416.1| pseudouridine synthase family protein [Arabidop... 487 e-136
emb|CAC07917.1| putative protein [Arabidopsis thaliana] gi|11358... 480 e-134
gb|AAW28554.1| At3g52260 [Arabidopsis thaliana] gi|22331748|ref|... 390 e-107
gb|AAL47436.1| AT3g52260/T25B15_30 [Arabidopsis thaliana] 387 e-106
ref|ZP_00517595.1| Pseudouridine synthase, RluD [Crocosphaera wa... 248 2e-64
ref|ZP_00521998.1| Pseudouridine synthase, RluD [Solibacter usit... 215 2e-54
ref|NP_923129.1| probable pseudouridine synthase [Gloeobacter vi... 212 2e-53
ref|ZP_00396840.1| Pseudouridine synthase, RluD [Deinococcus geo... 204 3e-51
gb|AAF11241.1| pseudouridine synthase, putative [Deinococcus rad... 201 3e-50
ref|NP_440308.1| hypothetical protein slr1592 [Synechocystis sp.... 194 3e-48
emb|CAE22216.1| Pseudouridine synthase [Prochlorococcus marinus ... 187 3e-46
emb|CAE08803.1| probable pseudouridine synthase [Synechococcus s... 137 4e-31
emb|CAD64195.1| pseudouridylate synthase [Lactobacillus plantaru... 128 3e-28
ref|NP_441760.1| hypothetical protein slr1629 [Synechocystis sp.... 126 9e-28
ref|NP_471677.1| hypothetical protein lin2346 [Listeria innocua ... 125 2e-27
ref|YP_155572.1| Pseudouridylate synthase [Idiomarina loihiensis... 124 6e-27
ref|NP_465768.1| hypothetical protein lmo2244 [Listeria monocyto... 124 6e-27
ref|NP_267154.1| pseudouridine synthase [Lactococcus lactis subs... 124 6e-27
ref|YP_014866.1| ribosomal large subunit pseudouridine synthase,... 124 6e-27
ref|ZP_00234442.1| ribosomal large subunit pseudouridine synthas... 124 6e-27
>ref|NP_974416.1| pseudouridine synthase family protein [Arabidopsis thaliana]
Length = 369
Score = 487 bits (1254), Expect = e-136
Identities = 240/366 (65%), Positives = 300/366 (81%), Gaps = 10/366 (2%)
Query: 4 PKKIGIPWPEFNDGLSYDDVVRPSDAGLTLI-EFYSTKYKSSAPLQGWLQRIKSDQITLD 62
P++IG+PWPE NDGL+Y D + S++ LT + EFY TKYKSSAPL GW+QRI++ QI +D
Sbjct: 5 PQRIGLPWPELNDGLAYKDAISSSESELTTVSEFYFTKYKSSAPLLGWIQRIQNGQIQID 64
Query: 63 ARVVTDPNTILRVGSKLVYHRIPWKEPDAPHTIEVLYEDDDIIALNKPSGLQVLPGGLFQ 122
VV DPNT+LR GSKLVY R+PWKEPD P+++EVLYEDDD+IALNKPSGLQVLPGGLFQ
Sbjct: 65 GEVVKDPNTLLRSGSKLVYSRLPWKEPDTPYSLEVLYEDDDLIALNKPSGLQVLPGGLFQ 124
Query: 123 QRTVLTQLQW-ETNNKSTIEAHQKPHPVPVHRLGRGTSGILLCAKTKLARARLASYFADG 181
QRTVLTQLQW N S I + + PHPVPVHRLGRGTSGILLCAKTKLA+ +LA+YFA+G
Sbjct: 125 QRTVLTQLQWCFGKNDSYIGSRESPHPVPVHRLGRGTSGILLCAKTKLAKTKLAAYFAEG 184
Query: 182 TSRIEGKGRHTNQELG---KIAKIYRALVSGIINNDQVTINQPIGMVKYPGVAKGLYVAS 238
TS + G G + +QE G K++KIYRAL GI+ D+V I QPIG+V+YPGVA+GLYVAS
Sbjct: 185 TSLV-GSG-NLDQECGTGRKLSKIYRALADGIVEEDEVVIKQPIGVVRYPGVAQGLYVAS 242
Query: 239 QSGKPALSVVHVLERNVKENSTLVQVKIQSGRPHQIRIHLSFIGHPLLGDPLYAVGGQPK 298
GKPA S V VLER+ ++N +LV+V+IQSGRPHQIRIHL+++GHPL+GDPLY GGQPK
Sbjct: 243 PEGKPAFSKVFVLERDREKNCSLVKVEIQSGRPHQIRIHLAYMGHPLVGDPLYVAGGQPK 302
Query: 299 CLDCDFLDE--SFAEDGGYTRPSKPVPGDCGYHLHAHRLFLSHPI-TNEIIEMIAPLPSI 355
C D D +D+ +FAEDGGY RP++ VPGDCGYHLHAH++ L + + T+++++++APLP I
Sbjct: 303 CFDPDLVDDAAAFAEDGGYRRPNQAVPGDCGYHLHAHQVELPNLLNTHKVVKIVAPLPPI 362
Query: 356 LQTHEE 361
LQT E
Sbjct: 363 LQTRCE 368
>emb|CAC07917.1| putative protein [Arabidopsis thaliana] gi|11358237|pir||T46096
hypothetical protein T25B15.30 - Arabidopsis thaliana
Length = 376
Score = 480 bits (1236), Expect = e-134
Identities = 240/373 (64%), Positives = 300/373 (80%), Gaps = 17/373 (4%)
Query: 4 PKKIGIPWPEFNDGLSYDDVVRPSDAGLTLI-EFYSTKYKSSAPLQGWLQRIKSDQITLD 62
P++IG+PWPE NDGL+Y D + S++ LT + EFY TKYKSSAPL GW+QRI++ QI +D
Sbjct: 5 PQRIGLPWPELNDGLAYKDAISSSESELTTVSEFYFTKYKSSAPLLGWIQRIQNGQIQID 64
Query: 63 ARVVTDPNTILRVGSKLVYHRIPWKEPDAPHTIEVLYEDDDIIALNKPSGLQVLPGGLFQ 122
VV DPNT+LR GSKLVY R+PWKEPD P+++EVLYEDDD+IALNKPSGLQVLPGGLFQ
