

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124970.4 + phase: 0 /pseudo
(204 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
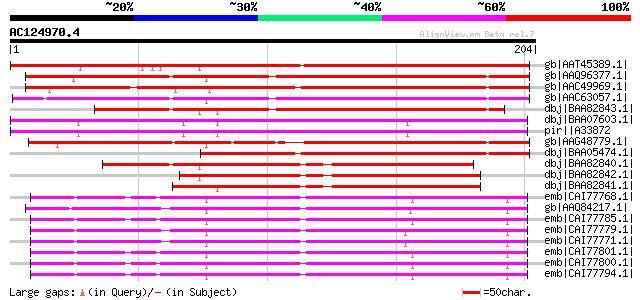
Score E
Sequences producing significant alignments: (bits) Value
gb|AAT45389.1| pathogen-inducible trypsin-inhibitor-like protein... 186 3e-46
gb|AAQ96377.1| miraculin-like protein [Solanum brevidens] 171 9e-42
gb|AAC49969.1| tumor-related protein [Nicotiana tabacum] gi|7438... 169 3e-41
gb|AAC63057.1| Lemir [Lycopersicon esculentum] gi|7438248|pir||T... 169 4e-41
dbj|BAA82843.1| miraculin homologue [Solanum melongena] 160 3e-38
dbj|BAA07603.1| miraculin precursor [Richadella dulcifica] gi|61... 152 6e-36
pir||A33872 miraculin precursor - sweet berry 151 9e-36
gb|AAG48779.1| putative lemir (miraculin) protein [Arabidopsis t... 143 3e-33
dbj|BAA05474.1| tumor-related protein [Nicotiana glauca x Nicoti... 142 4e-33
dbj|BAA82840.1| miraculin homologue [Youngia japonica] 127 2e-28
dbj|BAA82842.1| miraculin homologue [Taraxacum officinale] 119 5e-26
dbj|BAA82841.1| miraculin homologue [Youngia japonica] 115 1e-24
emb|CAI77768.1| kunitz trypsin inhibitor [Populus tremula] 113 3e-24
gb|AAQ84217.1| Kunitz trypsin inhibitor 4 [Populus balsamifera s... 112 6e-24
emb|CAI77785.1| kunitz trypsin inhibitor [Populus tremula] 111 1e-23
emb|CAI77779.1| kunitz trypsin inhibitor [Populus tremula] gi|65... 111 1e-23
emb|CAI77771.1| kunitz trypsin inhibitor [Populus tremula] 111 1e-23
emb|CAI77801.1| kunitz trypsin inhibitor [Populus tremula] gi|65... 111 1e-23
emb|CAI77800.1| kunitz trypsin inhibitor [Populus tremula] 111 1e-23
emb|CAI77794.1| kunitz trypsin inhibitor [Populus tremula] gi|65... 111 1e-23
>gb|AAT45389.1| pathogen-inducible trypsin-inhibitor-like protein [Medicago
truncatula]
Length = 213
Score = 186 bits (472), Expect = 3e-46
Identities = 102/214 (47%), Positives = 138/214 (63%), Gaps = 13/214 (6%)
Query: 1 MKLHFPLLFLLLTLFTTKPLQGAEQP--EEVRDTSGNLVRNSINYFILPSSI-QCGT--R 55
MK F L T +++PL GA + E+V DT G +R NY+I+P I +CG +
Sbjct: 1 MKNTLLAFFFLFTFLSSQPLLGAAEASNEQVVDTLGKKLRADANYYIIPVPIYKCGPYGK 60
Query: 56 CE-----MALLNTNKTCPLDVV--EEEEAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNC 108
C +AL + KTCPLDVV + +A+ +F+P N KKGVIRVSTDLN+ S C
Sbjct: 61 CRSSGSSLALASIGKTCPLDVVVVDRYQALPLTFIPVNPKKGVIRVSTDLNIKFSSRATC 120
Query: 109 STSSVTVWKVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESGYKLVFCPTVCRE 168
S+ VWK+D+ +V+ Q F+T GGV GNPG ET++NWFKIE++ YKLVFCP+V +
Sbjct: 121 LHHSM-VWKLDRFNVSKRQWFITIGGVAGNPGWETINNWFKIEKYGDAYKLVFCPSVVQS 179
Query: 169 CEVVCKDIGIFLDENRNTRFVLSDFPFGVKFQRA 202
