

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
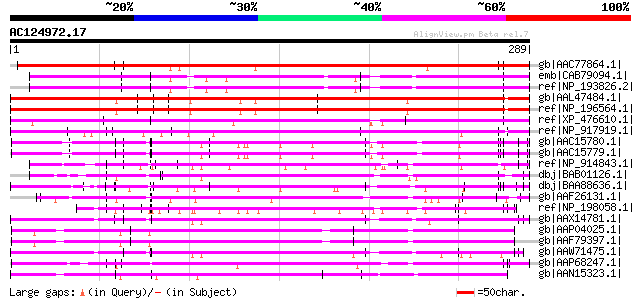
Query= AC124972.17 + phase: 0 /pseudo/partial
(289 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
gb|AAC77864.1| putative receptor-like protein kinase [Arabidopsi... 311 2e-83
emb|CAB79094.1| putative protein [Arabidopsis thaliana] gi|52627... 209 6e-53
ref|NP_193826.2| leucine-rich repeat family protein [Arabidopsis... 209 6e-53
gb|AAL47484.1| AT5g10020/T31P16_9 [Arabidopsis thaliana] 202 7e-51
ref|NP_196564.1| leucine-rich repeat transmembrane protein kinas... 202 7e-51
ref|XP_476610.1| putative phytosulfokine receptor [Oryza sativa ... 199 8e-50
ref|NP_917919.1| putative receptor-like protein kinase [Oryza sa... 187 4e-46
gb|AAC15780.1| Cf-2.2 [Lycopersicon pimpinellifolium] 124 4e-27
gb|AAC15779.1| Cf-2.1 [Lycopersicon pimpinellifolium] gi|7489083... 124 4e-27
ref|NP_914843.1| putative receptor-like protein [Oryza sativa (j... 118 2e-25
dbj|BAB01126.1| receptor protein kinase [Arabidopsis thaliana] g... 116 8e-25
dbj|BAA88636.1| elicitor-inducible LRR receptor-like protein EIL... 114 2e-24
gb|AAF26131.1| putative disease resistance protein [Arabidopsis ... 114 3e-24
ref|NP_198058.1| disease resistance family protein [Arabidopsis ... 113 7e-24
gb|AAX14781.1| RLP1 leucine-rich repeat receptor-like protein [M... 110 4e-23
gb|AAP04025.1| putative disease resistance protein [Arabidopsis ... 110 5e-23
gb|AAF79397.1| F16A14.12 [Arabidopsis thaliana] gi|25402771|pir|... 110 5e-23
gb|AAW71475.1| CLV1-like receptor kinase [Medicago truncatula] 110 5e-23
gb|AAP68247.1| At1g28440 [Arabidopsis thaliana] gi|20260672|gb|A... 110 6e-23
gb|AAN15323.1| Cf-5 disease resistance protein-like [Arabidopsis... 110 6e-23
>gb|AAC77864.1| putative receptor-like protein kinase [Arabidopsis thaliana]
gi|25407852|pir||C84668 probable receptor-like protein
kinase [imported] - Arabidopsis thaliana
gi|15225843|ref|NP_180274.1| leucine-rich repeat
transmembrane protein kinase, putative [Arabidopsis
thaliana]
Length = 1007
Score = 311 bits (796), Expect = 2e-83
Identities = 160/286 (55%), Positives = 208/286 (71%), Gaps = 1/286 (0%)
Query: 5 LVNTAFGNRDIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILCSEGNVISI 64
+V G D +ALLELKKG Q DP VL SWD+K+L S+ CP NWYG+ CS G V SI
Sbjct: 1 MVMKVSGFSDFEALLELKKGFQGDPSRKVLTSWDAKALSSDRCPLNWYGVTCSSGGVTSI 60
Query: 65 TLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKFNGSLP 124
L+ L+G F+F I L ML NLS+ NN F+G++ +I + SLK+LD+S N F+G+LP
Sbjct: 61 DLNGFGLLGSFSFPVIVGLRMLQNLSIANNQFSGTLSNIGSLTSLKYLDVSGNLFHGALP 120
Query: 125 PSFVELRSLVYLNLS-LNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLH 183
LR+L ++NLS N G +P+ F L +L+YLD NSFSG++M +F Q+ SV +
Sbjct: 121 SGIENLRNLEFVNLSGNNNLGGVIPSGFGSLAKLKYLDLQGNSFSGEVMSLFSQLISVEY 180
Query: 184 VDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQ 243
VD+S N FSG+LDLGL SF+ SI+HLNVS NSLVGELFAHDG+P+ D+LEVFDAS+NQ
Sbjct: 181 VDISRNNFSGSLDLGLAKSSFVSSIRHLNVSGNSLVGELFAHDGIPFFDSLEVFDASSNQ 240
Query: 244 LVGNIPSFTFVVSLRILRLACNQLTGSLPETLLKESSMMLSELDLS 289
L G++P F+FVVSL+ILRL NQL+ SLP LL+ESS +L++LDLS
Sbjct: 241 LSGSVPVFSFVVSLKILRLQDNQLSASLPPGLLQESSTILTDLDLS 286
Score = 81.6 bits (200), Expect = 2e-14
Identities = 65/222 (29%), Positives = 105/222 (47%), Gaps = 11/222 (4%)
Query: 59 GNVISITLDNASLVGEFNFLAISNLPMLHN----LSVVNNHFTGSMLHISPM-KSLKFLD 113
G++ S TL+ +L N L+ +LP+ + + NN +G + I S++ +
Sbjct: 295 GSITSSTLEKLNLSS--NRLS-GSLPLKVGHCAIIDLSNNKISGELSRIQNWGDSVEIIR 351
Query: 114 LSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIME 173
LS N G+LP + L L + N G +P + +L+ +D N SG I
Sbjct: 352 LSSNSLTGTLPGQTSQFLRLTSLKAANNSLQGVLPFILGTYPELKEIDLSHNQLSGVIPS 411
Query: 174 IFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDN 233
+ + ++LSNN FSG+L L S+ ++ +SHNSL G L + + N
Sbjct: 412 NLFISAKLTELNLSNNNFSGSLPLQDASTVGNLSLTNIGLSHNSLGGVL--SEELTRFHN 469
Query: 234 LEVFDASNNQLVGNIPSFTFVVSLRILRLACNQLTGSLPETL 275
L D S N GNIP SL++ ++ N L+G++PE L
Sbjct: 470 LISLDLSYNNFEGNIPD-GLPDSLKMFTVSANNLSGNVPENL 510
Score = 74.7 bits (182), Expect = 3e-12
Identities = 73/241 (30%), Positives = 106/241 (43%), Gaps = 46/241 (19%)
Query: 64 ITLDNASLVGEFNFLAISNLPMLHNLSVVN---NHFTGSMLHISPMKSLKFLDLSLNKFN 120
+ + SLVGE A +P +L V + N +GS+ S + SLK L L N+ +
Sbjct: 208 LNVSGNSLVGEL--FAHDGIPFFDSLEVFDASSNQLSGSVPVFSFVVSLKILRLQDNQLS 265
Query: 121 GSLPPSFVELRSLVY------------------------LNLSLNEFSGTVPNVFHKLDQ 156
SLPP ++ S + LNLS N SG++P K+
Sbjct: 266 ASLPPGLLQESSTILTDLDLSLNQLEGPIGSITSSTLEKLNLSSNRLSGSLP---LKVGH 322
Query: 157 LEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHN 216
+D +N SG++ I SV + LS+N +G L G S + L ++N
Sbjct: 323 CAIIDLSNNKISGELSRIQNWGDSVEIIRLSSNSLTGTLP---GQTSQFLRLTSLKAANN 379
Query: 217 SLVGELFAHDGMPYL----DNLEVFDASNNQLVGNIPSFTFV-VSLRILRLACNQLTGSL 271
SL G L P++ L+ D S+NQL G IPS F+ L L L+ N +GSL
Sbjct: 380 SLQGVL------PFILGTYPELKEIDLSHNQLSGVIPSNLFISAKLTELNLSNNNFSGSL 433
Query: 272 P 272
P
Sbjct: 434 P 434
>emb|CAB79094.1| putative protein [Arabidopsis thaliana] gi|5262784|emb|CAB45889.1|
putative protein [Arabidopsis thaliana]
gi|7486968|pir||T10636 hypothetical protein T13K14.100 -
Arabidopsis thaliana
Length = 1143
Score = 209 bits (533), Expect = 6e-53
Identities = 127/330 (38%), Positives = 176/330 (52%), Gaps = 59/330 (17%)
Query: 12 NRDIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILCSEGNVISITLDNASL 71
++DI ALLE KKGI++DP G VLNSW+ +S++ NGCP +W GI+C+ GNV + LDN L
Sbjct: 6 SQDIMALLEFKKGIKHDPTGFVLNSWNDESIDFNGCPSSWNGIVCNGGNVAGVVLDNLGL 65
Query: 72 VGEFNFLAISNLPMLHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKF----------- 119
+ +F SNL L LS+ NN +G + + + KSL+FLDLS N F
Sbjct: 66 TADADFSLFSNLTKLVKLSMSNNSLSGVLPNDLGSFKSLQFLDLSDNLFSSSLPKEIGRS 125
Query: 120 -------------------------------------NGSLPPSFVELRSLVYLNLSLNE 142
+G LP S L L+YLNLS N
Sbjct: 126 VSLRNLSLSGNNFSGEIPESMGGLISLQSLDMSSNSLSGPLPKSLTRLNDLLYLNLSSNG 185
Query: 143 FSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKF---SGALDLGL 199
F+G +P F + LE LD H NS G++ F+ + + +VD+S N+ SG L G+
Sbjct: 186 FTGKMPRGFELISSLEVLDLHGNSIDGNLDGEFFLLTNASYVDISGNRLVTTSGKLLPGV 245
Query: 200 GDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIPSFTFVVSLRI 259
+ SI+HLN+SHN L G L + G NL+V D S N L G +P F +V L +
Sbjct: 246 SE-----SIKHLNLSHNQLEGSLTS--GFQLFQNLKVLDLSYNMLSGELPGFNYVYDLEV 298
Query: 260 LRLACNQLTGSLPETLLKESSMMLSELDLS 289
L+L+ N+ +GSLP LLK S++L+ LDLS
Sbjct: 299 LKLSNNRFSGSLPNNLLKGDSLLLTTLDLS 328
Score = 91.7 bits (226), Expect = 2e-17
Identities = 73/220 (33%), Positives = 112/220 (50%), Gaps = 21/220 (9%)
Query: 63 SITLDNASLVGEFNFLAISNLPMLHN----LSVVNNHFTGSMLHISPMKSLKFLDLSLNK 118
++ L + SL GE LP+L L + NN F G++ S +++++LDLS N
Sbjct: 346 TLDLSSNSLTGE--------LPLLTGGCVLLDLSNNQFEGNLTRWSKWENIEYLDLSQNH 397
Query: 119 FNGSLPPSFVELRSLVYLNLSLNEFSGTVP-NVFHKLDQLEYLDFHSNSFSGDIMEIFYQ 177
F GS P + +L +LNLS N+ +G++P + +L LD SNS G I
Sbjct: 398 FTGSFPDATPQLLRANHLNLSYNKLTGSLPERIPTHYPKLRVLDISSNSLEGPIPGALLS 457
Query: 178 MGSVLHVDLSNNKFSGALDLGLGDVSFLFS-IQHLNVSHNSLVGELFAHDGMPYLDNLEV 236
M ++ + L NN +G +G + S I+ L++SHN G+L G L NL+V
Sbjct: 458 MPTLEEIHLQNNGMTG----NIGPLPSSGSRIRLLDLSHNRFDGDLPGVFGS--LTNLQV 511
Query: 237 FDASNNQLVGNIP-SFTFVVSLRILRLACNQLTGSLPETL 275
+ + N L G++P S +VSL L ++ N TG LP L
Sbjct: 512 LNLAANNLSGSLPSSMNDIVSLSSLDVSQNHFTGPLPSNL 551
Score = 62.8 bits (151), Expect = 1e-08
Identities = 39/121 (32%), Positives = 65/121 (53%), Gaps = 9/121 (7%)
Query: 79 AISNLPMLHNLSVVNNHFTGSMLHISPMKS----LKFLDLSLNKFNGSLPPSFVELRSLV 134
A+ ++P L + + NN TG+ I P+ S ++ LDLS N+F+G LP F L +L
Sbjct: 454 ALLSMPTLEEIHLQNNGMTGN---IGPLPSSGSRIRLLDLSHNRFDGDLPGVFGSLTNLQ 510
Query: 135 YLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGA 194
LNL+ N SG++P+ + + L LD N F+G + +++ ++S N SG
Sbjct: 511 VLNLAANNLSGSLPSSMNDIVSLSSLDVSQNHFTGPLPSNL--SSNIMAFNVSYNDLSGT 568
Query: 195 L 195
+
Sbjct: 569 V 569
>ref|NP_193826.2| leucine-rich repeat family protein [Arabidopsis thaliana]
Length = 977
Score = 209 bits (533), Expect = 6e-53
Identities = 127/330 (38%), Positives = 176/330 (52%), Gaps = 59/330 (17%)
Query: 12 NRDIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILCSEGNVISITLDNASL 71
++DI ALLE KKGI++DP G VLNSW+ +S++ NGCP +W GI+C+ GNV + LDN L
Sbjct: 6 SQDIMALLEFKKGIKHDPTGFVLNSWNDESIDFNGCPSSWNGIVCNGGNVAGVVLDNLGL 65
Query: 72 VGEFNFLAISNLPMLHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKF----------- 119
+ +F SNL L LS+ NN +G + + + KSL+FLDLS N F
Sbjct: 66 TADADFSLFSNLTKLVKLSMSNNSLSGVLPNDLGSFKSLQFLDLSDNLFSSSLPKEIGRS 125
Query: 120 -------------------------------------NGSLPPSFVELRSLVYLNLSLNE 142
+G LP S L L+YLNLS N
Sbjct: 126 VSLRNLSLSGNNFSGEIPESMGGLISLQSLDMSSNSLSGPLPKSLTRLNDLLYLNLSSNG 185
Query: 143 FSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKF---SGALDLGL 199
F+G +P F + LE LD H NS G++ F+ + + +VD+S N+ SG L G+
Sbjct: 186 FTGKMPRGFELISSLEVLDLHGNSIDGNLDGEFFLLTNASYVDISGNRLVTTSGKLLPGV 245
Query: 200 GDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIPSFTFVVSLRI 259
+ SI+HLN+SHN L G L + G NL+V D S N L G +P F +V L +
Sbjct: 246 SE-----SIKHLNLSHNQLEGSLTS--GFQLFQNLKVLDLSYNMLSGELPGFNYVYDLEV 298
Query: 260 LRLACNQLTGSLPETLLKESSMMLSELDLS 289
L+L+ N+ +GSLP LLK S++L+ LDLS
Sbjct: 299 LKLSNNRFSGSLPNNLLKGDSLLLTTLDLS 328
Score = 91.7 bits (226), Expect = 2e-17
Identities = 73/220 (33%), Positives = 112/220 (50%), Gaps = 21/220 (9%)
Query: 63 SITLDNASLVGEFNFLAISNLPMLHN----LSVVNNHFTGSMLHISPMKSLKFLDLSLNK 118
++ L + SL GE LP+L L + NN F G++ S +++++LDLS N
Sbjct: 346 TLDLSSNSLTGE--------LPLLTGGCVLLDLSNNQFEGNLTRWSKWENIEYLDLSQNH 397
Query: 119 FNGSLPPSFVELRSLVYLNLSLNEFSGTVP-NVFHKLDQLEYLDFHSNSFSGDIMEIFYQ 177
F GS P + +L +LNLS N+ +G++P + +L LD SNS G I
Sbjct: 398 FTGSFPDATPQLLRANHLNLSYNKLTGSLPERIPTHYPKLRVLDISSNSLEGPIPGALLS 457
Query: 178 MGSVLHVDLSNNKFSGALDLGLGDVSFLFS-IQHLNVSHNSLVGELFAHDGMPYLDNLEV 236
M ++ + L NN +G +G + S I+ L++SHN G+L G L NL+V
Sbjct: 458 MPTLEEIHLQNNGMTG----NIGPLPSSGSRIRLLDLSHNRFDGDLPGVFGS--LTNLQV 511
Query: 237 FDASNNQLVGNIP-SFTFVVSLRILRLACNQLTGSLPETL 275
+ + N L G++P S +VSL L ++ N TG LP L
Sbjct: 512 LNLAANNLSGSLPSSMNDIVSLSSLDVSQNHFTGPLPSNL 551
Score = 62.8 bits (151), Expect = 1e-08
Identities = 39/121 (32%), Positives = 65/121 (53%), Gaps = 9/121 (7%)
Query: 79 AISNLPMLHNLSVVNNHFTGSMLHISPMKS----LKFLDLSLNKFNGSLPPSFVELRSLV 134
A+ ++P L + + NN TG+ I P+ S ++ LDLS N+F+G LP F L +L
Sbjct: 454 ALLSMPTLEEIHLQNNGMTGN---IGPLPSSGSRIRLLDLSHNRFDGDLPGVFGSLTNLQ 510
Query: 135 YLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGA 194
LNL+ N SG++P+ + + L LD N F+G + +++ ++S N SG
Sbjct: 511 VLNLAANNLSGSLPSSMNDIVSLSSLDVSQNHFTGPLPSNL--SSNIMAFNVSYNDLSGT 568
Query: 195 L 195
+
Sbjct: 569 V 569
>gb|AAL47484.1| AT5g10020/T31P16_9 [Arabidopsis thaliana]
Length = 1048
Score = 202 bits (515), Expect = 7e-51
Identities = 123/294 (41%), Positives = 180/294 (60%), Gaps = 6/294 (2%)
Query: 1 MLLLLVNTAFGNRDIDALLELKKGIQNDPFGLVLNSWDSKSL-ESNGCPQNWYGILCSE- 58
+LLL A ++ +LLE +KGI+++ ++ D+ SL + + CP +W GI C
Sbjct: 13 LLLLHGANAVTETELRSLLEFRKGIRDETSHQRISWSDTSSLTDPSTCPNDWPGISCDPE 72
Query: 59 -GNVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSML-HISPMKSLKFLDLSL 116
G++I+I LD L GE F +S L L NLS+ N F+G ++ + + SL+ LDLS
Sbjct: 73 TGSIIAINLDRRGLSGELKFSTLSGLTRLRNLSLSGNSFSGRVVPSLGGISSLQHLDLSD 132
Query: 117 NKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFY 176
N F G +P EL SL +LNLS N+F G P+ F L QL LD H N GD+ EIF
Sbjct: 133 NGFYGPIPGRISELWSLNHLNLSSNKFEGGFPSGFRNLQQLRSLDLHKNEIWGDVGEIFT 192
Query: 177 QMGSVLHVDLSNNKFSGALDLGLGDVSFLF-SIQHLNVSHNSLVGELFAHDGMPYLDNLE 235
++ +V VDLS N+F+G L L + ++S + +++HLN+SHN+L G+ F+ + + NLE
Sbjct: 193 ELKNVEFVDLSCNRFNGGLSLPMENISSISNTLRHLNLSHNALNGKFFSEESIGSFKNLE 252
Query: 236 VFDASNNQLVGNIPSFTFVVSLRILRLACNQLTGSLPETLLKESSMMLSELDLS 289
+ D NNQ+ G +P F SLRIL+LA N+L G +P+ LL +SS+ L ELDLS
Sbjct: 253 IVDLENNQINGELPHFGSQPSLRILKLARNELFGLVPQELL-QSSIPLLELDLS 305
Score = 85.9 bits (211), Expect = 1e-15
Identities = 77/242 (31%), Positives = 117/242 (47%), Gaps = 45/242 (18%)
Query: 81 SNLPMLHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKFNGSLPPSF-----VELRSLVY 135
S++P+L L + N FTGS+ I+ +L L+LS N +G LP SF ++L +