Sbjct: 65 GEVVKDPNTLLRSGSKLVYSRLPWKEPDTPYSLEVLYEDDDLIALNKPSGLQVLPGGLFQ 124
Query: 123 QRTVLTQLQW-ETNNKSTIEAHQKPHPVPVHRLGRGTSGILLCAKTKLARARLASYFADG 181
QRTVLTQLQW N S I + + PHPVPVHRLGRGTSGILLCAKTKLA+ +LA+YFA+G
Sbjct: 125 QRTVLTQLQWCFGKNDSYIGSRESPHPVPVHRLGRGTSGILLCAKTKLAKTKLAAYFAEG 184
Query: 182 TSRIEGKGRHTNQELG---KIAKIYRALVSGIINNDQ-------VTINQPIGMVKYPGVA 231
TS + G G + +QE G K++KIYRAL GI+ D+ V I QPIG+V+YPGVA
Sbjct: 185 TSLV-GSG-NLDQECGTGRKLSKIYRALADGIVEEDEARPFLLFVVIKQPIGVVRYPGVA 242
Query: 232 KGLYVASQSGKPALSVVHVLERNVKENSTLVQVKIQSGRPHQIRIHLSFIGHPLLGDPLY 291
+GLYVAS GKPA S V VLER+ ++N +LV+V+IQSGRPHQIRIHL+++GHPL+GDPLY
Sbjct: 243 QGLYVASPEGKPAFSKVFVLERDREKNCSLVKVEIQSGRPHQIRIHLAYMGHPLVGDPLY 302
Query: 292 AVGGQPKCLDCDFLDE--SFAEDGGYTRPSKPVPGDCGYHLHAHRLFLSHPI-TNEIIEM 348
GGQPKC D D +D+ +FAEDGGY RP++ VPGDCGYHLHAH++ L + + T++++++
Sbjct: 303 VAGGQPKCFDPDLVDDAAAFAEDGGYRRPNQAVPGDCGYHLHAHQVELPNLLNTHKVVKI 362
Query: 349 IAPLPSILQTHEE 361
+APLP ILQT E
Sbjct: 363 VAPLPPILQTRCE 375
>gb|AAW28554.1| At3g52260 [Arabidopsis thaliana] gi|22331748|ref|NP_190794.2|
pseudouridine synthase family protein [Arabidopsis
thaliana]
Length = 294
Score = 390 bits (1002), Expect = e-107
Identities = 194/287 (67%), Positives = 237/287 (81%), Gaps = 7/287 (2%)
Query: 4 PKKIGIPWPEFNDGLSYDDVVRPSDAGLTLI-EFYSTKYKSSAPLQGWLQRIKSDQITLD 62
P++IG+PWPE NDGL+Y D + S++ LT + EFY TKYKSSAPL GW+QRI++ QI +D
Sbjct: 5 PQRIGLPWPELNDGLAYKDAISSSESELTTVSEFYFTKYKSSAPLLGWIQRIQNGQIQID 64
Query: 63 ARVVTDPNTILRVGSKLVYHRIPWKEPDAPHTIEVLYEDDDIIALNKPSGLQVLPGGLFQ 122
VV DPNT+LR GSKLVY R+PWKEPD P+++EVLYEDDD+IALNKPSGLQVLPGGLFQ
Sbjct: 65 GEVVKDPNTLLRSGSKLVYSRLPWKEPDTPYSLEVLYEDDDLIALNKPSGLQVLPGGLFQ 124
Query: 123 QRTVLTQLQW-ETNNKSTIEAHQKPHPVPVHRLGRGTSGILLCAKTKLARARLASYFADG 181
QRTVLTQLQW N S I + + PHPVPVHRLGRGTSGILLCAKTKLA+ +LA+YFA+G
Sbjct: 125 QRTVLTQLQWCFGKNDSYIGSRESPHPVPVHRLGRGTSGILLCAKTKLAKTKLAAYFAEG 184
Query: 182 TSRIEGKGRHTNQELG---KIAKIYRALVSGIINNDQVTINQPIGMVKYPGVAKGLYVAS 238
TS + G G + +QE G K++KIYRAL GI+ D+V I QPIG+V+YPGVA+GLYVAS
Sbjct: 185 TSLV-GSG-NLDQECGTGRKLSKIYRALADGIVEEDEVVIKQPIGVVRYPGVAQGLYVAS 242
Query: 239 QSGKPALSVVHVLERNVKENSTLVQVKIQSGRPHQIRIHLSFIGHPL 285
GKPA S V VLER+ ++N +LV+V+IQSGRPHQIRIHL+++GHPL
Sbjct: 243 PEGKPAFSKVFVLERDREKNCSLVKVEIQSGRPHQIRIHLAYMGHPL 289
>gb|AAL47436.1| AT3g52260/T25B15_30 [Arabidopsis thaliana]
Length = 294
Score = 387 bits (994), Expect = e-106
Identities = 193/287 (67%), Positives = 236/287 (81%), Gaps = 7/287 (2%)
Query: 4 PKKIGIPWPEFNDGLSYDDVVRPSDAGLTLI-EFYSTKYKSSAPLQGWLQRIKSDQITLD 62
P++IG+PWPE NDGL+Y D + S++ LT + EFY TKYKSSAPL GW+QRI++ QI +D
Sbjct: 5 PQRIGLPWPELNDGLAYKDAISSSESELTTVSEFYFTKYKSSAPLLGWIQRIQNGQIQID 64
Query: 63 ARVVTDPNTILRVGSKLVYHRIPWKEPDAPHTIEVLYEDDDIIALNKPSGLQVLPGGLFQ 122
VV DPNT+LR GSKLVY R+PWKEPD P+++EVLYEDDD+IALNKPSGLQVLPGGLFQ
Sbjct: 65 GEVVKDPNTLLRSGSKLVYSRLPWKEPDTPYSLEVLYEDDDLIALNKPSGLQVLPGGLFQ 124
Query: 123 QRTVLTQLQW-ETNNKSTIEAHQKPHPVPVHRLGRGTSGILLCAKTKLARARLASYFADG 181
QRTVLTQLQW N S I + + PHPVPVHRLGRGTSGILLCAKTKLA+ +LA+YFA+G
Sbjct: 125 QRTVLTQLQWCFGKNDSYIGSRESPHPVPVHRLGRGTSGILLCAKTKLAKTKLAAYFAEG 184
Query: 182 TSRIEGKGRHTNQELG---KIAKIYRALVSGIINNDQVTINQPIGMVKYPGVAKGLYVAS 238
TS + G G + +QE G K++KIYRAL GI+ D+V I QPIG+V+YPGVA+GLYVAS
Sbjct: 185 TSLV-GSG-NLDQECGTGRKLSKIYRALADGIVEEDEVVIKQPIGVVRYPGVAQGLYVAS 242
Query: 239 QSGKPALSVVHVLERNVKENSTLVQVKIQSGRPHQIRIHLSFIGHPL 285
KPA S V VLER+ ++N +LV+V+IQSGRPHQIRIHL+++GHPL
Sbjct: 243 PEWKPAFSKVFVLERDREKNCSLVKVEIQSGRPHQIRIHLAYMGHPL 289
>ref|ZP_00517595.1| Pseudouridine synthase, RluD [Crocosphaera watsonii WH 8501]
gi|67854003|gb|EAM49318.1| Pseudouridine synthase, RluD
[Crocosphaera watsonii WH 8501]
Length = 302
Score = 248 bits (634), Expect = 2e-64