+ +CKD+G+F+DEN N R LSD P VKFQ+A
Sbjct: 180 FKHMCKDVGVFVDENGNKRLALSDVPLKVKFQQA 213
>gb|AAQ96377.1| miraculin-like protein [Solanum brevidens]
Length = 209
Score = 171 bits (434), Expect = 9e-42
Identities = 94/202 (46%), Positives = 123/202 (60%), Gaps = 10/202 (4%)
Query: 7 LLFLLLTLFTTKPLQGA-EQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNK 65
L FL+L + + L A E P EV D G ++R ++Y+ILP G M + +K
Sbjct: 12 LPFLILVISSNSFLSSAAESPPEVVDIDGKILRTGVDYYILPVVRGRGGGLTMDSIG-DK 70
Query: 66 TCPLDVVEEE-----EAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDK 120
CPLD V +E + + +F P N KKGVIR STDLN+I F N T WK+D
Sbjct: 71 ICPLDAVVQEHQEIDQGLPLTFTPVNPKKGVIRESTDLNII--FSANSICVQTTQWKLDD 128
Query: 121 VDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESGYKLVFCPTVCRECEVVCKDIGIFL 180
D T Q F+T GG QGNPGRET+ NWFKIE+F+ YKL++CPTVC C+V+CK+IGIF+
Sbjct: 129 FDETTGQYFITLGGNQGNPGRETISNWFKIEKFDRDYKLLYCPTVCDFCKVICKEIGIFI 188
Query: 181 DENRNTRFVLSDFPFGVKFQRA 202
+ R LSD PF V F++A
Sbjct: 189 QDGVR-RLALSDVPFKVMFKKA 209
>gb|AAC49969.1| tumor-related protein [Nicotiana tabacum] gi|7438249|pir||T03803
tumor-related protein, clone NF34 - common tobacco
Length = 210
Score = 169 bits (429), Expect = 3e-41
Identities = 94/203 (46%), Positives = 128/203 (62%), Gaps = 12/203 (5%)
Query: 7 LLFLLLTL-FTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNT-N 64
L FL+ T+ F + AE P V D +G +R I+Y+ILP + G + L +T N
Sbjct: 8 LPFLIFTISFNSFLSSSAEAPPAVVDIAGKKLRTGIDYYILP--VVRGRGGGLTLDSTGN 65
Query: 65 KTCPLDVVEEEE-----AMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVD 119
++CPLD V +E+ + +F P N KKGVIR STDLN+ S + C + T+WK+D
Sbjct: 66 ESCPLDAVVQEQQEIKNGLPLTFTPVNPKKGVIRESTDLNIKFSAASICVQT--TLWKLD 123
Query: 120 KVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESGYKLVFCPTVCRECEVVCKDIGIF 179
D T + F+T GG +GNPGRET+ NWFKIE+FE YKLV+CPTVC C+V+CKD+GIF
Sbjct: 124 DFDETTGKYFITIGGNEGNPGRETISNWFKIEKFERDYKLVYCPTVCNFCKVICKDVGIF 183
Query: 180 LDENRNTRFVLSDFPFGVKFQRA 202
+ + R LSD PF V F++A
Sbjct: 184 IQDGIR-RLALSDVPFKVMFKKA 205
>gb|AAC63057.1| Lemir [Lycopersicon esculentum] gi|7438248|pir||T07871 miraculin
homolog, root-knot nematode-induced - tomato
Length = 205
Score = 169 bits (428), Expect = 4e-41
Identities = 92/206 (44%), Positives = 124/206 (59%), Gaps = 10/206 (4%)
Query: 2 KLHFPLLFLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALL 61
+L FP L L ++ F + AE P EV D G ++R ++Y+ILP G M +
Sbjct: 5 QLFFPFLILAIS-FNSLLSSAAESPPEVVDIDGKILRTGVDYYILPVVRGRGGGLTMDSI 63
Query: 62 NTNKTCPLDVVEEE-----EAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVW 116
+K CPLD V +E + + +F P + KKGVIR STDLN+I F N T W