Sbjct: 295 SSIPLLE-LDLSRNGFTGSISEINS-STLTMLNLSSNGLSGDLPSSFKSCSVIDLSGNTF 352
Query: 136 ----------------LNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMG 179
L+LS N SG++PN +L L +NS SG + ++
Sbjct: 353 SGDVSVVQKWEATPDVLDLSSNNLSGSLPNFTSAFSRLSVLSIRNNSVSGSLPSLWGD-S 411
Query: 180 SVLHVDLSNNKFSGALDLGLGDVSFLF-SIQHLNVSHNSLVG----------ELFAHDGM 228
+DLS+NKFSG + + F F S++ LN+S N+L G EL +
Sbjct: 412 QFSVIDLSSNKFSGFIPVSF----FTFASLRSLNLSRNNLEGPIPFRGSRASELLVLNSY 467
Query: 229 PYLDNLEVFDASNNQLVGNIP-SFTFVVSLRILRLACNQLTGSLPETLLKESSMMLSELD 287
P +E+ D S N L G +P + +++L LA N+L+G LP L K S ++ LD
Sbjct: 468 P---QMELLDLSTNSLTGMLPGDIGTMEKIKVLNLANNKLSGELPSDLNKLSGLLF--LD 522
Query: 288 LS 289
LS
Sbjct: 523 LS 524
Score = 73.9 bits (180), Expect = 5e-12
Identities = 60/187 (32%), Positives = 88/187 (46%), Gaps = 21/187 (11%)
Query: 89 LSVVNNHFTGSMLHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVP 148
LS+ NN +GS+ + +DLS NKF+G +P SF SL LNLS N G +P
Sbjct: 393 LSIRNNSVSGSLPSLWGDSQFSVIDLSSNKFSGFIPVSFFTFASLRSLNLSRNNLEGPIP 452
Query: 149 NVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSI 208
+ +L L+ + ME+ +DLS N +G L GD+ + I
Sbjct: 453 FRGSRASELLVLNSYPQ------MEL---------LDLSTNSLTGMLP---GDIGTMEKI 494
Query: 209 QHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIPSFTFVVSLRILRLACNQLT 268
+ LN+++N L GEL + L L D SNN G IP+ + ++ N L+
Sbjct: 495 KVLNLANNKLSGEL--PSDLNKLSGLLFLDLSNNTFKGQIPN-KLPSQMVGFNVSYNDLS 551
Query: 269 GSLPETL 275
G +PE L
Sbjct: 552 GIIPEDL 558
Score = 52.8 bits (125), Expect = 1e-05
Identities = 33/104 (31%), Positives = 52/104 (49%), Gaps = 4/104 (3%)
Query: 72 VGEFNFLAISNLPMLHNLSVVNNHFTGS----MLHISPMKSLKFLDLSLNKFNGSLPPSF 127
V F F ++ +L + N F GS +L ++ ++ LDLS N G LP
Sbjct: 429 VSFFTFASLRSLNLSRNNLEGPIPFRGSRASELLVLNSYPQMELLDLSTNSLTGMLPGDI 488
Query: 128 VELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDI 171
+ + LNL+ N+ SG +P+ +KL L +LD +N+F G I
Sbjct: 489 GTMEKIKVLNLANNKLSGELPSDLNKLSGLLFLDLSNNTFKGQI 532
>ref|NP_196564.1| leucine-rich repeat transmembrane protein kinase, putative
[Arabidopsis thaliana]
Length = 1048
Score = 202 bits (515), Expect = 7e-51
Identities = 123/294 (41%), Positives = 180/294 (60%), Gaps = 6/294 (2%)
Query: 1 MLLLLVNTAFGNRDIDALLELKKGIQNDPFGLVLNSWDSKSL-ESNGCPQNWYGILCSE- 58
+LLL A ++ +LLE +KGI+++ ++ D+ SL + + CP +W GI C
Sbjct: 13 LLLLHGANAVTETELRSLLEFRKGIRDETSHQRISWSDTSSLTDPSTCPNDWPGISCDPE 72
Query: 59 -GNVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSML-HISPMKSLKFLDLSL 116
G++I+I LD L GE F +S L L NLS+ N F+G ++ + + SL+ LDLS
Sbjct: 73 TGSIIAINLDRRGLSGELKFSTLSGLTRLRNLSLSGNSFSGRVVPSLGGISSLQHLDLSD 132
Query: 117 NKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFY 176
N F G +P EL SL +LNLS N+F G P+ F L QL LD H N GD+ EIF
Sbjct: 133 NGFYGPIPGRISELWSLNHLNLSSNKFEGGFPSGFRNLQQLRSLDLHKNEIWGDVGEIFT 192
Query: 177 QMGSVLHVDLSNNKFSGALDLGLGDVSFLF-SIQHLNVSHNSLVGELFAHDGMPYLDNLE 235
++ +V VDLS N+F+G L L + ++S + +++HLN+SHN+L G+ F+ + + NLE
Sbjct: 193 ELKNVEFVDLSCNRFNGGLSLPMENISSISNTLRHLNLSHNALNGKFFSEESIGSFKNLE 252
Query: 236 VFDASNNQLVGNIPSFTFVVSLRILRLACNQLTGSLPETLLKESSMMLSELDLS 289
+ D NNQ+ G +P F SLRIL+LA N+L G +P+ LL +SS+ L ELDLS
Sbjct: 253 IVDLENNQINGELPHFGSQPSLRILKLARNELFGLVPQELL-QSSIPLLELDLS 305
Score = 85.9 bits (211), Expect = 1e-15
Identities = 77/242 (31%), Positives = 117/242 (47%), Gaps = 45/242 (18%)
Query: 81 SNLPMLHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKFNGSLPPSF-----VELRSLVY 135
S++P+L L + N FTGS+ I+ +L L+LS N +G LP SF ++L +
Sbjct: 295 SSIPLLE-LDLSRNGFTGSISEINS-STLTMLNLSSNGLSGDLPSSFKSCSVIDLSGNTF 352
Query: 136 ----------------LNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMG 179
L+LS N SG++PN +L L +NS SG + ++
Sbjct: 353 SGDVSVVQKWEATPDVLDLSSNNLSGSLPNFTSAFSRLSVLSIRNNSVSGSLPSLWGD-S 411
Query: 180 SVLHVDLSNNKFSGALDLGLGDVSFLF-SIQHLNVSHNSLVG----------ELFAHDGM 228
+DLS+NKFSG + + F F S++ LN+S N+L G EL +
Sbjct: 412 QFSVIDLSSNKFSGFIPVSF----FTFASLRSLNLSRNNLEGPIPFRGSRASELLVLNSY 467
Query: 229 PYLDNLEVFDASNNQLVGNIP-SFTFVVSLRILRLACNQLTGSLPETLLKESSMMLSELD 287
P +E+ D S N L G +P + +++L LA N+L+G LP L K S ++ LD
Sbjct: 468 P---QMELLDLSTNSLTGMLPGDIGTMEKIKVLNLANNKLSGELPSDLNKLSGLLF--LD 522
Query: 288 LS 289
LS
Sbjct: 523 LS 524
Score = 73.9 bits (180), Expect = 5e-12
Identities = 60/187 (32%), Positives = 88/187 (46%), Gaps = 21/187 (11%)
Query: 89 LSVVNNHFTGSMLHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVP 148
LS+ NN +GS+ + +DLS NKF+G +P SF SL LNLS N G +P
Sbjct: 393 LSIRNNSVSGSLPSLWGDSQFSVIDLSSNKFSGFIPVSFFTFASLRSLNLSRNNLEGPIP 452
Query: 149 NVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSI 208
+ +L L+ + ME+ +DLS N +G L GD+ + I
Sbjct: 453 FRGSRASELLVLNSYPQ------MEL---------LDLSTNSLTGMLP---GDIGTMEKI 494
Query: 209 QHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIPSFTFVVSLRILRLACNQLT 268
+ LN+++N L GEL + L L D SNN G IP+ + ++ N L+
Sbjct: 495 KVLNLANNKLSGEL--PSDLNKLSGLLFLDLSNNTFKGQIPN-KLPSQMVGFNVSYNDLS 551
Query: 269 GSLPETL 275
G +PE L
Sbjct: 552 GIIPEDL 558
Score = 52.8 bits (125), Expect = 1e-05
Identities = 33/104 (31%), Positives = 52/104 (49%), Gaps = 4/104 (3%)
Query: 72 VGEFNFLAISNLPMLHNLSVVNNHFTGS----MLHISPMKSLKFLDLSLNKFNGSLPPSF 127
V F F ++ +L + N F GS +L ++ ++ LDLS N G LP
Sbjct: 429 VSFFTFASLRSLNLSRNNLEGPIPFRGSRASELLVLNSYPQMELLDLSTNSLTGMLPGDI 488
Query: 128 VELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDI 171
+ + LNL+ N+ SG +P+ +KL L +LD +N+F G I
Sbjct: 489 GTMEKIKVLNLANNKLSGELPSDLNKLSGLLFLDLSNNTFKGQI 532
>ref|XP_476610.1| putative phytosulfokine receptor [Oryza sativa (japonica
cultivar-group)] gi|34394890|dbj|BAC84362.1| putative
phytosulfokine receptor [Oryza sativa (japonica
cultivar-group)]
Length = 1065
Score = 199 bits (506), Expect = 8e-50
Identities = 124/342 (36%), Positives = 181/342 (52%), Gaps = 53/342 (15%)
Query: 1 MLLLLVNTAFG---NRDIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILCS 57
+LLLL AFG ++DI ALL KKGI +DP G + +SW+ +S++ NGCP +W GI+C+
Sbjct: 9 VLLLLAAPAFGQLPSQDILALLAFKKGITHDPAGFITDSWNDESIDFNGCPASWNGIVCN 68
Query: 58 EGNVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSL 116
NV + LD + G + NL ML LS+ NN+ +GS+ ++ +KSLKF+D+S
Sbjct: 69 GANVAGVVLDGHGISGVADLSVFVNLTMLVKLSMANNNLSGSLPSNVGSLKSLKFMDISN 128
Query: 117 NKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFY 176
N+F+G +P + LRSL L+L+ N FSG +P+ L L+ LD NS SG +
Sbjct: 129 NRFSGPIPDNIGNLRSLQNLSLARNNFSGPLPDSIDGLASLQSLDVSGNSLSGPLPSSLK 188
Query: 177 QMGSVLHVDLSNNKFSGALDLGLG-------------------DVSFLF----------- 206
+ S++ ++LS N F+ + GLG D FL
Sbjct: 189 GLRSMVALNLSYNAFTKGIPSGLGLLVNLQSLDLSWNQLEGGVDWKFLIESTVAHVDFSG 248
Query: 207 -------------------SIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGN 247
++ +LN+S+N L G L + L+V D S+NQL G+
Sbjct: 249 NLLTSTTPKELKFLADISETVLYLNLSNNKLTGSLIDGVELSTFGRLKVLDLSHNQLSGD 308
Query: 248 IPSFTFVVSLRILRLACNQLTGSLPETLLKESSMMLSELDLS 289
+P F +V L +LRLA N TG +P LLK S++LSELDLS
Sbjct: 309 LPGFNYVYDLEVLRLANNAFTGFVPSGLLKGDSLVLSELDLS 350
Score = 89.0 bits (219), Expect = 1e-16
Identities = 77/248 (31%), Positives = 117/248 (47%), Gaps = 32/248 (12%)
Query: 53 GILCSEGNVIS-ITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLHISPMKSLKF 111
G+L + V+S + L +L G N + + L ++ NLS +N G + ++ S
Sbjct: 335 GLLKGDSLVLSELDLSANNLTGHINMITSTTLQVI-NLS--SNALFGDLPMLAG--SCTV 389
Query: 112 LDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDI 171
LDLS NKF G+L L Y++LS N +GT+P+V + +L YL+ NS + I
Sbjct: 390 LDLSNNKFKGNLSVIAKWSNDLEYVDLSQNNLTGTIPDVSSQFLRLNYLNLSHNSLADTI 449
Query: 172 MEIFYQMGSVLHVDLSNNKFSGALDLGL-----------------------GDVSFLFSI 208
E Q + +DLS+N+F G + L G S S+
Sbjct: 450 PEAVVQYPKLTVLDLSSNQFRGPIPANLLTSSMLQELYIHDNMLSGGLSFPGSSSKNLSL 509
Query: 209 QHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIP-SFTFVVSLRILRLACNQL 267
Q L++S N G L D + L +L+ D S N G +P S T + +L L ++ NQ
Sbjct: 510 QVLDISGNHFNGSL--PDEIASLSSLQALDISTNNFSGPLPASITKLAALTALDISINQF 567
Query: 268 TGSLPETL 275
TGSLP+ L
Sbjct: 568 TGSLPDAL 575
Score = 61.6 bits (148), Expect = 2e-08
Identities = 44/145 (30%), Positives = 76/145 (52%), Gaps = 8/145 (5%)
Query: 79 AISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNKFNGSL--PPSFVELRSLVY 135
A+ P L L + +N F G + ++ L+ L + N +G L P S + SL
Sbjct: 452 AVVQYPKLTVLDLSSNQFRGPIPANLLTSSMLQELYIHDNMLSGGLSFPGSSSKNLSLQV 511
Query: 136 LNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGAL 195
L++S N F+G++P+ L L+ LD +N+FSG + ++ ++ +D+S N+F+G+L
Sbjct: 512 LDISGNHFNGSLPDEIASLSSLQALDISTNNFSGPLPASITKLAALTALDISINQFTGSL 571
Query: 196 DLGLGDVSFLFSIQHLNVSHNSLVG 220
L D ++Q N S+N L G
Sbjct: 572 PDALPD-----TLQSFNASYNDLSG 591
>ref|NP_917919.1| putative receptor-like protein kinase [Oryza sativa (japonica
cultivar-group)] gi|22093779|dbj|BAC07070.1| putative
receptor-like protein kinase [Oryza sativa (japonica
cultivar-group)]
Length = 1059
Score = 187 bits (474), Expect = 4e-46
Identities = 111/297 (37%), Positives = 168/297 (56%), Gaps = 9/297 (3%)
Query: 1 MLLLLVNTAFGNRDIDALLELKKGIQNDPFGLVLNSWDSKSLES-----NGCPQNWYGIL 55
+L++L D+ ALLE KKGI + VL SW + GCP W G++
Sbjct: 9 VLVVLGGGGAAGDDVAALLEFKKGISDRGRDPVLGSWSPPATPDAGGGGGGCPSGWRGVV 68
Query: 56 CSEGNVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLH-ISPMKSLKFLDL 114
C G V+ + LD L GE + +S + L NLS+ N F+G + I + SL+ LDL
Sbjct: 69 CDGGAVVGVALDGLGLAGELKLVTLSGMRALQNLSLAGNAFSGRLPPGIGYLSSLRHLDL 128
Query: 115 SLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVP-NVFHKLDQLEYLDFHSNSFSGDIME 173
S N+F G +P +L LV+LNLS N FS P + +L L +D SNSF G+ +
Sbjct: 129 SGNRFYGPIPGRLADLSGLVHLNLSHNNFSSGFPTDGIRQLQNLRRIDLRSNSFWGNAGD 188
Query: 174 IFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFS-IQHLNVSHNSLVGELFAHDGMPYLD 232
+ ++ + ++DLS+N F+GA+DL L +S + + +++LN+SHN L G F ++ +
Sbjct: 189 LLAELRNAEYIDLSDNLFTGAVDLELESLSSIGNTVKYLNLSHNKLQGGFFRNETVGAFK 248
Query: 233 NLEVFDASNNQLVGNIPSFTFVVSLRILRLACNQLTGSLPETLLKESSMMLSELDLS 289
NLEV D SN+ + G +P SL + R+A N L+G +PE +L ++SM L E+DLS
Sbjct: 249 NLEVLDLSNSGIAGMVPQIDAWFSLAVFRVAGNALSGVMPEAML-QNSMRLVEVDLS 304
Score = 81.6 bits (200), Expect = 2e-14
Identities = 71/221 (32%), Positives = 107/221 (48%), Gaps = 24/221 (10%)
Query: 58 EGNVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLHI-SPMKSLKFLDLSL 116
+G V +I L + L G + A S L +L + NN +GS+ + + L+FLDLSL
Sbjct: 362 DGTVETIDLSSNKLEGSYPNDA-SQFQNLVSLKLRNNLLSGSIPSVLGTYQKLQFLDLSL 420
Query: 117 NKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFY 176
N G + P F +L LNLS N F+GT+P + HS I +
Sbjct: 421 NALGGPVLPFFFLSPTLTVLNLSGNNFTGTIP----------FQSTHSTESIALIQPV-- 468
Query: 177 QMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEV 236
+ VDLS+N SG L D+S L ++ L ++ N L GE+ + + L LE
Sbjct: 469 ----LRIVDLSSNSLSGPLP---PDISNLQRVEFLTLAMNELSGEIPSE--ISKLQGLEY 519
Query: 237 FDASNNQLVGNIPSFTFVVSLRILRLACNQLTGSLPETLLK 277
D S+N G IP SL+I ++ N L G++P+++ K
Sbjct: 520 LDLSHNHFTGRIPDMP-QASLKIFNVSYNDLQGTVPKSVEK 559
Score = 70.5 bits (171), Expect = 5e-11
Identities = 59/185 (31%), Positives = 90/185 (47%), Gaps = 13/185 (7%)
Query: 91 VVNNHFTGSMLHISPMKSLKFL--DLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVP 148
V N +G M S++ + DLS N F+GS+P V +L LNLS N FSG++P
Sbjct: 278 VAGNALSGVMPEAMLQNSMRLVEVDLSRNGFSGSVP--VVNSTTLKLLNLSSNTFSGSLP 335
Query: 149 NVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSI 208
+ K +D N SG++ + G+V +DLS+NK G+ D S ++
Sbjct: 336 STVGKCSS---VDLSGNQLSGELAILRAWDGTVETIDLSSNKLEGSYP---NDASQFQNL 389
Query: 209 QHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNI-PSFTFVVSLRILRLACNQL 267
L + +N L G + + G L+ D S N L G + P F +L +L L+ N
Sbjct: 390 VSLKLRNNLLSGSIPSVLGT--YQKLQFLDLSLNALGGPVLPFFFLSPTLTVLNLSGNNF 447
Query: 268 TGSLP 272
TG++P
Sbjct: 448 TGTIP 452
Score = 65.1 bits (157), Expect = 2e-09
Identities = 57/201 (28%), Positives = 90/201 (44%), Gaps = 42/201 (20%)
Query: 33 VLNSWDSK----SLESNGCPQNWYGILCSEGNVISITLDNASLVG------------EFN 76
+L +WD L SN ++ N++S+ L N L G +F
Sbjct: 357 ILRAWDGTVETIDLSSNKLEGSYPNDASQFQNLVSLKLRNNLLSGSIPSVLGTYQKLQFL 416
Query: 77 FLAISNL-----------PMLHNLSVVNNHFTG-----------SMLHISPMKSLKFLDL 114
L+++ L P L L++ N+FTG S+ I P+ L+ +DL
Sbjct: 417 DLSLNALGGPVLPFFFLSPTLTVLNLSGNNFTGTIPFQSTHSTESIALIQPV--LRIVDL 474