Identities = 143/339 (42%), Positives = 200/339 (58%), Gaps = 42/339 (12%)
Query: 15 NDGLSYDDVVRPSDAGLTLIEFYSTKYKSSAPLQGWLQRIKSDQITLDARVVTDPNTILR 74
N G Y + ++ + AGLTLIE+Y TK+ + Q WL+RI S Q+ +D + V DP T+L+
Sbjct: 2 NKGWIYREKIKQNHAGLTLIEYY-TKHYHHSNYQQWLERIISGQVLVDEQTV-DPQTVLK 59
Query: 75 VGSKLVYHRIPWKEPDAPHTIEVLYEDDDIIALNKPSGLQVLPGGLFQQRTVLTQLQWET 134
G +L YHR PW+EP+ P + E++YEDD+++ +NKPSGL VLPGG F + T+L QL+
Sbjct: 60 TGQRLSYHRPPWQEPEVPLSFEIIYEDDNLLIVNKPSGLPVLPGGGFLEHTLLWQLK--- 116
Query: 135 NNKSTIEAHQKPHPVPVHRLGRGTSGILLCAKTKLARARLASYFADGTSRIEGKGRHTNQ 194
+ + + P+P+HRLGRGTSG++L K+ +AR+ L +
Sbjct: 117 ------KQYPQNEPIPIHRLGRGTSGLMLMGKSAIARSHLTQQMRE-------------- 156
Query: 195 ELGKIAKIYRALVSGIINNDQVTINQPIGMVKYPGVAKGLYVASQSGKPALSVVHVLERN 254
KI+K+Y+ L+ D+ +I+ PIG + P V +Y A GK A S VL R
Sbjct: 157 --QKISKVYQGLIGKNNLPDKFSIDHPIGKIPDP-VLGYIYGAKVDGKYAYSECRVLGRY 213
Query: 255 VKENSTLVQVKIQSGRPHQIRIHLSFIGHPLLGDPLYAVGGQPKCLDCDFLDESFAEDGG 314
V + TL++V I +GRPHQIRIHL+ IG+PLLGDPLY VGG PK F ED
Sbjct: 214 V--DKTLLEVTILTGRPHQIRIHLATIGYPLLGDPLYVVGGIPK--------TGFKEDND 263
Query: 315 YTRPSKPVPGDCGYHLHAHRLFLSHPITNEIIEMIAPLP 353
VPGDCGY LHA++L HP+TN+ + P
Sbjct: 264 TIN----VPGDCGYFLHAYQLSFVHPVTNQCLTFTCERP 298
>ref|ZP_00521998.1| Pseudouridine synthase, RluD [Solibacter usitatus Ellin6076]
gi|67864041|gb|EAM59049.1| Pseudouridine synthase, RluD
[Solibacter usitatus Ellin6076]
Length = 304
Score = 215 bits (547), Expect = 2e-54
Identities = 132/347 (38%), Positives = 183/347 (52%), Gaps = 50/347 (14%)
Query: 15 NDGLSYDDVVRPSDAGLTLIEFYSTKYKSSAPLQGWLQRIKSDQITLDARVVTDPNTILR 74
N G +Y ++ G TL+ ++ Y S + W Q + + ++TL+ T ++ R
Sbjct: 3 NRGYAYTTIISSKYPGQTLLSHLASLYPHSTS-EAWQQNLNNGEVTLNGVAATGSESV-R 60
Query: 75 VGSKLVYHRIPWKEPDAPHTIEVLYEDDDIIALNKPSGLQVLPGGLFQQRTVLTQLQWET 134
G L+++R PW EPD P EVL+ED ++A+NKPSGL LPGG F + T+L +Q T
Sbjct: 61 SGQILIWNRPPWIEPDVPKHFEVLFEDPHLLAVNKPSGLPTLPGGGFLENTLLRLVQKHT 120
Query: 135 NNKSTIEAHQKPHPVPVHRLGRGTSGILLCAKTKLARARLASYFADGTSRIEGKGRHTNQ 194
P+ PVHRLGR TSGI+L A+T A + L N
Sbjct: 121 -----------PNANPVHRLGRATSGIVLFARTPQAASNL----------------FANW 153
Query: 195 ELGKIAKIYRALVSGIINNDQVTINQPIGMVKYPGVAKGLYVASQSGKPALSVVHVLERN 254
KI KIYRAL I +D I PIG+V +P + ++ A+ SGKP+ S+ V+ R+
Sbjct: 154 NTPKIRKIYRALAQNIAQHDAYEILTPIGLVPHPRIG-SVWAANPSGKPSKSLAKVISRS 212
Query: 255 VKENSTLVQVKIQSGRPHQIRIHLSFIGHPLLGDPLYAVGGQPKCLDCDFLDESFAEDGG 314
++T +V + SGRPHQIRIHL+ IGHPL+GDPLY + GQP
Sbjct: 213 T--STTTFEVSLNSGRPHQIRIHLASIGHPLVGDPLYGLTGQP----------------- 253
Query: 315 YTRPSKPVPGDCGYHLHAHRLFLSHPITNEIIEMIAPLPSILQTHEE 361
+PGD GY LHA L HPIT E I + A LPS +H++
Sbjct: 254 -LENLPGLPGDGGYFLHAQFLQFQHPITGEQINLEAALPSGFSSHQQ 299
>ref|NP_923129.1| probable pseudouridine synthase [Gloeobacter violaceus PCC 7421]
gi|35210743|dbj|BAC88124.1| gll0183 [Gloeobacter
violaceus PCC 7421]
Length = 314
Score = 212 bits (539), Expect = 2e-53
Identities = 134/339 (39%), Positives = 184/339 (53%), Gaps = 46/339 (13%)
Query: 15 NDGLSYDDVVRPSDAGLTLIEFYSTKYKSSAPLQGWLQRIKSDQITLDARVVTDPNTILR 74
N G +Y D + P AGLT++E+Y +Y+ S + W RI++ Q+ LD R T +LR
Sbjct: 18 NQGWTYCDRIGPESAGLTVLEYYLLRYRHSGDHE-WQARIETGQVLLDHRS-TSSTELLR 75
Query: 75 VGSKLVYHRIPWKEPDAPHTIEVLYEDDDIIALNKPSGLQVLPGGLFQQRTVLTQLQWET 134
L Y R PW+EP P I ++Y D+ ++ + KP GL VLPGG F + T+L W+
Sbjct: 76 SAQVLTYCRPPWQEPSVPLDIVIIYADEQVLVVAKPGGLPVLPGGGFLEHTLL----WQV 131
Query: 135 NNKSTIEAHQKPHPVPVHRLGRGTSGILLCAKTKLARARLASYFADGTSRIEGKGRHTNQ 194
+ A ++P PVP+HRLGRGTSG++L A+T +RA L+
Sbjct: 132 RRRF---AGEEP-PVPIHRLGRGTSGLVLLARTHSSRAALSE----------------QM 171
Query: 195 ELGKIAKIYRALVSGIINNDQVTINQPIGMVKYPGVAKGLYVASQSGKPALSVVHVLERN 254
+I K+YRAL SG D+ TI QPIG V +P + L+ A+ G PA S VL R