Sbjct: 64 G-DKMCPLDAVVQEHNEIDQGLPLTFTPVDPKKGVIRESTDLNII--FSANSICVQTTQW 120
Query: 117 KVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESGYKLVFCPTVCRECEVVCKDI 176
K+D D T Q F+T GG QGNPG ET+ NWFKIE+++ YKL++CPTVC C+V+C+DI
Sbjct: 121 KLDDFDETTGQYFITLGGDQGNPGVETISNWFKIEKYDRDYKLLYCPTVCDFCKVICRDI 180
Query: 177 GIFLDENRNTRFVLSDFPFGVKFQRA 202
GIF+ + R LSD PF V F++A
Sbjct: 181 GIFIQDGVR-RLALSDVPFKVMFKKA 205
>dbj|BAA82843.1| miraculin homologue [Solanum melongena]
Length = 160
Score = 160 bits (404), Expect = 3e-38
Identities = 87/164 (53%), Positives = 105/164 (63%), Gaps = 9/164 (5%)
Query: 34 GNLVRNSINYFILPSSIQCGTRCEMALLNTNKTCPLDVV--EEEEAMQ---FSFVPFNFK 88
G ++R I+Y+ILP G M + NKTCPLD V E+EE Q +F PFN K
Sbjct: 1 GKILRTGIDYYILPVVRGRGGGLTMDSIG-NKTCPLDAVVQEQEEVKQGLPLTFTPFNPK 59
Query: 89 KGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVATSQRFVTTGGVQGNPGRETVDNWF 148
KGVIR STDLN+I F N T WK+D D T + F+T GG QGNPGRET+ NWF
Sbjct: 60 KGVIRESTDLNII--FSANSICVQTTQWKLDNFDETTGKYFITLGGNQGNPGRETISNWF 117
Query: 149 KIERFESGYKLVFCPTVCRECEVVCKDIGIFLDENRNTRFVLSD 192
KIE+FE YKLV+CPTVC C+V+CKDIGIF+ + R LSD
Sbjct: 118 KIEKFERDYKLVYCPTVCDFCKVICKDIGIFIQDGVR-RLALSD 160
>dbj|BAA07603.1| miraculin precursor [Richadella dulcifica]
gi|6166552|sp|P13087|MIRA_RICDU Miraculin precursor
(MIR)
Length = 220
Score = 152 bits (384), Expect = 6e-36
Identities = 87/210 (41%), Positives = 116/210 (54%), Gaps = 9/210 (4%)
Query: 1 MKLHFPLLFLLLTLFTTKPLQGAEQ-PEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMA 59
+ L F + LL L A+ P V D G +R NY+I+P G ++
Sbjct: 7 LSLSFFFVSALLAAAANPLLSAADSAPNPVLDIDGEKLRTGTNYYIVPVLRDHGGGLTVS 66
Query: 60 LLNTNKT--CPLDVVEEEEAMQ----FSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSV 113
N T CP VV+ + + +F P N K+ V+RVSTDLN+ S C +S
Sbjct: 67 ATTPNGTFVCPPRVVQTRKEVDHDRPLAFFPENPKEDVVRVSTDLNINFSAFMPCRWTSS 126
Query: 114 TVWKVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERF--ESGYKLVFCPTVCRECEV 171
TVW++DK D +T Q FVT GGV+GNPG ET+ +WFKIE F YKLVFCPTVC C+V
Sbjct: 127 TVWRLDKYDESTGQYFVTIGGVKGNPGPETISSWFKIEEFCGSGFYKLVFCPTVCGSCKV 186
Query: 172 VCKDIGIFLDENRNTRFVLSDFPFGVKFQR 201
C D+GI++D+ R LSD PF +F +
Sbjct: 187 KCGDVGIYIDQKGRRRLALSDKPFAFEFNK 216
>pir||A33872 miraculin precursor - sweet berry
Length = 220
Score = 151 bits (382), Expect = 9e-36
Identities = 87/210 (41%), Positives = 116/210 (54%), Gaps = 9/210 (4%)
Query: 1 MKLHFPLLFLLLTLFTTKPLQGAEQ-PEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMA 59
+ L F + LL L A+ P V D G +R NY+I+P G ++
Sbjct: 7 LSLSFFFVSGLLAAAANPRLSAADSAPNPVLDIDGEKLRTGTNYYIVPVLRDHGGGLTVS 66
Query: 60 LLNTNKT--CPLDVVEEEEAMQ----FSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSV 113