Query: 115 SLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEI 174
S N +G LPP L+ + +L L++NE SG +P+ KL LEYLD N F+G I ++
Sbjct: 475 SSNSLSGPLPPDISNLQRVEFLTLAMNELSGEIPSEISKLQGLEYLDLSHNHFTGRIPDM 534
Query: 175 FYQMGSVLHVDLSNNKFSGAL 195
S+ ++S N G +
Sbjct: 535 --PQASLKIFNVSYNDLQGTV 553
Score = 58.9 bits (141), Expect = 2e-07
Identities = 68/242 (28%), Positives = 105/242 (43%), Gaps = 44/242 (18%)
Query: 55 LCSEGNVIS-ITLDNASLVGEF-NFLAISNLPMLHNLSVVNNHFTGSMLHISPMKSLKFL 112
L S GN + + L + L G F + L L + N+ G + I SL
Sbjct: 217 LSSIGNTVKYLNLSHNKLQGGFFRNETVGAFKNLEVLDLSNSGIAGMVPQIDAWFSLAVF 276
Query: 113 DLSLNKFNGSLPPSFVE-LRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDI 171
++ N +G +P + ++ LV ++LS N FSG+VP V L+ L+ SN+FSG +
Sbjct: 277 RVAGNALSGVMPEAMLQNSMRLVEVDLSRNGFSGSVPVV--NSTTLKLLNLSSNTFSGSL 334
Query: 172 MEIFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYL 231
+G VDLS N+ SG L + L A DG
Sbjct: 335 PST---VGKCSSVDLSGNQLSGELAI------------------------LRAWDG---- 363
Query: 232 DNLEVFDASNNQLVGNIPS----FTFVVSLRILRLACNQLTGSLPETLLKESSMMLSELD 287
+E D S+N+L G+ P+ F +VSL++ N L+GS+P L + +L
Sbjct: 364 -TVETIDLSSNKLEGSYPNDASQFQNLVSLKLRN---NLLSGSIPSVLGTYQKLQFLDLS 419
Query: 288 LS 289
L+
Sbjct: 420 LN 421
>gb|AAC15780.1| Cf-2.2 [Lycopersicon pimpinellifolium]
Length = 1112
Score = 124 bits (310), Expect = 4e-27
Identities = 97/322 (30%), Positives = 159/322 (49%), Gaps = 55/322 (17%)
Query: 2 LLLLVNTAFGN-RDIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILCSEGN 60
L L AF + + ALL+ K +N NS+ + + S+ ++WYG++C G
Sbjct: 17 LFYLFTVAFASTEEATALLKWKATFKNQN-----NSFLASWIPSSNACKDWYGVVCFNGR 71
Query: 61 VISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSML-HISPMKSLKFLDLSLNKF 119
V ++ + NAS++G S+LP L NL + N+ G++ I + +L +LDL+ N+
Sbjct: 72 VNTLNITNASVIGTLYAFPFSSLPSLENLDLSKNNIYGTIPPEIGNLTNLVYLDLNNNQI 131
Query: 120 NGSLPPS-------------------FVE-----LRSLVYLNLSLNEFSGTVPNVFHKLD 155
+G++PP F+ LRSL L+L +N SG++P L+
Sbjct: 132 SGTIPPQIGLLAKLQIIRIFHNQLNGFIPKEIGYLRSLTKLSLGINFLSGSIPASVGNLN 191
Query: 156 QLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLG---DVSFLF------ 206
L +L ++N SG I E + S+ +DLS+N +G++ LG ++SFLF
Sbjct: 192 NLSFLYLYNNQLSGSIPEEISYLRSLTELDLSDNALNGSIPASLGNMNNLSFLFLYGNQL 251
Query: 207 ------------SIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIP-SFTF 253
S+ +L++S N+L G + A G L+NL NQL G+IP +
Sbjct: 252 SGSIPEEICYLRSLTYLDLSENALNGSIPASLG--NLNNLSFLFLYGNQLSGSIPEEIGY 309
Query: 254 VVSLRILRLACNQLTGSLPETL 275
+ SL +L L+ N L GS+P +L
Sbjct: 310 LRSLNVLGLSENALNGSIPASL 331
Score = 101 bits (252), Expect = 2e-20
Identities = 74/218 (33%), Positives = 116/218 (52%), Gaps = 8/218 (3%)
Query: 60 NVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNK 118
N+ + L N L G ++ NL L L + NN +GS+ I + SL +LDLS N
Sbjct: 384 NLSMLYLYNNQLSGSIP-ASLGNLNNLSRLYLYNNQLSGSIPEEIGYLSSLTYLDLSNNS 442
Query: 119 FNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQM 178
NG +P SF + +L +L L N+ + +VP L L LD N+ +G I F +
Sbjct: 443 INGFIPASFGNMSNLAFLFLYENQLASSVPEEIGYLRSLNVLDLSENALNGSIPASFGNL 502
Query: 179 GSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFD 238
++ ++L NN+ SG++ ++ +L S+ L++S N+L G + A G L+NL +
Sbjct: 503 NNLSRLNLVNNQLSGSIP---EEIGYLRSLNVLDLSENALNGSIPASFG--NLNNLSRLN 557
Query: 239 ASNNQLVGNIP-SFTFVVSLRILRLACNQLTGSLPETL 275
NNQL G+IP ++ SL L L+ N L GS+P +L
Sbjct: 558 LVNNQLSGSIPEEIGYLRSLNDLGLSENALNGSIPASL 595
Score = 100 bits (249), Expect = 5e-20
Identities = 69/207 (33%), Positives = 112/207 (53%), Gaps = 7/207 (3%)
Query: 79 AISNLPMLHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLN 137
+ NL L L++VNN +GS+ I ++SL LDLS N NGS+P SF L +L LN
Sbjct: 498 SFGNLNNLSRLNLVNNQLSGSIPEEIGYLRSLNVLDLSENALNGSIPASFGNLNNLSRLN 557
Query: 138 LSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDL 197
L N+ SG++P L L L N+ +G I + ++ + L NN+ SG++
Sbjct: 558 LVNNQLSGSIPEEIGYLRSLNDLGLSENALNGSIPASLGNLNNLSMLYLYNNQLSGSIP- 616
Query: 198 GLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIPSFTF-VVS 256
++ +L S+ +L++ +NSL G + A G + NL+ ++N L+G IPS + S
Sbjct: 617 --EEIGYLSSLTYLSLGNNSLNGLIPASFG--NMRNLQALILNDNNLIGEIPSSVCNLTS 672
Query: 257 LRILRLACNQLTGSLPETLLKESSMML 283
L +L + N L G +P+ L S++ +
Sbjct: 673 LEVLYMPRNNLKGKVPQCLGNISNLQV 699
Score = 99.0 bits (245), Expect = 1e-19
Identities = 73/211 (34%), Positives = 109/211 (51%), Gaps = 8/211 (3%)
Query: 64 ITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKFNGS 122
+ L N S+ G F + N+ L L + N S+ I ++SL LDLS N NGS
Sbjct: 436 LDLSNNSING-FIPASFGNMSNLAFLFLYENQLASSVPEEIGYLRSLNVLDLSENALNGS 494
Query: 123 LPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVL 182
+P SF L +L LNL N+ SG++P L L LD N+ +G I F + ++
Sbjct: 495 IPASFGNLNNLSRLNLVNNQLSGSIPEEIGYLRSLNVLDLSENALNGSIPASFGNLNNLS 554
Query: 183 HVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNN 242
++L NN+ SG++ ++ +L S+ L +S N+L G + A G L+NL + NN
Sbjct: 555 RLNLVNNQLSGSIP---EEIGYLRSLNDLGLSENALNGSIPASLG--NLNNLSMLYLYNN 609
Query: 243 QLVGNIP-SFTFVVSLRILRLACNQLTGSLP 272
QL G+IP ++ SL L L N L G +P
Sbjct: 610 QLSGSIPEEIGYLSSLTYLSLGNNSLNGLIP 640
Score = 98.6 bits (244), Expect = 2e-19
Identities = 72/198 (36%), Positives = 108/198 (54%), Gaps = 7/198 (3%)
Query: 80 ISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNL 138
IS L L L + +N GS+ + M +L FL L N+ +GS+P LRSL YL+L
Sbjct: 211 ISYLRSLTELDLSDNALNGSIPASLGNMNNLSFLFLYGNQLSGSIPEEICYLRSLTYLDL 270
Query: 139 SLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLG 198
S N +G++P L+ L +L + N SG I E + S+ + LS N +G++
Sbjct: 271 SENALNGSIPASLGNLNNLSFLFLYGNQLSGSIPEEIGYLRSLNVLGLSENALNGSIPAS 330
Query: 199 LGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIP-SFTFVVSL 257
LG+ L ++ LN+ +N L G + A G L+NL + NNQL G+IP S + +L
Sbjct: 331 LGN---LKNLSRLNLVNNQLSGSIPASLG--NLNNLSMLYLYNNQLSGSIPASLGNLNNL 385
Query: 258 RILRLACNQLTGSLPETL 275
+L L NQL+GS+P +L
Sbjct: 386 SMLYLYNNQLSGSIPASL 403
Score = 94.7 bits (234), Expect = 3e-18
Identities = 74/213 (34%), Positives = 112/213 (51%), Gaps = 9/213 (4%)
Query: 79 AISNLPMLHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLN 137
++ NL L L + N +GS+ I ++SL L LS N NGS+P S L++L LN
Sbjct: 282 SLGNLNNLSFLFLYGNQLSGSIPEEIGYLRSLNVLGLSENALNGSIPASLGNLKNLSRLN 341
Query: 138 LSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDL 197
L N+ SG++P L+ L L ++N SG I + ++ + L NN+ SG++
Sbjct: 342 LVNNQLSGSIPASLGNLNNLSMLYLYNNQLSGSIPASLGNLNNLSMLYLYNNQLSGSIPA 401
Query: 198 GLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIP-SFTFVVS 256
LG+++ ++ L + +N L G + G YL +L D SNN + G IP SF + +
Sbjct: 402 SLGNLN---NLSRLYLYNNQLSGSIPEEIG--YLSSLTYLDLSNNSINGFIPASFGNMSN 456
Query: 257 LRILRLACNQLTGSLPETLLKESSMMLSELDLS 289
L L L NQL S+PE + S L+ LDLS
Sbjct: 457 LAFLFLYENQLASSVPEEIGYLRS--LNVLDLS 487
Score = 93.2 bits (230), Expect = 8e-18
Identities = 74/219 (33%), Positives = 112/219 (50%), Gaps = 25/219 (11%)
Query: 80 ISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNL 138
I L L LS+ N +GS+ + + +L FL L N+ +GS+P LRSL L+L
Sbjct: 163 IGYLRSLTKLSLGINFLSGSIPASVGNLNNLSFLYLYNNQLSGSIPEEISYLRSLTELDL 222
Query: 139 SLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLG 198
S N +G++P ++ L +L + N SG I E + S+ ++DLS N +G++
Sbjct: 223 SDNALNGSIPASLGNMNNLSFLFLYGNQLSGSIPEEICYLRSLTYLDLSENALNGSIPAS 282
Query: 199 LG---DVSFLF------------------SIQHLNVSHNSLVGELFAHDGMPYLDNLEVF 237
LG ++SFLF S+ L +S N+L G + A G L NL
Sbjct: 283 LGNLNNLSFLFLYGNQLSGSIPEEIGYLRSLNVLGLSENALNGSIPASLG--NLKNLSRL 340
Query: 238 DASNNQLVGNIP-SFTFVVSLRILRLACNQLTGSLPETL 275
+ NNQL G+IP S + +L +L L NQL+GS+P +L
Sbjct: 341 NLVNNQLSGSIPASLGNLNNLSMLYLYNNQLSGSIPASL 379
Score = 91.7 bits (226), Expect = 2e-17
Identities = 70/212 (33%), Positives = 108/212 (50%), Gaps = 9/212 (4%)
Query: 79 AISNLPMLHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLN 137
++ NL L L + NN +GS+ I + SL +L L N NG +P SF +R+L L
Sbjct: 594 SLGNLNNLSMLYLYNNQLSGSIPEEIGYLSSLTYLSLGNNSLNGLIPASFGNMRNLQALI 653
Query: 138 LSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDL 197
L+ N G +P+ L LE L N+ G + + + ++ + +S+N FSG L
Sbjct: 654 LNDNNLIGEIPSSVCNLTSLEVLYMPRNNLKGKVPQCLGNISNLQVLSMSSNSFSGELP- 712
Query: 198 GLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIP-SFTFVVS 256
+S L S+Q L+ N+L G + G + +LEVFD NN+L G +P +F+ S
Sbjct: 713 --SSISNLTSLQILDFGRNNLEGAIPQCFG--NISSLEVFDMQNNKLSGTLPTNFSIGCS 768
Query: 257 LRILRLACNQLTGSLPETLLKESSMMLSELDL 288
L L L N+L +P +L ++ L LDL
Sbjct: 769 LISLNLHGNELEDEIPRSL--DNCKKLQVLDL 798
Score = 79.7 bits (195), Expect = 9e-14
Identities = 67/228 (29%), Positives = 107/228 (46%), Gaps = 8/228 (3%)
Query: 64 ITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNKFNGS 122
++L N SL G + N+ L L + +N+ G + + + SL+ L + N G
Sbjct: 628 LSLGNNSLNGLIP-ASFGNMRNLQALILNDNNLIGEIPSSVCNLTSLEVLYMPRNNLKGK 686
Query: 123 LPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVL 182
+P + +L L++S N FSG +P+ L L+ LDF N+ G I + F + S+
Sbjct: 687 VPQCLGNISNLQVLSMSSNSFSGELPSSISNLTSLQILDFGRNNLEGAIPQCFGNISSLE 746
Query: 183 HVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNN 242
D+ NNK SG L + S S+ LN+ N L E+ + L+V D +N
Sbjct: 747 VFDMQNNKLSGTLPT---NFSIGCSLISLNLHGNELEDEI--PRSLDNCKKLQVLDLGDN 801
Query: 243 QLVGNIPSFTFVV-SLRILRLACNQLTGSLPETLLKESSMMLSELDLS 289
QL P + + LR+LRL N+L G + + + L +DLS
Sbjct: 802 QLNDTFPMWLGTLPELRVLRLTSNKLHGPIRSSRAEIMFPDLRIIDLS 849
Score = 65.1 bits (157), Expect = 2e-09
Identities = 65/206 (31%), Positives = 94/206 (45%), Gaps = 34/206 (16%)
Query: 80 ISNLPMLHNLSVVNNHFTGSMLHISP---MKSLKFLDLSLNKFNGSLPPS-FVELRSLVY 135
+ LP L L + +N G + L+ +DLS N F+ LP S F L+ +
Sbjct: 811 LGTLPELRVLRLTSNKLHGPIRSSRAEIMFPDLRIIDLSRNAFSQDLPTSLFEHLKGMRT 870
Query: 136 LNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSF----SGDIMEIFYQMGSVLHVDLSNNKF 191
++ ++ E S Y ++ +S G +EI + +DLS+NKF
Sbjct: 871 VDKTMEEPS--------------YESYYDDSVVVVTKGLELEIVRILSLYTVIDLSSNKF 916
Query: 192 SGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIP-- 249
G + LGD L +I+ LNVSHN+L G + + L LE D S NQL G IP
Sbjct: 917 EGHIPSVLGD---LIAIRILNVSHNALQG--YIPSSLGSLSILESLDLSFNQLSGEIPQQ 971
Query: 250 --SFTFVVSLRILRLACNQLTGSLPE 273
S TF L L L+ N L G +P+
Sbjct: 972 LASLTF---LEFLNLSHNYLQGCIPQ 994
Score = 54.7 bits (130), Expect = 3e-06
Identities = 71/274 (25%), Positives = 120/274 (42%), Gaps = 52/274 (18%)
Query: 34 LNSWDSKSLESNGCPQNWYGILCSEGNVISITLDNASLVGEFNFLAISNLPMLHNLSVVN 93
L +D ++ + +G + I CS +IS+ L L E ++ N L L + +
Sbjct: 745 LEVFDMQNNKLSGTLPTNFSIGCS---LISLNLHGNELEDEIP-RSLDNCKKLQVLDLGD 800
Query: 94 NHFTGSM-LHISPMKSLKFLDLSLNKFNGSLPPSFVELR--SLVYLNLSLNEFSGTVPN- 149
N + + + + L+ L L+ NK +G + S E+ L ++LS N FS +P
Sbjct: 801 NQLNDTFPMWLGTLPELRVLRLTSNKLHGPIRSSRAEIMFPDLRIIDLSRNAFSQDLPTS 860
Query: 150 ----------VFHKLDQLEYLDFHSNSF----SGDIMEIFYQMGSVLHVDLSNNKFSGAL 195
V +++ Y ++ +S G +EI + +DLS+NKF G +
Sbjct: 861 LFEHLKGMRTVDKTMEEPSYESYYDDSVVVVTKGLELEIVRILSLYTVIDLSSNKFEGHI 920
Query: 196 DLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIPSFTFVV 255
LGD L +I+ LNVSHN+L G + + L +L + ++
Sbjct: 921 PSVLGD---LIAIRILNVSHNALQGYIPSS-----LGSLSILES---------------- 956
Query: 256 SLRILRLACNQLTGSLPETLLKESSMMLSELDLS 289
L L+ NQL+G +P+ L S L L+LS
Sbjct: 957 ----LDLSFNQLSGEIPQQL--ASLTFLEFLNLS 984
Score = 51.6 bits (122), Expect = 3e-05
Identities = 29/87 (33%), Positives = 45/87 (51%)
Query: 112 LDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDI 171
+DLS NKF G +P +L ++ LN+S N G +P+ L LE LD N SG+I
Sbjct: 909 IDLSSNKFEGHIPSVLGDLIAIRILNVSHNALQGYIPSSLGSLSILESLDLSFNQLSGEI 968
Query: 172 MEIFYQMGSVLHVDLSNNKFSGALDLG 198
+ + + ++LS+N G + G
Sbjct: 969 PQQLASLTFLEFLNLSHNYLQGCIPQG 995
Score = 42.4 bits (98), Expect = 0.016
Identities = 29/95 (30%), Positives = 47/95 (48%), Gaps = 11/95 (11%)
Query: 76 NFLAISNLPMLHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVY 135
+ +AI L + HN + + S+ +S ++SL DLS N+ +G +P L L +
Sbjct: 926 DLIAIRILNVSHN--ALQGYIPSSLGSLSILESL---DLSFNQLSGEIPQQLASLTFLEF 980
Query: 136 LNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGD 170
LNLS N G +P ++ F SNS+ G+
Sbjct: 981 LNLSHNYLQGCIP------QGPQFRTFESNSYEGN 1009
>gb|AAC15779.1| Cf-2.1 [Lycopersicon pimpinellifolium] gi|7489083|pir||T10504