Sbjct: 172 RRQRIGKLYRALASGTGMRDRFTIRQPIGKVPHP-LLGTLWAAAADGLPARSDCVVLRRQ 230
Query: 255 VKENSTLVQVKIQSGRPHQIRIHLSFIGHPLLGDPLYAVGGQPKCLDCDFLDESFAEDGG 314
+ +L+ V I +GRPHQIRIHL+ G+PL+GDPL+ GG + LD D
Sbjct: 231 CTD--SLLAVAISTGRPHQIRIHLAAAGYPLVGDPLFGPGGVAR-LDND----------- 276
Query: 315 YTRPSKPVPGDCGYHLHAHRLFLSHPITNEIIEMIAPLP 353
VPGD GY LHAHRL HP + + + P P
Sbjct: 277 -----ASVPGDGGYFLHAHRLSFLHPTSGRRLVLECPPP 310
>ref|ZP_00396840.1| Pseudouridine synthase, RluD [Deinococcus geothermalis DSM 11300]
gi|66781450|gb|EAL82418.1| Pseudouridine synthase, RluD
[Deinococcus geothermalis DSM 11300]
Length = 301
Score = 204 bits (519), Expect = 3e-51
Identities = 134/347 (38%), Positives = 183/347 (52%), Gaps = 50/347 (14%)
Query: 15 NDGLSYDDVVRPSDAGLTLIEFYSTKYKSSAPLQGWLQRIKSDQITLDARVVTDPNTILR 74
NDG +Y + + GLTL+ + + Y S+ W R++ ++ LD P T LR
Sbjct: 4 NDGYTYREELGRRACGLTLLAYLTRHYLHSSE-SDWQARLERGEVCLDGLPAYGPET-LR 61
Query: 75 VGSKLVYHRIPWKEPDAPHTIEVLYEDDDIIALNKPSGLQVLPGGLFQQRTVLTQLQWET 134
G L +HR PW+E DAP V+Y D ++A+ KPSGL LPGG F + T+L Q++ +
Sbjct: 62 PGQVLEWHRPPWEEEDAPLDFAVVYRDAALLAVAKPSGLPTLPGGGFLKHTLLAQVRTDF 121
Query: 135 NNKSTIEAHQKPHPVPVHRLGRGTSGILLCAKTKLARARLASYFADGTSRIEGKGRHTNQ 194
P P+HRLGRGTSG++L A+T A A LA +
Sbjct: 122 -----------PEARPLHRLGRGTSGLVLFARTHEAGAILARAW---------------- 154
Query: 195 ELGKIAKIYRALVSGIINNDQVTINQPIGMVKYPGVAKGLYVASQSGKPALSVVHVLERN 254
+ + K YRAL G ++ I PIG V +P + ++ AS +GK A S+ HVLER
Sbjct: 155 RVRAVQKRYRALAVGTPLQEEYAITTPIGPVPHPRLGS-VFAASAAGKAASSLAHVLERR 213
Query: 255 VKENSTLVQVKIQSGRPHQIRIHLSFIGHPLLGDPLYAVGGQPKCLDCDFLDESFAEDGG 314
+ L V I +GRPHQIRIHL+ IGHPL+GDPLYA GG P+ A+ G
Sbjct: 214 AA--TALFAVDIHTGRPHQIRIHLAAIGHPLVGDPLYASGGLPR-----------ADLPG 260
Query: 315 YTRPSKPVPGDCGYHLHAHRLFLSHPITNEIIEMIAPLPSILQTHEE 361
+PGD GY LHA RL HP+T E + + AP P L+ +E
Sbjct: 261 -------LPGDLGYLLHAERLTFLHPLTGERLSLKAPPPPELRRADE 300
>gb|AAF11241.1| pseudouridine synthase, putative [Deinococcus radiodurans]
gi|7473651|pir||B75366 probable pseudouridine synthase -
Deinococcus radiodurans (strain R1)
gi|15806687|ref|NP_295407.1| pseudouridine synthase,
putative [Deinococcus radiodurans R1]
Length = 321
Score = 201 bits (511), Expect = 3e-50
Identities = 128/353 (36%), Positives = 187/353 (52%), Gaps = 54/353 (15%)
Query: 4 PKKIGIPWPEFNDGLSYDDVVRPSDAGLTLIEFYSTKYKSSAPLQGWLQRIKSDQITLDA 63
P+ G P N G +Y ++ G +++F + +++ S W R+ + ++T+D
Sbjct: 16 PQMSGTPTSTLNSGYAYRTTLQRG--GQRVLDFLAQEFRHSDRAT-WAARLAAGEVTVDG 72
Query: 64 RVVTDPNTILRVGSKLVYHRIPWKEPDAPHTIEVLYEDDDIIALNKPSGLQVLPGGLFQQ 123
V T LR G LV+ R PW EPDAP +V+YED D++A++KPSGL +PGG F +
Sbjct: 73 LTVRGDET-LRPGQMLVWQRPPWAEPDAPLHFDVIYEDADLLAVSKPSGLPTMPGGGFLE 131
Query: 124 RTVLTQLQWETNNKSTIEAHQKPHPVPVHRLGRGTSGILLCAKTKLARARLASYFADGTS 183
T+L Q++ + P P+HRLGRGTSG++L + T A A L
Sbjct: 132 HTLLHQVR-----------GRWPDASPLHRLGRGTSGLVLFSLTSQAGAALLK------- 173
Query: 184 RIEGKGRHTNQELGKIAKIYRALVSGIINNDQVTINQPIGMVKYPGVAKGLYVASQSGKP 243
+ G++ K+YRAL +G+ D+ I PIG V +P + + ++ AS +GK
Sbjct: 174 ---------DWRAGRVQKVYRALSAGVAEQDEYDITVPIGPVAHPRLGQ-VFAASTAGKT 223
Query: 244 ALSVVHVLERNVKENSTLVQVKIQSGRPHQIRIHLSFIGHPLLGDPLYAVGGQPKCLDCD 303
A S VLER +E++TL +V+I +GRPHQIRIHL+ +G PL DPLY GG P
Sbjct: 224 AHSRARVLER--REDTTLFEVEIFTGRPHQIRIHLASLGLPLAVDPLYVSGGLP------ 275
Query: 304 FLDESFAEDGGYTRPSKPVPGDCGYHLHAHRLFLSHPITNEIIEMIAPLPSIL 356
RP +PGD GY LHA L HP + + AP P++L
Sbjct: 276 -------------RPD-ALPGDLGYALHASTLTFQHPAHGAALTLSAPAPTLL 314
>ref|NP_440308.1| hypothetical protein slr1592 [Synechocystis sp. PCC 6803]
gi|3915463|sp|P72970|Y1592_SYNY3 Hypothetical RNA
pseudouridine synthase slr1592 (RNA-uridine isomerase)
(RNA pseudouridylate synthase)
gi|1652063|dbj|BAA16988.1| slr1592 [Synechocystis sp.