N T CP VV+ + + +F P N K+ V+RVSTDLN+ S C +S
Sbjct: 67 ATTPNGTFVCPPRVVQTRKEVDHDRPLAFFPENPKEDVVRVSTDLNINFSAFMPCRWTSS 126
Query: 114 TVWKVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERF--ESGYKLVFCPTVCRECEV 171
TVW++DK D +T Q FVT GGV+GNPG ET+ +WFKIE F YKLVFCPTVC C+V
Sbjct: 127 TVWRLDKYDESTGQYFVTIGGVKGNPGPETISSWFKIEEFCGSGFYKLVFCPTVCGSCKV 186
Query: 172 VCKDIGIFLDENRNTRFVLSDFPFGVKFQR 201
C D+GI++D+ R LSD PF +F +
Sbjct: 187 KCGDVGIYIDQKGRRRLALSDKPFAFEFNK 216
>gb|AAG48779.1| putative lemir (miraculin) protein [Arabidopsis thaliana]
gi|20453293|gb|AAM19885.1| At1g17860/F2H15_8
[Arabidopsis thaliana] gi|20148401|gb|AAM10091.1|
unknown protein [Arabidopsis thaliana]
gi|15220874|ref|NP_173228.1| trypsin and protease
inhibitor family protein / Kunitz family protein
[Arabidopsis thaliana] gi|15294166|gb|AAK95260.1|
At1g17860/F2H15_8 [Arabidopsis thaliana]
gi|13899081|gb|AAK48962.1| Unknown protein [Arabidopsis
thaliana] gi|25294073|pir||G86313 hypothetical protein
F2H15.9 - Arabidopsis thaliana gi|9665064|gb|AAF97266.1|
Contains similarity to a tumor-related protein from
Nicotiana tabacum gb|U66263 and contains a trypsin and
protease inhibitor PF|00197 domain. ESTs gb|AV561824,
gb|T44961, gb|H36186, gb|T45060, gb|N38006, gb|F19847
come from this gene. [Arabidopsis thaliana]
Length = 196
Score = 143 bits (361), Expect = 3e-33
Identities = 79/200 (39%), Positives = 122/200 (60%), Gaps = 16/200 (8%)
Query: 8 LFLLLTLFTT-KPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKT 66
+FLLL +F + + + E V+D +G + +NY+ILP G M+ L T +T
Sbjct: 7 IFLLLAVFISHRGVTTEAAVEPVKDINGKSLLTGVNYYILPVIRGRGGGLTMSNLKT-ET 65
Query: 67 CPLDVVEEE----EAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVD 122
CP V++++ + + F P++ K I VSTD+N+ S PT+ +W++ D
Sbjct: 66 CPTSVIQDQFEVSQGLPVKFSPYD-KSRTIPVSTDVNIKFS-PTS-------IWELANFD 116
Query: 123 VATSQRFVTTGGVQGNPGRETVDNWFKIERFESGYKLVFCPTVCRECEVVCKDIGIFLDE 182
T Q F++T GV+GNPG++TVDNWFKI++FE YK+ FCPTVC C+V+C+D+G+F+ +
Sbjct: 117 ETTKQWFISTCGVEGNPGQKTVDNWFKIDKFEKDYKIRFCPTVCNFCKVICRDVGVFVQD 176
Query: 183 NRNTRFVLSDFPFGVKFQRA 202
+ R LSD P V F+RA
Sbjct: 177 GKR-RLALSDVPLKVMFKRA 195
>dbj|BAA05474.1| tumor-related protein [Nicotiana glauca x Nicotiana langsdorffii]
Length = 125
Score = 142 bits (359), Expect = 4e-33
Identities = 68/128 (53%), Positives = 88/128 (68%), Gaps = 3/128 (2%)
Query: 75 EEAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVATSQRFVTTGG 134
E + +F P N KKGVIR STDLN+ S + C + T+WK+D D T + F+T GG
Sbjct: 1 ENGLPLTFTPVNPKKGVIRESTDLNIKFSAASICVQT--TLWKLDDFDETTGKYFITIGG 58
Query: 135 VQGNPGRETVDNWFKIERFESGYKLVFCPTVCRECEVVCKDIGIFLDENRNTRFVLSDFP 194