disease resistance protein Cf-2.1 - currant tomato
gi|1587673|prf||2207203A Cf-2 gene
Length = 1112
Score = 124 bits (310), Expect = 4e-27
Identities = 97/322 (30%), Positives = 159/322 (49%), Gaps = 55/322 (17%)
Query: 2 LLLLVNTAFGN-RDIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILCSEGN 60
L L AF + + ALL+ K +N NS+ + + S+ ++WYG++C G
Sbjct: 17 LFYLFTVAFASTEEATALLKWKATFKNQN-----NSFLASWIPSSNACKDWYGVVCFNGR 71
Query: 61 VISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSML-HISPMKSLKFLDLSLNKF 119
V ++ + NAS++G S+LP L NL + N+ G++ I + +L +LDL+ N+
Sbjct: 72 VNTLNITNASVIGTLYAFPFSSLPSLENLDLSKNNIYGTIPPEIGNLTNLVYLDLNNNQI 131
Query: 120 NGSLPPS-------------------FVE-----LRSLVYLNLSLNEFSGTVPNVFHKLD 155
+G++PP F+ LRSL L+L +N SG++P L+
Sbjct: 132 SGTIPPQIGLLAKLQIIRIFHNQLNGFIPKEIGYLRSLTKLSLGINFLSGSIPASVGNLN 191
Query: 156 QLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLG---DVSFLF------ 206
L +L ++N SG I E + S+ +DLS+N +G++ LG ++SFLF
Sbjct: 192 NLSFLYLYNNQLSGSIPEEISYLRSLTELDLSDNALNGSIPASLGNMNNLSFLFLYGNQL 251
Query: 207 ------------SIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIP-SFTF 253
S+ +L++S N+L G + A G L+NL NQL G+IP +
Sbjct: 252 SGSIPEEICYLRSLTYLDLSENALNGSIPASLG--NLNNLSFLFLYGNQLSGSIPEEIGY 309
Query: 254 VVSLRILRLACNQLTGSLPETL 275
+ SL +L L+ N L GS+P +L
Sbjct: 310 LRSLNVLGLSENALNGSIPASL 331
Score = 101 bits (252), Expect = 2e-20
Identities = 74/218 (33%), Positives = 116/218 (52%), Gaps = 8/218 (3%)
Query: 60 NVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNK 118
N+ + L N L G ++ NL L L + NN +GS+ I + SL +LDLS N
Sbjct: 384 NLSMLYLYNNQLSGSIP-ASLGNLNNLSRLYLYNNQLSGSIPEEIGYLSSLTYLDLSNNS 442
Query: 119 FNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQM 178
NG +P SF + +L +L L N+ + +VP L L LD N+ +G I F +
Sbjct: 443 INGFIPASFGNMSNLAFLFLYENQLASSVPEEIGYLRSLNVLDLSENALNGSIPASFGNL 502
Query: 179 GSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFD 238
++ ++L NN+ SG++ ++ +L S+ L++S N+L G + A G L+NL +
Sbjct: 503 NNLSRLNLVNNQLSGSIP---EEIGYLRSLNVLDLSENALNGSIPASFG--NLNNLSRLN 557
Query: 239 ASNNQLVGNIP-SFTFVVSLRILRLACNQLTGSLPETL 275
NNQL G+IP ++ SL L L+ N L GS+P +L
Sbjct: 558 LVNNQLSGSIPEEIGYLRSLNDLGLSENALNGSIPASL 595
Score = 100 bits (249), Expect = 5e-20
Identities = 69/207 (33%), Positives = 112/207 (53%), Gaps = 7/207 (3%)
Query: 79 AISNLPMLHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLN 137
+ NL L L++VNN +GS+ I ++SL LDLS N NGS+P SF L +L LN
Sbjct: 498 SFGNLNNLSRLNLVNNQLSGSIPEEIGYLRSLNVLDLSENALNGSIPASFGNLNNLSRLN 557
Query: 138 LSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDL 197
L N+ SG++P L L L N+ +G I + ++ + L NN+ SG++
Sbjct: 558 LVNNQLSGSIPEEIGYLRSLNDLGLSENALNGSIPASLGNLNNLSMLYLYNNQLSGSIP- 616
Query: 198 GLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIPSFTF-VVS 256
++ +L S+ +L++ +NSL G + A G + NL+ ++N L+G IPS + S
Sbjct: 617 --EEIGYLSSLTYLSLGNNSLNGLIPASFG--NMRNLQALILNDNNLIGEIPSSVCNLTS 672
Query: 257 LRILRLACNQLTGSLPETLLKESSMML 283
L +L + N L G +P+ L S++ +
Sbjct: 673 LEVLYMPRNNLKGKVPQCLGNISNLQV 699
Score = 99.0 bits (245), Expect = 1e-19
Identities = 73/211 (34%), Positives = 109/211 (51%), Gaps = 8/211 (3%)
Query: 64 ITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKFNGS 122
+ L N S+ G F + N+ L L + N S+ I ++SL LDLS N NGS
Sbjct: 436 LDLSNNSING-FIPASFGNMSNLAFLFLYENQLASSVPEEIGYLRSLNVLDLSENALNGS 494
Query: 123 LPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVL 182
+P SF L +L LNL N+ SG++P L L LD N+ +G I F + ++
Sbjct: 495 IPASFGNLNNLSRLNLVNNQLSGSIPEEIGYLRSLNVLDLSENALNGSIPASFGNLNNLS 554
Query: 183 HVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNN 242
++L NN+ SG++ ++ +L S+ L +S N+L G + A G L+NL + NN
Sbjct: 555 RLNLVNNQLSGSIP---EEIGYLRSLNDLGLSENALNGSIPASLG--NLNNLSMLYLYNN 609
Query: 243 QLVGNIP-SFTFVVSLRILRLACNQLTGSLP 272
QL G+IP ++ SL L L N L G +P
Sbjct: 610 QLSGSIPEEIGYLSSLTYLSLGNNSLNGLIP 640
Score = 98.6 bits (244), Expect = 2e-19
Identities = 72/198 (36%), Positives = 108/198 (54%), Gaps = 7/198 (3%)
Query: 80 ISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNL 138
IS L L L + +N GS+ + M +L FL L N+ +GS+P LRSL YL+L
Sbjct: 211 ISYLRSLTELDLSDNALNGSIPASLGNMNNLSFLFLYGNQLSGSIPEEICYLRSLTYLDL 270
Query: 139 SLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLG 198
S N +G++P L+ L +L + N SG I E + S+ + LS N +G++
Sbjct: 271 SENALNGSIPASLGNLNNLSFLFLYGNQLSGSIPEEIGYLRSLNVLGLSENALNGSIPAS 330
Query: 199 LGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIP-SFTFVVSL 257
LG+ L ++ LN+ +N L G + A G L+NL + NNQL G+IP S + +L
Sbjct: 331 LGN---LKNLSRLNLVNNQLSGSIPASLG--NLNNLSMLYLYNNQLSGSIPASLGNLNNL 385
Query: 258 RILRLACNQLTGSLPETL 275
+L L NQL+GS+P +L
Sbjct: 386 SMLYLYNNQLSGSIPASL 403
Score = 94.7 bits (234), Expect = 3e-18
Identities = 74/213 (34%), Positives = 112/213 (51%), Gaps = 9/213 (4%)
Query: 79 AISNLPMLHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLN 137
++ NL L L + N +GS+ I ++SL L LS N NGS+P S L++L LN
Sbjct: 282 SLGNLNNLSFLFLYGNQLSGSIPEEIGYLRSLNVLGLSENALNGSIPASLGNLKNLSRLN 341
Query: 138 LSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDL 197
L N+ SG++P L+ L L ++N SG I + ++ + L NN+ SG++
Sbjct: 342 LVNNQLSGSIPASLGNLNNLSMLYLYNNQLSGSIPASLGNLNNLSMLYLYNNQLSGSIPA 401
Query: 198 GLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIP-SFTFVVS 256
LG+++ ++ L + +N L G + G YL +L D SNN + G IP SF + +
Sbjct: 402 SLGNLN---NLSRLYLYNNQLSGSIPEEIG--YLSSLTYLDLSNNSINGFIPASFGNMSN 456
Query: 257 LRILRLACNQLTGSLPETLLKESSMMLSELDLS 289
L L L NQL S+PE + S L+ LDLS
Sbjct: 457 LAFLFLYENQLASSVPEEIGYLRS--LNVLDLS 487
Score = 93.2 bits (230), Expect = 8e-18
Identities = 74/219 (33%), Positives = 112/219 (50%), Gaps = 25/219 (11%)
Query: 80 ISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNL 138
I L L LS+ N +GS+ + + +L FL L N+ +GS+P LRSL L+L
Sbjct: 163 IGYLRSLTKLSLGINFLSGSIPASVGNLNNLSFLYLYNNQLSGSIPEEISYLRSLTELDL 222
Query: 139 SLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLG 198
S N +G++P ++ L +L + N SG I E + S+ ++DLS N +G++
Sbjct: 223 SDNALNGSIPASLGNMNNLSFLFLYGNQLSGSIPEEICYLRSLTYLDLSENALNGSIPAS 282
Query: 199 LG---DVSFLF------------------SIQHLNVSHNSLVGELFAHDGMPYLDNLEVF 237
LG ++SFLF S+ L +S N+L G + A G L NL
Sbjct: 283 LGNLNNLSFLFLYGNQLSGSIPEEIGYLRSLNVLGLSENALNGSIPASLG--NLKNLSRL 340
Query: 238 DASNNQLVGNIP-SFTFVVSLRILRLACNQLTGSLPETL 275
+ NNQL G+IP S + +L +L L NQL+GS+P +L
Sbjct: 341 NLVNNQLSGSIPASLGNLNNLSMLYLYNNQLSGSIPASL 379
Score = 91.7 bits (226), Expect = 2e-17
Identities = 70/212 (33%), Positives = 108/212 (50%), Gaps = 9/212 (4%)
Query: 79 AISNLPMLHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLN 137
++ NL L L + NN +GS+ I + SL +L L N NG +P SF +R+L L
Sbjct: 594 SLGNLNNLSMLYLYNNQLSGSIPEEIGYLSSLTYLSLGNNSLNGLIPASFGNMRNLQALI 653
Query: 138 LSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDL 197
L+ N G +P+ L LE L N+ G + + + ++ + +S+N FSG L
Sbjct: 654 LNDNNLIGEIPSSVCNLTSLEVLYMPRNNLKGKVPQCLGNISNLQVLSMSSNSFSGELP- 712
Query: 198 GLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIP-SFTFVVS 256
+S L S+Q L+ N+L G + G + +LEVFD NN+L G +P +F+ S
Sbjct: 713 --SSISNLTSLQILDFGRNNLEGAIPQCFG--NISSLEVFDMQNNKLSGTLPTNFSIGCS 768
Query: 257 LRILRLACNQLTGSLPETLLKESSMMLSELDL 288
L L L N+L +P +L ++ L LDL
Sbjct: 769 LISLNLHGNELEDEIPRSL--DNCKKLQVLDL 798
Score = 79.7 bits (195), Expect = 9e-14
Identities = 67/228 (29%), Positives = 107/228 (46%), Gaps = 8/228 (3%)
Query: 64 ITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNKFNGS 122
++L N SL G + N+ L L + +N+ G + + + SL+ L + N G
Sbjct: 628 LSLGNNSLNGLIP-ASFGNMRNLQALILNDNNLIGEIPSSVCNLTSLEVLYMPRNNLKGK 686
Query: 123 LPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVL 182
+P + +L L++S N FSG +P+ L L+ LDF N+ G I + F + S+
Sbjct: 687 VPQCLGNISNLQVLSMSSNSFSGELPSSISNLTSLQILDFGRNNLEGAIPQCFGNISSLE 746
Query: 183 HVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNN 242
D+ NNK SG L + S S+ LN+ N L E+ + L+V D +N
Sbjct: 747 VFDMQNNKLSGTLPT---NFSIGCSLISLNLHGNELEDEI--PRSLDNCKKLQVLDLGDN 801
Query: 243 QLVGNIPSFTFVV-SLRILRLACNQLTGSLPETLLKESSMMLSELDLS 289
QL P + + LR+LRL N+L G + + + L +DLS
Sbjct: 802 QLNDTFPMWLGTLPELRVLRLTSNKLHGPIRSSRAEIMFPDLRIIDLS 849
Score = 65.1 bits (157), Expect = 2e-09
Identities = 65/206 (31%), Positives = 94/206 (45%), Gaps = 34/206 (16%)
Query: 80 ISNLPMLHNLSVVNNHFTGSMLHISP---MKSLKFLDLSLNKFNGSLPPS-FVELRSLVY 135
+ LP L L + +N G + L+ +DLS N F+ LP S F L+ +
Sbjct: 811 LGTLPELRVLRLTSNKLHGPIRSSRAEIMFPDLRIIDLSRNAFSQDLPTSLFEHLKGMRT 870
Query: 136 LNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSF----SGDIMEIFYQMGSVLHVDLSNNKF 191
++ ++ E S Y ++ +S G +EI + +DLS+NKF
Sbjct: 871 VDKTMEEPS--------------YESYYDDSVVVVTKGLELEIVRILSLYTVIDLSSNKF 916
Query: 192 SGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIP-- 249
G + LGD L +I+ LNVSHN+L G + + L LE D S NQL G IP
Sbjct: 917 EGHIPSVLGD---LIAIRILNVSHNALQG--YIPSSLGSLSILESLDLSFNQLSGEIPQQ 971
Query: 250 --SFTFVVSLRILRLACNQLTGSLPE 273
S TF L L L+ N L G +P+
Sbjct: 972 LASLTF---LEFLNLSHNYLQGCIPQ 994
Score = 54.7 bits (130), Expect = 3e-06
Identities = 71/274 (25%), Positives = 120/274 (42%), Gaps = 52/274 (18%)
Query: 34 LNSWDSKSLESNGCPQNWYGILCSEGNVISITLDNASLVGEFNFLAISNLPMLHNLSVVN 93
L +D ++ + +G + I CS +IS+ L L E ++ N L L + +
Sbjct: 745 LEVFDMQNNKLSGTLPTNFSIGCS---LISLNLHGNELEDEIP-RSLDNCKKLQVLDLGD 800
Query: 94 NHFTGSM-LHISPMKSLKFLDLSLNKFNGSLPPSFVELR--SLVYLNLSLNEFSGTVPN- 149
N + + + + L+ L L+ NK +G + S E+ L ++LS N FS +P
Sbjct: 801 NQLNDTFPMWLGTLPELRVLRLTSNKLHGPIRSSRAEIMFPDLRIIDLSRNAFSQDLPTS 860
Query: 150 ----------VFHKLDQLEYLDFHSNSF----SGDIMEIFYQMGSVLHVDLSNNKFSGAL 195
V +++ Y ++ +S G +EI + +DLS+NKF G +
Sbjct: 861 LFEHLKGMRTVDKTMEEPSYESYYDDSVVVVTKGLELEIVRILSLYTVIDLSSNKFEGHI 920
Query: 196 DLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIPSFTFVV 255
LGD L +I+ LNVSHN+L G + + L +L + ++
Sbjct: 921 PSVLGD---LIAIRILNVSHNALQGYIPSS-----LGSLSILES---------------- 956
Query: 256 SLRILRLACNQLTGSLPETLLKESSMMLSELDLS 289
L L+ NQL+G +P+ L S L L+LS
Sbjct: 957 ----LDLSFNQLSGEIPQQL--ASLTFLEFLNLS 984
Score = 51.6 bits (122), Expect = 3e-05
Identities = 29/87 (33%), Positives = 45/87 (51%)
Query: 112 LDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDI 171
+DLS NKF G +P +L ++ LN+S N G +P+ L LE LD N SG+I
Sbjct: 909 IDLSSNKFEGHIPSVLGDLIAIRILNVSHNALQGYIPSSLGSLSILESLDLSFNQLSGEI 968
Query: 172 MEIFYQMGSVLHVDLSNNKFSGALDLG 198
+ + + ++LS+N G + G
Sbjct: 969 PQQLASLTFLEFLNLSHNYLQGCIPQG 995
Score = 42.4 bits (98), Expect = 0.016
Identities = 29/95 (30%), Positives = 47/95 (48%), Gaps = 11/95 (11%)
Query: 76 NFLAISNLPMLHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVY 135
+ +AI L + HN + + S+ +S ++SL DLS N+ +G +P L L +
Sbjct: 926 DLIAIRILNVSHN--ALQGYIPSSLGSLSILESL---DLSFNQLSGEIPQQLASLTFLEF 980
Query: 136 LNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGD 170
LNLS N G +P ++ F SNS+ G+
Sbjct: 981 LNLSHNYLQGCIP------QGPQFRTFESNSYEGN 1009
>ref|NP_914843.1| putative receptor-like protein [Oryza sativa (japonica
cultivar-group)] gi|33383178|dbj|BAC81207.1| putative
leucin-rich repeat protein kinase [Oryza sativa
(japonica cultivar-group)] gi|19386763|dbj|BAB86144.1|
putative extra sporogenous cells [Oryza sativa (japonica
cultivar-group)]
Length = 1294
Score = 118 bits (296), Expect = 2e-25
Identities = 95/303 (31%), Positives = 146/303 (47%), Gaps = 38/303 (12%)
Query: 12 NRDIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILCSEGNVISITLDNASL 71
+RDI L L+ I G + N +DS++ P +W GI C NV++I L + L
Sbjct: 24 SRDISTLFTLRDSITEGK-GFLRNWFDSETP-----PCSWSGITCIGHNVVAIDLSSVPL 77
Query: 72 -------VGEFNFL----------------AISNLPMLHNLSVVNNHFTGSM-LHISPMK 107
+G F L A+ NL L L + NN TG + + + +K
Sbjct: 78 YAPFPLCIGAFQSLVRLNFSGCGFSGELPEALGNLQNLQYLDLSNNELTGPIPISLYNLK 137
Query: 108 SLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSF 167
LK + L N +G L P+ +L+ L L++S+N SG++P L LE LD N+F
Sbjct: 138 MLKEMVLDYNSLSGQLSPAIAQLQHLTKLSISMNSISGSLPPDLGSLKNLELLDIKMNTF 197
Query: 168 SGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDG 227
+G I F + +LH D S N +G++ G+ ++ L + L++S NS G + G
Sbjct: 198 NGSIPATFGNLSCLLHFDASQNNLTGSIFPGITSLTNLLT---LDLSSNSFEGTIPREIG 254
Query: 228 MPYLDNLEVFDASNNQLVGNIP-SFTFVVSLRILRLACNQLTGSLPETLLKESSMMLSEL 286