PCC 6803]
Length = 291
Score = 194 bits (494), Expect = 3e-48
Identities = 125/328 (38%), Positives = 180/328 (54%), Gaps = 48/328 (14%)
Query: 33 LIEFYSTKYKSSAPLQGWLQRIKSDQITLDARVVTDPNTILRVGSKLVYHRIPWKEPDAP 92
+++FYS +Y S Q W +RI+ QI +D + V DP L+ G L Y R PW EP+ P
Sbjct: 1 MLDFYSQRYLHSER-QAWQERIEQGQIEVDGKTV-DPTYPLQAGQTLTYWRSPWVEPEVP 58
Query: 93 HTIEVLYEDDDIIALNKPSGLQVLPGGLFQQRTVLTQLQWETNNKSTIEAHQKPHPVPVH 152
+ VL+ D+D+ A++KPSGL VLPGG F TVL QL+ + +S P+H
Sbjct: 59 LQLSVLHRDEDLWAIDKPSGLPVLPGGDFVYHTVLEQLKIQYPQESLF---------PLH 109
Query: 153 RLGRGTSGILLCAKTKLARARLASYFADGTSRIEGKGRHTNQELGKIAKIYRALVSGIIN 212
RLG GTSG+LL ++ L R L+ F + R KIYR ++
Sbjct: 110 RLGTGTSGLLLLGRSTLGRRELSRQFREHICR----------------KIYRTVIGPCDL 153
Query: 213 NDQVTINQPIGMVKYPGVAKGLYVASQSGKPALSVVHVLERNVKENSTLVQVKIQSGRPH 272
++ +Q IG + YP + +Y AS GKPA S +L R+ ++ T ++V+I++GRPH
Sbjct: 154 PEKFECHQAIGKIPYPQLGY-IYAASSHGKPAQSYGRILARSPEK--TWLEVEIRTGRPH 210
Query: 273 QIRIHLSFIGHPLLGDPLYAVGGQPKCLDCDFLDESFAEDGGYTRPSKPVPGDCGYHLHA 332
QIRIHL+ +G+PLLGD LY GG P ++C R ++ P D GY LH+
Sbjct: 211 QIRIHLASLGYPLLGDRLYGPGGVP--INC--------------RTAR--PSDGGYTLHS 252
Query: 333 HRLFLSHPITNEIIEMIAPLPSILQTHE 360
++L H T + I++ APLP L T E
Sbjct: 253 YQLGFLHLHTQQAIKLTAPLPPELITPE 280
>emb|CAE22216.1| Pseudouridine synthase [Prochlorococcus marinus str. MIT 9313]
gi|33864307|ref|NP_895867.1| Pseudouridine synthase
[Prochlorococcus marinus str. MIT 9313]
Length = 338
Score = 187 bits (476), Expect = 3e-46
Identities = 133/365 (36%), Positives = 190/365 (51%), Gaps = 61/365 (16%)
Query: 14 FNDGLSYDDVVRPSDAGLTLIEFYSTKYKSSAPLQGWLQRIKSDQITLDARVVTDPNTIL 73
FN G +Y + DAG L+E + Y S+ W QR+++ QI L+ +V + +L
Sbjct: 17 FNHGWTYSQRIAAVDAGRGLVELLAVHYTHSSR-DVWQQRLQAGQIRLNGELVR-LDQLL 74
Query: 74 RVGSKLVYHRIPWKEPDAPHTIEVLYEDDDIIALNKPSGLQVLPGGLFQQRTV--LTQLQ 131
G ++ + R PW E P + EV+++D D++ +NKPSGL V+PGG F T+ L Q +
Sbjct: 75 ASGDQVCWQRPPWLEAAVPASWEVIFDDGDLLVVNKPSGLPVIPGGGFLDHTLTGLLQRR 134
Query: 132 WETNNKSTIEAHQKPHPVPVHRLGRGTSGILLCAKTKLARARLASYFADGTSRIEGKGRH 191
+ + +S + P PVHRLGR TSG+L+CA+ + RA+L++ F T+ +G
Sbjct: 135 CQLDGESRL-------PRPVHRLGRFTSGLLICARQRETRAKLSAMFRRATAGDQG---- 183
Query: 192 TNQELGKIAKIYRALVSGIIN---NDQVTINQPIGMVKYPGVAKGLYVASQSGKP----- 243
KIYRAL +N ++ TI PI +P V K ++VA+ S +
Sbjct: 184 -------CKKIYRALAHRNLNVHCDEVATIRVPIVQTAHPMVGK-VWVAADSSRQHDSKQ 235
Query: 244 -------ALSVVHVLERNVKENSTLVQVKIQSGRPHQIRIHLSFIGHPLLGDPLYAVGGQ 296
ALS V +LER ++++ L++V IQ+GRPHQIRIHL+ IG PL+GDPLY GQ
Sbjct: 236 SESRVLTALSTVRLLER--RQDADLLEVAIQTGRPHQIRIHLAAIGSPLIGDPLYGFDGQ 293
Query: 297 PKCLDCDFLDESFAEDGGYTRPSKPVPGDCGYHLHAHRLFLSHPITNEIIEMIAPLPSIL 356
S PGD GY LHAHRL L P N + A P L
Sbjct: 294 IS--------------------SAATPGDGGYCLHAHRL-LDVPYRNRLHGFTACSPRCL 332
Query: 357 QTHEE 361
Q E
Sbjct: 333 QMKTE 337
>emb|CAE08803.1| probable pseudouridine synthase [Synechococcus sp. WH 8102]
gi|33866818|ref|NP_898377.1| probable pseudouridine
synthase [Synechococcus sp. WH 8102]
Length = 315
Score = 137 bits (346), Expect = 4e-31
Identities = 110/323 (34%), Positives = 155/323 (47%), Gaps = 45/323 (13%)
Query: 14 FNDGLSYDDVVRPSDAGLTLIEFYSTKYKSSAPLQGWLQRIKSDQITLDARVVTDPNTIL 73
FNDG +Y D V AG L + + +Y+ SA + W QRI S ++ + + +T L
Sbjct: 6 FNDGWTYRDRVPVGAAGTWLSCWLAERYQHSAG-EVWRQRIHSGELDCNGQRLTADRR-L 63
Query: 74 RVGSKLVYHRIPWKEPDAPHTIEVLYEDDDIIALNKPSGLQVLPGGLFQQRTVLTQLQWE 133
G +++ R PW E P E +++D D++ +NKPSGL V+PGG F T LT L
Sbjct: 64 AGGEAVLWRRPPWLEEAIPDQWETIHDDGDLLVINKPSGLPVMPGGGFLLHT-LTALLEP 122
Query: 134 TNNKSTIEAHQKPHPVPVHRLGRGTSGILLCAKTKLARARLASYFADGTSRIEGKGRHTN 193
T + PVHRLGR TSG+ +CA+ + RA L+ F R EG R
Sbjct: 123 TGAR------------PVHRLGRFTSGLQVCARDPMTRAALSKQF-----RPEGSCRKVY 165
Query: 194 QELGKIAKIYRALVSGIINNDQVTINQP-IGMVKYPGVAKGLYVASQSGKPALSVVHVLE 252
Q + + + ND V P +G V P K + + A S + +LE
Sbjct: 166 QAWAQRVPGLEQGQTLTVTNDVVERQHPLLGWVWGPEPLKQEPIRKR--LTARSELELLE 223
Query: 253 RNVKENSTLVQVKIQSGRPHQIRIHLSFIGHPLLGDPLYAVGGQPKCLDCDFLDESFAED 312
R + + +QV I +GRPHQIRIHL+ +G PLLGDPLY + +
Sbjct: 224 RTAEGDR--LQVTITTGRPHQIRIHLAQLGSPLLGDPLYLLNREIS-------------- 267
Query: 313 GGYTRPSKPVPGDCGYHLHAHRL 335
+ PGD GY LHA +L