+GNPGRET+ NWFKIE+FE YKLV+CPTVC C+V+CKD+GIF+ + R LSD P
Sbjct: 59 NEGNPGRETISNWFKIEKFERDYKLVYCPTVCNFCKVICKDVGIFIQDGIR-RLALSDVP 117
Query: 195 FGVKFQRA 202
F V F++A
Sbjct: 118 FKVMFKKA 125
>dbj|BAA82840.1| miraculin homologue [Youngia japonica]
Length = 142
Score = 127 bits (318), Expect = 2e-28
Identities = 69/148 (46%), Positives = 90/148 (60%), Gaps = 10/148 (6%)
Query: 37 VRNSINYFILPSSIQCGTRCEMALLNTNKTCPLDVV----EEEEAMQFSFVPFNFKKGVI 92
+R+ Y+ILP G +A N TCPLDVV E + + +F P + KKGVI
Sbjct: 1 LRSGTEYYILPVFRGMGGGLTLASTR-NDTCPLDVVQADLEVDNGLPLTFTPVDPKKGVI 59
Query: 93 RVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIER 152
R STDLN+I S + C S+V W +++ D Q V+ GV GNPGRET+ NWFKIE+
Sbjct: 60 RESTDLNIIFSASSICIQSNV--WNLEEYD---GQLIVSAHGVAGNPGRETISNWFKIEK 114
Query: 153 FESGYKLVFCPTVCRECEVVCKDIGIFL 180
+E YK+VFCPTVC C VC DIG+ +
Sbjct: 115 YEDDYKIVFCPTVCDFCRPVCGDIGVLI 142
>dbj|BAA82842.1| miraculin homologue [Taraxacum officinale]
Length = 136
Score = 119 bits (298), Expect = 5e-26
Identities = 60/121 (49%), Positives = 79/121 (64%), Gaps = 9/121 (7%)
Query: 67 CPLDVV----EEEEAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVD 122
CPLDVV E + + +F P + KKGVIR STDLN+I S + C S+V W +++ D
Sbjct: 18 CPLDVVQANQELDNGLPLTFTPVDPKKGVIRESTDLNIIFSASSICIQSNV--WMIEEYD 75
Query: 123 VATSQRFVTTGGVQGNPGRETVDNWFKIERFESGYKLVFCPTVCRECEVVCKDIGIFLDE 182
V+ GVQGNPG+ET+ NWFKIE+F+ YK+VFCP VC C +C DIG+ +DE
Sbjct: 76 ---GSLIVSAHGVQGNPGQETLSNWFKIEKFDDDYKIVFCPAVCDVCRPLCGDIGVEIDE 132
Query: 183 N 183
N
Sbjct: 133 N 133
>dbj|BAA82841.1| miraculin homologue [Youngia japonica]
Length = 136
Score = 115 bits (287), Expect = 1e-24
Identities = 58/124 (46%), Positives = 79/124 (62%), Gaps = 9/124 (7%)
Query: 64 NKTCPLDVVEEEEAMQ----FSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVD 119
N+ CP DVV+ ++ + +F P + KKGVIR STDLN+I + C S+V W ++
Sbjct: 15 NELCPPDVVQADQELDNGLPLTFTPVDPKKGVIRESTDLNIIFRAYSICIQSNV--WMLE 72
Query: 120 KVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESGYKLVFCPTVCRECEVVCKDIGIF 179
+ D V+ GV GNPG+ET+ NWFKIE++E YKLVFCPTVC C +C DIG+
Sbjct: 73 EYD---GSLIVSGHGVAGNPGQETISNWFKIEKYEDDYKLVFCPTVCDTCRPICGDIGVE 129
Query: 180 LDEN 183
+ EN
Sbjct: 130 ISEN 133
>emb|CAI77768.1| kunitz trypsin inhibitor [Populus tremula]
Length = 213
Score = 113 bits (283), Expect = 3e-24
Identities = 80/201 (39%), Positives = 105/201 (51%), Gaps = 14/201 (6%)
Query: 9 FLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKTCP 68
FLL + P A E V D G ++ Y I SSI G + TNKTCP
Sbjct: 11 FLLFAFVLSVPSIEAST-EPVLDIQGEELKAGTEYII--SSIFWGAGGG-DVSATNKTCP 66
Query: 69 LDVVEEE----EAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVA 124