L+NLE+ N L G IP + L++L L Q TG +P ++ SS L+EL
Sbjct: 255 Q--LENLELLILGKNDLTGRIPQEIGSLKQLKLLHLEECQFTGKIPWSISGLSS--LTEL 310
Query: 287 DLS 289
D+S
Sbjct: 311 DIS 313
Score = 87.8 bits (216), Expect = 3e-16
Identities = 69/242 (28%), Positives = 120/242 (49%), Gaps = 20/242 (8%)
Query: 55 LCSEGNVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLHISPMKSLKFLDL 114
+C ++ S+ L + +L G + A L L++++NH G + L L+L
Sbjct: 443 ICQANSLHSLLLHHNNLTGTIDE-AFKGCTNLTELNLLDNHIHGEVPGYLAELPLVTLEL 501
Query: 115 SLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEI 174
S NKF G LP E ++L+ ++LS NE +G +P KL L+ L +N G I +
Sbjct: 502 SQNKFAGMLPAELWESKTLLEISLSNNEITGPIPESIGKLSVLQRLHIDNNLLEGPIPQS 561
Query: 175 FYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNL 234
+ ++ ++ L N+ SG + L L + L + L++S+N+L G + + +L L
Sbjct: 562 VGDLRNLTNLSLRGNRLSGIIPLALFNCRKLAT---LDLSYNNLTGNI--PSAISHLTLL 616
Query: 235 EVFDASNNQLVGNIPS-------------FTFVVSLRILRLACNQLTGSLPETLLKESSM 281
+ S+NQL G+IP+ F+ +L L+ NQLTG +P T +K +M
Sbjct: 617 DSLILSSNQLSGSIPAEICVGFENEAHPDSEFLQHHGLLDLSYNQLTGQIP-TSIKNCAM 675
Query: 282 ML 283
++
Sbjct: 676 VM 677
Score = 73.9 bits (180), Expect = 5e-12
Identities = 60/230 (26%), Positives = 102/230 (44%), Gaps = 25/230 (10%)
Query: 82 NLPMLHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSL 140
NL L + N+ TGS+ I+ + +L LDLS N F G++P +L +L L L
Sbjct: 207 NLSCLLHFDASQNNLTGSIFPGITSLTNLLTLDLSSNSFEGTIPREIGQLENLELLILGK 266
Query: 141 NEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLG 200
N+ +G +P L QL+ L F+G I + S+ +D+S+N F L +G
Sbjct: 267 NDLTGRIPQEIGSLKQLKLLHLEECQFTGKIPWSISGLSSLTELDISDNNFDAELPSSMG 326
Query: 201 DVSFLF---------------------SIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDA 239
++ L + +N+S N+L+G + + L+ + F
Sbjct: 327 ELGNLTQLIAKNAGLSGNMPKELGNCKKLTVINLSFNALIGPI--PEEFADLEAIVSFFV 384
Query: 240 SNNQLVGNIPSFTFV-VSLRILRLACNQLTGSLPETLLKESSMMLSELDL 288
N+L G +P + + R +RL N+ +G LP L+ +E +L
Sbjct: 385 EGNKLSGRVPDWIQKWKNARSIRLGQNKFSGPLPVLPLQHLLSFAAESNL 434
Score = 72.0 bits (175), Expect = 2e-11
Identities = 70/249 (28%), Positives = 110/249 (44%), Gaps = 40/249 (16%)
Query: 79 AISNLPMLHNLSVVNNHFTGSM-----------LHISP--MKSLKFLDLSLNKFNGSLPP 125
AIS+L +L +L + +N +GS+ H ++ LDLS N+ G +P
Sbjct: 609 AISHLTLLDSLILSSNQLSGSIPAEICVGFENEAHPDSEFLQHHGLLDLSYNQLTGQIPT 668
Query: 126 SFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVD 185
S ++ LNL N +GT+P +L L ++ N F G ++ + + +
Sbjct: 669 SIKNCAMVMVLNLQGNLLNGTIPVELGELTNLTSINLSFNEFVGPMLPWSGPLVQLQGLI 728
Query: 186 LSNNKFSGALDLGLGDV--------------------SFLFS--IQHLNVSHNSLVG--E 221
LSNN G++ +G + S L + + HL+VS+N L G +
Sbjct: 729 LSNNHLDGSIPAKIGQILPKIAVLDLSSNALTGTLPQSLLCNNYLNHLDVSNNHLSGHIQ 788
Query: 222 LFAHDGMPYLDNLEVFDASNNQLVGNI-PSFTFVVSLRILRLACNQLTGSLPETLLKESS 280
DG Y L F++S+N G++ S + L L + N LTG LP L SS
Sbjct: 789 FSCPDGKEYSSTLLFFNSSSNHFSGSLDESISNFTQLSTLDIHNNSLTGRLPSALSDLSS 848
Query: 281 MMLSELDLS 289
L+ LDLS
Sbjct: 849 --LNYLDLS 855
Score = 63.5 bits (153), Expect = 7e-09
Identities = 63/223 (28%), Positives = 96/223 (42%), Gaps = 33/223 (14%)
Query: 94 NHFTGSMLHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFH- 152
N F+G L + P++ L N +GS+P + SL L L N +GT+ F
Sbjct: 411 NKFSGP-LPVLPLQHLLSFAAESNLLSGSIPSHICQANSLHSLLLHHNNLTGTIDEAFKG 469
Query: 153 --KLDQLEYLDFH--------------------SNSFSGDIMEIFYQMGSVLHVDLSNNK 190
L +L LD H N F+G + ++ ++L + LSNN+
Sbjct: 470 CTNLTELNLLDNHIHGEVPGYLAELPLVTLELSQNKFAGMLPAELWESKTLLEISLSNNE 529
Query: 191 FSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIPS 250
+G + +G +S L Q L++ +N L G + + L NL N+L G IP
Sbjct: 530 ITGPIPESIGKLSVL---QRLHIDNNLLEGPI--PQSVGDLRNLTNLSLRGNRLSGIIPL 584
Query: 251 FTF-VVSLRILRLACNQLTGSLPET---LLKESSMMLSELDLS 289
F L L L+ N LTG++P L S++LS LS
Sbjct: 585 ALFNCRKLATLDLSYNNLTGNIPSAISHLTLLDSLILSSNQLS 627
Score = 58.9 bits (141), Expect = 2e-07
Identities = 45/158 (28%), Positives = 72/158 (45%), Gaps = 8/158 (5%)
Query: 64 ITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLHISPMKS-LKFLDLSLNKFNG- 121
+ L N L G LP + L + +N TG++ + L LD+S N +G
Sbjct: 727 LILSNNHLDGSIPAKIGQILPKIAVLDLSSNALTGTLPQSLLCNNYLNHLDVSNNHLSGH 786
Query: 122 ---SLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQM 178
S P +L++ N S N FSG++ QL LD H+NS +G + +
Sbjct: 787 IQFSCPDGKEYSSTLLFFNSSSNHFSGSLDESISNFTQLSTLDIHNNSLTGRLPSALSDL 846
Query: 179 GSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHN 216
S+ ++DLS+N GA+ G+ ++ F + N S N
Sbjct: 847 SSLNYLDLSSNNLYGAIPCGICNI---FGLSFANFSGN 881
>dbj|BAB01126.1| receptor protein kinase [Arabidopsis thaliana]
gi|30689028|ref|NP_189443.2| leucine-rich repeat
transmembrane protein kinase, putative [Arabidopsis
thaliana]
Length = 1016
Score = 116 bits (290), Expect = 8e-25
Identities = 94/282 (33%), Positives = 141/282 (49%), Gaps = 16/282 (5%)
Query: 12 NRDIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILCSE--GNVISITLDNA 69
N D+ L+ K + NDPF L SW E + P +W + C+ VI ++LD
Sbjct: 34 NDDVLGLIVFKSDL-NDPFSH-LESWT----EDDNTPCSWSYVKCNPKTSRVIELSLDGL 87
Query: 70 SLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKFNGSLPPSFVE 129
+L G+ N I L L LS+ NN+FTG++ +S L+ LDLS N +G +P S
Sbjct: 88 ALTGKIN-RGIQKLQRLKVLSLSNNNFTGNINALSNNNHLQKLDLSHNNLSGQIPSSLGS 146
Query: 130 LRSLVYLNLSLNEFSGTV-PNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSN 188
+ SL +L+L+ N FSGT+ ++F+ L YL N G I ++ + ++LS
Sbjct: 147 ITSLQHLDLTGNSFSGTLSDDLFNNCSSLRYLSLSHNHLEGQIPSTLFRCSVLNSLNLSR 206
Query: 189 NKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNI 248
N+FSG G + L ++ L++S NSL G + G+ L NL+ NQ G +
Sbjct: 207 NRFSGNPSFVSG-IWRLERLRALDLSSNSLSGSIPL--GILSLHNLKELQLQRNQFSGAL 263
Query: 249 PS-FTFVVSLRILRLACNQLTGSLPETLLKESSMMLSELDLS 289
PS L + L+ N +G LP TL K S L+ D+S
Sbjct: 264 PSDIGLCPHLNRVDLSSNHFSGELPRTLQKLKS--LNHFDVS 303
Score = 103 bits (257), Expect = 6e-21
Identities = 88/257 (34%), Positives = 125/257 (48%), Gaps = 32/257 (12%)
Query: 55 LCSEGNVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLH-ISPMKSLKFLD 113
LC N + ++ ++ S GE + L L++ V NN +G I M L LD
Sbjct: 269 LCPHLNRVDLSSNHFS--GELP-RTLQKLKSLNHFDVSNNLLSGDFPPWIGDMTGLVHLD 325
Query: 114 LSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIME 173
S N+ G LP S LRSL LNLS N+ SG VP +L + N FSG+I +
Sbjct: 326 FSSNELTGKLPSSISNLRSLKDLNLSENKLSGEVPESLESCKELMIVQLKGNDFSGNIPD 385
Query: 174 IFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLF-SIQHLNVSHNSLVGE------LFAH- 225
F+ +G + +D S N +G++ G S LF S+ L++SHNSL G LF H
Sbjct: 386 GFFDLG-LQEMDFSGNGLTGSIPRGS---SRLFESLIRLDLSHNSLTGSIPGEVGLFIHM 441
Query: 226 ---------------DGMPYLDNLEVFDASNNQLVGNIPS-FTFVVSLRILRLACNQLTG 269
+ +L NL V D N+ L+G++P+ SL+IL+L N LTG
Sbjct: 442 RYLNLSWNHFNTRVPPEIEFLQNLTVLDLRNSALIGSVPADICESQSLQILQLDGNSLTG 501
Query: 270 SLPETLLKESSMMLSEL 286
S+PE + SS+ L L
Sbjct: 502 SIPEGIGNCSSLKLLSL 518
Score = 98.2 bits (243), Expect = 2e-19
Identities = 66/209 (31%), Positives = 111/209 (52%), Gaps = 12/209 (5%)
Query: 85 MLHNLSVVNNHFTGSMLHISP---MKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLN 141
+L++L++ N F+G+ +S ++ L+ LDLS N +GS+P + L +L L L N
Sbjct: 198 VLNSLNLSRNRFSGNPSFVSGIWRLERLRALDLSSNSLSGSIPLGILSLHNLKELQLQRN 257
Query: 142 EFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLGD 201
+FSG +P+ L +D SN FSG++ ++ S+ H D+SNN SG +GD
Sbjct: 258 QFSGALPSDIGLCPHLNRVDLSSNHFSGELPRTLQKLKSLNHFDVSNNLLSGDFPPWIGD 317
Query: 202 VSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIP-SFTFVVSLRIL 260
++ + HL+ S N L G+L + L +L+ + S N+L G +P S L I+
Sbjct: 318 MT---GLVHLDFSSNELTGKL--PSSISNLRSLKDLNLSENKLSGEVPESLESCKELMIV 372
Query: 261 RLACNQLTGSLPETLLKESSMMLSELDLS 289
+L N +G++P+ + L E+D S
Sbjct: 373 QLKGNDFSGNIPDGFF---DLGLQEMDFS 398
Score = 74.3 bits (181), Expect = 4e-12
Identities = 57/190 (30%), Positives = 92/190 (48%), Gaps = 8/190 (4%)
Query: 86 LHNLSVVNNHFTGSMLHISP--MKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEF 143
L + N TGS+ S +SL LDLS N GS+P + YLNLS N F
Sbjct: 392 LQEMDFSGNGLTGSIPRGSSRLFESLIRLDLSHNSLTGSIPGEVGLFIHMRYLNLSWNHF 451
Query: 144 SGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLGDVS 203
+ VP L L LD +++ G + + S+ + L N +G++ G+G+ S
Sbjct: 452 NTRVPPEIEFLQNLTVLDLRNSALIGSVPADICESQSLQILQLDGNSLTGSIPEGIGNCS 511
Query: 204 FLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIP-SFTFVVSLRILRL 262
S++ L++SHN+L G + + L L++ N+L G IP + +L ++ +
Sbjct: 512 ---SLKLLSLSHNNLTGPI--PKSLSNLQELKILKLEANKLSGEIPKELGDLQNLLLVNV 566
Query: 263 ACNQLTGSLP 272
+ N+L G LP
Sbjct: 567 SFNRLIGRLP 576
>dbj|BAA88636.1| elicitor-inducible LRR receptor-like protein EILP [Nicotiana
tabacum]
Length = 861
Score = 114 bits (286), Expect = 2e-24
Identities = 84/276 (30%), Positives = 138/276 (49%), Gaps = 34/276 (12%)
Query: 1 MLLLLVNTAFGNRDIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILCSEGN 60
+ L T ++ ALL+ K +QN L++ SW S C ++WYG++C G
Sbjct: 16 LFCLFTVTFASTKEATALLKWKATLQNQSNSLLV-SWTPSS---KAC-KSWYGVVCFNGR 70
Query: 61 VISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKFN 120
V + + A ++G N S+LP L +++DLS+N+
Sbjct: 71 VSKLDIPYAGVIGTLNNFPFSSLPFL-----------------------EYIDLSMNQLF 107
Query: 121 GSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGS 180
GS+PP +L +LVYL+LS N+ SGT+P L +L+ L N +G I + S
Sbjct: 108 GSIPPEIGKLTNLVYLDLSFNQISGTIPPQIGSLAKLQTLHILDNHLNGSIPGEIGHLRS 167
Query: 181 VLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDAS 240
+ +DLS N +G++ LG+ L ++ L + N++ G F + + YL +L D +
Sbjct: 168 LTELDLSINTLNGSIPPSLGN---LHNLSLLCLYKNNISG--FIPEEIGYLSSLIQLDLN 222
Query: 241 NNQLVGNIP-SFTFVVSLRILRLACNQLTGSLPETL 275
N L G+IP S + +L +L L NQL+GS+P+ +
Sbjct: 223 TNFLNGSIPASLENLHNLSLLYLYENQLSGSIPDEI 258
Score = 90.1 bits (222), Expect = 6e-17
Identities = 71/225 (31%), Positives = 113/225 (49%), Gaps = 11/225 (4%)
Query: 54 ILCSEGNVISIT---LDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSM-LHISPMKSL 109
I S GN+ S++ L++ L G I L L LS+ N GS+ + + + SL
Sbjct: 278 IPASLGNLTSLSILQLEHNQLSGSIPE-EIGYLRTLAVLSLYTNFLNGSIPISLGNLTSL 336
Query: 110 KFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSG 169
L L N +G +P S L +LVYL L N+ SG +P+ L L Y+ H N +G
Sbjct: 337 SSLSLYENHLSGPIPSSLGNLDNLVYLYLYANQLSGPIPSELGNLKNLNYMKLHDNQLNG 396
Query: 170 DIMEIFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMP 229
I F + ++ ++ L +N +G + L + + L S++ L++ NSL G++ +
Sbjct: 397 SIPASFGNLRNMQYLFLESNNLTGEIPLSICN---LMSLKVLSLGRNSLKGDIL--QCLI 451
Query: 230 YLDNLEVFDASNNQLVGNIP-SFTFVVSLRILRLACNQLTGSLPE 273
+ L+V +N L IP S + SLRIL L+ N L GS+P+
Sbjct: 452 NISRLQVLKIPDNNLSEEIPSSICNLTSLRILDLSRNNLKGSIPQ 496
Score = 78.6 bits (192), Expect = 2e-13
Identities = 65/211 (30%), Positives = 105/211 (48%), Gaps = 10/211 (4%)
Query: 81 SNLPMLHNLSVV---NNHFTGSML-HISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYL 136
++L LHNLS++ N +GS+ I +++L + L+ N GS+P S L SL L
Sbjct: 232 ASLENLHNLSLLYLYENQLSGSIPDEIGQLRTLTDIRLNTNFLTGSIPASLGNLTSLSIL 291
Query: 137 NLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALD 196
L N+ SG++P L L L ++N +G I + S+ + L N SG +
Sbjct: 292 QLEHNQLSGSIPEEIGYLRTLAVLSLYTNFLNGSIPISLGNLTSLSSLSLYENHLSGPIP 351
Query: 197 LGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIP-SFTFVV 255
LG+ L ++ +L + N L G + + G L NL +NQL G+IP SF +
Sbjct: 352 SSLGN---LDNLVYLYLYANQLSGPIPSELG--NLKNLNYMKLHDNQLNGSIPASFGNLR 406
Query: 256 SLRILRLACNQLTGSLPETLLKESSMMLSEL 286
+++ L L N LTG +P ++ S+ + L
Sbjct: 407 NMQYLFLESNNLTGEIPLSICNLMSLKVLSL 437
Score = 78.2 bits (191), Expect = 3e-13
Identities = 73/236 (30%), Positives = 113/236 (46%), Gaps = 27/236 (11%)
Query: 60 NVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSML----HISPMKSLKFLDLS 115
N+ + L++ +L GE L+I NL L LS+ N G +L +IS ++ LK D
Sbjct: 407 NMQYLFLESNNLTGEIP-LSICNLMSLKVLSLGRNSLKGDILQCLINISRLQVLKIPD-- 463
Query: 116 LNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLD-QLEYLDFHSNSFSGDIMEI 174
N + +P S L SL L+LS N G++P F + LE LD H N SG +
Sbjct: 464 -NNLSEEIPSSICNLTSLRILDLSRNNLKGSIPQCFGDMGGHLEVLDIHKNGISGTLPTT 522
Query: 175 FYQMGSVLH-VDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPY--- 230
F ++GSVL L N+ G + L + +Q L++ N L +D P
Sbjct: 523 F-RIGSVLRSFTLHENELEGKIPRSLANCK---ELQVLDLGDNLL------NDTFPMWLG 572