Sbjct: 268 ------ATATPGDGGYRLHAWQL 284
>emb|CAD64195.1| pseudouridylate synthase [Lactobacillus plantarum WCFS1]
gi|28378455|ref|NP_785347.1| pseudouridylate synthase
[Lactobacillus plantarum WCFS1]
Length = 307
Score = 128 bits (321), Expect = 3e-28
Identities = 96/307 (31%), Positives = 148/307 (47%), Gaps = 52/307 (16%)
Query: 54 IKSDQITLDARVVTDPNTILRVGSKLVYHRIPWKEPDAPHTI--EVLYEDDDIIALNKPS 111
IK+DQ+T++ ++ + +V + + P I +++YEDDD++ +NKP
Sbjct: 39 IKTDQVTVNGQLEASKYHVKPGDEIVVAAQTVQQTTIEPENIPLDIVYEDDDVLVVNKPQ 98
Query: 112 GLQVLPGGLFQQRTVLTQLQWETNNKSTIEAHQKPHPVPVHRLGRGTSGILLCAKTKLAR 171
G+ V P T++ L + + STI +P V HR+ + TSG+L+ AK A
Sbjct: 99 GMVVHPAPGHPNHTLVNALLYHSP-LSTINGEFRPGIV--HRIDKDTSGLLMVAKNDRAH 155
Query: 172 ARLASYFADGTSRIEGKGRHTNQELGKIAKIYRALVSGIINNDQVTINQPIGMVKYPGVA 231
LA+ T+ E Y ALV G+I D TIN P+G + P
Sbjct: 156 QSLAAQLKAKTNLRE----------------YVALVHGVIKEDTGTINAPLG--RSPKDR 197
Query: 232 KGLYVASQSGKPALSVVHVLERNVKENSTLVQVKIQSGRPHQIRIHLSFIGHPLLGDPLY 291
K + ++G+PA++ VL+R ++ TL+ ++++GR HQIR+HL IGHPL+GDPLY
Sbjct: 198 KKQAIV-KTGRPAVTHFKVLKRY--QHYTLISCRLETGRTHQIRVHLQSIGHPLVGDPLY 254
Query: 292 AVGGQPKCLDCDFLDESFAEDGGYTRPSKPVPGDCGYHLHAHRLFLSHPITNEIIEMIAP 351
P K + G G LHA L HP+T +++ AP
Sbjct: 255 G-------------------------PKKTIKGH-GQFLHAKLLGFKHPVTGDLLTFEAP 288
Query: 352 LPSILQT 358
LP I T
Sbjct: 289 LPPIFTT 295
>ref|NP_441760.1| hypothetical protein slr1629 [Synechocystis sp. PCC 6803]
gi|3915482|sp|P74346|Y1629_SYNY3 Hypothetical RNA
pseudouridine synthase slr1629 (RNA-uridine isomerase)
(RNA pseudouridylate synthase)
gi|1653527|dbj|BAA18440.1| slr1629 [Synechocystis sp.
PCC 6803]
Length = 327
Score = 126 bits (317), Expect = 9e-28
Identities = 96/306 (31%), Positives = 149/306 (48%), Gaps = 51/306 (16%)
Query: 54 IKSDQITLDARVVTDPNTILRVGSKLVYH---RIPWKEPDAPHTIEVLYEDDDIIALNKP 110
I+S Q+ + V D N IL+ G +LV + IP +++LYED+ +I +NKP
Sbjct: 54 IESGQVQRNGLVCQDKNLILKTGDRLVVNIPELIPLDVVAQNIPLDILYEDEQLIIINKP 113
Query: 111 SGLQVLPGGLFQQRTVLTQLQWETNNKSTIEAHQKPHPVPVHRLGRGTSGILLCAKTKLA 170
+GL V PG TV+ L + + I Q+P V HRL + T+G ++ AKT+LA
Sbjct: 114 AGLVVHPGPGHPDGTVVNALLAHCPDLAGIGGVQRPGIV--HRLDKDTTGAMVVAKTELA 171
Query: 171 RARLASYFADGTSRIEGKGRHTNQELGKIAKIYRALVSGIINNDQVTINQPIGMVKYPGV 230
L + T+R ++Y +V G Q T+N P+G ++PG
Sbjct: 172 LHNLQVQLKEKTAR----------------RLYWGIVYGSPKEIQGTVNLPVG--RHPGD 213
Query: 231 AK--GLYVASQSGKPALSVVHVLERNVKENSTLVQVKIQSGRPHQIRIHLSFIGHPLLGD 288
+ G+ + G+ A++ +LER N + ++ ++++GR HQIR+H +GHPL+GD
Sbjct: 214 RQKMGIVPVEKGGREAVTHWRLLERI--GNHSWLEFQLETGRTHQIRVHSKQMGHPLVGD 271
Query: 289 PLYAVGGQPKCLDCDFLDESFAEDGGYTRPSKPVPGDCGYHLHAHRLFLSHPITNEIIEM 348
L YT P G LHAH+L L HP++ EII
Sbjct: 272 NL------------------------YTSPGSVNVNLPGQALHAHQLSLIHPVSGEIITA 307
Query: 349 IAPLPS 354
IAP+P+
Sbjct: 308 IAPMPA 313
>ref|NP_471677.1| hypothetical protein lin2346 [Listeria innocua Clip11262]
gi|16414869|emb|CAC97573.1| lin2346 [Listeria innocua]
gi|25322411|pir||AE1725 probable ribosomal large chain
pseudouridine synthase homolog lin2346 [imported] -
Listeria innocua (strain Clip11262)
Length = 289
Score = 125 bits (314), Expect = 2e-27
Identities = 99/327 (30%), Positives = 161/327 (48%), Gaps = 63/327 (19%)
Query: 30 GLTLIEFYSTKYKSSAPLQGWLQRIKSDQITLDARVVTDPNTILRVGSKL-----VYHRI 84
GLTL K+ + ++ S + L+ + + D NT L VG+ L ++ +I
Sbjct: 14 GLTLTAILQEKFHLGKKARHEIRM--SQDVLLNNKPLDDWNTPLFVGASLSLPIILHDKI 71
Query: 85 PWKEPDAPHTIEVLYEDDDIIALNKPSGLQVLPGGLFQQRTVLTQLQWETNNKSTIEAHQ 144
P E D +E++YEDD ++ +NKP G++ ++ T +Q+ Q
Sbjct: 72 PVYEYD----LEIIYEDDFLLVVNKPVGMKTHANDSYETDTCANAVQYYLEKTG-----Q 122
Query: 145 KPHPVPVHRLGRGTSGILLCAKTKLARARLASYFADGTSRIEGKGRHTNQELGKIAKIYR 204
+ + + VHRL + TSG++L AK KLA A L+ ++E + H + Y
Sbjct: 123 EANALAVHRLDQTTSGLVLFAKNKLAIAGLSW-------QMENRAIH---------RTYL 166
Query: 205 ALVSGIINNDQVTINQPIGMVKYPGVAKGLYVASQSGKPALSVVHVLERNVKENSTLVQV 264
A V G TIN+PIG ++ G + + S G+PA++ V VLE + + N++LV+
Sbjct: 167 AAVDGTWGLVMQTINKPIGEDRHHGSRRQI---SPYGQPAVTHVKVLENDNENNTSLVEC 223
Query: 265 KIQSGRPHQIRIHLSFIGHPLLGDPLYAVGGQPKCLDCDFLDESFAEDGGYTRPSKPVPG 324
+I++GR HQIR+HLS IGHP++GD LY GG +R ++ +