DV++ + + +F P + + VIRVSTDLN+ S C SSV WK+ K +
Sbjct: 67 DDVIQYSLDLLQGLPVTFSPASSEDDVIRVSTDLNIKFSIKKACDHSSV--WKIQKSSNS 124
Query: 125 TSQRFVTTGGVQGNPGRETVDNWFKIERFES--GYKLVFCPTVCRECEVVCKDIGIFLDE 182
QR VTTGG +GNPG +T NWFKIE+ GYKLV+CP +C+D+GI+ +
Sbjct: 125 EVQRLVTTGGEEGNPGCDTFTNWFKIEKGAGILGYKLVYCPEDICPSVGLCRDVGIYFES 184
Query: 183 NRNTRFVLSD--FPFGVKFQR 201
NR LSD PF V F++
Sbjct: 185 NRGRILSLSDKLSPFLVLFKK 205
>gb|AAQ84217.1| Kunitz trypsin inhibitor 4 [Populus balsamifera subsp. trichocarpa
x Populus deltoides]
Length = 212
Score = 112 bits (280), Expect = 6e-24
Identities = 80/202 (39%), Positives = 103/202 (50%), Gaps = 13/202 (6%)
Query: 7 LLFLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKT 66
L FLL + P A E V D G ++ Y I G A TNKT
Sbjct: 9 LSFLLFAFVLSVPSIEA-YTEPVLDIQGEELKAGTEYIIGSIFFGAGGGDVSA---TNKT 64
Query: 67 CPLDVVEEE----EAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVD 122
CP DV++ + + +F P + VIRVSTDLN+ S C SSV WK+ K
Sbjct: 65 CPDDVIQYSSDLLQGLPVTFSPASSDDDVIRVSTDLNIKFSIKKACDHSSV--WKIQKSS 122
Query: 123 VATSQRFVTTGGVQGNPGRETVDNWFKIERFE-SGYKLVFCPTVCRECEVVCKDIGIFLD 181
+ Q FVTTGG +GNPG +T+ NWFKIE+ GYKLV CP C V+C+DIGI+ +
Sbjct: 123 NSEVQWFVTTGGEEGNPGIDTLTNWFKIEKAGILGYKLVSCPEGICHCGVLCRDIGIYRE 182
Query: 182 ENRNTRFVLSD--FPFGVKFQR 201
+ LSD PF V F++
Sbjct: 183 NDGRRILSLSDKLSPFLVLFKK 204
>emb|CAI77785.1| kunitz trypsin inhibitor [Populus tremula]
Length = 213
Score = 111 bits (278), Expect = 1e-23
Identities = 79/201 (39%), Positives = 104/201 (51%), Gaps = 14/201 (6%)
Query: 9 FLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKTCP 68
FLL + P A E V D G ++ Y I SSI G + TNKTCP
Sbjct: 11 FLLFAFVLSVPSIEAST-ESVLDIQGEELKAGTEYII--SSIFWGAGGG-DVAATNKTCP 66
Query: 69 LDVVEEE----EAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVA 124
DV++ + + +F P + + VIRVSTDLN+ S C SSV WK+ K +
Sbjct: 67 DDVIQYSLDLLQGLPVTFSPASSEDDVIRVSTDLNIKFSIKKACDRSSV--WKIQKSSNS 124
Query: 125 TSQRFVTTGGVQGNPGRETVDNWFKIERFES--GYKLVFCPTVCRECEVVCKDIGIFLDE 182
Q VTTGG +GNPG +T NWFKIE+ GYKLV+CP +C+D+GI+ +
Sbjct: 125 EVQWLVTTGGEEGNPGCDTFTNWFKIEKGAGILGYKLVYCPEDICPSVGLCRDVGIYFES 184
Query: 183 NRNTRFVLSD--FPFGVKFQR 201
NR LSD PF V F++
Sbjct: 185 NRGRILSLSDKLSPFLVLFKK 205
>emb|CAI77779.1| kunitz trypsin inhibitor [Populus tremula]
gi|65305817|emb|CAI77778.1| kunitz trypsin inhibitor
[Populus tremula] gi|65305795|emb|CAI77767.1| kunitz
trypsin inhibitor [Populus tremula]
Length = 215
Score = 111 bits (278), Expect = 1e-23
Identities = 78/203 (38%), Positives = 102/203 (49%), Gaps = 16/203 (7%)
Query: 9 FLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKTCP 68