Query: 231 -LDNLEVFDASNNQLVGNIPSF---TFVVSLRILRLACNQLTGSLPETLLKESSMM 282
L L+V +N+L G+I + + LRI+ L+ N TG++P +L ++ M
Sbjct: 573 TLPKLQVLRLKSNKLYGSIRTSKDENMFLELRIINLSYNAFTGNIPTSLFQQLKAM 628
Score = 66.2 bits (160), Expect = 1e-09
Identities = 57/214 (26%), Positives = 90/214 (41%), Gaps = 6/214 (2%)
Query: 42 LESNGCPQNWYGILCSEGNVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSM- 100
LESN +C+ ++ ++L SL G+ + N+ L L + +N+ + +
Sbjct: 413 LESNNLTGEIPLSICNLMSLKVLSLGRNSLKGDI-LQCLINISRLQVLKIPDNNLSEEIP 471
Query: 101 LHISPMKSLKFLDLSLNKFNGSLPPSFVELRS-LVYLNLSLNEFSGTVPNVFHKLDQLEY 159
I + SL+ LDLS N GS+P F ++ L L++ N SGT+P F L
Sbjct: 472 SSICNLTSLRILDLSRNNLKGSIPQCFGDMGGHLEVLDIHKNGISGTLPTTFRIGSVLRS 531
Query: 160 LDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLV 219
H N G I + +DL +N + + LG L +Q L + N L
Sbjct: 532 FTLHENELEGKIPRSLANCKELQVLDLGDNLLNDTFPMWLGT---LPKLQVLRLKSNKLY 588
Query: 220 GELFAHDGMPYLDNLEVFDASNNQLVGNIPSFTF 253
G + L + + S N GNIP+ F
Sbjct: 589 GSIRTSKDENMFLELRIINLSYNAFTGNIPTSLF 622
Score = 64.3 bits (155), Expect = 4e-09
Identities = 58/220 (26%), Positives = 94/220 (42%), Gaps = 30/220 (13%)
Query: 79 AISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLN 137
++ NL L L + N +G + + +K+L ++ L N+ NGS+P SF LR++ YL
Sbjct: 353 SLGNLDNLVYLYLYANQLSGPIPSELGNLKNLNYMKLHDNQLNGSIPASFGNLRNMQYLF 412
Query: 138 LSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMG------------------ 179
L N +G +P L L+ L NS GDI++ +
Sbjct: 413 LESNNLTGEIPLSICNLMSLKVLSLGRNSLKGDILQCLINISRLQVLKIPDNNLSEEIPS 472
Query: 180 ------SVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDN 233
S+ +DLS N G++ GD+ ++ L++ N + G L + +
Sbjct: 473 SICNLTSLRILDLSRNNLKGSIPQCFGDMG--GHLEVLDIHKNGISGTLPTTFRIGSV-- 528
Query: 234 LEVFDASNNQLVGNIP-SFTFVVSLRILRLACNQLTGSLP 272
L F N+L G IP S L++L L N L + P
Sbjct: 529 LRSFTLHENELEGKIPRSLANCKELQVLDLGDNLLNDTFP 568
Score = 54.3 bits (129), Expect = 4e-06
Identities = 68/260 (26%), Positives = 112/260 (42%), Gaps = 62/260 (23%)
Query: 59 GNVI-SITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSL 116
G+V+ S TL L G+ +++N L L + +N + + + + L+ L L
Sbjct: 526 GSVLRSFTLHENELEGKIP-RSLANCKELQVLDLGDNLLNDTFPMWLGTLPKLQVLRLKS 584
Query: 117 NKFNGSLPPS-----FVELRSLVYLNLSLNEFSGTVPN-------VFHKLDQLEYLDFHS 164
NK GS+ S F+ELR +NLS N F+G +P K+DQ +
Sbjct: 585 NKLYGSIRTSKDENMFLELR---IINLSYNAFTGNIPTSLFQQLKAMRKIDQTVKEPTYL 641
Query: 165 NSFSGDIMEIFYQMGSV---------------LHVDLSNNKFSGALDLGLGDVSFLFSIQ 209
F DI E Y + + +DLS+N+F G + +G+ L +++
Sbjct: 642 GKFGADIREYNYSVTVTTKGLELKLVRILTVYIIIDLSSNRFEGHVPSIMGE---LIALR 698
Query: 210 HLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIPSFTFVVSLRILRLACNQLTG 269
LN+S N L G + P L NL V ++ L L+ NQL+G
Sbjct: 699 VLNLSRNGLQGHI-----PPSLGNLFVIES--------------------LDLSFNQLSG 733
Query: 270 SLPETLLKESSMMLSELDLS 289
+P+ + + + L+ L+LS
Sbjct: 734 EIPQQIASQLT-SLAVLNLS 752
Score = 52.0 bits (123), Expect = 2e-05
Identities = 33/88 (37%), Positives = 46/88 (51%), Gaps = 1/88 (1%)
Query: 112 LDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDI 171
+DLS N+F G +P EL +L LNLS N G +P L +E LD N SG+I
Sbjct: 676 IDLSSNRFEGHVPSIMGELIALRVLNLSRNGLQGHIPPSLGNLFVIESLDLSFNQLSGEI 735
Query: 172 -MEIFYQMGSVLHVDLSNNKFSGALDLG 198
+I Q+ S+ ++LS N G + G
Sbjct: 736 PQQIASQLTSLAVLNLSYNHLQGCIPQG 763
Score = 42.7 bits (99), Expect = 0.012
Identities = 28/87 (32%), Positives = 44/87 (50%), Gaps = 2/87 (2%)
Query: 134 VYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSG 193
+ ++LS N F G VP++ +L L L+ N G I + + +DLS N+ SG
Sbjct: 674 IIIDLSSNRFEGHVPSIMGELIALRVLNLSRNGLQGHIPPSLGNLFVIESLDLSFNQLSG 733
Query: 194 ALDLGLGDVSFLFSIQHLNVSHNSLVG 220
+ + S L S+ LN+S+N L G
Sbjct: 734 EIPQQI--ASQLTSLAVLNLSYNHLQG 758
>gb|AAF26131.1| putative disease resistance protein [Arabidopsis thaliana]
gi|15230024|ref|NP_187217.1| disease resistance family
protein [Arabidopsis thaliana]
Length = 883
Score = 114 bits (285), Expect = 3e-24
Identities = 97/268 (36%), Positives = 133/268 (49%), Gaps = 20/268 (7%)
Query: 16 DALLELKKG--IQNDPFGLVLNSWDSKSLESNGCPQNWYGILCSE--GNVISITLDNASL 71
DALLE K I+ FG + +KS E+ +W GI C G VI I L + L
Sbjct: 36 DALLEFKNEFKIKKPCFGCP-SPLKTKSWENGSDCCHWDGITCDAKTGEVIEIDLMCSCL 94
Query: 72 VGEFNFLAISNLPMLHN------LSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKFNGSLP 124
G F+ + SNL ML N L + NH +G + I + L LDLS N F+G +P
Sbjct: 95 HGWFH--SNSNLSMLQNFHFLTTLDLSYNHLSGQISSSIGNLSHLTTLDLSGNNFSGWIP 152
Query: 125 PSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHV 184
S L L L+L N F G +P+ L L +LD +N+F G+I F + + +
Sbjct: 153 SSLGNLFHLTSLHLYDNNFGGEIPSSLGNLSYLTFLDLSTNNFVGEIPSSFGSLNQLSIL 212
Query: 185 DLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQL 244
L NNK SG L L +V L + +++SHN G L + L LE F AS N
Sbjct: 213 RLDNNKLSGNLPL---EVINLTKLSEISLSHNQFTGTL--PPNITSLSILESFSASGNNF 267
Query: 245 VGNIPSFTFVV-SLRILRLACNQLTGSL 271
VG IPS F + S+ ++ L NQL+G+L
Sbjct: 268 VGTIPSSLFTIPSITLIFLDNNQLSGTL 295
Score = 74.3 bits (181), Expect = 4e-12
Identities = 61/214 (28%), Positives = 100/214 (46%), Gaps = 7/214 (3%)
Query: 79 AISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLN 137
++ NL L L + N+F G + + L L L NK +G+LP + L L ++
Sbjct: 178 SLGNLSYLTFLDLSTNNFVGEIPSSFGSLNQLSILRLDNNKLSGNLPLEVINLTKLSEIS 237
Query: 138 LSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDL 197
LS N+F+GT+P L LE N+F G I + + S+ + L NN+ SG L+
Sbjct: 238 LSHNQFTGTLPPNITSLSILESFSASGNNFVGTIPSSLFTIPSITLIFLDNNQLSGTLE- 296
Query: 198 GLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIP--SFTFVV 255
G++S ++ L + N+L G + + L NL D S+ + G + F+ +
Sbjct: 297 -FGNISSPSNLLVLQLGGNNLRGPI--PTSISRLVNLRTLDLSHFNIQGQVDFNIFSHLK 353
Query: 256 SLRILRLACNQLTGSLPETLLKESSMMLSELDLS 289
L L L+ + T ++ + ML LDLS
Sbjct: 354 LLGNLYLSHSNTTTTIDLNAVLSCFKMLISLDLS 387
Score = 74.3 bits (181), Expect = 4e-12
Identities = 76/279 (27%), Positives = 123/279 (43%), Gaps = 33/279 (11%)
Query: 18 LLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWY-GILCSEGNVISITLDNASLVGEFN 76
L+ K + + P GL+ SL +GC + IL ++ + ++ + N + G+
Sbjct: 392 LVTNKSSVSDPPLGLI------GSLNLSGCGITEFPDILRTQRQMRTLDISNNKIKGQVP 445
Query: 77 FLAISNLPMLHNLSVVNNHFTGSMLH------ISPMKSLKFLDLSLNKFNGSLPPSFVEL 130
+ L +H + NN+F G + P S+K S N F+G +P L
Sbjct: 446 SWLLLQLEYMH---ISNNNFIGFERSTKLEKTVVPKPSMKHFFGSNNNFSGKIPSFICSL 502
Query: 131 RSLVYLNLSLNEFSGTVPNVFHKL-DQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNN 189
RSL+ L+LS N FSG +P K L L+ N SG + + + S+ +D+S+N
Sbjct: 503 RSLIILDLSNNNFSGAIPPCVGKFKSTLSDLNLRRNRLSGSLPKTIIK--SLRSLDVSHN 560
Query: 190 KFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPY----LDNLEVFDASNNQLV 245
+ G L L S +++ LNV N + +D P+ L L+V +N
Sbjct: 561 ELEGKLPRSLIHFS---TLEVLNVESNRI------NDTFPFWLSSLKKLQVLVLRSNAFH 611
Query: 246 GNIPSFTFVVSLRILRLACNQLTGSLPETLLKESSMMLS 284
G I F LRI+ ++ N G+LP E + M S
Sbjct: 612 GRIHKTRF-PKLRIIDISRNHFNGTLPSDCFVEWTGMHS 649
Score = 73.6 bits (179), Expect = 6e-12
Identities = 71/252 (28%), Positives = 118/252 (46%), Gaps = 24/252 (9%)
Query: 55 LCSEGNVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLHISPMKSLKFLDL 114
+CS ++I + L N + G L +L++ N +GS L + +KSL+ LD+
Sbjct: 499 ICSLRSLIILDLSNNNFSGAIPPCVGKFKSTLSDLNLRRNRLSGS-LPKTIIKSLRSLDV 557
Query: 115 SLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEI 174
S N+ G LP S + +L LN+ N + T P L +L+ L SN+F G I +
Sbjct: 558 SHNELEGKLPRSLIHFSTLEVLNVESNRINDTFPFWLSSLKKLQVLVLRSNAFHGRIHKT 617
Query: 175 FYQMGSVLHVDLSNNKFSGALDLG-LGDVSFLFSIQHLNVSHN-SLVGELFAHDGMPYLD 232
+ + +D+S N F+G L + + + S++ N +G + HD M ++
Sbjct: 618 RFPKLRI--IDISRNHFNGTLPSDCFVEWTGMHSLEKNEDRFNEKYMGSGYYHDSMVLMN 675
Query: 233 N---------LEVF---DASNNQLVGNIP-SFTFVVSLRILRLACNQLTGSLPETL--LK 277
L+++ D S N+ G IP S + L IL L+ N TG +P ++ L+
Sbjct: 676 KGLEMELVRILKIYTALDFSGNKFEGEIPRSIGLLKELHILNLSSNGFTGHIPSSMGNLR 735
Query: 278 ESSMMLSELDLS 289
E L LD+S
Sbjct: 736 E----LESLDVS 743
Score = 62.0 bits (149), Expect = 2e-08
Identities = 56/192 (29%), Positives = 88/192 (45%), Gaps = 25/192 (13%)
Query: 80 ISNLPMLHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKFNGSLPPS-FVELRSLVYLNL 138
+S+L L L + +N F G +H + L+ +D+S N FNG+LP FVE + L
Sbjct: 594 LSSLKKLQVLVLRSNAFHGR-IHKTRFPKLRIIDISRNHFNGTLPSDCFVEWTGMHSLEK 652
Query: 139 SLNEFS------------------GTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGS 180
+ + F+ G + L LDF N F G+I +
Sbjct: 653 NEDRFNEKYMGSGYYHDSMVLMNKGLEMELVRILKIYTALDFSGNKFEGEIPRSIGLLKE 712
Query: 181 VLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDAS 240
+ ++LS+N F+G + +G+ L ++ L+VS N L GE+ G L L + S
Sbjct: 713 LHILNLSSNGFTGHIPSSMGN---LRELESLDVSRNKLSGEIPQELGN--LSYLAYMNFS 767
Query: 241 NNQLVGNIPSFT 252
+NQLVG +P T
Sbjct: 768 HNQLVGQVPGGT 779
>ref|NP_198058.1| disease resistance family protein [Arabidopsis thaliana]
gi|5732036|gb|AAD48937.1| similar to disease resistance
proteins; contains similarity ot Pfam family PF00560 -
Leucine Rich Repeat; score=166.7, E=4e-46, N=24
[Arabidopsis thaliana]
Length = 957
Score = 113 bits (282), Expect = 7e-24
Identities = 88/274 (32%), Positives = 133/274 (48%), Gaps = 36/274 (13%)
Query: 38 DSKSLESNGCPQNWYGILCS--EGNVISITLDNASLVGEFN---------FL-------- 78
DS S+ C NW G+ C+ G VI + L +SL G F+ FL
Sbjct: 74 DSWGNNSDCC--NWEGVTCNAKSGEVIELDLSCSSLHGRFHSNSSIRNLHFLTTLDLSFN 131
Query: 79 --------AISNLPMLHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKFNGSLPPSFVE 129
+I NL L L + +NHF+G +L+ I + L +L+L N+F+G P S
Sbjct: 132 DFKGQITSSIENLSHLTYLDLSSNHFSGQILNSIGNLSRLTYLNLFDNQFSGQAPSSICN 191
Query: 130 LRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNN 189
L L +L+LS N F G P+ L L L SN FSG I + ++ +DLSNN
Sbjct: 192 LSHLTFLDLSYNRFFGQFPSSIGGLSHLTTLSLFSNKFSGQIPSSIGNLSNLTTLDLSNN 251
Query: 190 KFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIP 249
FSG + +G++S + L + N+ VGE+ + G L+ L +N+L GN P
Sbjct: 252 NFSGQIPSFIGNLS---QLTFLGLFSNNFVGEIPSSFG--NLNQLTRLYVDDNKLSGNFP 306
Query: 250 SFTF-VVSLRILRLACNQLTGSLPETLLKESSMM 282
+ + L +L L+ N+ TG+LP + S++M
Sbjct: 307 NVLLNLTGLSLLSLSNNKFTGTLPPNITSLSNLM 340
Score = 84.0 bits (206), Expect = 5e-15
Identities = 60/177 (33%), Positives = 85/177 (47%), Gaps = 13/177 (7%)
Query: 79 AISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLN 137
+I NL L L + NN+F+G + I + L FL L N F G +P SF L L L
Sbjct: 236 SIGNLSNLTTLDLSNNNFSGQIPSFIGNLSQLTFLGLFSNNFVGEIPSSFGNLNQLTRLY 295
Query: 138 LSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDL 197
+ N+ SG PNV L L L +N F+G + + +++ D S+N F+G
Sbjct: 296 VDDNKLSGNFPNVLLNLTGLSLLSLSNNKFTGTLPPNITSLSNLMDFDASDNAFTGTFP- 354
Query: 198 GLGDVSFLF---SIQHLNVSHNSLVGEL-FAHDGMPYLDNLEVFDASNNQLVGNIPS 250
SFLF S+ ++ ++ N L G L F + P NL D NN +G IPS
Sbjct: 355 -----SFLFTIPSLTYIRLNGNQLKGTLEFGNISSP--SNLYELDIGNNNFIGPIPS 404
Score = 75.5 bits (184), Expect = 2e-12
Identities = 70/227 (30%), Positives = 107/227 (46%), Gaps = 31/227 (13%)
Query: 74 EFNFLAISN-------------LPMLHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKFN 120
E FL ISN LP+L+ +++ NN G P SL +L S N F
Sbjct: 511 ELGFLDISNNKIKGQVPDWLWRLPILYYVNLSNNTLIGFQRPSKPEPSLLYLLGSNNNFI 570
Query: 121 GSLPPSFVELRSLVYLNLSLNEFSGTVPNVF-HKLDQLEYLDFHSNSFSGDIMEIFYQMG 179
G +P LRSL L+LS N F+G++P H L L+ N SG + + +++
Sbjct: 571 GKIPSFICGLRSLNTLDLSDNNFNGSIPRCMGHLKSTLSVLNLRQNHLSGGLPKQIFEI- 629
Query: 180 SVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPY----LDNLE 235
+ +D+ +N+ G L L SF +++ LNV N + +D P+ L L+
Sbjct: 630 -LRSLDVGHNQLVGKLPRSL---SFFSTLEVLNVESNRI------NDTFPFWLSSLPKLQ 679
Query: 236 VFDASNNQLVGNIPSFTFVVSLRILRLACNQLTGSLP-ETLLKESSM 281
V +N G I TF LRI+ ++ N+ G+LP E +K S+M
Sbjct: 680 VLVLRSNAFHGPIHEATFP-ELRIIDISHNRFNGTLPTEYFVKWSAM 725
Score = 65.9 bits (159), Expect = 1e-09
Identities = 58/219 (26%), Positives = 95/219 (42%), Gaps = 54/219 (24%)
Query: 80 ISNLPMLHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKFNGSLPPS-FVELR------- 131
+S+LP L L + +N F G +H + L+ +D+S N+FNG+LP FV+
Sbjct: 672 LSSLPKLQVLVLRSNAFHGP-IHEATFPELRIIDISHNRFNGTLPTEYFVKWSAMSSLGK 730
Query: 132 ------------------SLVYLN------------------LSLNEFSGTVPNVFHKLD 155
S+V +N S N F G +P L
Sbjct: 731 NEDQSNEKYMGSGLYYQDSMVLMNKGVAMELVRILTIYTAVDFSGNRFEGEIPKSIGLLK 790