Sbjct: 224 QIETGRTHQIRVHLSGIGHPIIGDTLY---------------------GGSSRANRIM-- 260
Query: 325 DCGYHLHAHRLFLSHPITNEIIEMIAP 351
LHA +L L+HP + + AP
Sbjct: 261 -----LHAEKLHLNHPFKGKEMNFSAP 282
>ref|YP_155572.1| Pseudouridylate synthase [Idiomarina loihiensis L2TR]
gi|56179301|gb|AAV82023.1| Pseudouridylate synthase
[Idiomarina loihiensis L2TR]
Length = 322
Score = 124 bits (310), Expect = 6e-27
Identities = 106/348 (30%), Positives = 172/348 (48%), Gaps = 48/348 (13%)
Query: 15 NDGLSYDDVVRPSDAGLTLIEFYSTKYK--SSAPLQGWLQRIKSDQITLDARVVTDPNTI 72
N + + +V S AG L + + + S + L+ W I ++T+D +V+ P
Sbjct: 2 NKSIELEAIVTESLAGKRLDQALAELFPDYSRSRLKSW---ITDGKVTIDTKVIEKPREK 58
Query: 73 LRVGSKL-VYHRIPWKEPDA--PHTIEVLYEDDDIIALNKPSGLQVLPGGLFQQRTVLTQ 129
+ G+ + V I +E A P ++++YEDDDI+ +NKP+GL V PG RT+L
Sbjct: 59 VVAGALVAVDAEIEGEERHAAQPVELDIVYEDDDILVINKPAGLVVHPGAGAPDRTLLNG 118
Query: 130 LQWETNNKSTIEAHQKPHPVPVHRLGRGTSGILLCAKTKLARARLASYFADGTSRIEGKG 189
L + P +HRL + T+G+++ A+ A+ +L S D
Sbjct: 119 LLHYDPQLDLV-----PRAGIIHRLDKDTTGLMVVARNVPAQTQLVSMMQDRD------- 166
Query: 190 RHTNQELGKIAKIYRALVSGIINNDQVTINQPIGMVKYPGVAKGLYVASQSGKPALSVVH 249
I + Y A+ +G++ + ++QPIG ++P + V QSGKPA++
Sbjct: 167 ---------ITREYEAVCNGVMTAGGM-VDQPIG--RHPTKRTHMAVV-QSGKPAVTHYR 213
Query: 250 VLERNVKENSTLVQVKIQSGRPHQIRIHLSFIGHPLLGDPLYAVGGQPKCLDCDFLDESF 309
VL+R TL++++++SGR HQIR+H++ I HPL+GD Y GG+P+
Sbjct: 214 VLDRY--RAHTLLRLRLESGRTHQIRVHMAHIRHPLVGDLTY--GGRPR-------PPKQ 262
Query: 310 AEDGGYTRPSKPVPGDCGYHLHAHRLFLSHPITNEIIEMIAPLPSILQ 357
AED R + + G LHA L L HPIT + +E P P+ Q
Sbjct: 263 AED----RLLQTLRGFNRQALHAASLGLHHPITGDWLEFNVPRPTDFQ 306
>ref|NP_465768.1| hypothetical protein lmo2244 [Listeria monocytogenes EGD-e]
gi|16411714|emb|CAD00322.1| lmo2244 [Listeria
monocytogenes] gi|25322409|pir||AD1355 probable
ribosomal large chain pseudouridine synthase homolog
lmo2244 [imported] - Listeria monocytogenes (strain
EGD-e)
Length = 289
Score = 124 bits (310), Expect = 6e-27
Identities = 94/301 (31%), Positives = 151/301 (49%), Gaps = 61/301 (20%)
Query: 56 SDQITLDARVVTDPNTILRVGSKL-----VYHRIPWKEPDAPHTIEVLYEDDDIIALNKP 110
S + L+ + + D NT L VG+ L ++ +IP E D +E++YEDD ++ +NKP
Sbjct: 38 SQNVLLNNKPLDDWNTPLFVGASLSLPIFLHDKIPVYEYD----LEIIYEDDFLLVVNKP 93
Query: 111 SGLQVLPGGLFQQRTVLTQLQWETNNKSTIEAHQKPHPVPVHRLGRGTSGILLCAKTKLA 170
++ P ++ T +Q+ Q + + VHRL + TSG++L AK KLA
Sbjct: 94 VDMKTHPNDSYETDTCANAVQYYLEKTG-----QDANALAVHRLDQTTSGLVLFAKNKLA 148
Query: 171 RARLASYFADGTSRIEGKGRHTNQELGKIAKIYRALVSGIINNDQVTINQPIGMVKYPGV 230
A L+ ++E + H + Y A V G TIN+PIG ++ G
Sbjct: 149 IAGLSW-------QMENRAIH---------RTYLAAVDGTWGLVMQTINKPIGEDRHHGS 192
Query: 231 AKGLYVASQSGKPALSVVHVLERNVKENSTLVQVKIQSGRPHQIRIHLSFIGHPLLGDPL 290
+ + S G+PA++ V VLE + ++N++LV+ +I++GR HQIR+HLS IGHP++GD L
Sbjct: 193 RRQI---SPYGQPAVTHVKVLENDNEKNTSLVECQIETGRTHQIRVHLSGIGHPIIGDTL 249
Query: 291 YAVGGQPKCLDCDFLDESFAEDGGYTRPSKPVPGDCGYHLHAHRLFLSHPITNEIIEMIA 350
Y GG R ++ + LHA +L L+HP + + A
Sbjct: 250 Y---------------------GGSPRANRIM-------LHAEKLHLNHPFKGKEMNFSA 281
Query: 351 P 351
P
Sbjct: 282 P 282
>ref|NP_267154.1| pseudouridine synthase [Lactococcus lactis subsp. lactis Il1403]
gi|12723945|gb|AAK05096.1| pseudouridine synthase
[Lactococcus lactis subsp. lactis Il1403]
gi|25322325|pir||F86749 pseudouridine synthase
[imported] - Lactococcus lactis subsp. lactis (strain
IL1403)
Length = 301
Score = 124 bits (310), Expect = 6e-27
Identities = 86/275 (31%), Positives = 127/275 (45%), Gaps = 56/275 (20%)
Query: 95 IEVLYEDDDIIALNKPSGLQVLPGGLFQQRTVLTQLQWETNNKSTIEAHQKPHPVPVHRL 154
+E++YED D+ +NKP G+ V P T++ + + + S+I +P V HR+
Sbjct: 77 LEIIYEDSDVAVVNKPQGMVVHPAAGHASGTLVNAIMYHIKDLSSINGVIRPGIV--HRI 134
Query: 155 GRGTSGILLCAKTKLARARLASYFADGTSRIEGKGRHTNQELGKIAKIYRALVSGIINND 214
+ TSG+L+ AK A LA+ D TS+ + Y A+V G I ND
Sbjct: 135 DKDTSGLLMIAKNDKAHESLAAQLKDKTSK----------------RRYLAIVHGEIAND 178
Query: 215 QVTINQPIGMV----KYPGVAKGLYVASQSGKPALSVVHVLERNVKENSTLVQVKIQSGR 270
+ TI PIG K V G GKPA++ VLER TL+ +++++GR
Sbjct: 179 KGTIEAPIGRNLKDRKKQAVVSG-------GKPAVTHFEVLERFY--GYTLIALRLETGR 229
Query: 271 PHQIRIHLSFIGHPLLGDPLYAVGGQPKCLDCDFLDESFAEDGGYTRPSKPVPGDCGYHL 330
HQIR+H+++IGHP+ GDPLY P+K + + G L