FLL + P A+ E V D G ++ Y I G A TNKTCP
Sbjct: 11 FLLFAFVLSVPSIEADT-EPVLDIQGEELKAGTEYIITSIFRGAGGGDVAA---TNKTCP 66
Query: 69 LDVVEEE----EAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVA 124
DV++ + + +F P + + VIRVSTDLN+ S C SSV WK+ K +
Sbjct: 67 DDVIQYSLDVFQGLPVTFSPASSEDDVIRVSTDLNIKFSIKKACDRSSV--WKIQKSSNS 124
Query: 125 TSQRFVTTGGVQGNPGRETVDNWFKIER----FESGYKLVFCPTVCRECEVVCKDIGIFL 180
Q VTTGG +GNPG +T NWFKIE+ F YKLVFCP +C+D+GI+
Sbjct: 125 EVQWLVTTGGEEGNPGCDTFTNWFKIEKGAGTFGYNYKLVFCPEDICPSVGLCRDVGIYF 184
Query: 181 DENRNTRFVLSD--FPFGVKFQR 201
+ NR LSD PF V F++
Sbjct: 185 ESNRGRILSLSDKLSPFLVVFRK 207
>emb|CAI77771.1| kunitz trypsin inhibitor [Populus tremula]
Length = 215
Score = 111 bits (278), Expect = 1e-23
Identities = 78/203 (38%), Positives = 102/203 (49%), Gaps = 16/203 (7%)
Query: 9 FLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKTCP 68
FLL + P A+ E V D G ++ Y I G A TNKTCP
Sbjct: 11 FLLFAFVLSVPSIEADT-EPVLDIQGEELKAGTEYIITSIFRGAGGGDVAA---TNKTCP 66
Query: 69 LDVVEEE----EAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVA 124
DV++ + + +F P + + VIRVSTDLN+ S C SSV WK+ K +
Sbjct: 67 DDVIQYSLDVFQGLPVTFSPASSEDDVIRVSTDLNIKFSIKKACDRSSV--WKIQKSSNS 124
Query: 125 TSQRFVTTGGVQGNPGRETVDNWFKIER----FESGYKLVFCPTVCRECEVVCKDIGIFL 180
Q VTTGG +GNPG +T NWFKIE+ F YKLVFCP +C+D+GI+
Sbjct: 125 EVQWLVTTGGEEGNPGCDTFTNWFKIEKGAGTFGYNYKLVFCPEDICPSVGLCRDVGIYF 184
Query: 181 DENRNTRFVLSD--FPFGVKFQR 201
+ NR LSD PF V F++
Sbjct: 185 ESNRGRILSLSDKLSPFLVVFRK 207
>emb|CAI77801.1| kunitz trypsin inhibitor [Populus tremula]
gi|65305855|emb|CAI77797.1| kunitz trypsin inhibitor
[Populus tremula] gi|65305853|emb|CAI77796.1| kunitz
trypsin inhibitor [Populus tremula]
gi|65305845|emb|CAI77792.1| kunitz trypsin inhibitor
[Populus tremula] gi|65305791|emb|CAI77765.1| kunitz
trypsin inhibitor [Populus tremula]
Length = 213
Score = 111 bits (277), Expect = 1e-23
Identities = 79/201 (39%), Positives = 104/201 (51%), Gaps = 14/201 (6%)
Query: 9 FLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKTCP 68
FLL + P A E V D G ++ Y I SSI G + TNKTCP
Sbjct: 11 FLLFAFVLSVPSIEAST-EPVLDIQGEELKAGTEYII--SSIFWGAGGG-DVAATNKTCP 66
Query: 69 LDVVEEE----EAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVA 124
DV++ + + +F P + + VIRVSTDLN+ S C SSV WK+ K +
Sbjct: 67 DDVIQYSLDLLQGLPVTFSPASSEDDVIRVSTDLNIKFSIKKACDRSSV--WKIQKSSNS 124
Query: 125 TSQRFVTTGGVQGNPGRETVDNWFKIERFES--GYKLVFCPTVCRECEVVCKDIGIFLDE 182
Q VTTGG +GNPG +T NWFKIE+ GYKLV+CP +C+D+GI+ +
Sbjct: 125 EVQWLVTTGGEEGNPGCDTFTNWFKIEKGAGILGYKLVYCPEDICPSVGLCRDVGIYFES 184
Query: 183 NRNTRFVLSD--FPFGVKFQR 201
NR LSD PF V F++
Sbjct: 185 NRGRILSLSDKLSPFLVLFKK 205