Query: 156 QLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSH 215
+L L +N+FSG + + ++ +D+S NK +G + LGD+SFL ++N SH
Sbjct: 791 ELLVLSLSNNAFSGHMPSSMGNLTALESLDVSKNKLTGEIPQELGDLSFL---AYMNFSH 847
Query: 216 NSLVG------ELFAHDGMPYLDNLEVFDASNNQLVGNI 248
N L G + + + DNL +F +S ++ +I
Sbjct: 848 NQLAGLVPGGQQFLTQNCSAFEDNLGLFGSSLEEVCRDI 886
Score = 56.6 bits (135), Expect = 8e-07
Identities = 58/225 (25%), Positives = 105/225 (45%), Gaps = 22/225 (9%)
Query: 64 ITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSM---LHISPMKSLKFLDLS----- 115
+ + + + G +F S+L L +L++ + + T + +S K L LDLS
Sbjct: 415 LDISHLNTQGPVDFSIFSHLKSLLDLNISHLNTTTRIDLNYFLSYFKRLLLLDLSGNHVS 474
Query: 116 -LNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEI 174
NK + S PPS + ++SL + EF P +L +LD +N G + +
Sbjct: 475 ATNKSSVSDPPSQL-IQSLYLSGCGITEF----PEFVRTQHELGFLDISNNKIKGQVPDW 529
Query: 175 FYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNL 234
+++ + +V+LSNN G + S L+ L S+N+ +G++ + + L +L
Sbjct: 530 LWRLPILYYVNLSNNTLIGFQRPSKPEPSLLY----LLGSNNNFIGKIPSF--ICGLRSL 583
Query: 235 EVFDASNNQLVGNIPSFT--FVVSLRILRLACNQLTGSLPETLLK 277
D S+N G+IP +L +L L N L+G LP+ + +
Sbjct: 584 NTLDLSDNNFNGSIPRCMGHLKSTLSVLNLRQNHLSGGLPKQIFE 628
Score = 56.2 bits (134), Expect = 1e-06
Identities = 63/232 (27%), Positives = 96/232 (41%), Gaps = 50/232 (21%)
Query: 89 LSVVN---NHFTGSMLHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSG 145
LSV+N NH +G L + L+ LD+ N+ G LP S +L LN+ N +
Sbjct: 608 LSVLNLRQNHLSGG-LPKQIFEILRSLDVGHNQLVGKLPRSLSFFSTLEVLNVESNRIND 666
Query: 146 TVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGAL---------- 195
T P L +L+ L SN+F G I E + + +D+S+N+F+G L
Sbjct: 667 TFPFWLSSLPKLQVLVLRSNAFHGPIHEATFPELRI--IDISHNRFNGTLPTEYFVKWSA 724
Query: 196 --DLGLGD-----------------------------VSFLFSIQHLNVSHNSLVGELFA 224
LG + V L ++ S N GE+
Sbjct: 725 MSSLGKNEDQSNEKYMGSGLYYQDSMVLMNKGVAMELVRILTIYTAVDFSGNRFEGEIPK 784
Query: 225 HDGMPYLDNLEVFDASNNQLVGNIPS-FTFVVSLRILRLACNQLTGSLPETL 275
G+ L L V SNN G++PS + +L L ++ N+LTG +P+ L
Sbjct: 785 SIGL--LKELLVLSLSNNAFSGHMPSSMGNLTALESLDVSKNKLTGEIPQEL 834
Score = 47.4 bits (111), Expect = 5e-04
Identities = 68/276 (24%), Positives = 109/276 (38%), Gaps = 62/276 (22%)
Query: 55 LCSEGNVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSML--HISPMKSLKFL 112
+ S N++ + + G F + +P L + + N G++ +IS +L L
Sbjct: 333 ITSLSNLMDFDASDNAFTGTFPSFLFT-IPSLTYIRLNGNQLKGTLEFGNISSPSNLYEL 391
Query: 113 DLSLNKFNGSLPPS-------------------------FVELRSLVYLNLS-LNEFSGT 146
D+ N F G +P S F L+SL+ LN+S LN +
Sbjct: 392 DIGNNNFIGPIPSSISKLVKLFRLDISHLNTQGPVDFSIFSHLKSLLDLNISHLNTTTRI 451
Query: 147 VPNVF-HKLDQLEYLDFHSNSFS-----------GDIMEIFYQMGSVL------------ 182
N F +L LD N S +++ Y G +
Sbjct: 452 DLNYFLSYFKRLLLLDLSGNHVSATNKSSVSDPPSQLIQSLYLSGCGITEFPEFVRTQHE 511
Query: 183 --HVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDAS 240
+D+SNNK G + L + L+ ++N+S+N+L+G F P L + S
Sbjct: 512 LGFLDISNNKIKGQVPDWLWRLPILY---YVNLSNNTLIG--FQRPSKPEPSLLYLL-GS 565
Query: 241 NNQLVGNIPSFTF-VVSLRILRLACNQLTGSLPETL 275
NN +G IPSF + SL L L+ N GS+P +
Sbjct: 566 NNNFIGKIPSFICGLRSLNTLDLSDNNFNGSIPRCM 601
>gb|AAX14781.1| RLP1 leucine-rich repeat receptor-like protein [Medicago
truncatula]
Length = 671
Score = 110 bits (276), Expect = 4e-23
Identities = 91/314 (28%), Positives = 153/314 (47%), Gaps = 39/314 (12%)
Query: 1 MLLLLVNTAFG-NRDIDALLELKKGIQNDPF-GLVLNSWDSKSLESNGCPQNWYGILCS- 57
+L +L T + N D+DALL+LKK ++ + L W + S C ++ G+ C
Sbjct: 10 LLCMLFTTCYSLNNDLDALLKLKKSMKGEKAKDDALKDWKFSTSASGHC--SFSGVKCDG 67
Query: 58 EGNVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSM------------LHISP 105
E VI++ + L G + I L ML +L++ ++ TG + L+IS
Sbjct: 68 EQRVIALNVTQVPLFGHLS-KEIGELNMLESLTITMDNLTGELPTELSKLTSLRILNISH 126
Query: 106 --------------MKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVF 151
MK L+ LD N F G LP V L L YL+ + N FSGT+P +
Sbjct: 127 NLFSGNFPGNITFGMKKLEALDAYDNNFEGPLPEEIVSLMKLKYLSFAGNFFSGTIPESY 186
Query: 152 HKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLS-NNKFSGALDLGLGDVSFLFSIQH 210
+ +LE L + NS +G I + ++ + + L +N ++G + G + S+++
Sbjct: 187 SEFQKLEILRLNYNSLTGKIPKSLSKLKKLKELCLGYDNAYAGGIPPEFGSIK---SLRY 243
Query: 211 LNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNI-PSFTFVVSLRILRLACNQLTG 269
L++S+++L GE+ + L+NL+ N L G I P + + SL +L L+ N+L+G
Sbjct: 244 LDISNSNLTGEI--PPSLGNLENLDYLFLQMNYLTGKIPPELSSMRSLMMLDLSINELSG 301
Query: 270 SLPETLLKESSMML 283
+PET K + L
Sbjct: 302 EIPETFSKLKHLTL 315
Score = 79.0 bits (193), Expect = 1e-13
Identities = 67/245 (27%), Positives = 116/245 (47%), Gaps = 23/245 (9%)
Query: 64 ITLDNASLVGEFNFLAISNLPMLHNLSV-VNNHFTGSML-HISPMKSLKFLDLSLNKFNG 121
+ L+ SL G+ ++S L L L + +N + G + +KSL++LD+S + G
Sbjct: 195 LRLNYNSLTGKIP-KSLSKLKKLKELCLGYDNAYAGGIPPEFGSIKSLRYLDISNSNLTG 253
Query: 122 SLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSV 181
+PPS L +L YL L +N +G +P + L LD N SG+I E F ++ +
Sbjct: 254 EIPPSLGNLENLDYLFLQMNYLTGKIPPELSSMRSLMMLDLSINELSGEIPETFSKLKHL 313
Query: 182 LHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDG-MPYLD-------- 232
++ NK G++ +GD+ L ++Q + + +S++ + +G Y D
Sbjct: 314 TLINFFQNKLCGSIPAFVGDLPNLETLQVWDNNFSSVLPQNLGSNGKFIYFDVTKNHLTG 373
Query: 233 ----------NLEVFDASNNQLVGNIPS-FTFVVSLRILRLACNQLTGSLPETLLKESSM 281
L+ F S+N L G IP+ SL +R+A N L G +P + + S+
Sbjct: 374 LIPPELCKSKKLKTFIVSDNFLSGPIPNGIGACKSLEKIRVANNYLDGLVPPGIFQLPSV 433
Query: 282 MLSEL 286
+ EL
Sbjct: 434 TMMEL 438
Score = 76.6 bits (187), Expect = 7e-13
Identities = 53/171 (30%), Positives = 82/171 (46%), Gaps = 5/171 (2%)
Query: 80 ISNLPMLHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLS 139
I LP + + + NN F G + SL L LS N F G + S LRSL L L
Sbjct: 427 IFQLPSVTMMELRNNRFNGQLPSEISGNSLGILALSNNLFTGRISASMKNLRSLQTLLLD 486
Query: 140 LNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGL 199
N+F G +P L L ++ N+ +G I + Q ++ VD S N +G + G+
Sbjct: 487 ANQFVGEIPTEVFALPVLTRINISGNNLTGGIPKTVTQCSTLTAVDFSLNMLTGEVPKGM 546
Query: 200 GDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIPS 250
++ L LNVSHNS+ G++ + + ++ +L D S N G +P+
Sbjct: 547 KNLKVL---NILNVSHNSISGQI--PNDIRFMMSLTTLDLSYNNFTGIVPT 592
Score = 76.6 bits (187), Expect = 7e-13
Identities = 57/212 (26%), Positives = 102/212 (47%), Gaps = 8/212 (3%)
Query: 80 ISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNL 138
+ +LP L L V +N+F+ + ++ + D++ N G +PP + + L +
Sbjct: 331 VGDLPNLETLQVWDNNFSSVLPQNLGSNGKFIYFDVTKNHLTGLIPPELCKSKKLKTFIV 390
Query: 139 SLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLG 198
S N SG +PN LE + +N G + +Q+ SV ++L NN+F+G L
Sbjct: 391 SDNFLSGPIPNGIGACKSLEKIRVANNYLDGLVPPGIFQLPSVTMMELRNNRFNGQLPSE 450
Query: 199 LGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIPSFTFVVS-L 257
+ S+ L +S+N G + A M L +L+ NQ VG IP+ F + L
Sbjct: 451 ISG----NSLGILALSNNLFTGRISA--SMKNLRSLQTLLLDANQFVGEIPTEVFALPVL 504
Query: 258 RILRLACNQLTGSLPETLLKESSMMLSELDLS 289
+ ++ N LTG +P+T+ + S++ + L+
Sbjct: 505 TRINISGNNLTGGIPKTVTQCSTLTAVDFSLN 536
Score = 56.6 bits (135), Expect = 8e-07
Identities = 42/142 (29%), Positives = 65/142 (45%), Gaps = 3/142 (2%)
Query: 59 GNVISI-TLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSL 116
GN + I L N G + ++ NL L L + N F G + + + L +++S
Sbjct: 453 GNSLGILALSNNLFTGRIS-ASMKNLRSLQTLLLDANQFVGEIPTEVFALPVLTRINISG 511
Query: 117 NKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFY 176
N G +P + + +L ++ SLN +G VP L L L+ NS SG I
Sbjct: 512 NNLTGGIPKTVTQCSTLTAVDFSLNMLTGEVPKGMKNLKVLNILNVSHNSISGQIPNDIR 571
Query: 177 QMGSVLHVDLSNNKFSGALDLG 198
M S+ +DLS N F+G + G
Sbjct: 572 FMMSLTTLDLSYNNFTGIVPTG 593
>gb|AAP04025.1| putative disease resistance protein [Arabidopsis thaliana]
gi|26450219|dbj|BAC42228.1| putative disease resistance
protein [Arabidopsis thaliana]
gi|15222979|ref|NP_172844.1| leucine-rich repeat family
protein [Arabidopsis thaliana]
Length = 330
Score = 110 bits (275), Expect = 5e-23
Identities = 91/289 (31%), Positives = 138/289 (47%), Gaps = 16/289 (5%)
Query: 2 LLLLVNTAFGN---RDIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILCSE 58
LLLL N +F RD+ AL E+KK + ++ SW +G W G+ CS+
Sbjct: 17 LLLLFNVSFAKTLKRDMKALNEIKKLVG----WRLVYSWVGDDPCGDGVLPPWSGVTCSK 72
Query: 59 GN----VISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSML-HISPMKSLKFLD 113
V+ + + + S+VG F AI+ L L L + NN TG + I +K L L+
Sbjct: 73 VGDYRVVVKLEVYSMSIVGNFP-KAITKLLDLTVLDMHNNKLTGPIPPEIGRLKRLITLN 131
Query: 114 LSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIME 173
L NK +LPP L+SL YL LS N F G +P L +L+YL N F+G I
Sbjct: 132 LRWNKLQQALPPEIGGLKSLTYLYLSFNNFKGEIPKELANLHELQYLHIQENHFTGRIPA 191
Query: 174 IFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDN 233
+ + H+D NN G++ ++++L +++N L G L + + L N
Sbjct: 192 ELGTLQKLRHLDAGNNNLVGSISDLFRIEGCFPALRNLFLNNNYLTGGL--PNKLANLTN 249
Query: 234 LEVFDASNNQLVGNIP-SFTFVVSLRILRLACNQLTGSLPETLLKESSM 281
LE+ S N++ G IP + + L L L N GS+PE K ++
Sbjct: 250 LEILYLSFNKMTGAIPAALASIPRLTNLHLDHNLFNGSIPEAFYKHPNL 298
Score = 60.8 bits (146), Expect = 4e-08
Identities = 39/127 (30%), Positives = 67/127 (52%), Gaps = 7/127 (5%)
Query: 68 NASLVGEFN--FLAISNLPMLHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKFNGSLP 124
N +LVG + F P L NL + NN+ TG + + ++ + +L+ L LS NK G++P
Sbjct: 206 NNNLVGSISDLFRIEGCFPALRNLFLNNNYLTGGLPNKLANLTNLEILYLSFNKMTGAIP 265
Query: 125 PSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHV 184
+ + L L+L N F+G++P F+K L+ + N+F D+ I G+ +
Sbjct: 266 AALASIPRLTNLHLDHNLFNGSIPEAFYKHPNLKDMYIEGNAFKSDVKAI----GAHKVL 321
Query: 185 DLSNNKF 191
+LS+ F
Sbjct: 322 ELSDTDF 328
>gb|AAF79397.1| F16A14.12 [Arabidopsis thaliana] gi|25402771|pir||B86272 protein
F16A14.12 [imported] - Arabidopsis thaliana
Length = 383
Score = 110 bits (275), Expect = 5e-23
Identities = 91/289 (31%), Positives = 138/289 (47%), Gaps = 16/289 (5%)
Query: 2 LLLLVNTAFGN---RDIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILCSE 58
LLLL N +F RD+ AL E+KK + ++ SW +G W G+ CS+
Sbjct: 70 LLLLFNVSFAKTLKRDMKALNEIKKLVG----WRLVYSWVGDDPCGDGVLPPWSGVTCSK 125
Query: 59 GN----VISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSML-HISPMKSLKFLD 113
V+ + + + S+VG F AI+ L L L + NN TG + I +K L L+
Sbjct: 126 VGDYRVVVKLEVYSMSIVGNFP-KAITKLLDLTVLDMHNNKLTGPIPPEIGRLKRLITLN 184
Query: 114 LSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIME 173
L NK +LPP L+SL YL LS N F G +P L +L+YL N F+G I
Sbjct: 185 LRWNKLQQALPPEIGGLKSLTYLYLSFNNFKGEIPKELANLHELQYLHIQENHFTGRIPA 244
Query: 174 IFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDN 233
+ + H+D NN G++ ++++L +++N L G L + + L N
Sbjct: 245 ELGTLQKLRHLDAGNNNLVGSISDLFRIEGCFPALRNLFLNNNYLTGGL--PNKLANLTN 302
Query: 234 LEVFDASNNQLVGNIP-SFTFVVSLRILRLACNQLTGSLPETLLKESSM 281
LE+ S N++ G IP + + L L L N GS+PE K ++
Sbjct: 303 LEILYLSFNKMTGAIPAALASIPRLTNLHLDHNLFNGSIPEAFYKHPNL 351
Score = 60.8 bits (146), Expect = 4e-08
Identities = 39/127 (30%), Positives = 67/127 (52%), Gaps = 7/127 (5%)
Query: 68 NASLVGEFN--FLAISNLPMLHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKFNGSLP 124
N +LVG + F P L NL + NN+ TG + + ++ + +L+ L LS NK G++P
Sbjct: 259 NNNLVGSISDLFRIEGCFPALRNLFLNNNYLTGGLPNKLANLTNLEILYLSFNKMTGAIP 318
Query: 125 PSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHV 184
+ + L L+L N F+G++P F+K L+ + N+F D+ I G+ +
Sbjct: 319 AALASIPRLTNLHLDHNLFNGSIPEAFYKHPNLKDMYIEGNAFKSDVKAI----GAHKVL 374
Query: 185 DLSNNKF 191
+LS+ F
Sbjct: 375 ELSDTDF 381
>gb|AAW71475.1| CLV1-like receptor kinase [Medicago truncatula]
Length = 974
Score = 110 bits (275), Expect = 5e-23
Identities = 96/317 (30%), Positives = 151/317 (47%), Gaps = 45/317 (14%)
Query: 1 MLLLLVNTAFG-NRDIDALLELKKGIQNDPF-GLVLNSWDSKSLESNGCPQNWYGILCSE 58
+L +L T + N D+DALL+LKK ++ + L W + S C ++ G+ C E
Sbjct: 10 LLCMLFTTCYSLNNDLDALLKLKKSMKGEKAKDDALKDWKFSTSASAHC--SFSGVKCDE 67
Query: 59 GN-VISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSM------------LHISP 105
VI++ + L G + I L ML +L++ ++ TG + L+IS
Sbjct: 68 DQRVIALNVTQVPLFGHLS-KEIGELNMLESLTITMDNLTGELPTELSKLTSLRILNISH 126
Query: 106 --------------MKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVF 151
MK L+ LD N F G LP V L L YL+ + N FSGT+P +
Sbjct: 127 NLFSGNFPGNITFGMKKLEALDAYDNNFEGPLPEEIVSLMKLKYLSFAGNFFSGTIPESY 186