Sbjct: 230 THQIRVHMNYIGHPVAGDPLYG-------------------------PNKTLTPNKGQFL 264
Query: 331 HAHRLFLSHPITNEIIEMIAPLPSILQTHEEAIAQ 365
HA L HP T E +E P I E + +
Sbjct: 265 HAETLGFDHPTTGEFMEFQVDAPEIFNEQLEKLRE 299
>ref|YP_014866.1| ribosomal large subunit pseudouridine synthase, RluD subfamily
[Listeria monocytogenes str. 4b F2365]
gi|47093359|ref|ZP_00231127.1| ribosomal large subunit
pseudouridine synthase, RluD subfamily [Listeria
monocytogenes str. 4b H7858] gi|47018278|gb|EAL09043.1|
ribosomal large subunit pseudouridine synthase, RluD
subfamily [Listeria monocytogenes str. 4b H7858]
gi|46881748|gb|AAT05043.1| ribosomal large subunit
pseudouridine synthase, RluD subfamily [Listeria
monocytogenes str. 4b F2365]
Length = 289
Score = 124 bits (310), Expect = 6e-27
Identities = 94/301 (31%), Positives = 151/301 (49%), Gaps = 61/301 (20%)
Query: 56 SDQITLDARVVTDPNTILRVGSKL-----VYHRIPWKEPDAPHTIEVLYEDDDIIALNKP 110
S + L+ + + D NT L VG+ L ++ +IP E D +E++YEDD ++ +NKP
Sbjct: 38 SQNVLLNNKPLDDWNTPLFVGASLSLPVFLHDKIPVYEYD----LEIIYEDDFLLVVNKP 93
Query: 111 SGLQVLPGGLFQQRTVLTQLQWETNNKSTIEAHQKPHPVPVHRLGRGTSGILLCAKTKLA 170
++ P ++ T +Q+ Q + + VHRL + TSG++L AK KLA
Sbjct: 94 VDMKTHPNDSYETDTCANAVQYYLEKTG-----QDANALAVHRLDQTTSGLVLFAKNKLA 148
Query: 171 RARLASYFADGTSRIEGKGRHTNQELGKIAKIYRALVSGIINNDQVTINQPIGMVKYPGV 230
A L+ ++E + H + Y A V G TIN+PIG ++ G
Sbjct: 149 IAGLSW-------QMENRAIH---------RTYLAAVDGTWGLVMQTINKPIGEDRHHGS 192
Query: 231 AKGLYVASQSGKPALSVVHVLERNVKENSTLVQVKIQSGRPHQIRIHLSFIGHPLLGDPL 290
+ + S G+PA++ V VLE + ++N++LV+ +I++GR HQIR+HLS IGHP++GD L
Sbjct: 193 RRQI---SPYGQPAVTHVKVLENDNEKNTSLVECQIETGRTHQIRVHLSGIGHPIIGDTL 249
Query: 291 YAVGGQPKCLDCDFLDESFAEDGGYTRPSKPVPGDCGYHLHAHRLFLSHPITNEIIEMIA 350
Y GG R ++ + LHA +L L+HP + + A
Sbjct: 250 Y---------------------GGSPRANRIM-------LHAEKLHLNHPFKGKEMNFSA 281
Query: 351 P 351
P
Sbjct: 282 P 282
>ref|ZP_00234442.1| ribosomal large subunit pseudouridine synthase, RluD subfamily
[Listeria monocytogenes str. 1/2a F6854]
gi|47014781|gb|EAL05734.1| ribosomal large subunit
pseudouridine synthase, RluD subfamily [Listeria
monocytogenes str. 1/2a F6854]
Length = 289
Score = 124 bits (310), Expect = 6e-27
Identities = 94/301 (31%), Positives = 151/301 (49%), Gaps = 61/301 (20%)
Query: 56 SDQITLDARVVTDPNTILRVGSKL-----VYHRIPWKEPDAPHTIEVLYEDDDIIALNKP 110
S + L+ + + D NT L VG+ L ++ +IP E D +E++YEDD ++ +NKP
Sbjct: 38 SQNVLLNNKPLDDWNTPLFVGASLSLPIFLHDKIPVYEYD----LEIIYEDDFLLVVNKP 93
Query: 111 SGLQVLPGGLFQQRTVLTQLQWETNNKSTIEAHQKPHPVPVHRLGRGTSGILLCAKTKLA 170
++ P ++ T +Q+ Q + + VHRL + TSG++L AK KLA
Sbjct: 94 VDMKTHPNDSYETDTCANAVQYYLEKTG-----QDTNALAVHRLDQTTSGLVLFAKNKLA 148
Query: 171 RARLASYFADGTSRIEGKGRHTNQELGKIAKIYRALVSGIINNDQVTINQPIGMVKYPGV 230
A L+ ++E + H + Y A V G TIN+PIG ++ G
Sbjct: 149 IAGLSW-------QMENRAIH---------RTYLAAVDGTWGLVMQTINKPIGEDRHHGS 192
Query: 231 AKGLYVASQSGKPALSVVHVLERNVKENSTLVQVKIQSGRPHQIRIHLSFIGHPLLGDPL 290
+ + S G+PA++ V VLE + ++N++LV+ +I++GR HQIR+HLS IGHP++GD L
Sbjct: 193 RRQI---SPYGQPAVTHVKVLENDNEKNTSLVECQIETGRTHQIRVHLSGIGHPIIGDTL 249
Query: 291 YAVGGQPKCLDCDFLDESFAEDGGYTRPSKPVPGDCGYHLHAHRLFLSHPITNEIIEMIA 350
Y GG R ++ + LHA +L L+HP + + A
Sbjct: 250 Y---------------------GGSPRANRIM-------LHAEKLHLNHPFKGKEMNFSA 281
Query: 351 P 351
P
Sbjct: 282 P 282
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.318 0.137 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 677,127,701
Number of Sequences: 2540612
Number of extensions: 30904430
Number of successful extensions: 61659
Number of sequences better than 10.0: 955
Number of HSP's better than 10.0 without gapping: 823
Number of HSP's successfully gapped in prelim test: 132
Number of HSP's that attempted gapping in prelim test: 58297
Number of HSP's gapped (non-prelim): 1295
length of query: 371
length of database: 863,360,394
effective HSP length: 129
effective length of query: 242
effective length of database: 535,621,446
effective search space: 129620389932
effective search space used: 129620389932
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 76 (33.9 bits)
Medicago: description of AC126015.11