>emb|CAI77800.1| kunitz trypsin inhibitor [Populus tremula]
Length = 213
Score = 111 bits (277), Expect = 1e-23
Identities = 79/201 (39%), Positives = 104/201 (51%), Gaps = 14/201 (6%)
Query: 9 FLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKTCP 68
FLL + P A E V D G ++ Y I SSI G + TNKTCP
Sbjct: 11 FLLFAFVLSVPSIEAST-EPVLDIQGEELKAGTEYII--SSIFWGAGGG-DVAATNKTCP 66
Query: 69 LDVVEEE----EAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVA 124
DV++ + + +F P + + VIRVSTDLN+ S C SSV WK+ K +
Sbjct: 67 DDVIQYSLDLLQVLPVTFSPASSEDDVIRVSTDLNIKFSIKKACDRSSV--WKIQKSSNS 124
Query: 125 TSQRFVTTGGVQGNPGRETVDNWFKIERFES--GYKLVFCPTVCRECEVVCKDIGIFLDE 182
Q VTTGG +GNPG +T NWFKIE+ GYKLV+CP +C+D+GI+ +
Sbjct: 125 EVQWLVTTGGEEGNPGCDTFTNWFKIEKGAGILGYKLVYCPEDICPSVGLCRDVGIYFES 184
Query: 183 NRNTRFVLSD--FPFGVKFQR 201
NR LSD PF V F++
Sbjct: 185 NRGRILSLSDKLSPFLVLFKK 205
>emb|CAI77794.1| kunitz trypsin inhibitor [Populus tremula]
gi|65305825|emb|CAI77782.1| kunitz trypsin inhibitor
[Populus tremula]
Length = 213
Score = 111 bits (277), Expect = 1e-23
Identities = 79/201 (39%), Positives = 104/201 (51%), Gaps = 14/201 (6%)
Query: 9 FLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKTCP 68
FLL + P A E V D G ++ Y I SSI G + TNKTCP
Sbjct: 11 FLLFAFVLSVPSIEAST-EPVLDIQGEELKAGTEYII--SSIFWGAGGG-DVAATNKTCP 66
Query: 69 LDVVEEE----EAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVA 124
DV++ + + +F P + + VIRVSTDLN+ S C SSV WK+ K +
Sbjct: 67 DDVIQYSLDLLQGLPVTFSPASSEDDVIRVSTDLNIKFSIKKACDRSSV--WKIQKSSNS 124
Query: 125 TSQRFVTTGGVQGNPGRETVDNWFKIERFES--GYKLVFCPTVCRECEVVCKDIGIFLDE 182
Q VTTGG +GNPG +T NWFKIE+ GYKLV+CP +C+D+GI+ +
Sbjct: 125 EVQWLVTTGGEEGNPGCDTFTNWFKIEKGAGILGYKLVYCPEDICPSVGLCRDVGIYFES 184
Query: 183 NRNTRFVLSD--FPFGVKFQR 201
NR LSD PF V F++
Sbjct: 185 NRGRILSLSDKLSPFLVVFKK 205
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.324 0.139 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 349,387,683
Number of Sequences: 2540612
Number of extensions: 13859743
Number of successful extensions: 33974
Number of sequences better than 10.0: 294
Number of HSP's better than 10.0 without gapping: 75
Number of HSP's successfully gapped in prelim test: 219
Number of HSP's that attempted gapping in prelim test: 33569
Number of HSP's gapped (non-prelim): 305
length of query: 204
length of database: 863,360,394
effective HSP length: 122
effective length of query: 82
effective length of database: 553,405,730
effective search space: 45379269860
effective search space used: 45379269860
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 72 (32.3 bits)
Medicago: description of AC124970.4