Query: 152 HKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLS-NNKFSGALDLGLGDVSFLFSIQH 210
+ +LE L + NS +G I + ++ + + L N +SG + LG + S+++
Sbjct: 187 SEFQKLEILRLNYNSLTGKIPKSLSKLKMLKELQLGYENAYSGGIPPELGSIK---SLRY 243
Query: 211 LNVSHNSLVGELFAHDGMPYLDNLEVFDA---SNNQLVGNI-PSFTFVVSLRILRLACNQ 266
L +S+ +L GE+ P L NLE D+ N L G I P + + SL L L+ N
Sbjct: 244 LEISNANLTGEI-----PPSLGNLENLDSLFLQMNNLTGTIPPELSSMRSLMSLDLSING 298
Query: 267 LTGSLPETLLKESSMML 283
L+G +PET K ++ L
Sbjct: 299 LSGEIPETFSKLKNLTL 315
Score = 84.3 bits (207), Expect = 4e-15
Identities = 68/257 (26%), Positives = 122/257 (47%), Gaps = 40/257 (15%)
Query: 64 ITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSML-HISPMKSLKFLDLSLNKFNGS 122
+ + NA+L GE ++ NL L +L + N+ TG++ +S M+SL LDLS+N +G
Sbjct: 244 LEISNANLTGEIP-PSLGNLENLDSLFLQMNNLTGTIPPELSSMRSLMSLDLSINGLSGE 302
Query: 123 LPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVL 182
+P +F +L++L +N N+ G++P L LE L N+FS + + G +
Sbjct: 303 IPETFSKLKNLTLINFFQNKLRGSIPAFIGDLPNLETLQVWENNFSFVLPQNLGSNGKFI 362
Query: 183 HVDLSNNK------------------------FSGALDLGLGDVSFLFSIQHLNVSHNSL 218
+ D++ N F G + G+G S++ + V++N L
Sbjct: 363 YFDVTKNHLTGLIPPELCKSKKLKTFIVTDNFFRGPIPNGIGPCK---SLEKIRVANNYL 419
Query: 219 VGELFAHDGMPYLDNLEVFDASNNQLVGNIPSFTFVVSLRILRLACNQLTGSLP------ 272
G + G+ L ++++ + NN+ G +P+ SL L L+ N TG +P
Sbjct: 420 DGPV--PPGIFQLPSVQIIELGNNRFNGQLPTEISGNSLGNLALSNNLFTGRIPASMKNL 477
Query: 273 ---ETLLKESSMMLSEL 286
+TLL +++ L E+
Sbjct: 478 RSLQTLLLDANQFLGEI 494
Score = 80.5 bits (197), Expect = 5e-14
Identities = 55/171 (32%), Positives = 82/171 (47%), Gaps = 5/171 (2%)
Query: 80 ISNLPMLHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLS 139
I LP + + + NN F G + SL L LS N F G +P S LRSL L L
Sbjct: 427 IFQLPSVQIIELGNNRFNGQLPTEISGNSLGNLALSNNLFTGRIPASMKNLRSLQTLLLD 486
Query: 140 LNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGL 199
N+F G +P L L ++ N+ +G I + Q S+ VD S N +G + G+
Sbjct: 487 ANQFLGEIPAEVFALPVLTRINISGNNLTGGIPKTVTQCSSLTAVDFSRNMLTGEVPKGM 546
Query: 200 GDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIPS 250
++ L NVSHNS+ G++ D + ++ +L D S N G +P+
Sbjct: 547 KNLKVL---SIFNVSHNSISGKI--PDEIRFMTSLTTLDLSYNNFTGIVPT 592
Score = 77.8 bits (190), Expect = 3e-13
Identities = 66/250 (26%), Positives = 115/250 (45%), Gaps = 33/250 (13%)
Query: 64 ITLDNASLVGEFNFLAISNLPMLHNLSV-VNNHFTGSML-HISPMKSLKFLDLSLNKFNG 121
+ L+ SL G+ ++S L ML L + N ++G + + +KSL++L++S G
Sbjct: 195 LRLNYNSLTGKIP-KSLSKLKMLKELQLGYENAYSGGIPPELGSIKSLRYLEISNANLTG 253
Query: 122 SLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSV 181
+PPS L +L L L +N +GT+P + L LD N SG+I E F ++ ++
Sbjct: 254 EIPPSLGNLENLDSLFLQMNNLTGTIPPELSSMRSLMSLDLSINGLSGEIPETFSKLKNL 313
Query: 182 LHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASN 241
++ NK G++ +GD+ L ++Q + + ++ + +G FD +
Sbjct: 314 TLINFFQNKLRGSIPAFIGDLPNLETLQVWENNFSFVLPQNLGSNG-----KFIYFDVTK 368
Query: 242 NQLVGNIPS--------FTFVV-----------------SLRILRLACNQLTGSLPETLL 276
N L G IP TF+V SL +R+A N L G +P +
Sbjct: 369 NHLTGLIPPELCKSKKLKTFIVTDNFFRGPIPNGIGPCKSLEKIRVANNYLDGPVPPGIF 428
Query: 277 KESSMMLSEL 286
+ S+ + EL
Sbjct: 429 QLPSVQIIEL 438
Score = 76.3 bits (186), Expect = 1e-12
Identities = 56/204 (27%), Positives = 99/204 (48%), Gaps = 8/204 (3%)
Query: 80 ISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNL 138
I +LP L L V N+F+ + ++ + D++ N G +PP + + L +
Sbjct: 331 IGDLPNLETLQVWENNFSFVLPQNLGSNGKFIYFDVTKNHLTGLIPPELCKSKKLKTFIV 390
Query: 139 SLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLG 198
+ N F G +PN LE + +N G + +Q+ SV ++L NN+F+G L
Sbjct: 391 TDNFFRGPIPNGIGPCKSLEKIRVANNYLDGPVPPGIFQLPSVQIIELGNNRFNGQLPTE 450
Query: 199 LGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIPSFTFVVS-L 257
+ S+ +L +S+N G + A M L +L+ NQ +G IP+ F + L
Sbjct: 451 ISG----NSLGNLALSNNLFTGRIPA--SMKNLRSLQTLLLDANQFLGEIPAEVFALPVL 504
Query: 258 RILRLACNQLTGSLPETLLKESSM 281
+ ++ N LTG +P+T+ + SS+
Sbjct: 505 TRINISGNNLTGGIPKTVTQCSSL 528
Score = 52.8 bits (125), Expect = 1e-05
Identities = 35/121 (28%), Positives = 56/121 (45%), Gaps = 1/121 (0%)
Query: 79 AISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLN 137
++ NL L L + N F G + + + L +++S N G +P + + SL ++
Sbjct: 473 SMKNLRSLQTLLLDANQFLGEIPAEVFALPVLTRINISGNNLTGGIPKTVTQCSSLTAVD 532
Query: 138 LSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDL 197
S N +G VP L L + NS SG I + M S+ +DLS N F+G +
Sbjct: 533 FSRNMLTGEVPKGMKNLKVLSIFNVSHNSISGKIPDEIRFMTSLTTLDLSYNNFTGIVPT 592
Query: 198 G 198
G
Sbjct: 593 G 593
>gb|AAP68247.1| At1g28440 [Arabidopsis thaliana] gi|20260672|gb|AAM13234.1|
putative receptor protein kinase [Arabidopsis thaliana]
gi|15218660|ref|NP_174166.1| leucine-rich repeat
transmembrane protein kinase, putative [Arabidopsis
thaliana] gi|6560764|gb|AAF16764.1| F3M18.12
[Arabidopsis thaliana] gi|25287711|pir||F86410 protein
F3M18.12 [imported] - Arabidopsis thaliana
Length = 996
Score = 110 bits (274), Expect = 6e-23
Identities = 95/305 (31%), Positives = 145/305 (47%), Gaps = 40/305 (13%)
Query: 2 LLLLVNTAFGNRDIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILCSE--G 59
L LL T F +L+ K +DP L+SW+S ++ P W G+ C+
Sbjct: 6 LFLLFPTVFSLNQDGFILQQVKLSLDDPDSY-LSSWNS----NDASPCRWSGVSCAGDFS 60
Query: 60 NVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNK 118
+V S+ L +A+L G F + I L L +LS+ NN ++ L+I+ KSL+ LDLS N
Sbjct: 61 SVTSVDLSSANLAGPFPSV-ICRLSNLAHLSLYNNSINSTLPLNIAACKSLQTLDLSQNL 119
Query: 119 FNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQM 178
G LP + ++ +LV+L+L+ N FSG +P F K + LE L N G I +
Sbjct: 120 LTGELPQTLADIPTLVHLDLTGNNFSGDIPASFGKFENLEVLSLVYNLLDGTIPPFLGNI 179
Query: 179 GSVLHVDLSNNKFS-------------------------GALDLGLGDVSFLFSIQHLNV 213
++ ++LS N FS G + LG +S L L++
Sbjct: 180 STLKMLNLSYNPFSPSRIPPEFGNLTNLEVMWLTECHLVGQIPDSLGQLSKLVD---LDL 236
Query: 214 SHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNI-PSFTFVVSLRILRLACNQLTGSLP 272
+ N LVG + + L N+ + NN L G I P + SLR+L + NQLTG +P
Sbjct: 237 ALNDLVGHI--PPSLGGLTNVVQIELYNNSLTGEIPPELGNLKSLRLLDASMNQLTGKIP 294
Query: 273 ETLLK 277
+ L +
Sbjct: 295 DELCR 299
Score = 89.0 bits (219), Expect = 1e-16
Identities = 65/225 (28%), Positives = 107/225 (46%), Gaps = 7/225 (3%)
Query: 55 LCSEGNVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLD 113
LC++G + + + + S G ++++ L + + N F+GS+ + + L+
Sbjct: 368 LCAKGELEELLIIHNSFSGVIPE-SLADCRSLTRIRLAYNRFSGSVPTGFWGLPHVNLLE 426
Query: 114 LSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIME 173
L N F+G + S +L L LS NEF+G++P LD L L N FSG + +
Sbjct: 427 LVNNSFSGEISKSIGGASNLSLLILSNNEFTGSLPEEIGSLDNLNQLSASGNKFSGSLPD 486
Query: 174 IFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDN 233
+G + +DL N+FSG L G+ + LN++ N G++ D + L
Sbjct: 487 SLMSLGELGTLDLHGNQFSGELTSGIKSWK---KLNELNLADNEFTGKI--PDEIGSLSV 541
Query: 234 LEVFDASNNQLVGNIPSFTFVVSLRILRLACNQLTGSLPETLLKE 278
L D S N G IP + L L L+ N+L+G LP +L K+
Sbjct: 542 LNYLDLSGNMFSGKIPVSLQSLKLNQLNLSYNRLSGDLPPSLAKD 586
Score = 87.4 bits (215), Expect = 4e-16
Identities = 77/253 (30%), Positives = 118/253 (46%), Gaps = 34/253 (13%)
Query: 63 SITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLHISPMKS-LKFLDLSLNKFNG 121
S+ L +L GE +I+ P L+ + + N TG + + S L++LD+S N+F+G
Sbjct: 304 SLNLYENNLEGELP-ASIALSPNLYEIRIFGNRLTGGLPKDLGLNSPLRWLDVSENEFSG 362
Query: 122 SLPP------------------------SFVELRSLVYLNLSLNEFSGTVPNVFHKLDQL 157
LP S + RSL + L+ N FSG+VP F L +
Sbjct: 363 DLPADLCAKGELEELLIIHNSFSGVIPESLADCRSLTRIRLAYNRFSGSVPTGFWGLPHV 422
Query: 158 EYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNS 217
L+ +NSFSG+I + ++ + LSNN+F+G+L +G L ++ L+ S N
Sbjct: 423 NLLELVNNSFSGEISKSIGGASNLSLLILSNNEFTGSLPEEIGS---LDNLNQLSASGNK 479
Query: 218 LVGELFAHDGMPYLDNLEVFDASNNQLVGNIPS-FTFVVSLRILRLACNQLTGSLPETLL 276
G L D + L L D NQ G + S L L LA N+ TG +P+ +
Sbjct: 480 FSGSL--PDSLMSLGELGTLDLHGNQFSGELTSGIKSWKKLNELNLADNEFTGKIPDEI- 536
Query: 277 KESSMMLSELDLS 289
S +L+ LDLS
Sbjct: 537 -GSLSVLNYLDLS 548
Score = 75.9 bits (185), Expect = 1e-12
Identities = 65/218 (29%), Positives = 98/218 (44%), Gaps = 9/218 (4%)
Query: 60 NVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKF 119
NV+ I L N SL GE + NL L L N TG + L+ L+L N
Sbjct: 254 NVVQIELYNNSLTGEIP-PELGNLKSLRLLDASMNQLTGKIPDELCRVPLESLNLYENNL 312
Query: 120 NGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMG 179
G LP S +L + + N +G +P L +LD N FSGD+ G
Sbjct: 313 EGELPASIALSPNLYEIRIFGNRLTGGLPKDLGLNSPLRWLDVSENEFSGDLPADLCAKG 372
Query: 180 SVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGEL-FAHDGMPYLDNLEVFD 238
+ + + +N FSG + L D S+ + +++N G + G+P+++ LE+
Sbjct: 373 ELEELLIIHNSFSGVIPESLADCR---SLTRIRLAYNRFSGSVPTGFWGLPHVNLLELV- 428
Query: 239 ASNNQLVGNI-PSFTFVVSLRILRLACNQLTGSLPETL 275
NN G I S +L +L L+ N+ TGSLPE +
Sbjct: 429 --NNSFSGEISKSIGGASNLSLLILSNNEFTGSLPEEI 464
>gb|AAN15323.1| Cf-5 disease resistance protein-like [Arabidopsis thaliana]
gi|22135942|gb|AAM91553.1| Cf-5 disease resistance
protein-like [Arabidopsis thaliana]
gi|9757845|dbj|BAB08479.1| leucine-rich repeat disease
resistance protein-like [Arabidopsis thaliana]
Length = 326
Score = 110 bits (274), Expect = 6e-23
Identities = 90/284 (31%), Positives = 140/284 (48%), Gaps = 15/284 (5%)
Query: 1 MLLLLVNTAFGNRDIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILCS-EG 59
+L+ ++ RD+ AL E+K + V+ SW +G W G+ CS +G
Sbjct: 15 LLIAFAHSKTLKRDVKALNEIKASLG----WRVVYSWVGDDPCGDGDLPPWSGVTCSTQG 70
Query: 60 N---VISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSML-HISPMKSLKFLDLS 115
+ V + + S+VG F +A++NL L L + NN TG + I +K LK L+L
Sbjct: 71 DYRVVTELEVYAVSIVGPFP-IAVTNLLDLTRLDLHNNKLTGPIPPQIGRLKRLKVLNLR 129
Query: 116 LNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIF 175
NK +PP EL+ L +L LS N F G +P L +L YL N G I
Sbjct: 130 WNKLQDVIPPEIGELKRLTHLYLSFNSFKGEIPKELAALPELRYLYLQENRLIGRIPAEL 189
Query: 176 YQMGSVLHVDLSNNKFSGAL-DLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNL 234
+ ++ H+D+ NN G + +L D SF ++++L +++N L G + A + L NL
Sbjct: 190 GTLQNLRHLDVGNNHLVGTIRELIRFDGSFP-ALRNLYLNNNYLSGGIPAQ--LSNLTNL 246
Query: 235 EVFDASNNQLVGNIP-SFTFVVSLRILRLACNQLTGSLPETLLK 277
E+ S N+ +GNIP + + L L L NQ TG +P+ K
Sbjct: 247 EIVYLSYNKFIGNIPFAIAHIPKLTYLYLDHNQFTGRIPDAFYK 290
Score = 58.5 bits (140), Expect = 2e-07
Identities = 36/118 (30%), Positives = 61/118 (51%), Gaps = 3/118 (2%)
Query: 60 NVISITLDNASLVGEFNFLAI--SNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSL 116
N+ + + N LVG L + P L NL + NN+ +G + +S + +L+ + LS
Sbjct: 194 NLRHLDVGNNHLVGTIRELIRFDGSFPALRNLYLNNNYLSGGIPAQLSNLTNLEIVYLSY 253
Query: 117 NKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEI 174
NKF G++P + + L YL L N+F+G +P+ F+K L+ + N F + I
Sbjct: 254 NKFIGNIPFAIAHIPKLTYLYLDHNQFTGRIPDAFYKHPFLKEMYIEGNMFKSGVNPI 311
Score = 55.1 bits (131), Expect = 2e-06
Identities = 33/121 (27%), Positives = 57/121 (46%), Gaps = 4/121 (3%)
Query: 80 ISNLPMLHNLSVVNNHFTGSMLHI----SPMKSLKFLDLSLNKFNGSLPPSFVELRSLVY 135
+ L L +L V NNH G++ + +L+ L L+ N +G +P L +L
Sbjct: 189 LGTLQNLRHLDVGNNHLVGTIRELIRFDGSFPALRNLYLNNNYLSGGIPAQLSNLTNLEI 248
Query: 136 LNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGAL 195
+ LS N+F G +P + +L YL N F+G I + FY+ + + + N F +
Sbjct: 249 VYLSYNKFIGNIPFAIAHIPKLTYLYLDHNQFTGRIPDAFYKHPFLKEMYIEGNMFKSGV 308
Query: 196 D 196
+
Sbjct: 309 N 309
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.320 0.138 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 484,823,131
Number of Sequences: 2540612
Number of extensions: 20746285
Number of successful extensions: 82454
Number of sequences better than 10.0: 3641
Number of HSP's better than 10.0 without gapping: 2162
Number of HSP's successfully gapped in prelim test: 1530
Number of HSP's that attempted gapping in prelim test: 47010
Number of HSP's gapped (non-prelim): 14369
length of query: 289
length of database: 863,360,394
effective HSP length: 127
effective length of query: 162
effective length of database: 540,702,670
effective search space: 87593832540
effective search space used: 87593832540
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 74 (33.1 bits)
Medicago: description of AC124972.17


